SUPPLEMENTARY INFORMATION
|
|
|
- Phyllis James
- 5 years ago
- Views:
Transcription
1 doi:1.138/nature111 cytosol Model: PILS function in cellular auxin homeostasis ER nucleus IAA degradation? sequestration? conjugation? storage? signalling? PILS IAA ER cytosol Supplemental Figure 1 Model on cellular PILS function. PILS proteins localize to the endoplasmic reticulum (ER) and regulate cellular auxin accumulation, presumably by regulating auxin transport from the cytosol into the ER lumen. PILS function increases auxin conjugation-based inactivation of auxin and negatively regulates nuclear auxin signaling. We assume that both auxin sequestration into the ER and auxin conjugation might affect the availability of auxin for nuclear signalling. While our data may support a potential auxin conjugation in the ER, it remains to be seen whether known or yet to be discovered auxin amide conjugate synthetases function in the ER. To our knowledge, potential candidates of the GRETCHEN HAGEN (GH3) IAA amido synthetase family have not been localized in Arabidopsis, but in silico analysis of GH3 sequences do not suggest their residence in the ER 36. In Physcomitrella, cytoplasmic localization was suggested for GH3-1 tagged with GFP 37. Nevertheless, auxin metabolism is likely to be compartmentalized in the ER. Most known auxin conjugate hydrolyses, such as IAA-LEUCINE RESTISTANT1 LIKE (ILL) proteins, contain ER retention motifs 38,39, but their localization remains to be experimentally proven. Moreover, isoforms of YUCCA4 catalyzes auxin biosynthesis at the outer surface of the ER 4, possibly providing substrate for the auxin transport into the ER. Notably, also the auxin receptor AUXIN BINDING PROTEIN1 (ABP1) mainly resides in the lumen of the ER 41,42. These results strongly suggest that auxin compartmentalization is important for plant development, but the unifying mechanism remains to be discovered. 1
2 SUPPLEMENTARY INFORMATION PILS1 PILS5 PILS2 PILS6 PILS3 PILS7 PILS4 Supplemental Figure 2 PILS protein topologies. In silico predicted transmembrane domain organization of PILS1-7 proteins. 2
3 Supplemental Figure 3 Phylogenetic tree of the PILS orthologues. PILS orthologues in several plant species are identified and represented in a 3
4 SUPPLEMENTARY INFORMATION phylogenetic tree using the Plaza2. online platform ( 31,43 ). Bootstrap values are given as percentages on the nodes. Taxonomy abbreviations: aly: Arabidopsis lyrata (light green) ; ath: Arabidopsis thaliana (light green); cpa: Carica papaya (green); gma: Glycine max (dark purple); lja: Lotus japonicas (light purple); mes: Manihot esculenta (light brown); mtr: Medicago truncatula (purple); osa: Oryza sativa (blue) spp. Japonica; osaindica: Oryza sativa spp. Indica (dark blue); ppa: Physcomitrella patens (red); ptr: Populus trichocarpa (dark brown); rco: Ricinus communis; sbi: Sorghum bicolor (yellow); vvi: Vitis vinifera (dark green); zma: Zea mays (yellow); mdo: Malus domestica; cre: Chlamidomonas reinhardtii (dark gray); mrcc299: Micromonas sp. RCC299 (dark bordeaux); ota: Ostreococcus tauri (bordeaux); smo: Selaginella moellendorfii (orange); vca: Volvox carteri (light gray); olu: Ostreococcus lucimarinus (light bordeaux); bdi: Brachypodium distachyon (dark yellow) 3 2 PILS expression 1h NAA 3h NAA 6h NAA Fold change 1 PILS2 PILS3 PILS4 PILS5 PILS6 PILS7 Supplemental Figure 4 qpcr analysis of auxin induced PILS expression. The effect of 1μM NAA treatment for 1, 3 and 6 h on PILS expression levels was analyzed via quantitative RT-PCR. Graph depicts fold change in PILS expression upon auxin treatment compared to PILS expression levels in untreated seedlings. (n=3 repetitions) Error bars depict s.e.m.. 4
5 pils2-1 pils2-2 ATG TGA pils2-1 pils2-2 WT 5 3 PILS 2 PILS2 EIF4a pils5-1 ATG pils5-2 TGA pils5-2 WT 5 3 PILS5 PILS 5.5 kb EIF4a Promoter UTR Exon Intron Supplemental Figure 5 PILS2 and PILS5 gene structure. The PILS2 and PILS5 genes contain 1 and 9 exons, respectively. The analyzed T- DNA insertions are depicted in the figure (arrow heads). The PILS2 and PILS5 expression levels in the respective insertion lines were analysed by RT-PCR (given in the right panels). Arrows depict the annealing position of the used RT-PCR primers. pils2-1 and pils2-2 showed a strong reduction in transcription level. We did not detect PILS5 mrna in the pils5-2, suggesting a full knock-out mutant. In contrast, PILS5 mrna levels in the pils5-1 mutant where similar to wild type (not shown). Scale bar, 5bp. 5
6 SUPPLEMENTARY INFORMATION Col--WT pils2-2 a b Lateral root pdr5::rfp Main root pdr5::gfp WT pils2-2 pils5-2 WT p35s::pils5::gfp 25 Supplemental Figure 6 Nuclear auxin signaling in pils loss- and gain-of-function mutants. a,b Analysis of the auxin reporter pdr5rev::gfp (a) or pdr5rev::mrfp1er (b) expression reveals no change in auxin signaling in the main root of pils2-2 and pils5-2 single mutants compared to wild type (WT) (a). Lateral roots of seedlings overexpressing PILS5 display decreased auxin signaling compared to WT. Representative pictures are shown and a color-coded heat map (black to white) was used to visualize (low to high) pdr5rev::mrfp1er and pdr5rev::gfp signal intensity. 6
7 a pexp7a::gfp/gus pexp7a::pils1::gfp pexp7a::pils3::gfp b pexp7a::gfp/gus pexp7a::pils1:gfp pexp7a::pils3:gfp c.5.4 Root hair length (mm).3.2 *** ***.1 pexp7a:: GFP/GUS pexp7a:: PILS1::GFP pexp7a:: PILS3::GFP Supplemental Figure 7: Root hair cell-specific PILS activity represses root hair growth. a, NLS-GFP::GUS (left), PILS1::GFP (middle) and PILS3::GFP (right) in root hairs (green). Propidium iodide staining in red. b-c, Root hair specific pexp7a::pils1::gfp and pexp7a::pils3::gfp expression represses root hair growth. c, Graph depicts the root hair length of seedlings expressing pexp7a::gfp/gus or pexp7a::pils1/3::gfp in the root hair cells (n=2 seedlings with 4 counted root hairs in total). Error bars represent s.e.m.. 7
