Computational engineering of cellulase Cel9A-68 functional motions through mutations in its linker region. WT 1TF4 (crystal) -90 ERRAT PROVE VERIFY3D
|
|
|
- Laurel Smith
- 5 years ago
- Views:
Transcription
1 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 218 Supplementary Material: Computational engineering of cellulase Cel9-68 functional motions through mutations in its linker region M. G. S. Costa, a Y. F. Silva a and P. R. atista* a a. Programa de Computação Científica (PROCC), Fundação Oswaldo Cruz, Rio de Janeiro, razil. pbatista@fiocruz.br Fig. S1. 1TF4 (crystal) G46P General Glycine General Glycine Proline -9 9 Pre-Proline -9 9 Proline -9 9 Pre-Proline Overall quality factor: Overall quality factor: % of buried protein atoms are outliers 3.9 % of buried protein atoms are outliers 97.19% of the residues had an averaged 3D-1D score >= % of the residues had an averaged 3D-1D score >=.2 G46+ Overall quality factor: % of buried protein atoms are outliers 97.64% of the residues had an averaged 3D-1D score >=.2 Overall quality factor: % of buried protein atoms are outliers 97.98% of the residues had an averaged 3D-1D score >=.2 Fig.S1 Validation of the models. The Ramachandran plots and the results returned by, and VERIFY 3D are given for each system considered in this work, including the crystallographic Cel9-68 structure (PD ID: 1TF4).
2 Fig. S2. cum. contribu,on (%) collec,vity G46P G mode # C D RMSF (Å) Δ RMSF (Å) CD residue # linker CM G46P - G Fig.S2 Normal mode analysis of Cel9-68 variants. In, cumulative contribution of the first ten low-frequency normal modes to the overall atomic fluctuations. In, Collectivity of motions described by low frequency modes. In C, absolute fluctuations per residue. In D, relative fluctuations (taking as reference) per mode coloured as indicated in the legend. The Cel9-68 domain definitions are indicated above the plot. Fig. S3. G46+ G46P run 1 run 2 run 3 Fig.S3 Time evolution of backbone RMSD per replica. The simulated systems are identified above each plot and the results obtained for each independent run are coloured as indicated in the legend.
3 Fig. S4. G46P G46+ run 1 run 2 run 3 radius of gyra7on (Å) distance between domains (Å) Time (ns) Fig.S4 Time evolution of the interdomain distances (in ) and the radius of gyration of Cel9-68 (in ), per replica, along the MD simulations. The simulated systems are identified above each plot and the results obtained for each independent run are coloured as indicated in the legend. Fig. S5. Fig.S5 Distribution of RMSD distances for all pairs of conformations obtained in the concatenated trajectories for each simulated system. Coloured as indicated in the legend.
4 Fig. S6. mode 7 mode 8 mode 9 Fig.S6 Results of population analysis based on projections of the conformations sampled during explicit solvent MD onto the three low-frequency normal modes calculated for the system (top) or (bottom). Coloured as indicated in the legend.
5 Captions of supplementary movies Supplementary movie 1: Displacements along normal mode 7 calculated for the Cel9-68 wild type system. Each structural domain was differentially coloured as in Figure 1: catalytic domain (red), linker (cyan) and carbohydrate binding module (green). Supplementary movie 2: Displacements along normal mode 8 calculated for the Cel9-68 wild type system. The protein is coloured as in supplementary movie 1. Supplementary movie 3: Displacements along normal mode 9 calculated for the Cel9-68 wild type system. The protein is coloured as in supplementary movie 1. Supplementary movie 4: Differences between the lowest frequency normal mode calculated for the and systems (related to Figure 3). The Cel9-68 structural domains are indicated in the left part of the video. Different colouring of the CM was applied to highlight the conformational states reached after displacements along each sense of the lowest frequency modes.
