Supplemental Materials Molecular Biology of the Cell
|
|
|
- Shawn Reeves
- 5 years ago
- Views:
Transcription
1 Supplemental Materials Molecular iology of the Cell Figure S1 Krüger et al. Arabidopsis Plasmodium H. sapiens* 1) Xenopus* 1) Drosophila C.elegans S.cerevisae S.pombe L.major T.cruzi T.brucei DCP5 CITH RAP55/Lsm14 RAP55/Lsm14 Tra1 (Trailer hitch) Car1 Sum2 Lsm domain DFDF FFD-TFG all Q and N outside other domains amino acids RGG *1) isoform a Figure S1: Schematics of orthologues to /RAP55 from various organisms Domains, RGG motifs and Q or N residues outside of other domains are marked.
2 Figure S2 A kda / + -eyfp / - anti- -eyfp / + -eyfp cummulative cell density [cells/ml] 1* *10 9 1*10 7 1* time [h] +/+ -eyfp / - Figure S2: -eyfp is functional A cell line was made that had one allele replaced by a blasticidin resistence gene using 5 and 3 UTR sequences for homologous recombination. The second allele was endogenously tagged with a C-terminal eyfp tag, as described in Kelly et al., (A) Western blot with cell lysates of the cell lines + / +, -eyfp/- and -eyfp / -. () Growth of the - eyfp/- cells in comparison to wild type cells ( +/+) Kelly, S., Reed, J., Kramer, S., Ellis, L., Webb, H., Sunter, J., Salje, J., Marinsek, N., Gull, K., Wickstead,. et al. (2007). Functional genomics in Trypanosoma brucei: a collection of vectors for the expression of tagged proteins from endogenous and ectopic gene loci. Mol iochem Parasitol 154,
3 A Figure S3 eyfp DAPI eyfp DAPI no expression of (0 hours TET) mchfp-dhh1 expression of (24 hours TET) mchfp-dhh1 10 µm Figure S3: A) Inducible expression of eyfp in procyclic trypanosomes does not result in the formation of granules. The total fluorescence intensity of this particular cell was and therefore well above the treshhold expression level required for granule formation at inducible -eyfp overexpression. ) Inducible expression of (without eyfp tag) causes recruitment of mchfp-dhh1 to granules in a similar way as does the expression of -eyfp.
4 Figure S4 cell lysates (No. of cells) recombinant (ng) 5*10 5 1* anti Figure S4: A titration of recombinant protein purified from E. coli was used to estimate the amount of molecules in a procyclic trypanosome cell. There are about 3 ng in 1x10 6 cells, equivalent to ~60,000 molecules per cell.
5 A -eyfp DIC normal morphology Figure S5 high expression of -eyfp low expression of -eyfp 1K2N pointed posterior end 1K2N 1K0N 1K0N D 100% 80% 60% 40% 20% 0% TET (h) N>400 for each time-point normal division (no TET) 2K2N 1K1N others 1K2N 1K0N 2K2N 2K1N 1K1N C percentage pointed posterior end K1N 2K1N 2K2N N>20 for each time-point cells with a division furrow resulting in a daughter cell with abnormal karyotype + 1K1N 2K1N with division furrow (abnormal) 1K1N with dividing nucleus (abnormal) 1K0N (abnormal) + impaired cell divisions (24 h TET) 2K2N with nuclei in wrong positions 1K2N (abnormal) 1K0N (abnormal) + Figure S5: Over-expression of -eyfp results in a block in mitosis Over-expression of -eyfp causes a change in cell morphology and an increase in cells with abnormal karyotypes. Cells become pointed at both ends (A) and an increasing number of cells has abnormal karyotypes (A and ). After mitosis, procyclic trypanosomes have one nucleus and one kinetoplast (1K1N), then first divide the kinetoplast (2K1N) and subsequently the nucleus (2K2N) that will move in between the two kinetoplasts to allow longitudal cell division. On over-expression of -eyfp, abnormal 1K0N and 1K2N cells increased (A and ). In addition, the majority of cells with normal karyotypes were abnormal in size and shape of the nuclei and position/presence of the division furrow: 1K1N cells often had a large and bilobed nucleus and sometimes a division furrow; the majority of 2K1N cells had large nuclei and a division furrow in a position to produce a 1K0N cell as well as a 1K1N cells with a bilobed nucleus; many of the 2K2N were about to divide into a 1K2N and 1K0N cell and often nuclei did not become separated and appeared different in size. In fact, more than half of all 2K1N and 2K2N cells with a division furrow present had the division furrow wrongly positioned (C). Taken together, the data can be explained with untimed cell division prior to either the division of the nucleus or the correct positioning of the two daughter nuclei: 2K1N cells divide to a 1K0N cell and a 1K1N cell with a dividing nucleus and 2K2N cells give rise to 1K0N and 1K2N cells (D). (A) Typical fluorescent microscopy image (single plane) at 24 hours tetracyclin induction showing two cells with high - eyfp expression (including one with the abnormal 1K2N karyotype), one cell with low -eyfp expression and one 1K0N cell. lack arrows in the DIC image point to the spindle-shaped cell poles that are absent in the cell with low expression level. () Karyotype analysis over a time-course of induction of -eyfp over-expression. Normal karyotypes are shown in three different shades of grey, abnormal karyotypes in colour. (C) Percentage of cells with normal karyotypes that had a division furrow resulting in daughter cells with abnormal karyotypes. The analysis was done after 24 hours induction with tetracyclin. (D) Two models of an asymmetric cell division caused by -eyfp overexpression that are in agreement with the data. A normal cell division is shown on top. All images were taken using a Zeiss Axioskop microscope equipped with a Plan- Apochromat 100x/1.4 Oil DIC objective and a monochrome CCD camera AxioCam MR.
