SUPPLEMENTARY INFORMATION
|
|
|
- Erik Norman
- 6 years ago
- Views:
Transcription
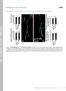
1 SUPPLMTARY IFORMATIO a doi:10.108/nature10402 b 100 nm 100 nm c SAXS Model d ulers assigned to reference- Back-projected free class averages class averages Refinement against single particles Reconstructed density 1.0 fourier shell correlation Unbound Target-bound fsc = fsc= Figure10.0 Supplementary legends resolution (Angstroms) 6.66 Supplementary Figure 1 ryo electron microscopy and three dimensional reconstruction. Images of the unbound (a) and RA bound ascade complex (b) collected at 100,000X magnification. c, Initial models for three dimensional reconstruction were determined using a low resolution SAXS reconstruction14. The SAXS reconstruction was low pass filtered to 60Å resolution, forward projected at an angular increment of 1 degrees, and a multi reference alignment was performed using,000 reference free class averages of each of the ascade complexes. The WWW. AT U R. O M / AT U R aligned class averages were back projected to generate a new density, which was 1
2 RSARH SUPPLMTARY IFORMATIO using,000 reference free class averages of each of the ascade complexes. The aligned class averages were back projected to generate a new density, which was then used for another iteration of projection matching. The unbound and targetbound datasets were processed separately, each using their corresponding initial model. Three dimensional refinements of the starting densities were performed using an iterative projection matching and Fourier reconstruction approach using libraries from the MA2 and SPARX software packages 24,2. d, The resolution of each structure was estimated using Fourier shell correlations 2. Resolutions reported in the paper were made using a conservative 0. Fourier shell correlation criterion. 2
3 Supplementary Figure legends SUPPLMTARY IFORMATIO RSARH microscopy and b a Supplementary Figure 1 ryo electron c three dimensional crra crra reconstruction. Images of the unbound (a) and RA bound ascade complex (b) r s eq u e n c, Initial models for three dimensional collected at 100,000X magnification. e c stem-loop reconstruction were determined using a low resolution SAXS reconstruction14. The ace sp A SAXS reconstruction was low pass filtered to 60Å resolution, forward projected at an angular increment of 1 degrees, and a multi reference alignment was performed Seed cr R using,000 reference free class averages of each of the ascade complexes. The -handle aligned class averages were back projected to generate a new density, which was d then used for another iteration of projection matching. The unbound and target e as bound datasets were processed separately, each using their corresponding initial sa2 model. Three dimensional refinements of the starting densities were performed using an iterative projection matching and Fourier reconstruction approach using 24,2 libraries from the MA2 and SPARX software 90 packages 90. d, The resolution of 90 each structure was estimated using Fourier shell correlations2. Resolutions reported in the paper were made using a conservative 0. Fourier shell correlation criterion. Supplementary Figure 2 Three dimensional docking of crystal structures into ascade. a, Atomic coordinates for homologous structures of as, the crra stem loop and asb from T. thermophilus HB8 were docked in the ~8 Å cryo M map. b, rystal structure of a RISPR specific endoribonuclease (magenta) and the crra stem loop (red) docked in the segmented density for as (gray mesh) and the crra (green surface) (PDB: 2Y8W) 11. c, crra sequence (61nt) modeled into the crra density. d, rystal structure of a asb homolog (se2) form T. thermophilus (yellow) docked in the segmented density for asb (gray mesh) (PDB: 2ZA). Surface charge representation of the. coli asb sequence threaded onto the se2 crystal structure. e, rystal structure of a distant as homolog from the P2 strain of Sulfolobus solfataricus (PDB: PS0)4. The sa2 structure is significantly different form the as density and the two structures do not superimpose with high fidelity. However, shared secondary structure elements are detectable (blue, yellow, magenta and orange) and slight modifications to the pitch or angle of these elements reveal a similar architecture. W W W. A T U R. O M / A T U R
4 RSARH SUPPLMTARY IFORMATIO a b ascade ascade spacer as asa asb as asd as as1 as2 se1 se2 se4 ase se crra ascade ascade ascade as c DA target RA target Spacer Spacer Self PAM min A Self PAM min D B A D B 6 Supplementary Figure Structural interrogation of ascade by limited proteolysis. a, Schematic of the RISPR/as locus in the. coli K12 genome. ascade assembles from an unequal number of as proteins and a crra (asa 1B 2 6D 1 1crRA 1). b c) Limited trypsin digestion of ascade bound to a nucleic acid substrate complementary to the spacer, the spacer plus the handle (self) or to an oligo that contains a protospacer adjacent motif (PAM). A proteolytic product appears at the first time point (green arrow) in all samples. The asb subunits are sensitive to proteolysis 20 minutes after binding to DA or RA targets (red arrow). 4
5 SUPPLMTARY IFORMATIO RSARH Supplementary Figure 4 The as subunits are superimposable between the target bound and unbound structures. The as hexamer from the target bound (surface) and unbound structures (mesh) of ascade. The red arrow indicates additional density for 6 in the unbound structure.