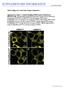
8 Supplemental Figure 8 SUPPLEMENTARY INFORMATION Tobacco BY-2 cells A.thaliana p35s::pils1::gfp p35s::pils1::rfp p35s::pils2::gfp p35s::pils2::rfp p35s::pils3::gfp p35s::pils3::rfp p35s::pils5::gfp p35s::pils5::rfp used to visualize (low to high) pdr5rev::mrfp1er and pdr5rev::gfp signal intensity. p35s::pils6::gfp p35s::pils6::rfp Supplemental Figure 7: Root hair cell-specific PILS activity represses root hair growth. a, NLS-GFP::GUS (left), PILS1::GFP (middle) and PILS3::GFP (right) in root hairs (green). Propidium p35s::pils7::gfp iodide staining in red. p35s::pils7::rfp b-c, Root hair specific pexp7a::pils1::gfp and pexp7a::pils3::gfp expression represses root hair growth. c, Graph depicts the root hair length of seedlings expressing pexp7a::gfp/gus or pexp7a::pils1/3::gfp in the root hair cells (n=2 seedlings with 4 counted root hairs in total). Error bars represent s.e.m.. Supplemental Figure 8 PILS intracellular localization. PILS(1-3)::GFP and PILS(5-7)::GFP fusion proteins reveal intracellular PILS localization to the ER as shown by live cell imaging of p35s::pils::gfp in A.thaliana root cells (left panel). Propidium iodide stained cell walls in red. Transient co-expression of PILS(1-3)::RFP and PILS(5-7)::RFP protein fusions and the ER- 8 W W W. N A T U R E. C O M / N A T U R E
9 A.thaliana root cells (left panel). Propidium iodide stained cell walls in red. Transient co-expression of PILS(1-3)::RFP and PILS(5-7)::RFP protein fusions and the ERmarker in tobacco BY-2 cells shows complete colocalization (right panel). PILS4::GFP/RFP did not show detectable fluorescence (not shown). Scale bar, 1µm. 4 *** 3 LR/cm 2 1 pils2-2 pils2-2 pils5-2 pils2-2 pils5-2 +pp5p5::gfp Supplemental Figure 9 pils2-2 pils5-2 mutant complementation. PILS5 expression under its endogenous promoter complements the enhanced lateral root density phenotype of the pils2-2 pils5-2 loss-of-function mutant (n=4 seedlings). Error bars represent s.e.m. Student t-test P-values: *P<.5, **P<.1 ***P<
10 SUPPLEMENTARY INFORMATION a A. thaliana protoplasts [ H]IAA export (% of initial export) WT pils2-2 pils5-2 pils2-2 pils t (min.) b S.cerevisiae Organic acid accumulation vector control PILS3-HA PILS7-HA [ 3 14 H ]IAA [ C ]BA c pgpd::pils3::gfp pgpd::pils7::gfp Supplemental Figure 1 PILS-dependent auxin accumulation in plant and non-plant cells a, Arabidopsis protoplasts of pils2-2 pils5-2 mutant leaves show higher 3 H-IAA export compared to protoplasts from wild type (WT) leaves (n=3 repetitions). Error bars represent s.e.m. b, Auxin accumulation assay in the Saccharomyces cerevisiae yeast cells. Cells transformed with pgpd::pils3::ha or pgpd::pils7::ha display higher auxin but not benzoic acid accumulation compared to cells transformed with an empty vector (n=3 repetitions). c, PILS3::GFP and PILS7::GFP localization in Saccharomyces cerevisiae. Error bars represent s.e.m.. 1
11 References for supplemental figures: Ludwig-Müller, J., Auxin conjugates: their role for plant development and in the evolution of land plants. J. Exp. Bot 62, (211). Ludwig-Muller, J., Julke, S., Bierfreund, N.M., Decker, E.L., & Reski, R., Moss (Physcomitrella patens) GH3 proteins act in auxin homeostasis. New Phytol 181, (29). LeClere, S., Tellez, R., Rampey, R.A., Matsuda, S.P., & Bartel, B., Characterization of a family of IAA-amino acid conjugate hydrolases from Arabidopsis. J Biol Chem 277, (22). Campanella, J.J., Larko, D., & Smalley, J., A molecular phylogenomic analysis of the ILR1-like family of IAA amidohydrolase genes. Comp Funct Genomics 4, (23). Kriechbaumer, V., Wang, P., Hawes, C., & Abell, B.M., Alternative splicing of the auxin biosynthesis gene YUCCA4 determines its subcellular compartmentation, Plant J. doi: /j X x (212) Jones, A.M. & Herman, E.M., KDEL-Containing Auxin-Binding Protein Is Secreted to the Plasma Membrane and Cell Wall. Plant Physiol 11, (1993). Henderson, J., Bauly, JM., Ashford DA, Oliver SC, Hawes CR et al., Retention of maize auxin-binding protein in the endoplasmic reticulum: quantifying escape and the role of auxin. planta 22, (1997). 43 Zmasek, CM., Eddy, SR., ATV: display and manipulation of annotated phylogenetic trees. Bioinformatics. 17, (21). 11
Cao, J, K Schneeberger, S Ossowski, et al Whole genome sequencing of multiple Arabidopsis thaliana populations. Nat Genet 43:
 Figure S1. Syntenic map of SAE1B duplication. We have used the nucleotide sequences of Arabidopsis thaliana Col-0 gene tandem duplicates AT5G50580 and AT5G506800 as queries in independent BLASTN searches
Figure S1. Syntenic map of SAE1B duplication. We have used the nucleotide sequences of Arabidopsis thaliana Col-0 gene tandem duplicates AT5G50580 and AT5G506800 as queries in independent BLASTN searches
Supplemental Figure 1. Comparison of Tiller Bud Formation between the Wild Type and d27. (A) and (B) Longitudinal sections of shoot apex in wild-type
 A B 2 3 3 2 1 1 Supplemental Figure 1. Comparison of Tiller Bud Formation between the Wild Type and d27. (A) and (B) Longitudinal sections of shoot apex in wild-type (A) and d27 (B) seedlings at the four
A B 2 3 3 2 1 1 Supplemental Figure 1. Comparison of Tiller Bud Formation between the Wild Type and d27. (A) and (B) Longitudinal sections of shoot apex in wild-type (A) and d27 (B) seedlings at the four
Supplemental Data. Perea-Resa et al. Plant Cell. (2012) /tpc
 Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature13082 Supplementary Table 1. Examination of nectar production in wild-type and atsweet9 flowers. No. of flowers with detectable nectar out of the total observed
SUPPLEMENTARY INFORMATION doi:10.1038/nature13082 Supplementary Table 1. Examination of nectar production in wild-type and atsweet9 flowers. No. of flowers with detectable nectar out of the total observed
