In order to confirm the contribution of diffusion to the FRAP recovery curves of
|
|
|
- Claud Ray
- 6 years ago
- Views:
Transcription
1 Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP and PC::GFP (Fig. 4) we performed curve smoothing tests for these recovery curves and compared them to GFPnls (diffusion dependent) and H2A::RFP (diffusion independent) recovery curves confirming a contribution of diffusion to recovery for all PC::GFP and PH::GFP data sets (Fig. S). 2) Parameter extraction and cross validation 2.) Extraction of kinetic parameters from FRAP data The FRAP recovery data were analysed by fitting kinetic models (Mueller et al. 28) to averaged FRAP recovery data shown in Figure 4. This fitting procedure enables the extraction of values for diffusion coefficient (Df), the pseudo first order association rate k* on and the dissociation rate k off. 2.2) Adaptation of model for optimal parameter combination An additional step was performed to optimise extracted kinetic parameters. After the calculation of the radius of the model nucleus (RM) (Mueller et al. 28) an additional set of radii was defined, composed of radii - pixels from RM to +2 pixels. These 3 radii were used as input values for the reaction-diffusion or pure-difusion model fit to the experimental data. The resulting set of individual extracted kinetic parameters and their confidence intervals as well as the goodness of fit was used to select the optimal radius for the experiment. This selection consisted of a weighted search with /3 of the
2 2 weight being given to the goodness of the confidence intervals of association and dissociation constants, /3 to the goodness of the extracted diffusion constant confidence interval, /6 to the size of the squared sum of residuals and /6 to the distance from the initial RM, with smaller distances being favoured. MATLAB files are available on request. 2.3) Contribution of binding to FRAP recovery curves To evaluate the role of binding in the recovery kinetics we compared reaction-diffusion (3 extracted parameters: Df, k* on and k off ) and pure-difusion model fits (single extracted parameter: Df) as described in (Mueller et al. 28) to our experimental data. In all cases shown in Figure 4, the best fit was given by the full reaction-diffusion model, indicating the presence of a bound fraction, and giving extracted values for Df, k* on and k off. 2.4) Cross-validation of extracted Df In addition to the extracted values for Df from fitting the reaction- diffusion model, the Df for each protein in each cell type was measured independently. This was achieved by performing FRAP on the region of the metaphase cell that is outside chromatin and fitting the pure diffusion model (Mueller et al. 28) to the recovery data, giving an independent and direct measure of Df. Interphase values were calculated by conversion via diffusion coefficients measured for GFP by fitting the pure diffusion model to FRAP recovery curves measured in both interphase and metaphase, (Table S). The values of Df thus measured showed excellent agreement with those extracted from fitting the full model (Figure S3).
3 3 2.5) Robustness of extracted k* on, k off The robustness of the extracted k* on and k off values was examined by simulations performed at the value of Df that was extracted from the reaction-diffusion model fit, and in which k* on and k off were varied, and the fit to experimental data was evaluated (Fig. S4). This analysis showed that for most data sets, a limited range of k* on and k off values gave optimal fits to the data (Fig. S4). 3) Other models 3.) Localised binding sites: metaphase The effect of localized binding sites in metaphase was examined using the local binding site model described in (Beaudouin et al. 26; Sprague et al. 26), showing that both the improved global binding (Mueller et al. 28) and the localized binding (Beaudouin et al. 26; Sprague et al. 26) models give essentially identical results in conditions of low binding, as is the case for the metaphase data shown here (data not shown). Unlike the Müller model (Mueller et al. 28) the Sprague model (Beaudouin et al. 26; Sprague et al. 26) does not include consideration of the radial bleach profile. Thus in order to achieve consistency of analysis, the Müller Model(Mueller et al. 28) was used for analysis of all data sets. 3.2) Non homogeneous distribution of proteins: interphase To test for the effect of non-homogeneity in protein distribution observed in interphase (Figure 2 and 3) on extracted kinetic parameters, we adapted the model described in (Mueller et al. 28) from its original application to redistribution of photoactivatable
4 4 GFP, to render it applicable to the analysis of FRAP recovery curves, described here. Fitting this model to interphase data for individual nuclei gave similar values for the three extracted parameters whether initial distribution was assumed to be heterogenous or homogenous (Figure S2). 3.2.) Generation of images of single nuclei. In order to construct input protein distribution images for parameter extraction, all prebleach images of a single nucleus (25) were averaged and used to threshold the region of the nucleus in the total image. This region was selected to define the nucleus within the average image of 2s before photobleaching. Due to the speed of scanning, it was not feasible to image the entire nucleus. The shape of NB and SOP nuclei approximates well to a circle, thus the initial binding site distribution in the entire nucleus was reconstructed from the image of the "equatorial" region, covering approximately 2/3 of the nucleus. On the resulting image a circle of radius RM (model nucleus radius calculated as described in (Mueller et al. 28) with adaptation as described in 2. above) was defined with the bleach region centered. This image was used to give the initial distribution of binding sites in the nucleus. In order to produce the first postbleach image, a bleach pattern with parameters describing the bleach spot profile was calculated from the experimental data (Mueller et al. 28) and was superimposed on the prebleach image. Matlab files for image processing are available on request ) Extraction of kinetic parameters from FRAP data, taking non homogeneous protein distribution into account. The intensity distribution images generated as described above were used as input for
5 5 fitting the spatial model described below to the individual FRAP recovery curve for each nucleus, and extraction of parameters. The spatial model was implemented in Mathematica (Wolfram) and is available on request. The reaction-diffusion system is simulated on a 2D circular domain, with a Neumann no-flux condition imposed on the boundary. The method-of-lines is used to numerically solve the resulting partial-differential equation, where a second-order finite difference method is used to discretize the diffusion operator on a uniform mesh. The spatial discretization gives rise to a coupled system of ordinary differential equations for the free and bound concentrations at each mesh point, which is then numerically integrated using an implicit solution scheme. The unknown parameters in the model consist of: the diffusion constant Df, the off-rate of the reaction k off, and the ratio of the total amount of free molecules to bound molecules, Free. Given a value for the free fraction, Free, the initial conditions for the free and bound proteins are obtained from the smoothed, pre-bleached images. Given the values of k off and Free, the spatially varying k on [C] is computed from the intensity distribution of the averaged chromatin images, following the methodology of (Mueller et al. 28). In order to ensure the positivity of k on [C] in the model, a lower bound on the free fraction is imposed, whose value is required to be greater than the minimum chromatin intensity over its average for the circular domain. The unknown parameters (Df, k off, Free) are estimated from the measured fluorescence recovery curve for each individual nucleus by solving the inequality constrained optimization problem using the interior point method. As starting values for these three parameters, the extracted values from averaged data were used (Fig. 4, Table S).