6 RSARH SUPPLMTARY IFORMATIO 8-nt -Handle 2-nt Spacer 21-nt -Stem asa as6 asd A U A A A A U A U U U U A A U U U U A A A 1 Seed T A T T T T A T A A as as4 T A T A A A A T T A as as as2 as as1 A A A T T A A A A T T T T A A A A A T T T A U U as ascade ascade ascade ascade ascade target -8 to 8 1 to 16 9 to to 2 2 to 40 Supplementary Figure Molecular recognition of foreign nucleic acids by ascade. Top, Schematic of the ascade architecture highlighting the relative position of each as protein and the three distinct segments of the mature crra (8 nt handle / 2 nt spacer / 21 nt stem loop). The spacer sequence is shown in red and the protein subunits are colored according to Fig 1. Bottom, Target binding affinities at discrete regions along the crra were determined by electrophoretic mobility shift assays with increasing concentrations of ascade ( nm) in each panel. Sixteen nucleotide ssda oligos that tile along the length of the crra were end labeled with 2P. The last lane in each panel includes 00 nm ascade and a ssda oligo that is not complementary to the crra (). ascade makes low affinity, non sequence specific interactions with DA 14. Supplemental references 2 van Heel, M. & Schatz, M. Fourier shell correlation threshold criteria. J Struct Biol 11, (200). Agari, Y., Yokoyama, S., Kuramitsu, S. & Shinkai, A. X ray crystal structure of a RISPR associated protein, se2, from Thermus thermophilus HB8. Proteins Structure Function and Bioinformatics, (2008). 4 Lintner,.. et al. Structural and functional characterization of an archaeal ASAD complex for RISPR mediated viral defense. J Biol hem (2011). 6
Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples.
 Supplementary Figure 1 Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples. (a-b) SDS-PAGE analysis of the hexamer and heptamer samples. The eluted hexamer
Supplementary Figure 1 Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples. (a-b) SDS-PAGE analysis of the hexamer and heptamer samples. The eluted hexamer
 Supplemental Figure and Movie Legends Figure S1. Time course experiments on the thermal stability of apo AT cpn-α by native PAGE (related to Figure 2B). The samples were heated, respectively, to (A) 45
Supplemental Figure and Movie Legends Figure S1. Time course experiments on the thermal stability of apo AT cpn-α by native PAGE (related to Figure 2B). The samples were heated, respectively, to (A) 45
Cryo-EM data collection, refinement and validation statistics
 1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
Supplementary Figure 1. Aligned sequences of yeast IDH1 (top) and IDH2 (bottom) with isocitrate
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Aligned sequences of yeast IDH1 (top) and IDH2 (bottom) with isocitrate dehydrogenase from Escherichia coli [ICD, pdb 1PB1, Mesecar, A. D., and Koshland,
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1. Aligned sequences of yeast IDH1 (top) and IDH2 (bottom) with isocitrate dehydrogenase from Escherichia coli [ICD, pdb 1PB1, Mesecar, A. D., and Koshland,
SUPPLEMENTARY FIGURES. Structure of the cholera toxin secretion channel in its. closed state
 SUPPLEMENTARY FIGURES Structure of the cholera toxin secretion channel in its closed state Steve L. Reichow 1,3, Konstantin V. Korotkov 1,3, Wim G. J. Hol 1$ and Tamir Gonen 1,2$ 1, Department of Biochemistry
SUPPLEMENTARY FIGURES Structure of the cholera toxin secretion channel in its closed state Steve L. Reichow 1,3, Konstantin V. Korotkov 1,3, Wim G. J. Hol 1$ and Tamir Gonen 1,2$ 1, Department of Biochemistry
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Three-dimensional structure of a viral genome-delivery portal vertex
