Camello, a novel family of Histone Acetyltransferases that acetylate histone H4 and is essential for zebrafish development
|
|
|
- Charles Riley
- 5 years ago
- Views:
Transcription
1 Supplementary Information: Camello, a novel family of Histone Acetyltransferases that acetylate histone H4 and is essential for zebrafish development Krishanpal Karmodiya 1, Krishanpal Anamika 1,2, Vijaykumar Muley 1, Saurabh J. Pradhan 1, Yoshita Bhide 1 and Sanjeev Galande 1 1 Center of Excellence in Epigenetics, Indian Institute of Science Education and Research, Pashan, Pune , India 2 Present address: Persistent Systems Limited Pingala - Aryabhata, Erandwane, Pune , India. Corresponding author Running title: Genome-wide survey of HATs Keywords: histone acetyltransferase/camello proteins/histone acetylation/histone 4/ evolution/zebrafish/.
2 Supplementary Information Methods Homologs of mouse CMLO3 protein and phylogenetic analysis Homologs of CMLO3 protein of mouse were fished by searching against the database of protein sequences from 23 completely sequenced eukaryotic organisms (Supplementary Table 1). Splice isoforms were removed from ENSEMBL genome datasets prior to search for homologs. Jackhmmer program of HMMER package was used to search database for 3 iterations. The e- value for reporting homologs of CMLO3 protein sequence was set to the 1 and e-value significance for including sequences in next round of search was This search yielded 450 proteins as probable homologs of CMLO3 protein after search of three iterations. The alignments generated for homologs of CMLO3 were retained and used for further analysis. These proteins were then subjected to the hmmscan program of HMMER package to identify domains and superfamilies reported in Pfam and Superfamily databases [1] Phylogenetic analysis of CMLO3 homologs Mulitple sequence alignments (MSAs) were constructed for Pfam and Superfamily detected acetyltransferase domains only and also for complete 450 sequences. These MSAs were subjected to the phyml software to reconstruct Maximum-likelihood Phylogenetic trees using default parameters [2]. Cell culture and transfection HeLa cells were grown in DMEM medium supplemented with 10% fetal bovine serum at 37 0 C with 5 % CO 2. For transient transfections, 0.5 x 10 6 cells were transfected with 2.5 and 5.0 µg of plasmid DNA by Lipofectamine 2000 (Invitrogen). Transfection was performed according to the manufacturer s instructions. Immunostaining and microscopy HeLa cells were stained with anti-calnexin (ab22595, Abcam) and anti-lamin B1 (Ab16048, Abcam) antibodies after fixation and permeabilization. DNA counterstaining was performed using DAPI (4, 6 -diamidino-2-phenylindole). Z stacks of the images were collected as 0.2 µm
3 optical slices by confocal microscope (Carl Zeiss). Images were processed using ImageJ software. References: 1. Bateman A, Birney E, Cerruti L, Durbin R, Etwiller L, Eddy SR, Griffiths-Jones S, Howe KL, Marshall M, Sonnhammer EL (2002) The Pfam protein families database. Nucleic Acids Res, 30: Guindon S., Dufayard J.F., Lefort V., Anisimova M., Hordijk W., Gascuel O (2010) New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Systematic Biology, 59(3):
4 Supplementary Figure legends: Additional file 1: Figure S1. Complement of KAT homologs in Zebrafish genome. Dendrogram of KATs in mouse classified into known and new family containing camello proteins. Homologs of different KAT families are represented in different colors. The new family of KAT Camello family identified in this study is highlighted in blue color. Presence of Camello proteins in zebrafish suggest that this family of proteins is conserved across genomes.
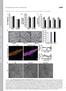
5 Additional file2: Figure S2. Purification of camello proteins, CMLO3, E0CYR6 and CMLO2 by Ni-NTA chromatography. 12 % SDS PAGE of recombinant camello proteins. Protein molecular weight markers is from MBI Fermentas. Lane 1: purified CMLO3. Lane 2: E0CYR6. Lane 3: CMLO2. CMLO3, E0CYR6 and CMLO2 (~28 kda proteins) are purified to homogeneity.
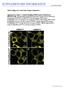
6 Additional file3: Figure S3. CMLO3 is expressed as GFP fusion protein in HeLa cells. Colocalization of CMLO3-GFP fusion protein (green, channel 2) was compared with endoplasmic reticulum marker, calnexin (red, channel 3). CMLO3 is not co-localized with endoplasmic reticulum marker.
7 Additional file4: Figure S4. CMLO3 is expressed as GFP fusion protein in HeLa cells. Colocalization of CMLO3-GFP fusion protein (green, channel 1) was compared with marker of inner nuclear membrane, β-lamin (red, channel 2). CMLO3 is co-localized with marker of inner nuclear membrane lamin B1.
8 Additional file5: Figure S5. (A) C-terminal his-myc tagging of the CMLO3 recapitulate the localization profile of the CMLO3-GFP fusion protein and its overlap with lamin B1, marker of inner nuclear membrane. (B) N-terminal tagging of CMLO3 with his-myc affects the localization of the CMLO3 and was found mostly in the cytoplasm.
