Table S1. Sequence of primers used in RT-qPCR
|
|
|
- Aldous Henry
- 5 years ago
- Views:
Transcription
1 Table S1. Sequence of primers used in RT-qPCR Primer Name P16Ink4a-F P16Ink4a-R P15Ink4b-F P15Ink4b-R P19Arf-F P19Arf-R P53-F P53-R P21cip1-F P21cip1-R P27kip1-F P27kip1-R P18Ink4c-F P18Ink4c-R P19Ink4d-F P19Ink4d-R P57Kip2-F P57kip2-R Kdm2b-F Kdm2b-R GAPDH-F GAPDH-R Primer Sequence G A AC T C T T T C G G T C G T A C C C C G A A T C T G C A C C G T A G T T G A C C A C C C T T A C C A G A C C T G T G A G G C G T C A C A C A C A T C C A G G C T C T G G C T T T C G T G A A C A T G C A G A A G A G C T G C T A C G T G A T G G A A G A C T C C A G T G G G A A T C T T C T G T A C G G C G G T C T C T T T G C A C T C T G G T G T C T G A G C T G C G C T T G G A G T G A T A G A A A A A C T A A C C C G G G A C T T G G A G C C A G G G G C T T A T G A T T C T G A T G G A A C T G G T T T T G C T G T C A G G G C A G G T T C C C T T C A T T A T G C A G G T C A T G A T G T T T G G A A T A G T A C C G G A G G C A T C T T G G G G A G C A G G A C G A G A A T C A A G G A A G A A G T C G T T C G C A T T G G T A A A G G A T G C C C A G A T G A G G A G G C G C A G C T C T A C A A T G T T G C A G T G G C A A A G T G G A G A T T G A A T T T G C C G T G A G T G G A G T Table S2. Sequence of primers used in p15 Ink4b ChIP assay Primer Name Ink4bAmp1-F Ink4bAmp1-R Ink4bAmp2-F Ink4bAmp2-R Ink4bAmp3-F Ink4bAmp3-R Primer Sequence C C G C C T A G A G A T G A A C T A G C C C G C T T T T G C A A T T G A C T G A C C A C C G A A G C T A C T G G G T C T C C T G T G G C A G A A A T G G T C C T T A T G T T C T A A G A G G C T T T G T T T C C A C A T T T G T G C A T A G G A G A T C A G G
2 A Relative Jhdm1b mrna Level Jhdm1b OE Figure S1. Over-expression of Kdm2b/Jhdm1b in hematopoietic progenitors (A) Efficient transduction of hematopoietic progenitor cells by lentiviral vectors.shown are the photographs of bright (left panel) and flurorescent (right panel) fields of hematopoietic progenitor cells at 48 hours after lenti- transduction. () Kdm2b/Jhdm1b-transduced hematopoietic progenitor cells express much higher levels of Kdm2b/Jhdm1b compared to control. Relative mrna levels are measured by RT-qPCR and normalized to the Gapdh. The mrna level of control cells is set as 1.
3 Relative Expression Level KD-HM A CKD-HMG J1bKD-HMG Control shrna Jhdm1b shrna Hoxa9 Meis1 2A 2A Hoxa9 Meis1 2A 2A G G FP C G J1bKD-HM FSC 1.2 Nontransduced CDK-HMG J1bKD-HMG.4.2 Figure S2. Genetic modifications of hematopoietic progenitor cells (A) Schematic representation of lentiviral vectors used in the experiments. Shown are diagrams of polycistronic lentiviral vectors that express alone, or Hoxa9, Meis1 and coupled with control shrna (CKD-HMG) or shrna targeting Kdm2b/Jhdm1b (J1bKD-HMG). Transgenes are linked by P2A sequence of foot-to-mouth disease virus. shrna driven by the U6 promoter is used for Kdm2b/Jhdm1b knocking down (Left panel). Transduced + cells are sorted by FACS and used for functional analysis (right panel). () RT-qPCR analysis demonstrating that Kdm2b/Jhdm1b is efficiently knocked down after transducing with a lentiviral vector expressing Kdm1b/ (J1bKD-HMG). Relative mrna levels are normalized to the Gapdh. The mrna level of Kdm2b/Jhdm2b in non-trasduced cells is set as 1.
4 c-kit c-kit A 2X 4X IgG IgG Gr-1 Mac-1 Figure S3. Characterization of Hoxa9/Meis1 induced murine AML (A) May-Grunwald/Giemsa staining showing typical leukoblasts in the bone marrow of recipient mice transplanted with CKD-HMG cells (left, 2x magnification, ar =2μm; right, 4x magnification, ar = 1μm). () Flow cytometry analysis demonstrates that Hoxa9/Meis1 induced leukemic cells expressing both c-kit and myeloid lineage markers Gr-1/Mac-1.
