The structural basis of Edc3- and Scd6-mediated activation of the Dcp1:Dcp2 mrna decapping complex
|
|
|
- Jody Boyd
- 6 years ago
- Views:
Transcription
1 The EMO Journal (2012) 31, & 2012 European Molecular iology Organization ll Rights Reserved /12 The structural basis of - and Scd6-mediated activation of the Dcp1: mrn decapping complex THE EMO JOURNL Simon Fromm 1, Vincent Truffault 2, Julia Kamenz 3, Joerg E raun 2, Niklas Hoffmann 1, Elisa Izaurralde 2 and Remco Sprangers 1, * 1 Max Planck Institute for Developmental iology, Tübingen, Germany, 2 Department of iochemistry, Max Planck Institute for Developmental iology, Tübingen, Germany and 3 Friedrich Miescher Laboratory of the Max Planck Society, Tübingen, Germany The Dcp1: decapping complex catalyses the removal of the mrn 5 0 cap structure. ctivator proteins, including (enhancer of decapping 3), modulate its activity. Here, we solved the structure of the yeast LSm domain in complex with a short helical leucine-rich motif (HLM) from. The motif interacts with the monomeric LSm domain in an unprecedented manner and recognizes a noncanonical binding surface. ased on the structure, we identified additional HLMs in the disordered -terminal extension of that can interact with. Moreover, the LSm domain of the -related protein Scd6 competes with for the interaction with these HLMs. We show that both and Scd6 stimulate decapping in vitro, presumably by preventing the Dcp1: complex from adopting an inactive conformation. In addition, we show that the -terminal HLMs in are necessary for the localization of the Dcp1: decapping complex to P-bodies in vivo. Unexpectedly, in contrast to yeast, in metazoans the HLM is found in Dcp1, suggesting that details underlying the regulation of mrn decapping changed throughout evolution. The EMO Journal (2012) 31, doi: / emboj ; Published online 15 November 2011 Subject ategories: RN; structural biology Keywords: mrn decapping; proteins; RN; structural biology Introduction The cleavage of the mrn 5 0 cap structure (decapping) is an important step in gene expression, as it removes a transcript from the translational pool. Therefore, it is essential to tightly regulate the activity of the enzymes that perform the decapping reaction as premature or incomplete decapping could interfere with cellular homeostasis. The mrn decapping reaction is catalysed by the enzyme (van Dijk et al, 2002; Wang et al, 2002), which works in concert with its *orresponding author. Max Planck Institute for Developmental iology, Spemannstr. 35, Tübingen 72076, Germany. Tel.: þ ; Fax: þ ; remco.sprangers@tuebingen.mpg.de Received: 26 pril 2011; accepted: 12 October 2011; published online: 15 November 2011 prime activator Dcp1. Dcp1: activity is regulated in vivo and in vitro by several proteins, including the enhancers of decapping 1 4 (Edc1 4), Pat1 and the LSm1 7 complex (oeck et al, 1998; onnerot et al, 2000; ouveret et al, 2000; Kshirsagar and Parker, 2004; Fenger-Gron et al, 2005; Haas et al, 2010; orja et al, 2011). Initial insights into the mechanism and regulation of the decapping complex have been provided by a number of highresolution crystal structures. Dcp1 (She et al, 2004) folds into an EVH1 domain (allebaut, 2002) and interacts tightly with (She et al, 2006, 2008) via an N-terminal helical extension that is unique to Dcp1. The structure of displays two folded domains, an N-terminal a-helical regulatory domain followed by a catalytic or Nudix domain (She et al, 2006) (Figure 1). Interestingly, the three different crystal structures of (She et al, 2006, 2008) (Supplementary Figure S1) show that the catalytic domain can adopt multiple, significantly different orientations with respect to the regulatory domain. In general, the available Dcp1: structures can be divided into open and closed forms. In solution, it has been shown that different structural states are sampled during the enzymatic cycle, and the compactness of the decapping complex increases when nucleotides are added (She et al, 2008). These observations suggest that the active state is more closed, whereas the inactive state is in a more open conformation. More recently, however, it was noted that the residues involved in substrate binding are not clustered in any of the known structures (Floor et al, 2010). This suggests that the closed structure, as observed in the high-resolution crystal structure, does not represent the catalytically active form of the decapping complex (Floor et al, 2010) and that additional structural states of the enzyme must exist. The residues -terminal to the structured catalytic domain are rich in proline residues (Gaudon et al, 1999), harbour low overall sequence conservation, and are predicted to be largely intrinsically disordered (Figure 1). No functional data are available for this region, apart from a 50 amino acid long sequence close to the -terminal part of the catalytic domain that has been reported to interact with in yeast (Harigaya et al, 2010). In a cellular context, the Dcp1: decapping enzyme is part of an RNP (ribonucleo protein) complex that can vary in composition (Franks and Lykke-ndersen, 2008). Under certain conditions, it has been observed that these decapping complexes can cluster into large assemblies called mrn processing bodies or P-bodies, which can be visualized using fluorescence microscopy (Sheth and Parker, 2003; Franks and Lykke-ndersen, 2008). How the clustering of the mrn degradation machinery influences cellular function is unclear, but most of the proteins found in P-bodies are involved in translational repression and mrn degradation (Eulalio et al, 2007). However, the molecular details of the inter- & 2012 European Molecular iology Organization The EMO Journal VOL 31 NO
2 1 127 Dcp1 EVH LSm FDF HLM HLM-1 to HLM Regulatory atalytic Proline rich -terminal extension Scd6 YjeF-N LSm Proline rich FDF RGG β2 β5 β3 β4 β1 N α1 Variable loop D MW His Dcp E 15 N (p.p.m.) R38 V48 L35 L37 G8 K45 L G23 D54 4 V3 Q36 I57 L55 L58 N25 D29 N27 I52 Y H (p.p.m.) Figure 1 Interaction between and. () Domain organization of the S. pombe Dcp1,,, and Scd6 (Tral/Sum2) proteins. Folded protein domains are coloured in yellow (Dcp1), light green ( regulatory domain), dark green ( catalytic domain), cyan (YjeF-N domain), and blue (Lsm domain). The FDF motif is coloured orange, the linear sequence motifs that interact with the and Scd6 LSm domains (HLM-1, HLM-1 to HLM-5; see below) are coloured red. () The solution structure of the S. pombe LSm domain. Secondary structure elements are indicated. The short N-terminal helical turn is coloured green. ll figures displaying protein structures are made using Pymol ( () n overlay of the structures of the S. pombe LSm domain (blue/green) with the LSm domains from D. melanogaster (brown, PD: 2RM4) and H. sapiens (yellow, PD: 2V8). The variable region that is absent in the yeast LSm domain is indicated. (D) Identification of the residues required for interaction with. His 6 Dcp1, was co-expressed with different versions of (see Supplementary data) and supplemented with separately expressed LSm domain. The mixed cell lysates were purified using Ni-affinity chromatography. Dcp1 interacts with in all cases, indicating that all constructs were properly folded. Only the Dcp1: complex that contains residues binds the LSm domain (lane 3). (E) 1 H- 15 N NMR spectra of the 15 N-labelled S. pombe LSm domain in the absence (blue) and presence (red) of an unlabelled peptide corresponding to residues in. Some assignments in the isolated LSm domain (blue) are indicated. molecular interactions within these large protein RN complexes are poorly understood. One of the proteins that influences P-body formation is the enhancer of decapping 3 (), and deletion results in a reduction in the number and size of these cellular foci (Decker et al, 2007; Ling et al, 2008). In yeast, both the LSm domain and the YjeF-N dimerization domain of have been shown to be functionally important for P-body formation (Decker et al, 2007). Proteins of the Scd6 family (Lsm13 15; Trailer hitch/tral in Drosophila melanogaster; R-1 in aenorhabditis elegans; Sum2 in Schizosaccharomyces pombe) share a number of features with (Figure 1). First, both proteins localize to P-bodies (Tritschler et al, 2007, 2008). Second, both and Scd6 contain an FDF motif that directly interacts with the DED box helicase Dhh1 in a competitive fashion (Tritschler et al, 2008, 2009a). Third, both proteins contain an N-terminal LSm domain (lbrecht and Lengauer, 2004). Finally, both proteins are part of the same RNP and interact with in yeast (Decourty et al, 2008; Tarassov et al, 2008; Nissan et al, 2010). The cellular function of Scd6 is not well established, although Scd6 seems to play a role in inhibiting translation initiation in yeast (Nissan et al, 2010), and Scd6 orthologues have been shown to associate with translationally repressed mrns (Decker and Parker, 2006). Interestingly, a functional redundancy between and Scd6 has been proposed (Decourty et al, 2008). However, in contrast to, Scd6 has not been shown to modulate the catalytic activity of the Dcp1: decapping complex in vitro. Here, we address how and Scd6 interact with the Dcp1: decapping complex and thereby influence its enzymatic activity. To this end, we solved the structure of the S. pombe LSm domain in complex with a sequence motif derived from. ased on this structure, we identified additional conserved sequence motifs in that interact with the LSm domains of Scd6 and in a mutually exclusive manner. We show that deletion of the -terminal motifs of that interact with the and Scd6 LSm domains results in the failure of the Dcp1: decapping complex to localize to P-bodies. In summary, our data reveal the molecular details of a network of competing interactions that can modulate Dcp1: decapping activity and cellular localization. Results S. pombe residues interact with the LSm domain The protein is composed of an N-terminal LSm domain (Like-Sm) (lbrecht and Lengauer, 2004; Tritschler et al, 2007) that is connected to a -terminal YjeF-N domain (Ling et al, 2008) (Figure 1). The linker region between the two folded domains is predicted to be unstructured and contains an FDF motif that interacts with the RN helicase Dhh1/Me31b (Tritschleret al, 2007, 2009a). In yeast, the LSm domain has been shown to interact with a 50 amino acid long region in that follows the catalytic domain 280 The EMO Journal VOL 31 NO & 2012 European Molecular iology Organization