9 Additional file7: Figure S6. Multiple sequence alignment of CMLO3 homologs. 450 proteins detected by jackhmmer in 23 genomes aligned using clustalo program. Green color box shows sequence region unique to the CMLO3 proteins and red color box shows approximate location of acetyl transferase domain. I could detect homologs in the unicellular yeast, monosiga, and sponge for acetyl transferase domain but Cnidarian Nematostella encodes full length CMLO3- like protein suggesting that the N-terminus (green box) region of CMLO3 proteins is acquired along with the tool kit necessary for two germ layer formations. Cnidarians are the early branching multicellular organisms with two germ layers and body axis.
10 Additional file6: Table S1. Details of eukaryotic organisms used for CMLO3 protein homologs. Complete sets of protein sequences for all organisms were downloaded from Ensembl, NCBI, and JGI databases in January, Ensembl sequences were updated in April, Name of the organism Phylum Data source Saccharomyces cerevisiae Ascomycota Ensembl Monosiga brevicollis Craspedida JGI Amphimedon queenslandica Porifera Ensembl Hydra magnipapillata Cnidaria NCBI Nematostella vectensis Cnidaria Ensembl Drosophila melanogaster Arthropoda Ensembl Caenorhabditis elegans Nematoda Ensembl Strongylocentrotus purpuratus Echinodermata Ensembl Saccoglossus kowalevskii Hemichordata NCBI Branchiostoma floridae Chordata NCBI Ciona intestinalis Chordata Ensembl Petromyzon marinus Chordata Ensembl Danio rerio Chordata Ensembl Xenopus tropicalis Chordata Ensembl Gallus gallus Chordata Ensembl Anolis carolinensis Chordata Ensembl Ornithorhynchus anatinus Chordata Ensembl Monodelphis domestica Chordata Ensembl Bos Taurus Chordata Ensembl Oryctolagus cuniculus Chordata Ensembl Mus musculus Chordata Ensembl Pan troglodytes Chordata Ensembl Homo sapiens Chordata Ensembl
11 Additional file8: Figure S7: Camello family proteins CMLO3 and E0CYR6 are active histone acetyltransferases. (A) HAT assay using baculo produced recombinant histones, H3 and H4. E0CYR6 and CMLO3 are showing histone H4 acetylation (lanes 2 and 3, respectively).
12 Additional file9: Figure S8: Western blot analysis of CMLO3-GFP overexpressing HeLa cells after cytoplasmic and nuclear fractionation. Lamin B1 and tubulin were used as markers of nuclear and cytoplasmic fractions respectively. CMLO3-GFP was mainly observed in nuclear fraction. C - Cytoplasmic fraction; N- Nuclear fraction.
Comparing Genomes! Homologies and Families! Sequence Alignments!
 Comparing Genomes! Homologies and Families! Sequence Alignments! Allows us to achieve a greater understanding of vertebrate evolution! Tells us what is common and what is unique between different species
Comparing Genomes! Homologies and Families! Sequence Alignments! Allows us to achieve a greater understanding of vertebrate evolution! Tells us what is common and what is unique between different species
Advanced Cell Biology. Lecture 2
 Advanced Cell Biology. Lecture 2 Alexey Shipunov Minot State University January 13, 2012 Outline Questions and answers Microscopy Prokaryotic and eukaryotic cells Outline Questions and answers Microscopy
Advanced Cell Biology. Lecture 2 Alexey Shipunov Minot State University January 13, 2012 Outline Questions and answers Microscopy Prokaryotic and eukaryotic cells Outline Questions and answers Microscopy
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Reassessing Domain Architecture Evolution of Metazoan Proteins: Major Impact of Gene Prediction Errors
 Genes 2011, 2, 449-501; doi:10.3390/genes2030449 Article OPEN ACCESS genes ISSN 2073-4425 www.mdpi.com/journal/genes Reassessing Domain Architecture Evolution of Metazoan Proteins: Major Impact of Gene
Genes 2011, 2, 449-501; doi:10.3390/genes2030449 Article OPEN ACCESS genes ISSN 2073-4425 www.mdpi.com/journal/genes Reassessing Domain Architecture Evolution of Metazoan Proteins: Major Impact of Gene
Biased amino acid composition in warm-blooded animals
 Biased amino acid composition in warm-blooded animals Guang-Zhong Wang and Martin J. Lercher Bioinformatics group, Heinrich-Heine-University, Düsseldorf, Germany Among eubacteria and archeabacteria, amino
Biased amino acid composition in warm-blooded animals Guang-Zhong Wang and Martin J. Lercher Bioinformatics group, Heinrich-Heine-University, Düsseldorf, Germany Among eubacteria and archeabacteria, amino
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1213 Supplementary Table 1. The Taxonomy of the Organisms Used in this Study Organism (acronym) Taxonomy Yeasts Zygosacharomyces rouxii (Zrou) Verterbrates Xenopus tropicalis (Xtro) Gallus
doi:1.138/nature1213 Supplementary Table 1. The Taxonomy of the Organisms Used in this Study Organism (acronym) Taxonomy Yeasts Zygosacharomyces rouxii (Zrou) Verterbrates Xenopus tropicalis (Xtro) Gallus
Supplemental Figure 1.