5 A Control KD Control shrna +LacZ LacZ-Flag +wt Jhdm1b-F wt Jhdm1b-Flag +mut Jhdm1b-F Mut Jhdm1b-Flag Relative Jhdm1b mrna Level Control KD + LacZ +wt Jhdm1b-F +mut Jhdm1b-F C Jhdm1b-F -2a-Lacz-F LacZ-F * * Non-specific Figure S4. Genetic modifications of leukemia stem cells (A) Schematic representation of lentiviral vectors used for genetic modification of leukemia stem cells isolated from primary AML mouse. The polycistronic lentiviral vectors express LacZ-Flag, shrna-resistant wild type and mutant Kdm2b/Jhdm1b- Flag and a drug selection marker (). Transgenes are linked by P2A sequence of foot-to-mouth disease virus. shrna driven by U6 promoter is used for Kdm2b/Jhdm1b knockdown. () RT-qPCR analysis shows the relative Kdm2b/Jhdm1b levels in the cells with control knockdown, Kdm2b/Jhdm1b knockdown, Kdm2b/Jhdm1b knockdown rescued with lacz, wild-type or mutant Kdm2b/Jhdm1b. Relative mrna levels are normalized to the Gapdh. The mrna level of Kdm2b/Jhdm2b in control knockdown cells is set as 1. (C) Western blot analysis using anti-flag antibody demonstrates the expression of transgenes. Lane 1: Mock control; Lane 2: Kdm2b/Jdhm1bKD+LacZ-F, * denotes LacZ-Flag, denotes uncleaved -2a-LacZ-Flag fusion protein; * Lane 3: Kdm2b/Jdhm1bKD+wild-type Kdm2b/Jdhm1b-F; Lane 4: Kdm2b/Jdhm1bKD +wild-type Kdm2b/Jdhm1b-F.
6 % of Input % of Input A H3K36me2 H3 CKD J1bKD p15 Expression Level 1.2 Mock (LacZ) Jhdm1b OE C 5bp p15 Ink4b anti-flag IP LacZ-Flag Jhdm1b-Flag anti-h3k36me2 IP LacZ-Flag Jhdm1b-Flag Amplicon Amplicon D Mock Jhdm1b OE Hoxa9/MeisI MeCpG 3.3% 2.9% 2.1% Figure S5. Epigenetic analysis of p15ink4b locus in Kdm2b/Jhdm1b modified cells (A) Western blot analysis using anti-h3k36me2 and anti-h3 antibodies demonstrates the global H3K36me2 expression level upon Kdm2b/Jhdm1b knockdown. () RT-qPCR analysis shows the relative p15ink4b levels in the mock and Kdm2b/Jhdm1b over-expressing cultured for 2 weeks. Relative mrna levels are normalized to the Gapdh. The mrna level of mock HPCs is set as 1.
7 (C) ChIP experiments using chromatin prepared from HPCs over-expressing LacZ-Flag and Kdm2b/Jhdm1b-Flag were carried out using anti-flag and anti-h3k36me2 antibodies. Kdm2b/Jhdm1b-Flag binding and H3K36me2 level was assayed by qpcr at the three genomic regions depicted in upper panel. (D) isulfite analysis demonstrates the DNA methylation levels in controlled HPCs, Kdm2b/Jhdm1b immortalized HPCs and Hoxa9/MeisI transformed cells. Empty circle represents unmethylated CpG dinucleotides and black circle represents methylated CpG dinucleotides. The percentage of methylated CpG dinucleotides = total methylated CpG dinucleotides/ total CpG dinucleotides x 1%.
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice Supplementary Figure 2.
 Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
FSC-W FSC-H CD4 CD62-L
 Supplementary Fig. 1 a SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H CD4 CD62-L b SSC-A FSC-A FSC-W FSC-A FSC-A 7-AAD FSC-A CD4 IL-9 CD4 c SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H 7-AAD KI67 Annexin-V 7-AAD d I L -5
Supplementary Fig. 1 a SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H CD4 CD62-L b SSC-A FSC-A FSC-W FSC-A FSC-A 7-AAD FSC-A CD4 IL-9 CD4 c SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H 7-AAD KI67 Annexin-V 7-AAD d I L -5
Supplementary Figure 1. Nature Genetics: doi: /ng.3848
 Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
ydci GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC
 Table S1. DNA primers used in this study. Name ydci P1ydcIkd3 Sequence GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC Kd3ydcIp2 lacz fusion YdcIendP1 YdcItrgP2 GAC AGC
Table S1. DNA primers used in this study. Name ydci P1ydcIkd3 Sequence GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC Kd3ydcIp2 lacz fusion YdcIendP1 YdcItrgP2 GAC AGC
TNFα 18hr. Control. CHX 18hr. TNFα+ CHX 18hr. TNFα: 18 18hr (KDa) PARP. Cleaved. Cleaved. Cleaved. Caspase3. Pellino3 shrna. Control shrna.
 Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
JMJ14-HA. Col. Col. jmj14-1. jmj14-1 JMJ14ΔFYR-HA. Methylene Blue. Methylene Blue
 Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11419 Supplementary Figure 1 Schematic representation of innate immune signaling pathways induced by intracellular Salmonella in cultured macrophages. a, During the infection Salmonella
doi:10.1038/nature11419 Supplementary Figure 1 Schematic representation of innate immune signaling pathways induced by intracellular Salmonella in cultured macrophages. a, During the infection Salmonella
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
downstream (0.8 kb) homologous sequences to the genomic locus of DIC. A DIC mutant strain (ro- 6
 A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. NAD + -related genes expression in mouse tissues and in cells overexpressing NRK1. mrna (a) and protein (b) levels of NRK1, NRK2 and other enzymes involved
Supplementary Figure 1 Supplementary Figure 1. NAD + -related genes expression in mouse tissues and in cells overexpressing NRK1. mrna (a) and protein (b) levels of NRK1, NRK2 and other enzymes involved
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
SUPPLEMENTARY INFORMATION
 reverse 3175 3175 F L C 318 318 3185 3185 319 319 3195 3195 315 8 1 315 3155 315 317 Supplementary Figure 3. Stability of expression of the GFP sensor constructs return to warm conditions. Semi-quantitative
reverse 3175 3175 F L C 318 318 3185 3185 319 319 3195 3195 315 8 1 315 3155 315 317 Supplementary Figure 3. Stability of expression of the GFP sensor constructs return to warm conditions. Semi-quantitative
Gene Expression. Molecular Genetics, March, 2018
 Gene Expression Molecular Genetics, March, 2018 Gene Expression Control of Protein Levels Bacteria Lac Operon Promoter mrna Inducer CAP Control Trp Operon RepressorOperator Control Attenuation Riboswitches
Gene Expression Molecular Genetics, March, 2018 Gene Expression Control of Protein Levels Bacteria Lac Operon Promoter mrna Inducer CAP Control Trp Operon RepressorOperator Control Attenuation Riboswitches
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Generation of paraxial mesoderm from the H7 hesc line.
 Supplementary Figure 1 Generation of paraxial mesoderm from the H7 hesc line. H7 hescs were differentiated as shown in Figure 1a. (a) Flow cytometric analyses of the proportion of CD56+, PDGFRα+, and KDR+
Supplementary Figure 1 Generation of paraxial mesoderm from the H7 hesc line. H7 hescs were differentiated as shown in Figure 1a. (a) Flow cytometric analyses of the proportion of CD56+, PDGFRα+, and KDR+
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures
 Nature Methods Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures Yannick Doyon, Thuy D Vo, Matthew C Mendel, Shon G Greenberg, Jianbin Wang, Danny F Xia, Jeffrey
Nature Methods Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures Yannick Doyon, Thuy D Vo, Matthew C Mendel, Shon G Greenberg, Jianbin Wang, Danny F Xia, Jeffrey
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1
 Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Supplemental Data. Chen and Thelen (2010). Plant Cell /tpc
 Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Origin of mesenchymal stem cell
 Origin of mesenchymal stem cell Takumi Era Department of Cell Modulation, Institute of Molecular embryology and Genetics (IMEG) Kumamoto University Differentiation pathways of in vitro ES cell culture
Origin of mesenchymal stem cell Takumi Era Department of Cell Modulation, Institute of Molecular embryology and Genetics (IMEG) Kumamoto University Differentiation pathways of in vitro ES cell culture
Supplemental Figure 1: Increased Fe deficiency gene expression in roots of nas4x-2
 Supplemental Figure 1: Increased Fe deficiency gene expression in roots of nas4x-2 IRT1, FRO2 and FIT expression levels in roots of the wild-type, nas4x- 1 and nas4x-2, showing that in both nas mutants
Supplemental Figure 1: Increased Fe deficiency gene expression in roots of nas4x-2 IRT1, FRO2 and FIT expression levels in roots of the wild-type, nas4x- 1 and nas4x-2, showing that in both nas mutants
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09414 Supplementary Figure 1 FACS-isolated 8c hepatocytes are highly pure. a, Gating strategy for identifying hepatocyte populations based on DNA content. b, Detection of mchry and mchr9