 Three-dimensional structure of a viral genome-delivery portal vertex Adam S. Olia 1, Peter E. Prevelige Jr. 2, John E. Johnson 3 and Gino Cingolani 4 1 Department of Biological Sciences, Purdue University,
Three-dimensional structure of a viral genome-delivery portal vertex Adam S. Olia 1, Peter E. Prevelige Jr. 2, John E. Johnson 3 and Gino Cingolani 4 1 Department of Biological Sciences, Purdue University,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
ParM filament images were extracted and from the electron micrographs and
 Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11991 Supplementary Figure 1 - Refinement strategy for PIC intermediate assemblies by negative stain EM. The cryo-negative stain structure of free Pol II 1 (a) was used as initial reference
doi:10.1038/nature11991 Supplementary Figure 1 - Refinement strategy for PIC intermediate assemblies by negative stain EM. The cryo-negative stain structure of free Pol II 1 (a) was used as initial reference
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R)
 Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
of the Guanine Nucleotide Exchange Factor FARP2
 Structure, Volume 21 Supplemental Information Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2 Xiaojing He, Yi-Chun Kuo, Tyler J. Rosche, and Xuewu Zhang Inventory of
Structure, Volume 21 Supplemental Information Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2 Xiaojing He, Yi-Chun Kuo, Tyler J. Rosche, and Xuewu Zhang Inventory of
SUPPLEMENTARY INFORMATION
 Dph2 SeMet (iron-free) # Dph2 (iron-free) Dph2-[4Fe-4S] Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 58.26, 82.08, 160.42 58.74, 81.87, 160.01 55.70, 80.53,
Dph2 SeMet (iron-free) # Dph2 (iron-free) Dph2-[4Fe-4S] Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 58.26, 82.08, 160.42 58.74, 81.87, 160.01 55.70, 80.53,
Structure and RNA-binding properties. of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex
 Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
SUPPLEMENTARY INFORMATION
 www.nature.com/nature 1 Figure S1 Sequence alignment. a Structure based alignment of the plgic of E. chrysanthemi (ELIC), the acetylcholine binding protein from the snail Lymnea stagnalis (AchBP, PDB code
www.nature.com/nature 1 Figure S1 Sequence alignment. a Structure based alignment of the plgic of E. chrysanthemi (ELIC), the acetylcholine binding protein from the snail Lymnea stagnalis (AchBP, PDB code
Table 1. Crystallographic data collection, phasing and refinement statistics. Native Hg soaked Mn soaked 1 Mn soaked 2
 Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/6/e1700147/dc1 Supplementary Materials for Ribosome rearrangements at the onset of translational bypassing Xabier Agirrezabala, Ekaterina Samatova, Mariia Klimova,
advances.sciencemag.org/cgi/content/full/3/6/e1700147/dc1 Supplementary Materials for Ribosome rearrangements at the onset of translational bypassing Xabier Agirrezabala, Ekaterina Samatova, Mariia Klimova,
Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome
 Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome Ahmad Jomaa 1, Yu-Hsien Hwang Fu 2, Daniel Boehringer 1, Marc Leibundgut 1, Shu-ou Shan 2, and Nenad
Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome Ahmad Jomaa 1, Yu-Hsien Hwang Fu 2, Daniel Boehringer 1, Marc Leibundgut 1, Shu-ou Shan 2, and Nenad
SUPPLEMENTARY INFORMATION
 Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
SUPPLEMENTARY INFORMATION
 Supplementary Results DNA binding property of the SRA domain was examined by an electrophoresis mobility shift assay (EMSA) using synthesized 12-bp oligonucleotide duplexes containing unmodified, hemi-methylated,