Insights into the Biotransformation of 2,4,6- Trinitrotoluene by the Old Yellow Enzyme Family of Flavoproteins. A Computational Study
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2019 Supporting Information for Insights into the Biotransformation of 2,4,6- Trinitrotoluene
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2019 Supporting Information for Insights into the Biotransformation of 2,4,6- Trinitrotoluene
Supplementary Information
 Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
Supporting Information
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supporting Information Lipid molecules can induce an opening of membrane-facing
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supporting Information Lipid molecules can induce an opening of membrane-facing
Supporting Information How does Darunavir prevent HIV-1 protease dimerization?
 Supporting Information How does Darunavir prevent HIV- protease dimerization? Danzhi Huang and Amedeo Caflisch a Department of Biochemistry University of Zürich, Winterthurerstrasse 9 CH-7 Zürich, Switzerland
Supporting Information How does Darunavir prevent HIV- protease dimerization? Danzhi Huang and Amedeo Caflisch a Department of Biochemistry University of Zürich, Winterthurerstrasse 9 CH-7 Zürich, Switzerland
Structural and mechanistic insight into the substrate. binding from the conformational dynamics in apo. and substrate-bound DapE enzyme
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 215 Structural and mechanistic insight into the substrate binding from the conformational
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 215 Structural and mechanistic insight into the substrate binding from the conformational
Experimental and Computational Mutagenesis to Investigate the. Positioning of a General Base within an Enzyme Active Site
 Experimental and Computational Mutagenesis to Investigate the Positioning of a General Base within an Enzyme Active Site Jason P. Schwans, Philip Hanoian, Benjamin J. Lengerich, Fanny Sunden, Ana Gonzalez
Experimental and Computational Mutagenesis to Investigate the Positioning of a General Base within an Enzyme Active Site Jason P. Schwans, Philip Hanoian, Benjamin J. Lengerich, Fanny Sunden, Ana Gonzalez
T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y. jgp
 S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
Supporting information for: Mechanism of lignin inhibition of enzymatic. biomass deconstruction
 Supporting information for: Mechanism of lignin inhibition of enzymatic biomass deconstruction Josh V. Vermaas,, Loukas Petridis, Xianghong Qi,, Roland Schulz,, Benjamin Lindner, and Jeremy C. Smith,,
Supporting information for: Mechanism of lignin inhibition of enzymatic biomass deconstruction Josh V. Vermaas,, Loukas Petridis, Xianghong Qi,, Roland Schulz,, Benjamin Lindner, and Jeremy C. Smith,,
Mechanism of water/ion exchange at a protein surface: a weakly bound chloride in Helicobacter pylori apoflavodoxin.
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 25 ELECTRONIC SUPPLEMENTARY INFORMATION Mechanism of water/ion exchange at a protein
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 25 ELECTRONIC SUPPLEMENTARY INFORMATION Mechanism of water/ion exchange at a protein
IgE binds asymmetrically to its B cell receptor CD23
 Supplementary Information IgE binds asymmetrically to its B cell receptor CD23 Balvinder Dhaliwal 1*, Marie O. Y. Pang 2, Anthony H. Keeble 2,3, Louisa K. James 2,4, Hannah J. Gould 2, James M. McDonnell
Supplementary Information IgE binds asymmetrically to its B cell receptor CD23 Balvinder Dhaliwal 1*, Marie O. Y. Pang 2, Anthony H. Keeble 2,3, Louisa K. James 2,4, Hannah J. Gould 2, James M. McDonnell
Structural basis for catalytically restrictive dynamics of a high-energy enzyme state
 Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Assessing the potential of atomistic molecular dynamics simulations to probe reversible. protein-protein recognition and binding
 Supporting information for Assessing the potential of atomistic molecular dynamics simulations to probe reversible protein-protein recognition and binding Luciano A. Abriata and Matteo Dal Peraro Laboratory