6 Figure S6 no TET TET (expression of -CerFP) PAP2 PAP2 eif2α eif2α eif4e1 eif4e1 eif4e3 eif4e3 eif4e4 eif4e4 eif4g1 eif4g1 eif4g2 eif4g2 eif4g3 eif4g3 eif4g4 eif4g4 eif4g5 eif4g5 eyfp DAPI eyfp CerFP Figure S6: Localization of other proteins to induced granules Trypanosome homologues to translation initiation factors were expressed as eyfp fusion proteins from their endogenous loci in a cell line engineered to over-express -CerFP at induction with tetracyclin (TET). Fluorescent microscopy images (Z-stack projection of deconvolved Z-stacks, sum slices) are shown for cells not expressing -CerFP (no TET) and after 24 hours of induction.
7 A Lsm14 N-rich C...MYSGG DHH1 merged Figure S7 sinefungin single plane C DHH1 merged starvation Lsm14 N-rich...NNNMR C* DHH1 merged sinefungin single plane C* DHH1 merged starvation C MPQSP... RGG FDF-TFG RGG sinefungin N single plane DHH1 merged Figure S7: Cells expressing a C-terminally or N-terminally truncated mutant as indicated for 24 hours were treated with sinefungin (60 min) or starvation (120 min PS). Projections of deconvolved Z-stacks of one representative cell, or single plane images are shown. N DHH1 merged starvation
8 A Figure S8 RNAi over-expression RNAi + over-expression clone 1 clone 2 kda h TET PFR (loading) -eyfp PFR (loading) C-eYFP anti- and anti-pfr -eyfp over-expression mchfp- DHH1 merged RNAi + over-expression -eyfp mchfp- DHH1 merged C-eYFP mchfp- DHH1 merged C-eYFP mchfp- DHH1 merged Figure S8: Simultanous RNAi knock-down of and over-expression of -eyfp or C-eYFP Procyclic trypanosome cell lines were engineered for tetracyclin induced RNAi knock-down of, tetracyclin induced over-expression of either -eyfp or C-eYFP as well as both RNAi and over-expression together. Part of the 3 UTR sequence of was used as RNAi target; the over-expression vectors contain the modified aldolase 3 UTR (Sunter et. al., ), and expression is therefore not affected by RNAi. (A) Western blot with cell lysates taken over a time-course of RNAi, over-expression or RNAi + over-expression probed with a polyclonal serum rised against (Kramer et al., 2008); results from two different clones are shown. PFR served as loading control. The polyclonal antiserume preferentially recognizes the C-terminal part of, thus, the apparent lower expression level of the C-eYFP protein in comparison to the -eyfp protein is not real; when expression levels are compared using anti-gfp, expression of C-eYFP is in fact higher than the expression of -eyfp (data not shown here). () Single plane fluorescence microscopy images of cells overexpressing -eyfp or C-eYFP in the absence or presence of RNAi knock-down of the endogenous. All images were taken using a Zeiss Axioskop microscope equipped with a Plan-Apochromat 100x/1.4 Oil DIC objective and a monochrome CCD camera AxioCam MR. Sunter, J., Wickstead,., Gull, K. and Carrington, M. (). A new generation of T7 RNA polymerase-independent inducible expression plasmids for Trypanosoma brucei. PLoS One 7, e Kramer, S., Queiroz, R., Ellis, L., Webb, H., Hoheisel, J. D., Clayton, C. and Carrington, M. (2008). Heat shock causes a decrease in polysomes and the appearance of stress granules in trypanosomes independently of eif2(alpha) phosphorylation at Thr169. J Cell Sci 121,
9 Table S1: Comprehensive list of plasmids used in this work (including supplementary figures) Name of plasmid ID of the trypanosome protein Name of protein Description ased on mother plasmid Reference for mother plasmid Selection 3924 Tb eyfp Inducible overexpression 3888 Sunter et al., SD 3342 Tb PAP2-mChFP Endogenous expression 3086 (=2705 with Hygro) / Hygro 3925 Tb mchfp-dhh1 Endogenous expression 2679 Kelly 2007 Puromycin SK85 Tb eif4e1-mchfp Endogenous expression 3086 (=2705 with Hygro) / Hygro 2792 Tb eyfp Endogenous expression 2710 Kelly 2007 NEO 2913 Tb Deletion of one / / SD allele by replacing the ORF by SD cassette 3351 Tb PAP2-eYFP Endogenous expression 3345 / Puro SK53 Tb eyfp-eif2a Endogenous expression / / Puro SK54 Tb eif4e1-eyfp Endogenous expression 2948 / Hygro SK56 Tb eif4e3-eyfp Endogenous expression 2948 / Hygro SK57 Tb eif4e4-eyfp Endogenous expression 2948 / Hygro SK60 Tb eif4g1-eyfp Endogenous expression 2948 / Hygro SK62 Tb eif4g2-eyfp Endogenous expression 2948 / Hygro SK58 Tb eif4g3-eyfp Endogenous expression 2948 / Hygro SK64 Tb eif4g4-eyfp Endogenous expression 2948 / Hygro SK66 Tb eif4g5-eyfp Endogenous expression 2948 / Hygro SK69 Tb CerFP Inducible overexpression 4256 (=3927 with CerFP) Sunter et al., SD 4209, Tb and Inducible overexpression 3888 Sunter et al., SD 4191, 4009, SK29, SK23, SK27, SK25, 4260, 4175, 4007, 4008, 4176, 4010 of truncations (see Figure 6 for details; plasmid numbers are in the same order as in Figure 6) 4296 Tb NLSeYFP Inducible overexpression 4259 (=3888 with 2 NLS) / SD 4297 Tb C- Inducible overexpression 4259 (=3888 with 2 NLS) / SD 2NLS-eYFP SK91 Tb eif4e1-nls- Inducible overexpression SK89 (=4256 (=3927 with / Puro CerFP CerFP) with one NLS) 4286 Tb RNAi RNAi, targeting the Sunter et al., SD UTR of 4287 Tb eyfp Inducible overexpression 4275 (=Hygro in 3888) / Hygro SK122 eyfp Inducible overexpression 3383 Sunter et al., SD SK123 Tb Inducible overexpression 3383 Sunter et al., SD
TNFα 18hr. Control. CHX 18hr. TNFα+ CHX 18hr. TNFα: 18 18hr (KDa) PARP. Cleaved. Cleaved. Cleaved. Caspase3. Pellino3 shrna. Control shrna.
 Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Seminar 1 Components and Regulation of Initiation of Translation
 Institut für Biochemie und Molekulare Medizin Seminar 1 Components and Regulation of Initiation of Translation Michael Altmann FS 2015 Seminar 1 - What are the biol. consequences of mrna transport and
Institut für Biochemie und Molekulare Medizin Seminar 1 Components and Regulation of Initiation of Translation Michael Altmann FS 2015 Seminar 1 - What are the biol. consequences of mrna transport and
JMJ14-HA. Col. Col. jmj14-1. jmj14-1 JMJ14ΔFYR-HA. Methylene Blue. Methylene Blue
 Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Dictyostelium Aurora Kinase Has Properties of both Aurora A and Aurora B Kinases
 EUKARYOTIC CELL, May 2008, p. 894 905 Vol. 7, No. 5 1535-9778/08/$08.00 0 doi:10.1128/ec.00422-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Dictyostelium Aurora Kinase Has
EUKARYOTIC CELL, May 2008, p. 894 905 Vol. 7, No. 5 1535-9778/08/$08.00 0 doi:10.1128/ec.00422-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Dictyostelium Aurora Kinase Has
In order to confirm the contribution of diffusion to the FRAP recovery curves of
 Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
4) Please cite Dagda et al J Biol Chem 284: , for any publications or presentations resulting from use or modification of the macro.
 Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Topic 3: Genetics (Student) Essential Idea: Chromosomes carry genes in a linear sequence that is shared by members of a species.
 Topic 3: Genetics (Student) 3.2 Essential Idea: Chromosomes carry genes in a linear sequence that is shared by members of a species. 3.2 Chromosomes 3.2.U1 Prokaryotes have one chromosome consisting of
Topic 3: Genetics (Student) 3.2 Essential Idea: Chromosomes carry genes in a linear sequence that is shared by members of a species. 3.2 Chromosomes 3.2.U1 Prokaryotes have one chromosome consisting of
CONJOINT 541. Translating a Transcriptome at Specific Times and Places. David Morris. Department of Biochemistry
 CONJOINT 541 Translating a Transcriptome at Specific Times and Places David Morris Department of Biochemistry http://faculty.washington.edu/dmorris/ Lecture 1 The Biology and Experimental Analysis of mrna
CONJOINT 541 Translating a Transcriptome at Specific Times and Places David Morris Department of Biochemistry http://faculty.washington.edu/dmorris/ Lecture 1 The Biology and Experimental Analysis of mrna
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Topic 8 Mitosis & Meiosis Ch.12 & 13. The Eukaryotic Genome. The Eukaryotic Genome. The Eukaryotic Genome
 Topic 8 Mitosis & Meiosis Ch.12 & 13 The Eukaryotic Genome pp. 244-245,268-269 Genome All of the genes in a cell. Eukaryotic cells contain their DNA in long linear pieces. In prokaryotic cells, there is
Topic 8 Mitosis & Meiosis Ch.12 & 13 The Eukaryotic Genome pp. 244-245,268-269 Genome All of the genes in a cell. Eukaryotic cells contain their DNA in long linear pieces. In prokaryotic cells, there is
 DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Structure and RNA-binding properties. of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex
 Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
SUPPLEMENTARY INFORMATION
 med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
A synthetic oscillatory network of transcriptional regulators
 A synthetic oscillatory network of transcriptional regulators Michael B. Elowitz & Stanislas Leibler, Nature, 403, 2000 igem Team Heidelberg 2008 Journal Club Andreas Kühne Introduction Networks of interacting
A synthetic oscillatory network of transcriptional regulators Michael B. Elowitz & Stanislas Leibler, Nature, 403, 2000 igem Team Heidelberg 2008 Journal Club Andreas Kühne Introduction Networks of interacting
Supplementary Materials for
 www.advances.sciencemag.org/cgi/content/full/1/5/e1500358/dc1 Supplementary Materials for Transplantability of a circadian clock to a noncircadian organism Anna H. Chen, David Lubkowicz, Vivian Yeong,
www.advances.sciencemag.org/cgi/content/full/1/5/e1500358/dc1 Supplementary Materials for Transplantability of a circadian clock to a noncircadian organism Anna H. Chen, David Lubkowicz, Vivian Yeong,
Reading: Chapter 5, pp ; Reference chapter D, pp Problem set F
 Mosaic Analysis Reading: Chapter 5, pp140-141; Reference chapter D, pp820-823 Problem set F Twin spots in Drosophila Although segregation and recombination in mitosis do not occur at the same frequency
Mosaic Analysis Reading: Chapter 5, pp140-141; Reference chapter D, pp820-823 Problem set F Twin spots in Drosophila Although segregation and recombination in mitosis do not occur at the same frequency
Molecular Biology (9)
 Molecular Biology (9) Translation Mamoun Ahram, PhD Second semester, 2017-2018 1 Resources This lecture Cooper, Ch. 8 (297-319) 2 General information Protein synthesis involves interactions between three
Molecular Biology (9) Translation Mamoun Ahram, PhD Second semester, 2017-2018 1 Resources This lecture Cooper, Ch. 8 (297-319) 2 General information Protein synthesis involves interactions between three
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Molecular Biology of the Cell
 Alberts Johnson Lewis Morgan Raff Roberts Walter Molecular Biology of the Cell Sixth Edition Chapter 6 (pp. 333-368) How Cells Read the Genome: From DNA to Protein Copyright Garland Science 2015 Genetic
Alberts Johnson Lewis Morgan Raff Roberts Walter Molecular Biology of the Cell Sixth Edition Chapter 6 (pp. 333-368) How Cells Read the Genome: From DNA to Protein Copyright Garland Science 2015 Genetic
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct
 SUPPLEMENTAL FIGURE LEGENDS Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct waves. (A) A representative sub-lineage (ABala) of the C. elegans lineage tree (adapted from
SUPPLEMENTAL FIGURE LEGENDS Figure S1. Programmed cell death in the AB lineage occurs in temporally distinct waves. (A) A representative sub-lineage (ABala) of the C. elegans lineage tree (adapted from
Fertilization of sperm and egg produces offspring
 In sexual reproduction Fertilization of sperm and egg produces offspring In asexual reproduction Offspring are produced by a single parent, without the participation of sperm and egg CONNECTIONS BETWEEN
In sexual reproduction Fertilization of sperm and egg produces offspring In asexual reproduction Offspring are produced by a single parent, without the participation of sperm and egg CONNECTIONS BETWEEN
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Cell Division: the process of copying and dividing entire cells The cell grows, prepares for division, and then divides to form new daughter cells.