 Supplemental Material: Annu. Rev. Genet. 2015. 49:213 42 doi: 10.1146/annurev-genet-120213-092023 A Uniform System for the Annotation of Vertebrate microrna Genes and the Evolution of the Human micrornaome
Supplemental Material: Annu. Rev. Genet. 2015. 49:213 42 doi: 10.1146/annurev-genet-120213-092023 A Uniform System for the Annotation of Vertebrate microrna Genes and the Evolution of the Human micrornaome
Master Biomedizin ) UCSC & UniProt 2) Homology 3) MSA 4) Phylogeny. Pablo Mier
 Master Biomedizin 2018 1) UCSC & UniProt 2) Homology 3) MSA 4) 1 12 a. All of the sequences in file1.fasta (https://cbdm.uni-mainz.de/mb18/) are homologs. How many groups of orthologs would you say there
Master Biomedizin 2018 1) UCSC & UniProt 2) Homology 3) MSA 4) 1 12 a. All of the sequences in file1.fasta (https://cbdm.uni-mainz.de/mb18/) are homologs. How many groups of orthologs would you say there
Computational Structural Bioinformatics
 Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny
 Phylogenetics and chromosomal synteny of the GATAs 1273 GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny CHUNJIANG HE, HANHUA CHENG* and RONGJIA ZHOU* Department
Phylogenetics and chromosomal synteny of the GATAs 1273 GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny CHUNJIANG HE, HANHUA CHENG* and RONGJIA ZHOU* Department
Supplementary Information. Recognition of the pre-mirna structure by Drosophila Dicer-1
 Supplementary Information Recognition of the pre-mirna structure by Drosophila Dicer-1 Akihisa Tsutsumi 1,2, Tomoko Kawamata 1, Natsuko Izumi 1 Hervé Seitz 3,4 and Yukihide Tomari 1,2 Nature Structural
Supplementary Information Recognition of the pre-mirna structure by Drosophila Dicer-1 Akihisa Tsutsumi 1,2, Tomoko Kawamata 1, Natsuko Izumi 1 Hervé Seitz 3,4 and Yukihide Tomari 1,2 Nature Structural
Cryo-EM data collection, refinement and validation statistics
 1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
Supplemental Data. Gao et al. (2012). Plant Cell /tpc
 Supplemental Figure 1. Plant EMP Proteins. (A) The Accession numbers of the 12 EMP members from Arabidopsis. (B) Phylogenetic analysis of EMP proteins from Arabidopsis, human and yeast using the Mac Vector
Supplemental Figure 1. Plant EMP Proteins. (A) The Accession numbers of the 12 EMP members from Arabidopsis. (B) Phylogenetic analysis of EMP proteins from Arabidopsis, human and yeast using the Mac Vector
Supplemental Materials Molecular Biology of the Cell
 Supplemental Materials Molecular iology of the Cell Figure S1 Krüger et al. Arabidopsis Plasmodium H. sapiens* 1) Xenopus* 1) Drosophila C.elegans S.cerevisae S.pombe L.major T.cruzi T.brucei DCP5 CITH
Supplemental Materials Molecular iology of the Cell Figure S1 Krüger et al. Arabidopsis Plasmodium H. sapiens* 1) Xenopus* 1) Drosophila C.elegans S.cerevisae S.pombe L.major T.cruzi T.brucei DCP5 CITH
BIOINFORMATICS LAB AP BIOLOGY
 BIOINFORMATICS LAB AP BIOLOGY Bioinformatics is the science of collecting and analyzing complex biological data. Bioinformatics combines computer science, statistics and biology to allow scientists to
BIOINFORMATICS LAB AP BIOLOGY Bioinformatics is the science of collecting and analyzing complex biological data. Bioinformatics combines computer science, statistics and biology to allow scientists to
Introduction to cells
 Almen Cellebiologi Introduction to cells 1. Unity and diversity of cells 2. Microscopes and visualization of cells 3. Prokaryotic cells, eubacteria and archaea 4. Eucaryotic cells, nucleus, mitochondria
Almen Cellebiologi Introduction to cells 1. Unity and diversity of cells 2. Microscopes and visualization of cells 3. Prokaryotic cells, eubacteria and archaea 4. Eucaryotic cells, nucleus, mitochondria
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Marine medaka ATP-binding cassette (ABC) superfamily and new insight into teleost Abch nomenclature
 Supplementary file for: Marine medaka ATP-binding cassette (ABC) superfamily and new insight into teleost Abch nomenclature Chang-Bum Jeong a,b,#, Bo-Mi Kim a,#, Hye-Min Kang a, Ik-Young Choi c, Jae-Sung
Supplementary file for: Marine medaka ATP-binding cassette (ABC) superfamily and new insight into teleost Abch nomenclature Chang-Bum Jeong a,b,#, Bo-Mi Kim a,#, Hye-Min Kang a, Ik-Young Choi c, Jae-Sung
Sheet1. Page 1. protein
 build 1 version 1 Ab initio model Ab initio GPIPE/6085/1.1/gnomon_prot build 1 version 1 Build GPIPE/6085/1.1/ Acyrthosiphon pisum build 2 version 1 Build GPIPE/7029/2.1/ Acyrthosiphon pisum build 2.1
build 1 version 1 Ab initio model Ab initio GPIPE/6085/1.1/gnomon_prot build 1 version 1 Build GPIPE/6085/1.1/ Acyrthosiphon pisum build 2 version 1 Build GPIPE/7029/2.1/ Acyrthosiphon pisum build 2.1
TNFα 18hr. Control. CHX 18hr. TNFα+ CHX 18hr. TNFα: 18 18hr (KDa) PARP. Cleaved. Cleaved. Cleaved. Caspase3. Pellino3 shrna. Control shrna.
 Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
4) Please cite Dagda et al J Biol Chem 284: , for any publications or presentations resulting from use or modification of the macro.
 Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Exceptionally high cumulative percentage of NUMTs originating from linear mitochondrial DNA molecules in the Hydra magnipapillata genome
 Song et al. BMC Genomics 2013, 14:447 RESEARCH ARTICLE Open Access Exceptionally high cumulative percentage of NUMTs originating from linear mitochondrial DNA molecules in the Hydra magnipapillata genome
Song et al. BMC Genomics 2013, 14:447 RESEARCH ARTICLE Open Access Exceptionally high cumulative percentage of NUMTs originating from linear mitochondrial DNA molecules in the Hydra magnipapillata genome
The Diffusion Coefficient for PGK Folding in Eukaryotic Cells
 Biophysical Journal, Volume 99 Supporting Material The Diffusion Coefficient for PGK Folding in Eukaryotic Cells Apratim Dhar, Simon Ebbinghaus, Zhen Shen, Tripta Mishra, and Martin Gruebele 1 Supplementary
Biophysical Journal, Volume 99 Supporting Material The Diffusion Coefficient for PGK Folding in Eukaryotic Cells Apratim Dhar, Simon Ebbinghaus, Zhen Shen, Tripta Mishra, and Martin Gruebele 1 Supplementary
Small RNA in rice genome
 Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis
 Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09606 Chen Li-Qing et al. A novel class of sugar transporters. Supplementary Figure 1. Confocal imaging of FRET sensor localization in HEK293T cells. The subcellular localization of the
doi:10.1038/nature09606 Chen Li-Qing et al. A novel class of sugar transporters. Supplementary Figure 1. Confocal imaging of FRET sensor localization in HEK293T cells. The subcellular localization of the
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES
 Molecular Biology-2018 1 Definitions: RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES Heterologues: Genes or proteins that possess different sequences and activities. Homologues: Genes or proteins that
Molecular Biology-2018 1 Definitions: RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES Heterologues: Genes or proteins that possess different sequences and activities. Homologues: Genes or proteins that
Comparative study of Hippo pathway genes in cellular conveyor belts of a ctenophore and a cnidarian
 DOI 10.1186/s13227-016-0041-y EvoDevo RESEARCH Open Access Comparative study of Hippo pathway genes in cellular conveyor belts of a ctenophore and a cnidarian Alicia Coste, Muriel Jager, Jean Philippe
DOI 10.1186/s13227-016-0041-y EvoDevo RESEARCH Open Access Comparative study of Hippo pathway genes in cellular conveyor belts of a ctenophore and a cnidarian Alicia Coste, Muriel Jager, Jean Philippe
PGA: A Program for Genome Annotation by Comparative Analysis of. Maximum Likelihood Phylogenies of Genes and Species
 PGA: A Program for Genome Annotation by Comparative Analysis of Maximum Likelihood Phylogenies of Genes and Species Paulo Bandiera-Paiva 1 and Marcelo R.S. Briones 2 1 Departmento de Informática em Saúde
PGA: A Program for Genome Annotation by Comparative Analysis of Maximum Likelihood Phylogenies of Genes and Species Paulo Bandiera-Paiva 1 and Marcelo R.S. Briones 2 1 Departmento de Informática em Saúde
Deciphering the role of chondroitin sulphate in increasing the transfection efficiency of amphipathic peptide basednanocomplexes
 Deciphering the role of chondroitin sulphate in increasing the transfection efficiency of amphipathic peptide basednanocomplexes aniel Nisakar 1#, Manika Vij 1,2#, Tanuja Pandey 1, Poornemaa Natarajan
Deciphering the role of chondroitin sulphate in increasing the transfection efficiency of amphipathic peptide basednanocomplexes aniel Nisakar 1#, Manika Vij 1,2#, Tanuja Pandey 1, Poornemaa Natarajan
Introductory Microbiology Dr. Hala Al Daghistani
 Introductory Microbiology Dr. Hala Al Daghistani Why Study Microbes? Microbiology is the branch of biological sciences concerned with the study of the microbes. 1. Microbes and Man in Sickness and Health
Introductory Microbiology Dr. Hala Al Daghistani Why Study Microbes? Microbiology is the branch of biological sciences concerned with the study of the microbes. 1. Microbes and Man in Sickness and Health
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/312/5780/1653/dc1 Supporting Online Material for The Xist RNA Gene Evolved in Eutherians by Pseudogenization of a Protein-Coding Gene Laurent Duret,* Corinne Chureau,
www.sciencemag.org/cgi/content/full/312/5780/1653/dc1 Supporting Online Material for The Xist RNA Gene Evolved in Eutherians by Pseudogenization of a Protein-Coding Gene Laurent Duret,* Corinne Chureau,
a-dB. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Unit 5: Cell Division and Development Guided Reading Questions (45 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 12 The Cell Cycle Unit 5: Cell Division and Development Guided
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 12 The Cell Cycle Unit 5: Cell Division and Development Guided
CGS 5991 (2 Credits) Bioinformatics Tools
 CAP 5991 (3 Credits) Introduction to Bioinformatics CGS 5991 (2 Credits) Bioinformatics Tools Giri Narasimhan 8/26/03 CAP/CGS 5991: Lecture 1 1 Course Schedules CAP 5991 (3 credit) will meet every Tue
CAP 5991 (3 Credits) Introduction to Bioinformatics CGS 5991 (2 Credits) Bioinformatics Tools Giri Narasimhan 8/26/03 CAP/CGS 5991: Lecture 1 1 Course Schedules CAP 5991 (3 credit) will meet every Tue
Supplementary Information
 Supplementary Information Supplementary Figure 1: MEIOC is a novel evolutionarily conserved protein Alignments of some representative MEIOC amino acid sequences encoded by orthologs were processed with