3 (Harigaya et al, 2010). To obtain insight into this interaction, we solved the solution structure of the S. pombe LSm domain. The LSm domain folds into a canonical five-stranded b-sheet that is preceded by a short a-helical turn (Figure 1; Supplementary Figure S2; Supplementary Table SI). The yeast LSm structure is similar to the structures of the D. melanogaster and Homo sapiens LSm domains (Figure 1) (Tritschler et al, 2007) although the variable loop between b-strands 3 and 4 is shorter in the yeast protein. Many (L)Sm proteins form hexa- or heptameric rings (e.g. LSm1 7 and Hfq) (Wilusz and Wilusz, 2005). However, the S. pombe LSm domain is monomeric in solution, as previously described for the D. melanogaster and H. sapiens LSm domains (Tritschler et al, 2007). To identify which residues in are responsible for the interaction with, we performed pull-down experiments with His 6 Dcp1: decapping complexes. We used constructs of increasing length (Figure 1D) and found that a region located between residues 255 and 266 is required for the interaction with (Figure 1D, lane 3 versus lanes 1 and 2). oth in Saccharomyces cerevisiae (Harigaya et al, 2010) and in S. pombe, a region -terminal of the catalytic domain is thus important for the interaction with. In the reverse experiment, we performed NMR titration experiments to identify residues in the LSm domain that interact with. To this end, we added NMR-inactive to 15 N-labelled LSm domain (Figure 1E, blue). residues that come in close spatial proximity with are expected to experience chemical shift perturbations upon binding. Indeed, 450% of the residues in the LSm domain sense the presence of (Figure 1E, blue versus red). The large number of chemical shift changes, however, prevented the accurate identification of an binding site for residues This could reflect a small secondary conformational effect in the LSm domain upon binding, but given the marked sensitivity of amide group chemical shifts to even small structural movements, any such change may be very small. The LSm domain interacts with in an unprecedented manner Previous studies have shown that LSm domains use two distinct modes to interact with their binding partners. In one mode, they establish intermolecular interactions between b-strand 4 of one monomer and b-strand 5 of another monomer (Kambach et al, 1999), resulting in the assembly of a sixor seven-member LSm ring (Wilusz and Wilusz, 2005). The other mode of interaction is between LSm rings and RN, in which binding sites on both the proximal and distal faces have been reported and often involve residues in the loop connecting b-strands 2 and 3 and residues in the loop that connects b-strands 4 and 5 (Toro et al, 2001; Thore et al, 2003; Link et al, 2009; Leung et al, 2011). s such, we expected that the monomeric LSm domain would exploit one of these binding sites to interact with. The NMR titration experiments (Figure 1E) did not reveal which residues in contact directly. To determine the molecular details of this interaction, we assigned the resonances of the complex of the LSm domain and residues and solved the solution structure (Figure 2 and ; Supplementary Figure S2; Supplementary Table SI). In the complex, the LSm domain fold remains unchanged (Supplementary Figure S2 E), whereas residues fold into an amphipathic a-helical structure LSm β5 β2 β2 β3 β5 N N β4 α1 β1 β1 N L261 N S257 L260 D54 L264 F6 I52 β5 L55 I57 F28 α1 β1 β2 T49 β3 N α1 LSm β4 N F6 α1 β2 L260 F28 I57 T49 L264 L55 β1 I52 Hfq LSm S257 L261 D54 β5 Figure 2 The structure of the : complex. () The solution structure of the S. pombe LSm domain in complex with residues residues are in blue and green (see Figure 1), and residues are in red. () view into the binding pocket of the : complex. Sidechains that provide the main intermolecular contacts are displayed with sticks. () n overlay of the : complex (blue, green, and red) with a structure of a canonical Hfq LSm domain (yellow, PD: 1KQ1). The N-terminal helix is significantly shorter in the LSm domain (green) compared with the canonical (L)Sm fold (yellow). The resultant space on the surface that becomes available is used for interaction with the -terminal end of the helix. & 2012 European Molecular iology Organization The EMO Journal VOL 31 NO
4 (shown in red in Figure 2). Due to the interaction, the hydrophobic core of the LSm domain increases significantly in size, which explains the large number of chemical shift changes in the NMR spectrum of the LSm domain upon complex formation with (Figure 1E). The residues in the -binding site that are closest to are located in the N-terminal a-helix (F6), b-strand 2 (F28), the loop between b-strands 4 and 5 (T49 and I52), and b-strand 5 (D54, L55, and I57) (Figure 2). These residues form a hydrophobic surface on the LSm domain that contacts four residues on one side of the helix (S257, L260, L261, and L264) (Figure 2). In particular, the methyl groups of the leucine residues make ample interactions with the aromatic and long hydrophobic residues in and contribute significantly to the tight binding. Indeed, mutating either L260 or L264 in to alanine reduces the strength of the interaction, whereas a double mutation in which both of these leucines are replaced with alanines completely abolishes the : interaction (Supplementary Figure S3). In summary, we identified that residues S257 to L264 form a linear motif that interacts tightly with (Figures 1D, 2 and ). The mode of interaction of the LSm domain with differs significantly from previously reported LSm interactions and presents a novel mode of binding between LSm domains and protein ligands. In canonical (L)Sm proteins an N-terminal a-helix covers the surface of the protein formed by b-strands 2 and 5, and the loop preceding b-strand 5. In the LSm domain of, the corresponding helix is much shorter and contains only a single turn (shown in green for and yellow for one monomer of the hexameric Hfq protein in Figure 2). s a consequence, an interaction surface becomes available that is used to contact the -terminal half of the helix. In addition, the N-terminal half of the helix binds to the central residues in b-strand 5, which is used to contact the neighbouring monomer in multimeric LSm ringlike structures. Therefore, both the shorter N-terminal helix and the monomeric nature of the LSm domain are important characteristics that allow the unprecedented mode of interaction with. lthough a fragment comprising residues possess helical propensity, it is disordered in isolation in solution and only folds into a stable helical conformation in the presence of the LSm domain (Supplementary Figure S3). To probe the relevance of our structure in the context of the active Dcp1: decapping complex, we compared NMR spectra of the Dcp1: (residues 1 289) decapping complex in the absence and presence of the LSm domain. fter the formation of the trimeric complex containing Dcp1: and, resonances that correspond to the helix appear at the same frequency as observed in the dimeric : complex (Supplementary Figure S4). This clearly indicates that the LSm interaction motif is structurally identical in the Dcp1: decapping complex and in the structure of the minimal complex that we present here. contains multiple linear motifs that interact with lose inspection of the residues that interact with revealed that the primary amino acid sequence has a helical propensity. From the structure, it can be inferred that the amino acids of importance are SxxLLxxL (Figure 2). Interestingly, the -terminal extension of contains multiple similar sequence motifs that are conserved between different yeast species (Supplementary Figure S5). In the -terminal extension of S. pombe (residue ), there are at least five of these conserved motifs (Figures 1 and 3; Supplementary Figure S5) located between residues 556 and 563 (SQLLQL), 581 and 589 (SLSLLTLL), 648 and 655 (QFDLLKVS), 693 and 700 (SPGFVKIL), and 721 and 729 (DDHFLSYL). Similar linear sequence motifs were previously identified in from S. cerevisiae (Gaudon et al, 1999). These motifs were termed helical leucine-rich motifs (HLMs) and in analogy we will refer to the linear motifs we identified here in S. pombe as HLM-1 for the first motif (residues ) and HLM-1 to HLM-5 for the -terminal motifs. part from these motifs, the sequence conservation of the -terminus is very low. In addition, the -terminal extension is predicted to be unstructured in isolation, which is confirmed by NMR spectra of residues (Supplementary Figure S6). The presence of several motifs in the -terminal extension that resembled the sequence that interacts with the LSm domain prompted us to probe for interactions between this region of and. To this end, we performed pull-down experiments using the His 6 -tagged LSm domain and the -terminal extension (residues ; Figure 3; Supplementary Figure S6). s a positive control, we used a construct of residues that includes HLM-1 and that strongly interacts