Supplementary Results DNA binding property of the SRA domain was examined by an electrophoresis mobility shift assay (EMSA) using synthesized 12-bp oligonucleotide duplexes containing unmodified, hemi-methylated,
Cks1 CDK1 CDK1 CDK1 CKS1. are ice- lobe. conserved. conserved
 Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Supplementary Figure 3 a. Structural comparison between the two determined structures for the IL 23:MA12 complex. The overall RMSD between the two
 Supplementary Figure 1. Biopanningg and clone enrichment of Alphabody binders against human IL 23. Positive clones in i phage ELISA with optical density (OD) 3 times higher than background are shown for
Supplementary Figure 1. Biopanningg and clone enrichment of Alphabody binders against human IL 23. Positive clones in i phage ELISA with optical density (OD) 3 times higher than background are shown for
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Supplementary figure 1. Comparison of unbound ogm-csf and ogm-csf as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine
 Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Supplemental Materials and Methods
 Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Diphthamide biosynthesis requires a radical iron-sulfur enzyme. Pennsylvania State University, University Park, Pennsylvania 16802, USA
 Diphthamide biosynthesis requires a radical iron-sulfur enzyme Yang Zhang, 1,4 Xuling Zhu, 1,4 Andrew T. Torelli, 1 Michael Lee, 2 Boris Dzikovski, 1 Rachel Koralewski, 1 Eileen Wang, 1 Jack Freed, 1 Carsten
Diphthamide biosynthesis requires a radical iron-sulfur enzyme Yang Zhang, 1,4 Xuling Zhu, 1,4 Andrew T. Torelli, 1 Michael Lee, 2 Boris Dzikovski, 1 Rachel Koralewski, 1 Eileen Wang, 1 Jack Freed, 1 Carsten
Time-dependence of key H-bond/electrostatic interaction distances in the sirna5-hago2 complexes... Page S14
 Supporting Information Probing the Binding Interactions between Chemically Modified sirnas and Human Argonaute 2 Using Microsecond Molecular Dynamics Simulations S. Harikrishna* and P. I. Pradeepkumar*
Supporting Information Probing the Binding Interactions between Chemically Modified sirnas and Human Argonaute 2 Using Microsecond Molecular Dynamics Simulations S. Harikrishna* and P. I. Pradeepkumar*
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Cryo-EM structure and model of the C. thermophilum 90S preribosome. a, Gold standard FSC curve showing the average resolution of the 90S preribosome masked and unmasked (left). FSC
Supplementary Figure 1 Cryo-EM structure and model of the C. thermophilum 90S preribosome. a, Gold standard FSC curve showing the average resolution of the 90S preribosome masked and unmasked (left). FSC
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Definition and assessment of ciap1 constructs.
 Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/310/5753/1513/dc1 Supporting Online Material for Structural Roles for Human Translation Factor eif3 in Initiation of Protein Synthesis Bunpote Siridechadilok, Christopher
www.sciencemag.org/cgi/content/full/310/5753/1513/dc1 Supporting Online Material for Structural Roles for Human Translation Factor eif3 in Initiation of Protein Synthesis Bunpote Siridechadilok, Christopher
Supplemental Data SUPPLEMENTAL FIGURES
 Supplemental Data CRYSTAL STRUCTURE OF THE MG.ADP-INHIBITED STATE OF THE YEAST F 1 C 10 ATP SYNTHASE Alain Dautant*, Jean Velours and Marie-France Giraud* From Université Bordeaux 2, CNRS; Institut de
Supplemental Data CRYSTAL STRUCTURE OF THE MG.ADP-INHIBITED STATE OF THE YEAST F 1 C 10 ATP SYNTHASE Alain Dautant*, Jean Velours and Marie-France Giraud* From Université Bordeaux 2, CNRS; Institut de
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Measuring quaternary structure similarity using global versus local measures.