Supplementary Information Supplementary Figure 1: MEIOC is a novel evolutionarily conserved protein Alignments of some representative MEIOC amino acid sequences encoded by orthologs were processed with
Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc
 OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
Phylogenetic analysis of uroporphyrinogen III synthase (UROS) gene
 www.bioinformation.net Hypothesis Volume 8(25) Phylogenetic analysis of uroporphyrinogen III synthase (UROS) gene Abjal Pasha Shaik 1,$, *, Abbas H Alsaeed 1$ & Asma Sultana 2$ 1Department of Clinical
www.bioinformation.net Hypothesis Volume 8(25) Phylogenetic analysis of uroporphyrinogen III synthase (UROS) gene Abjal Pasha Shaik 1,$, *, Abbas H Alsaeed 1$ & Asma Sultana 2$ 1Department of Clinical
19/10/2015. Response to environmental manipulation. Its not as simple as it seems!
 Jackie Mitchell Department of Basic and Clinical Neuroscience - IoPPN Why do we need model genetic systems? What does it take to be a good model? Common in vivo model species Genetic manipulations (Lecture
Jackie Mitchell Department of Basic and Clinical Neuroscience - IoPPN Why do we need model genetic systems? What does it take to be a good model? Common in vivo model species Genetic manipulations (Lecture
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
RNA polymerase II clustering through carboxyterminal domain phase separation
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41594-018-0112-y In the format provided by the authors and unedited. RNA polymerase II clustering through carboxyterminal domain phase separation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41594-018-0112-y In the format provided by the authors and unedited. RNA polymerase II clustering through carboxyterminal domain phase separation
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Computational Biology, Part 24 Clustering and Unmixing of Subcellular Patterns
 Computational Biology, Part 24 Clustering and Unmixing of Subcellular Patterns Robert F. Murphy Copyright 1996, 1999, 2000-2009. All rights reserved. Unsupervised Learning to Identify High-Resolution Protein
Computational Biology, Part 24 Clustering and Unmixing of Subcellular Patterns Robert F. Murphy Copyright 1996, 1999, 2000-2009. All rights reserved. Unsupervised Learning to Identify High-Resolution Protein
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission Name: Email: Lab Antibody Name: Target: Company/ Source: Catalog Number, database ID, laboratory Lot Number Antibody Description: Target Description:
ENCODE DCC Antibody Validation Document Date of Submission Name: Email: Lab Antibody Name: Target: Company/ Source: Catalog Number, database ID, laboratory Lot Number Antibody Description: Target Description:
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Pyrobayes: an improved base caller for SNP discovery in pyrosequences
 Pyrobayes: an improved base caller for SNP discovery in pyrosequences Aaron R Quinlan, Donald A Stewart, Michael P Strömberg & Gábor T Marth Supplementary figures and text: Supplementary Figure 1. The
Pyrobayes: an improved base caller for SNP discovery in pyrosequences Aaron R Quinlan, Donald A Stewart, Michael P Strömberg & Gábor T Marth Supplementary figures and text: Supplementary Figure 1. The
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supporting Information
 Supporting Information Wang et al. 10.1073/pnas.0804871105 SI Materials and Methods Cell Culture and Transfection. Human neuroblastoma M17 cells were grown as described before (1). Transfection was performed
Supporting Information Wang et al. 10.1073/pnas.0804871105 SI Materials and Methods Cell Culture and Transfection. Human neuroblastoma M17 cells were grown as described before (1). Transfection was performed
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Procedure to Create NCBI KOGS
 Procedure to Create NCBI KOGS full details in: Tatusov et al (2003) BMC Bioinformatics 4:41. 1. Detect and mask typical repetitive domains Reason: masking prevents spurious lumping of non-orthologs based
Procedure to Create NCBI KOGS full details in: Tatusov et al (2003) BMC Bioinformatics 4:41. 1. Detect and mask typical repetitive domains Reason: masking prevents spurious lumping of non-orthologs based
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Cellular Signalling 24 (2012) Contents lists available at SciVerse ScienceDirect. Cellular Signalling
 Cellular Signalling 24 (2012) 758 769 Contents lists available at SciVerse ScienceDirect Cellular Signalling journal homepage: www.elsevier.com/locate/cellsig Unexpected diversity in Shisa-like proteins
Cellular Signalling 24 (2012) 758 769 Contents lists available at SciVerse ScienceDirect Cellular Signalling journal homepage: www.elsevier.com/locate/cellsig Unexpected diversity in Shisa-like proteins
Tiffany Samaroo MB&B 452a December 8, Take Home Final. Topic 1
 Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Exploring metazoan evolution through dynamic and holistic changes in protein families and domains
 Wang et al. BMC Evolutionary Biology 2012, 12:138 RESEARCH ARTICLE Open Access Exploring metazoan evolution through dynamic and holistic changes in protein families and domains Zhengyuan Wang 1, Dante
Wang et al. BMC Evolutionary Biology 2012, 12:138 RESEARCH ARTICLE Open Access Exploring metazoan evolution through dynamic and holistic changes in protein families and domains Zhengyuan Wang 1, Dante
a-fB. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Reconstructing Mitochondrial Evolution?? Morphological Diversity. Mitochondrial Diversity??? What is your definition of a mitochondrion??