with (Figures 1D and 3, lane 1). Interestingly, the -terminal extension shows a strong interaction with the LSm domain (Figure 3, lane 2), which corroborates our hypothesis that this region of contains at least one motif that strongly interacts with. To probe which of the HLM-1 to HLM-5 sequences is responsible for binding, we performed co-purifications with His 6 -tagged and five protein fragments, each of which contained a potential interaction motif (Figure 3, lanes 3 7; Supplementary Figure S6). We observed that the fragment containing HLM-2 (lane 4) interacted with the LSm domain to a similar extent as HLM-1 (lane 1). In addition, a weaker interaction between and a fragment containing HLM-1 was observed (lane 3). In contrast, interactions with the other motifs were not detected in this assay. Hence, we conclude that HLM-2 is the main contributor to the binding between the -terminal extension and. To investigate whether the HLMs that did not interact with in pull-down assays exhibited weak affinity for, we used NMR titration experiments. We followed changes in the spectrum of the 15 N-labelled LSm domain upon addition of the different -terminal motifs (Figure 3). In agreement with the pull-down experiments, both the complete -terminal region (HLM-1 5) and the two isolated motifs (HLM-1 and HLM-2) induce large chemical shift changes in the spectrum (Figure 3, top panels). In addition, we observed chemical shift changes upon the addition of the most -terminal motif (HLM-5), indicating that the last motif also interacts with. The other two motifs in the -terminal tail do not interact significantly with, which can be judged from the very slight (HLM-4) or nonexistent chemical shift changes (HLM-3) in the NMR spectra of the LSm domain. The absence of binding for 282 The EMO Journal VOL 31 NO & 2012 European Molecular iology Organization
5 HLM-1 ( ) HLM-1 ( ) HLM-2 ( ) HLM-3 ( ) HLM-4 ( ) HLM-5 ( ) MW HLM-1 HLM-1 5 HLM-1 HLM-2 HLM-3 HLM-4 HLM-5 MP sequences 15 N (p.p.m.) terminal HLMs N10 R Q K V48 HLM HLM R38 N10 Q36 K45 V48 HLM N R R38 Q36 10 D MW Input PD Input PD Input PD LSm -His FL E 9.1 HLM HLM H (p.p.m.) K45 V48 HLM Yjef Dcp His Dcp1 Dcp1 LSm Dcp1 LSm HLM-1 Yjef-N HLM-1 5 Figure 3 The -terminal region interacts with. () Sequence alignment of different HLMs in S. pombe. The top sequence (HLM-1) is located close to the catalytic domain and is used to solve the : structure (Figure 2). The residues that contact the LSm domain are marked with a red dot. The lower five sequences (HLM-1 to HLM-5) are located in the -terminal extension (Supplementary Figure S5). The residue numbers of the fragments are indicated in brackets. () Pull-down experiments with the - terminally His 6 -tagged LSm domain and MP-tagged LSm interaction motifs. The LSm domain interacts strongly with the HLM-1 sequence (lane 1, Figures 1 and 2). strong interaction is also observed between the last 200 residues (HLM-1 5, lane 2). The latter interaction is mediated through the HLM-2 sequence (lane 4) and through the HLM-1 sequence (lane 3). The interaction with the other HLMs in the -terminus are too weak to be detected under the experimental conditions. The corresponding MP HLM proteins were present in the input of all pull-down experiments (Supplementary Figure S5). () NMR spectra of 15 N-labelled LSm domain without (blue) and with (red) different HLMs derived from the -terminus. Specific interactions are observed upon addition of the complete -terminal extension (HLM-1 5) or upon addition of three (LSM-1, LSM-2, and LSM-5) of the five fragments. Note that the same residues in the LSm are affected upon addition of the HLM-1 sequence (Figure 1E). (D) can interact with two decapping complexes simultaneously. Purified His 6 -tagged Dcp1: (residues 1 289) and untagged Dcp1: were mixed together with different constructs (LSm domain, YjeF-N domain, or FL). The protein mixture was applied to Ni-affinity resin that selected for His 6 -tagged Dcp1 (underlined). The bound protein was eluted from the matrix prior to SDS PGE analysis. Lanes 5 and 6 show that untagged Dcp1 co-purified with His 6 -tagged Dcp1 in the presence of FL. (E) Summary of the : interactions we identified here. The -terminus can interact with the LSm domain (and the Scd6 LSm domain, see below) at multiple sites. The dimer can bind two decapping complexes simultaneously. Scd6 these two motifs (HLM-3 and HLM-4) correlates well with the lack of helical propensity in these sequences (Figure 3). ll motifs that interact with induce similar changes in the NMR spectrum, indicating that all HLMs interact with the LSm domain in a structurally similar way, as displayed in the structure of the :HLM-1 complex (Figure 2 and ). In the reverse experiment, we probed the effect of the LSm domain addition on the spectrum of the disordered 15 N-labelled -terminal residues. s expected, we & 2012 European Molecular iology Organization The EMO Journal VOL 31 NO
6 clearly observed binding and resonances indicative of a- helical structure appeared (Supplementary Figure S6). This shows that the leucine-rich motifs in the disordered -terminus fold into an a-helical conformation when interacting with the LSm domains, as we observed for the HLM-1 sequence. The interaction can promote clustering of decapping complexes Our findings show that contains multiple HLMs that can interact with the LSm domain (Figure 3 and ). Interestingly, forms dimers in solution that are mediated by the -terminal YjeF-N domain (Ling et al, 2008). This observation prompted us to probe if one dimer could also interact with two Dcp1: decapping complexes simultaneously. To test this, we incubated equal amounts of purified His 6 - tagged Dcp1: and untagged Dcp1: in the presence of different versions of the protein. We then purified the protein mixture over Ni-NT resin to select for protein complexes that contained at least one copy of His 6 -tagged Dcp1. When the decapping complex was supplemented with the LSm domain (Figure 3D, lane 1), a complex consisting of His 6 -tagged Dcp1,, and the LSm domain was purified (Figure 3D, lane 2). This result is fully consistent with the experiment shown in Figure 1D (lane 3), in which the LSm domain is shown to co-purify with His 6 Dcp1:. When a mixture of decapping complexes consisting of His 6 Dcp1: and untagged Dcp1: was supplemented with the YjeF-N dimerization domain (Figure 3D, lane 3), the Ni-NT purified complex only contained the His 6 Dcp1: complex (Figure 3D, lane 4). This indicates that the YjeF-N dimerization domain does not directly interact with the Dcp1: decapping complex. However, when full-length (FL) was added to the mixture of His 6 Dcp1: and Dcp1: (Figure 3D, lane 5), the Ni-NT purified complex did contain untagged Dcp1 (Figure 3D, lane 6). This clearly shows that untagged Dcp1 co-purifies with His 6 -tagged Dcp1 in the presence of and FL. This result can be explained by the formation of a complex that contains His 6 Dcp1: coupled to an untagged Dcp1: complex through a FL dimer. These in vitro experiments indicate that one dimer is able to act as a scaffold that can bind two decapping complexes simultaneously. This finding, taken together with the fact that can interact with multiple proteins, suggests a redundant network of interactions between and (Figure 3E). The Scd6 LSm domain interacts with In addition to, other proteins that interact with yeast have been identified (Fromont-Racine et al, 2000; Decourty et al, 2008; Tarassov et al, 2008; Nissan et al, 2010). These include the DED box helicase Dhh1, Pat1, and the LSm domain containing protein Scd6 (Figure 1). Given the similarities between the LSm domains of and Scd6, we tested whether the Scd6: interaction is direct and mediated through the Scd6 LSm domain and the HLMs. To this end, we performed pull-down assays (Figure 4) and found that, in contrast to the LSm domain (Figure 3), the interaction of the Scd6 LSm domain with HLM-1 in was too weak to be detected in this assay. However, using NMR spectroscopy as a more sensitive technique, we observed chemical shift changes in the NMR spectrum of the Scd6 LSm domain upon addition of HLM-1, HLM-1, HLM-2, and HLM-5. This indicates that the Scd6 LSm domain specifically interacts with (Figure 4), albeit with a lower affinity than the LSm domain. MW Input PD Scd6 LSm domain 121 HLM-1 Scd6 LSm domain + HLMs 121 HLM HLM Scd6: MP HLM-1 Scd6 LSm -His 15 N (p.p.m.) HLM HLM H (p.p.m.) HLM Scd Scd Scd6: Figure 4 Interaction between the Scd6 LSm domain and the HLMs. () Pull-down experiment of the Scd6 LSm domain with MP oth proteins are present in the E. coli cell lysate (Input, lane 1); however, the interaction is too weak to allow for co-purification (PD, pull-down, lane 2). s the LSm domain: complex co-purified under the same conditions (Figure 3), we conclude that interacts stronger with than Scd6. () NMR spectra of the 15 N-labelled Scd6 LSm domain in the absence (blue) and presence (red) of HLM-1 and HLM-1 to HLM-5. Specific binding between the Scd6 LSm domains is observed for the same motifs that interact with the LSm domain (Figure 3). () The LSm domain competes with the Scd6 LSm domain for residues (HLM-1). The NMR spectra of the 15 N-labelled Scd6 LSm domain in isolation (blue) and in complex with the HLM-1 sequence (red) are shown. Unlabelled LSm domain was added to the sample (orange, cyan, and green), which resulted in the dissociation of the Scd6: complex due to the formation of the NMR-invisible : complex. This indicates a much higher affinity for the : complex than for the Scd6: complex, as observed in the pull-down assays (Figures 3 and 4) (see also Supplementary Figure S7). 284 The EMO Journal VOL 31 NO & 2012 European Molecular iology Organization