Supplemental Table 1. Primers used for cloning and PCR amplification in this study
 Supplemental Table 1. Primers used for cloning and PCR amplification in this study Target Gene Primer sequence NATA1 (At2g393) forward GGG GAC AAG TTT GTA CAA AAA AGC AGG CTT CAT GGC GCC TCC AAC CGC AGC
Supplemental Table 1. Primers used for cloning and PCR amplification in this study Target Gene Primer sequence NATA1 (At2g393) forward GGG GAC AAG TTT GTA CAA AAA AGC AGG CTT CAT GGC GCC TCC AAC CGC AGC
Supplemental Data. Gao et al. (2012). Plant Cell /tpc
 Supplemental Figure 1. Plant EMP Proteins. (A) The Accession numbers of the 12 EMP members from Arabidopsis. (B) Phylogenetic analysis of EMP proteins from Arabidopsis, human and yeast using the Mac Vector
Supplemental Figure 1. Plant EMP Proteins. (A) The Accession numbers of the 12 EMP members from Arabidopsis. (B) Phylogenetic analysis of EMP proteins from Arabidopsis, human and yeast using the Mac Vector
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 3
 Supplementary Figure 3 7.0 Col Kas-1 Line FTH1A 8.4 F3PII3 8.9 F26H11 ATQ1 T9I22 PLS8 F26B6-B 9.6 F27L4 9.81 F27D4 9.92 9.96 10.12 10.14 10.2 11.1 0.5 Mb T1D16 Col % RGR 83.3 101 227 93.5 75.9 132 90 375
Supplementary Figure 3 7.0 Col Kas-1 Line FTH1A 8.4 F3PII3 8.9 F26H11 ATQ1 T9I22 PLS8 F26B6-B 9.6 F27L4 9.81 F27D4 9.92 9.96 10.12 10.14 10.2 11.1 0.5 Mb T1D16 Col % RGR 83.3 101 227 93.5 75.9 132 90 375
Genome-Wide Computational Prediction and Analysis of Core Promoter Elements across Plant Monocots and Dicots
 Genome-Wide Computational Prediction and Analysis of Core Promoter Elements across Plant Monocots and Dicots Sunita Kumari 1, Doreen Ware 1,2 * 1 Cold Spring Harbor Laboratory, Cold Spring Harbor, New
Genome-Wide Computational Prediction and Analysis of Core Promoter Elements across Plant Monocots and Dicots Sunita Kumari 1, Doreen Ware 1,2 * 1 Cold Spring Harbor Laboratory, Cold Spring Harbor, New
SUPPLEMENTARY INFORMATION
 reverse 3175 3175 F L C 318 318 3185 3185 319 319 3195 3195 315 8 1 315 3155 315 317 Supplementary Figure 3. Stability of expression of the GFP sensor constructs return to warm conditions. Semi-quantitative
reverse 3175 3175 F L C 318 318 3185 3185 319 319 3195 3195 315 8 1 315 3155 315 317 Supplementary Figure 3. Stability of expression of the GFP sensor constructs return to warm conditions. Semi-quantitative
AtTIL-P91V. AtTIL-P92V. AtTIL-P95V. AtTIL-P98V YFP-HPR
 Online Resource 1. Primers used to generate constructs AtTIL-P91V, AtTIL-P92V, AtTIL-P95V and AtTIL-P98V and YFP(HPR) using overlapping PCR. pentr/d- TOPO-AtTIL was used as template to generate the constructs
Online Resource 1. Primers used to generate constructs AtTIL-P91V, AtTIL-P92V, AtTIL-P95V and AtTIL-P98V and YFP(HPR) using overlapping PCR. pentr/d- TOPO-AtTIL was used as template to generate the constructs
AUXIN CONJUGATE HYDROLYSIS DURING PLANT-MICROBE INTERACTION AND EVOLUTION
 AUXIN CONJUGATE HYDROLYSIS DURING PLANT-MICROBE INTERACTION AND EVOLUTION J. Ludwig-Müller 1, A. Schuller 1, A.F. Olajide 2, V. Bakllamaja 2, J.J. Campanella 2 ABSTRACT Plants regulate auxin balance through
AUXIN CONJUGATE HYDROLYSIS DURING PLANT-MICROBE INTERACTION AND EVOLUTION J. Ludwig-Müller 1, A. Schuller 1, A.F. Olajide 2, V. Bakllamaja 2, J.J. Campanella 2 ABSTRACT Plants regulate auxin balance through
Bioinformatics tools to analyze complex genomes. Yves Van de Peer Ghent University/VIB
 Bioinformatics tools to analyze complex genomes Yves Van de Peer Ghent University/VIB Detecting colinearity and large-scale gene duplications A 1 2 3 4 5 6 7 8 9 10 11 Speciation/Duplicatio n S1 S2 1
Bioinformatics tools to analyze complex genomes Yves Van de Peer Ghent University/VIB Detecting colinearity and large-scale gene duplications A 1 2 3 4 5 6 7 8 9 10 11 Speciation/Duplicatio n S1 S2 1
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
Genome-wide discovery of G-quadruplex forming sequences and their functional
 *Correspondence and requests for materials should be addressed to R.G. (rohini@nipgr.ac.in) Genome-wide discovery of G-quadruplex forming sequences and their functional relevance in plants Rohini Garg*,
*Correspondence and requests for materials should be addressed to R.G. (rohini@nipgr.ac.in) Genome-wide discovery of G-quadruplex forming sequences and their functional relevance in plants Rohini Garg*,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09606 Chen Li-Qing et al. A novel class of sugar transporters. Supplementary Figure 1. Confocal imaging of FRET sensor localization in HEK293T cells. The subcellular localization of the
doi:10.1038/nature09606 Chen Li-Qing et al. A novel class of sugar transporters. Supplementary Figure 1. Confocal imaging of FRET sensor localization in HEK293T cells. The subcellular localization of the
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011) /tpc
 Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011). 10.1105/tpc.110.080788 Supplemental Figure S1. Phylogenetic tree of MYC2-related proteins from Arabidopsis and other plants. Phenogram representation
Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011). 10.1105/tpc.110.080788 Supplemental Figure S1. Phylogenetic tree of MYC2-related proteins from Arabidopsis and other plants. Phenogram representation
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1121356/dc1 Supporting Online Material for Polar PIN Localization Directs Auxin Flow in Plants Justyna Wiśniewska, Jian Xu, Daniela Seifertová, Philip B. Brewer, Kamil
www.sciencemag.org/cgi/content/full/1121356/dc1 Supporting Online Material for Polar PIN Localization Directs Auxin Flow in Plants Justyna Wiśniewska, Jian Xu, Daniela Seifertová, Philip B. Brewer, Kamil
Nature Genetics: doi: /ng Supplementary Figure 1. ssp mutant phenotypes in a functional SP background.
