SUPPLEMENTARY INFORMATION
|
|
|
- Silas Hardy
- 6 years ago
- Views:
Transcription
1 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 2 ARTICLE NUMBER: Structure of the mycobacterial ESX-5 type VII secretion system membrane complex by single particle analysis Katherine S.H. Beckham 1#, Luciano Ciccarelli 2,3,4#, Catalin M. Bunduc 5#, Haydyn D.T. Mertens 1, Roy Ummels 6, Wolfgang Lugmayr 2,3,4, Julia Mayr 2,3,4, Mandy Rettel 7, Mikhail M. Savitski 7, Dmitri I. Svergun 1, Wilbert Bitter 5,6, Matthias Wilmanns 1,4,8, Thomas C. Marlovits 2,3,4,8,9*, Annabel H.A. Parret 1*, Edith N.G. Houben 5* # equal contribution * Correspondence: Thomas.Marlovits@imba.oeaw.ac.at, parret@embl-hamburg.de, e.n.g.houben@vu.nl 1 European Molecular Biology Laboratory, Hamburg Unit, Notkestraße 85, 22607, Hamburg, Germany 2 Research Institute of Molecular Pathology, Dr. Bohr-Gasse 7, 1030 Vienna, Austria 3 Institute of Molecular Biotechnology, Austrian Academy of Sciences, Dr. Bohr-Gasse 3, 1030 Vienna, Austria 4 University Medical Centre Hamburg-Eppendorf, Martinistraße 52, Hamburg, Germany 5 Vrije Universiteit Amsterdam, De Boelelaan 1108, 1081 HZ Amsterdam, The Netherlands 6 VU University Medical Center, De Boelelaan 1108, 1081 HZ Amsterdam, The Netherlands 7 European Molecular Biology Laboratory, Meyerhofstraße 1, Heidelberg, Germany 8 Centre for Structural Systems Biology, Notkestraße 85, Hamburg, Germany 9 Deutsches Elektronen-Synchrotron, Notkestraße 85, Hamburg, Germany This file contains Supplementary Figure 1-9 and Supplementary Tables 1-3. NATURE MICROBIOLOGY DOI: /nmicrobiol Macmillan Publishers Limited, part of Springer Nature. All rights reserved.
2 Supplementary Figure 1. Conserved gene architecture and expression of esx-5 loci. a, Gene organization of the esx-5 locus in M. tuberculosis H37Rv. The homologous esx-5 locus from M. xenopi was expressed in M. smegmatis from the pmv-esx-5 construct. Genes have been coloured according the scheme shown in Fig. 3g. The Strep-tag II on eccc5 is shown as a black box. To test the effect of secreted effectors on complex formation, all genes encoding for substrates (two pe, two ppe and two esx genes) were deleted to generate pmv-esx-5 substr as indicated by the dotted lines. b, Immunoblot of detergent-solubilized cell envelope fractions of M. smegmatis expressing pmv-esx-5 and pmv-esx-5 substr analysed by blue native PAGE. Intact ESX-5 complex is indicated by a black arrow. Representative immunoblot using anti-eccb 5 antibodies from at least three independent experiments.
3 a b DDM A8-35 c A8-35 DDM standards Normalized mau (280 nm) d Volume (ml) 66 - Supplementary Figure 2. Improved stability of the ESX-5 complex in amphipols. a, Analytical size exclusion chromatography analysis of the ESX-5 complex in DDM (grey line) versus amphipol A8-35 (black line). The arrow indicates the elution fraction used for blue native PAGE and EM analysis. The molecular weight of a standard set of protein markers is indicated on top of the corresponding peak (dotted line; in kilodaltons). b. Coomassiestained blue native PAGE gel of ESX-5 complex solubilized in DDM or A8-35. Representative gel from at least three independent experiments. c, and d, Representative negative stain EM showing ESX-5 particles solubilized in DDM and A8-35, respectively. Scale bar is 50 nm.
4 a b c 1.0 d 50º 0º CC FSC FSC = Å Degrees Resolution (1/Å) e f g 90º 150º 100º 50º 180º 0º Untilted Tilted Untilted Tilted Untilted Tilted 270º Supplementary Figure 3. Single particle analysis of ESX-5 complexes and map validation. a, Gallery of representative 2D class averages. b, Rotational cross-correlation (CC) from a representative 2D class average. c, Fourier Shell Correlation curve of the ESX-5 complex reconstruction. The resolution at was 13.1Å. d, Upper panel shows a representative tilt-pair of ESX-5 particles at 0 and 50 tilting. Below are the corresponding views of the 3D map. e, Angular distribution applying C6 symmetry is shown by the graphical display of e2eulerxplor.py. Projections in top, intermediate and side views are shown on the left row adjacent to the corresponding 2D class average. Particle orientation is evenly distributed in the 3D space extending from the equator to the pole, enabling a robust negative stain 3D reconstruction with no missing information. f, Representative tilt-pairs collected at 0 and 50 and subjected to tilt-pair analysis. g, Tilt-pair parameter plot (TPPP) 1 of tilt-pairs shows a clear clustering around the 50 tilt angle corresponding to the angle that was used to record the second micrograph of the tilt pair.
5 a Absorbance at 280 nm (mau) EccC 5 99 Theoretical MW (Da) Experimental MW (Da) Monomer 142, ,971 Dimer 284, , Molecular weight (Da) Retention volume (ml) b c P(r) P(r) log log I(s) I(S) DUF A1 A2 A3 Χ D max (nm) r (nm) s (nm -1 ) Supplementary Figure 4. Analysis of truncated EccC5 (EccC5 99 ) by SEC-MALS and SAXS. a, Molecular weights were calculated from analytical size-exclusion chromatography analysis coupled with static light scattering and plotted across the peaks (blue). The derived molar weights of 147 and 310 match the expected molecular weights of a monomeric and dimeric protein, respectively. b, Pair-wise distance distribution (P(r)) of the scattering data. c, Fits of model ensemble 3 to the raw SAXS data. Inset shows the schematic overview of modelling approaches used with the Ensemble Optimization Method (EOM). Residue boundaries of EccC 5 domains are as follows: DUF, ; A1, ; A2, ; A3, Domains boxed by dotted lines were treated as rigid bodies. The chi-squared ( 2 ) values of each of the EOM models are indicated.