7 Interestingly, motifs that interact with the LSm domain also bind to the Scd6 LSm domain, whereas the motifs that are not recognized by the LSm domain also fail to interact with the Scd6 LSm domain. In addition, the number of residues in Scd6 that experience shift changes during the NMR titration experiments are comparable to the results obtained for. These results strongly suggest a structurally similar mode of interaction. In conclusion, the LSm domains of and Scd6 can interact with the same HLMs in. The and Scd6 LSm domains compete for binding to HLM sequences To confirm the finding that and the Scd6 LSm domains recognize the exact same binding sites in, we performed competition assays (Figure 4). To this end, we prepared a complex of the 15 N-labelled Scd6 LSm domain and the NMRinactive HLM-1 ( residues ). NMR spectra clearly show that the Scd6 LSm domain (Figure 4, blue spectrum) formed a complex with the HLM-1 sequence (Figure 4, red spectrum). stepwise addition of the tighter binding LSm domain resulted in a release of HLM-1 from the Scd6 LSm domain (Figure 4, orange, blue, and green). In a control experiment, where we added the Yjef-N domain, the Scd6: complex does not dissociate (Supplementary Figure S7). In summary, our data show that the and Scd6 LSm domains compete for the same - binding motifs and that both interactions are mutually exclusive. oth and Scd6 stimulate mrn decapping in vitro Previous studies have shown that the interaction of with the Dcp1: decapping complex results in increased catalytic activity (Harigaya et al, 2010; Nissan et al, 2010). Given that the Scd6 LSm domain interacts with in a structurally similar way, we hypothesized that the Scd6 LSm domain should also stimulate Dcp1: decapping activity. To test this, we performed in vitro decapping assays using purified proteins. s expected, the decapping activity of a Dcp1: complex containing FL Dcp1 and residues of (which include the regulatory and catalytic domains and the HLM-1 sequence) was enhanced by addition of the LSm domain (Figure 5, lane 1). Interestingly, the Scd6 Lsm domain also stimulates Dcp1: decapping activity (lane 2). Nevertheless, the activation mediated by Scd6 was weaker, reflecting its lower affinity for (Figures 3 and 4). The stimulatory effect of the LSm domains of or Scd6 required the interaction with HLM-1 in, as that lacks HLM-1 ( 1 243) and that carried mutations in HLM-1 that abolished and Scd6 binding (L260 L264) were not stimulated (Figure 5, lanes 3 6). It should be noted that the decapping activation capability of Scd6 was not previously detected when the decapping complex was supplemented with an equimolar amount of Scd6 (Nissan et al, 2010). This finding reflects the low affinity of the Scd6 LSm domain for. For that reason, we used a 10-fold excess of Scd6 in order to occupy the HLM-1 to a larger extend. To be consistent, we also used a 10-fold molar excess of, although this was not required and a similar stimulation was observed with equimolar amounts. can adopt distinct conformations, termed open and closed, in which the relative domain orientation of the regulatory and catalytic domains varies significantly (Supplementary Figure S1). Interestingly, we noticed that superimposing the LSm domain onto the closed crystal form of Dcp1: (Figure 5) results in extensive steric clashes between and the regulatory domain (Figure 5). In order to interact with or Scd6, the HLM-1 must thus dissociate from the regulatory domain. The loss of this intramolecular interaction most likely results in the opening of the decapping complex (Supplementary Figure S1). In the open conformation the HLM-1 region is freely available for the interaction with the /Scd6 LSm domain (Supplementary Figure S3). Decapping activity compared with unstimulated Dcp1: (%) Dcp1: Dcp1: Dcp1: L2 Scd6 Scd6 Scd Dcp1 HLM-1 LSm Figure 5 mrn decapping activity is stimulated by the interaction of the and Scd6 LSm domains with. () Different versions of the Dcp1: decapping complex were incubated with or without a 10-fold molar excess of the or Scd6 LSm domains. The increase in activity with respect to the corresponding isolated decapping complexes (set to 100% activity) is reported. The activity of the Dcp1: decapping complex that contains the LSm interaction motif (HLM-1) is increased after addition of the (lane 1) or Scd6 (lane 2) LSm domains. If the HLM-1 sequence is deleted (lanes 3 and 4) or mutated such that the LSm domain interaction is abolished (lanes 5 and 6), decapping is no longer stimulated. This clearly shows that the stimulation of mrn decapping is a direct consequence of :LSm domain interaction. () The structure of the Dcp1: complex in the closed and inactive conformation (Supplementary Figure S1, 2QKM) (She et al, 2008). The HLM-1 sequence is disordered in the open conformations of the decapping complex and not visible in the structure (Supplementary Figure S1). () The : structure (blue and red, same orientation as shown in Figure 2) is docked onto the closed Dcp1: structure. This results in extensive steric clashing between the LSm domain and the regulatory domain. The Dcp1: complex can thus not adopt the closed conformation displayed in the crystal structure when bound to /Scd6. & 2012 European Molecular iology Organization The EMO Journal VOL 31 NO
8 Interestingly, it has been suggested that the closed conformation is not the catalytically active conformation (Floor et al, 2010). The binding of activators to the decapping complex could thus destabilize an inactive conformation, which would lead to a higher enzymatic activity. The -terminal HLMs are important for P-body localization in vivo In vivo, components of the mrn degradation machinery localize to P-bodies (Eulalio et al, 2007). has been shown to play an important role in the formation of P-bodies, in addition to its role in activating the Dcp1: decapping complex. udding yeast strains that lack show a strong reduction in the size and number of P-bodies (Decker et al, 2007), suggesting that the protein functions as a scaffold on which components of the mrn degradation machinery assemble. To probe the importance of the interaction between the HLMs and for the proper localization of the Dcp1: complex to P-bodies, we performed fluorescence microscopy studies in the fission yeast S. pombe. We replaced the endogenous dcp2 þ gene either by a -terminally GFPtagged version (dcp2 þ GFP) or by truncated constructs, where HLM-1 5 (dcp2 D GFP), the entire -terminus except the first HLM (dcp2 D GFP), or the -terminus including all HLMs (dcp2 D GFP) was deleted. These versions were integrated in a strain expressing dcp1 þ mherry. Dcp1 mherry and GFP localized to the same cytoplasmic foci, but this localization was abolished in all three truncations and both Dcp1 as well as the truncated proteins were homogenously distributed throughout the cytoplasm (Figure 6). dditionally, we observed that D GFP and D GFP were no longer excluded from the nucleus (Figure 6 and ). These changes in localization were not caused by differences in abundance, as FL and all truncation mutants were expressed to similar levels (Supplementary Figure S8). In order to address whether the absence of the HLM motifs from generally influenced the formation of P-bodies, we asked whether other known components of the mrn degradation machinery were still able to assemble in such foci. However, neither for mherry (Figure 6) nor for Lsm7 mherry (Supplementary Figure S8) we were able to observe an obvious difference in the number or size of cytoplasmic foci when the dcp2 truncation mutants were expressed in place of endogenous dcp2 þ. Furthermore, the absence of Dcp1 and from P-bodies did not impair cell growth (Supplementary Figure S8). Only the truncation of that also removed HLM-1 (dcp2 D GFP) caused a slight but reproducible temperature-dependent growth defect (Supplementary Figure S8). In summary, we conclude that the -terminal HLMs (HLM- 1 5) of, some of which interact strongly with (Figure 3), are essential for the recruitment of the Dcp1: decapping complex to P-bodies, but localization of the decapping complex to P-bodies is dispensable for P-body formation. The deletion of all HLMs, including HLM-1, shows an effect on cellular growth, possibly because in this mutant decapping activity cannot be modulated by and/or Scd6. GFP Dcp1 mherry Merge GFP mherry Merge GFP GFP Δ GFP Δ GFP Δ GFP Δ GFP Δ GFP Δ GFP Figure 6 The -terminal HLMs of are required for P-body localization of Dcp1 and. Fluorescence micrographs of S. pombe cells expressing dcp2 þ GFP or truncated versions thereof in combination with dcp1 þ mherry () or edc3 þ mherry (). GFP co-localizes with Dcp1 mherry () and with mherry () in distinct cytoplasmic foci (P-bodies). () oth Dcp1 and fail to localize to P- bodies, if the -terminal HLMs of are deleted. () mherry still forms P-bodies in all dcp2 mutant strains, indicating that the redundant process of P-body formation is not significantly disrupted by the truncations. ll images were scaled and processed in the same way. The length of the scale bars corresponds to 10 mm. 286 The EMO Journal VOL 31 NO & 2012 European Molecular iology Organization