 Mitosis & Meiosis SC.912.L.16.17 Compare and contrast mitosis and meiosis and relate to the processes of sexual and asexual reproduction and their consequences for genetic variation. 1. Students will describe
Mitosis & Meiosis SC.912.L.16.17 Compare and contrast mitosis and meiosis and relate to the processes of sexual and asexual reproduction and their consequences for genetic variation. 1. Students will describe
Chapter 17 The Mechanism of Translation I: Initiation
 Chapter 17 The Mechanism of Translation I: Initiation Focus only on experiments discussed in class. Completely skip Figure 17.36 Read pg 521-527 up to the sentence that begins "In 1969, Joan Steitz..."
Chapter 17 The Mechanism of Translation I: Initiation Focus only on experiments discussed in class. Completely skip Figure 17.36 Read pg 521-527 up to the sentence that begins "In 1969, Joan Steitz..."
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
WD Repeat Domain of Dictyostelium Myosin Heavy Chain Kinase C Functions in both Substrate Targeting and Cellular Localization,
 WD Repeat Domain of Dictyostelium Myosin Heavy Chain Kinase C Functions in both Substrate Targeting and Cellular Localization, By: Atiya Franklin, Linzi Hyatt, Alyssa Chowdhury, and Paul A. Steimle* Franklin,
WD Repeat Domain of Dictyostelium Myosin Heavy Chain Kinase C Functions in both Substrate Targeting and Cellular Localization, By: Atiya Franklin, Linzi Hyatt, Alyssa Chowdhury, and Paul A. Steimle* Franklin,
Chapter 13: Meiosis & Sexual Life Cycles
 Chapter 13: Meiosis & Sexual Life Cycles What you must know The difference between asexual and sexual reproduction. The role of meiosis and fertilization in sexually reproducing organisms. The importance
Chapter 13: Meiosis & Sexual Life Cycles What you must know The difference between asexual and sexual reproduction. The role of meiosis and fertilization in sexually reproducing organisms. The importance
Lipid transfer proteins confer resistance to trichothecenes
 Lipid transfer proteins confer resistance to trichothecenes John McLaughlin and Anwar Bin-Umer Tumer Laboratory National Fusarium Head Blight Forum December 6th, 2012 FY09-11: Identify trichothecene resistance
Lipid transfer proteins confer resistance to trichothecenes John McLaughlin and Anwar Bin-Umer Tumer Laboratory National Fusarium Head Blight Forum December 6th, 2012 FY09-11: Identify trichothecene resistance
Supplemental Data. Chen and Thelen (2010). Plant Cell /tpc
 Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
A diploid somatic cell from a rat has a total of 42 chromosomes (2n = 42). As in humans, sex chromosomes determine sex: XX in females and XY in males.