Supporting information for Assessing the potential of atomistic molecular dynamics simulations to probe reversible protein-protein recognition and binding Luciano A. Abriata and Matteo Dal Peraro Laboratory
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Tu 1,*, , Sweden
 Supplementary Material Computational studiess of the binding profile of phosphoinositide PtdIns(,4,5)P with the pleckstrin homology domain d of an oomycetee cellulose synthase Guanglin Kuang 1, Vincent
Supplementary Material Computational studiess of the binding profile of phosphoinositide PtdIns(,4,5)P with the pleckstrin homology domain d of an oomycetee cellulose synthase Guanglin Kuang 1, Vincent
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
Supporting Information
 Supporting Information Critical role of inter-domain interactions on the conformational change and catalytic mechanism of Endoplasmic Reticulum Aminopeptidase 1 Athanasios Stamogiannos 1, Zachary Maben
Supporting Information Critical role of inter-domain interactions on the conformational change and catalytic mechanism of Endoplasmic Reticulum Aminopeptidase 1 Athanasios Stamogiannos 1, Zachary Maben
Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine. JAK2 Selective Mechanism Combined Molecular Dynamics Simulation
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Supporting Information Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Supporting Information Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine
NB-DNJ/GCase-pH 7.4 NB-DNJ+/GCase-pH 7.4 NB-DNJ+/GCase-pH 4.5
 SUPPLEMENTARY TABLES Suppl. Table 1. Protonation states at ph 7.4 and 4.5. Protonation states of titratable residues in GCase at ph 7.4 and 4.5. Histidine: HID, H at δ-nitrogen; HIE, H at ε-nitrogen; HIP,
SUPPLEMENTARY TABLES Suppl. Table 1. Protonation states at ph 7.4 and 4.5. Protonation states of titratable residues in GCase at ph 7.4 and 4.5. Histidine: HID, H at δ-nitrogen; HIE, H at ε-nitrogen; HIP,
Carbazole Derivatives Binding to c-kit G-quadruplex DNA
 Supplementary Materials Carbazole Derivatives Binding to c-kit G-quadruplex DNA Agata Głuszyńska 1, *, Bernard Juskowiak 1, Martyna Kuta-Siejkowska 2, Marcin Hoffmann 2 and Shozeb Haider 3 1 Laboratory
Supplementary Materials Carbazole Derivatives Binding to c-kit G-quadruplex DNA Agata Głuszyńska 1, *, Bernard Juskowiak 1, Martyna Kuta-Siejkowska 2, Marcin Hoffmann 2 and Shozeb Haider 3 1 Laboratory
Nature Structural and Molecular Biology: doi: /nsmb.2938
 Supplementary Figure 1 Characterization of designed leucine-rich-repeat proteins. (a) Water-mediate hydrogen-bond network is frequently visible in the convex region of LRR crystal structures. Examples
Supplementary Figure 1 Characterization of designed leucine-rich-repeat proteins. (a) Water-mediate hydrogen-bond network is frequently visible in the convex region of LRR crystal structures. Examples
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1737 Supplementary Table 1 variant Description FSEC - 2B12 a FSEC - 6A1 a K d (leucine) c Leucine uptake e K (wild-type like) K (Y18F) K (TS) K (TSY) K288A mutant, lipid facing side chain
doi:1.138/nature1737 Supplementary Table 1 variant Description FSEC - 2B12 a FSEC - 6A1 a K d (leucine) c Leucine uptake e K (wild-type like) K (Y18F) K (TS) K (TSY) K288A mutant, lipid facing side chain
Supplementary information for cloud computing approaches for prediction of ligand binding poses and pathways
 Supplementary information for cloud computing approaches for prediction of ligand binding poses and pathways Morgan Lawrenz 1, Diwakar Shukla 1,2 & Vijay S. Pande 1,2 1 Department of Chemistry, Stanford
Supplementary information for cloud computing approaches for prediction of ligand binding poses and pathways Morgan Lawrenz 1, Diwakar Shukla 1,2 & Vijay S. Pande 1,2 1 Department of Chemistry, Stanford
Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate
 Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate Shidi Zhao a, Laura Restrepo-Pérez b, Misha Soskine c, Giovanni Maglia c, Chirlmin Joo b, Cees Dekker b and Aleksei Aksimentiev
Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate Shidi Zhao a, Laura Restrepo-Pérez b, Misha Soskine c, Giovanni Maglia c, Chirlmin Joo b, Cees Dekker b and Aleksei Aksimentiev
A dynamic interaction process between KaiA and KaiC is critical to
 A dynamic interaction process between KaiA and KaiC is critical to the cyanobacterial circadian oscillator Pei Dong 1,2, Ying Fan 2, Jianqiang Sun 3, Mengting Lv 2, Ming Yi 4, Xiao Tan 1,2, Sen Liu 1,2,*
A dynamic interaction process between KaiA and KaiC is critical to the cyanobacterial circadian oscillator Pei Dong 1,2, Ying Fan 2, Jianqiang Sun 3, Mengting Lv 2, Ming Yi 4, Xiao Tan 1,2, Sen Liu 1,2,*
SUPPLEMENTARY INFORMATION
 Table of Contents Page Supplementary Table 1. Diffraction data collection statistics 2 Supplementary Table 2. Crystallographic refinement statistics 3 Supplementary Fig. 1. casic1mfc packing in the R3
Table of Contents Page Supplementary Table 1. Diffraction data collection statistics 2 Supplementary Table 2. Crystallographic refinement statistics 3 Supplementary Fig. 1. casic1mfc packing in the R3
Supporting Information. Accurate prediction of the structure and vibrational spectra of ionic
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Supporting Information Accurate prediction of the structure and vibrational spectra
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Supporting Information Accurate prediction of the structure and vibrational spectra
FW 1 CDR 1 FW 2 CDR 2
 Supplementary Figure 1 Supplementary Figure 1: Interface of the E9:Fas structure. The two interfaces formed by V H and V L of E9 with Fas are shown in stereo. The Fas receptor is represented as a surface
Supplementary Figure 1 Supplementary Figure 1: Interface of the E9:Fas structure. The two interfaces formed by V H and V L of E9 with Fas are shown in stereo. The Fas receptor is represented as a surface
Structural Insights from Molecular Dynamics. Simulations of Tryptophan 7-Halogenase and
 Supporting Information Structural Insights from Molecular Dynamics Simulations of Tryptophan 7-Halogenase and Tryptophan 5-halogenase Jon Ainsley 1, Adrian J. Mulholland 2, Gary W. Black 1, Olivier Sparagano
Supporting Information Structural Insights from Molecular Dynamics Simulations of Tryptophan 7-Halogenase and Tryptophan 5-halogenase Jon Ainsley 1, Adrian J. Mulholland 2, Gary W. Black 1, Olivier Sparagano
SUPPLEMENTARY INFORMATION
 Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
L718Q mutant EGFR escapes covalent inhibition by stabilizing. a non-reactive conformation of the lung cancer drug. osimertinib
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Electronic Supplementary Information (ESI) for L718Q mutant EGFR escapes covalent inhibition
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Electronic Supplementary Information (ESI) for L718Q mutant EGFR escapes covalent inhibition
Structural basis of PROTAC cooperative recognition for selective protein degradation
 SUPPLEMENTARY INFORMATION Structural basis of PROTAC cooperative recognition for selective protein degradation Morgan S. Gadd 1, Andrea Testa 1, Xavier Lucas 1, Kwok-Ho Chan, Wenzhang Chen, Douglas J.
SUPPLEMENTARY INFORMATION Structural basis of PROTAC cooperative recognition for selective protein degradation Morgan S. Gadd 1, Andrea Testa 1, Xavier Lucas 1, Kwok-Ho Chan, Wenzhang Chen, Douglas J.
HOMOLOGY MODELING. The sequence alignment and template structure are then used to produce a structural model of the target.
 HOMOLOGY MODELING Homology modeling, also known as comparative modeling of protein refers to constructing an atomic-resolution model of the "target" protein from its amino acid sequence and an experimental
HOMOLOGY MODELING Homology modeling, also known as comparative modeling of protein refers to constructing an atomic-resolution model of the "target" protein from its amino acid sequence and an experimental
A conserved P-loop anchor limits the structural dynamics that mediate. nucleotide dissociation in EF-Tu.