 Supplementary Figure 1 ssp mutant phenotypes in a functional SP background. (a,b) Statistical comparisons of primary and sympodial shoot flowering times as determined by mean values for leaf number on
Supplementary Figure 1 ssp mutant phenotypes in a functional SP background. (a,b) Statistical comparisons of primary and sympodial shoot flowering times as determined by mean values for leaf number on
23-. Shoot and root development depend on ratio of IAA/CK
 Balance of Hormones regulate growth and development Environmental factors regulate hormone levels light- e.g. phototropism gravity- e.g. gravitropism temperature Mode of action of each hormone 1. Signal
Balance of Hormones regulate growth and development Environmental factors regulate hormone levels light- e.g. phototropism gravity- e.g. gravitropism temperature Mode of action of each hormone 1. Signal
Supplemental Data. Chen and Thelen (2010). Plant Cell /tpc
 Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Biological Roles of Cytokinins
 Direct Control of Shoot Meristem Activity by a Cytokinin-Activating Enzyme By Kurakawa et. Al. Published in Nature Presented by Boyana Grigorova Biological Roles of Cytokinins Cytokinins are positive regulators
Direct Control of Shoot Meristem Activity by a Cytokinin-Activating Enzyme By Kurakawa et. Al. Published in Nature Presented by Boyana Grigorova Biological Roles of Cytokinins Cytokinins are positive regulators
SUPPLEMENTARY INFORMATION
 doi:1.138/nature11394 Supplementary Note Our study is based on two different tracriptome indices. Both combine tracriptional information with an important evolutionary parameter. For the tracriptome age
doi:1.138/nature11394 Supplementary Note Our study is based on two different tracriptome indices. Both combine tracriptional information with an important evolutionary parameter. For the tracriptome age
CHAPTER 3. Cell Structure and Genetic Control. Chapter 3 Outline
 CHAPTER 3 Cell Structure and Genetic Control Chapter 3 Outline Plasma Membrane Cytoplasm and Its Organelles Cell Nucleus and Gene Expression Protein Synthesis and Secretion DNA Synthesis and Cell Division
CHAPTER 3 Cell Structure and Genetic Control Chapter 3 Outline Plasma Membrane Cytoplasm and Its Organelles Cell Nucleus and Gene Expression Protein Synthesis and Secretion DNA Synthesis and Cell Division
Supplementary Information for: The genome of the extremophile crucifer Thellungiella parvula
 Supplementary Information for: The genome of the extremophile crucifer Thellungiella parvula Maheshi Dassanayake 1,9, Dong-Ha Oh 1,9, Jeffrey S. Haas 1,2, Alvaro Hernandez 3, Hyewon Hong 1,4, Shahjahan
Supplementary Information for: The genome of the extremophile crucifer Thellungiella parvula Maheshi Dassanayake 1,9, Dong-Ha Oh 1,9, Jeffrey S. Haas 1,2, Alvaro Hernandez 3, Hyewon Hong 1,4, Shahjahan
Evaluation of Genome Sequencing Quality in Selected Plant Species Using Expressed Sequence Tags
 Evaluation of Genome Sequencing Quality in Selected Plant Species Using Expressed Sequence Tags Lingfei Shangguan 1, Jian Han 1, Emrul Kayesh 1, Xin Sun 1, Changqing Zhang 2, Tariq Pervaiz 1, Xicheng Wen
Evaluation of Genome Sequencing Quality in Selected Plant Species Using Expressed Sequence Tags Lingfei Shangguan 1, Jian Han 1, Emrul Kayesh 1, Xin Sun 1, Changqing Zhang 2, Tariq Pervaiz 1, Xicheng Wen
** LCA LCN PCA
 % of wild type value % of wild type value a 12 1 8 2 b 12 1 8 2 LCA LCN PCA Col- sod3-1 Supplementary Figure 1 sod3-1 influences cell proliferation. (a) Fifth leaf cell area (LCA) and leaf cell number
% of wild type value % of wild type value a 12 1 8 2 b 12 1 8 2 LCA LCN PCA Col- sod3-1 Supplementary Figure 1 sod3-1 influences cell proliferation. (a) Fifth leaf cell area (LCA) and leaf cell number
Supplemental Data. Yang et al. (2012). Plant Cell /tpc
 Supplemental Figure 1. Mature flowers of P. heterotricha. (A) An inflorescence of P. heterotricha showing the front view of a zygomorphic flower characterized by two small dorsal petals and only two fertile
Supplemental Figure 1. Mature flowers of P. heterotricha. (A) An inflorescence of P. heterotricha showing the front view of a zygomorphic flower characterized by two small dorsal petals and only two fertile
. Supplementary Information
 . Supplementary Information Supplementary Figure S1. Mature embryo sac observations. Supplementary Figure S2. STT observations. Supplementary Figure S3. Comparison of the PTB1 cdna with that of the mutant.
. Supplementary Information Supplementary Figure S1. Mature embryo sac observations. Supplementary Figure S2. STT observations. Supplementary Figure S3. Comparison of the PTB1 cdna with that of the mutant.
Supporting Information
 Supporting Information Rossig et al. 10.1073/pnas.1319648110 SI Materials and Methods Athp20 and Athp30 Mutant Characterization and Analysis of the Athp30/30-2 RNAi Lines. Athp20;1 and Athp20;2 (SALK_020671
Supporting Information Rossig et al. 10.1073/pnas.1319648110 SI Materials and Methods Athp20 and Athp30 Mutant Characterization and Analysis of the Athp30/30-2 RNAi Lines. Athp20;1 and Athp20;2 (SALK_020671
Supplementary Figure 1. Phenotype of the HI strain.