doi:10.1038/nature09414 Supplementary Figure 1 FACS-isolated 8c hepatocytes are highly pure. a, Gating strategy for identifying hepatocyte populations based on DNA content. b, Detection of mchry and mchr9
Supplemental Information
 Supplemental Information Combinatorial Readout of Unmodified H3R2 and Acetylated H3K14 by the Tandem PHD Finger of MOZ Reveals a Regulatory Mechanism for HOXA9 Transcription Yu Qiu 1, Lei Liu 1, Chen Zhao
Supplemental Information Combinatorial Readout of Unmodified H3R2 and Acetylated H3K14 by the Tandem PHD Finger of MOZ Reveals a Regulatory Mechanism for HOXA9 Transcription Yu Qiu 1, Lei Liu 1, Chen Zhao
Waithe et al Supplementary Figures
 Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
RANK. Alternative names. Discovery. Structure. William J. Boyle* SUMMARY BACKGROUND
 RANK William J. Boyle* Department of Cell Biology, Amgen, Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA * corresponding author tel: 805-447-4304, fax: 805-447-1982, e-mail: bboyle@amgen.com
RANK William J. Boyle* Department of Cell Biology, Amgen, Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA * corresponding author tel: 805-447-4304, fax: 805-447-1982, e-mail: bboyle@amgen.com
Actinobacteria Relative abundance (%) Co-housed CD300f WT. CD300f KO. Colon length (cm) Day 9. Microscopic inflammation score
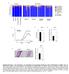 y groups y individuals 9 Actinobacteria Relative abundance (%) acteroidetes Cyanobacteria Deferribacteres Firmicutes Proteobacteria TM Tenericutes Unclassified CDf CDf Co-housed CDf Co-housed CDf CDf CDf
y groups y individuals 9 Actinobacteria Relative abundance (%) acteroidetes Cyanobacteria Deferribacteres Firmicutes Proteobacteria TM Tenericutes Unclassified CDf CDf Co-housed CDf Co-housed CDf CDf CDf
Supplemental Information
 Supplemental Information Figure S. Regions recognized by the specific antibodies. The amino acid sequence of the major p3-alc species, p3-alc α 35, p3-alc β 37 and p3-alc γ 3, are indicated as red letters.
Supplemental Information Figure S. Regions recognized by the specific antibodies. The amino acid sequence of the major p3-alc species, p3-alc α 35, p3-alc β 37 and p3-alc γ 3, are indicated as red letters.
Arabidopsis COMPASS-Like Complexes Mediate Histone H3 Lysine-4 Trimethylation to Control Floral Transition and Plant Development
 Arabidopsis COMPASS-Like Complexes Mediate Histone H3 Lysine-4 Trimethylation to Control Floral Transition and Plant Development Danhua Jiang 1,2, Nicholas C. Kong 1,2, Xiaofeng Gu 2, Zicong Li 1, Yuehui
Arabidopsis COMPASS-Like Complexes Mediate Histone H3 Lysine-4 Trimethylation to Control Floral Transition and Plant Development Danhua Jiang 1,2, Nicholas C. Kong 1,2, Xiaofeng Gu 2, Zicong Li 1, Yuehui
23-. Shoot and root development depend on ratio of IAA/CK
 Balance of Hormones regulate growth and development Environmental factors regulate hormone levels light- e.g. phototropism gravity- e.g. gravitropism temperature Mode of action of each hormone 1. Signal
Balance of Hormones regulate growth and development Environmental factors regulate hormone levels light- e.g. phototropism gravity- e.g. gravitropism temperature Mode of action of each hormone 1. Signal
4) Please cite Dagda et al J Biol Chem 284: , for any publications or presentations resulting from use or modification of the macro.
 Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Regulation of gene Expression in Prokaryotes & Eukaryotes
 Regulation of gene Expression in Prokaryotes & Eukaryotes 1 The trp Operon Contains 5 genes coding for proteins (enzymes) required for the synthesis of the amino acid tryptophan. Also contains a promoter
Regulation of gene Expression in Prokaryotes & Eukaryotes 1 The trp Operon Contains 5 genes coding for proteins (enzymes) required for the synthesis of the amino acid tryptophan. Also contains a promoter
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Supplemental material
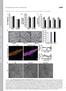 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Nature Medicine: doi: /nm.3776
 C terminal Hsp90 inhibitors restore glucocorticoid sensitivity and relieve a mouse allograft model of Cushing s disease Mathias Riebold, Christian Kozany, Lee Freiburger, Michael Sattler, Michael Buchfelder,
C terminal Hsp90 inhibitors restore glucocorticoid sensitivity and relieve a mouse allograft model of Cushing s disease Mathias Riebold, Christian Kozany, Lee Freiburger, Michael Sattler, Michael Buchfelder,
0! control! 4-OHCY! control! 4-OHCY!
