Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
|
|
|
- Candice Sullivan
- 6 years ago
- Views:
Transcription
1 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the wild type, where differentiating neurons reside, is free of ph3 labeling. In the mutant, ph3 labeling is detectable in all regions and the ratio of ph3-positive nuclei is increased (see Fig. 1G). (C,D) BrdU labeling for 2 hours in wild-type (C) and mutant (D) retinae at 2.8 dpf. In the wild type, a small region of the central retina, where presumptive ganglion cells are differentiating, is free of BrdU labeling (arrowhead), whereas the mutant retina is ubiquitously labeled. (E,F) BrdU incorporation for 12 hours starting at 4 dpf. In the wild type (E), BrdU incorporation is restricted to the peripheral retina where proliferating RPCs reside, whereas in the mutant BrdU incorporation is detectable in all regions of the retina. (G,H) BrdU (green) and ph3 (red) double labeling at stage 27 to examine S- to M-phase cell cycle progression. BrdU was injected at stage 27 and after 9 hours embryos were fixed and stained for ph3 and BrdU. Examples of double-positive nuclei (yellow) are indicated (arrowheads). (I) The ratio of ph3 and BrdU double-positive to total ph3-positive nuclei was analyzed to determine the relative number of cells that progress from S to M phase during the 9 hours of BrdU incorporation. This ratio is similar in both wild type and mutant, indicating that cell cycle progression proceeds normally in mutant RPCs. (J,K) The expression of ath5 was used to visualize RPCs undergoing the final cell division prior to neuronal differentiation (Poggi et al., 2005; Yamaguchi et al., 2010) by crossing a stable transgenic line expressing GFP under the control of the ath5 regulatory region [ath5::gfp (del Bene et al., 2007)] into the med mutant. In this context, cells undergoing a neurogenic division are positive for both ph3 (red, M phase) and GFP (green, ath5). In both mutant and wild type, GFP expression was first detectable in central RPCs at stage 26 and subsequently spread across the retina to more peripheral regions (del Bene et al., 2007) (data not shown). This indicates that neurogenesis is normally initiated in the mutant retina. At stage 27, we quantified the ratio of ph3/gfp double-positive nuclei (arrowhead) to total ph3 nuclei to analyse cell cycle exit. (L) Quantification the ratio of ph3/gfp double-positive to total ph3-positive cells to analyze the relative number of cells exiting the cell cycle by neurogenic divisions. In the wild type, 33% of the ph3-positive cells are also GFP positive, in the mutant this ratio is significantly reduced to 6.8%, indicating an almost fivefold reduction of cell cycle exit. (A-H,J,K) Confocal transversal sections, nuclei are stained with DAPI (blue). Scale bars: 25 µm in A-H,J,K.
2 Fig. S2. Vertebrate ArhGEF18 homologues are conserved. Alignments of medaka (Oryzias latipes, OlArhGEF18: GenBank JN868075), zebrafish (Danio rerio, DrArhGEF18: Uniprot Q6NSP2), mouse (Mus musculus, MmsArhGEF18: GenBank EDL21902), macaca (Macaca mulatta, MmaArhGEF18: NCBI XP_ ) and human (Homo sapiens, HsArhGEF18: NCBI NP_056133) proteins. The conserved DH (blue) and PH (red) domains are indicated. The DH domain is 60.2% identical to that of the zebrafish homologue and 65.8% to the DH domain of the murine and human homologue. The PH domain is 67.3 identical to that of zebrafish and 71.2% and 69.2% to that of mouse and humans, respectively. The C-terminal part of 604 amino acids is less well conserved, ranging from 39.5% (zebrafish) to 41.2% (human) identity.
3 Fig. S3. OlArhGEF18 morpholino injection phenocopies the med mutation. (A-C) Dorsal views of control-injected embryos (A), morpholino 3+4-injected embryos with weak phenotype (B) and morpholino 3+4-injected embryos with strong phenotype (C) at 4 dpf. (D) RT-PCR analysis of OlArhGEF18 mrna of morpholino and uninjected embryos. Uninjected and control morpholino-injected embryos show the wild-type transcript (lower band: wt); injection of morpholino 3, morpholino 4 and morpholino 3+4 results in splicing inhibition. The amount of unspliced transcript (upper band: mo) correlates with the severity of the phenotype. (E,F) Position of M-phase nuclei as visualized by a-ph3 antibody-labeling in control morpholino- (E) and morpholino 3+4- (F) injected embryos at stage 25 in the retina. Morpholino 3+4 injection results in mislocalized nuclei (F, arrowheads), whereas in the wild type, nuclei are apically localized (E). (G,H) apkc localization in control morpholino- (G) and morpholino 3+4- (H) injected embryos at stage 27. Apical apkc localization in the retina is severely perturbed in morpholino 3+4-injected embryos (arrowheads). (E-H) Nuclei are stained with DAPI (blue). Scale bars: 50 µm in E,F. Morpholino sequences: MO3, TGAATGCAAATGTGGGTGCTTACCA; MO3C, TCAATGGAAATCTGGGTCCTTAGCA; MO4, CAGCTCTGTGAGACGACACCAGTGT; MO4C, CACCTCTCTGACACGAGACCACTGT.