6 6 Supplementary Legends Figure S. Diffusion influences FRAP recovery for GFPnls, PC::GFP, PH::GFP and H2A::RFP. Diffusion test was performed using an adaptation of the method of curve smoothing (Mueller et al. 28). (A-F) Radial intensity profiles of FRAP experiments at four different time points after photobleaching (time in seconds is shown at the right of each plot). The gaussian edges of intensity profiles normalized to prebleach levels (Mueller et al. 28) are plotted (symbols) and were fitted using linear regression (solid lines). The gray background indicates the bleach region. (A) H2A::RFP recovery is not affected by diffusion, indicated by similar slopes of lines at all four time points. Comparison of the extracted slopes was performed using ANCOVA (p-value given on each plot represents significance of difference between slopes at the four time points). (B-F) GFP-nls, PC::GFP and PH::GFP FRAP recovery shows an influence of diffusion, indicated by gradual flattening of radial profiles at later time points. (E) Comparative summary plot. For data in (A-F), the value /slope was calculated for each linear fit and normalized to the slope at time. These values are plotted for each data set for consecutive time points, showing a gradual increase in (/slope) at later time points for all experiments with the exception of H2A::RFP (black) for which little change was detected. Figure S2. Comparison of the effects of binding site non-homogeneity on parameters extracted from FRAP experiments. Extracted diffusion (A,D,G and J), free fraction (B,E,H and K) and dissociation rates (k off, C,F,I and L) of PH::GFP (A-C, G-I) and PC::GFP (D-F, J-L) FRAP experiments in neuroblast interphase (A-F) and sensory organ precursor cell interphase (G-L) were analysed using an adaptation of the model described in (Mueller et al. 28). (See Supplementary Information, FRAP Data
7 7 Analysis, for detailed description). Black bars represent the mean and 95% confidence intervals of the extracted parameters using the same model with an initial homogeneous distribution of binding sites and grey bars represent the mean and 95% confidence intervals of the extracted parameters using the image-based heterogeneous distribution of binding sites for each nucleus. n represents number of nuclei used in each experiment. 2-tailed paired t-tests were performed for each comparison resulting in p-values >.5 with the exception of B (p=.), C (p=.69), D (p=.282) and E (p=.63). Dashed lines represent parameters extracted using the FRAP model described in (Mueller et al. 28) and shown in Figure 4 and Table S. Figure S3. Cross validation of extracted diffusion constants by independent measurements. (A) Comparison of diffusion constants extracted from fitting 3 parameter FRAP model in all cell types (Df (), black) to diffusion constants calculated by fitting single parameter FRAP model (diffusion only) to FRAP recovery performed on the non-chromatin volume in metaphase (Df (2), grey). The interphase Df values (grey) were calculated using GFPnls for calibration as described in Supplementary Information. The Df values calculated by the two procedures show good agreement. NB and SOP indicate neuroblast and SOP interphase and NBmet and SOPmet indicate neuroblast and SOP metaphase. piia and piib indicate the interphase of the respective cells. Data show mean of at least four measurements for each cell type. Error bars represent 95% confidence intervals. (B) Estimated molecular weight of PH::GFP and PC::GFP in neuroblasts (black) and SOPs (grey). Estimations were based on the extracted Df for GFPnls, PH::GFP and PC::GFP in regions outside chromatin at metaphase in neuroblasts and SOPs and calculated using the following
8 8 equation: Mw protein = Mw GFP /(Df protein /Df GFP )^3. The Mw estimated for PC::GFP is consistent with the predicted size of the PRC complex. In contrast, the Mw estimated for PH::GFP is approximately 5MDa. The PH protein has not been reported to participate in such large complexes, thus this result suggests that the extracted diffusion constant for PH::GFP may comprise both the true diffusion and a binding component (Mueller et al. 28). Figure S4. Parameter space for best fits of FRAP model to recovery data. For each FRAP recovery data set shown in Figure 4, simulations were performed in which Df was fixed to the value extracted from the 3 parameter fit (Fig. S3a, grey bars; Table S), and k* on and k off were varied between -4 and. For each simulation, the fit to the experimental data was evaluated as squared sum of residuals (ssrs). The white, red or black lines delineate ssrs.25 times larger than the minimum ssr found. Top row: interphase and metaphase best fit regions from each data set as indicated above the plots, are superimposed for comparison. Below: ssrs for each data set are plotted individually as heat maps (colour scale for ssrs is shown at the right of the plot.) Figure S5. Dot blot analysis of α-h3k27me3s28ph antibody. Serial dilutions of synthetic peptides corresponding to N-terminal sequence of histone H3 (amino acids 9-37), with different S28 phosphorylation and K27 methylation status as indicated above the figure, were spotted on a PVDF membrane and probed with the α- H3K27me3S28ph antibody (dilution : 2). For detection a secondary anti rabbit horseradish peroxidase-conjugated antibody and the Enhanced Chemiluminescence (ECL) detection system were used. To ensure equal peptide loading, a duplicate membrane was stained with Ponceau S.
9 9 Movie S. PH::GFP in neuroblast. Green channel: PH::GFP under the worniu-gal4 driver is visualized in neuroblast and ganglion mother cells (GMCs). Red channel: Histone H2A::RFP is expressed under the ubiqutin promoter and visualized in all cells. RFP marked chromatin becomes visible in the neuroblast at mitosis. The movie starts at interphase; the largest cell is the neuroblast. One mitotic division up to the next telophase is shown. Movie S2. PC::GFP in neuroblast. Green channel: PC::GFP is expressed under the Pc promoter and is visualized in all cells. Red channel: Histone H2A::RFP is expressed under the ubiqutin promoter and visualized in all cells. RFP marked chromatin becomes visible in the neuroblast at mitosis. The movie starts at interphase; the largest cell is the neuroblast. One mitotic division up to the next telophase is shown. Movie S3. PH::GFP in SOP. Both PH::GFP (green channel) and histone H2A::RFP were expressed under the neuralized-gal4 driver and are visible in specifically in the SOP and its daughter cells piia and piib. RFP marked chromatin is visible at all stages. The movie starts at SOP interphase. One mitotic division up to the next interphase is shown. At the end of the movie, the two daughter cells piia and piib are seen. Movie S4. PC::GFP in SOP. Green channel: PC::GFP is expressed under the Pc promoter and is visualized in all cells. Red channel: histone H2A::RFP was expressed under the neuralized-gal4 driver and is visible in specifically in the SOP and its
10 daughter cells piia and piib. RFP marked chromatin is visible at all stages. The movie starts at SOP interphase. The SOP is the largest cell and the only one showing red signal. One mitotic division up to the next interphase is shown. At the end of the movie, the two daughter cells piia and piib are seen. Table S. Compilation of measured and extracted parameters of PH::GFP, PC::GFP and GFPnls in Neuroblast interphase (NB) and metaphase (NBmet), SOP interphase (SOP) and metaphase (SOPmet), piia interphase (piia) and piib interphase (piib). For the quantification parameters, volume measurements in cubic micrometers, determined by GFP fluorescence (Blue masks in Figure 2 and 3), are shown (A) as well as number and micromolar concentrations of GFP-fused (B, C), endogenous (D, E) and endogenous in yw flies (F,G) molecules of PH and PC. Kinetic parameters extracted using the method described in (Mueller et al. 28) are shown in the section Kinetic parameters. The radius used for the parameter extraction is shown in µm (H). (I) represents the extracted diffusion from the full model (3 parameter fit) for PH::GFP and PC::GFP and from the pure diffusion model (single parameter fit) for GFPnls. (J) represents the extracted diffusion from the pure diffusion model in regions outside chromatin in NBmet and SOPmet, and the estimated PH::GFP and PC::GFP diffusions in interphase of all other cell types through the comparison with GFPnls diffusions (Fig.S3). Residence time (M) was calculated as (/K). The fraction of bound molecules in the chromatin region (N) was calculated by the following equation: *(L)/(L+M). The total fraction of bound molecules (O) was calculated with the following equation: (N)*(T)/(B). (T) represents the number of GFP-fused proteins that are localized in the region determined by H2A::RFP fluorescence (Yellow masks in Figures 2 and 3). Number of bound GFP-fused (P), GFP-fused and endogenous (Q) and endogenous in
11 yw flies (R) molecules are shown. In the Image-based parameters section are listed the volume in cubic micrometers occupied by chromatin (S) and the number of GFP proteins that are in this volume (T). As (T), (S) was determined by H2A::RFP fluorescence. The calculated fraction of bound molecules in the chromatin region, without the assumption of equilibrium, according to equation 8 of Supplementary Information Mathematical modeling is listed as (U). The number of GFP-fused proteins bound to chromatin is listed as (W). The total fraction of bound molecules (V) was calculated with the ratio W/B. In the Modelling parameters section are listed the parameters used for the model shown in Figure 5: pseudo-first order association rate (X), the dissociation rate (Y) the number of endogenous Polycomb proteins (Z), as well as the cell (AA) and chromatin (AB) volumes. These parameters were selected from the experimentally determined values listed in (L), (K), (F), (A) and (S). Also shown are the assumed number of binding sites (AC), representing the maximum possible number of H3K27 methylated tails in the diploid genome based on H3K27me3 distributions in polytene chromosomes and genome-wide ChIP profiles, assuming methylation of all H3 tails within a region of H3K27me3 signal. Based on this number of binding sites, the calculated micromolar dissociation constant (AD) is shown.