 Supplementary Figure 1 Measuring quaternary structure similarity using global versus local measures. (a) Structural similarity of two protein complexes can be inferred from a global superposition, which
Supplementary Figure 1 Measuring quaternary structure similarity using global versus local measures. (a) Structural similarity of two protein complexes can be inferred from a global superposition, which
SUPPLEMENTARY INFORMATION
 Data collection Supplementary Table 1 Statistics of data collection, phasing and refinement Native Se-MAD Space group P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 50.4, 94.2, 115.4 49.8, 94.2,
Data collection Supplementary Table 1 Statistics of data collection, phasing and refinement Native Se-MAD Space group P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 50.4, 94.2, 115.4 49.8, 94.2,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12045 Supplementary Table 1 Data collection and refinement statistics. Native Pt-SAD X-ray source SSRF BL17U SPring-8 BL41XU Wavelength (Å) 0.97947 1.07171 Space group P2 1 2 1 2 1 P2
doi:10.1038/nature12045 Supplementary Table 1 Data collection and refinement statistics. Native Pt-SAD X-ray source SSRF BL17U SPring-8 BL41XU Wavelength (Å) 0.97947 1.07171 Space group P2 1 2 1 2 1 P2
Introduction to Comparative Protein Modeling. Chapter 4 Part I
 Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Detection of Protein Binding Sites II
 Detection of Protein Binding Sites II Goal: Given a protein structure, predict where a ligand might bind Thomas Funkhouser Princeton University CS597A, Fall 2007 1hld Geometric, chemical, evolutionary
Detection of Protein Binding Sites II Goal: Given a protein structure, predict where a ligand might bind Thomas Funkhouser Princeton University CS597A, Fall 2007 1hld Geometric, chemical, evolutionary
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
SUPPLEMENTARY INFORMATION. doi: /nature07461
 Figure S1 Electrophysiology. a ph-activation of. Two-electrode voltage clamp recordings of Xenopus oocytes expressing in comparison to waterinjected oocytes. Currents were recorded at 40 mv. The ph of
Figure S1 Electrophysiology. a ph-activation of. Two-electrode voltage clamp recordings of Xenopus oocytes expressing in comparison to waterinjected oocytes. Currents were recorded at 40 mv. The ph of
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.1290 Metal-directed, chemically tunable assembly of one-, two- and threedimensional crystalline protein arrays 1 2 1 1 1,3 Jeffrey D. Brodin 1, X. I. Ambroggio 2, Chunyan Tang 1, Kristin
DOI: 10.1038/NCHEM.1290 Metal-directed, chemically tunable assembly of one-, two- and threedimensional crystalline protein arrays 1 2 1 1 1,3 Jeffrey D. Brodin 1, X. I. Ambroggio 2, Chunyan Tang 1, Kristin
Structural insights into bacterial flagellar hooks similarities and specificities
 Supplementary information Structural insights into bacterial flagellar hooks similarities and specificities Young-Ho Yoon, Clive S. Barker, Paula V. Bulieris, Hideyuki Matsunami, Fadel A. Samatey* Affiliation:
Supplementary information Structural insights into bacterial flagellar hooks similarities and specificities Young-Ho Yoon, Clive S. Barker, Paula V. Bulieris, Hideyuki Matsunami, Fadel A. Samatey* Affiliation:
SUPPLEMENTARY FIGURES. Figure S1
 SUPPLEMENTARY FIGURES Figure S1 The substrate for DH domain (2R,3R,4R,6R,7S,8S,9R)-3,7,9-trihydroxy-5-oxo-2,4,6,8 tetramethylundecanoate) was docked as two separate fragments shown in magenta and blue
SUPPLEMENTARY FIGURES Figure S1 The substrate for DH domain (2R,3R,4R,6R,7S,8S,9R)-3,7,9-trihydroxy-5-oxo-2,4,6,8 tetramethylundecanoate) was docked as two separate fragments shown in magenta and blue
SUPPLEMENTARY INFORMATION
 Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Supplementary Figure 1
 Supplementary Figure 1 The correlation of n-score cutoff and FDR in both CID-only and CID-ETD fragmentation strategies. A bar diagram of different n-score thresholds applied in the search, plotted against
Supplementary Figure 1 The correlation of n-score cutoff and FDR in both CID-only and CID-ETD fragmentation strategies. A bar diagram of different n-score thresholds applied in the search, plotted against
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Chemical structure of LPS and LPS biogenesis in Gram-negative bacteria. a. Chemical structure of LPS. LPS molecule consists of Lipid A, core oligosaccharide and O-antigen. The polar
Supplementary Figure 1 Chemical structure of LPS and LPS biogenesis in Gram-negative bacteria. a. Chemical structure of LPS. LPS molecule consists of Lipid A, core oligosaccharide and O-antigen. The polar
SI Text S1 Solution Scattering Data Collection and Analysis. SI references
 SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
Acta Crystallographica Section D
 Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: Structural characterization of the virulence factor Nuclease A from Streptococcus agalactiae
Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: Structural characterization of the virulence factor Nuclease A from Streptococcus agalactiae
Supplemental Information. Molecular Basis of Spectral Diversity. in Near-Infrared Phytochrome-Based. Fluorescent Proteins
 Chemistry & Biology, Volume 22 Supplemental Information Molecular Basis of Spectral Diversity in Near-Infrared Phytochrome-Based Fluorescent Proteins Daria M. Shcherbakova, Mikhail Baloban, Sergei Pletnev,
Chemistry & Biology, Volume 22 Supplemental Information Molecular Basis of Spectral Diversity in Near-Infrared Phytochrome-Based Fluorescent Proteins Daria M. Shcherbakova, Mikhail Baloban, Sergei Pletnev,
Protein Structure Prediction, Engineering & Design CHEM 430
 Protein Structure Prediction, Engineering & Design CHEM 430 Eero Saarinen The free energy surface of a protein Protein Structure Prediction & Design Full Protein Structure from Sequence - High Alignment
Protein Structure Prediction, Engineering & Design CHEM 430 Eero Saarinen The free energy surface of a protein Protein Structure Prediction & Design Full Protein Structure from Sequence - High Alignment
Spliceosome and Localization of Its Catalytic Core
 Molecular Cell, Volume 40 Supplemental Information 3D Cryo-EM Structure of an Active Step I Spliceosome and Localization of Its Catalytic Core Monika M. Golas, Bjoern Sander, Sergey Bessonov, Michael Grote,
Molecular Cell, Volume 40 Supplemental Information 3D Cryo-EM Structure of an Active Step I Spliceosome and Localization of Its Catalytic Core Monika M. Golas, Bjoern Sander, Sergey Bessonov, Michael Grote,
Department of Biochemistry, University of Zürich, Winterthurerstrasse 190, CH-8057 Zürich, Switzerland
 Supporting information Twenty crystal structures of bromodomain and PHD finger containing protein 1 (BRPF1)/ligand complexes reveal conserved binding motifs and rare interactions Jian Zhu and Amedeo Caflisch*
Supporting information Twenty crystal structures of bromodomain and PHD finger containing protein 1 (BRPF1)/ligand complexes reveal conserved binding motifs and rare interactions Jian Zhu and Amedeo Caflisch*
Homology Modeling. Roberto Lins EPFL - summer semester 2005
 Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Nature Methods: doi: /nmeth Supplementary Figure 1. Overview of the density-guided rebuilding and refinement protocol.
 Supplementary Figure 1 Overview of the density-guided rebuilding and refinement protocol. (a) Flowchart of the protocol. The protocol uses 250 cycles of local backbone rebuilding, followed by five cycles
Supplementary Figure 1 Overview of the density-guided rebuilding and refinement protocol. (a) Flowchart of the protocol. The protocol uses 250 cycles of local backbone rebuilding, followed by five cycles
Three-Dimensional Electron Microscopy of Macromolecular Assemblies
 Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Structure Comparison
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits. AhpC and AhpF from Escherichia coli
 Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits AhpC and AhpF from Escherichia coli Phat Vinh Dip 1,#, Neelagandan Kamariah 2,#, Malathy Sony Subramanian Manimekalai
Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits AhpC and AhpF from Escherichia coli Phat Vinh Dip 1,#, Neelagandan Kamariah 2,#, Malathy Sony Subramanian Manimekalai
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Data collection, phasing and refinement statistics ChbC/Ta 6 Br 12 Native ChbC Data collection Space group P4 3 2 1 2 P4 3 2 1 2 Cell dimensions a, c (Å) 132.75, 453.57 132.81, 452.95
Supplementary Table 1: Data collection, phasing and refinement statistics ChbC/Ta 6 Br 12 Native ChbC Data collection Space group P4 3 2 1 2 P4 3 2 1 2 Cell dimensions a, c (Å) 132.75, 453.57 132.81, 452.95
Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
Supplementary Figure 1 Crystal packing of ClR and electron density maps. Crystal packing of type A crystal (a) and type B crystal (b).
 Supplementary Figure 1 Crystal packing of ClR and electron density maps. Crystal packing of type A crystal (a) and type B crystal (b). Crystal contacts at B-C loop are magnified and stereo view of A-weighted
Supplementary Figure 1 Crystal packing of ClR and electron density maps. Crystal packing of type A crystal (a) and type B crystal (b). Crystal contacts at B-C loop are magnified and stereo view of A-weighted
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
Structural characterization of NiV N 0 P in solution and in crystal.
 Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Table S1. Primers used for the constructions of recombinant GAL1 and λ5 mutants. GAL1-E74A ccgagcagcgggcggctgtctttcc ggaaagacagccgcccgctgctcgg
 SUPPLEMENTAL DATA Table S1. Primers used for the constructions of recombinant GAL1 and λ5 mutants Sense primer (5 to 3 ) Anti-sense primer (5 to 3 ) GAL1 mutants GAL1-E74A ccgagcagcgggcggctgtctttcc ggaaagacagccgcccgctgctcgg
SUPPLEMENTAL DATA Table S1. Primers used for the constructions of recombinant GAL1 and λ5 mutants Sense primer (5 to 3 ) Anti-sense primer (5 to 3 ) GAL1 mutants GAL1-E74A ccgagcagcgggcggctgtctttcc ggaaagacagccgcccgctgctcgg
Probabilistic Arithmetic Automata
 Probabilistic Arithmetic Automata Applications of a Stochastic Computational Framework in Biological Sequence Analysis Inke Herms PhD thesis defense Overview 1 Probabilistic Arithmetic Automata 2 Application
Probabilistic Arithmetic Automata Applications of a Stochastic Computational Framework in Biological Sequence Analysis Inke Herms PhD thesis defense Overview 1 Probabilistic Arithmetic Automata 2 Application
Transmembrane Domains (TMDs) of ABC transporters
 Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Pymol Practial Guide
 Pymol Practial Guide Pymol is a powerful visualizor very convenient to work with protein molecules. Its interface may seem complex at first, but you will see that with a little practice is simple and powerful
Pymol Practial Guide Pymol is a powerful visualizor very convenient to work with protein molecules. Its interface may seem complex at first, but you will see that with a little practice is simple and powerful
Image Assessment San Diego, November 2005
 Image Assessment San Diego, November 005 Pawel A. Penczek The University of Texas Houston Medical School Department of Biochemistry and Molecular Biology 6431 Fannin, MSB6.18, Houston, TX 77030, USA phone:
Image Assessment San Diego, November 005 Pawel A. Penczek The University of Texas Houston Medical School Department of Biochemistry and Molecular Biology 6431 Fannin, MSB6.18, Houston, TX 77030, USA phone:
Supplementary materials. Crystal structure of the carboxyltransferase domain. of acetyl coenzyme A carboxylase. Department of Biological Sciences
 Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Kasprowicz et al., http://www.jcb.org/cgi/content/full/jcb.201310090/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. NMJ morphology of shi 12-12B ; shi-4c not treated
Supplemental material Kasprowicz et al., http://www.jcb.org/cgi/content/full/jcb.201310090/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. NMJ morphology of shi 12-12B ; shi-4c not treated
SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition SUPPLEMENTARY INFORMATION
 SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition Nadia J. Kershaw 1,2, James M. Murphy 1,2, Nicholas P.D. Liau 1,2, Leila N. Varghese 1,2, Artem Laktyushin
SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition Nadia J. Kershaw 1,2, James M. Murphy 1,2, Nicholas P.D. Liau 1,2, Leila N. Varghese 1,2, Artem Laktyushin
Expanded View Figures
 The EMBO Journal Structure of a Dm peptide bound to the OT module Tobias Raisch et al Expanded View Figures A Hs Dm 262 297 685 8 HEAT HEAT MIF4G 9BD 1SHD 761 91 193 169 1152 1317 16 1376 1467 HEAT HEAT
The EMBO Journal Structure of a Dm peptide bound to the OT module Tobias Raisch et al Expanded View Figures A Hs Dm 262 297 685 8 HEAT HEAT MIF4G 9BD 1SHD 761 91 193 169 1152 1317 16 1376 1467 HEAT HEAT
Nature Structural and Molecular Biology: doi: /nsmb.2938
 Supplementary Figure 1 Characterization of designed leucine-rich-repeat proteins. (a) Water-mediate hydrogen-bond network is frequently visible in the convex region of LRR crystal structures. Examples
Supplementary Figure 1 Characterization of designed leucine-rich-repeat proteins. (a) Water-mediate hydrogen-bond network is frequently visible in the convex region of LRR crystal structures. Examples
Structure of the SPRY domain of human DDX1 helicase, a putative interaction platform within a DEAD-box protein
 Supporting information Volume 71 (2015) Supporting information for article: Structure of the SPRY domain of human DDX1 helicase, a putative interaction platform within a DEAD-box protein Julian Kellner
Supporting information Volume 71 (2015) Supporting information for article: Structure of the SPRY domain of human DDX1 helicase, a putative interaction platform within a DEAD-box protein Julian Kellner
The Fic protein Doc uses an inverted substrate to phosphorylate and. inactivate EF-Tu
 The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
Supplementary information
 Supplementary information The structural basis of modularity in ECF-type ABC transporters Guus B. Erkens 1,2, Ronnie P-A. Berntsson 1,2, Faizah Fulyani 1,2, Maria Majsnerowska 1,2, Andreja Vujičić-Žagar
Supplementary information The structural basis of modularity in ECF-type ABC transporters Guus B. Erkens 1,2, Ronnie P-A. Berntsson 1,2, Faizah Fulyani 1,2, Maria Majsnerowska 1,2, Andreja Vujičić-Žagar
Bioinformatics. Macromolecular structure
 Bioinformatics Macromolecular structure Contents Determination of protein structure Structure databases Secondary structure elements (SSE) Tertiary structure Structure analysis Structure alignment Domain
Bioinformatics Macromolecular structure Contents Determination of protein structure Structure databases Secondary structure elements (SSE) Tertiary structure Structure analysis Structure alignment Domain
Protein synthesis I Biochemistry 302. Bob Kelm February 23, 2004
 Protein synthesis I Biochemistry 302 Bob Kelm February 23, 2004 Key features of protein synthesis Energy glutton Essential metabolic activity of the cell. Consumes 90% of the chemical energy (ATP,GTP).