4 Fig. S4. Vertebrate RhoA, Rac1 and Cdc42 GTPases are highly conserved and ubiquitously expressed in medaka embryos at neurula and organogenesis stages. (A) Protein sequence comparison of human RhoA (Hs RhoA), Rac1 (HsRac1), Cdc42 (HsCdc42); zebrafish RhoAa (DrRhoAa), Rac1 (DrRac1), Cdc42 and Cdc42-like (DrCdc42, DrCdc42-like); and medaka RhoAa and RhoAb (OlRhoAa, OlRhoAb), Rac1a and Rac1b (OlRac1a, OlRac1b), and Cdc42 and Cdc42-like (OlCdc42, OlCdc42-like) are shown. Medaka RhoAa and RhoAb are 95% and 96% identical to human RhoA. Medaka Rac1a and Rac1b share 97% and 96% identity with human Rac1. Medaka Cdc42 and Cdc42-like are 96% and 99% identical, respectively, to human Cdc42. Most variable amino acids are located in the C-terminal region of the respective GTPases. All amino acids that have been implicated in GEF binding (Karnoub et al., 2001; Oleksy et al., 2006; Rossman et al., 2002; Worthylake et al., 2000) are completely conserved in these GTPases. OlRhoAa FOE002-P00007-DPE-F_I14 (GenBank: AM298929), OlRhoAb FOE002-P00007-DPE-F_M18 (GenBank: AM299029), OlRac1a FOE002-P00022-DPE-F_N10 (GenBank: AM304480), OlRac1b FOE002-P00013-DPE-F_P20 (GenBank: AM301286), OlCdc42 FOE002-P00024-DPE-F_A03 (GenBank: AM304876) and OlCdc42-like FOE002-P00008-DPE-F_C14 (GenBank: AM299156) were identified by BLAST searches from a medaka unigene cdna library (Souren et al., 2009) using sequences of zebrafish homologues. (B) Whole-mount in situ hybridization expression analysis of RhoAa, RhoAb, Rac1a, Rac1b, Cdc42 and Cdc42-like GTPases in wild-type medaka embryos at neurula (stage 18) and organogenesis (stage 24) stages. With the exception of Cdc42-like, all GTPases are ubiquitously expressed in the developing medaka embryo.
5 Fig. S5. T2A peptide separates GFP and a C-terminal polypeptide into two functional proteins. (A) Scheme of the GFP-T2A expression constructs. N-terminal GFP and the C-terminal dominant-negative protein were linked in-frame by the Thosea asigna virus T2A peptide, where a ribosomal skip mechanism results in the cleavage of GFP and dominant-negative protein, respectively. This allows us to co-express these two proteins from a single vector (Szymczak et al., 2004). The expected sizes of the respective cleavage -products in kda is indicated. The amino acid sequence of the Thosea asigna virus T2A peptide and the cleavage -site is shown. (B-D) Western blot analysis of 24 hpf medaka embryos injected at the one-cell stage with RNA encoding GFP-T2A-dominantnegative-RhoA (GFP-T2A-dnRhoA; B), GFP-T2A- dominant-negative-rac1 (GFP-T2A-dnRac1; C) and GFP-T2A- dominantnegative-rock2 (GFP-T2A-dnRock2; D). Lane 1, lysate of injected embryos probed with the indicated antibodies; lane 2, lysate of uninjected embryos probed with the indicated antibodies; lane 3, lysate of injected embryos probed with a-gfp antibody. The protein sizes are indicated in kda. Expression of GFP-T2A-dnRhoA (B), GFP-T2A-dnRac1 (C) and GFP-T2A-dnRock2 (D) in medaka embryos results in the cleavage of GFP and the respective C terminal protein. (E,F) GTPase activation in HEK293 cells in response to GFP-T2A-dnRhoA and constitutive activated RhoA (E,E ) and GFP-T2A-dnRac1 and constitutive activated Rac1 (F,F ). Lanes are as follows: 1, untransfected; 2-4, GFP-T2A-dominant negative GTPase; 5-6, constitutive activated GTPase. (E) GFP-T2A-dnRhoA is inactive as determined by the absence of binding to GST-RBD (lanes 2-4), whereas activated RhoA strongly binds GST-RBD (lanes 5-7). (E ) Total amount of RhoA in the cell lysate. (F) GFP-T2A-dnRac1 is inactive as determined by the absence of binding to GST-Pak1 (lanes 2-4), whereas activated Rac1 shows strong binding. (F ) Total amount of Rac1 in the cell lysate. (G) HEK298 cells expressing GFP-T2A (negative control), GFP-T2A-dnRhoA and GFP-T2A-dnRock2 were stained for GFP (a-gfp) and actin (phalloidin), respectively. Cells expressing GFP-T2A-dnRhoA and GFP-T2A-dnRock2 delaminate and round up, thus indicating that dnrhoa and dnrock2 efficiently block RhoA and Rock2 activity, respectively (Barberis et al., 2005).