9 n LSm-interacting motif that mediates binding is present in metazoan Dcp1 The LSm interaction motifs that are found in yeast sequences are not readily identifiable in metazoan sequences. Indeed, the LSm interaction motif that is -terminal to the catalytic domain (HLM-1) is absent in metazoan, and the -terminal extension contains only a few less conserved regions that not clearly resemble HLMs. On the other hand, it has been reported that the D. melanogaster Dcp1 protein interacts with the LSm domains of and Tral (Tritschler et al, 2007, 2008), where the :Dcp1 interaction is mediated through a conserved Dcp1 sequence motif that was termed motif-1 (Tritschler et al, 2009b). This finding might seem to contradict the results we obtained here for the yeast decapping complex, in which interacts with and Scd6. However, close inspection of metazoan Dcp1 sequences (Supplementary Figure S9) revealed that motif-1 (SIFNMLT) has the propensity to fold into an amphipatic helical structure, as observed for the HLMs. To investigate whether the D. melanogaster Dcp1 motif-1 interacts with the D. melanogaster LSm domain, we performed NMR titration experiments. Interestingly, we observed significant chemical shift perturbations in the LSm domain upon addition of a peptide containing the Dcp1 motif-1, which demonstrates a direct interaction (Figure 7). Mapping these changes onto the D. melanogaster LSm structure (Tritschler et al, 2007) clearly shows that the conserved Dcp1 motif-1 binds to the same surface as the LSm interaction motif in the S. pombe : complex (Figure 7). Taken together, these findings show that motif-1 functions like a HLM in metazoan Dcp1. Discussion The decapping enzyme requires the interaction with its protein partners for full decapping activity. These partners, or decapping activators, include Dcp1, which interacts with directly to form the conserved core of the decapping complex and Edc1 4, Scd6/Tral, Pat1, and the Lsm1 7 ring (Parker and Song, 2004; Franks and Lykke-ndersen, 2008). The detailed mechanism by which these activators stimulate decapping has remained largely unknown. Here, we identify the molecular basis of a network of interactions between the Dcp1: decapping complex and the decapping components and Scd6. In this interaction network, both and Scd6 use an N-terminal LSm domain to interact with helical leucine-rich sequence motifs in (HLMs) (Gaudon et al, 1999) (Figures 2 4). Multiple HLM sequences are present in, close to the catalytic domain and in the disordered -terminal extension (Figures 1 and 3). The presence of multiple HLMs suggests that multiple activators can bind at various positions in the decapping complex, resulting in the assembly of decapping complexes that differ in /Scd6 content and that potentially differ in substrate specificity. In addition, we show that multiple decapping complexes can be clustered through. Our results thus reveal a plastic network of intermolecular interactions that can result in mrn decapping clusters of varying composition. We unravelled the structural basis of the interactions between the decapping complex and the activator proteins by solving the structure of the LSm domain in complex 15 N (p.p.m.) D.m. LSm domain D.m. LSm domain + Dcp1 Motif I W8 10 I15 R63 G10 V13 L27 D7 S32 I31 59 with HLM-1 from (Figure 2). The structure of the complex displays a novel mode of LSm domain interaction with ligands. Upon complex formation, the disordered HLM sequence adopts a helical conformation that binds close to the N-terminal ends of b-strands 2 and 5. This novel mode of interaction is possible only as a result of two specific features of the and Scd6 LSm domains: (1) the monomeric nature of the LSm domain that makes b-strand 5, that is normally used for intermolecular contacts, available for the interaction with the N-terminal end of the HLM helix and (2) the very short N-terminal helix in the and Scd6 LSm domain compared with canonical (L)Sm folds, which makes space available for the interaction with the -terminal end of the HLM helix. ased on the finding that two different LSm domains are able to interact with four different HLM sequences in yeast, it is clear that a certain degree of sequence variation is tolerated I68 D66 12 L57 D.m. LSm T4 E6 W8 E H (p.p.m.) N 11 D.m. Dcp1 N E35 Figure 7 D. melanogaster motif-1 in Dcp1 resembles a metazoan HLM. () NMR spectra of the LSm domain from D. melanogaster in the absence (brown) and presence (red) of the D. melanogaster Dcp1 motif-1. Residues that experience significant chemical shift changes are indicated. () Model of the D. melanogaster LSm domain in complex with Dcp1 motif-1. The model was constructed by superimposing the D. melanogaster LSm domain (brown, PD:2RM4; see Figure 1) onto the yeast LSm domain complex structure. Residues of the D. melanogaster LSm domain that are affected by the interaction with Dcp1 motif-1 (Figure 7) are coloured blue. The location of the helix in the S. pombe : structure is indicated in red (Figure 2). The chemical shift changes observed for the D. melanogaster Dcp1 Motif-1: complex agree well with the S. pombe : structure and suggest a similar mode of binding. I62 7 & 2012 European Molecular iology Organization The EMO Journal VOL 31 NO
10 in both the interaction domains and the recognition sequences (Figure 3). This plasticity is especially true for the HLM sequence that, according to the structure of the complex, must possess a helical propensity. ased on the interacting motifs, the HLMs preferably contain an SxxLLxLL motif; however, more than a single leucine-toalanine mutation is required to abolish binding, indicating the lack of a very stringent requirement for these amino acids (Figure 3; Supplementary Figure S3). s such, it is tempting to speculate that additional LSm-interacting HLMs are present in other components of the mrn degradation machinery. One such protein is Rps28 that interacts with (Ito et al, 2001; Decourty et al, 2008) and contains a sequence that is reminiscent of an HLM (EDDILVLM between residues 51 and 58 in S. pombe). Interestingly, in S. cerevisiae, the deadenylation-independent degradation of the Rps28 mrn depends on one hand on the protein and on the other hand on an interaction between the Rps28 mrn 3 0 UTR and the Rps28 protein (adis et al, 2004). It is thus tempting to speculate that the Dcp1: decapping complex can recruit the Rps28 mrn through, where the dimeric protein uses one LSm domain to interact with and uses the other LSm domain to interact with the HLM in the Rps28 protein mrn complex. We show that there are two main biological consequences of the interaction of /Scd6 with the decapping complex. First, the interaction between the or Scd6 LSm domains and the first HLM sequence in (HLM-1, residues ) results in an increase in the catalytic activity of the decapping complex (Figure 5). We argue that the molecular basis of this modulation of decapping activity results from the destabilization of the inactive closed conformation of the enzyme. Indeed, the interaction of the decapping complex with the LSm domain is incompatible with the closed conformation of the decapping complex that has been suggested to be catalytically inactive (Floor et al, 2010). The interaction of or Scd6 with HLM-1 in thus induces a shift of the structural equilibrium away from catalytically inactive conformations (Figure 5). In general, the level of -induced activation of the Dcp1: decapping complex is less than twofold (Figure 5) (Harigaya et al, 2010; Nissan et al, 2010). From a structural point of view, this is not surprising, as the population of the closed inactive state that is prevented by the interaction of the decapping complex with the /Scd6 LSm domain is only one of the possible conformations that the decapping complex can adopt. Interestingly, removal of all HLMs from fission yeast (Figure 6) results in a growth defect at higher temperatures (Supplementary Figure S8). s the truncated decapping complex still possesses catalytic activity (Floor et al, 2010) (Figure 5), this suggests that cellular function is impaired due to a loss of the regulation of the decapping complex activity. To what extend individual mrn degradation rates are influenced remains to be determined. It is, however, interesting to note that removal of all -terminal residues from S. cerevisiae results in increased Rps28 and MF2pG mrn levels (Harigaya et al, 2010), suggesting that HLM-1 not only has an important role in modulating the activity of the decapping complex in vitro, but also in vivo. The second biological function of the HLMs results from the LSm interaction sites located in the -terminal region of. Using fluorescence microscopy of fission yeast strains that carry different versions of the protein, we show that HLM-1 to HLM-5 are required for the localization of the decapping complex to processing bodies (Figure 6). Interestingly, a single HLM (HLM-1) is not sufficient to target the decapping complex to P-bodies (Figure 6). Upon deletion of all HLMs from,, and Lsm7 still localized to cytoplasmic foci. This shows that is not essential for P- body formation and is in agreement with the fact that no single protein has been identified that is absolutely essential for the formation of these cytoplasmic foci in yeast (Decker et al, 2007; Teixeira and Parker, 2007). Nevertheless, it has been shown that contributes to the assembly or maintenance of P-bodies (Teixeira and Parker, 2007). This correlates well with our in vitro findings that show that is able to promote a clustering of decapping complexes through HLMs in (Figure 3D and E). The basic set of proteins that regulate mrn degradation is conserved from yeast to humans. However, in higher eukaryotes, additional proteins (e.g. Edc4/Ge-1) are required for Dcp1: catalytic activity (Fenger-Gron et al, 2005), indicating that differences in the regulation of mrn decapping may have evolved. The HLMs in yeast are not present in metazoan sequences. However, we showed that the conserved motif-1 in the metazoan Dcp1 resembles an HLM-like motif that directly interacts with metazoan LSm domain. The observation that both yeast and metazoan are able to interact with Dcp1: indicates that the basic interactions between decapping complex components are conserved but that the details have