6 a b Approach 1 Approach 2 i) Modelling EccB 5 hexamer i) Segmentation analysis Top Side Bottom Top Side Side cross-section ii) Docking EccB 5 hexamer ii) Docking EccB 5 monomer 90 Side Top Top Side Side cross-section iii) Applying 6-fold symmetry Top Side Side cross-section Supplementary Figure 5. Overview outlining EccB5 docking procedures. a, Approach 1 used a hexameric model of EccB5 of M. xenopi (i) generated using the method described by Zhang et al. 2. Subsequently, the EccB5 hexamer was docked into the EM map as a rigid body (ii). Top, side and cross section views are indicated. b, In approach 2, the EM map was segmented using the program Segger revealing a feature consistent with an EccB5 hexamer (i). Next, individual EccB5 monomers were docked into the segmented volume (ii) and a sixfold symmetry was applied (iii). Both docking approaches place the N-terminus of EccB5 at the apex of the map, but differ in their positioning of the C-terminal region of EccB5 preceding the transmembrane domain.
7 a b c EccC 5 strep EccB 5 EccE 5 his EccD Supplementary Figure 6. Purification of M. xenopi ESX-5 EccE-His complex. a, Coomassiestained SDS-PAGE gel of purified ESX-5 complex following affinity purification using the Strep-tag II on EccC5 and size-exclusion chromatography. Representative gel from at least three independent experiments. b, Coomassie-stained blue native PAGE gel of the purified complex as indicated by the black arrow. Representative gel from at least three independent experiments. c, Representative negative stain EM of the ESX-5 complex. Visual inspection of the particles suggests that the overall architecture of ESX-5 EccE-His is comparable to the ESX-5 complex, indicating that the addition of a C-terminal His6-tag on EccE5 has no impact on complex formation or stability. Scale bar is 50 nm.
8 M. xenopi - DTT + DTT 75 - DTT + DTT 50 M. tuberculosis αeccb5 50 αecce5 37 Supplementary Figure 7. Disulphide bridge formation of EccE5 indicative of its periplasmic location. Immunoblot analysis of purified ESX-5 complexes under reducing (+ DTT) and nonreducing (- DTT) conditions to analyse disulphide bridge formation. This experiment was also performed on the ESX-5 system from M. tuberculosis reconstituted in M. smegmatis because of the higher number of cysteines in EccE5 from M. tuberculosis (6) versus EccE5 from M. xenopi (3). As a positive control EccB5, that is known to form a disulfide bridge 3, was also examined. Both EccB5 and EccE5 showed a difference in mobility depending on the presence of DTT. Representative immunoblots using anti-eccb5 and anti- EccE5 antibodies from three independent experiments.
9 Supplementary Figure 8. Purification of M. xenopi ESX-5 ΔEccE complex. a, Coomassie-stained SDS-PAGE gel of purified complex following affinity purification and size-exclusion chromatography, confirming the presence of the three components EccB5, EccC5 strep, EccD5 and the absence of EccE5. Representative gel from at least three independent experiments. b, Immunoblot of blue native PAGE gel of purified ESX-5 ΔEccE complex indicating a molecular mass of ~ 1,3 MDa. Representative immunoblot using anti-eccb5 antibodies from at least three independent experiments. c, Representative size exclusion chromatography profile of ESX-5 ΔEccE. The yield of the ESX-5 ΔEccE complex is considerably lower than that of ESX-5. The black arrow indicates the fraction shown in panel d. d, Representative electron micrograph showing ESX-5 ΔEccE particles in several orientations. The overall diameter of the particles is ~22-25nm. Scale bar is 50 nm. Representative particles of the ESX-5 ΔEccE (e) and the ESX-5 complex (f) (box size = 180 x 180 pixels). g, Gallery of class averages of 684 ESX-5 ΔEccE particles. h, Four class averages of ESX-5 and ESX-5 ΔEccE complexes rotationally aligned are shown at same magnification. i, Histogram showing the average diameter for both ESX-5 and ESX-5 ΔEccE complexes. Particles were measured along the horizontal axis, the average diameter of the ESX-5 and ESX-5 ΔEccE complexes is 28.9 ± 1.2 nm and 23.8 ± 1.3, respectively. Error bars indicate the standard deviation of 50 particles. T-test analysis shows a significant difference with a p-value
10 Full blots of results Figure 1a P S ESX-5 P S ESX αgroel αeccb5 αppe18-ha sec exposure αesxn sec exposure Figure 1b
11 Full blots of results (continued) Supplementary Fig. 1b ESX-5 ESX-5 Δsubstr αeccb Supplementary Fig. 2b A8-35 DDM
12 Full blots of results (continued) Supplementary Fig. 6a Supplementary Fig. 6b
13 Full blots of results (continued) Supplementary Fig. 7 M. xenopi - + DTT - + DTT αeccb5 αecce5 M. tuberculosis - + DTT - + DTT αeccb5-37 αecce5
14 Full blots of results (continued) Supplementary Fig. 8a Supplementary Fig. 8b
15 Supplementary Table 1 Quantitative mass-spectrometry analysis of ESX-5 to determine subunit stoichiometry. Average relative protein concentrations of the four subunits was determined using ibaq-based quantitative mass spectrometry and calculated across four biological replicates (1-4). These date indicate a 1:1:1:1 stoichiometry for EccB 5:EccC 5 :EccD 5:EccE 5. Proteins ibaq Relative protein concentration Average relative protein concentration S.E. Razor + unique peptides Sequence coverage (%) MW () EccB 5 3.5E E E E EccC 5 4.5E E E E EccD 5 5.2E E E E EccE 5 4.1E E E E
16 Supplementary Table 2 EM data collection and structural analysis of ESX-5. Micrographs 65 Particles 1,418 Defocus um Detector FEI BM Eagle Pixel size 2.18 Å Reconstruction method EMAN2 - e2initialmodel Symmetry 6-fold Resolution FSC gold standard 13.1 Å
17 Supplementary Table 3 SAXS data collection and derived parameters for EccC5 99. Data collection parameters Instrument EMBL P12 beam line (PETRA-III, DESY, Hamburg) Beam geometry 0.2 x 0.12 mm 2 Wavelength (Å) 1.24 s range (Å -1 ) a Exposure time (s) 1 (20 x 0.05) Concentration range (mg/ml) Temperature (K) 283 Structural parameters b I(0) (relative) (from p(r)) ± Rg (Å) (from p(r)) 65 ± 1 I(0) (relative) (from Guinier) ± Rg (Å) (from Guinier) 62 ± 1 Dmax (Å) 230 Porod volume estimate (Å 3 ) 255,000 ± 30,000 Excluded volume estimate (Å 3 ) 340,000 ± 50,000 Dry volume calculated c from sequence (Å 3 ) 172,365 Molecular mass determination I(0) (cm -1 ) BSA (72,000 Da) ± Molecular mass Mr (Da) (from I(0)) 141,000 ± 15,000 Molecular mass Mr (Da) (from Porod volume (Vp/1.7)) 159,000 ± 15,000 Molecular mass Mr (Da) (from excluded volume (Vex/2)) 173,500 ± 20,000 Calculated monomeric Mr (from sequence) 142,450 Software employed Primary data reduction Data processing Equilibrium analysis Computation of model intensities 3D graphics representations a Momentum transfer s = 4πsin(θ)/λ b Values reported for 0.4 mg ml -1 RADAVER PRIMUS/Qt EOM CRYSOL PyMOL, UCSF Chimera c Dry volume from: Abbreviations: Mr : Molecular mass Rg : Radius of gyration Dmax : Maximal particle dimension Vp : Porod volume Vex : Particle excluded volume
18 Supplementary References 1. Henderson, R. et al. Tilt-pair analysis of images from a range of different specimens in single-particle electron cryomicroscopy. J. Mol. Biol. 413, (2011). 2. Zhang, X.-L. et al. Core component EccB1 of the Mycobacterium tuberculosis type VII secretion system is a periplasmic ATPase. FASEB J. 29, (2015). 3. Wagner, J. M. et al. Structures of EccB1 and EccD1 from the core complex of the mycobacterial ESX-1 type VII secretion system. BMC Struct. Biol. 16, 5 (2016).