6 Fig. S6. RhoA and Rock2 but not Rac1 are required for cortical actin localization and apical localization of M-phase nuclei. (A-D) Representative examples of confocal transversal sections of embryos at stage 25 expressing GFP (A), dominant-negative RhoA (B), dominant-negative Rock2 (C) or dominant-negative Rac1 (D) under the control of a heat-shock promoter. Expressing cells are labeled with GFP (green). The retinal neuroepithelium is outlined by a broken line in column PH3. In cells expressing dominantnegative RhoA (B) and dominant-negative Rock2 (C) cortical actin localization is disturbed. Apical actin bundles are mislocalized or absent (arrowheads in column actin of B,C). In addition, apical localization of M-phase nuclei is affected by expression of dominant-negative RhoA (B) and dominant negative Rock2 (C). Nuclei of mitotic cells visualized by a-ph3 antibody staining are often displaced basally in expressing cells (arrowheads in column PH3 of B,C). Cortical actin localization and apical localization of M-phase nuclei are not affected by expression of dominant-negative Rac1 (D). Scale bars: 25 µm.
 DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
SUPPLEMENTARY FIGURES AND TABLES AND THEIR LEGENDS. Transient infection of the zebrafish notochord triggers chronic inflammation
 SUPPLEMENTARY FIGURES AND TABLES AND THEIR LEGENDS Transient infection of the zebrafish notochord triggers chronic inflammation Mai Nguyen-Chi 1,2,5, Quang Tien Phan 1,2,5, Catherine Gonzalez 1,2, Jean-François
SUPPLEMENTARY FIGURES AND TABLES AND THEIR LEGENDS Transient infection of the zebrafish notochord triggers chronic inflammation Mai Nguyen-Chi 1,2,5, Quang Tien Phan 1,2,5, Catherine Gonzalez 1,2, Jean-François
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
Supporting Information
 Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supplementary Information
 Supplementary Information MAP2/Hoechst Hyp.-AP ph 6.5 Hyp.-SD ph 7.2 Norm.-SD ph 7.2 Supplementary Figure 1. Mitochondrial elongation in cortical neurons by acidosis. Representative images of neuronal
Supplementary Information MAP2/Hoechst Hyp.-AP ph 6.5 Hyp.-SD ph 7.2 Norm.-SD ph 7.2 Supplementary Figure 1. Mitochondrial elongation in cortical neurons by acidosis. Representative images of neuronal
Translation and Operons
 Translation and Operons You Should Be Able To 1. Describe the three stages translation. including the movement of trna molecules through the ribosome. 2. Compare and contrast the roles of three different
Translation and Operons You Should Be Able To 1. Describe the three stages translation. including the movement of trna molecules through the ribosome. 2. Compare and contrast the roles of three different
SUPPLEMENTARY INFORMATION
 GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and
 Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figures
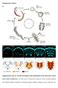 Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Gene Duplication of the Zebrafish kit ligand and Partitioning of Melanocyte Development Functions to kit ligand a
 Gene Duplication of the Zebrafish kit ligand and Partitioning of Melanocyte Development Functions to kit ligand a Keith A. Hultman 1, Nathan Bahary 2, Leonard I. Zon 3,4, Stephen L. Johnson 1* 1 Department
Gene Duplication of the Zebrafish kit ligand and Partitioning of Melanocyte Development Functions to kit ligand a Keith A. Hultman 1, Nathan Bahary 2, Leonard I. Zon 3,4, Stephen L. Johnson 1* 1 Department
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
10/2/2015. Chapter 4. Determination and Differentiation. Neuroanatomical Diversity
 Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes
 9 The Nucleus Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes Explain general structures of Nuclear Envelope, Nuclear Lamina, Nuclear Pore Complex Explain movement of proteins
9 The Nucleus Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes Explain general structures of Nuclear Envelope, Nuclear Lamina, Nuclear Pore Complex Explain movement of proteins
Supporting Information
 Supporting Information Cao et al. 10.1073/pnas.1306220110 Gram - bacteria Hemolymph Cytoplasm PGRP-LC TAK1 signalosome Imd dfadd Dredd Dnr1 Ikk signalosome P Relish Nucleus AMP and effector genes Fig.