changed during evolution. These changes can be explained by the low evolutionary pressure on the location of small linear motifs in the protein complexes, as only a small number of point mutations in a disordered region of a protein are required to relocate these motifs (Neduva and Russell, 2005). urrently, there is no high-resolution structure of the Dcp1:Ge-1: decapping complex available. Thus, it is not possible to speculate whether the interaction of the /Scd6 Lsm domain with Dcp1 will be able to enhance the catalytic activity by inducing conformational changes in, similar to the observation presented here for the yeast decapping machinery. It is worth noting, however, that it was previously shown that metazoan Dcp1 trimerizes through a unique domain at the -terminus (Tritschler et al, 2009b). s a result, the metazoan Dcp1: complex contains multiple binding motifs for, similar to the multiple binding motifs in the monomeric yeast protein. Interestingly, the removal of the trimerization domain from metazoan Dcp1 results in the failure of this decapping complex to localize to P-bodies (Tritschler et al, 2007, 2009b), indicating that a single :Dcp1 binding event is not sufficient for the P- body localization. This correlates with our findings here that show that a single HLM is not sufficient for the P-body localization of the fission yeast decapping complex (Figure 6). In summary, we provide the first structural insights into the interactions that regulate the activity and cellular localization of the mrn decapping complex. ased on these and previous results, it appears that proteins involved in mrn degradation have multiple functions. not only enhances the catalytic activity of the Dcp1: mrn decapping complex but also is responsible for the localization of the complex to P-bodies. Scd6 was previously reported to repress 288 The EMO Journal VOL 31 NO & 2012 European Molecular iology Organization
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Structure and RNA-binding properties. of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex
 Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Chapter 6. The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR
 The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Transmembrane Domains (TMDs) of ABC transporters
 Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Gene regulation I Biochemistry 302. Bob Kelm February 25, 2005
 Gene regulation I Biochemistry 302 Bob Kelm February 25, 2005 Principles of gene regulation (cellular versus molecular level) Extracellular signals Chemical (e.g. hormones, growth factors) Environmental
Gene regulation I Biochemistry 302 Bob Kelm February 25, 2005 Principles of gene regulation (cellular versus molecular level) Extracellular signals Chemical (e.g. hormones, growth factors) Environmental
Three types of RNA polymerase in eukaryotic nuclei
 Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Gene Control Mechanisms at Transcription and Translation Levels
 Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Table 1. Crystallographic data collection, phasing and refinement statistics. Native Hg soaked Mn soaked 1 Mn soaked 2
 Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
RNA Polymerase I Contains a TFIIF-Related DNA-Binding Subcomplex
 Molecular Cell, Volume 39 Supplemental Information RNA Polymerase I Contains a TFIIFRelated DNABinding Subcomplex Sebastian R. Geiger, Kristina Lorenzen, Amelie Schreieck, Patrizia Hanecker, Dirk Kostrewa,
Molecular Cell, Volume 39 Supplemental Information RNA Polymerase I Contains a TFIIFRelated DNABinding Subcomplex Sebastian R. Geiger, Kristina Lorenzen, Amelie Schreieck, Patrizia Hanecker, Dirk Kostrewa,
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/4/e1600663/dc1 Supplementary Materials for A dynamic hydrophobic core orchestrates allostery in protein kinases Jonggul Kim, Lalima G. Ahuja, Fa-An Chao, Youlin
advances.sciencemag.org/cgi/content/full/3/4/e1600663/dc1 Supplementary Materials for A dynamic hydrophobic core orchestrates allostery in protein kinases Jonggul Kim, Lalima G. Ahuja, Fa-An Chao, Youlin
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
Gene regulation II Biochemistry 302. February 27, 2006
 Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Supplemental Materials Molecular Biology of the Cell
 Supplemental Materials Molecular iology of the Cell Figure S1 Krüger et al. Arabidopsis Plasmodium H. sapiens* 1) Xenopus* 1) Drosophila C.elegans S.cerevisae S.pombe L.major T.cruzi T.brucei DCP5 CITH
Supplemental Materials Molecular iology of the Cell Figure S1 Krüger et al. Arabidopsis Plasmodium H. sapiens* 1) Xenopus* 1) Drosophila C.elegans S.cerevisae S.pombe L.major T.cruzi T.brucei DCP5 CITH
Initiation of translation in eukaryotic cells:connecting the head and tail
 Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Structural characterization of NiV N 0 P in solution and in crystal.
 Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
SUPPLEMENTARY INFORMATION
 Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
DNA Structure. Voet & Voet: Chapter 29 Pages Slide 1
 DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10955 Supplementary Figures Supplementary Figure 1. Electron-density maps and crystallographic dimer structures of the motor domain. (a f) Stereo views of the final electron-density maps
doi:10.1038/nature10955 Supplementary Figures Supplementary Figure 1. Electron-density maps and crystallographic dimer structures of the motor domain. (a f) Stereo views of the final electron-density maps
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
CH 3 CH 2 OH +H 2 O CHO. 2e + 2H + + O 2 H 2 O +HCOOH
 2 4 H CH 3 2e + 2H + + 2 H 2 2 H CH 2 H 2e + 2H + + 2 H 2 2 H +H 2 CH 2e + 2H + + 2 H 2 2 H +HCH Supplemental Figure S. The three-step 4DM reaction, each step requires two reducing equivalents from ADPH
2 4 H CH 3 2e + 2H + + 2 H 2 2 H CH 2 H 2e + 2H + + 2 H 2 2 H +H 2 CH 2e + 2H + + 2 H 2 2 H +HCH Supplemental Figure S. The three-step 4DM reaction, each step requires two reducing equivalents from ADPH
Protein Structure. W. M. Grogan, Ph.D. OBJECTIVES
 Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
SUPPLEMENTARY INFORMATION
 Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
Nature Structural and Molecular Biology: doi: /nsmb.2938
 Supplementary Figure 1 Characterization of designed leucine-rich-repeat proteins. (a) Water-mediate hydrogen-bond network is frequently visible in the convex region of LRR crystal structures. Examples
Supplementary Figure 1 Characterization of designed leucine-rich-repeat proteins. (a) Water-mediate hydrogen-bond network is frequently visible in the convex region of LRR crystal structures. Examples
Reading Assignments. A. Genes and the Synthesis of Polypeptides. Lecture Series 7 From DNA to Protein: Genotype to Phenotype
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Introduction to" Protein Structure
 Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN
 THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1737 Supplementary Table 1 variant Description FSEC - 2B12 a FSEC - 6A1 a K d (leucine) c Leucine uptake e K (wild-type like) K (Y18F) K (TS) K (TSY) K288A mutant, lipid facing side chain
doi:1.138/nature1737 Supplementary Table 1 variant Description FSEC - 2B12 a FSEC - 6A1 a K d (leucine) c Leucine uptake e K (wild-type like) K (Y18F) K (TS) K (TSY) K288A mutant, lipid facing side chain
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
Review Article Insights from a Paradigm Shift: How the Poly(A)-Binding Protein Brings Translating mrnas Full Circle
 New Journal of Science, Article ID 873084, 16 pages http://dx.doi.org/10.1155/2014/873084 Review Article Insights from a Paradigm Shift: How the Poly(A)-Binding Protein Brings Translating mrnas Full Circle
New Journal of Science, Article ID 873084, 16 pages http://dx.doi.org/10.1155/2014/873084 Review Article Insights from a Paradigm Shift: How the Poly(A)-Binding Protein Brings Translating mrnas Full Circle
Cks1 CDK1 CDK1 CDK1 CKS1. are ice- lobe. conserved. conserved
 Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Introduction to Comparative Protein Modeling. Chapter 4 Part I
 Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
eif1a/eif5b interaction network and its functions in translation initiation complex assembly and remodeling
 Published online 20 June 2016 Nucleic Acids Research, 2016, Vol. 44, No. 15 7441 7456 doi: 10.1093/nar/gkw552 eif1a/eif5b interaction network and its functions in translation initiation complex assembly
Published online 20 June 2016 Nucleic Acids Research, 2016, Vol. 44, No. 15 7441 7456 doi: 10.1093/nar/gkw552 eif1a/eif5b interaction network and its functions in translation initiation complex assembly
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
SUPPLEMENTARY INFORMATION
 Dph2 SeMet (iron-free) # Dph2 (iron-free) Dph2-[4Fe-4S] Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 58.26, 82.08, 160.42 58.74, 81.87, 160.01 55.70, 80.53,
Dph2 SeMet (iron-free) # Dph2 (iron-free) Dph2-[4Fe-4S] Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 58.26, 82.08, 160.42 58.74, 81.87, 160.01 55.70, 80.53,
Bi 8 Lecture 11. Quantitative aspects of transcription factor binding and gene regulatory circuit design. Ellen Rothenberg 9 February 2016
 Bi 8 Lecture 11 Quantitative aspects of transcription factor binding and gene regulatory circuit design Ellen Rothenberg 9 February 2016 Major take-home messages from λ phage system that apply to many
Bi 8 Lecture 11 Quantitative aspects of transcription factor binding and gene regulatory circuit design Ellen Rothenberg 9 February 2016 Major take-home messages from λ phage system that apply to many
Translation. A ribosome, mrna, and trna.