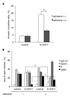 A 25 Annexin-V-positive cells (%) 20 15 10 5 * adhesion ( ) adhesion (+) 0 control 4-OHCY B 80 size of each fraction (%) 60 40 20 * * sub-g1 G0/G1 S G2/M 0 control 4-OHCY control 4-OHCY adhesion + Supplemental
A 25 Annexin-V-positive cells (%) 20 15 10 5 * adhesion ( ) adhesion (+) 0 control 4-OHCY B 80 size of each fraction (%) 60 40 20 * * sub-g1 G0/G1 S G2/M 0 control 4-OHCY control 4-OHCY adhesion + Supplemental
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2017-2018 GENETICS BIO-5009A Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2017-2018 GENETICS BIO-5009A Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section
Supplemental material
 Supplemental material Table 1- Segregation analysis of sgt1a sgt1b double mutant plants Parental genotype No. of lines Normal seeds Phenotype Abnormal seeds Ratio of abnormal seeds (%) WT 3 171 2 1.16
Supplemental material Table 1- Segregation analysis of sgt1a sgt1b double mutant plants Parental genotype No. of lines Normal seeds Phenotype Abnormal seeds Ratio of abnormal seeds (%) WT 3 171 2 1.16
Name KEY. Biology Developmental Biology Winter Quarter Midterm 1
 Name KEY 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2008 Midterm 1 25 questions - 4 pts each 1 extra-credit question - 4 points Provide answers using full sentences,
Name KEY 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2008 Midterm 1 25 questions - 4 pts each 1 extra-credit question - 4 points Provide answers using full sentences,
Biological Roles of Cytokinins
 Direct Control of Shoot Meristem Activity by a Cytokinin-Activating Enzyme By Kurakawa et. Al. Published in Nature Presented by Boyana Grigorova Biological Roles of Cytokinins Cytokinins are positive regulators
Direct Control of Shoot Meristem Activity by a Cytokinin-Activating Enzyme By Kurakawa et. Al. Published in Nature Presented by Boyana Grigorova Biological Roles of Cytokinins Cytokinins are positive regulators
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Genome 541! Unit 4, lecture 2! Transcription factor binding using functional genomics
 Genome 541 Unit 4, lecture 2 Transcription factor binding using functional genomics Slides vs chalk talk: I m not sure why you chose a chalk talk over ppt. I prefer the latter no issues with readability
Genome 541 Unit 4, lecture 2 Transcription factor binding using functional genomics Slides vs chalk talk: I m not sure why you chose a chalk talk over ppt. I prefer the latter no issues with readability
Supplementary Figure 1. Confirmation of REF52-hE2F1p::4NLS-d4Venus reporter cells and characterization of E2F dynamics. (a) Alignment of E2F dynamics
 Supplementary Figure 1. Confirmation of REF52-hE2F1p::4NLS-d4Venus reporter cells and characterization of E2F dynamics. (a) Alignment of E2F dynamics trajectories to endogenous E2F1 mrna expression. Gray
Supplementary Figure 1. Confirmation of REF52-hE2F1p::4NLS-d4Venus reporter cells and characterization of E2F dynamics. (a) Alignment of E2F dynamics trajectories to endogenous E2F1 mrna expression. Gray
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL Figure S1. Mitochondrial morphology in Fis1-null, Mff-null and Fis1/Mff-null MEF cells. (A) Western blotting of lysates from Fis1-null, Mff-null and Fis1/Mff-null cells. Lysates were
SUPPLEMENTAL MATERIAL Figure S1. Mitochondrial morphology in Fis1-null, Mff-null and Fis1/Mff-null MEF cells. (A) Western blotting of lysates from Fis1-null, Mff-null and Fis1/Mff-null cells. Lysates were
Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome
 Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome Marcos S. Simões-Costa 1., Sonja J. McKeown 1., Joanne Tan-Cabugao 1, Tatjana Sauka-Spengler
Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome Marcos S. Simões-Costa 1., Sonja J. McKeown 1., Joanne Tan-Cabugao 1, Tatjana Sauka-Spengler
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
 DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
CJ LI LRRK2 MOUSE COMPARISON STUDY. Phenotyping Data Results
 CJ LI MOUSE COMPARISON STUDY Phenotyping Data Results NOTE FOR USE This comparison study was run as an opportunistic look at the phenotype of multiple CJ Li-generated mouse lines. The primary purpose of
CJ LI MOUSE COMPARISON STUDY Phenotyping Data Results NOTE FOR USE This comparison study was run as an opportunistic look at the phenotype of multiple CJ Li-generated mouse lines. The primary purpose of
Gene Regula*on, ChIP- X and DNA Mo*fs. Statistics in Genomics Hongkai Ji
 Gene Regula*on, ChIP- X and DNA Mo*fs Statistics in Genomics Hongkai Ji (hji@jhsph.edu) Genetic information is stored in DNA TCAGTTGGAGCTGCTCCCCCACGGCCTCTCCTCACATTCCACGTCCTGTAGCTCTATGACCTCCACCTTTGAGTCCCTCCTC
Gene Regula*on, ChIP- X and DNA Mo*fs Statistics in Genomics Hongkai Ji (hji@jhsph.edu) Genetic information is stored in DNA TCAGTTGGAGCTGCTCCCCCACGGCCTCTCCTCACATTCCACGTCCTGTAGCTCTATGACCTCCACCTTTGAGTCCCTCCTC
Unit 4 Evaluation Question 1:
 Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
SUPPLEMENTARY INFORMATION
 PRC2 represses dedifferentiation of mature somatic cells in Arabidopsis Momoko Ikeuchi 1 *, Akira Iwase 1 *, Bart Rymen 1, Hirofumi Harashima 1, Michitaro Shibata 1, Mariko Ohnuma 1, Christian Breuer 1,
PRC2 represses dedifferentiation of mature somatic cells in Arabidopsis Momoko Ikeuchi 1 *, Akira Iwase 1 *, Bart Rymen 1, Hirofumi Harashima 1, Michitaro Shibata 1, Mariko Ohnuma 1, Christian Breuer 1,
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Supplementary Figure 1. Phenotype of the HI strain.