 Multiple Choice Use the following information for questions 1-3. A diploid somatic cell from a rat has a total of 42 chromosomes (2n = 42). As in humans, sex chromosomes determine sex: XX in females and
Multiple Choice Use the following information for questions 1-3. A diploid somatic cell from a rat has a total of 42 chromosomes (2n = 42). As in humans, sex chromosomes determine sex: XX in females and
The structural basis of Edc3- and Scd6-mediated activation of the Dcp1:Dcp2 mrna decapping complex
 The EMO Journal (2012) 31, 279 290 & 2012 European Molecular iology Organization ll Rights Reserved 0261-4189/12 www.embojournal.org The structural basis of - and Scd6-mediated activation of the Dcp1:
The EMO Journal (2012) 31, 279 290 & 2012 European Molecular iology Organization ll Rights Reserved 0261-4189/12 www.embojournal.org The structural basis of - and Scd6-mediated activation of the Dcp1:
Supplemental Data. Perea-Resa et al. Plant Cell. (2012) /tpc
 Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant
 9 10 11 Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant Makoto Kazama 1 *, Sanae Ogiwara, Takashi Makiuchi 1, Kazuhiro Yoshida 1, Kumiko Nakada-Tsukui, Tomoyoshi
9 10 11 Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant Makoto Kazama 1 *, Sanae Ogiwara, Takashi Makiuchi 1, Kazuhiro Yoshida 1, Kumiko Nakada-Tsukui, Tomoyoshi
Supplemental Information. Inferring Cell-State Transition. Dynamics from Lineage Trees. and Endpoint Single-Cell Measurements
 Cell Systems, Volume 3 Supplemental Information Inferring Cell-State Transition Dynamics from Lineage Trees and Endpoint Single-Cell Measurements Sahand Hormoz, Zakary S. Singer, James M. Linton, Yaron
Cell Systems, Volume 3 Supplemental Information Inferring Cell-State Transition Dynamics from Lineage Trees and Endpoint Single-Cell Measurements Sahand Hormoz, Zakary S. Singer, James M. Linton, Yaron
Translational Initiation
 Translational Initiation Lecture Outline 1. Process of Initiation. Alternative mechanisms of Initiation 3. Key Experiments on Initiation 4. Regulation of Initiation Translation is a process with three
Translational Initiation Lecture Outline 1. Process of Initiation. Alternative mechanisms of Initiation 3. Key Experiments on Initiation 4. Regulation of Initiation Translation is a process with three
Supplementary Information
 Supplementary Information An engineered protein antagonist of K-Ras/B-Raf interaction Monique J. Kauke, 1,2 Michael W. Traxlmayr 1,2, Jillian A. Parker 3, Jonathan D. Kiefer 4, Ryan Knihtila 3, John McGee
Supplementary Information An engineered protein antagonist of K-Ras/B-Raf interaction Monique J. Kauke, 1,2 Michael W. Traxlmayr 1,2, Jillian A. Parker 3, Jonathan D. Kiefer 4, Ryan Knihtila 3, John McGee
Biology, 7e (Campbell) Chapter 13: Meiosis and Sexual Life Cycles
 Biology, 7e (Campbell) Chapter 13: Meiosis and Sexual Life Cycles Chapter Questions 1) What is a genome? A) the complete complement of an organism's genes B) a specific sequence of polypeptides within
Biology, 7e (Campbell) Chapter 13: Meiosis and Sexual Life Cycles Chapter Questions 1) What is a genome? A) the complete complement of an organism's genes B) a specific sequence of polypeptides within
Which of these best predicts the outcome of the changes illustrated in the diagrams?
 1. The diagrams below show two different scenarios for a pair of homologous chromosomes, known as a tetrad, undergoing a change where segments of DNA switch on parts of the chromosomes. In each scenario,
1. The diagrams below show two different scenarios for a pair of homologous chromosomes, known as a tetrad, undergoing a change where segments of DNA switch on parts of the chromosomes. In each scenario,
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN
 THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
Describe the process of cell division in prokaryotic cells. The Cell Cycle
 The Cell Cycle Objective # 1 In this topic we will examine the cell cycle, the series of changes that a cell goes through from one division to the next. We will pay particular attention to how the genetic
The Cell Cycle Objective # 1 In this topic we will examine the cell cycle, the series of changes that a cell goes through from one division to the next. We will pay particular attention to how the genetic
Supplementary material
 Supplementary material Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins Vladimir V. Rogov 1,*, Hironori Suzuki 2,3,*, Mija Marinković 4, Verena Lang
Supplementary material Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins Vladimir V. Rogov 1,*, Hironori Suzuki 2,3,*, Mija Marinković 4, Verena Lang
Welcome to Class 21!
 Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Supplementary Information for. Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria
 Supplementary Information for Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria Isabella Santi 1 *, Neeraj Dhar 1, Djenet Bousbaine 1, Yuichi Wakamoto, John D.
Supplementary Information for Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria Isabella Santi 1 *, Neeraj Dhar 1, Djenet Bousbaine 1, Yuichi Wakamoto, John D.
-14. -Abdulrahman Al-Hanbali. -Shahd Alqudah. -Dr Ma mon Ahram. 1 P a g e
 -14 -Abdulrahman Al-Hanbali -Shahd Alqudah -Dr Ma mon Ahram 1 P a g e In this lecture we will talk about the last stage in the synthesis of proteins from DNA which is translation. Translation is the process
-14 -Abdulrahman Al-Hanbali -Shahd Alqudah -Dr Ma mon Ahram 1 P a g e In this lecture we will talk about the last stage in the synthesis of proteins from DNA which is translation. Translation is the process
Meiosis and Sexual Life Cycles
 Chapter 13 Meiosis and Sexual Life Cycles Lecture Outline Overview Living organisms are distinguished by their ability to reproduce their own kind. Offspring resemble their parents more than they do less
Chapter 13 Meiosis and Sexual Life Cycles Lecture Outline Overview Living organisms are distinguished by their ability to reproduce their own kind. Offspring resemble their parents more than they do less
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
BIOLOGY. Chapter 10 CELL REPRODUCTION PowerPoint Image Slideshow
 BIOLOGY Chapter 10 CELL REPRODUCTION PowerPoint Image Slideshow FIGURE 10.1 A sea urchin begins life as a single cell that (a) divides to form two cells, visible by scanning electron microscopy. After
BIOLOGY Chapter 10 CELL REPRODUCTION PowerPoint Image Slideshow FIGURE 10.1 A sea urchin begins life as a single cell that (a) divides to form two cells, visible by scanning electron microscopy. After
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb97 P ( μm, hours) 1 2 4 P DMSO Figure S1 unningham et al. E 97 65 27 MFN1 GFP-Parkin Opa1 ctin GPDH HEK293 GFP-Parkin 19 115 97 65 27 Mitochondrial Fraction SH-SY5Y GFP-Parkin Mito DMSO Mito
DOI:.38/ncb97 P ( μm, hours) 1 2 4 P DMSO Figure S1 unningham et al. E 97 65 27 MFN1 GFP-Parkin Opa1 ctin GPDH HEK293 GFP-Parkin 19 115 97 65 27 Mitochondrial Fraction SH-SY5Y GFP-Parkin Mito DMSO Mito
Table S1. Aspergillus nidulans strains used in this study Strain Genotype Derivation
 Supplemental Material De Souza et al., 211 Table S1. Aspergillus nidulans strains used in this study Strain Genotype Derivation CDS295 pyrg89; pyroa4; pyrg Af ::son promotor::gfp-son nup98/nup96 ; chaa1
Supplemental Material De Souza et al., 211 Table S1. Aspergillus nidulans strains used in this study Strain Genotype Derivation CDS295 pyrg89; pyroa4; pyrg Af ::son promotor::gfp-son nup98/nup96 ; chaa1
Three types of RNA polymerase in eukaryotic nuclei
 Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
Biology I Level - 2nd Semester Final Review
 Biology I Level - 2nd Semester Final Review The 2 nd Semester Final encompasses all material that was discussed during second semester. It s important that you review ALL notes and worksheets from the
Biology I Level - 2nd Semester Final Review The 2 nd Semester Final encompasses all material that was discussed during second semester. It s important that you review ALL notes and worksheets from the
Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION
 http://smtom.lecture.ub.ac.id/ Password: https://syukur16tom.wordpress.com/ Password: Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/translation2.gif
http://smtom.lecture.ub.ac.id/ Password: https://syukur16tom.wordpress.com/ Password: Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/translation2.gif
Prof. Fahd M. Nasr. Lebanese university Faculty of sciences I Department of Natural Sciences.