 Translation The basic processes of translation are conserved among prokaryotes and eukaryotes. Prokaryotic Translation A ribosome, mrna, and trna. In the initiation of translation in prokaryotes, the Shine-Dalgarno
Translation The basic processes of translation are conserved among prokaryotes and eukaryotes. Prokaryotic Translation A ribosome, mrna, and trna. In the initiation of translation in prokaryotes, the Shine-Dalgarno
Welcome to Class 21!
 Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
eif1a/eif5b Interaction Network and its Functions in Translation Initiation Complex Assembly and Remodeling
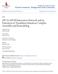 Bridgewater State University Virtual Commons - Bridgewater State University Biological Sciences Faculty Publications Biological Sciences Department 2016 eif1a/eif5b Interaction Network and its Functions
Bridgewater State University Virtual Commons - Bridgewater State University Biological Sciences Faculty Publications Biological Sciences Department 2016 eif1a/eif5b Interaction Network and its Functions
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R)
 Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Supplemental Data SUPPLEMENTAL FIGURES
 Supplemental Data CRYSTAL STRUCTURE OF THE MG.ADP-INHIBITED STATE OF THE YEAST F 1 C 10 ATP SYNTHASE Alain Dautant*, Jean Velours and Marie-France Giraud* From Université Bordeaux 2, CNRS; Institut de
Supplemental Data CRYSTAL STRUCTURE OF THE MG.ADP-INHIBITED STATE OF THE YEAST F 1 C 10 ATP SYNTHASE Alain Dautant*, Jean Velours and Marie-France Giraud* From Université Bordeaux 2, CNRS; Institut de
Bioinformatics Practical for Biochemists
 Bioinformatics Practical for Biochemists Andrei Lupas, Birte Höcker, Steffen Schmidt WS 2013/14 03. Sequence Features Targeting proteins signal peptide targets proteins to the secretory pathway N-terminal
Bioinformatics Practical for Biochemists Andrei Lupas, Birte Höcker, Steffen Schmidt WS 2013/14 03. Sequence Features Targeting proteins signal peptide targets proteins to the secretory pathway N-terminal
Transcription Regulation And Gene Expression in Eukaryotes UPSTREAM TRANSCRIPTION FACTORS
 Transcription Regulation And Gene Expression in Eukaryotes UPSTREAM TRANSCRIPTION FACTORS RG. Clerc March 26. 2008 UPSTREAM TRANSCRIPTION FACTORS Experimental approaches DNA binding domains (DBD) Transcription
Transcription Regulation And Gene Expression in Eukaryotes UPSTREAM TRANSCRIPTION FACTORS RG. Clerc March 26. 2008 UPSTREAM TRANSCRIPTION FACTORS Experimental approaches DNA binding domains (DBD) Transcription
BA, BSc, and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
Lecture 7: Simple genetic circuits I
 Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
BSc and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BSc and MSc Degree Examinations 2018-9 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes Marking
Examination Candidate Number: Desk Number: BSc and MSc Degree Examinations 2018-9 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes Marking
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
ASSESSING TRANSLATIONAL EFFICIACY THROUGH POLY(A)- TAIL PROFILING AND IN VIVO RNA SECONDARY STRUCTURE DETERMINATION
 ASSESSING TRANSLATIONAL EFFICIACY THROUGH POLY(A)- TAIL PROFILING AND IN VIVO RNA SECONDARY STRUCTURE DETERMINATION Journal Club, April 15th 2014 Karl Frontzek, Institute of Neuropathology POLY(A)-TAIL
ASSESSING TRANSLATIONAL EFFICIACY THROUGH POLY(A)- TAIL PROFILING AND IN VIVO RNA SECONDARY STRUCTURE DETERMINATION Journal Club, April 15th 2014 Karl Frontzek, Institute of Neuropathology POLY(A)-TAIL
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Definition and assessment of ciap1 constructs.
 Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2720 1 2 3 Tuning underwater adhesion with cation-π interactions Matthew A. Gebbie, Wei Wei, Alex M. Schrader,
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2720 1 2 3 Tuning underwater adhesion with cation-π interactions Matthew A. Gebbie, Wei Wei, Alex M. Schrader,
From gene to protein. Premedical biology
 From gene to protein Premedical biology Central dogma of Biology, Molecular Biology, Genetics transcription replication reverse transcription translation DNA RNA Protein RNA chemically similar to DNA,
From gene to protein Premedical biology Central dogma of Biology, Molecular Biology, Genetics transcription replication reverse transcription translation DNA RNA Protein RNA chemically similar to DNA,
Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6
 Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Acta Crystallographica Section D
 Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: Structural basis of the heterodimerization of the MST and RASSF SARAH domains in the Hippo signalling
Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: Structural basis of the heterodimerization of the MST and RASSF SARAH domains in the Hippo signalling
Diphthamide biosynthesis requires a radical iron-sulfur enzyme. Pennsylvania State University, University Park, Pennsylvania 16802, USA
 Diphthamide biosynthesis requires a radical iron-sulfur enzyme Yang Zhang, 1,4 Xuling Zhu, 1,4 Andrew T. Torelli, 1 Michael Lee, 2 Boris Dzikovski, 1 Rachel Koralewski, 1 Eileen Wang, 1 Jack Freed, 1 Carsten
Diphthamide biosynthesis requires a radical iron-sulfur enzyme Yang Zhang, 1,4 Xuling Zhu, 1,4 Andrew T. Torelli, 1 Michael Lee, 2 Boris Dzikovski, 1 Rachel Koralewski, 1 Eileen Wang, 1 Jack Freed, 1 Carsten
Supplementary Information. Overlap between folding and functional energy landscapes for. adenylate kinase conformational change
 Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Full-length GlpG sequence was generated by PCR from E. coli genomic DNA. (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
 Supplementary Methods Protein expression and purification Full-length GlpG sequence was generated by PCR from E. coli genomic DNA (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
Supplementary Methods Protein expression and purification Full-length GlpG sequence was generated by PCR from E. coli genomic DNA (with two sequence variations, D51E/L52V, from the gene bank entry aac28166),
Activation of a receptor. Assembly of the complex
 Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Copyright Mark Brandt, Ph.D A third method, cryogenic electron microscopy has seen increasing use over the past few years.
 Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Cryo-EM data collection, refinement and validation statistics
 1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
1 Table S1 Cryo-EM data collection, refinement and validation statistics Data collection and processing CPSF-160 WDR33 (EMDB-7114) (PDB 6BM0) CPSF-160 WDR33 (EMDB-7113) (PDB 6BLY) CPSF-160 WDR33 CPSF-30
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12242 C. thermophilum 666 RPAVLDNVYIRPALE-GKRVPGKVEIHQNGIRYQSPLSTTQRVDVLFSNIRHLFFQPCQN S. pombe 659 RPAHINDVYVRPAID-GKRLPGFIEIHQNGIRYQSPLRSDSHIDLLFSNMKHLFFQPCEG
SUPPLEMENTARY INFORMATION doi:10.1038/nature12242 C. thermophilum 666 RPAVLDNVYIRPALE-GKRVPGKVEIHQNGIRYQSPLSTTQRVDVLFSNIRHLFFQPCQN S. pombe 659 RPAHINDVYVRPAID-GKRLPGFIEIHQNGIRYQSPLRSDSHIDLLFSNMKHLFFQPCEG
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
ml. ph 7.5 ph 6.5 ph 5.5 ph 4.5. β 2 AR-Gs complex + GDP β 2 AR-Gs complex + GTPγS
 a UV28 absorption (mau) 9 8 7 5 3 β 2 AR-Gs complex β 2 AR-Gs complex + GDP β 2 AR-Gs complex + GTPγS β 2 AR-Gs complex dissociated complex excess nucleotides b 9 8 7 5 3 β 2 AR-Gs complex β 2 AR-Gs complex
a UV28 absorption (mau) 9 8 7 5 3 β 2 AR-Gs complex β 2 AR-Gs complex + GDP β 2 AR-Gs complex + GTPγS β 2 AR-Gs complex dissociated complex excess nucleotides b 9 8 7 5 3 β 2 AR-Gs complex β 2 AR-Gs complex
Nature Structural & Molecular Biology doi: /nsmb Supplementary Figure 1. CRBN binding assay with thalidomide enantiomers.