Supplementary figure 1. Comparison of unbound ogm-csf and ogm-csf as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine
 Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Definition and assessment of ciap1 constructs.
 Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplemental Information. Structural and Mechanistic Paradigm. of Leptin Receptor Activation Revealed
 Structure, Volume 22 Supplemental Information Structural and Mechanistic Paradigm of Leptin Receptor Activation Revealed by Complexes with Wild-Type and Antagonist Leptins Kedar Moharana, Lennart Zabeau,
Structure, Volume 22 Supplemental Information Structural and Mechanistic Paradigm of Leptin Receptor Activation Revealed by Complexes with Wild-Type and Antagonist Leptins Kedar Moharana, Lennart Zabeau,
Watanabe'et'al.' Supplementary'Figure'1'
 MW IN O 80 2 IN O 80 SW 3 R -C 3 MW SW R -C SW 1 R -C 2 IN O 80 1 Supplementary'Figure'1' Supplementary Figure 1. SDS-PAGE image of INO80-C and SWR-C. Three independent preparations of each TAP-tagged
MW IN O 80 2 IN O 80 SW 3 R -C 3 MW SW R -C SW 1 R -C 2 IN O 80 1 Supplementary'Figure'1' Supplementary Figure 1. SDS-PAGE image of INO80-C and SWR-C. Three independent preparations of each TAP-tagged
SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition SUPPLEMENTARY INFORMATION
 SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition Nadia J. Kershaw 1,2, James M. Murphy 1,2, Nicholas P.D. Liau 1,2, Leila N. Varghese 1,2, Artem Laktyushin
SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition Nadia J. Kershaw 1,2, James M. Murphy 1,2, Nicholas P.D. Liau 1,2, Leila N. Varghese 1,2, Artem Laktyushin
SI Text S1 Solution Scattering Data Collection and Analysis. SI references
 SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
BM29 biosaxs data processing tutorial
 HERCULES 2014 BM29 biosaxs data processing tutorial Page 2 OUTLINE Sample Changer Primary Data Processing Model Validation HPLC-SAXS Primary Data Processing Model Validation Ab Initio Model Software in
HERCULES 2014 BM29 biosaxs data processing tutorial Page 2 OUTLINE Sample Changer Primary Data Processing Model Validation HPLC-SAXS Primary Data Processing Model Validation Ab Initio Model Software in
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013
 Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013 Structural Biology Shape Dynamic Light Scattering Electron Microscopy Small Angle X-ray Scattering Cryo-Electron Microscopy Wide Angle X- ray
Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013 Structural Biology Shape Dynamic Light Scattering Electron Microscopy Small Angle X-ray Scattering Cryo-Electron Microscopy Wide Angle X- ray
ID14-EH3. Adam Round
 Bio-SAXS @ ID14-EH3 Adam Round Contents What can be obtained from Bio-SAXS Measurable parameters Modelling strategies How to collect data at Bio-SAXS Procedure Data collection tests Data Verification and
Bio-SAXS @ ID14-EH3 Adam Round Contents What can be obtained from Bio-SAXS Measurable parameters Modelling strategies How to collect data at Bio-SAXS Procedure Data collection tests Data Verification and
Sample preparation and characterization around SAXS
 Sample preparation and characterization around SAXS Experimental verification and validation? Rob Meijers EMBL Hamburg Garbage in? The right stuff Molecular weight Oligomerization state Monodispersity
Sample preparation and characterization around SAXS Experimental verification and validation? Rob Meijers EMBL Hamburg Garbage in? The right stuff Molecular weight Oligomerization state Monodispersity
Small-Angle Scattering Atomic Structure Based Modeling
 Small-Angle Scattering Atomic Structure Based Modeling Alejandro Panjkovich EMBL Hamburg 07.12.2017 A. Panjkovich (EMBL) BioSAS atomic modeling 07.12.2017 1 / 49 From the forest to the particle accelerator
Small-Angle Scattering Atomic Structure Based Modeling Alejandro Panjkovich EMBL Hamburg 07.12.2017 A. Panjkovich (EMBL) BioSAS atomic modeling 07.12.2017 1 / 49 From the forest to the particle accelerator
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
The Fic protein Doc uses an inverted substrate to phosphorylate and. inactivate EF-Tu
 The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/310/5753/1513/dc1 Supporting Online Material for Structural Roles for Human Translation Factor eif3 in Initiation of Protein Synthesis Bunpote Siridechadilok, Christopher
www.sciencemag.org/cgi/content/full/310/5753/1513/dc1 Supporting Online Material for Structural Roles for Human Translation Factor eif3 in Initiation of Protein Synthesis Bunpote Siridechadilok, Christopher
Acta Cryst. (2017). D73, , doi: /s
 Supporting information Volume 73 (2017) Supporting information for article: 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update
Supporting information Volume 73 (2017) Supporting information for article: 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update
Modelling against small angle scattering data. Al Kikhney EMBL Hamburg, Germany
 Modelling against small angle scattering data Al Kikhney EMBL Hamburg, Germany Validation of atomic models CRYSOL Rigid body modelling SASREF BUNCH CORAL Oligomeric mixtures OLIGOMER Flexible systems EOM