 Supplementary Figure 1. Phenotype of the HI strain. (A) Phenotype of the HI and wild type plant after flowering (~1month). Wild type plant is tall with well elongated inflorescence. All four HI plants
Supplementary Figure 1. Phenotype of the HI strain. (A) Phenotype of the HI and wild type plant after flowering (~1month). Wild type plant is tall with well elongated inflorescence. All four HI plants
Supplemental Data. Wang et al. (2014). Plant Cell /tpc
 Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Lipid transfer proteins confer resistance to trichothecenes
 Lipid transfer proteins confer resistance to trichothecenes John McLaughlin and Anwar Bin-Umer Tumer Laboratory National Fusarium Head Blight Forum December 6th, 2012 FY09-11: Identify trichothecene resistance
Lipid transfer proteins confer resistance to trichothecenes John McLaughlin and Anwar Bin-Umer Tumer Laboratory National Fusarium Head Blight Forum December 6th, 2012 FY09-11: Identify trichothecene resistance
Genome-wide Identification of Lineage Specific Genes in Arabidopsis, Oryza and Populus
 Genome-wide Identification of Lineage Specific Genes in Arabidopsis, Oryza and Populus Xiaohan Yang Sara Jawdy Timothy Tschaplinski Gerald Tuskan Environmental Sciences Division Oak Ridge National Laboratory
Genome-wide Identification of Lineage Specific Genes in Arabidopsis, Oryza and Populus Xiaohan Yang Sara Jawdy Timothy Tschaplinski Gerald Tuskan Environmental Sciences Division Oak Ridge National Laboratory
7.06 Cell Biology EXAM #3 April 21, 2005
 7.06 Cell Biology EXAM #3 April 21, 2005 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #3 April 21, 2005 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
Origin and diversification of leucine-rich repeat receptor-like protein kinase (LRR-RLK) genes in plants
 Liu et al. BMC Evolutionary Biology (2017) 17:47 DOI 10.1186/s12862-017-0891-5 RESEARCH ARTICLE Origin and diversification of leucine-rich repeat receptor-like protein kinase (LRR-RLK) genes in plants
Liu et al. BMC Evolutionary Biology (2017) 17:47 DOI 10.1186/s12862-017-0891-5 RESEARCH ARTICLE Origin and diversification of leucine-rich repeat receptor-like protein kinase (LRR-RLK) genes in plants
Cellular Neuroanatomy I The Prototypical Neuron: Soma. Reading: BCP Chapter 2
 Cellular Neuroanatomy I The Prototypical Neuron: Soma Reading: BCP Chapter 2 Functional Unit of the Nervous System The functional unit of the nervous system is the neuron. Neurons are cells specialized
Cellular Neuroanatomy I The Prototypical Neuron: Soma Reading: BCP Chapter 2 Functional Unit of the Nervous System The functional unit of the nervous system is the neuron. Neurons are cells specialized
JMJ14-HA. Col. Col. jmj14-1. jmj14-1 JMJ14ΔFYR-HA. Methylene Blue. Methylene Blue
 Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated
 Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated form by recombinant UGT74E2. The naturally occurring auxin
Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated form by recombinant UGT74E2. The naturally occurring auxin
Penghui Li, Beibei Chen, Gaoyang Zhang, Longxiang Chen, Qiang Dong, Jiangqi Wen, Kirankumar S. Mysore and Jian Zhao
 New Phytologist Supporting Information Regulation of anthocyanin and proanthocyanidin biosynthesis by Medicago truncatula bhlh transcription factor MtTT8 Penghui Li, Beibei Chen, Gaoyang Zhang, Longxiang
New Phytologist Supporting Information Regulation of anthocyanin and proanthocyanidin biosynthesis by Medicago truncatula bhlh transcription factor MtTT8 Penghui Li, Beibei Chen, Gaoyang Zhang, Longxiang
** * * * Col-0 cau1 CAU1. Actin2 CAS. Actin2. Supplemental Figure 1. CAU1 affects calcium accumulation.
 Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Supplemental Data. Fu et al. Plant Cell. (213). 1.115/tpc.113.113886 A 5 4 3 * Col- cau1 B 4 3 2 Col- cau1 ** * * ** C 2 1 25 2 15 1 5 Shoots Roots *
Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Supplemental Data. Fu et al. Plant Cell. (213). 1.115/tpc.113.113886 A 5 4 3 * Col- cau1 B 4 3 2 Col- cau1 ** * * ** C 2 1 25 2 15 1 5 Shoots Roots *
Regulatory Change in YABBY-like Transcription Factor Led to Evolution of Extreme Fruit Size during Tomato Domestication
 SUPPORTING ONLINE MATERIALS Regulatory Change in YABBY-like Transcription Factor Led to Evolution of Extreme Fruit Size during Tomato Domestication Bin Cong, Luz Barrero, & Steven Tanksley 1 SUPPORTING
SUPPORTING ONLINE MATERIALS Regulatory Change in YABBY-like Transcription Factor Led to Evolution of Extreme Fruit Size during Tomato Domestication Bin Cong, Luz Barrero, & Steven Tanksley 1 SUPPORTING
Potato Genome Analysis
 Potato Genome Analysis Xin Liu Deputy director BGI research 2016.1.21 WCRTC 2016 @ Nanning Reference genome construction???????????????????????????????????????? Sequencing HELL RIEND WELCOME BGI ZHEN LLOFRI
Potato Genome Analysis Xin Liu Deputy director BGI research 2016.1.21 WCRTC 2016 @ Nanning Reference genome construction???????????????????????????????????????? Sequencing HELL RIEND WELCOME BGI ZHEN LLOFRI
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Name: TF: Section Time: LS1a ICE 5. Practice ICE Version B
 Name: TF: Section Time: LS1a ICE 5 Practice ICE Version B 1. (8 points) In addition to ion channels, certain small molecules can modulate membrane potential. a. (4 points) DNP ( 2,4-dinitrophenol ), as
Name: TF: Section Time: LS1a ICE 5 Practice ICE Version B 1. (8 points) In addition to ion channels, certain small molecules can modulate membrane potential. a. (4 points) DNP ( 2,4-dinitrophenol ), as
Browsing Genomic Information with Ensembl Plants
 Browsing Genomic Information with Ensembl Plants Etienne de Villiers, PhD (Adapted from slides by Bert Overduin EMBL-EBI) Outline of workshop Brief introduction to Ensembl Plants History Content Tutorial
Browsing Genomic Information with Ensembl Plants Etienne de Villiers, PhD (Adapted from slides by Bert Overduin EMBL-EBI) Outline of workshop Brief introduction to Ensembl Plants History Content Tutorial
Development 143: doi: /dev : Supplementary information
 Supplementary Materials and Methods Plant materials The mutants and transgenic plants used in the present study were as follows: E361 (from Alex Webb s laboratory); tmm-1, ptmm::tmm-gfp and flp-1 (from
Supplementary Materials and Methods Plant materials The mutants and transgenic plants used in the present study were as follows: E361 (from Alex Webb s laboratory); tmm-1, ptmm::tmm-gfp and flp-1 (from
Introduction to cells
 Almen Cellebiologi Introduction to cells 1. Unity and diversity of cells 2. Microscopes and visualization of cells 3. Prokaryotic cells, eubacteria and archaea 4. Eucaryotic cells, nucleus, mitochondria
Almen Cellebiologi Introduction to cells 1. Unity and diversity of cells 2. Microscopes and visualization of cells 3. Prokaryotic cells, eubacteria and archaea 4. Eucaryotic cells, nucleus, mitochondria
PLAZA: A Comparative Genomics Resource to Study Gene and Genome Evolution in Plants W
 The Plant Cell, Vol. 21: 3718 3731, December 2009, www.plantcell.org ã 2009 American Society of Plant Biologists Special Series on Large-Scale Biology PLAZA: A Comparative Genomics Resource to Study Gene
The Plant Cell, Vol. 21: 3718 3731, December 2009, www.plantcell.org ã 2009 American Society of Plant Biologists Special Series on Large-Scale Biology PLAZA: A Comparative Genomics Resource to Study Gene
Supplementary Figure 1. Number of CC- and TIR- type NBS- LRR genes and presence of mir482/2118 on sequenced plant genomes.