Supporting Information Cao et al. 10.1073/pnas.1306220110 Gram - bacteria Hemolymph Cytoplasm PGRP-LC TAK1 signalosome Imd dfadd Dredd Dnr1 Ikk signalosome P Relish Nucleus AMP and effector genes Fig.
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
5- Semaphorin-Plexin-Neuropilin
 5- Semaphorin-Plexin-Neuropilin 1 SEMAPHORINS-PLEXINS-NEUROPILINS ligands receptors co-receptors semaphorins and their receptors are known signals for: -axon guidance -cell migration -morphogenesis -immune
5- Semaphorin-Plexin-Neuropilin 1 SEMAPHORINS-PLEXINS-NEUROPILINS ligands receptors co-receptors semaphorins and their receptors are known signals for: -axon guidance -cell migration -morphogenesis -immune
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
Videos. Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.
 Translation Translation Videos Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.be/itsb2sqr-r0 Translation Translation The
Translation Translation Videos Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.be/itsb2sqr-r0 Translation Translation The
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1 Sns and Duf co-localise in embryonic nephrocytes a-c, Wild-type stage 11 (a),14 (b) and 16 (c) embryos stained with anti-duf (green) and anti-sns (red). Higher magnification images
Supplementary Figure 1 Sns and Duf co-localise in embryonic nephrocytes a-c, Wild-type stage 11 (a),14 (b) and 16 (c) embryos stained with anti-duf (green) and anti-sns (red). Higher magnification images
TNFα 18hr. Control. CHX 18hr. TNFα+ CHX 18hr. TNFα: 18 18hr (KDa) PARP. Cleaved. Cleaved. Cleaved. Caspase3. Pellino3 shrna. Control shrna.
 Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Biological Roles of Cytokinins
 Direct Control of Shoot Meristem Activity by a Cytokinin-Activating Enzyme By Kurakawa et. Al. Published in Nature Presented by Boyana Grigorova Biological Roles of Cytokinins Cytokinins are positive regulators
Direct Control of Shoot Meristem Activity by a Cytokinin-Activating Enzyme By Kurakawa et. Al. Published in Nature Presented by Boyana Grigorova Biological Roles of Cytokinins Cytokinins are positive regulators
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1
 Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
SUPPLEMENTARY INFORMATION
 med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
elipsa is an early determinant of ciliogenesis that links the IFT particle to membrane-associated small GTPase
 letters elipsa is an early determinant of ciliogenesis that links the IFT particle to membrane-associated small GTPase Rab8 Yoshihiro Omori 1, Chengtian Zhao 1, Arunesh Saras 1, Saikat Mukhopadhyay 2,
letters elipsa is an early determinant of ciliogenesis that links the IFT particle to membrane-associated small GTPase Rab8 Yoshihiro Omori 1, Chengtian Zhao 1, Arunesh Saras 1, Saikat Mukhopadhyay 2,
1. In most cases, genes code for and it is that
 Name Chapter 10 Reading Guide From DNA to Protein: Gene Expression Concept 10.1 Genetics Shows That Genes Code for Proteins 1. In most cases, genes code for and it is that determine. 2. Describe what Garrod
Name Chapter 10 Reading Guide From DNA to Protein: Gene Expression Concept 10.1 Genetics Shows That Genes Code for Proteins 1. In most cases, genes code for and it is that determine. 2. Describe what Garrod
Supplemental Data. Wang et al. (2014). Plant Cell /tpc
 Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio
 Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
4) Please cite Dagda et al J Biol Chem 284: , for any publications or presentations resulting from use or modification of the macro.
 Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct
 SUPPLEMENTAL FIGURE LEGENDS Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct waves. (A) A representative sub-lineage (ABala) of the C. elegans lineage tree (adapted from
SUPPLEMENTAL FIGURE LEGENDS Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct waves. (A) A representative sub-lineage (ABala) of the C. elegans lineage tree (adapted from
Rho1 binding site PtdIns(4,5)P2 binding site Both sites
 localization Mutation site DMSO LatB WT F77A I115A I131A K134A Rho1 binding site PtdIns(4,5)P2 binding site Both sites E186A E199A N201A R84A-E186A-E199A L131A-K136A-E186A L131A-E186A-E199A K136A-E186A-E199A
localization Mutation site DMSO LatB WT F77A I115A I131A K134A Rho1 binding site PtdIns(4,5)P2 binding site Both sites E186A E199A N201A R84A-E186A-E199A L131A-K136A-E186A L131A-E186A-E199A K136A-E186A-E199A
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
FSC-W FSC-H CD4 CD62-L
 Supplementary Fig. 1 a SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H CD4 CD62-L b SSC-A FSC-A FSC-W FSC-A FSC-A 7-AAD FSC-A CD4 IL-9 CD4 c SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H 7-AAD KI67 Annexin-V 7-AAD d I L -5
Supplementary Fig. 1 a SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H CD4 CD62-L b SSC-A FSC-A FSC-W FSC-A FSC-A 7-AAD FSC-A CD4 IL-9 CD4 c SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H 7-AAD KI67 Annexin-V 7-AAD d I L -5
Ab1 (57-68) Ab2 ( ) Ab3 ( ) Ab4 ( ) GBP-N domain GBP-C domain CARD domain
 MDKPVCLIDTGSDGKLCVQQAALQVLQQIQQPVVVVAVVGLYRTGKSFLMNRLAG 55 KRTGFALSSNIKPKTEGIWMWCVPHPTKAGTSLVLLDTKGLGDVEKGDSKRDTYI 110 FSLTVLLSSTLVYNSRGVIDNKAMEELQYVTELIEHIKVTPDEDADDCTAFAKFF 165 PHFIWCLRDFTLELKLDGKDLTEDEYLEFALKLRPGTLKKVMMYNLPRECIQKFF
MDKPVCLIDTGSDGKLCVQQAALQVLQQIQQPVVVVAVVGLYRTGKSFLMNRLAG 55 KRTGFALSSNIKPKTEGIWMWCVPHPTKAGTSLVLLDTKGLGDVEKGDSKRDTYI 110 FSLTVLLSSTLVYNSRGVIDNKAMEELQYVTELIEHIKVTPDEDADDCTAFAKFF 165 PHFIWCLRDFTLELKLDGKDLTEDEYLEFALKLRPGTLKKVMMYNLPRECIQKFF
Structure and RNA-binding properties. of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex
 Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
In order to confirm the contribution of diffusion to the FRAP recovery curves of
 Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Fig. S1. Expression pattern of moody-gal4 in third instar. Maximum projection illustrating a dissected moody-gal4>ngfp L3 larva stained for Repo
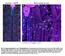 Fig. S1. Expression pattern of moody-gal4 in third instar. Maximum projection illustrating a dissected moody-gal4>ngfp L3 larva stained for Repo (magenta), Fas2 (blue) and GFP (green) in overview (A) and
Fig. S1. Expression pattern of moody-gal4 in third instar. Maximum projection illustrating a dissected moody-gal4>ngfp L3 larva stained for Repo (magenta), Fas2 (blue) and GFP (green) in overview (A) and
Essential roles of zebrafish rtn4/nogo paralogues in embryonic development
 Essential roles of zebrafish rtn4/nogo paralogues in embryonic development Pinzón-Olejua et al. Pinzón-Olejua et al. Neural Development 2014, 9:8 Pinzón-Olejua et al. Neural Development 2014, 9:8 RESEARCH
Essential roles of zebrafish rtn4/nogo paralogues in embryonic development Pinzón-Olejua et al. Pinzón-Olejua et al. Neural Development 2014, 9:8 Pinzón-Olejua et al. Neural Development 2014, 9:8 RESEARCH
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11419 Supplementary Figure 1 Schematic representation of innate immune signaling pathways induced by intracellular Salmonella in cultured macrophages. a, During the infection Salmonella
doi:10.1038/nature11419 Supplementary Figure 1 Schematic representation of innate immune signaling pathways induced by intracellular Salmonella in cultured macrophages. a, During the infection Salmonella