12 84648, FigureS, Fonseca A H2A::RFP - SOP E Diffusion test Normalised Intensity.2. P= Radius ( m) normalised /slope time points H2A::RFP (SOP) GFPnls (SOP) PC::GFP (SOP) PH::GFP (SOP) PC::GFP (NB) PH::GFP (NB) B GFPnls - SOP F GFPnls - NB Normalised Intensity..4 P=.3..5 Radius ( m) Normalised Intensity. P= Radius (um).4.2 C PC::GFP - SOP G PC::GFP - NB Normalised Intensity..4 P< Radius (um).6.3 Normalised Intensity. P= Radius (um)..8 D PH::GFP - SOP H PH::GFP - NB Normalised Intensity. P< Radius (um) 8 5 Normalised Intensity.2. P=.2..5 Radius (um).2
13 84648, Figure S2, Fonseca NB PH Diffusion Free Fraction k off A B C D f (µm 2.s - ) n = 5 Homogeneous Heterogeneous Free Fraction..4.2 n = 5. Homogeneous Heterogeneous k off (s - ).. Homogeneous Heterogeneous n = 5 D E F. NB PC D f (µm 2.s - ) Homogeneous Heterogeneous n = 4 Free Fraction.4.2. Homogeneous Heterogeneous n = 4 k off (s - ). Homogeneous Heterogeneous n = 4 G H I.5 SOP PH D f (µm 2.s - )..5. Homogeneous Heterogeneous n = 6 Free Fraction.4.2. Homogeneous Heterogeneous n = 6 k off (s - ).. Homogeneous Heterogeneous n = 6 SOP PC D f (µm 2.s - ) J K L n = 6 Homogeneous Heterogeneous Free Fraction..4.2 n = 6. Homogeneous Heterogeneous k off (s - )... Homogeneous Heterogeneous n = 6
14 84648, Figure S3, Fonseca A D f (µm 2.s - ) B Estimated MW (kda) PH NB PH NBmet PH SOP PH SOPmet PH piia PH piib PC NB PC NBmet PC SOP PC SOPmet PC piia Df () Df (2) PC piib NB SOP Polycomb Polyhomeotic
15 84648, Figure S4, Fonseca
16 84648, Figure S5, Fonseca H3 unmodified H3 S28ph H3K27me3 H3K27meS28ph H3K27me2S28ph H3K27me3S28ph H3K9me3Sph 5 pmol pmol 2 pmol Ponceau S (5 pmol)
17 PH::GFP PC::GFP GFPnls Section Variable ID Variable NB NBmet SOP SOPmet piia piib NB Nbmet SOP SOPmet piia piib NB Nbmet SOP SOPmet piia piib A Volume (µm 3 ) ± ± ± ± ± ± ± ± ± ± ± ± 9.76 B # GFP 3949 ± ± ± ± ± ± ± ± ± ± ± ± 484 Image-based parameters Kinetic parameters Quantification C µm GFP.98 ±.6.9 ±.3.36 ± ±.4 6 ±.2 ±..7 ±.8.27 ±.2.36 ±.2. ±..36 ±.3.36 ±.3 D # end ± ± ± ± ± ± 87 E µm end.23 ±.3.9 ±..2 ±..6 ±.2.2 ±..2 ±.2 F # end yw 465 ± ± ± ± ± ± 5355 G µm end yw.38 ±.9.5 ±.3.48 ±.3.4 ±.4.48 ±.3.48 ±.3 H Radius (µm) I Df (µm 2.s - ).5 ±.7.26 ±.5.76 ± ±.56.2 ±.3. ± ± ± ± ± ± ± ±.96.5 ± ± ± ± ± J Df 2 (µm 2.s - ).4 ±.7.5 ±.6 8 ±.5.77 ±.6.57 ±.4.48 ± ± ± ± ± ±.4.97 ±.2 K k off (s - ).23 ±.5.2 ±.7. ±..24 ±.6.5 ±..3 ±. 2.9 ±.75.3 ±.8.72 ±.4.2 ±.7.24 ±..27 ±.4 L k* on (s - ). ±.4. ±.5.3 ±.4.5 ±.8.29 ±.5.27 ±.6.5 ±.38.6 ±.6.8 ±.8.3 ±.3.7 ±.4.4 ±.3 M Rtime (s) N Fbound chr (%) O Fbound total (%) P # GFP bound 436 ± ± ± ± ± ± ± ± ± ± ± ±97 Q #GFP + end bound 8498 ± ± ± ± ± ± 34 R # end bound yw 7688 ± ± ± ± ± ± 255 S Volume chr (µm 3 ) T # GFP chr 3,977 6,365 3,562 3,75 U Fbound chr (%) V Fbound total (%) W # GFP chr bound ,82 X k* (s - ) Y k - (s - ) Modeling parameters Z # PC AA Volume Cell (µm3) AB Volume Chr (µm3) AC # chromatin sites AD Kd (µm)
Fluorescence fluctuation microscopy: FRAP and FCS
 Fluorescence fluctuation microscopy: FRAP and FCS Confocal laser scanning fluorescence microscope for the mobility analysis in living cells Ensemble measurements of protein mobility and interactions pre
Fluorescence fluctuation microscopy: FRAP and FCS Confocal laser scanning fluorescence microscope for the mobility analysis in living cells Ensemble measurements of protein mobility and interactions pre
Figure 1. Automated 4D detection and tracking of endosomes and polarity markers.
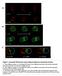 A t = 0 z t = 90 B t = 0 z t = 90 C t = 0-90 t = 0-180 t = 0-300 Figure 1. Automated 4D detection and tracking of endosomes and polarity markers. A: Three different z planes of a dividing SOP shown at
A t = 0 z t = 90 B t = 0 z t = 90 C t = 0-90 t = 0-180 t = 0-300 Figure 1. Automated 4D detection and tracking of endosomes and polarity markers. A: Three different z planes of a dividing SOP shown at
The Process of Cell Division. Lesson Overview. Lesson Overview The Process of Cell Division
 Lesson Overview 10.2 The Process of Cell Division Chromosomes genetic information passed from parent to offspring is carried by chromosomes. Chromosomes enable precise DNA separation during cell division.
Lesson Overview 10.2 The Process of Cell Division Chromosomes genetic information passed from parent to offspring is carried by chromosomes. Chromosomes enable precise DNA separation during cell division.
BIOLOGY. Chapter 10 CELL REPRODUCTION PowerPoint Image Slideshow
 BIOLOGY Chapter 10 CELL REPRODUCTION PowerPoint Image Slideshow FIGURE 10.1 A sea urchin begins life as a single cell that (a) divides to form two cells, visible by scanning electron microscopy. After
BIOLOGY Chapter 10 CELL REPRODUCTION PowerPoint Image Slideshow FIGURE 10.1 A sea urchin begins life as a single cell that (a) divides to form two cells, visible by scanning electron microscopy. After
10.2 The Process of Cell Division
 10.2 The Process of Cell Division Lesson Objectives Describe the role of chromosomes in cell division. Name the main events of the cell cycle. Describe what happens during the four phases of mitosis. Describe
10.2 The Process of Cell Division Lesson Objectives Describe the role of chromosomes in cell division. Name the main events of the cell cycle. Describe what happens during the four phases of mitosis. Describe
Cell Growth, Division, and Reproduction
 Cell Growth, Division, and Reproduction Human Development: Mitosis and Meiosis Division of the Cell Before a cell grows too large, it divides into two new daughter cells in a process called cell division.
Cell Growth, Division, and Reproduction Human Development: Mitosis and Meiosis Division of the Cell Before a cell grows too large, it divides into two new daughter cells in a process called cell division.
Meiosis produces haploid gametes.