Modelling against small angle scattering data Al Kikhney EMBL Hamburg, Germany Validation of atomic models CRYSOL Rigid body modelling SASREF BUNCH CORAL Oligomeric mixtures OLIGOMER Flexible systems EOM
Use of SEC-MALS. (Size Exclusion Chromatography - Multi Angle. Light Scattering) for protein quality and characterization
 Use of SEC-MALS (Size Exclusion Chromatography - Multi Angle Light Scattering) for protein quality and characterization Methods for protein characterization Analytical SEC is a common method to characterize
Use of SEC-MALS (Size Exclusion Chromatography - Multi Angle Light Scattering) for protein quality and characterization Methods for protein characterization Analytical SEC is a common method to characterize
Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins. Michael Kachala EMBL-Hamburg, Germany
 Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins Michael Kachala EMBL-Hamburg, Germany 60 mkl >1 mg/ml Monocromatic X-ray beam Sample Mono- or polydisperse
Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins Michael Kachala EMBL-Hamburg, Germany 60 mkl >1 mg/ml Monocromatic X-ray beam Sample Mono- or polydisperse
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
SUPPLEMENTARY INFORMATION
 SUPPLMTARY IFORMATIO a doi:10.108/nature10402 b 100 nm 100 nm c SAXS Model d ulers assigned to reference- Back-projected free class averages class averages Refinement against single particles Reconstructed
SUPPLMTARY IFORMATIO a doi:10.108/nature10402 b 100 nm 100 nm c SAXS Model d ulers assigned to reference- Back-projected free class averages class averages Refinement against single particles Reconstructed
depends on the translocation assembly module nanomachine
 ARTICLE NUMBER: 16064 DOI: 10.1038/NMICROBIOL.16.64 Effective assembly of fimbriae in Escherichia coli depends on the translocation assembly module nanomachine Christopher Stubenrauch 1, Matthew J. Belousoff
ARTICLE NUMBER: 16064 DOI: 10.1038/NMICROBIOL.16.64 Effective assembly of fimbriae in Escherichia coli depends on the translocation assembly module nanomachine Christopher Stubenrauch 1, Matthew J. Belousoff
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Data reduction and processing tutorial
 Data reduction and processing tutorial Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group EMBL BioSAXS beamline X33, 2012 Optics Vacuum cell Completely redesigned 2005-2012
Data reduction and processing tutorial Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group EMBL BioSAXS beamline X33, 2012 Optics Vacuum cell Completely redesigned 2005-2012
BSA pegylation PG Journal of chromatographic A, 1147 (2007)
 BSA pegylation BSA 5mg/mL, pi= 4.7, Sigma A796, 65-7% free sulfyhydryl, cysteine 34 of BSA is not paired with another cysteine in the protein structure 1. PEG 1kD, kd, 3 kd 1mg/mL PEG:BSA=1.2:1 (molar
BSA pegylation BSA 5mg/mL, pi= 4.7, Sigma A796, 65-7% free sulfyhydryl, cysteine 34 of BSA is not paired with another cysteine in the protein structure 1. PEG 1kD, kd, 3 kd 1mg/mL PEG:BSA=1.2:1 (molar
Small Angle X-Ray Solution Scattering of Biological Macromolecules
 Small Angle X-Ray Solution Scattering of Biological Macromolecules Emre Brookes UltraScan Workshop 15 June 2014 Overview Experimental method Sample preparation Experimental data analysis Experimental method
Small Angle X-Ray Solution Scattering of Biological Macromolecules Emre Brookes UltraScan Workshop 15 June 2014 Overview Experimental method Sample preparation Experimental data analysis Experimental method
ml. ph 7.5 ph 6.5 ph 5.5 ph 4.5. β 2 AR-Gs complex + GDP β 2 AR-Gs complex + GTPγS
 a UV28 absorption (mau) 9 8 7 5 3 β 2 AR-Gs complex β 2 AR-Gs complex + GDP β 2 AR-Gs complex + GTPγS β 2 AR-Gs complex dissociated complex excess nucleotides b 9 8 7 5 3 β 2 AR-Gs complex β 2 AR-Gs complex
a UV28 absorption (mau) 9 8 7 5 3 β 2 AR-Gs complex β 2 AR-Gs complex + GDP β 2 AR-Gs complex + GTPγS β 2 AR-Gs complex dissociated complex excess nucleotides b 9 8 7 5 3 β 2 AR-Gs complex β 2 AR-Gs complex
 Supplemental Figure and Movie Legends Figure S1. Time course experiments on the thermal stability of apo AT cpn-α by native PAGE (related to Figure 2B). The samples were heated, respectively, to (A) 45
Supplemental Figure and Movie Legends Figure S1. Time course experiments on the thermal stability of apo AT cpn-α by native PAGE (related to Figure 2B). The samples were heated, respectively, to (A) 45
Cryo-EM data collection, refinement and validation statistics
 1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands
 Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome
 Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome Ahmad Jomaa 1, Yu-Hsien Hwang Fu 2, Daniel Boehringer 1, Marc Leibundgut 1, Shu-ou Shan 2, and Nenad
Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome Ahmad Jomaa 1, Yu-Hsien Hwang Fu 2, Daniel Boehringer 1, Marc Leibundgut 1, Shu-ou Shan 2, and Nenad
ParM filament images were extracted and from the electron micrographs and
 Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples.