 Prof. Fahd M. Nasr Lebanese university Faculty of sciences I Department of Natural Sciences fnasr@ul.edu.lb B3206 Microbial Genetics Eukaryotic M. G. The yeast Saccharomyces cerevisiae as a genetic model
Prof. Fahd M. Nasr Lebanese university Faculty of sciences I Department of Natural Sciences fnasr@ul.edu.lb B3206 Microbial Genetics Eukaryotic M. G. The yeast Saccharomyces cerevisiae as a genetic model
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10534 Supplementary Fig. 1. Diagrammatic representation of the N-end rule pathway of targeted proteolysis (after Graciet and Wellmer 2010 9 ). Tertiary, secondary
SUPPLEMENTARY INFORMATION doi:10.1038/nature10534 Supplementary Fig. 1. Diagrammatic representation of the N-end rule pathway of targeted proteolysis (after Graciet and Wellmer 2010 9 ). Tertiary, secondary
Interactions among Ytm1, Erb1, and Nop7 required for assembly of the Nop7-subcomplex in yeast preribosomes.
 Carnegie Mellon University Research Showcase @ CMU Department of Biological Sciences Mellon College of Science 7-1-2008 Interactions among Ytm1, Erb1, and Nop7 required for assembly of the Nop7-subcomplex
Carnegie Mellon University Research Showcase @ CMU Department of Biological Sciences Mellon College of Science 7-1-2008 Interactions among Ytm1, Erb1, and Nop7 required for assembly of the Nop7-subcomplex
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10244 a O07391_MYCAV/127-243 NLPC_HAEIN/80-181 SPR_SHIFL/79-183 P74160_SYNY3/112-245 O24914_HELPY/301-437 Q51835_PORGI/68-178 DPP6_BACSH/163-263 YKFC_BACSU/185-292 YDHO_ECOLI/153-263
doi:10.1038/nature10244 a O07391_MYCAV/127-243 NLPC_HAEIN/80-181 SPR_SHIFL/79-183 P74160_SYNY3/112-245 O24914_HELPY/301-437 Q51835_PORGI/68-178 DPP6_BACSH/163-263 YKFC_BACSU/185-292 YDHO_ECOLI/153-263
Nature Genetics: doi: /ng Supplementary Figure 1. ssp mutant phenotypes in a functional SP background.
 Supplementary Figure 1 ssp mutant phenotypes in a functional SP background. (a,b) Statistical comparisons of primary and sympodial shoot flowering times as determined by mean values for leaf number on
Supplementary Figure 1 ssp mutant phenotypes in a functional SP background. (a,b) Statistical comparisons of primary and sympodial shoot flowering times as determined by mean values for leaf number on
downstream (0.8 kb) homologous sequences to the genomic locus of DIC. A DIC mutant strain (ro- 6
 A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
Reading Assignments. A. Systems of Cell Division. Lecture Series 5 Cell Cycle & Cell Division
 Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Death Read Chapter 19 Cell Division Read Chapter 20 pages 659-672 672 only (Benefits of Sex & Meiosis sections)
Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Death Read Chapter 19 Cell Division Read Chapter 20 pages 659-672 672 only (Benefits of Sex & Meiosis sections)
Lecture Series 5 Cell Cycle & Cell Division
 Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Death Read Chapter 19 Cell Division Read Chapter 20 pages 659-672 672 only (Benefits of Sex & Meiosis sections)
Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Death Read Chapter 19 Cell Division Read Chapter 20 pages 659-672 672 only (Benefits of Sex & Meiosis sections)
MEIOSIS LAB INTRODUCTION PART I: SIMULATION OF MEIOSIS EVOLUTION. Activity #9
 AP BIOLOGY EVOLUTION Unit 1 Part 7 Chapter 13 Activity #9 NAME DATE PERIOD MEIOSIS LAB INTRODUCTION Meiosis involves two successive nuclear divisions that produce four haploid cells. Meiosis I is the reduction
AP BIOLOGY EVOLUTION Unit 1 Part 7 Chapter 13 Activity #9 NAME DATE PERIOD MEIOSIS LAB INTRODUCTION Meiosis involves two successive nuclear divisions that produce four haploid cells. Meiosis I is the reduction
Meiosis and Sexual Life Cycles
 Chapter 13 Meiosis and Sexual Life Cycles PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 13 Meiosis and Sexual Life Cycles PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Induction of cap-independent BiP (hsp-3) and Bcl-2 (ced-9) translation in response to eif4g (IFG-1) depletion in C. elegans
 Research Paper Translation 2, e28935; April; 2014 Landes Bioscience Research Paper Induction of cap-independent BiP (hsp-3) and Bcl-2 (ced-9) translation in response to eif4g (IFG-1) depletion in C. elegans
Research Paper Translation 2, e28935; April; 2014 Landes Bioscience Research Paper Induction of cap-independent BiP (hsp-3) and Bcl-2 (ced-9) translation in response to eif4g (IFG-1) depletion in C. elegans
Camello, a novel family of Histone Acetyltransferases that acetylate histone H4 and is essential for zebrafish development
 Supplementary Information: Camello, a novel family of Histone Acetyltransferases that acetylate histone H4 and is essential for zebrafish development Krishanpal Karmodiya 1, Krishanpal Anamika 1,2, Vijaykumar
Supplementary Information: Camello, a novel family of Histone Acetyltransferases that acetylate histone H4 and is essential for zebrafish development Krishanpal Karmodiya 1, Krishanpal Anamika 1,2, Vijaykumar
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Expanded View Figures
 The EMBO Journal Structure of a Dm peptide bound to the OT module Tobias Raisch et al Expanded View Figures A Hs Dm 262 297 685 8 HEAT HEAT MIF4G 9BD 1SHD 761 91 193 169 1152 1317 16 1376 1467 HEAT HEAT