 Section 1: produces haploid gametes. K What I Know W What I Want to Find Out L What I Learned Essential Questions How does the reduction in chromosome number occur during meiosis? What are the stages of
Section 1: produces haploid gametes. K What I Know W What I Want to Find Out L What I Learned Essential Questions How does the reduction in chromosome number occur during meiosis? What are the stages of
Cellular Reproduction = Cell Division. Passes on Genes from Cells to Cells Reproduction of Organisms
 Cellular Reproduction = Cell Division Passes on Genes from Cells to Cells Reproduction of Organisms Genes DNA Chromatin fiber Chromosomes Fig. 9.6 Genes, the segments of DNA, are part of chromatin fiber
Cellular Reproduction = Cell Division Passes on Genes from Cells to Cells Reproduction of Organisms Genes DNA Chromatin fiber Chromosomes Fig. 9.6 Genes, the segments of DNA, are part of chromatin fiber
Cell Division. Genetic info must be copied. Each cell gets a complete copy of that info. It occurs in two main stages:
 10-2 Cell Division Key Questions: 1)What is the role of chromosomes in cell division? 2) What are the main events of the cell cycle? 3) What events occur during each of the four phases of mitosis? 4) How
10-2 Cell Division Key Questions: 1)What is the role of chromosomes in cell division? 2) What are the main events of the cell cycle? 3) What events occur during each of the four phases of mitosis? 4) How
Cell Division. Binary Fission, Mitosis & Meiosis 2/9/2016. Dr. Saud Alamri
 Cell Division Binary Fission, Mitosis & Meiosis 1 Prokaryotic cells reproduce asexually by a type of cell division called binary fission 2 Prokaryotic chromosome Division into two daughter cells Plasma
Cell Division Binary Fission, Mitosis & Meiosis 1 Prokaryotic cells reproduce asexually by a type of cell division called binary fission 2 Prokaryotic chromosome Division into two daughter cells Plasma
The Cell Cycle. Chapter 12
 The Cell Cycle Chapter 12 Why are cells small? As cells get bigger they don t work as well WHY? Difficulties Larger Cells Have: More demands on its DNA Less efficient in moving nutrients/waste across its
The Cell Cycle Chapter 12 Why are cells small? As cells get bigger they don t work as well WHY? Difficulties Larger Cells Have: More demands on its DNA Less efficient in moving nutrients/waste across its
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09414 Supplementary Figure 1 FACS-isolated 8c hepatocytes are highly pure. a, Gating strategy for identifying hepatocyte populations based on DNA content. b, Detection of mchry and mchr9
doi:10.1038/nature09414 Supplementary Figure 1 FACS-isolated 8c hepatocytes are highly pure. a, Gating strategy for identifying hepatocyte populations based on DNA content. b, Detection of mchry and mchr9
Meiosis. Bởi: OpenStaxCollege
 Meiosis Bởi: OpenStaxCollege Sexual reproduction requires fertilization, a union of two cells from two individual organisms. If those two cells each contain one set of chromosomes, then the resulting cell
Meiosis Bởi: OpenStaxCollege Sexual reproduction requires fertilization, a union of two cells from two individual organisms. If those two cells each contain one set of chromosomes, then the resulting cell
10.1 Cell Growth, Division, and Reproduction
 10.1 Cell Growth, Division, and Reproduction Lesson Objectives Explain the problems that growth causes for cells. Compare asexual and sexual reproduction. Lesson Summary Limits to Cell Size There are two
10.1 Cell Growth, Division, and Reproduction Lesson Objectives Explain the problems that growth causes for cells. Compare asexual and sexual reproduction. Lesson Summary Limits to Cell Size There are two
Topic 8 Mitosis & Meiosis Ch.12 & 13. The Eukaryotic Genome. The Eukaryotic Genome. The Eukaryotic Genome
 Topic 8 Mitosis & Meiosis Ch.12 & 13 The Eukaryotic Genome pp. 244-245,268-269 Genome All of the genes in a cell. Eukaryotic cells contain their DNA in long linear pieces. In prokaryotic cells, there is
Topic 8 Mitosis & Meiosis Ch.12 & 13 The Eukaryotic Genome pp. 244-245,268-269 Genome All of the genes in a cell. Eukaryotic cells contain their DNA in long linear pieces. In prokaryotic cells, there is
Interphase When a cell isn t dividing, the nucleus usually just looks grainy, and chromosomes are present but not visible.
 The dance that chromosomes go through when cells divide is known as... Interphase When a cell isn t dividing, the nucleus usually just looks grainy, and chromosomes are present but not visible. Prophase
The dance that chromosomes go through when cells divide is known as... Interphase When a cell isn t dividing, the nucleus usually just looks grainy, and chromosomes are present but not visible. Prophase
Introduction to Botany
 Introduction to Botany Alexey Shipunov Minot State University Lecture 13 Shipunov (MSU) Introduction to Botany Lecture 13 1 / 27 Outline 1 Questions and answers Quiz 2 Plant cell Cell boundaries Protein
Introduction to Botany Alexey Shipunov Minot State University Lecture 13 Shipunov (MSU) Introduction to Botany Lecture 13 1 / 27 Outline 1 Questions and answers Quiz 2 Plant cell Cell boundaries Protein
Bio 10: 10.1 Cell Growth, Division, and Reproduction
 Bio 10: 10.1 Cell Growth, Division, and Reproduction Lesson Objectives Explain the problems that growth causes for cells. Compare asexual and sexual reproduction. Lesson Summary Limits to Cell Size There
Bio 10: 10.1 Cell Growth, Division, and Reproduction Lesson Objectives Explain the problems that growth causes for cells. Compare asexual and sexual reproduction. Lesson Summary Limits to Cell Size There
Chapter 2: Chromosomes and cellular reproduction
 Chapter 2: Chromosomes and cellular reproduction I. Contrast between prokaryotes and eukaryotes. See Figure 2.1 Nucleus absent Small diameter 1 to 10 µm Genome usually 1 circular molecule Small genome;
Chapter 2: Chromosomes and cellular reproduction I. Contrast between prokaryotes and eukaryotes. See Figure 2.1 Nucleus absent Small diameter 1 to 10 µm Genome usually 1 circular molecule Small genome;
Biology Notes 2. Mitosis vs Meiosis
 Biology Notes 2 Mitosis vs Meiosis Diagram Booklet Cell Cycle (bottom corner draw cell in interphase) Mitosis Meiosis l Meiosis ll Cell Cycle Interphase Cell spends the majority of its life in this phase
Biology Notes 2 Mitosis vs Meiosis Diagram Booklet Cell Cycle (bottom corner draw cell in interphase) Mitosis Meiosis l Meiosis ll Cell Cycle Interphase Cell spends the majority of its life in this phase
Describe the process of cell division in prokaryotic cells. The Cell Cycle
 The Cell Cycle Objective # 1 In this topic we will examine the cell cycle, the series of changes that a cell goes through from one division to the next. We will pay particular attention to how the genetic
The Cell Cycle Objective # 1 In this topic we will examine the cell cycle, the series of changes that a cell goes through from one division to the next. We will pay particular attention to how the genetic
Unit 2: Characteristics of Living Things Lesson 25: Mitosis
 Name Unit 2: Characteristics of Living Things Lesson 25: Mitosis Objective: Students will be able to explain the phases of Mitosis. Date Essential Questions: 1. What are the phases of the eukaryotic cell
Name Unit 2: Characteristics of Living Things Lesson 25: Mitosis Objective: Students will be able to explain the phases of Mitosis. Date Essential Questions: 1. What are the phases of the eukaryotic cell
Name: Date: Period: Cell Cycles and DNA Study Guide
 Name: Date: Period: DNA (Deoxyribonucleic Acid) is the chemical inside the nucleus of cells that contains hereditary information. DNA is shaped like a double helix/twisted ladder. The sides of the ladder
Name: Date: Period: DNA (Deoxyribonucleic Acid) is the chemical inside the nucleus of cells that contains hereditary information. DNA is shaped like a double helix/twisted ladder. The sides of the ladder
Cell Growth and Division
 Cell Growth and Division Growth, Development, and Reproduction Q: How does a cell produce a new cell? 10.1 Why do cells divide? WHAT I KNOW SAMPLE ANSWER: Cells divide to produce more cells. WHAT I LEARNED
Cell Growth and Division Growth, Development, and Reproduction Q: How does a cell produce a new cell? 10.1 Why do cells divide? WHAT I KNOW SAMPLE ANSWER: Cells divide to produce more cells. WHAT I LEARNED
Key Concepts. n Cell Cycle. n Interphase. n Mitosis. n Cytokinesis
 The Cell Cycle B-2.6: Summarize the characteristics of the cell cycle: interphase (G 1, S, G 2 ); the phases of mitosis (prophase, metaphase, anaphase, telophase); and plant and animal cytokinesis. Key
The Cell Cycle B-2.6: Summarize the characteristics of the cell cycle: interphase (G 1, S, G 2 ); the phases of mitosis (prophase, metaphase, anaphase, telophase); and plant and animal cytokinesis. Key
Sexual Reproduction. The two parent cells needed for sexual reproduction are called gametes. They are formed during a process known as meiosis.