 Supplementary Figure 1 Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples. (a-b) SDS-PAGE analysis of the hexamer and heptamer samples. The eluted hexamer
Supplementary Figure 1 Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples. (a-b) SDS-PAGE analysis of the hexamer and heptamer samples. The eluted hexamer
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Introduction to Biological Small Angle Scattering
 Introduction to Biological Small Angle Scattering Tom Grant, Ph.D. Staff Scientist BioXFEL Science and Technology Center Hauptman-Woodward Institute Buffalo, New York, USA tgrant@hwi.buffalo.edu SAXS Literature
Introduction to Biological Small Angle Scattering Tom Grant, Ph.D. Staff Scientist BioXFEL Science and Technology Center Hauptman-Woodward Institute Buffalo, New York, USA tgrant@hwi.buffalo.edu SAXS Literature
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN
 THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
Supplemental Information. Expanded Coverage of the 26S Proteasome. Conformational Landscape Reveals. Mechanisms of Peptidase Gating
 Cell Reports, Volume 24 Supplemental Information Expanded Coverage of the 26S Proteasome Conformational Landscape Reveals Mechanisms of Peptidase Gating Markus R. Eisele, Randi G. Reed, Till Rudack, Andreas
Cell Reports, Volume 24 Supplemental Information Expanded Coverage of the 26S Proteasome Conformational Landscape Reveals Mechanisms of Peptidase Gating Markus R. Eisele, Randi G. Reed, Till Rudack, Andreas
Structure of the α-helix
 Structure of the α-helix Structure of the β Sheet Protein Dynamics Basics of Quenching HDX Hydrogen exchange of amide protons is catalyzed by H 2 O, OH -, and H 3 O +, but it s most dominated by base
Structure of the α-helix Structure of the β Sheet Protein Dynamics Basics of Quenching HDX Hydrogen exchange of amide protons is catalyzed by H 2 O, OH -, and H 3 O +, but it s most dominated by base
V19 Topology of Protein Complexes on Graphs
 V19 Topology of Protein Complexes on Graphs (1) CombDock (2) Nuclear Pore Complex 19. Lecture WS 2013/14 Bioinformatics III 1 Prediction Structures of Protein Complexes from Connectivities CombDock: automated
V19 Topology of Protein Complexes on Graphs (1) CombDock (2) Nuclear Pore Complex 19. Lecture WS 2013/14 Bioinformatics III 1 Prediction Structures of Protein Complexes from Connectivities CombDock: automated
The Effect of Linker DNA on the Structure and Interaction of Nucleosome Core Particles
 Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 SUPPLEMENTARY INFORMATION The Effect of Linker DNA on the Structure and Interaction of Nucleosome
Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 SUPPLEMENTARY INFORMATION The Effect of Linker DNA on the Structure and Interaction of Nucleosome
Size Exclusion Chromatography in the Presence of an Anionic Surfactant
 Size Exclusion Chromatography in the Presence of an Anionic Surfactant Intact Protein Profiling Application Note Author Andy Coffey Agilent Technologies, Inc Abstract Sodium dodecyl sulfate (SDS, or SLS)
Size Exclusion Chromatography in the Presence of an Anionic Surfactant Intact Protein Profiling Application Note Author Andy Coffey Agilent Technologies, Inc Abstract Sodium dodecyl sulfate (SDS, or SLS)
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R)
 Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Biological Sciences 11 Spring Experiment 4. Protein crosslinking
 Biological Sciences 11 Spring 2000 Experiment 4. Protein crosslinking = C - CH 2 - CH 2 - CH 2 - C = H H GA Cl - H 2 N N H 2 Cl - C - CH 2 - CH 2 - CH 2 - CH 2 - CH 2 - CH 2 - C DMS CH 3 CH 3 N - - C -
Biological Sciences 11 Spring 2000 Experiment 4. Protein crosslinking = C - CH 2 - CH 2 - CH 2 - C = H H GA Cl - H 2 N N H 2 Cl - C - CH 2 - CH 2 - CH 2 - CH 2 - CH 2 - CH 2 - C DMS CH 3 CH 3 N - - C -
SUPPLEMENTARY FIGURES. Structure of the cholera toxin secretion channel in its. closed state
 SUPPLEMENTARY FIGURES Structure of the cholera toxin secretion channel in its closed state Steve L. Reichow 1,3, Konstantin V. Korotkov 1,3, Wim G. J. Hol 1$ and Tamir Gonen 1,2$ 1, Department of Biochemistry
SUPPLEMENTARY FIGURES Structure of the cholera toxin secretion channel in its closed state Steve L. Reichow 1,3, Konstantin V. Korotkov 1,3, Wim G. J. Hol 1$ and Tamir Gonen 1,2$ 1, Department of Biochemistry
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb97 P ( μm, hours) 1 2 4 P DMSO Figure S1 unningham et al. E 97 65 27 MFN1 GFP-Parkin Opa1 ctin GPDH HEK293 GFP-Parkin 19 115 97 65 27 Mitochondrial Fraction SH-SY5Y GFP-Parkin Mito DMSO Mito
DOI:.38/ncb97 P ( μm, hours) 1 2 4 P DMSO Figure S1 unningham et al. E 97 65 27 MFN1 GFP-Parkin Opa1 ctin GPDH HEK293 GFP-Parkin 19 115 97 65 27 Mitochondrial Fraction SH-SY5Y GFP-Parkin Mito DMSO Mito
How to judge data quality
 SSRL Workshop: Small-Angle X-ray Scattering and Diffraction Studies, March 28-30, 2016 How to judge data quality Tsutomu Matsui SSRL Lab / Dept. of Chemistry Stanford University Subject of this session
SSRL Workshop: Small-Angle X-ray Scattering and Diffraction Studies, March 28-30, 2016 How to judge data quality Tsutomu Matsui SSRL Lab / Dept. of Chemistry Stanford University Subject of this session
Model building and validation for cryo-em maps at low resolution
 International Workshop of Advanced Image Processing of Cryo-Electron Microscopy 2015 Tsinghua University, Beijing, China June 3-7, 2015 Model building and validation for cryo-em maps at low resolution
International Workshop of Advanced Image Processing of Cryo-Electron Microscopy 2015 Tsinghua University, Beijing, China June 3-7, 2015 Model building and validation for cryo-em maps at low resolution
Supplementary Information
 Supplementary Information The direct role of selenocysteine in [NiFeSe] hydrogenase maturation and catalysis Marta C. Marques a, Cristina Tapia b, Oscar Gutiérrez-Sanz b, Ana Raquel Ramos a, Kimberly L.
