SUPPLEMENTARY INFORMATION
|
|
|
- Muriel McCormick
- 5 years ago
- Views:
Transcription
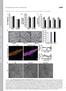
1 DOI:.38/ncb97 P ( μm, hours) P DMSO Figure S1 unningham et al. E MFN1 GFP-Parkin Opa1 ctin GPDH HEK293 GFP-Parkin Mitochondrial Fraction SH-SY5Y GFP-Parkin Mito DMSO Mito P yto DMSO SH-SY5Y ells DMSO 1 P 1 yto P and and and and D and and D F Relative bundance ytosol and 3 1 and and DMSO Mitochondria and DMSO Supplementary Figure 1 ubiquitin chains are enriched in HEK293 and SH-SY5Y whole cell lysates and mitochondria upon P treatment. () SH-SY5Y Western blot confirmation of mitochondrial depolarization upon P treatment over time. MFN1 levels decrease and Opa1 processing increased over the course of 4hr with μm P treatment in SH-SY5Y cells. () ase peak chromatograms of immunoaffinity purified K-GG peptides from SH-SY5Y cells following Parkin overexpression (oe) and/ or μm P treatment for 2hr. hromatographic features showing ubiquitin signature peptides for,,, and 3-linked chains are denoted by arrows. Inset shows extracted ion chromatograms from peptides in the treated and untreated samples. () SDS-PGE gel regions excised for ubiquitin linkage analysis in Figure S1D. HEK293 cells stably expressing GFP-Parkin were treated with either DMSO or μm P for 3hr, fractionated into mitochondria, and lysed for trypsinization P 3 3 P 3 3 / 3 Time (min) and and D ytosol and Mitochondria and Parkin oe ontrol Parkin oe +P 2hr and subsequent ubiquitin linkage analysis. Data represent one out of three independent experiments. (D) Ubiquitin linkage analysis on 4 gel regions ranging from 5-kDa. bundance (fmol) of each ubiquitin linkage is reported for DMSO (black circles) or P (red circles) treatment for each gel region. Data represent one out of three experiments. (E) Representative SDS-PGE gel showing excised regions for ubiquitin linkage analysis in Figure S1F. SH-SY5Y cells were treated with either DMSO or μm P for 3hr, fractionated into cytosolic and mitochondrial fractions, and lysed for trypsinization and subsequent ubiquitin linkage analysis. (F) Ubiquitin linkage analysis of cytosolic and mitochondrial fractions of 2 gel regions ranging from -kda. bundance (fmol) of each ubiquitin linkage is reported for DMSO (black circles) or P (red circles) treatment for each gel region from either the cytosolic or mitochondrial fractions of two replicate experiments in SH-SY5Y cells Macmillan Publishers Limited. ll rights reserved
2 OP1 Mfn1 GFP Tub ct GPDH Input + DMSO Input + P FT + P ug Input + P FT + P band 1 band fmol and PU (a.u.) Parkin Input Flow-thru and Supplementary Figure 2 Immunodepletion of GFP-Parkin confirms substrates other than Parkin carry -linked polyubiquitin chains. () HEK293 cells overexpressing GFP-Parkin were treated with um P or a DMSO vehicle control for 2hr. Lysates were immunodepleted with anti- GFP beads and input and flow through (FT) fractions were run on Western blot. MFN1 levels decreased and Opa1 processing increased confirming mitochondrial depolarization upon P treatment. This experiment was performed once. () Input and flow through (FT) fractions treated with P from () were run on SDS-PGE and excised into gel regions spanning 5-kDa. () Ubiquitin linkage analysis was used to measure the abundance of chains (fmol) for each gel region excised (left) and Parkin peptide abundance (U) was measured for each gel region excised (right) Macmillan Publishers Limited. ll rights reserved
3 v (nm/sec) USP2 USP UbM (μm) USP7 R eq (nm) oncentration ( μm) n.s. Monomer K6 K11 K27 K29 K48 K33 K63 Linear D kda 37 * 25 Ub 4 : ounts vs. Deconvoluted Mass (amu) E kda 1 75 F Fraction Top and Fraction ottom and Ratio (ottom/top) K K K N/ 37 K Top and ottom and Supplementary Figure 3 USP is an ubiquitin specific deubiquitinase that prefers compact chain types. () Michaelis-Menten plot of USP2 (blue), USP (orange), and USP7 (red) catalytic domain activity. Data were fit using the equation v = V max [S] /K M +[S]. () USP s preference for compact ubiquitin linkages is not driven by a hyper affinity for dimeric ubiquitin chain types in a biolayer interferometry experiment. Equilibrium response is shown as a function of di-ubiquitin concentration. ompact chains (,, and ) are shown in colors. Greater overall signal was observed with dimers, but individual kinetic curves displayed some non-specific adherence to the tip and fitting showed no significant (n.s.) decrease in dissociation constant (all K D s > 5μM). () Gel of,, and 3 purified tetramers. The lower band of (*) migrates most closely to 3 tetramers. (D) Intact mass spectrometry of tetramer shows a single molecular weight peak at the expected MW of tetrameric ubiquitin. (E) SDS-PGE gel of tetramers showing regions excised for ubiquitin linkage analysis. (F) Relative amounts of detected chain types for each excised band in (E). 3 make up a larger fraction of linkages in the lower band indicating this chain type as an off-target product during the enzymatic formation of K6 tetramers Macmillan Publishers Limited. ll rights reserved