 Reconstructing Mitochondrial Evolution?? What is your definition of a mitochondrion?? Morphological Diversity Mitochondria as we all know them: Suprarenal gland Liver cell Plasma cell Adrenal cortex Mitochondrial
Reconstructing Mitochondrial Evolution?? What is your definition of a mitochondrion?? Morphological Diversity Mitochondria as we all know them: Suprarenal gland Liver cell Plasma cell Adrenal cortex Mitochondrial
Cell Structure and Function. The Development of Cell Theory
 Cell Structure and Function The Development of Cell Theory Hooke's original microscope, that he used at Oxford University, in 1665. Robert Hooke first examined a thin piece of cork (see below), and coined
Cell Structure and Function The Development of Cell Theory Hooke's original microscope, that he used at Oxford University, in 1665. Robert Hooke first examined a thin piece of cork (see below), and coined
11/24/13. Science, then, and now. Computational Structural Bioinformatics. Learning curve. ECS129 Instructor: Patrice Koehl
 Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://www.cs.ucdavis.edu/~koehl/teaching/ecs129/index.html koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://www.cs.ucdavis.edu/~koehl/teaching/ecs129/index.html koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
SUPPLEMENTARY INFORMATION
 GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
Quantitative and qualitative analyses. of in-paralogs
 Quantitative and qualitative analyses of in-paralogs Dissertation zur Erlangung des naturwissentschaflichen Doktorgrades der Bayerischen Julius-Maximilians Universität Würzburg vorgelegt von Stanislav
Quantitative and qualitative analyses of in-paralogs Dissertation zur Erlangung des naturwissentschaflichen Doktorgrades der Bayerischen Julius-Maximilians Universität Würzburg vorgelegt von Stanislav
DNA. Announcements. Invertebrates DNA. DNA Code. DNA Molecule of inheritance. & Protein Synthesis. Midterm II is Friday
 Midterm II is Friday Announcements DNA & Protein Synthesis Shannon and Val Review session on Wednesday April 5 from 5:30 to 6:30pm in 2301 Tolman Invertebrates DNA Molecule of inheritance. Contains code
Midterm II is Friday Announcements DNA & Protein Synthesis Shannon and Val Review session on Wednesday April 5 from 5:30 to 6:30pm in 2301 Tolman Invertebrates DNA Molecule of inheritance. Contains code
Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction. Lesser Tenrec (Echinops telfairi)
 Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction Lesser Tenrec (Echinops telfairi) Goals: 1. Use phylogenetic experimental design theory to select optimal taxa to
Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction Lesser Tenrec (Echinops telfairi) Goals: 1. Use phylogenetic experimental design theory to select optimal taxa to
CHAPTER 1 INTRODUCTION TO CELLS 2009 Garland Science Publishing 3 rd Edition
 Unity and Diversity of Cells 1-1 The smallest unit of life is a(n) (a) DNA molecule. (b) cell. (c) organelle. (d) virus. (e) protein. CHAPTER 1 INTRODUCTION TO CELLS 2009 Garland Science Publishing 3 rd
Unity and Diversity of Cells 1-1 The smallest unit of life is a(n) (a) DNA molecule. (b) cell. (c) organelle. (d) virus. (e) protein. CHAPTER 1 INTRODUCTION TO CELLS 2009 Garland Science Publishing 3 rd
Synonymous Codon Substitution Matrices
 Synonymous Codon Substitution Matrices Adrian Schneider, Gaston H. Gonnet, and Gina M. Cannarozzi Computational Biology Research Group, Institute for Computational Science, ETH Zürich, Universitätstrasse
Synonymous Codon Substitution Matrices Adrian Schneider, Gaston H. Gonnet, and Gina M. Cannarozzi Computational Biology Research Group, Institute for Computational Science, ETH Zürich, Universitätstrasse
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Zn 2+ -binding sites in USP18. (a) The two molecules of USP18 present in the asymmetric unit are shown. Chain A is shown in blue, chain B in green. Bound Zn 2+ ions are shown as
Supplementary Figure 1 Zn 2+ -binding sites in USP18. (a) The two molecules of USP18 present in the asymmetric unit are shown. Chain A is shown in blue, chain B in green. Bound Zn 2+ ions are shown as
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools. Evaluation. Course Homepage.
 CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Biology 160 Cell Lab. Name Lab Section: 1:00pm 3:00 pm. Student Learning Outcomes:
 Biology 160 Cell Lab Name Lab Section: 1:00pm 3:00 pm Student Learning Outcomes: Upon completion of today s lab you will be able to do the following: Properly use a compound light microscope Discuss the
Biology 160 Cell Lab Name Lab Section: 1:00pm 3:00 pm Student Learning Outcomes: Upon completion of today s lab you will be able to do the following: Properly use a compound light microscope Discuss the
Application of new distance matrix to phylogenetic tree construction
 Application of new distance matrix to phylogenetic tree construction P.V.Lakshmi Computer Science & Engg Dept GITAM Institute of Technology GITAM University Andhra Pradesh India Allam Appa Rao Jawaharlal
Application of new distance matrix to phylogenetic tree construction P.V.Lakshmi Computer Science & Engg Dept GITAM Institute of Technology GITAM University Andhra Pradesh India Allam Appa Rao Jawaharlal
Multiple Sequence Alignment. Sequences
 Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Supplemental Table 1. Primers used for cloning and PCR amplification in this study
 Supplemental Table 1. Primers used for cloning and PCR amplification in this study Target Gene Primer sequence NATA1 (At2g393) forward GGG GAC AAG TTT GTA CAA AAA AGC AGG CTT CAT GGC GCC TCC AAC CGC AGC
Supplemental Table 1. Primers used for cloning and PCR amplification in this study Target Gene Primer sequence NATA1 (At2g393) forward GGG GAC AAG TTT GTA CAA AAA AGC AGG CTT CAT GGC GCC TCC AAC CGC AGC
Robust Classification of Subcellular Location Patterns in Fluorescence Microscope Images
 Robust Classification of Subcellular Location Patterns in Fluorescence Microscope Images Robert F. Murphy, Meel Velliste, and Gregory Porreca Departments of Biological Sciences and Biomedical Engineering,
Robust Classification of Subcellular Location Patterns in Fluorescence Microscope Images Robert F. Murphy, Meel Velliste, and Gregory Porreca Departments of Biological Sciences and Biomedical Engineering,
Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors
 Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors Arli Parikesit, Peter F. Stadler, Sonja J. Prohaska Bioinformatics Group Institute of Computer Science University
Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors Arli Parikesit, Peter F. Stadler, Sonja J. Prohaska Bioinformatics Group Institute of Computer Science University
Chapter # EVOLUTION AND ORIGIN OF NEUROFIBROMIN, THE PRODUCT OF THE NEUROFIBROMATOSIS TYPE 1 (NF1) TUMOR-SUPRESSOR GENE
 142 Part 5 Chapter # EVOLUTION AND ORIGIN OF NEUROFIBROMIN, THE PRODUCT OF THE NEUROFIBROMATOSIS TYPE 1 (NF1) TUMOR-SUPRESSOR GENE Golovnina K. *1, Blinov A. 1, Chang L.-S. 2 1 Institute of Cytology and
142 Part 5 Chapter # EVOLUTION AND ORIGIN OF NEUROFIBROMIN, THE PRODUCT OF THE NEUROFIBROMATOSIS TYPE 1 (NF1) TUMOR-SUPRESSOR GENE Golovnina K. *1, Blinov A. 1, Chang L.-S. 2 1 Institute of Cytology and
JEPSLD: A JUDGMENTAL EUKARYOTIC PROTEIN SUBCELLULAR LOCATION DATABASE. Sanjeev Patra
 JEPSLD: A JUDGMENTAL EUKARYOTIC PROTEIN SUBCELLULAR LOCATION DATABASE by Sanjeev Patra A thesis submitted to the Faculty of the University of Delaware in partial fulfillment of the requirements for the
JEPSLD: A JUDGMENTAL EUKARYOTIC PROTEIN SUBCELLULAR LOCATION DATABASE by Sanjeev Patra A thesis submitted to the Faculty of the University of Delaware in partial fulfillment of the requirements for the
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Last time: Obtaining information from a cloned gene
 Last time: Obtaining information from a cloned gene Objectives: 1. What is the biochemical role of the gene? 2. Where and when is the gene expressed (transcribed)? 3. Where and when is the protein made?
Last time: Obtaining information from a cloned gene Objectives: 1. What is the biochemical role of the gene? 2. Where and when is the gene expressed (transcribed)? 3. Where and when is the protein made?
Emergence of Xin Demarcates a Key Innovation in Heart Evolution
 Emergence of Xin Demarcates a Key Innovation in Heart Evolution Shaun E. Grosskurth, Debashish Bhattacharya, Qinchuan Wang, Jim Jung-Ching Lin* Department of Biology, University of Iowa, Iowa City, Iowa,
Emergence of Xin Demarcates a Key Innovation in Heart Evolution Shaun E. Grosskurth, Debashish Bhattacharya, Qinchuan Wang, Jim Jung-Ching Lin* Department of Biology, University of Iowa, Iowa City, Iowa,
What is a cell? (*Know the parts of the microscope!)
 Cells What is a cell? All living things have cells whether it is one or many! Therefore, a cell is the basic unit of all life. The invention of the microscope was pivotal to the study of cell biology.
Cells What is a cell? All living things have cells whether it is one or many! Therefore, a cell is the basic unit of all life. The invention of the microscope was pivotal to the study of cell biology.