Supplementary Information The direct role of selenocysteine in [NiFeSe] hydrogenase maturation and catalysis Marta C. Marques a, Cristina Tapia b, Oscar Gutiérrez-Sanz b, Ana Raquel Ramos a, Kimberly L.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10244 a O07391_MYCAV/127-243 NLPC_HAEIN/80-181 SPR_SHIFL/79-183 P74160_SYNY3/112-245 O24914_HELPY/301-437 Q51835_PORGI/68-178 DPP6_BACSH/163-263 YKFC_BACSU/185-292 YDHO_ECOLI/153-263
doi:10.1038/nature10244 a O07391_MYCAV/127-243 NLPC_HAEIN/80-181 SPR_SHIFL/79-183 P74160_SYNY3/112-245 O24914_HELPY/301-437 Q51835_PORGI/68-178 DPP6_BACSH/163-263 YKFC_BACSU/185-292 YDHO_ECOLI/153-263
Full-length GlpG sequence was generated by PCR from E. coli genomic DNA. (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
 Supplementary Methods Protein expression and purification Full-length GlpG sequence was generated by PCR from E. coli genomic DNA (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
Supplementary Methods Protein expression and purification Full-length GlpG sequence was generated by PCR from E. coli genomic DNA (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
Spliceosome and Localization of Its Catalytic Core
 Molecular Cell, Volume 40 Supplemental Information 3D Cryo-EM Structure of an Active Step I Spliceosome and Localization of Its Catalytic Core Monika M. Golas, Bjoern Sander, Sergey Bessonov, Michael Grote,
Molecular Cell, Volume 40 Supplemental Information 3D Cryo-EM Structure of an Active Step I Spliceosome and Localization of Its Catalytic Core Monika M. Golas, Bjoern Sander, Sergey Bessonov, Michael Grote,
Image Assessment San Diego, November 2005
 Image Assessment San Diego, November 005 Pawel A. Penczek The University of Texas Houston Medical School Department of Biochemistry and Molecular Biology 6431 Fannin, MSB6.18, Houston, TX 77030, USA phone:
Image Assessment San Diego, November 005 Pawel A. Penczek The University of Texas Houston Medical School Department of Biochemistry and Molecular Biology 6431 Fannin, MSB6.18, Houston, TX 77030, USA phone:
Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
Transmembrane Domains (TMDs) of ABC transporters
 Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System
 Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System Keywords: Hydrodynamic radius, diffusion coefficient, sizing, Taylor dispersion analysis, protein, antibodies,
Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System Keywords: Hydrodynamic radius, diffusion coefficient, sizing, Taylor dispersion analysis, protein, antibodies,
type GroEL-GroES complex. Crystals were grown in buffer D (100 mm HEPES, ph 7.5,
 Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Enhancing hydrogen production of microalgae by redirecting electrons from photosystem I to hydrogenase
 Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2014 Supplementary information for Enhancing hydrogen production of microalgae
Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2014 Supplementary information for Enhancing hydrogen production of microalgae
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
SUPPLEMENTARY INFORMATION
 Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
Crystal Structure of Fibroblast Growth Factor 9 (FGF9) Reveals Regions. Implicated in Dimerization and Autoinhibition
 JBC Papers in Press. Published on November 1, 2000 as Manuscript M006502200 Crystal Structure of Fibroblast Growth Factor 9 (FGF9) Reveals Regions Implicated in Dimerization and Autoinhibition 1 Copyright
JBC Papers in Press. Published on November 1, 2000 as Manuscript M006502200 Crystal Structure of Fibroblast Growth Factor 9 (FGF9) Reveals Regions Implicated in Dimerization and Autoinhibition 1 Copyright
SUPPLEMENTARY INFORMATION
 doi:10.108/nature11899 Supplementar Table 1. Data collection and refinement statistics (+TPMP, native) (-TPMP, native) (+TPMP, recombinant) (MgCl ) (MgSO ) Data collection Space group C P 1 C P 1 1 P 1
doi:10.108/nature11899 Supplementar Table 1. Data collection and refinement statistics (+TPMP, native) (-TPMP, native) (+TPMP, recombinant) (MgCl ) (MgSO ) Data collection Space group C P 1 C P 1 1 P 1
Three-Dimensional Electron Microscopy of Macromolecular Assemblies
 Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi
 Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi 1 Wikipedia Small-angle X-ray scattering (SAXS) is a small-angle scattering (SAS) technique where the elastic scattering of X-rays (wavelength
Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi 1 Wikipedia Small-angle X-ray scattering (SAXS) is a small-angle scattering (SAS) technique where the elastic scattering of X-rays (wavelength
New insight into the dynamical system of αb-crystallin oligomers
 Supplementary Materials New insight into the dynamical system of αb-crystallin oligomers Rintaro Inoue*, Takumi Takata, Norihiko Fujii, Kentaro Ishii, Susumu Uchiyama, Nobuhiro Sato, Yojiro Oba, Kathleen
Supplementary Materials New insight into the dynamical system of αb-crystallin oligomers Rintaro Inoue*, Takumi Takata, Norihiko Fujii, Kentaro Ishii, Susumu Uchiyama, Nobuhiro Sato, Yojiro Oba, Kathleen
Supplementary Information. Autocatalytic backbone N-methylation in a family of ribosomal peptide natural products
 Supplementary Information Autocatalytic backbone -methylation in a family of ribosomal peptide natural products iels S. van der Velden 1, oemi Kälin 1, Maximilian J. Helf 1, Jörn Piel 1, Michael F. Freeman
Supplementary Information Autocatalytic backbone -methylation in a family of ribosomal peptide natural products iels S. van der Velden 1, oemi Kälin 1, Maximilian J. Helf 1, Jörn Piel 1, Michael F. Freeman
Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6
 Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/6/e1700147/dc1 Supplementary Materials for Ribosome rearrangements at the onset of translational bypassing Xabier Agirrezabala, Ekaterina Samatova, Mariia Klimova,
advances.sciencemag.org/cgi/content/full/3/6/e1700147/dc1 Supplementary Materials for Ribosome rearrangements at the onset of translational bypassing Xabier Agirrezabala, Ekaterina Samatova, Mariia Klimova,
Supplementary Tables and Figures
 Supplementary Tables Supplementary Tables and Figures Supplementary Table 1: Tumor types and samples analyzed. Supplementary Table 2: Genes analyzed here. Supplementary Table 3: Statistically significant
Supplementary Tables Supplementary Tables and Figures Supplementary Table 1: Tumor types and samples analyzed. Supplementary Table 2: Genes analyzed here. Supplementary Table 3: Statistically significant
How to use GPC/SEC for compositional analysis
 How to use GPC/SEC for compositional analysis Determining the relative concentration of two components in a polymer sample MOLECULAR SIZE MOLECULAR STRUCTURE MOLECULAR WEIGHT Introduction Over the last
How to use GPC/SEC for compositional analysis Determining the relative concentration of two components in a polymer sample MOLECULAR SIZE MOLECULAR STRUCTURE MOLECULAR WEIGHT Introduction Over the last
Advantages of Agilent AdvanceBio SEC Columns for Biopharmaceutical Analysis
 Advantages of Agilent AdvanceBio SEC Columns for Biopharmaceutical Analysis Comparing Columns from Different Vendors to Improve Data Quality Technical Overview Introduction Size exclusion chromatography
Advantages of Agilent AdvanceBio SEC Columns for Biopharmaceutical Analysis Comparing Columns from Different Vendors to Improve Data Quality Technical Overview Introduction Size exclusion chromatography