The EMBO Journal Structure of a Dm peptide bound to the OT module Tobias Raisch et al Expanded View Figures A Hs Dm 262 297 685 8 HEAT HEAT MIF4G 9BD 1SHD 761 91 193 169 1152 1317 16 1376 1467 HEAT HEAT
Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with
 Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with DsRed-mito. Right panels are time-course enlarged images
Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with DsRed-mito. Right panels are time-course enlarged images
Chapter 2: Chromosomes and cellular reproduction
 Chapter 2: Chromosomes and cellular reproduction I. Contrast between prokaryotes and eukaryotes. See Figure 2.1 Nucleus absent Small diameter 1 to 10 µm Genome usually 1 circular molecule Small genome;
Chapter 2: Chromosomes and cellular reproduction I. Contrast between prokaryotes and eukaryotes. See Figure 2.1 Nucleus absent Small diameter 1 to 10 µm Genome usually 1 circular molecule Small genome;
GFP WxTP-GFP WxTP-GFP. stromules. DsRed WxTP-DsRed WxTP-DsRed. stromules
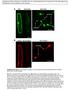 Supplemental Data. Kitajima et al. (2009). The rice -amylase glycoprotein is targeted from the Golgi apparatus through the secretory pathway to the plastids. A GFP WxTP-GFP WxTP-GFP stromules stromules
Supplemental Data. Kitajima et al. (2009). The rice -amylase glycoprotein is targeted from the Golgi apparatus through the secretory pathway to the plastids. A GFP WxTP-GFP WxTP-GFP stromules stromules
Meiosis and Sexual Life Cycles
 Chapter 13 Meiosis and Sexual Life Cycles PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 13 Meiosis and Sexual Life Cycles PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Ladies and Gentlemen.. The King of Rock and Roll
 Ladies and Gentlemen.. The King of Rock and Roll Learning Objectives: The student is able to construct an explanation, using visual representations or narratives, as to how DNA in chromosomes is transmitted
Ladies and Gentlemen.. The King of Rock and Roll Learning Objectives: The student is able to construct an explanation, using visual representations or narratives, as to how DNA in chromosomes is transmitted
EST1 Homology Domain. 100 aa. hest1a / SMG6 PIN TPR TPR. Est1-like DBD? hest1b / SMG5. TPR-like TPR. a helical. hest1c / SMG7.
 hest1a / SMG6 EST1 Homology Domain 100 aa 853 695 761 780 1206 hest1 / SMG5 -like? -like 109 145 214 237 497 165 239 1016 114 207 212 381 583 hest1c / SMG7 a helical 1091 Sc 57 185 267 284 699 Figure S1:
hest1a / SMG6 EST1 Homology Domain 100 aa 853 695 761 780 1206 hest1 / SMG5 -like? -like 109 145 214 237 497 165 239 1016 114 207 212 381 583 hest1c / SMG7 a helical 1091 Sc 57 185 267 284 699 Figure S1:
Lecture Series 5 Cell Cycle & Cell Division
 Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Division Read Chapter 19 pages 651-663 663 only (Benefits of Sex & Meiosis sections these are in Chapter
Lecture Series 5 Cell Cycle & Cell Division Reading Assignments Read Chapter 18 Cell Cycle & Cell Division Read Chapter 19 pages 651-663 663 only (Benefits of Sex & Meiosis sections these are in Chapter
Chromosome Chr Duplica Duplic t a ion Pixley
 Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and
 Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Chromosome duplication and distribution during cell division
 CELL DIVISION AND HEREDITY Student Packet SUMMARY IN EUKARYOTES, HERITABLE INFORMATION IS PASSED TO THE NEXT GENERATION VIA PROCESSES THAT INCLUDE THE CELL CYCLE, MITOSIS /MEIOSIS AND FERTILIZATION Mitosis
CELL DIVISION AND HEREDITY Student Packet SUMMARY IN EUKARYOTES, HERITABLE INFORMATION IS PASSED TO THE NEXT GENERATION VIA PROCESSES THAT INCLUDE THE CELL CYCLE, MITOSIS /MEIOSIS AND FERTILIZATION Mitosis
11-4 Meiosis. Chromosome Number
 11-4 Meiosis Chromosome Number Sexual reproduction shuffles and recombines genes from two parents. During gametogenesis, genes are segregated and assorted (shuffled) into gemetes, and at fertilization,
11-4 Meiosis Chromosome Number Sexual reproduction shuffles and recombines genes from two parents. During gametogenesis, genes are segregated and assorted (shuffled) into gemetes, and at fertilization,
Learning Objectives LO 3.7 The student can make predictions about natural phenomena occurring during the cell cycle. [See SP 6.4]
![Learning Objectives LO 3.7 The student can make predictions about natural phenomena occurring during the cell cycle. [See SP 6.4] Learning Objectives LO 3.7 The student can make predictions about natural phenomena occurring during the cell cycle. [See SP 6.4]](/thumbs/94/118687827.jpg) Big Ideas 3.A.2: In eukaryotes, heritable information is passed to the next generation via processes that include the cell cycle and mitosis or meiosis plus fertilization. CHAPTER 13 MEIOSIS AND SEXUAL
Big Ideas 3.A.2: In eukaryotes, heritable information is passed to the next generation via processes that include the cell cycle and mitosis or meiosis plus fertilization. CHAPTER 13 MEIOSIS AND SEXUAL
Supplementary Figure 1. Nature Genetics: doi: /ng.3848
 Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
Reconstructing Mitochondrial Evolution?? Morphological Diversity. Mitochondrial Diversity??? What is your definition of a mitochondrion??