4 E Figure S4 unningham et al. U Ratio (KO/WT) Relative bundance Relative bundance +P Usp (um) 1 15 kda HSPD1 WT TP5 USP_1 USP_2 USP_ KO Time (min) Time (min) Excised for linkage analysis Supplementary Figure 4 Mass spectrometry validation and ubiquitin linkage analysis of USP knockout cell line. (, ) Mass spectrometry confirmation of USP RISPR knockout. Ratios of area under the curve (U) between USP knockout HEK293 cells expressing GFP-Parkin and a wild type USP isogenic cell line for 2 mitochondrial matrix proteins (HSPD1, TP5) and three independent USP peptides (; see Table S2 for raw data). Ratios are not shown for the three USP signature peptides since no signal was detected in the USP knockout cell line, as represented in (). Error bars represent standard deviation from n=3 independent experiments (Table S2). () Representative SDS-PGE gel from D and and and and D HEK293 GFP-Parkin WT DMSO KO P WT P KO DMSO Mitochondrial Fraction WT P KO P and and and and D three independent experiments showing excision locations used for ubiquitin linkage analysis in Figures 3 and S3D. USP knockout (KO) or WT USP (WT) HEK293 cells stably expressing GFP-Parkin were treated with either DMSO or P (μm, 3hr), enriched for mitochondria, and lysed for trypsinization and subsequent ubiquitin linkage analysis. (D) Ubiquitin linkage analysis on 4 gel regions ranging from 5-kDa. fmol of each ubiquitin linkage is reported for P treated WT (black circles) or USP knockout (red circles) mitochondrial enriched fractions for each gel region. Data represent one out of two replicate experiments. (E) Representative SDS- PGE gel showing regions excised for ubiquitin linkage analysis in Figure Macmillan Publishers Limited. ll rights reserved
5 S U P P L E M E N T R Y I N F O R M T I O N P (2 hours, 5 μm) Doxycycline MFN1 GFP-Parkin Usp-3xFLG Opa1 ctin GPDH p-value 1e 1e 5 Experiment 1 MUL1 FKP8 TOM USP TDRKH MTX1 GDP1 VD3 VD ES Log2(USP + P vs. P alone) 3 Experiment 2 1e 1e 5 VD3 PS2 H12 TOM H31 ISD1 RL34 PN p-value.1.1 MID49 PSMG2 F188 SRSF9 USP MID49 VD3 PS2 H12 PSMG2 TOM F188 SRSF9 H31 ISD1 RL34 PN Log2(USP + P vs. P alone) Supplementary Figure 5 USP deubiquitinates outer membrane mitochondrial substrates. () Western blot confirmation of USP induction following doxycycline administration and mitochondrial depolarization upon P treatment. USP showed strong induction, while reduced MFN1 levels and increase in Opa1 processing were observed. Representative Western blot shown from experiment 1. (-) Volcano plots of ubiquitinated Figure S5 unningham et al. proteins identified in Experiments 1 and 2, respectively. The fold change of ubiquitination for each protein between ombo (USP overexpression and P) versus P is denoted on the x-axis with significance reported on the y-axis. oth values were determined based on the aggregate values of individual KGG peptides using LiME analysis. Proteins with p-value <.1 are labeled; mitochondria-associated proteins are colored red Macmillan Publishers Limited. ll rights reserved
6 DU-FLG HSP Transfection: TOM-MLS- TOM-MLS- TOM-MLS- TOM-MLS- I: USP-FLG USP-FLG USP2-FLG USP5-FLG TRID-FLG Transfection: GFP-Parkin + USP-FLG I: GFP-Parkin DU-FLG HSP TOM DU-FLG TOM-MLS USP-FLG HSP I: Transfection: GFP-Parkin + ß-Gal H-Ub wild type H-Ub K6R H-Ub K11R H-Ub K6R/K11R H-Ub K48R/K63R H-Ub KallR TOM-MLS USP2-FLG GFP- Parkin TOM-MLS TRID-FLG TOM TOM-MLS USP5-FLG H-Ub D * ß-Gal % of cells without mito (TOM staining) 24hr P ß-Gal WT Figure S6 unningham et al. Supplementary Figure 6 typical ubiquitin chain linkages signal for mitophagy. () Immunostaining of HeLa cells co-transfected with GFP- Parkin and mitochondrial-localized, FLG-tagged DUs. 24 hours after transfection cells were immunostained for FLG (green) and endogenous HSP (red). Each DU co-localizes with HSP indicating proper mitochondrial targeting (see merge). In addition, expression levels appear uniform across all DUs transfected. Representative cells are shown for each transfection. Scale bar, μm. () Large field images from Figure 5. Solid yellow outlines highlight cells with GFP-Parkin and DU-Flag expressing cells. Dashed yellow outlines highlight cells expressing only GFP- Parkin. Scale bar, μm. () Immunostaining of HeLa cells co-transfected with GFP-Parkin and H-tagged ubiquitin variants as listed in Figure 5. Only GFP-Parkin expressing cells are outlined. Solid lines indicate TOM positive staining cells; dashed lines indicate TOM negative staining cells. Scale bar, μm. (D) Quantification of TOM in HeLa cells treated with P (15μM, 24hr) and transfected with either beta-gal or H-ubiquitin. significant difference in TOM was observed with a 24 hour P treatment preventing analysis of individual H-ubiquitin mutants tested in Figure 5. Each point represents a single experiment with a mean intensity from 25- cells measured; the line represents the mean from n= 5 experiments and error bars represent S.E.M. Significance is reported as one-way NOV with Holm-Sidak s multiple comparison test (* p <.5) Macmillan Publishers Limited. ll rights reserved