 Supplementary Figure 1 CRBN binding assay with thalidomide enantiomers. (a) Competitive elution assay using thalidomide-immobilized beads coupled with racemic thalidomide. Beads were washed three times
Supplementary Figure 1 CRBN binding assay with thalidomide enantiomers. (a) Competitive elution assay using thalidomide-immobilized beads coupled with racemic thalidomide. Beads were washed three times
Genetics 304 Lecture 6
 Genetics 304 Lecture 6 00/01/27 Assigned Readings Busby, S. and R.H. Ebright (1994). Promoter structure, promoter recognition, and transcription activation in prokaryotes. Cell 79:743-746. Reed, W.L. and
Genetics 304 Lecture 6 00/01/27 Assigned Readings Busby, S. and R.H. Ebright (1994). Promoter structure, promoter recognition, and transcription activation in prokaryotes. Cell 79:743-746. Reed, W.L. and
CHAPTER 29 HW: AMINO ACIDS + PROTEINS
 CAPTER 29 W: AMI ACIDS + PRTEIS For all problems, consult the table of 20 Amino Acids provided in lecture if an amino acid structure is needed; these will be given on exams. Use natural amino acids (L)
CAPTER 29 W: AMI ACIDS + PRTEIS For all problems, consult the table of 20 Amino Acids provided in lecture if an amino acid structure is needed; these will be given on exams. Use natural amino acids (L)
Chapter 9 DNA recognition by eukaryotic transcription factors
 Chapter 9 DNA recognition by eukaryotic transcription factors TRANSCRIPTION 101 Eukaryotic RNA polymerases RNA polymerase RNA polymerase I RNA polymerase II RNA polymerase III RNA polymerase IV Function
Chapter 9 DNA recognition by eukaryotic transcription factors TRANSCRIPTION 101 Eukaryotic RNA polymerases RNA polymerase RNA polymerase I RNA polymerase II RNA polymerase III RNA polymerase IV Function
Regulation of Transcription in Eukaryotes
 Regulation of Transcription in Eukaryotes Leucine zipper and helix-loop-helix proteins contain DNA-binding domains formed by dimerization of two polypeptide chains. Different members of each family can
Regulation of Transcription in Eukaryotes Leucine zipper and helix-loop-helix proteins contain DNA-binding domains formed by dimerization of two polypeptide chains. Different members of each family can
Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises
 Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises six Orai subunits, each with identical amino acid sequences
Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises six Orai subunits, each with identical amino acid sequences
Identification of Functionally Important Residues of Arp2/3 Complex by Analysis of Homology Models from Diverse Species
 doi:10.1016/j.jmb.2003.12.017 J. Mol. Biol. (2004) 336, 551 565 Identification of Functionally Important Residues of Arp2/3 Complex by Analysis of Homology Models from Diverse Species Christopher C. Beltzner
doi:10.1016/j.jmb.2003.12.017 J. Mol. Biol. (2004) 336, 551 565 Identification of Functionally Important Residues of Arp2/3 Complex by Analysis of Homology Models from Diverse Species Christopher C. Beltzner
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Basics of protein structure
 Today: 1. Projects a. Requirements: i. Critical review of one paper ii. At least one computational result b. Noon, Dec. 3 rd written report and oral presentation are due; submit via email to bphys101@fas.harvard.edu
Today: 1. Projects a. Requirements: i. Critical review of one paper ii. At least one computational result b. Noon, Dec. 3 rd written report and oral presentation are due; submit via email to bphys101@fas.harvard.edu
NMR study of complexes between low molecular mass inhibitors and the West Nile virus NS2B-NS3 protease
 University of Wollongong Research Online Faculty of Science - Papers (Archive) Faculty of Science, Medicine and Health 2009 NMR study of complexes between low molecular mass inhibitors and the West Nile
University of Wollongong Research Online Faculty of Science - Papers (Archive) Faculty of Science, Medicine and Health 2009 NMR study of complexes between low molecular mass inhibitors and the West Nile
Detailed description of overall and active site architecture of PPDC- 3dThDP, PPDC-2HE3dThDP, PPDC-3dThDP-PPA and PPDC- 3dThDP-POVA
 Online Supplemental Results Detailed description of overall and active site architecture of PPDC- 3dThDP, PPDC-2HE3dThDP, PPDC-3dThDP-PPA and PPDC- 3dThDP-POVA Structure solution and overall architecture
Online Supplemental Results Detailed description of overall and active site architecture of PPDC- 3dThDP, PPDC-2HE3dThDP, PPDC-3dThDP-PPA and PPDC- 3dThDP-POVA Structure solution and overall architecture
Understanding Science Through the Lens of Computation. Richard M. Karp Nov. 3, 2007
 Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
Supplemental Information. Expanded Coverage of the 26S Proteasome. Conformational Landscape Reveals. Mechanisms of Peptidase Gating
 Cell Reports, Volume 24 Supplemental Information Expanded Coverage of the 26S Proteasome Conformational Landscape Reveals Mechanisms of Peptidase Gating Markus R. Eisele, Randi G. Reed, Till Rudack, Andreas
Cell Reports, Volume 24 Supplemental Information Expanded Coverage of the 26S Proteasome Conformational Landscape Reveals Mechanisms of Peptidase Gating Markus R. Eisele, Randi G. Reed, Till Rudack, Andreas
National de la Recherche Scientifique and Université Paris Descartes, Paris, France.
 FAST-RESPONSE CALMODULIN-BASED FLUORESCENT INDICATORS REVEAL RAPID INTRACELLULAR CALCIUM DYNAMICS Nordine Helassa a, Xiao-hua Zhang b, Ianina Conte a,c, John Scaringi b, Elric Esposito d, Jonathan Bradley
FAST-RESPONSE CALMODULIN-BASED FLUORESCENT INDICATORS REVEAL RAPID INTRACELLULAR CALCIUM DYNAMICS Nordine Helassa a, Xiao-hua Zhang b, Ianina Conte a,c, John Scaringi b, Elric Esposito d, Jonathan Bradley
Biomolecules: lecture 10
 Biomolecules: lecture 10 - understanding in detail how protein 3D structures form - realize that protein molecules are not static wire models but instead dynamic, where in principle every atom moves (yet
Biomolecules: lecture 10 - understanding in detail how protein 3D structures form - realize that protein molecules are not static wire models but instead dynamic, where in principle every atom moves (yet
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
The Fic protein Doc uses an inverted substrate to phosphorylate and. inactivate EF-Tu
 The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
SUPPLEMENTARY INFORMATION
 Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
Host-Pathogen interaction-ii. Pl Path 604 PN Sharma Department of Plant Pathology CSK HPKV, Palampur
 Host-Pathogen interaction-ii Pl Path 604 PN Sharma Department of Plant Pathology CSK HPKV, Palampur-176062 It was originally believed that gene-for-gene resistance was conferred by a direct interaction
Host-Pathogen interaction-ii Pl Path 604 PN Sharma Department of Plant Pathology CSK HPKV, Palampur-176062 It was originally believed that gene-for-gene resistance was conferred by a direct interaction
Structural basis for docking of peroxisomal membrane protein carrier Pex19p onto its receptor Pex3p
 Manuscript EMBO-2010-75031 Structural basis for docking of peroxisomal membrane protein carrier Pex19p onto its receptor Pex3p Yasuhiko Sato, Hiroyuki Shibata, Toru Nakatsu, Hiroaki Nakano, Yoshinori Kashiwayama,
Manuscript EMBO-2010-75031 Structural basis for docking of peroxisomal membrane protein carrier Pex19p onto its receptor Pex3p Yasuhiko Sato, Hiroyuki Shibata, Toru Nakatsu, Hiroaki Nakano, Yoshinori Kashiwayama,
MBLG lecture 5. The EGG! Visualising Molecules. Dr. Dale Hancock Lab 715
 MBLG lecture 5 Dr. Dale Hancock D.Hancock@mmb.usyd.edu.au Lab 715 The EGG! Visualising Molecules In molecular biology and biochemistry it is better to view molecules as killer pythons rather than smarties.
MBLG lecture 5 Dr. Dale Hancock D.Hancock@mmb.usyd.edu.au Lab 715 The EGG! Visualising Molecules In molecular biology and biochemistry it is better to view molecules as killer pythons rather than smarties.
A) at equilibrium B) endergonic C) endothermic D) exergonic E) exothermic.
 CHEM 2770: Elements of Biochemistry Mid Term EXAMINATION VERSION A Date: October 29, 2014 Instructor: H. Perreault Location: 172 Schultz Time: 4 or 6 pm. Duration: 1 hour Instructions Please mark the Answer
CHEM 2770: Elements of Biochemistry Mid Term EXAMINATION VERSION A Date: October 29, 2014 Instructor: H. Perreault Location: 172 Schultz Time: 4 or 6 pm. Duration: 1 hour Instructions Please mark the Answer
Chapter 16 Lecture. Concepts Of Genetics. Tenth Edition. Regulation of Gene Expression in Prokaryotes
 Chapter 16 Lecture Concepts Of Genetics Tenth Edition Regulation of Gene Expression in Prokaryotes Chapter Contents 16.1 Prokaryotes Regulate Gene Expression in Response to Environmental Conditions 16.2
Chapter 16 Lecture Concepts Of Genetics Tenth Edition Regulation of Gene Expression in Prokaryotes Chapter Contents 16.1 Prokaryotes Regulate Gene Expression in Response to Environmental Conditions 16.2
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