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/11/eaao4709/dc1 Supplementary Materials for Pushing the limits of photoreception in twilight conditions: The rod-like cone retina of the deep-sea pearlsides Fanny
advances.sciencemag.org/cgi/content/full/3/11/eaao4709/dc1 Supplementary Materials for Pushing the limits of photoreception in twilight conditions: The rod-like cone retina of the deep-sea pearlsides Fanny
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
Cryo-EM data collection, refinement and validation statistics
 1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
BCH 4054 Spring 2001 Chapter 33 Lecture Notes
 BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
Waithe et al Supplementary Figures
 Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
Maternal Control of GermLayer Formation in Xenopus
 Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Supplementary Figure 1
 Supplementary Figure 1 Single-cell RNA sequencing reveals unique sets of neuropeptides, transcription factors and receptors in specific types of sympathetic neurons (a) Dissection of paravertebral SGs
Supplementary Figure 1 Single-cell RNA sequencing reveals unique sets of neuropeptides, transcription factors and receptors in specific types of sympathetic neurons (a) Dissection of paravertebral SGs
Sonic hedgehog (Shh) signalling in the rabbit embryo
 Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
** LCA LCN PCA
 % of wild type value % of wild type value a 12 1 8 2 b 12 1 8 2 LCA LCN PCA Col- sod3-1 Supplementary Figure 1 sod3-1 influences cell proliferation. (a) Fifth leaf cell area (LCA) and leaf cell number
% of wild type value % of wild type value a 12 1 8 2 b 12 1 8 2 LCA LCN PCA Col- sod3-1 Supplementary Figure 1 sod3-1 influences cell proliferation. (a) Fifth leaf cell area (LCA) and leaf cell number
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The orientation of cell division influences cell-fate choice in the developing mammalian retina
 Development 130, 2329-2339 2003 The Company of Biologists Ltd doi:10.1242/dev.00446 2329 The orientation of cell division influences cell-fate choice in the developing mammalian retina Michel Cayouette*,
Development 130, 2329-2339 2003 The Company of Biologists Ltd doi:10.1242/dev.00446 2329 The orientation of cell division influences cell-fate choice in the developing mammalian retina Michel Cayouette*,
Section 7. Junaid Malek, M.D.
 Section 7 Junaid Malek, M.D. RNA Processing and Nomenclature For the purposes of this class, please do not refer to anything as mrna that has not been completely processed (spliced, capped, tailed) RNAs
Section 7 Junaid Malek, M.D. RNA Processing and Nomenclature For the purposes of this class, please do not refer to anything as mrna that has not been completely processed (spliced, capped, tailed) RNAs
Six3 inactivation reveals its essential role for the formation and patterning of
 Development 129, 4057-4063 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV2906 4057 Six3 inactivation reveals its essential role for the formation and patterning of the vertebrate
Development 129, 4057-4063 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV2906 4057 Six3 inactivation reveals its essential role for the formation and patterning of the vertebrate
Oligomerization of DH Domain Is Essential for Dbl-Induced Transformation
 MOLECULAR AND CELLULAR BIOLOGY, Jan. 2001, p. 425 437 Vol. 21, No. 2 0270-7306/01/$04.00 0 DOI: 10.1128/MCB.21.2.425 437.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Oligomerization
MOLECULAR AND CELLULAR BIOLOGY, Jan. 2001, p. 425 437 Vol. 21, No. 2 0270-7306/01/$04.00 0 DOI: 10.1128/MCB.21.2.425 437.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Oligomerization