Static and dynamic light scattering. Cy Jeffries EMBL Hamburg
 Static and dynamic light scattering. Cy Jeffries EMBL Hamburg Introduction. The electromagnetic spectrum. visible 10-16 10-10 10-8 10-4 10-2 10 4 (l m) g-rays X-rays UV IR micro wave Long radio waves 400
Static and dynamic light scattering. Cy Jeffries EMBL Hamburg Introduction. The electromagnetic spectrum. visible 10-16 10-10 10-8 10-4 10-2 10 4 (l m) g-rays X-rays UV IR micro wave Long radio waves 400
Supplementary information
 Supplementary information doi: 10.1038/nchem.1025 Platinum nanocrystals selectively shaped using facet-specific peptide sequences Chin-Yi Chiu 1, Yujing Li 1, Lingyan Ruan 1, Xingchen Ye 2, Christopher
Supplementary information doi: 10.1038/nchem.1025 Platinum nanocrystals selectively shaped using facet-specific peptide sequences Chin-Yi Chiu 1, Yujing Li 1, Lingyan Ruan 1, Xingchen Ye 2, Christopher
MEASURING PROTEIN AGGREGATION WITH THE VISCOTEK SEC-MALS 20
 MEASURING PROTEIN AGGREGATION WITH THE VISCOTEK SEC-MALS 20 Introduction Protein aggregation is recognised as a major issue in the biopharmaceutical industry. Proteins have a tendency to aggregate over
MEASURING PROTEIN AGGREGATION WITH THE VISCOTEK SEC-MALS 20 Introduction Protein aggregation is recognised as a major issue in the biopharmaceutical industry. Proteins have a tendency to aggregate over
targets. clustering show that different complex pathway
 Supplementary Figure 1. CLICR allows clustering and activation of cytoplasmic protein targets. (a, b) Upon light activation, the Cry2 (red) and LRP6c (green) components co-cluster due to the heterodimeric
Supplementary Figure 1. CLICR allows clustering and activation of cytoplasmic protein targets. (a, b) Upon light activation, the Cry2 (red) and LRP6c (green) components co-cluster due to the heterodimeric
SAXS/SANS data processing and overall parameters
 EMBO Global Exchange Lecture Course 30 November 2012 Hyderabad India SAXS/SANS data processing and overall parameters Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group
EMBO Global Exchange Lecture Course 30 November 2012 Hyderabad India SAXS/SANS data processing and overall parameters Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group
Measuring quaternary structure similarity using global versus local measures.
 Supplementary Figure 1 Measuring quaternary structure similarity using global versus local measures. (a) Structural similarity of two protein complexes can be inferred from a global superposition, which
Supplementary Figure 1 Measuring quaternary structure similarity using global versus local measures. (a) Structural similarity of two protein complexes can be inferred from a global superposition, which
Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and
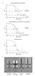 Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and comparisons to other published in situ T3SS structures. a, Resolution estimates after applying Fourier shell correlation
Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and comparisons to other published in situ T3SS structures. a, Resolution estimates after applying Fourier shell correlation
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. DOI: 10.1038/NCHEM.2675 Real-time molecular scale observation of crystal formation Roy E. Schreiber, 1 Lothar Houben, 2 Sharon G. Wolf, 2 Gregory Leitus,
In the format provided by the authors and unedited. DOI: 10.1038/NCHEM.2675 Real-time molecular scale observation of crystal formation Roy E. Schreiber, 1 Lothar Houben, 2 Sharon G. Wolf, 2 Gregory Leitus,
Supplementary Information. Cryo-EM Studies of Drp1 Reveal Cardiolipin Interactions that Activate
 Supplementary Information Cryo-EM Studies of Drp1 Reveal Cardiolipin Interactions that Activate the Helical Oligomer Christopher A. Francy 1, 2, 3, Ryan W. Clinton 1, 2, 3, Chris Fröhlich 4, 5, Colleen
Supplementary Information Cryo-EM Studies of Drp1 Reveal Cardiolipin Interactions that Activate the Helical Oligomer Christopher A. Francy 1, 2, 3, Ryan W. Clinton 1, 2, 3, Chris Fröhlich 4, 5, Colleen
Field-Flow Fractionation of Macromolecules and Structures That Cannot be Characterized by Conventional GPC/SEC Techniques
 The Field-Flow Fractionation Platform Field-Flow Fractionation of Macromolecules and Structures That Cannot be Characterized by Conventional GPC/SEC Techniques Trevor Havard, Evelin Moldenhaur, Soheyl
The Field-Flow Fractionation Platform Field-Flow Fractionation of Macromolecules and Structures That Cannot be Characterized by Conventional GPC/SEC Techniques Trevor Havard, Evelin Moldenhaur, Soheyl
Chris C. Broomell, Henrik Birkedal, Cristiano L. P. Oliveira, Jan Skov Pedersen, Jan-André Gertenbach, Mark Young, and Trevor Douglas*
 Chris C. Broomell, Henrik Birkedal, Cristiano L. P. Oliveira, Jan Skov Pedersen, Jan-André Gertenbach, Mark Young, and Trevor Douglas* Protein Cage Nanoparticles as Secondary Building Units for the Synthesis
Chris C. Broomell, Henrik Birkedal, Cristiano L. P. Oliveira, Jan Skov Pedersen, Jan-André Gertenbach, Mark Young, and Trevor Douglas* Protein Cage Nanoparticles as Secondary Building Units for the Synthesis
Impact of the crystallization condition on importin-β conformation
 Supporting information Volume 72 (2016) Supporting information for article: Impact of the crystallization condition on importin-β conformation Marcel J. Tauchert, Clément Hémonnot, Piotr Neumann, Sarah
Supporting information Volume 72 (2016) Supporting information for article: Impact of the crystallization condition on importin-β conformation Marcel J. Tauchert, Clément Hémonnot, Piotr Neumann, Sarah
SAXS Basics for BioSAXS. Robert P. Rambo Diamond Light Source B21
 SAXS Basics for BioSAXS Robert P. Rambo Diamond Light Source B21 Scattering and Size SAXS 0.11 nm (1.1 Å) smaller than C-C bond 6000x range 690 nm (6900 Å) 3.5x larger than E.coli 10x larger than a ribosome
SAXS Basics for BioSAXS Robert P. Rambo Diamond Light Source B21 Scattering and Size SAXS 0.11 nm (1.1 Å) smaller than C-C bond 6000x range 690 nm (6900 Å) 3.5x larger than E.coli 10x larger than a ribosome
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions
 Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions undergo endocytosis into clathrin-coated vesicles dependent
Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions undergo endocytosis into clathrin-coated vesicles dependent
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Data collection, phasing and refinement statistics ChbC/Ta 6 Br 12 Native ChbC Data collection Space group P4 3 2 1 2 P4 3 2 1 2 Cell dimensions a, c (Å) 132.75, 453.57 132.81, 452.95
Supplementary Table 1: Data collection, phasing and refinement statistics ChbC/Ta 6 Br 12 Native ChbC Data collection Space group P4 3 2 1 2 P4 3 2 1 2 Cell dimensions a, c (Å) 132.75, 453.57 132.81, 452.95
Tellurite resistance protein/ethidium efflux transporter/ proflavin transporter. Putative inner membrane protein: function unknown
 Additional file 1. Table S1 and Figures S1-4 of Zhang et al. High-level production of membrane proteins in E. coli BL21(DE3) by omitting the inducer IPTG Table S1. Properties of the membrane proteins used
Additional file 1. Table S1 and Figures S1-4 of Zhang et al. High-level production of membrane proteins in E. coli BL21(DE3) by omitting the inducer IPTG Table S1. Properties of the membrane proteins used
National de la Recherche Scientifique and Université Paris Descartes, Paris, France.