 Number of CC- NBS and CC- NBS- LRR R- genes Number of TIR- NBS and TIR- NBS- LRR R- genes 0 50 100 150 200 250 0 50 100 150 200 250 300 350 400 450 mir482 and mir2118 Cajanus cajan Glycine max Hevea brasiliensis
Number of CC- NBS and CC- NBS- LRR R- genes Number of TIR- NBS and TIR- NBS- LRR R- genes 0 50 100 150 200 250 0 50 100 150 200 250 300 350 400 450 mir482 and mir2118 Cajanus cajan Glycine max Hevea brasiliensis
Supplementary Material
 Supplementary Material Supplementary Table S1. Genomes available in build 47 Supplementary Table S2. Counts of putative contiguous gene split models in 39 plant reference genomes in build 47 Supplementary
Supplementary Material Supplementary Table S1. Genomes available in build 47 Supplementary Table S2. Counts of putative contiguous gene split models in 39 plant reference genomes in build 47 Supplementary
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Nature Neuroscience: doi: /nn.2717
 Supplementary Fig. 1. Dendrite length is not secondary to body length. Dendrite growth proceeds independently of the rate of body growth and decreases in rate in adults. n 20 on dendrite measurement, n
Supplementary Fig. 1. Dendrite length is not secondary to body length. Dendrite growth proceeds independently of the rate of body growth and decreases in rate in adults. n 20 on dendrite measurement, n
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Supporting Information
 Supporting Information Cao et al. 10.1073/pnas.1306220110 Gram - bacteria Hemolymph Cytoplasm PGRP-LC TAK1 signalosome Imd dfadd Dredd Dnr1 Ikk signalosome P Relish Nucleus AMP and effector genes Fig.
Supporting Information Cao et al. 10.1073/pnas.1306220110 Gram - bacteria Hemolymph Cytoplasm PGRP-LC TAK1 signalosome Imd dfadd Dredd Dnr1 Ikk signalosome P Relish Nucleus AMP and effector genes Fig.
Regulation of Phosphate Homeostasis by microrna in Plants
 Regulation of Phosphate Homeostasis by microrna in Plants Tzyy-Jen Chiou 1 *, Kyaw Aung 1,2, Shu-I Lin 1,3, Chia-Chune Wu 1, Su-Fen Chiang 1, and Chun-Lin Su 1 Abstract Upon phosphate (Pi) starvation,
Regulation of Phosphate Homeostasis by microrna in Plants Tzyy-Jen Chiou 1 *, Kyaw Aung 1,2, Shu-I Lin 1,3, Chia-Chune Wu 1, Su-Fen Chiang 1, and Chun-Lin Su 1 Abstract Upon phosphate (Pi) starvation,
7.06 Cell Biology EXAM #3 KEY
 7.06 Cell Biology EXAM #3 KEY May 2, 2006 This is an OPEN BOOK exam, and you are allowed access to books, a calculator, and notes BUT NOT computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #3 KEY May 2, 2006 This is an OPEN BOOK exam, and you are allowed access to books, a calculator, and notes BUT NOT computers or any other types of electronic devices. Please write
!"#$%&'%()*%+*,,%-&,./*%01%02%/*/3452*%3&.26%&4752*,,*1%%
 !"#$%&'%()*%+*,,%-&,./*%01%02%/*/3452*%3&.26%&4752*,,*1%% !"#$%&'(")*++*%,*'-&'./%/,*#01#%-2)#3&)/% 4'(")*++*% % %5"0)%-2)#3&) %%% %67'2#72'*%%%%%%%%%%%%%%%%%%%%%%%4'(")0/./% % 8$+&'&,+"/7 % %,$&7&/9)7$*/0/%%%%%%%%%%
!"#$%&'%()*%+*,,%-&,./*%01%02%/*/3452*%3&.26%&4752*,,*1%% !"#$%&'(")*++*%,*'-&'./%/,*#01#%-2)#3&)/% 4'(")*++*% % %5"0)%-2)#3&) %%% %67'2#72'*%%%%%%%%%%%%%%%%%%%%%%%4'(")0/./% % 8$+&'&,+"/7 % %,$&7&/9)7$*/0/%%%%%%%%%%
Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering
 Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering by Valverde et. Al Published in Science 2004 Presented by Boyana Grigorova CBMG 688R Feb. 12, 2007 Circadian Rhythms: The Clock Within
Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering by Valverde et. Al Published in Science 2004 Presented by Boyana Grigorova CBMG 688R Feb. 12, 2007 Circadian Rhythms: The Clock Within
Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport
 Ph.D. thesis Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport Zsigmond Laura Supervisor: Dr. Szabados László Arabidopsis Molecular Genetic Group Institute of Plant
Ph.D. thesis Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport Zsigmond Laura Supervisor: Dr. Szabados László Arabidopsis Molecular Genetic Group Institute of Plant
Chapter 12: Intracellular sorting
 Chapter 12: Intracellular sorting Principles of intracellular sorting Principles of intracellular sorting Cells have many distinct compartments (What are they? What do they do?) Specific mechanisms are
Chapter 12: Intracellular sorting Principles of intracellular sorting Principles of intracellular sorting Cells have many distinct compartments (What are they? What do they do?) Specific mechanisms are
Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field.
 Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field. A, Phenotypes of 30-day old wild-type (WT) and abci8 mutant plants grown in a paddy field under normal sunny
Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field. A, Phenotypes of 30-day old wild-type (WT) and abci8 mutant plants grown in a paddy field under normal sunny
L3.1: Circuits: Introduction to Transcription Networks. Cellular Design Principles Prof. Jenna Rickus
 L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
Nature Genetics: doi: /ng Supplementary Figure 1. Icm/Dot secretion system region I in 41 Legionella species.
 Supplementary Figure 1 Icm/Dot secretion system region I in 41 Legionella species. Homologs of the effector-coding gene lega15 (orange) were found within Icm/Dot region I in 13 Legionella species. In four
Supplementary Figure 1 Icm/Dot secretion system region I in 41 Legionella species. Homologs of the effector-coding gene lega15 (orange) were found within Icm/Dot region I in 13 Legionella species. In four
Cytokinin. Fig Cytokinin needed for growth of shoot apical meristem. F Cytokinin stimulates chloroplast development in the dark
 Cytokinin Abundant in young, dividing cells Shoot apical meristem Root apical meristem Synthesized in root tip, developing embryos, young leaves, fruits Transported passively via xylem into shoots from
Cytokinin Abundant in young, dividing cells Shoot apical meristem Root apical meristem Synthesized in root tip, developing embryos, young leaves, fruits Transported passively via xylem into shoots from
GFP GAL bp 3964 bp
 Supplemental Data. Møller et al. (2009) Shoot Na + exclusion and increased salinity tolerance engineered by cell type-specific alteration of Na + transport in Arabidopsis Supplemental Figure 1. Salt-sensitive
Supplemental Data. Møller et al. (2009) Shoot Na + exclusion and increased salinity tolerance engineered by cell type-specific alteration of Na + transport in Arabidopsis Supplemental Figure 1. Salt-sensitive
The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas
 The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas Is there a Role for the CCM in Increasing Biological CO 2 Capture? Generalized
The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas Is there a Role for the CCM in Increasing Biological CO 2 Capture? Generalized
Cells and Passive Transport Study Guide
 Cells and Passive Transport Study Guide Success Criteria: - Complete - If multiple choice, answer has explanations - Quality answers/best answer possible 1. List the 2 types of active transport and the
Cells and Passive Transport Study Guide Success Criteria: - Complete - If multiple choice, answer has explanations - Quality answers/best answer possible 1. List the 2 types of active transport and the
Supplemental Figure 1. Comparisons of GC3 distribution computed with raw EST data, bi-beta fits and complete genome sequences for 6 species.
 Supplemental Figure 1. Comparisons of GC3 distribution computed with raw EST data, bi-beta fits and complete genome sequences for 6 species. Filled distributions: GC3 computed with raw EST data. Dashed
Supplemental Figure 1. Comparisons of GC3 distribution computed with raw EST data, bi-beta fits and complete genome sequences for 6 species. Filled distributions: GC3 computed with raw EST data. Dashed
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
Supplementary Information
 Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Cover Page. The handle holds various files of this Leiden University dissertation
 Cover Page The handle http://hdl.handle.net/1887/41480 holds various files of this Leiden University dissertation Author: Tleis, Mohamed Title: Image analysis for gene expression based phenotype characterization
Cover Page The handle http://hdl.handle.net/1887/41480 holds various files of this Leiden University dissertation Author: Tleis, Mohamed Title: Image analysis for gene expression based phenotype characterization
Eukaryotic vs. Prokaryotic genes
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
Actions of auxin. Hormones: communicating with chemicals History: Discovery of a growth substance (hormone- auxin)
 Hormones: communicating with chemicals History- discovery of plant hormone. Auxin Concepts of hormones Auxin levels are regulated by synthesis/degradation, transport, compartmentation, conjugation. Polar
Hormones: communicating with chemicals History- discovery of plant hormone. Auxin Concepts of hormones Auxin levels are regulated by synthesis/degradation, transport, compartmentation, conjugation. Polar
GENETIC ANALYSES OF ROOT SYSTEM DEVELOPMENT IN THE TOMATO CROP MODEL
 GENETIC ANALYSES OF ROOT SYSTEM DEVELOPMENT IN THE TOMATO CROP MODEL Kelsey Hoth 1 Dr. Maria Ivanchenko 2 Bioresourse Research 1, Department of Botany and Plant Physiology 2, Oregon State University, Corvallis,
GENETIC ANALYSES OF ROOT SYSTEM DEVELOPMENT IN THE TOMATO CROP MODEL Kelsey Hoth 1 Dr. Maria Ivanchenko 2 Bioresourse Research 1, Department of Botany and Plant Physiology 2, Oregon State University, Corvallis,
GENES AND CHROMOSOMES III. Lecture 5. Biology Department Concordia University. Dr. S. Azam BIOL 266/
 GENES AND CHROMOSOMES III Lecture 5 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION OPERONS Introduction All cells in a multi-cellular
GENES AND CHROMOSOMES III Lecture 5 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION OPERONS Introduction All cells in a multi-cellular
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
SOMNUS, a CCCH-Type Zinc Finger Protein in Arabidopsis, Negatively Regulates Light-Dependent Seed Germination Downstream of PIL5 W
 The Plant Cell, Vol. 20: 1260 1277, May 2008, www.plantcell.org ª 2008 American Society of Plant Biologists SOMNUS, a CCCH-Type Zinc Finger Protein in Arabidopsis, Negatively Regulates Light-Dependent
The Plant Cell, Vol. 20: 1260 1277, May 2008, www.plantcell.org ª 2008 American Society of Plant Biologists SOMNUS, a CCCH-Type Zinc Finger Protein in Arabidopsis, Negatively Regulates Light-Dependent
02/02/ Living things are organized. Analyze the functional inter-relationship of cell structures. Learning Outcome B1
 Analyze the functional inter-relationship of cell structures Learning Outcome B1 Describe the following cell structures and their functions: Cell membrane Cell wall Chloroplast Cytoskeleton Cytoplasm Golgi
Analyze the functional inter-relationship of cell structures Learning Outcome B1 Describe the following cell structures and their functions: Cell membrane Cell wall Chloroplast Cytoskeleton Cytoplasm Golgi
Last time: Obtaining information from a cloned gene
 Last time: Obtaining information from a cloned gene Objectives: 1. What is the biochemical role of the gene? 2. Where and when is the gene expressed (transcribed)? 3. Where and when is the protein made?