Biology Tutorial. Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan
 Biology Tutorial Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan Viruses A T4 bacteriophage injecting DNA into a cell. Influenza A virus Electron micrograph of HIV. Cone-shaped cores are
Biology Tutorial Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan Viruses A T4 bacteriophage injecting DNA into a cell. Influenza A virus Electron micrograph of HIV. Cone-shaped cores are
Chapter 17. From Gene to Protein. Biology Kevin Dees
 Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Transitivity-dependent and transitivity-independent cell-to-cell movement of RNA
 Himber et al. Transitivity-dependent and transitivity-independent cell-to-cell movement of RNA silencing SUPPLEMENTAL MATERIAL Supplemental material 1. Short-range movement of GFP silencing affects a nearly
Himber et al. Transitivity-dependent and transitivity-independent cell-to-cell movement of RNA silencing SUPPLEMENTAL MATERIAL Supplemental material 1. Short-range movement of GFP silencing affects a nearly
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Mitochondrial Dynamics Is a Distinguishing Feature of Skeletal Muscle Fiber Types and Regulates Organellar Compartmentalization
 Cell Metabolism Supplemental Information Mitochondrial Dynamics Is a Distinguishing Feature of Skeletal Muscle Fiber Types and Regulates Organellar Compartmentalization Prashant Mishra, Grigor Varuzhanyan,
Cell Metabolism Supplemental Information Mitochondrial Dynamics Is a Distinguishing Feature of Skeletal Muscle Fiber Types and Regulates Organellar Compartmentalization Prashant Mishra, Grigor Varuzhanyan,
Table S1. Sequence of primers used in RT-qPCR
 Table S1. Sequence of primers used in RT-qPCR Primer Name P16Ink4a-F P16Ink4a-R P15Ink4b-F P15Ink4b-R P19Arf-F P19Arf-R P53-F P53-R P21cip1-F P21cip1-R P27kip1-F P27kip1-R P18Ink4c-F P18Ink4c-R P19Ink4d-F
Table S1. Sequence of primers used in RT-qPCR Primer Name P16Ink4a-F P16Ink4a-R P15Ink4b-F P15Ink4b-R P19Arf-F P19Arf-R P53-F P53-R P21cip1-F P21cip1-R P27kip1-F P27kip1-R P18Ink4c-F P18Ink4c-R P19Ink4d-F
Prokaryotic Regulation
 Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
GENE REGULATION AND PROBLEMS OF DEVELOPMENT
 GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
Supplemental Data. Hou et al. (2016). Plant Cell /tpc
 Supplemental Data. Hou et al. (216). Plant Cell 1.115/tpc.16.295 A Distance to 1 st nt of start codon Distance to 1 st nt of stop codon B Normalized PARE abundance 8 14 nt 17 nt Frame1 Arabidopsis inflorescence
Supplemental Data. Hou et al. (216). Plant Cell 1.115/tpc.16.295 A Distance to 1 st nt of start codon Distance to 1 st nt of stop codon B Normalized PARE abundance 8 14 nt 17 nt Frame1 Arabidopsis inflorescence
Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering
 Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering by Valverde et. Al Published in Science 2004 Presented by Boyana Grigorova CBMG 688R Feb. 12, 2007 Circadian Rhythms: The Clock Within
Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering by Valverde et. Al Published in Science 2004 Presented by Boyana Grigorova CBMG 688R Feb. 12, 2007 Circadian Rhythms: The Clock Within
Comparing Genomes! Homologies and Families! Sequence Alignments!
 Comparing Genomes! Homologies and Families! Sequence Alignments! Allows us to achieve a greater understanding of vertebrate evolution! Tells us what is common and what is unique between different species
Comparing Genomes! Homologies and Families! Sequence Alignments! Allows us to achieve a greater understanding of vertebrate evolution! Tells us what is common and what is unique between different species
GENETICS - CLUTCH CH.11 TRANSLATION.