7 Supplementary Figure 6 Full scans Macmillan Publishers Limited. ll rights reserved
8 Supplementary Table 1 Total peptide counts of selected proteins from isolated cytosolic (orange highlights) and mitochondrial fractions of HEK293T cells stably expressing GFP- Parkin confirm proper enrichment within each fraction. Upon P treatment Parkin is recruited to mitochondria and MFN1 and MFN2 are degraded. bbreviations are defined as: IMS inner membrane space; IM inner membrane; OM outer membrane. Supplementary Table 2 rea under the curve (U) measurements for a USP knockout HEK293 cell line expressing GFP-Parkin and a wild type USP isogenic cell line. Data are reported for two mitochondrial matrix proteins (HSPD1, STP5) and three independent USP peptides for three replicate experiments. Supplementary Table 3 Ubiquitin chain type analysis calculations for each chain type in four separate replicate experiments. ll amounts have been normalized by a panel of mitochondrial specific peptides to account for sample to sample variation. Supplementary Table 4 Mass spectrometry identified proteins after a K-GG immunoprecipitation (see methods) for two biological replicates. For each protein identified we are reporting the log2 fold change in ubiquitination during P treatment in the presence or absence of USP overexpression with the significance of the fold change (see Methods). Supplementary Table 5 Peptide abundance levels for proteins identified in the proteomic screen described in Figures 4, S4, and Table S Macmillan Publishers Limited. ll rights reserved
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with
 Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with DsRed-mito. Right panels are time-course enlarged images
Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with DsRed-mito. Right panels are time-course enlarged images
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Information
 Supplementary Information MAP2/Hoechst Hyp.-AP ph 6.5 Hyp.-SD ph 7.2 Norm.-SD ph 7.2 Supplementary Figure 1. Mitochondrial elongation in cortical neurons by acidosis. Representative images of neuronal
Supplementary Information MAP2/Hoechst Hyp.-AP ph 6.5 Hyp.-SD ph 7.2 Norm.-SD ph 7.2 Supplementary Figure 1. Mitochondrial elongation in cortical neurons by acidosis. Representative images of neuronal
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL Figure S1. Mitochondrial morphology in Fis1-null, Mff-null and Fis1/Mff-null MEF cells. (A) Western blotting of lysates from Fis1-null, Mff-null and Fis1/Mff-null cells. Lysates were
SUPPLEMENTAL MATERIAL Figure S1. Mitochondrial morphology in Fis1-null, Mff-null and Fis1/Mff-null MEF cells. (A) Western blotting of lysates from Fis1-null, Mff-null and Fis1/Mff-null cells. Lysates were
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
 DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
4) Please cite Dagda et al J Biol Chem 284: , for any publications or presentations resulting from use or modification of the macro.
 Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplementary Information
 Supplementary Information An engineered protein antagonist of K-Ras/B-Raf interaction Monique J. Kauke, 1,2 Michael W. Traxlmayr 1,2, Jillian A. Parker 3, Jonathan D. Kiefer 4, Ryan Knihtila 3, John McGee
Supplementary Information An engineered protein antagonist of K-Ras/B-Raf interaction Monique J. Kauke, 1,2 Michael W. Traxlmayr 1,2, Jillian A. Parker 3, Jonathan D. Kiefer 4, Ryan Knihtila 3, John McGee
Mitochondrial Dynamics Is a Distinguishing Feature of Skeletal Muscle Fiber Types and Regulates Organellar Compartmentalization
 Cell Metabolism Supplemental Information Mitochondrial Dynamics Is a Distinguishing Feature of Skeletal Muscle Fiber Types and Regulates Organellar Compartmentalization Prashant Mishra, Grigor Varuzhanyan,
Cell Metabolism Supplemental Information Mitochondrial Dynamics Is a Distinguishing Feature of Skeletal Muscle Fiber Types and Regulates Organellar Compartmentalization Prashant Mishra, Grigor Varuzhanyan,
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN
 THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
TNFα 18hr. Control. CHX 18hr. TNFα+ CHX 18hr. TNFα: 18 18hr (KDa) PARP. Cleaved. Cleaved. Cleaved. Caspase3. Pellino3 shrna. Control shrna.
 Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
FSC-W FSC-H CD4 CD62-L
 Supplementary Fig. 1 a SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H CD4 CD62-L b SSC-A FSC-A FSC-W FSC-A FSC-A 7-AAD FSC-A CD4 IL-9 CD4 c SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H 7-AAD KI67 Annexin-V 7-AAD d I L -5
Supplementary Fig. 1 a SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H CD4 CD62-L b SSC-A FSC-A FSC-W FSC-A FSC-A 7-AAD FSC-A CD4 IL-9 CD4 c SSC-A FSC-A FSC-W FSC-H SSC-W SSC-H 7-AAD KI67 Annexin-V 7-AAD d I L -5
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
SUPPLEMENTARY INFORMATION
 Supplementary Table S1 Kinetic Analyses of the AMSH-LP mutants AMSH-LP K M (μm) k cat x 10-3 (s -1 ) WT 71.8 ± 6.3 860 ± 65.4 T353A 76.8 ± 11.7 46.3 ± 3.7 F355A 58.9 ± 10.4 5.33 ± 0.30 proximal S358A 75.1
Supplementary Table S1 Kinetic Analyses of the AMSH-LP mutants AMSH-LP K M (μm) k cat x 10-3 (s -1 ) WT 71.8 ± 6.3 860 ± 65.4 T353A 76.8 ± 11.7 46.3 ± 3.7 F355A 58.9 ± 10.4 5.33 ± 0.30 proximal S358A 75.1
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
Supplementary material
 Supplementary material Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins Vladimir V. Rogov 1,*, Hironori Suzuki 2,3,*, Mija Marinković 4, Verena Lang
Supplementary material Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins Vladimir V. Rogov 1,*, Hironori Suzuki 2,3,*, Mija Marinković 4, Verena Lang
Supporting Information
 Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Structure and RNA-binding properties. of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex
 Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Waithe et al Supplementary Figures
 Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Role of Mitochondrial Remodeling in Programmed Cell Death in
 Developmental Cell, Vol. 12 Supplementary Data Role of Mitochondrial Remodeling in Programmed Cell Death in Drosophila melanogaster Gaurav Goyal, Brennan Fell, Apurva Sarin, Richard J. Youle, V. Sriram.
Developmental Cell, Vol. 12 Supplementary Data Role of Mitochondrial Remodeling in Programmed Cell Death in Drosophila melanogaster Gaurav Goyal, Brennan Fell, Apurva Sarin, Richard J. Youle, V. Sriram.
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Definition and assessment of ciap1 constructs.
 Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters
![Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters](/thumbs/89/100978514.jpg) Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters 20 February 2018
Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters 20 February 2018
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
A compound-based proteomic approach discloses 15-ketoatractyligenin. methyl ester as a new PPARγ partial agonist with anti-proliferative ability
 upplementary Information A compound-based proteomic approach discloses 15-ketoatractyligenin methyl ester as a new PPARγ partial agonist with anti-proliferative ability Michele Vasaturo 1, Lorenzo Fiengo
upplementary Information A compound-based proteomic approach discloses 15-ketoatractyligenin methyl ester as a new PPARγ partial agonist with anti-proliferative ability Michele Vasaturo 1, Lorenzo Fiengo
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Expanded View Figures
 The EMBO Journal Structure of a Dm peptide bound to the OT module Tobias Raisch et al Expanded View Figures A Hs Dm 262 297 685 8 HEAT HEAT MIF4G 9BD 1SHD 761 91 193 169 1152 1317 16 1376 1467 HEAT HEAT
The EMBO Journal Structure of a Dm peptide bound to the OT module Tobias Raisch et al Expanded View Figures A Hs Dm 262 297 685 8 HEAT HEAT MIF4G 9BD 1SHD 761 91 193 169 1152 1317 16 1376 1467 HEAT HEAT
Structural basis of PROTAC cooperative recognition for selective protein degradation
 SUPPLEMENTARY INFORMATION Structural basis of PROTAC cooperative recognition for selective protein degradation Morgan S. Gadd 1, Andrea Testa 1, Xavier Lucas 1, Kwok-Ho Chan, Wenzhang Chen, Douglas J.
SUPPLEMENTARY INFORMATION Structural basis of PROTAC cooperative recognition for selective protein degradation Morgan S. Gadd 1, Andrea Testa 1, Xavier Lucas 1, Kwok-Ho Chan, Wenzhang Chen, Douglas J.
downstream (0.8 kb) homologous sequences to the genomic locus of DIC. A DIC mutant strain (ro- 6
 A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and
 Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figures
 1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
Linear ubiquitination of cytosolic Salmonella Typhimurium activates NF-κB and restricts
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 2 ARTICLE NUMBER: 17066 Linear ubiquitination of cytosolic Salmonella Typhimurium activates NF-κB and restricts bacterial
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION VOLUME: 2 ARTICLE NUMBER: 17066 Linear ubiquitination of cytosolic Salmonella Typhimurium activates NF-κB and restricts bacterial
Figure S1: Extracellular nicotinic acid, but not tryptophan, is sufficient to maintain
 SUPPLEMENTAL INFORMATION Supplemental Figure Legends Figure S1: Extracellular nicotinic acid, but not tryptophan, is sufficient to maintain mitochondrial NAD +. A) Extracellular tryptophan, even at 5 µm,
SUPPLEMENTAL INFORMATION Supplemental Figure Legends Figure S1: Extracellular nicotinic acid, but not tryptophan, is sufficient to maintain mitochondrial NAD +. A) Extracellular tryptophan, even at 5 µm,
7.06 Problem Set
 7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
targets. clustering show that different complex pathway
 Supplementary Figure 1. CLICR allows clustering and activation of cytoplasmic protein targets. (a, b) Upon light activation, the Cry2 (red) and LRP6c (green) components co-cluster due to the heterodimeric
Supplementary Figure 1. CLICR allows clustering and activation of cytoplasmic protein targets. (a, b) Upon light activation, the Cry2 (red) and LRP6c (green) components co-cluster due to the heterodimeric
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11419 Supplementary Figure 1 Schematic representation of innate immune signaling pathways induced by intracellular Salmonella in cultured macrophages. a, During the infection Salmonella
doi:10.1038/nature11419 Supplementary Figure 1 Schematic representation of innate immune signaling pathways induced by intracellular Salmonella in cultured macrophages. a, During the infection Salmonella
Quantification of Protein Half-Lives in the Budding Yeast Proteome
 Supporting Methods Quantification of Protein Half-Lives in the Budding Yeast Proteome 1 Cell Growth and Cycloheximide Treatment Three parallel cultures (17 ml) of each TAP-tagged strain were grown in separate
Supporting Methods Quantification of Protein Half-Lives in the Budding Yeast Proteome 1 Cell Growth and Cycloheximide Treatment Three parallel cultures (17 ml) of each TAP-tagged strain were grown in separate
Supplementary Information. Recognition of the pre-mirna structure by Drosophila Dicer-1
 Supplementary Information Recognition of the pre-mirna structure by Drosophila Dicer-1 Akihisa Tsutsumi 1,2, Tomoko Kawamata 1, Natsuko Izumi 1 Hervé Seitz 3,4 and Yukihide Tomari 1,2 Nature Structural
Supplementary Information Recognition of the pre-mirna structure by Drosophila Dicer-1 Akihisa Tsutsumi 1,2, Tomoko Kawamata 1, Natsuko Izumi 1 Hervé Seitz 3,4 and Yukihide Tomari 1,2 Nature Structural
Reconstructing Mitochondrial Evolution?? Morphological Diversity. Mitochondrial Diversity??? What is your definition of a mitochondrion??