 Sexual Reproduction Recall that asexual reproduction involves only one parent cell. This parent cell divides to produce two daughter cells that are genetically identical to the parent. Sexual reproduction,
Sexual Reproduction Recall that asexual reproduction involves only one parent cell. This parent cell divides to produce two daughter cells that are genetically identical to the parent. Sexual reproduction,
LAB 8 EUKARYOTIC CELL DIVISION: MITOSIS AND MEIOSIS
 LAB 8 EUKARYOTIC CELL DIVISION: MITOSIS AND MEIOSIS Name: Date: INTRODUCTION BINARY FISSION: Prokaryotic cells (bacteria) reproduce asexually by binary fission. Bacterial cells have a single circular chromosome,
LAB 8 EUKARYOTIC CELL DIVISION: MITOSIS AND MEIOSIS Name: Date: INTRODUCTION BINARY FISSION: Prokaryotic cells (bacteria) reproduce asexually by binary fission. Bacterial cells have a single circular chromosome,
The Cell Cycle and Cell Division
 The Cell Cycle and Cell Division «The cell cycle is a regular pattern of growth, DNA replication, and cell division. The cell cycle has four main stages. «The main stages of the cell cycle are G1 (gap
The Cell Cycle and Cell Division «The cell cycle is a regular pattern of growth, DNA replication, and cell division. The cell cycle has four main stages. «The main stages of the cell cycle are G1 (gap
A diploid somatic cell from a rat has a total of 42 chromosomes (2n = 42). As in humans, sex chromosomes determine sex: XX in females and XY in males.
 Multiple Choice Use the following information for questions 1-3. A diploid somatic cell from a rat has a total of 42 chromosomes (2n = 42). As in humans, sex chromosomes determine sex: XX in females and
Multiple Choice Use the following information for questions 1-3. A diploid somatic cell from a rat has a total of 42 chromosomes (2n = 42). As in humans, sex chromosomes determine sex: XX in females and
Mitosis Verses Meiosis
 Mitosis Verses Meiosis Name LT: I can compare mitosis and meiosis using various resources. Standards: 4.1b, 4.1c Visit the following links: https://www.youtube.com/watch?v=f-ldpgefahi https://www.youtube.com/watch?v=vzdmg7ke69g
Mitosis Verses Meiosis Name LT: I can compare mitosis and meiosis using various resources. Standards: 4.1b, 4.1c Visit the following links: https://www.youtube.com/watch?v=f-ldpgefahi https://www.youtube.com/watch?v=vzdmg7ke69g
Cell Division. OpenStax College. 1 Genomic DNA
 OpenStax-CNX module: m44459 1 Cell Division OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will be
OpenStax-CNX module: m44459 1 Cell Division OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will be
Meiosis * OpenStax. This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0.
 OpenStax-CNX module: m45466 1 Meiosis * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will be able to: Abstract
OpenStax-CNX module: m45466 1 Meiosis * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will be able to: Abstract
Supplementary Figure 1. Phenotype of the HI strain.
 Supplementary Figure 1. Phenotype of the HI strain. (A) Phenotype of the HI and wild type plant after flowering (~1month). Wild type plant is tall with well elongated inflorescence. All four HI plants
Supplementary Figure 1. Phenotype of the HI strain. (A) Phenotype of the HI and wild type plant after flowering (~1month). Wild type plant is tall with well elongated inflorescence. All four HI plants
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
2. is the period of growth and development for a cell. 3. During interphase, most cells go through three stages rapid growth and
 Chapter 5 Lesson 1- General Lesson Outline Directions: Use the words below to fill in the outline of the text from lesson one. If the word is used more than once, it is followed by the number of times
Chapter 5 Lesson 1- General Lesson Outline Directions: Use the words below to fill in the outline of the text from lesson one. If the word is used more than once, it is followed by the number of times
AP Biology Fall Semester Set 1
 1. During which stage does DNA replication occur? A. Prophase B. Metaphase C. Anaphase D. none of these 2. At what phase in the cell cycle does DNA replication occur? A. G1 B. S C. G2 D. M 3. Which of
1. During which stage does DNA replication occur? A. Prophase B. Metaphase C. Anaphase D. none of these 2. At what phase in the cell cycle does DNA replication occur? A. G1 B. S C. G2 D. M 3. Which of
Fertilization of sperm and egg produces offspring
 In sexual reproduction Fertilization of sperm and egg produces offspring In asexual reproduction Offspring are produced by a single parent, without the participation of sperm and egg CONNECTIONS BETWEEN
In sexual reproduction Fertilization of sperm and egg produces offspring In asexual reproduction Offspring are produced by a single parent, without the participation of sperm and egg CONNECTIONS BETWEEN
11-4 Meiosis. Chromosome Number
 11-4 Meiosis Chromosome Number Sexual reproduction shuffles and recombines genes from two parents. During gametogenesis, genes are segregated and assorted (shuffled) into gemetes, and at fertilization,
11-4 Meiosis Chromosome Number Sexual reproduction shuffles and recombines genes from two parents. During gametogenesis, genes are segregated and assorted (shuffled) into gemetes, and at fertilization,
Dr. Mahmood S. Choudhery, PhD, Postdoc (USA) Assistant Professor Tissue Engineering & Regenerative Medicine King Edward Medical University
 CELL DIVISION Dr. Mahmood S. Choudhery, PhD, Postdoc (USA) Assistant Professor Tissue Engineering & Regenerative Medicine King Edward Medical University Cell Division The key roles of cell division Unicellular
CELL DIVISION Dr. Mahmood S. Choudhery, PhD, Postdoc (USA) Assistant Professor Tissue Engineering & Regenerative Medicine King Edward Medical University Cell Division The key roles of cell division Unicellular
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Mitosis and Meiosis Cell growth and division
 Mitosis and Meiosis Cell growth and division The larger the cell, the more trouble the cell has moving nutrients and waste across the cell membrane. 1. DNA/information overload As a cell increases in size,
Mitosis and Meiosis Cell growth and division The larger the cell, the more trouble the cell has moving nutrients and waste across the cell membrane. 1. DNA/information overload As a cell increases in size,
LIST of SUPPLEMENTARY MATERIALS
 LIST of SUPPLEMENTARY MATERIALS Mir et al., Dense Bicoid Hubs Accentuate Binding along the Morphogen Gradient Supplemental Movie S1 (Related to Figure 1). Movies corresponding to the still frames shown
LIST of SUPPLEMENTARY MATERIALS Mir et al., Dense Bicoid Hubs Accentuate Binding along the Morphogen Gradient Supplemental Movie S1 (Related to Figure 1). Movies corresponding to the still frames shown
Chromosome Chr Duplica Duplic t a ion Pixley
 Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
11.4 Meiosis. Vocabulary: Homologous Diploid Haploid Meiosis Crossing-over Tetrad
 11.4 Meiosis Vocabulary: Homologous Diploid Haploid Meiosis Crossing-over Tetrad Key Concept: What happens during the process of meiosis? How is MEIOSIS different than mitosis? Blast from the past What
11.4 Meiosis Vocabulary: Homologous Diploid Haploid Meiosis Crossing-over Tetrad Key Concept: What happens during the process of meiosis? How is MEIOSIS different than mitosis? Blast from the past What
AP Biology - Cell cycle / division
 AP Biology - Cell cycle / division Quiz Directions 1. During which stage does DNA replication occur? A. Prophase B. Metaphase C. Anaphase D. none of these 2. At what phase in the cell cycle does DNA replication
AP Biology - Cell cycle / division Quiz Directions 1. During which stage does DNA replication occur? A. Prophase B. Metaphase C. Anaphase D. none of these 2. At what phase in the cell cycle does DNA replication
2:1 Chromosomes DNA Genes Chromatin Chromosomes CHROMATIN: nuclear material in non-dividing cell, composed of DNA/protein in thin uncoiled strands
 Human Heredity Chapter 2 Chromosomes, Mitosis, and Meiosis 2:1 Chromosomes DNA Genes Chromatin Chromosomes CHROMATIN: nuclear material in non-dividing cell, composed of DNA/protein in thin uncoiled strands
Human Heredity Chapter 2 Chromosomes, Mitosis, and Meiosis 2:1 Chromosomes DNA Genes Chromatin Chromosomes CHROMATIN: nuclear material in non-dividing cell, composed of DNA/protein in thin uncoiled strands
Cell Growth and Reproduction Module B, Anchor 1
 Cell Growth and Reproduction Module B, Anchor 1 Key Concepts: - The larger a cell becomes, the more demands the cell places on its DNA. In addition, a larger cell is less efficient in moving nutrients
Cell Growth and Reproduction Module B, Anchor 1 Key Concepts: - The larger a cell becomes, the more demands the cell places on its DNA. In addition, a larger cell is less efficient in moving nutrients
Name Chapter 10: Chromosomes, Mitosis, and Meiosis Mrs. Laux Take home test #7 DUE: MONDAY, NOVEMBER 16, 2009 MULTIPLE CHOICE QUESTIONS
 MULTIPLE CHOICE QUESTIONS 1. A bacterial chromosome consists of: A. a linear DNA molecule many times larger than the cell. B. a circular DNA molecule many times larger than the cell. C. a circular DNA
MULTIPLE CHOICE QUESTIONS 1. A bacterial chromosome consists of: A. a linear DNA molecule many times larger than the cell. B. a circular DNA molecule many times larger than the cell. C. a circular DNA
Human Biology Chapter 13.4: Meiosis and Genetic Variation
 OpenStax-CNX module: m58013 1 Human Biology Chapter 13.4: Meiosis and Genetic Variation Willy Cushwa Based on Meiosis by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
OpenStax-CNX module: m58013 1 Human Biology Chapter 13.4: Meiosis and Genetic Variation Willy Cushwa Based on Meiosis by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
Anaphase, Telophase. Animal cells divide their cytoplasm by forming? Cleavage furrow. Bacteria, Paramecium, Amoeba, etc. reproduce by...