 Supplementary Figure 1. Phenotype of the HI strain. (A) Phenotype of the HI and wild type plant after flowering (~1month). Wild type plant is tall with well elongated inflorescence. All four HI plants
Supplementary Figure 1. Phenotype of the HI strain. (A) Phenotype of the HI and wild type plant after flowering (~1month). Wild type plant is tall with well elongated inflorescence. All four HI plants
** LCA LCN PCA
 % of wild type value % of wild type value a 12 1 8 2 b 12 1 8 2 LCA LCN PCA Col- sod3-1 Supplementary Figure 1 sod3-1 influences cell proliferation. (a) Fifth leaf cell area (LCA) and leaf cell number
% of wild type value % of wild type value a 12 1 8 2 b 12 1 8 2 LCA LCN PCA Col- sod3-1 Supplementary Figure 1 sod3-1 influences cell proliferation. (a) Fifth leaf cell area (LCA) and leaf cell number
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Molecular biology and biochemistry of archaeal DNA replication
 Molecular biology and biochemistry of archaeal DNA replication Isaac Cann Institute for Universal Biology, NASA Astrobiology Institute Department of Animal Science & Department of Microbiology University
Molecular biology and biochemistry of archaeal DNA replication Isaac Cann Institute for Universal Biology, NASA Astrobiology Institute Department of Animal Science & Department of Microbiology University
Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with
 Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with DsRed-mito. Right panels are time-course enlarged images
Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with DsRed-mito. Right panels are time-course enlarged images
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
2. Yeast two-hybrid system
 2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011) /tpc
 Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011). 10.1105/tpc.110.080788 Supplemental Figure S1. Phylogenetic tree of MYC2-related proteins from Arabidopsis and other plants. Phenogram representation
Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011). 10.1105/tpc.110.080788 Supplemental Figure S1. Phylogenetic tree of MYC2-related proteins from Arabidopsis and other plants. Phenogram representation
Statistical analysis of genomic binding sites using high-throughput ChIP-seq data
 Statistical analysis of genomic binding sites using high-throughput ChIP-seq data Ibrahim Ali H Nafisah Department of Statistics University of Leeds Submitted in accordance with the requirments for the
Statistical analysis of genomic binding sites using high-throughput ChIP-seq data Ibrahim Ali H Nafisah Department of Statistics University of Leeds Submitted in accordance with the requirments for the
Genes, Development, and Evolution
 14 Genes, Development, and Evolution Chapter 14 Genes, Development, and Evolution Key Concepts 14.1 Development Involves Distinct but Overlapping Processes 14.2 Changes in Gene Expression Underlie Cell
14 Genes, Development, and Evolution Chapter 14 Genes, Development, and Evolution Key Concepts 14.1 Development Involves Distinct but Overlapping Processes 14.2 Changes in Gene Expression Underlie Cell
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Information
 Supplementary Information MAP2/Hoechst Hyp.-AP ph 6.5 Hyp.-SD ph 7.2 Norm.-SD ph 7.2 Supplementary Figure 1. Mitochondrial elongation in cortical neurons by acidosis. Representative images of neuronal
Supplementary Information MAP2/Hoechst Hyp.-AP ph 6.5 Hyp.-SD ph 7.2 Norm.-SD ph 7.2 Supplementary Figure 1. Mitochondrial elongation in cortical neurons by acidosis. Representative images of neuronal
Supporting Information
 Supporting Information López et al. 10.1073/pnas.0810940106 1. Ivey DM, et al. (1993) Cloning and characterization of a putative Ca2 /H antiporter gene from Escherichia coli upon functional complementation
Supporting Information López et al. 10.1073/pnas.0810940106 1. Ivey DM, et al. (1993) Cloning and characterization of a putative Ca2 /H antiporter gene from Escherichia coli upon functional complementation