 Reconstructing Mitochondrial Evolution?? What is your definition of a mitochondrion?? Morphological Diversity Mitochondria as we all know them: Suprarenal gland Liver cell Plasma cell Adrenal cortex Mitochondrial
Reconstructing Mitochondrial Evolution?? What is your definition of a mitochondrion?? Morphological Diversity Mitochondria as we all know them: Suprarenal gland Liver cell Plasma cell Adrenal cortex Mitochondrial
The Fic protein Doc uses an inverted substrate to phosphorylate and. inactivate EF-Tu
 The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Breker et al., http://www.jcb.org/cgi/content/full/jcb.201301120/dc1 Figure S1. Single-cell proteomics of stress responses. (a) Using
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Breker et al., http://www.jcb.org/cgi/content/full/jcb.201301120/dc1 Figure S1. Single-cell proteomics of stress responses. (a) Using
Supplemental Information. Cand1-Mediated Adaptive Exchange Mechanism. Enables Variation in F-Box Protein Expression
 Molecular Cell, Volume 69 upplemental Information -Mediated Adaptive Exchange Mechanism Enables Variation in F-Box Protein Expression Xing Liu, Justin M. Reitsma, Jennifer L. Mamrosh, Yaru Zhang, Ronny
Molecular Cell, Volume 69 upplemental Information -Mediated Adaptive Exchange Mechanism Enables Variation in F-Box Protein Expression Xing Liu, Justin M. Reitsma, Jennifer L. Mamrosh, Yaru Zhang, Ronny
SUPPLEMENTARY INFORMATION
 GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
Heather Currinn, Benjamin Guscott, Zita Balklava, Alice Rothnie and Thomas Wassmer*
 Online Resources APP controls the formation of PI(3,5)P 2 vesicles through its binding of the PIKfyve complex. Cellular and Molecular Life Sciences Heather Currinn, Benjamin Guscott, Zita Balklava, Alice
Online Resources APP controls the formation of PI(3,5)P 2 vesicles through its binding of the PIKfyve complex. Cellular and Molecular Life Sciences Heather Currinn, Benjamin Guscott, Zita Balklava, Alice
Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures
 Nature Methods Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures Yannick Doyon, Thuy D Vo, Matthew C Mendel, Shon G Greenberg, Jianbin Wang, Danny F Xia, Jeffrey
Nature Methods Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures Yannick Doyon, Thuy D Vo, Matthew C Mendel, Shon G Greenberg, Jianbin Wang, Danny F Xia, Jeffrey
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
Supplemental Data. Chen and Thelen (2010). Plant Cell /tpc
 Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Nature Medicine: doi: /nm.3776
 C terminal Hsp90 inhibitors restore glucocorticoid sensitivity and relieve a mouse allograft model of Cushing s disease Mathias Riebold, Christian Kozany, Lee Freiburger, Michael Sattler, Michael Buchfelder,
C terminal Hsp90 inhibitors restore glucocorticoid sensitivity and relieve a mouse allograft model of Cushing s disease Mathias Riebold, Christian Kozany, Lee Freiburger, Michael Sattler, Michael Buchfelder,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
University of Groningen
 University of Groningen Role of mitochondrial inner membrane organizing system in protein biogenesis of the mitochondrial outer membrane Bohnert, Maria; Wenz, Lena-Sophie; Zerbes, Ralf M.; Horvath, Susanne
University of Groningen Role of mitochondrial inner membrane organizing system in protein biogenesis of the mitochondrial outer membrane Bohnert, Maria; Wenz, Lena-Sophie; Zerbes, Ralf M.; Horvath, Susanne
CHAPTER 1: ENZYME KINETICS AND APPLICATIONS
 CHAPTER 1: ENZYME KINETICS AND APPLICATIONS EM 1 2012/13 ERT 317 BIOCHEMICAL ENGINEERING Course details Credit hours/units : 4 Contact hours : 3 hr (L), 3 hr (P) and 1 hr (T) per week Evaluations Final
CHAPTER 1: ENZYME KINETICS AND APPLICATIONS EM 1 2012/13 ERT 317 BIOCHEMICAL ENGINEERING Course details Credit hours/units : 4 Contact hours : 3 hr (L), 3 hr (P) and 1 hr (T) per week Evaluations Final
Supplemental Data. Gao et al. (2012). Plant Cell /tpc
 Supplemental Figure 1. Plant EMP Proteins. (A) The Accession numbers of the 12 EMP members from Arabidopsis. (B) Phylogenetic analysis of EMP proteins from Arabidopsis, human and yeast using the Mac Vector
Supplemental Figure 1. Plant EMP Proteins. (A) The Accession numbers of the 12 EMP members from Arabidopsis. (B) Phylogenetic analysis of EMP proteins from Arabidopsis, human and yeast using the Mac Vector