 FAST-RESPONSE CALMODULIN-BASED FLUORESCENT INDICATORS REVEAL RAPID INTRACELLULAR CALCIUM DYNAMICS Nordine Helassa a, Xiao-hua Zhang b, Ianina Conte a,c, John Scaringi b, Elric Esposito d, Jonathan Bradley
FAST-RESPONSE CALMODULIN-BASED FLUORESCENT INDICATORS REVEAL RAPID INTRACELLULAR CALCIUM DYNAMICS Nordine Helassa a, Xiao-hua Zhang b, Ianina Conte a,c, John Scaringi b, Elric Esposito d, Jonathan Bradley
BCM Protein crystallography - I. Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS
 BCM 6200 - Protein crystallography - I Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS What SAXS can do Small-angle X-ray scattering (SAXS) yields information on biological
BCM 6200 - Protein crystallography - I Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS What SAXS can do Small-angle X-ray scattering (SAXS) yields information on biological
BA, BSc, and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS
 Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS CNS input file generatemetal.inp: remarks file generate/generatemetal.inp
Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS CNS input file generatemetal.inp: remarks file generate/generatemetal.inp
Supplementary Figure 1 Mycobacterium tuberculosis WhiB1 expressed in Mycobacterium smegmatis possesses an O 2 -stable [4Fe-4S] cluster.
![Supplementary Figure 1 Mycobacterium tuberculosis WhiB1 expressed in Mycobacterium smegmatis possesses an O 2 -stable [4Fe-4S] cluster. Supplementary Figure 1 Mycobacterium tuberculosis WhiB1 expressed in Mycobacterium smegmatis possesses an O 2 -stable [4Fe-4S] cluster.](/thumbs/80/81296835.jpg) Supplementary Figure 1 Mycobacterium tuberculosis WhiB1 expressed in Mycobacterium smegmatis possesses an O 2 -stable [4Fe-4S] cluster. a, Absorbance spectra of WhiB1 isolated from recombinant M. smegmatis
Supplementary Figure 1 Mycobacterium tuberculosis WhiB1 expressed in Mycobacterium smegmatis possesses an O 2 -stable [4Fe-4S] cluster. a, Absorbance spectra of WhiB1 isolated from recombinant M. smegmatis
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11744 Supplementary Table 1. Crystallographic data collection and refinement statistics. Wild-type Se-Met-BcsA-B SmCl 3 -soaked EMTS-soaked Data collection Space
SUPPLEMENTARY INFORMATION doi:10.1038/nature11744 Supplementary Table 1. Crystallographic data collection and refinement statistics. Wild-type Se-Met-BcsA-B SmCl 3 -soaked EMTS-soaked Data collection Space
What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil
 Review R45 What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil Chaperonins form large, cage-like structures that act as protein folding machines. Recent developments in
Review R45 What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil Chaperonins form large, cage-like structures that act as protein folding machines. Recent developments in
Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS)
 nature methods Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS) Greg L Hura, Angeli L Menon, Michal Hammel, Robert P Rambo, Farris L Poole II, Susan E Tsutakawa,
nature methods Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS) Greg L Hura, Angeli L Menon, Michal Hammel, Robert P Rambo, Farris L Poole II, Susan E Tsutakawa,
Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits. AhpC and AhpF from Escherichia coli
 Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits AhpC and AhpF from Escherichia coli Phat Vinh Dip 1,#, Neelagandan Kamariah 2,#, Malathy Sony Subramanian Manimekalai
Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits AhpC and AhpF from Escherichia coli Phat Vinh Dip 1,#, Neelagandan Kamariah 2,#, Malathy Sony Subramanian Manimekalai
Electric-Field-Assisted Assembly of. Nanopores
 Supporting Information for: Electric-Field-Assisted Assembly of Polymer-Tethered Gold Nanorods in Cylindrical Nanopores Ke Wang, Seon-Mi Jin, Jiangping Xu, Ruijing Liang, Khurram Shezad, Zhigang Xue,*,
Supporting Information for: Electric-Field-Assisted Assembly of Polymer-Tethered Gold Nanorods in Cylindrical Nanopores Ke Wang, Seon-Mi Jin, Jiangping Xu, Ruijing Liang, Khurram Shezad, Zhigang Xue,*,
Sample characterization: Quality control and sample handling prior to data collection
 Sample characterization: Quality control and sample handling prior to data collection Marc JAMIN UMI 3265 UJF-EMBL-CNRS Unit of Virus Host Cell interactions Grenoble, France jamin@embl.fr Take home message
Sample characterization: Quality control and sample handling prior to data collection Marc JAMIN UMI 3265 UJF-EMBL-CNRS Unit of Virus Host Cell interactions Grenoble, France jamin@embl.fr Take home message