** * * * Col-0 cau1 CAU1. Actin2 CAS. Actin2. Supplemental Figure 1. CAU1 affects calcium accumulation.
 Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Supplemental Data. Fu et al. Plant Cell. (213). 1.115/tpc.113.113886 A 5 4 3 * Col- cau1 B 4 3 2 Col- cau1 ** * * ** C 2 1 25 2 15 1 5 Shoots Roots *
Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Supplemental Data. Fu et al. Plant Cell. (213). 1.115/tpc.113.113886 A 5 4 3 * Col- cau1 B 4 3 2 Col- cau1 ** * * ** C 2 1 25 2 15 1 5 Shoots Roots *
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
Genome 541! Unit 4, lecture 3! Genomics assays
 Genome 541! Unit 4, lecture 3! Genomics assays Much easier to follow with slides. Good pace.! Having the slides was really helpful clearer to read and easier to follow the trajectory of the lecture.!!
Genome 541! Unit 4, lecture 3! Genomics assays Much easier to follow with slides. Good pace.! Having the slides was really helpful clearer to read and easier to follow the trajectory of the lecture.!!
Genome 541 Gene regulation and epigenomics Lecture 2 Transcription factor binding using functional genomics
 Genome 541 Gene regulation and epigenomics Lecture 2 Transcription factor binding using functional genomics I believe it is helpful to number your slides for easy reference. It's been a while since I took
Genome 541 Gene regulation and epigenomics Lecture 2 Transcription factor binding using functional genomics I believe it is helpful to number your slides for easy reference. It's been a while since I took
ChIP-seq analysis M. Defrance, C. Herrmann, S. Le Gras, D. Puthier, M. Thomas.Chollier
 ChIP-seq analysis M. Defrance, C. Herrmann, S. Le Gras, D. Puthier, M. Thomas.Chollier Data visualization, quality control, normalization & peak calling Peak annotation Presentation () Practical session
ChIP-seq analysis M. Defrance, C. Herrmann, S. Le Gras, D. Puthier, M. Thomas.Chollier Data visualization, quality control, normalization & peak calling Peak annotation Presentation () Practical session
Supplementary Figure 1. AnnexinV FITC and Sytox orange staining in wild type, Nlrp3 /, ASC / and casp1/11 / TEC treated with TNF /CHX.
 Supplementary Figure 1. AnnexinV FITC and Sytox orange staining in wild type, Nlrp3 /, ASC / and casp1/11 / TEC treated with TNF /CHX. Phase contrast and widefield fluorescence microscopy (20x magnification).
Supplementary Figure 1. AnnexinV FITC and Sytox orange staining in wild type, Nlrp3 /, ASC / and casp1/11 / TEC treated with TNF /CHX. Phase contrast and widefield fluorescence microscopy (20x magnification).
Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field.
 Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field. A, Phenotypes of 30-day old wild-type (WT) and abci8 mutant plants grown in a paddy field under normal sunny
Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field. A, Phenotypes of 30-day old wild-type (WT) and abci8 mutant plants grown in a paddy field under normal sunny
Supplementary Figure 1 Analysis of beige fat and cells and characteristics of exosome release, related to Figure 1
 Supplementary Figure 1 Analysis of beige fat and cells and characteristics of exosome release, related to Figure 1 (a) Fold-change in UCP-1 mrna abundance in white adipocytes upon β-adrenergic stimulation
Supplementary Figure 1 Analysis of beige fat and cells and characteristics of exosome release, related to Figure 1 (a) Fold-change in UCP-1 mrna abundance in white adipocytes upon β-adrenergic stimulation
Biology 218, practise Exam 2, 2011
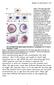 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated
 Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated form by recombinant UGT74E2. The naturally occurring auxin
Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated form by recombinant UGT74E2. The naturally occurring auxin
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Design, synthesis and binding properties of a fluorescent α 9 β 1 /α 4 β 1 integrin antagonist and its application as an in vivo probe for bone marrow haemopoietic stem cells Benjamin
SUPPORTING INFORMATION Design, synthesis and binding properties of a fluorescent α 9 β 1 /α 4 β 1 integrin antagonist and its application as an in vivo probe for bone marrow haemopoietic stem cells Benjamin
Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc
 OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1121356/dc1 Supporting Online Material for Polar PIN Localization Directs Auxin Flow in Plants Justyna Wiśniewska, Jian Xu, Daniela Seifertová, Philip B. Brewer, Kamil