Supplementary Figure 1 Analysis of beige fat and cells and characteristics of exosome release, related to Figure 1
 Supplementary Figure 1 Analysis of beige fat and cells and characteristics of exosome release, related to Figure 1 (a) Fold-change in UCP-1 mrna abundance in white adipocytes upon β-adrenergic stimulation
Supplementary Figure 1 Analysis of beige fat and cells and characteristics of exosome release, related to Figure 1 (a) Fold-change in UCP-1 mrna abundance in white adipocytes upon β-adrenergic stimulation
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R)
 Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1
 Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Yeast Genome-wide Screens to Ascertain the Genetic Landscape for Barth Syndrome. Christopher R. McMaster, PhD Dalhousie University
 Yeast Genome-wide Screens to Ascertain the Genetic Landscape for Barth Syndrome Christopher R. McMaster, PhD Dalhousie University Using Systematic Genetics to Identify Modifies Genes that Affect Fitness
Yeast Genome-wide Screens to Ascertain the Genetic Landscape for Barth Syndrome Christopher R. McMaster, PhD Dalhousie University Using Systematic Genetics to Identify Modifies Genes that Affect Fitness
Supplementary Information. Overlap between folding and functional energy landscapes for. adenylate kinase conformational change
 Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10244 a O07391_MYCAV/127-243 NLPC_HAEIN/80-181 SPR_SHIFL/79-183 P74160_SYNY3/112-245 O24914_HELPY/301-437 Q51835_PORGI/68-178 DPP6_BACSH/163-263 YKFC_BACSU/185-292 YDHO_ECOLI/153-263
doi:10.1038/nature10244 a O07391_MYCAV/127-243 NLPC_HAEIN/80-181 SPR_SHIFL/79-183 P74160_SYNY3/112-245 O24914_HELPY/301-437 Q51835_PORGI/68-178 DPP6_BACSH/163-263 YKFC_BACSU/185-292 YDHO_ECOLI/153-263
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Watanabe'et'al.' Supplementary'Figure'1'
 MW IN O 80 2 IN O 80 SW 3 R -C 3 MW SW R -C SW 1 R -C 2 IN O 80 1 Supplementary'Figure'1' Supplementary Figure 1. SDS-PAGE image of INO80-C and SWR-C. Three independent preparations of each TAP-tagged
MW IN O 80 2 IN O 80 SW 3 R -C 3 MW SW R -C SW 1 R -C 2 IN O 80 1 Supplementary'Figure'1' Supplementary Figure 1. SDS-PAGE image of INO80-C and SWR-C. Three independent preparations of each TAP-tagged
Lecture 13: Data Analysis and Interpretation of the Michaelis-Menten Parameters
 Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2019 Lecture 13: Data Analysis and Interpretation of the Michaelis-Menten Parameters 19 February 2019 c David P. Goldenberg University
Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2019 Lecture 13: Data Analysis and Interpretation of the Michaelis-Menten Parameters 19 February 2019 c David P. Goldenberg University
Tracking Protein Allostery in Evolution
 Tracking Protein Allostery in Evolution Glycogen phosphorylase frees sugars to provide energy GP orthologs diverged 600,000,000 years can respond to transcription controls, metabolite concentrations and
Tracking Protein Allostery in Evolution Glycogen phosphorylase frees sugars to provide energy GP orthologs diverged 600,000,000 years can respond to transcription controls, metabolite concentrations and
Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises
 Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises six Orai subunits, each with identical amino acid sequences
Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises six Orai subunits, each with identical amino acid sequences
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10458 Active Site Remodeling in the Bifunctional Fructose-1,6- bisphosphate aldolase/phosphatase Juan Du, Rafael F. Say, Wei Lü, Georg Fuchs & Oliver Einsle SUPPLEMENTARY FIGURES Figure
doi:10.1038/nature10458 Active Site Remodeling in the Bifunctional Fructose-1,6- bisphosphate aldolase/phosphatase Juan Du, Rafael F. Say, Wei Lü, Georg Fuchs & Oliver Einsle SUPPLEMENTARY FIGURES Figure
Membrane depolarization activates the mitochondrial protease OMA1 by stimulating self-cleavage
 Scientific Report Membrane depolarization activates the mitochondrial protease OMA1 by stimulating self-cleavage Kuan Zhang, Huihui Li & Zhiyin Song* Abstract Mitochondrial inner membrane fusion depends