Protein synthesis I Biochemistry 302 Bob Kelm February 23, 2004 Key features of protein synthesis Energy glutton Essential metabolic activity of the cell. Consumes 90% of the chemical energy (ATP,GTP).
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Tertiary Structure Prediction
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
CMPS 3110: Bioinformatics. Tertiary Structure Prediction
 CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
Visualization of Macromolecular Structures
 Visualization of Macromolecular Structures Present by: Qihang Li orig. author: O Donoghue, et al. Structural biology is rapidly accumulating a wealth of detailed information. Over 60,000 high-resolution
Visualization of Macromolecular Structures Present by: Qihang Li orig. author: O Donoghue, et al. Structural biology is rapidly accumulating a wealth of detailed information. Over 60,000 high-resolution
Three types of RNA polymerase in eukaryotic nuclei
 Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
RNA Polymerase I Contains a TFIIF-Related DNA-Binding Subcomplex
 Molecular Cell, Volume 39 Supplemental Information RNA Polymerase I Contains a TFIIFRelated DNABinding Subcomplex Sebastian R. Geiger, Kristina Lorenzen, Amelie Schreieck, Patrizia Hanecker, Dirk Kostrewa,
Molecular Cell, Volume 39 Supplemental Information RNA Polymerase I Contains a TFIIFRelated DNABinding Subcomplex Sebastian R. Geiger, Kristina Lorenzen, Amelie Schreieck, Patrizia Hanecker, Dirk Kostrewa,
Supporting Information
 Supporting Information Ottmann et al. 10.1073/pnas.0907587106 Fig. S1. Primary structure alignment of SBT3 with C5 peptidase from Streptococcus pyogenes. The Matchmaker tool in UCSF Chimera (http:// www.cgl.ucsf.edu/chimera)
Supporting Information Ottmann et al. 10.1073/pnas.0907587106 Fig. S1. Primary structure alignment of SBT3 with C5 peptidase from Streptococcus pyogenes. The Matchmaker tool in UCSF Chimera (http:// www.cgl.ucsf.edu/chimera)
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Structural Bioinformatics (C3210) Molecular Docking
 Structural Bioinformatics (C3210) Molecular Docking Molecular Recognition, Molecular Docking Molecular recognition is the ability of biomolecules to recognize other biomolecules and selectively interact
Structural Bioinformatics (C3210) Molecular Docking Molecular Recognition, Molecular Docking Molecular recognition is the ability of biomolecules to recognize other biomolecules and selectively interact
Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions
 Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions undergo endocytosis into clathrin-coated vesicles dependent
Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions undergo endocytosis into clathrin-coated vesicles dependent
Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08 Å resolution: comparison with the Azotobacter vinelandii MoFe protein
 Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Protein Structure Prediction
 Page 1 Protein Structure Prediction Russ B. Altman BMI 214 CS 274 Protein Folding is different from structure prediction --Folding is concerned with the process of taking the 3D shape, usually based on
Page 1 Protein Structure Prediction Russ B. Altman BMI 214 CS 274 Protein Folding is different from structure prediction --Folding is concerned with the process of taking the 3D shape, usually based on