 The 4 phases of mitosis Animal cells divide their cytoplasm by forming? Bacteria, Paramecium, Amoeba, etc. reproduce by... Cell which after division is identical to the original is called a Prophase, Metaphase,
The 4 phases of mitosis Animal cells divide their cytoplasm by forming? Bacteria, Paramecium, Amoeba, etc. reproduce by... Cell which after division is identical to the original is called a Prophase, Metaphase,
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Meiosis. The form of cell division by which gametes, with half the regular number of chromosomes, are produced.
 MEIOSIS Meiosis The form of cell division by which gametes, with half the regular number of chromosomes, are produced. diploid (2n) haploid (n) (complete set of chromosomes) (half the regular number of
MEIOSIS Meiosis The form of cell division by which gametes, with half the regular number of chromosomes, are produced. diploid (2n) haploid (n) (complete set of chromosomes) (half the regular number of
Purposes of Cell Division
 Purposes of Cell Division Increase the number of cells for growth and repair of worn out tissues What examples in the human body can you think of? Transmit genetic information to later generations Why
Purposes of Cell Division Increase the number of cells for growth and repair of worn out tissues What examples in the human body can you think of? Transmit genetic information to later generations Why
AP Biology. Biology is the only subject in which multiplication is the same thing as division. The Cell Cycle: Cell Growth, Cell Division
 QuickTime and and a TIFF TIFF (Uncompressed) decompressor are are needed needed to to see see this this picture. picture. Biology is the only subject in which multiplication is the same thing as division
QuickTime and and a TIFF TIFF (Uncompressed) decompressor are are needed needed to to see see this this picture. picture. Biology is the only subject in which multiplication is the same thing as division
GENERAL SAFETY: Follow your teacher s directions. Do not work in the laboratory without your teacher s supervision.
 Name: Bio AP Lab: Cell Division B: Mitosis & Meiosis (Modified from AP Biology Investigative Labs) BACKGROUND: One of the characteristics of living things is the ability to replicate and pass on genetic
Name: Bio AP Lab: Cell Division B: Mitosis & Meiosis (Modified from AP Biology Investigative Labs) BACKGROUND: One of the characteristics of living things is the ability to replicate and pass on genetic
Mitosis and Meiosis Cell growth and division
 LIMITS TO CELL GROWTH Mitosis and Meiosis Cell growth and division The larger the cell, the more trouble the cell has moving nutrients and waste across the cell membrane. LIMITS TO CELL GROWTH 1. DNA/information
LIMITS TO CELL GROWTH Mitosis and Meiosis Cell growth and division The larger the cell, the more trouble the cell has moving nutrients and waste across the cell membrane. LIMITS TO CELL GROWTH 1. DNA/information
Cellular Reproduction. MXMS 7th Grade Science
 Cellular Reproduction MXMS 7th Grade Science What is cell division? 2 primary methods allow for cells to divide and reproduce themselves: A. Mitosis: produces identical offspring B. Meiosis: produces genetically
Cellular Reproduction MXMS 7th Grade Science What is cell division? 2 primary methods allow for cells to divide and reproduce themselves: A. Mitosis: produces identical offspring B. Meiosis: produces genetically
Human biology Laboratory. Cell division. Lecturer Maysam A Mezher
 Human biology Laboratory Cell division Lecturer Maysam A Mezher CHROMOSOME STRUCTURE 1. During nuclear division, the DNA (as chromatin) in a Eukaryotic cell's nucleus is coiled into very tight compact
Human biology Laboratory Cell division Lecturer Maysam A Mezher CHROMOSOME STRUCTURE 1. During nuclear division, the DNA (as chromatin) in a Eukaryotic cell's nucleus is coiled into very tight compact
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/9/e1500511/dc1 Supplementary Materials for Contractility parameters of human -cardiac myosin with the hypertrophic cardiomyopathy mutation R403Q show loss of
advances.sciencemag.org/cgi/content/full/1/9/e1500511/dc1 Supplementary Materials for Contractility parameters of human -cardiac myosin with the hypertrophic cardiomyopathy mutation R403Q show loss of
CHAPTER 12 - THE CELL CYCLE (pgs )
 CHAPTER 12 - THE CELL CYCLE (pgs. 228-245) CHAPTER SEVEN TARGETS I. Describe the importance of mitosis in single-celled and multi-cellular organisms. II. Explain the organization of DNA molecules and their
CHAPTER 12 - THE CELL CYCLE (pgs. 228-245) CHAPTER SEVEN TARGETS I. Describe the importance of mitosis in single-celled and multi-cellular organisms. II. Explain the organization of DNA molecules and their
CELL REPRODUCTION. Mitotic M phase Mitosis. Chromosomes divide. Cytokinesis. Cytoplasm and cell membrane divide. Chromosomes as Packaged Genes
 CELL REPRODUCTION Kimberly Lozano Biology 490 Spring 2010 CELL CYCLE Interphase G1: Growth (1) New organelles form within the cell. S: Synthesis Cell duplicates its DNA. G2: Growth (2) Cell prepares for
CELL REPRODUCTION Kimberly Lozano Biology 490 Spring 2010 CELL CYCLE Interphase G1: Growth (1) New organelles form within the cell. S: Synthesis Cell duplicates its DNA. G2: Growth (2) Cell prepares for
Lecture Series 5 Cell Cycle & Cell Division
 Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Division Read Chapter 19 pages 651-663 663 only (Benefits of Sex & Meiosis sections these are in Chapter
Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Division Read Chapter 19 pages 651-663 663 only (Benefits of Sex & Meiosis sections these are in Chapter
Cell Division (Meiosis)
 Cell Division (Meiosis) Meiosis The form of cell division by which gametes, with half the number of chromosomes, are produced. Diploid (2n) haploid (n) Meiosis is sexual reproduction. Two divisions (meiosis
Cell Division (Meiosis) Meiosis The form of cell division by which gametes, with half the number of chromosomes, are produced. Diploid (2n) haploid (n) Meiosis is sexual reproduction. Two divisions (meiosis
Cell Cycle and Mitosis
 Cell Cycle and Mitosis THE CELL CYCLE The cell cycle, or cell-division cycle, is the series of events that take place in a eukaryotic cell between its formation and the moment it replicates itself. These
Cell Cycle and Mitosis THE CELL CYCLE The cell cycle, or cell-division cycle, is the series of events that take place in a eukaryotic cell between its formation and the moment it replicates itself. These
ACCELERATE ITS BIOCHEMICAL PROCESSES WHICH WERE SLOWED DOWN BY MITOSIS. THE LENGTH OF THE G1 PHASE CREATES THE DIFFERENCE BETWEEN FAST DIVIDING
 CHAPTER 1: OVERVIEW OF THE CELL CYCLE THE THREE STAGES OF INTERPHASE: INTERPHASE BEFORE A CELL CAN ENTER CELL DIVISION, IT NEEDS TO PREPARE ITSELF BY REPLICATING ITS GENETIC INFORMATION AND ALL OF THE
CHAPTER 1: OVERVIEW OF THE CELL CYCLE THE THREE STAGES OF INTERPHASE: INTERPHASE BEFORE A CELL CAN ENTER CELL DIVISION, IT NEEDS TO PREPARE ITSELF BY REPLICATING ITS GENETIC INFORMATION AND ALL OF THE
THE CELL CYCLE & MITOSIS. Asexual Reproduction: Production of genetically identical offspring from a single parent.