Scientific Report Membrane depolarization activates the mitochondrial protease OMA1 by stimulating self-cleavage Kuan Zhang, Huihui Li & Zhiyin Song* Abstract Mitochondrial inner membrane fusion depends
UCD Conway Institute of Biomolecular & Biomedical Research Graduate Education 2009/2010
 EMERGING PROTEOMIC TECHNOLOGIES - MODULE SCHEDULE & OUTLINE 2010 Course Organiser: Dr. Giuliano Elia Module Co-ordinator: Dr Giuliano Elia Credits: 5 Date & Time Session & Topic Coordinator 14th April
EMERGING PROTEOMIC TECHNOLOGIES - MODULE SCHEDULE & OUTLINE 2010 Course Organiser: Dr. Giuliano Elia Module Co-ordinator: Dr Giuliano Elia Credits: 5 Date & Time Session & Topic Coordinator 14th April
Biological Roles of Cytokinins
 Direct Control of Shoot Meristem Activity by a Cytokinin-Activating Enzyme By Kurakawa et. Al. Published in Nature Presented by Boyana Grigorova Biological Roles of Cytokinins Cytokinins are positive regulators
Direct Control of Shoot Meristem Activity by a Cytokinin-Activating Enzyme By Kurakawa et. Al. Published in Nature Presented by Boyana Grigorova Biological Roles of Cytokinins Cytokinins are positive regulators
The Role of GRASP65 in Golgi Cisternal Stacking and Cell Cycle Progression
 Traffic 2010; 11: 827 842 2010 John Wiley & Sons A/S doi:10.1111/j.1600-0854.2010.01055.x The Role of GRASP65 in Golgi Cisternal Stacking and Cell Cycle Progression Danming Tang, Hebao Yuan and Yanzhuang
Traffic 2010; 11: 827 842 2010 John Wiley & Sons A/S doi:10.1111/j.1600-0854.2010.01055.x The Role of GRASP65 in Golgi Cisternal Stacking and Cell Cycle Progression Danming Tang, Hebao Yuan and Yanzhuang
Biophysics Service at the MPIB Biochemistry Core Facility Stephan Uebel, Biochemistry Core Facility
 Biophysics Service at the MPIB Biochemistry Core Facility 30.11.2015 Stephan Uebel, Biochemistry Core Facility uebel@biochem.mpg.de Overview Peptide Chemistry - Peptide synthesis -Amino acid analysis -
Biophysics Service at the MPIB Biochemistry Core Facility 30.11.2015 Stephan Uebel, Biochemistry Core Facility uebel@biochem.mpg.de Overview Peptide Chemistry - Peptide synthesis -Amino acid analysis -
Absorbance (a. u.) Wavelength (nm) Wavelength (nm) Intensity (a. u.) Wavelength (nm) Wavelength (nm)
 Intensity (a. u.) Absorbance (a. u.) a UV UV b Wavelength (nm) Wavelength (nm) UV UV Wavelength (nm) Wavelength (nm) Supplementary Figure 1. UV-Vis absorbance spectral changes of (a) SP-Gal (left, 10 μm),
Intensity (a. u.) Absorbance (a. u.) a UV UV b Wavelength (nm) Wavelength (nm) UV UV Wavelength (nm) Wavelength (nm) Supplementary Figure 1. UV-Vis absorbance spectral changes of (a) SP-Gal (left, 10 μm),
Supplemental Information
 Supplemental Information Figure S. Regions recognized by the specific antibodies. The amino acid sequence of the major p3-alc species, p3-alc α 35, p3-alc β 37 and p3-alc γ 3, are indicated as red letters.
Supplemental Information Figure S. Regions recognized by the specific antibodies. The amino acid sequence of the major p3-alc species, p3-alc α 35, p3-alc β 37 and p3-alc γ 3, are indicated as red letters.
MS-based proteomics to investigate proteins and their modifications
 MS-based proteomics to investigate proteins and their modifications Francis Impens VIB Proteomics Core October th 217 Overview Mass spectrometry-based proteomics: general workflow Identification of protein
MS-based proteomics to investigate proteins and their modifications Francis Impens VIB Proteomics Core October th 217 Overview Mass spectrometry-based proteomics: general workflow Identification of protein
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/9/e1500511/dc1 Supplementary Materials for Contractility parameters of human -cardiac myosin with the hypertrophic cardiomyopathy mutation R403Q show loss of
advances.sciencemag.org/cgi/content/full/1/9/e1500511/dc1 Supplementary Materials for Contractility parameters of human -cardiac myosin with the hypertrophic cardiomyopathy mutation R403Q show loss of
Journal of Cell Science Accepted manuscript
 2015. Published by The Company of Biologists Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/3.0), which
2015. Published by The Company of Biologists Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/3.0), which
Nature Methods: doi: /nmeth Supplementary Figure 1. In vitro screening of recombinant R-CaMP2 variants.