Last time: Obtaining information from a cloned gene Objectives: 1. What is the biochemical role of the gene? 2. Where and when is the gene expressed (transcribed)? 3. Where and when is the protein made?
Supplementary Methods
 Supplementary Methods Microarray analysis Grains of 7 DAP of the wild-type and gif1 were harvested for RNA preparation. Microarray analysis was performed with the Affymetrix (Santa Clara, CA) GeneChip
Supplementary Methods Microarray analysis Grains of 7 DAP of the wild-type and gif1 were harvested for RNA preparation. Microarray analysis was performed with the Affymetrix (Santa Clara, CA) GeneChip
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10534 Supplementary Fig. 1. Diagrammatic representation of the N-end rule pathway of targeted proteolysis (after Graciet and Wellmer 2010 9 ). Tertiary, secondary
SUPPLEMENTARY INFORMATION doi:10.1038/nature10534 Supplementary Fig. 1. Diagrammatic representation of the N-end rule pathway of targeted proteolysis (after Graciet and Wellmer 2010 9 ). Tertiary, secondary
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
9/11/18. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Related Courses He who asks is a fool for five minutes, but he who does not ask remains a fool forever.
 CSE 527 Computational Biology http://www.cs.washington.edu/527 Lecture 1: Overview & Bio Review Autumn 2004 Larry Ruzzo Related Courses He who asks is a fool for five minutes, but he who does not ask remains
CSE 527 Computational Biology http://www.cs.washington.edu/527 Lecture 1: Overview & Bio Review Autumn 2004 Larry Ruzzo Related Courses He who asks is a fool for five minutes, but he who does not ask remains
7.06 Problem Set
 7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
Supplemental Data. Hou et al. (2016). Plant Cell /tpc
 Supplemental Data. Hou et al. (216). Plant Cell 1.115/tpc.16.295 A Distance to 1 st nt of start codon Distance to 1 st nt of stop codon B Normalized PARE abundance 8 14 nt 17 nt Frame1 Arabidopsis inflorescence
Supplemental Data. Hou et al. (216). Plant Cell 1.115/tpc.16.295 A Distance to 1 st nt of start codon Distance to 1 st nt of stop codon B Normalized PARE abundance 8 14 nt 17 nt Frame1 Arabidopsis inflorescence
NAME: PERIOD: DATE: A View of the Cell. Use Chapter 8 of your book to complete the chart of eukaryotic cell components.
 NAME: PERIOD: DATE: A View of the Cell Use Chapter 8 of your book to complete the chart of eukaryotic cell components. Cell Part Cell Wall Centriole Chloroplast Cilia Cytoplasm Cytoskeleton Endoplasmic
NAME: PERIOD: DATE: A View of the Cell Use Chapter 8 of your book to complete the chart of eukaryotic cell components. Cell Part Cell Wall Centriole Chloroplast Cilia Cytoplasm Cytoskeleton Endoplasmic
Ph.D. thesis. Study of proline accumulation and transcriptional regulation of genes involved in this process in Arabidopsis thaliana
 Ph.D. thesis Study of proline accumulation and transcriptional regulation of genes involved in this process in Arabidopsis thaliana Written by: Edit Ábrahám Temesváriné Supervisors: Dr. László Szabados
Ph.D. thesis Study of proline accumulation and transcriptional regulation of genes involved in this process in Arabidopsis thaliana Written by: Edit Ábrahám Temesváriné Supervisors: Dr. László Szabados
The sucrose non-fermenting-1-related protein kinases SAPK1 and SAPK2 function collaboratively as positive regulators of salt stress tolerance in rice
 Lou et al. BMC Plant Biology (2018) 18:203 https://doi.org/10.1186/s12870-018-1408-0 RESEARCH ARTICLE Open Access The sucrose non-fermenting-1-related protein kinases SAPK1 and SAPK2 function collaboratively
Lou et al. BMC Plant Biology (2018) 18:203 https://doi.org/10.1186/s12870-018-1408-0 RESEARCH ARTICLE Open Access The sucrose non-fermenting-1-related protein kinases SAPK1 and SAPK2 function collaboratively
9/2/17. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Supplemental Information. Inferring Cell-State Transition. Dynamics from Lineage Trees. and Endpoint Single-Cell Measurements
 Cell Systems, Volume 3 Supplemental Information Inferring Cell-State Transition Dynamics from Lineage Trees and Endpoint Single-Cell Measurements Sahand Hormoz, Zakary S. Singer, James M. Linton, Yaron
Cell Systems, Volume 3 Supplemental Information Inferring Cell-State Transition Dynamics from Lineage Trees and Endpoint Single-Cell Measurements Sahand Hormoz, Zakary S. Singer, James M. Linton, Yaron
Protein Sorting. By: Jarod, Tyler, and Tu
 Protein Sorting By: Jarod, Tyler, and Tu Definition Organizing of proteins Organelles Nucleus Ribosomes Endoplasmic Reticulum Golgi Apparatus/Vesicles How do they know where to go? Amino Acid Sequence
Protein Sorting By: Jarod, Tyler, and Tu Definition Organizing of proteins Organelles Nucleus Ribosomes Endoplasmic Reticulum Golgi Apparatus/Vesicles How do they know where to go? Amino Acid Sequence
Maria V. Yamburenko, Yan O. Zubo, Radomíra Vanková, Victor V. Kusnetsov, Olga N. Kulaeva, Thomas Börner
 ABA represses the transcription of chloroplast genes Maria V. Yamburenko, Yan O. Zubo, Radomíra Vanková, Victor V. Kusnetsov, Olga N. Kulaeva, Thomas Börner Supplementary data Supplementary tables Table
ABA represses the transcription of chloroplast genes Maria V. Yamburenko, Yan O. Zubo, Radomíra Vanková, Victor V. Kusnetsov, Olga N. Kulaeva, Thomas Börner Supplementary data Supplementary tables Table
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Protein Sorting, Intracellular Trafficking, and Vesicular Transport
 Protein Sorting, Intracellular Trafficking, and Vesicular Transport Noemi Polgar, Ph.D. Department of Anatomy, Biochemistry and Physiology Email: polgar@hawaii.edu Phone: 692-1422 Outline Part 1- Trafficking
Protein Sorting, Intracellular Trafficking, and Vesicular Transport Noemi Polgar, Ph.D. Department of Anatomy, Biochemistry and Physiology Email: polgar@hawaii.edu Phone: 692-1422 Outline Part 1- Trafficking