 THE CELL CYCLE & MITOSIS Asexual Reproduction: Production of genetically identical offspring from a single parent. Sexual Reproduction: The fusion of two separate parent cells that produce offspring with
THE CELL CYCLE & MITOSIS Asexual Reproduction: Production of genetically identical offspring from a single parent. Sexual Reproduction: The fusion of two separate parent cells that produce offspring with
Typical Life Cycle of Algae and Fungi. 5 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
 Module 3B Meiosis and Sexual Life Cycles In this module, we will examine a second type of cell division used by eukaryotic cells called meiosis. In addition, we will see how the 2 types of eukaryotic cell
Module 3B Meiosis and Sexual Life Cycles In this module, we will examine a second type of cell division used by eukaryotic cells called meiosis. In addition, we will see how the 2 types of eukaryotic cell
Chapter 5: Mitosis is the Basis of Asexual Reproduction
 Chapter 5: Mitosis is the Basis of Asexual Reproduction Section 5.1: The Cell Cycle and Mitosis Living things must be able to reproduce. For unicellular organisms, cell reproduction is necessary to maintain
Chapter 5: Mitosis is the Basis of Asexual Reproduction Section 5.1: The Cell Cycle and Mitosis Living things must be able to reproduce. For unicellular organisms, cell reproduction is necessary to maintain
LECTURE 10A: MEIO S S
 LECTURE 10A: MEIOSIS Meiosis Definition INTRODUCTION 1. Meiosis is the production of gametes, which is a reduction division which means a diploid gamete produces haploid gametes - from a full complement
LECTURE 10A: MEIOSIS Meiosis Definition INTRODUCTION 1. Meiosis is the production of gametes, which is a reduction division which means a diploid gamete produces haploid gametes - from a full complement
Reading Assignments. A. Systems of Cell Division. Lecture Series 5 Cell Cycle & Cell Division
 Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Death Read Chapter 19 Cell Division Read Chapter 20 pages 659-672 672 only (Benefits of Sex & Meiosis sections)
Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Death Read Chapter 19 Cell Division Read Chapter 20 pages 659-672 672 only (Benefits of Sex & Meiosis sections)
Lecture Series 5 Cell Cycle & Cell Division
 Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Death Read Chapter 19 Cell Division Read Chapter 20 pages 659-672 672 only (Benefits of Sex & Meiosis sections)
Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Death Read Chapter 19 Cell Division Read Chapter 20 pages 659-672 672 only (Benefits of Sex & Meiosis sections)
Bio 102 Practice Problems Cell Cycle and Cell Division
 Bio 102 Practice Problems Cell Cycle and Cell Division Multiple choice: Unless otherwise directed, circle the one best answer: 1. Which one of the following events does NOT occur during prophase of mitosis?
Bio 102 Practice Problems Cell Cycle and Cell Division Multiple choice: Unless otherwise directed, circle the one best answer: 1. Which one of the following events does NOT occur during prophase of mitosis?
4) Please cite Dagda et al J Biol Chem 284: , for any publications or presentations resulting from use or modification of the macro.
 Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Meiosis. Two distinct divisions, called meiosis I and meiosis II
 Meiosis A process in which the number of chromosomes per cell is cut in half through the separation of homologous chromosomes to form gametes, or sex cells Two distinct divisions, called meiosis I and
Meiosis A process in which the number of chromosomes per cell is cut in half through the separation of homologous chromosomes to form gametes, or sex cells Two distinct divisions, called meiosis I and
Honors Biology-CW/HW Cell Biology 2018
 Class: Date: Honors Biology-CW/HW Cell Biology 2018 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Hooke s discovery of cells was made observing a. living
Class: Date: Honors Biology-CW/HW Cell Biology 2018 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Hooke s discovery of cells was made observing a. living
Supplemental Materials Molecular Biology of the Cell
 Supplemental Materials Molecular iology of the Cell Figure S1 Krüger et al. Arabidopsis Plasmodium H. sapiens* 1) Xenopus* 1) Drosophila C.elegans S.cerevisae S.pombe L.major T.cruzi T.brucei DCP5 CITH
Supplemental Materials Molecular iology of the Cell Figure S1 Krüger et al. Arabidopsis Plasmodium H. sapiens* 1) Xenopus* 1) Drosophila C.elegans S.cerevisae S.pombe L.major T.cruzi T.brucei DCP5 CITH
WarmUp 1. C. a phosphate group is removed
 WarmUp 1 1. Energy is released from ATP when C. a phosphate group is removed 2. During normal mitotic cell division, a parent cell with four chromosomes will produce two daughter cells, each containing-
WarmUp 1 1. Energy is released from ATP when C. a phosphate group is removed 2. During normal mitotic cell division, a parent cell with four chromosomes will produce two daughter cells, each containing-
THE PROCESS OF LIVING THINGS CREATING OFFSPRING.
 REPRODUCTION 1 THE PROCESS OF LIVING THINGS CREATING OFFSPRING. Offspring are the next generation. It happens on multiple levels for multicellular organisms 2 SPECIES SURVIVAL Think back to 7th grade Organisms
REPRODUCTION 1 THE PROCESS OF LIVING THINGS CREATING OFFSPRING. Offspring are the next generation. It happens on multiple levels for multicellular organisms 2 SPECIES SURVIVAL Think back to 7th grade Organisms
Biology. Chapter 10 Cell Reproduction. I. Chromosomes
 Biology Chapter 10 Cell Reproduction I. Chromosomes Long thin molecules that store genetic information. A. Chromosome Structure 1. Rod shaped structure composed of DNA and protein. 2. DNA is wrapped around
Biology Chapter 10 Cell Reproduction I. Chromosomes Long thin molecules that store genetic information. A. Chromosome Structure 1. Rod shaped structure composed of DNA and protein. 2. DNA is wrapped around
Lesson Overview Meiosis
 11.4 THINK ABOUT IT As geneticists in the early 1900s applied Mendel s laws, they wondered where genes might be located. They expected genes to be carried on structures inside the cell, but which structures?
11.4 THINK ABOUT IT As geneticists in the early 1900s applied Mendel s laws, they wondered where genes might be located. They expected genes to be carried on structures inside the cell, but which structures?
11-4 Meiosis Meiosis. Slide 1 of 35. Copyright Pearson Prentice Hall
 11-4 Meiosis 1 of 35 Each organism must inherit a single copy of every gene from each of its parents. Gametes are formed by a process that separates the two sets of genes so that each gamete ends up with
11-4 Meiosis 1 of 35 Each organism must inherit a single copy of every gene from each of its parents. Gametes are formed by a process that separates the two sets of genes so that each gamete ends up with
3.2.2 All cells arise from other cells
 alevelbiology.co.uk SPECIFICATION Within multicellular organisms, not all cells retain the ability to divide. Eukaryotic cells that do retain the ability to divide show a cell cycle. DNA replication occurs
alevelbiology.co.uk SPECIFICATION Within multicellular organisms, not all cells retain the ability to divide. Eukaryotic cells that do retain the ability to divide show a cell cycle. DNA replication occurs
Cell Size. Cell Growth and Reproduction 12/3/14
 Cell Growth and Reproduction Cell Size Why are cells so small? Cells do not contain a circulatory system Cells receive nutrients and remove waste through diffusion Diffusion- movement of molecules from
Cell Growth and Reproduction Cell Size Why are cells so small? Cells do not contain a circulatory system Cells receive nutrients and remove waste through diffusion Diffusion- movement of molecules from
Topic 3: Genetics (Student) Essential Idea: Chromosomes carry genes in a linear sequence that is shared by members of a species.