 Supplementary Figure 1 In vitro screening of recombinant R-CaMP2 variants. Baseline fluorescence compared to R-CaMP1.07 at nominally zero calcium plotted versus dynamic range ( F/F) for 150 recombinant
Supplementary Figure 1 In vitro screening of recombinant R-CaMP2 variants. Baseline fluorescence compared to R-CaMP1.07 at nominally zero calcium plotted versus dynamic range ( F/F) for 150 recombinant
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/5/e1602426/dc1 Supplementary Materials for In vivo mapping of tissue- and subcellular-specific proteomes in Caenorhabditis elegans The PDF file includes: Aaron
advances.sciencemag.org/cgi/content/full/3/5/e1602426/dc1 Supplementary Materials for In vivo mapping of tissue- and subcellular-specific proteomes in Caenorhabditis elegans The PDF file includes: Aaron
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/1/e1500989/dc1 Supplementary Materials for An epidermis-driven mechanism positions and scales stem cell niches in plants Jérémy Gruel, Benoit Landrein, Paul Tarr,
advances.sciencemag.org/cgi/content/full/2/1/e1500989/dc1 Supplementary Materials for An epidermis-driven mechanism positions and scales stem cell niches in plants Jérémy Gruel, Benoit Landrein, Paul Tarr,
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.1244624/dc1 Supplementary Materials for Cytoneme-Mediated Contact-Dependent Transport of the Drosophila Decapentaplegic Signaling Protein Sougata Roy, Hai Huang,
www.sciencemag.org/cgi/content/full/science.1244624/dc1 Supplementary Materials for Cytoneme-Mediated Contact-Dependent Transport of the Drosophila Decapentaplegic Signaling Protein Sougata Roy, Hai Huang,
NB-DNJ/GCase-pH 7.4 NB-DNJ+/GCase-pH 7.4 NB-DNJ+/GCase-pH 4.5
 SUPPLEMENTARY TABLES Suppl. Table 1. Protonation states at ph 7.4 and 4.5. Protonation states of titratable residues in GCase at ph 7.4 and 4.5. Histidine: HID, H at δ-nitrogen; HIE, H at ε-nitrogen; HIP,
SUPPLEMENTARY TABLES Suppl. Table 1. Protonation states at ph 7.4 and 4.5. Protonation states of titratable residues in GCase at ph 7.4 and 4.5. Histidine: HID, H at δ-nitrogen; HIE, H at ε-nitrogen; HIP,
Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element
 Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element as determined in a previous study 1. The log K value
Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element as determined in a previous study 1. The log K value
Supplementary Information
 Supplementary Information Structural analysis of leader peptide binding enables leaderfree cyanobactin processing Jesko Koehnke 1,2, Greg Mann 1,2, Andrew F Bent 1,2, Hannes Ludewig 1, Sally Shirran 1,
Supplementary Information Structural analysis of leader peptide binding enables leaderfree cyanobactin processing Jesko Koehnke 1,2, Greg Mann 1,2, Andrew F Bent 1,2, Hannes Ludewig 1, Sally Shirran 1,
Supporting Information
 Supporting Information Wiley-VCH 2012 69451 Weinheim, Germany Bound Cations Significantly Stabilize the Structure of Multiprotein Complexes in the Gas Phase** Linjie Han, Suk-Joon Hyung, and Brandon T.
Supporting Information Wiley-VCH 2012 69451 Weinheim, Germany Bound Cations Significantly Stabilize the Structure of Multiprotein Complexes in the Gas Phase** Linjie Han, Suk-Joon Hyung, and Brandon T.
Quantitation of a target protein in crude samples using targeted peptide quantification by Mass Spectrometry
 Quantitation of a target protein in crude samples using targeted peptide quantification by Mass Spectrometry Jon Hao, Rong Ye, and Mason Tao Poochon Scientific, Frederick, Maryland 21701 Abstract Background:
Quantitation of a target protein in crude samples using targeted peptide quantification by Mass Spectrometry Jon Hao, Rong Ye, and Mason Tao Poochon Scientific, Frederick, Maryland 21701 Abstract Background:
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Previous Class. Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements. Today
 Previous Class Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements Today Spectrophotometry Spectrofluorimetry Radioactive Procedures ph dependency Spectrophotometry
Previous Class Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements Today Spectrophotometry Spectrofluorimetry Radioactive Procedures ph dependency Spectrophotometry
Supporting information for
 Supporting information for Rewiring multi-domain protein switches: transforming a fluorescent Zn 2+ -sensor into a light-responsive Zn 2+ binding protein Stijn J.A. Aper and Maarten Merkx Laboratory of
Supporting information for Rewiring multi-domain protein switches: transforming a fluorescent Zn 2+ -sensor into a light-responsive Zn 2+ binding protein Stijn J.A. Aper and Maarten Merkx Laboratory of
Previous Class. Today. Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements
 Previous Class Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements Today Practical Methods for Kinetics and Equilibria Spectrophotometry Radioactive Procedures Spectrofluorimetry
Previous Class Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements Today Practical Methods for Kinetics and Equilibria Spectrophotometry Radioactive Procedures Spectrofluorimetry
Supplementary figure 1. Comparison of unbound ogm-csf and ogm-csf as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine
 Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
JMJ14-HA. Col. Col. jmj14-1. jmj14-1 JMJ14ΔFYR-HA. Methylene Blue. Methylene Blue
 Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
Supplementary Tables and Figures
 Supplementary Tables Supplementary Tables and Figures Supplementary Table 1: Tumor types and samples analyzed. Supplementary Table 2: Genes analyzed here. Supplementary Table 3: Statistically significant
Supplementary Tables Supplementary Tables and Figures Supplementary Table 1: Tumor types and samples analyzed. Supplementary Table 2: Genes analyzed here. Supplementary Table 3: Statistically significant