 !! www.clutchprep.com CONCEPT: GENETIC CODE Nucleotides and amino acids are translated in a 1 to 1 method The triplet code states that three nucleotides codes for one amino acid - A codon is a term for
!! www.clutchprep.com CONCEPT: GENETIC CODE Nucleotides and amino acids are translated in a 1 to 1 method The triplet code states that three nucleotides codes for one amino acid - A codon is a term for
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1121356/dc1 Supporting Online Material for Polar PIN Localization Directs Auxin Flow in Plants Justyna Wiśniewska, Jian Xu, Daniela Seifertová, Philip B. Brewer, Kamil
www.sciencemag.org/cgi/content/full/1121356/dc1 Supporting Online Material for Polar PIN Localization Directs Auxin Flow in Plants Justyna Wiśniewska, Jian Xu, Daniela Seifertová, Philip B. Brewer, Kamil
Three different fusions led to three basic ideas: 1) If one fuses a cell in mitosis with a cell in any other stage of the cell cycle, the chromosomes
 Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
The Drosophila Pkn protein kinase is a Rho/Rac effector target required for dorsal closure during embryogenesis
 The Drosophila Pkn protein kinase is a Rho/Rac effector target required for dorsal closure during embryogenesis Yu Lu and Jeffrey Settleman 1 Massachusetts General Hospital Cancer Center and Harvard Medical
The Drosophila Pkn protein kinase is a Rho/Rac effector target required for dorsal closure during embryogenesis Yu Lu and Jeffrey Settleman 1 Massachusetts General Hospital Cancer Center and Harvard Medical
A MicroRNA as a Translational Repressor of APETALA2 in Arabidopsis Flower Development
 A MicroRNA as a Translational Repressor of APETALA2 in Arabidopsis Flower Development Xuemei Chen Waksman Institute, Rutgers University, Piscataway, NJ 08854, USA. E-mail: xuemei@waksman.rutgers.edu Plant
A MicroRNA as a Translational Repressor of APETALA2 in Arabidopsis Flower Development Xuemei Chen Waksman Institute, Rutgers University, Piscataway, NJ 08854, USA. E-mail: xuemei@waksman.rutgers.edu Plant
Supporting Information
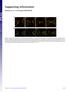 Supporting Information Fleissner et al. 10.1073/pnas.0907039106 Fig. S1. (A) MAK-2-GFP localized to CATs tips is not bound by membrane. his-3::pccg1 mak-2-gfp; mak-2 strain labeled with membrane dye FM4
Supporting Information Fleissner et al. 10.1073/pnas.0907039106 Fig. S1. (A) MAK-2-GFP localized to CATs tips is not bound by membrane. his-3::pccg1 mak-2-gfp; mak-2 strain labeled with membrane dye FM4
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Tumorhead distribution to cytoplasmic membrane of neural plate cells is positively regulated by Xenopus p21-activated kinase 1 (X-PAK1)
 Developmental Biology 308 (2007) 169 186 www.elsevier.com/locate/ydbio Tumorhead distribution to cytoplasmic membrane of neural plate cells is positively regulated by Xenopus p21-activated kinase 1 (X-PAK1)
Developmental Biology 308 (2007) 169 186 www.elsevier.com/locate/ydbio Tumorhead distribution to cytoplasmic membrane of neural plate cells is positively regulated by Xenopus p21-activated kinase 1 (X-PAK1)
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10923 Supplementary Figure 1 Ten-a and Ten-m antibody and cell type specificities. a c, Representative single confocal sections of a Drosophila NMJ stained with antibodies to Ten-a (red),
doi:10.1038/nature10923 Supplementary Figure 1 Ten-a and Ten-m antibody and cell type specificities. a c, Representative single confocal sections of a Drosophila NMJ stained with antibodies to Ten-a (red),
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
A Genome-Wide Analysis of Arabidopsis Rop-Interactive CRIB Motif Containing Proteins That Act as Rop GTPase Targets
 The Plant Cell, Vol. 13, 2841 2856, December 2001, www.plantcell.org 2001 American Society of Plant Biologists A Genome-Wide Analysis of Arabidopsis Rop-Interactive CRIB Motif Containing Proteins That
The Plant Cell, Vol. 13, 2841 2856, December 2001, www.plantcell.org 2001 American Society of Plant Biologists A Genome-Wide Analysis of Arabidopsis Rop-Interactive CRIB Motif Containing Proteins That
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Welcome to Class 21!
 Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice Supplementary Figure 2.
 Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
chokh/rx3 specifies the retinal pigment epithelium fate independently of eye morphogenesis
 Developmental Biology 288 (2005) 348 362 www.elsevier.com/locate/ydbio chokh/rx3 specifies the retinal pigment epithelium fate independently of eye morphogenesis Agustin Rojas-Muñoz*, Ralf Dahm 1, Christiane
Developmental Biology 288 (2005) 348 362 www.elsevier.com/locate/ydbio chokh/rx3 specifies the retinal pigment epithelium fate independently of eye morphogenesis Agustin Rojas-Muñoz*, Ralf Dahm 1, Christiane
downstream (0.8 kb) homologous sequences to the genomic locus of DIC. A DIC mutant strain (ro- 6
 A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
23-. Shoot and root development depend on ratio of IAA/CK
 Balance of Hormones regulate growth and development Environmental factors regulate hormone levels light- e.g. phototropism gravity- e.g. gravitropism temperature Mode of action of each hormone 1. Signal
Balance of Hormones regulate growth and development Environmental factors regulate hormone levels light- e.g. phototropism gravity- e.g. gravitropism temperature Mode of action of each hormone 1. Signal
7.06 Problem Set
 7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