SUPPLEMENTARY EXPERIMENTAL PROCEDURES
 SUPPLEMENTARY EXPERIMENTAL PROCEDURES Yeast two-hybrid assay The yeast two hybrid assay was performed according to described in Assmann (2006) [01] and was performed with the baits SCOCO (2-82) and SCOCO
SUPPLEMENTARY EXPERIMENTAL PROCEDURES Yeast two-hybrid assay The yeast two hybrid assay was performed according to described in Assmann (2006) [01] and was performed with the baits SCOCO (2-82) and SCOCO
CLASSIFICATION OF LIVING THINGS
 CLASSIFICATION OF LIVING THINGS 1. Taxonomy The branch of biology that deals with the classification of living organisms About 1.8 million species of plants and animals have been identified. Some scientists
CLASSIFICATION OF LIVING THINGS 1. Taxonomy The branch of biology that deals with the classification of living organisms About 1.8 million species of plants and animals have been identified. Some scientists
Full-length GlpG sequence was generated by PCR from E. coli genomic DNA. (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
 Supplementary Methods Protein expression and purification Full-length GlpG sequence was generated by PCR from E. coli genomic DNA (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
Supplementary Methods Protein expression and purification Full-length GlpG sequence was generated by PCR from E. coli genomic DNA (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
Warm Up. What are some examples of living things? Describe the characteristics of living things
 Warm Up What are some examples of living things? Describe the characteristics of living things Objectives Identify the levels of biological organization and explain their relationships Describe cell structure
Warm Up What are some examples of living things? Describe the characteristics of living things Objectives Identify the levels of biological organization and explain their relationships Describe cell structure
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10244 a O07391_MYCAV/127-243 NLPC_HAEIN/80-181 SPR_SHIFL/79-183 P74160_SYNY3/112-245 O24914_HELPY/301-437 Q51835_PORGI/68-178 DPP6_BACSH/163-263 YKFC_BACSU/185-292 YDHO_ECOLI/153-263
doi:10.1038/nature10244 a O07391_MYCAV/127-243 NLPC_HAEIN/80-181 SPR_SHIFL/79-183 P74160_SYNY3/112-245 O24914_HELPY/301-437 Q51835_PORGI/68-178 DPP6_BACSH/163-263 YKFC_BACSU/185-292 YDHO_ECOLI/153-263
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Civelekoglu-Scholey et al., http://www.jcb.org/cgi/content/full/jcb.200908150/dc1 Figure S1. Spindle dynamics in Drosophila embryos and in vitro gliding
Supplemental material THE JOURNAL OF CELL BIOLOGY Civelekoglu-Scholey et al., http://www.jcb.org/cgi/content/full/jcb.200908150/dc1 Figure S1. Spindle dynamics in Drosophila embryos and in vitro gliding
Introduction to protein alignments
 Introduction to protein alignments Comparative Analysis of Proteins Experimental evidence from one or more proteins can be used to infer function of related protein(s). Gene A Gene X Protein A compare
Introduction to protein alignments Comparative Analysis of Proteins Experimental evidence from one or more proteins can be used to infer function of related protein(s). Gene A Gene X Protein A compare
Evolutionary Rate Covariation of Domain Families
 Evolutionary Rate Covariation of Domain Families Author: Brandon Jernigan A Thesis Submitted to the Department of Chemistry and Biochemistry in Partial Fulfillment of the Bachelors of Science Degree in
Evolutionary Rate Covariation of Domain Families Author: Brandon Jernigan A Thesis Submitted to the Department of Chemistry and Biochemistry in Partial Fulfillment of the Bachelors of Science Degree in
7.06 Problem Set
 7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
 DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
7.06 Cell Biology EXAM #3 April 21, 2005
 7.06 Cell Biology EXAM #3 April 21, 2005 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #3 April 21, 2005 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
Genomes and Their Evolution
 Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
The MANTiS Manual. Contents. MANTiS Version 1.1
 The MANTiS Manual MANTiS Version 1.1 Contents Connection to the MANTiS database... 2 Memory settings... 2 Main functionalities... 2 Character Mapping View... 4 Genome content View... 5 Biological processes
The MANTiS Manual MANTiS Version 1.1 Contents Connection to the MANTiS database... 2 Memory settings... 2 Main functionalities... 2 Character Mapping View... 4 Genome content View... 5 Biological processes
Cellular basis of life History of cell Biology Year Name of the scientist Importance
 Cellular basis of life History of cell Biology Year Name of the scientist Importance 1590 Jansen 1650 Anton van Leeuwenhoek 1665 Robert Hooke 1831 Matthias Schleiden 1831 Theodore Schwann 1855 Rudolf Virchow
Cellular basis of life History of cell Biology Year Name of the scientist Importance 1590 Jansen 1650 Anton van Leeuwenhoek 1665 Robert Hooke 1831 Matthias Schleiden 1831 Theodore Schwann 1855 Rudolf Virchow
Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species
 Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species Zhiqiang Li 1 and Peter Z. Revesz 1,a 1 Department of Computer Science, University of Nebraska-Lincoln, Lincoln, NE,
Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species Zhiqiang Li 1 and Peter Z. Revesz 1,a 1 Department of Computer Science, University of Nebraska-Lincoln, Lincoln, NE,
Supporting Information
 Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