 Reconstructing Mitochondrial Evolution?? What is your definition of a mitochondrion?? Morphological Diversity Mitochondria as we all know them: Suprarenal gland Liver cell Plasma cell Adrenal cortex Mitochondrial
Reconstructing Mitochondrial Evolution?? What is your definition of a mitochondrion?? Morphological Diversity Mitochondria as we all know them: Suprarenal gland Liver cell Plasma cell Adrenal cortex Mitochondrial
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Arabidopsis COMPASS-Like Complexes Mediate Histone H3 Lysine-4 Trimethylation to Control Floral Transition and Plant Development
 Arabidopsis COMPASS-Like Complexes Mediate Histone H3 Lysine-4 Trimethylation to Control Floral Transition and Plant Development Danhua Jiang 1,2, Nicholas C. Kong 1,2, Xiaofeng Gu 2, Zicong Li 1, Yuehui
Arabidopsis COMPASS-Like Complexes Mediate Histone H3 Lysine-4 Trimethylation to Control Floral Transition and Plant Development Danhua Jiang 1,2, Nicholas C. Kong 1,2, Xiaofeng Gu 2, Zicong Li 1, Yuehui
Purposes of Cell Division
 Purposes of Cell Division Increase the number of cells for growth and repair of worn out tissues What examples in the human body can you think of? Transmit genetic information to later generations Why
Purposes of Cell Division Increase the number of cells for growth and repair of worn out tissues What examples in the human body can you think of? Transmit genetic information to later generations Why
STUDY UNIT 1 MITOSIS AND MEIOSIS. Klug, Cummings & Spencer Chapter 2. Morphology of eukaryotic metaphase chromosomes. Chromatids
 STUDY UNIT 1 MITOSIS AND MEIOSIS Klug, Cummings & Spencer Chapter 2 Life depends on cell division and reproduction of organisms. Process involves transfer of genetic material. New somatic (body) cells
STUDY UNIT 1 MITOSIS AND MEIOSIS Klug, Cummings & Spencer Chapter 2 Life depends on cell division and reproduction of organisms. Process involves transfer of genetic material. New somatic (body) cells
Supplemental Data. Wang et al. (2014). Plant Cell /tpc
 Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Supplemental Data. Hou et al. (2016). Plant Cell /tpc
 Supplemental Data. Hou et al. (216). Plant Cell 1.115/tpc.16.295 A Distance to 1 st nt of start codon Distance to 1 st nt of stop codon B Normalized PARE abundance 8 14 nt 17 nt Frame1 Arabidopsis inflorescence
Supplemental Data. Hou et al. (216). Plant Cell 1.115/tpc.16.295 A Distance to 1 st nt of start codon Distance to 1 st nt of stop codon B Normalized PARE abundance 8 14 nt 17 nt Frame1 Arabidopsis inflorescence
2012 Univ Aguilera Lecture. Introduction to Molecular and Cell Biology
 2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
Cell Division: the process of copying and dividing entire cells The cell grows, prepares for division, and then divides to form new daughter cells.
 Mitosis & Meiosis SC.912.L.16.17 Compare and contrast mitosis and meiosis and relate to the processes of sexual and asexual reproduction and their consequences for genetic variation. 1. Students will describe
Mitosis & Meiosis SC.912.L.16.17 Compare and contrast mitosis and meiosis and relate to the processes of sexual and asexual reproduction and their consequences for genetic variation. 1. Students will describe
CHAPTER 6. Chromosomes and Meiosis
 CHAPTER 6 Chromosomes and Meiosis CHROMOSOMES DNA (deoxyribonucleic acid) is a long, thin molecule that directs cellular functions and heredity. DNA contains information that is encoded in segments called
CHAPTER 6 Chromosomes and Meiosis CHROMOSOMES DNA (deoxyribonucleic acid) is a long, thin molecule that directs cellular functions and heredity. DNA contains information that is encoded in segments called
CELL CYCLE UNIT GUIDE- Due January 19, 2016
 CELL CYCLE UNIT GUIDE- Due January 19, 2016 Monday Tuesday Wednesday Thursday Friday January 4- No School 5-Cell Cycle/Mitosis 6-Cell Cycle/ Mitosis 7-Mitosis 8-Meiosis Reading Check Quiz #1 sections 5.1-5.5
CELL CYCLE UNIT GUIDE- Due January 19, 2016 Monday Tuesday Wednesday Thursday Friday January 4- No School 5-Cell Cycle/Mitosis 6-Cell Cycle/ Mitosis 7-Mitosis 8-Meiosis Reading Check Quiz #1 sections 5.1-5.5