 Topic 3: Genetics (Student) 3.2 Essential Idea: Chromosomes carry genes in a linear sequence that is shared by members of a species. 3.2 Chromosomes 3.2.U1 Prokaryotes have one chromosome consisting of
Topic 3: Genetics (Student) 3.2 Essential Idea: Chromosomes carry genes in a linear sequence that is shared by members of a species. 3.2 Chromosomes 3.2.U1 Prokaryotes have one chromosome consisting of
2 The Cell Cycle. TAKE A LOOK 2. Complete Prokaryotic cells divide by.
 CHAPTER 5 2 The Cell Cycle SECTION The Cell in Action BEFORE YOU READ After you read this section, you should be able to answer these questions: How are new cells made? What is mitosis? What happens when
CHAPTER 5 2 The Cell Cycle SECTION The Cell in Action BEFORE YOU READ After you read this section, you should be able to answer these questions: How are new cells made? What is mitosis? What happens when
MEIOSIS LAB INTRODUCTION PART I: MEIOSIS
 MEIOSIS LAB INTRODUCTION Meiosis involves two successive nuclear divisions that produce four haploid cells. Meiosis I is the reduction division. It is this first division that reduces the chromosome number
MEIOSIS LAB INTRODUCTION Meiosis involves two successive nuclear divisions that produce four haploid cells. Meiosis I is the reduction division. It is this first division that reduces the chromosome number
Study Guide A. Answer Key. Cell Growth and Division. SECTION 1. THE CELL CYCLE 1. a; d; b; c 2. gaps 3. c and d 4. c 5. b and d 6.
 Cell Growth and Division Answer Key SECTION 1. THE CELL CYCLE 1. a; d; b; c 2. gaps 3. c and d 4. c 5. b and d 6. G 1 7. G 0 8. c 9. faster; too large 10. volume 11. a and b 12. repeating pattern or repetition
Cell Growth and Division Answer Key SECTION 1. THE CELL CYCLE 1. a; d; b; c 2. gaps 3. c and d 4. c 5. b and d 6. G 1 7. G 0 8. c 9. faster; too large 10. volume 11. a and b 12. repeating pattern or repetition
Sexual Reproduction and Genetics
 10 Sexual Reproduction and Genetics section 1 Meiosis Before You Read Think about the traits that make people unique. Some people are tall, while others are short. People can have brown, blue, or green
10 Sexual Reproduction and Genetics section 1 Meiosis Before You Read Think about the traits that make people unique. Some people are tall, while others are short. People can have brown, blue, or green
BIOLOGY 111. CHAPTER 5: Chromosomes and Inheritance
 BIOLOGY 111 CHAPTER 5: Chromosomes and Inheritance Chromosomes and Inheritance Learning Outcomes 5.1 Differentiate between sexual and asexual reproduction in terms of the genetic variation of the offspring.
BIOLOGY 111 CHAPTER 5: Chromosomes and Inheritance Chromosomes and Inheritance Learning Outcomes 5.1 Differentiate between sexual and asexual reproduction in terms of the genetic variation of the offspring.
5.1. Cells have distinct phases of growth, reproduction, and normal functions. G 1. Cell Growth and Division CHAPTER 5 THE CELL CYCLE KEY CONCEPT
 SECTION 5.1 THE CELL CYCLE Study Guide KEY CONCEPT Cells have distinct phases of growth, reproduction, and normal functions. VOCABULARY cell cycle mitosis cytokinesis The cell cycle has four main stages.
SECTION 5.1 THE CELL CYCLE Study Guide KEY CONCEPT Cells have distinct phases of growth, reproduction, and normal functions. VOCABULARY cell cycle mitosis cytokinesis The cell cycle has four main stages.
Syllabus -- Objectives
 Cell Continuity Syllabus -- Objectives (OL) Explain the terms: cell continuity & chromosomes. Define the terms: haploid & diploid number. Describe the cell activities in he state of non-division: Interphase
Cell Continuity Syllabus -- Objectives (OL) Explain the terms: cell continuity & chromosomes. Define the terms: haploid & diploid number. Describe the cell activities in he state of non-division: Interphase
The division of a unicellular organism reproduces an entire organism, increasing the population. Here s one amoeba dividing into 2.
 1. Cell division functions in 3 things : reproduction, growth, and repair The division of a unicellular organism reproduces an entire organism, increasing the population. Here s one amoeba dividing into
1. Cell division functions in 3 things : reproduction, growth, and repair The division of a unicellular organism reproduces an entire organism, increasing the population. Here s one amoeba dividing into
Mitosis and Genetics Study Guide Answer Key
 Mitosis and Genetics Study Guide Answer Key 1. Which of the following is true of Interphase? a. It is part of Meiosis b. It occurs before Meiosis c. The cell does normal cell activities during interphase
Mitosis and Genetics Study Guide Answer Key 1. Which of the following is true of Interphase? a. It is part of Meiosis b. It occurs before Meiosis c. The cell does normal cell activities during interphase
Mitosis and Meiosis for AP Biology
 Mitosis and Meiosis for AP Biology by Mark Anestis Practice problems for these concepts can be found at : Cell Division Review Questions for AP Biology Mitosis During mitosis, the fourth stage of the cell
Mitosis and Meiosis for AP Biology by Mark Anestis Practice problems for these concepts can be found at : Cell Division Review Questions for AP Biology Mitosis During mitosis, the fourth stage of the cell
The Cell Cycle & Cell Division
 The Cell Cycle & Cell Division http://www.nobel.se/medicine/laureates/2001/press.html The Cell Cycle Animated Cycle http://www.cellsalive.com/cell_cycle.htm MITOSIS Mitosis The process of cell division
The Cell Cycle & Cell Division http://www.nobel.se/medicine/laureates/2001/press.html The Cell Cycle Animated Cycle http://www.cellsalive.com/cell_cycle.htm MITOSIS Mitosis The process of cell division
Cell Division. Mitosis
 Cell division consists of two phases, nuclear division followed by cytokinesis. Nuclear division divides the genetic material in the nucleus, while cytokinesis divides the cytoplasm. There are two kinds
Cell division consists of two phases, nuclear division followed by cytokinesis. Nuclear division divides the genetic material in the nucleus, while cytokinesis divides the cytoplasm. There are two kinds
9-4 Meiosis Meiosis. Slide 1 of 35
 9-4 Meiosis 11-4 Meiosis 1 of 35 11-4 Meiosis Each organism must inherit a single copy of every gene from each of its parents. Gametes are formed by a process that separates the two sets of genes so that
9-4 Meiosis 11-4 Meiosis 1 of 35 11-4 Meiosis Each organism must inherit a single copy of every gene from each of its parents. Gametes are formed by a process that separates the two sets of genes so that
How did that single cell develop into a body with more than a trillion cells?
 CELL DIVISION Cell division is a process which leads to cell multiplication. It occurs in both plants and animals. Original cells which undergo division are known as parent cells and the new on ones resulting
CELL DIVISION Cell division is a process which leads to cell multiplication. It occurs in both plants and animals. Original cells which undergo division are known as parent cells and the new on ones resulting
Chromosome Numbers of Some Common Organisms
 Meiosis Introduction Process of reduction division Purpose: Produces gametes (sex ) sperm & egg Meiosis is NOT a cycle like mitosis. 1 2 3 Diploid vs. Haploid Diploid a cell that contains homologous chromosomes
Meiosis Introduction Process of reduction division Purpose: Produces gametes (sex ) sperm & egg Meiosis is NOT a cycle like mitosis. 1 2 3 Diploid vs. Haploid Diploid a cell that contains homologous chromosomes
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