www.sciencemag.org/cgi/content/full/1121356/dc1 Supporting Online Material for Polar PIN Localization Directs Auxin Flow in Plants Justyna Wiśniewska, Jian Xu, Daniela Seifertová, Philip B. Brewer, Kamil
SUPPLEMENTARY INFORMATION
 GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
Measures of hydroxymethylation
 Measures of hydroxymethylation Alla Slynko Axel Benner July 22, 2018 arxiv:1708.04819v2 [q-bio.qm] 17 Aug 2017 Abstract Hydroxymethylcytosine (5hmC) methylation is well-known epigenetic mark impacting
Measures of hydroxymethylation Alla Slynko Axel Benner July 22, 2018 arxiv:1708.04819v2 [q-bio.qm] 17 Aug 2017 Abstract Hydroxymethylcytosine (5hmC) methylation is well-known epigenetic mark impacting
Supplemental Information. Inferring Cell-State Transition. Dynamics from Lineage Trees. and Endpoint Single-Cell Measurements
 Cell Systems, Volume 3 Supplemental Information Inferring Cell-State Transition Dynamics from Lineage Trees and Endpoint Single-Cell Measurements Sahand Hormoz, Zakary S. Singer, James M. Linton, Yaron
Cell Systems, Volume 3 Supplemental Information Inferring Cell-State Transition Dynamics from Lineage Trees and Endpoint Single-Cell Measurements Sahand Hormoz, Zakary S. Singer, James M. Linton, Yaron
Last time: Obtaining information from a cloned gene
 Last time: Obtaining information from a cloned gene Objectives: 1. What is the biochemical role of the gene? 2. Where and when is the gene expressed (transcribed)? 3. Where and when is the protein made?
Last time: Obtaining information from a cloned gene Objectives: 1. What is the biochemical role of the gene? 2. Where and when is the gene expressed (transcribed)? 3. Where and when is the protein made?
Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct
 SUPPLEMENTAL FIGURE LEGENDS Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct waves. (A) A representative sub-lineage (ABala) of the C. elegans lineage tree (adapted from
SUPPLEMENTAL FIGURE LEGENDS Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct waves. (A) A representative sub-lineage (ABala) of the C. elegans lineage tree (adapted from
REVIEW SESSION. Wednesday, September 15 5:30 PM SHANTZ 242 E
 REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
Eukaryotic Gene Expression
 Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Measuring TF-DNA interactions
 Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Discovering MultipleLevels of Regulatory Networks
 Discovering MultipleLevels of Regulatory Networks IAS EXTENDED WORKSHOP ON GENOMES, CELLS, AND MATHEMATICS Hong Kong, July 25, 2018 Gary D. Stormo Department of Genetics Outline of the talk 1. Transcriptional
Discovering MultipleLevels of Regulatory Networks IAS EXTENDED WORKSHOP ON GENOMES, CELLS, AND MATHEMATICS Hong Kong, July 25, 2018 Gary D. Stormo Department of Genetics Outline of the talk 1. Transcriptional
An Information-Theoretic Approach to Methylation Data Analysis
 An Information-Theoretic Approach to Methylation Data Analysis John Goutsias Whitaker Biomedical Engineering Institute The Johns Hopkins University Baltimore, MD 21218 Objective Present an information-theoretic
An Information-Theoretic Approach to Methylation Data Analysis John Goutsias Whitaker Biomedical Engineering Institute The Johns Hopkins University Baltimore, MD 21218 Objective Present an information-theoretic
Regulation of gene expression. Premedical - Biology
 Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Towards the Ultimate Solution: Genetic Resistance to HLB in Commercial Citrus. Greening Summit Florida Citrus Growers Institute 2008
 Towards the Ultimate Solution: Genetic Resistance to HLB in Commercial Citrus Greening Summit Florida Citrus Growers Institute 2008 Jude Grosser University of Florida, Citrus Research and Education Center,
Towards the Ultimate Solution: Genetic Resistance to HLB in Commercial Citrus Greening Summit Florida Citrus Growers Institute 2008 Jude Grosser University of Florida, Citrus Research and Education Center,
Amersham CyDye fluors
 GE Healthcare Life Sciences Amersham CyDye fluors Selection Guide Welcome to Amersham CyDye fluors Amersham CyDye fluors are versatile fluorophores designed for a broad range of applications. CyDye fluors
GE Healthcare Life Sciences Amersham CyDye fluors Selection Guide Welcome to Amersham CyDye fluors Amersham CyDye fluors are versatile fluorophores designed for a broad range of applications. CyDye fluors
Chromosome Chr Duplica Duplic t a ion Pixley
 Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence
 Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence www.plantcell.org/cgi/doi/10.1105/tpc.110.tt0110 Epigenetics Usually
Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence www.plantcell.org/cgi/doi/10.1105/tpc.110.tt0110 Epigenetics Usually