 Supplemental Material for A conserved P-loop anchor limits the structural dynamics that mediate nucleotide dissociation in EF-Tu. Evan Mercier 1,2, Dylan Girodat 1, and Hans-Joachim Wieden 1 * 1 Alberta
Supplemental Material for A conserved P-loop anchor limits the structural dynamics that mediate nucleotide dissociation in EF-Tu. Evan Mercier 1,2, Dylan Girodat 1, and Hans-Joachim Wieden 1 * 1 Alberta
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
β 2 -microglobulin fibrillation?
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Can small hydrophobic gold nanoparticles inhibit β 2 -microglobulin fibrillation? G. Brancolini,,
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Can small hydrophobic gold nanoparticles inhibit β 2 -microglobulin fibrillation? G. Brancolini,,
Supporting Information
 Supporting Information Micelle-Triggered b-hairpin to a-helix Transition in a 14-Residue Peptide from a Choline-Binding Repeat of the Pneumococcal Autolysin LytA HØctor Zamora-Carreras, [a] Beatriz Maestro,
Supporting Information Micelle-Triggered b-hairpin to a-helix Transition in a 14-Residue Peptide from a Choline-Binding Repeat of the Pneumococcal Autolysin LytA HØctor Zamora-Carreras, [a] Beatriz Maestro,
Intrinsic Dynamics of Restriction Endonuclease EcoO109I Studied by Molecular Dynamics Simulations and X-Ray Scattering Data Analysis
 2808 Biophysical Journal Volume 96 April 2009 2808 2822 Intrinsic Dynamics of Restriction Endonuclease EcoO109I Studied by Molecular Dynamics Simulations and X-Ray Scattering Data Analysis Tomotaka Oroguchi,
2808 Biophysical Journal Volume 96 April 2009 2808 2822 Intrinsic Dynamics of Restriction Endonuclease EcoO109I Studied by Molecular Dynamics Simulations and X-Ray Scattering Data Analysis Tomotaka Oroguchi,
Ensemble refinement of protein crystal structures in PHENIX. Tom Burnley Piet Gros
 Ensemble refinement of protein crystal structures in PHENIX Tom Burnley Piet Gros Incomplete modelling of disorder contributes to R factor gap Only ~5% of residues in the PDB are modelled with more than
Ensemble refinement of protein crystal structures in PHENIX Tom Burnley Piet Gros Incomplete modelling of disorder contributes to R factor gap Only ~5% of residues in the PDB are modelled with more than
Relative binding enthalpies from molecular dynamics simulations. using a direct method
 Relative binding enthalpies from molecular dynamics simulations using a direct method Amitava Roy, 1 Duy P. Hua, 1 Joshua M. Ward, 1 and Carol Beth Post* Department of Medicinal Chemistry, Markey Center
Relative binding enthalpies from molecular dynamics simulations using a direct method Amitava Roy, 1 Duy P. Hua, 1 Joshua M. Ward, 1 and Carol Beth Post* Department of Medicinal Chemistry, Markey Center
SUPPLEMENTARY MATERIAL. Supplementary material and methods:
 Electronic Supplementary Material (ESI) for Catalysis Science & Technology. This journal is The Royal Society of Chemistry 2015 SUPPLEMENTARY MATERIAL Supplementary material and methods: - Computational
Electronic Supplementary Material (ESI) for Catalysis Science & Technology. This journal is The Royal Society of Chemistry 2015 SUPPLEMENTARY MATERIAL Supplementary material and methods: - Computational
Analysis of MD trajectories in GROMACS David van der Spoel
 Analysis of MD trajectories in GROMACS David van der Spoel What does MD produce? Energy terms E(t) Coordinates x(t) Velocities v(t) Forces f(t) Managing your files trjcat - merging trajectories concatenating
Analysis of MD trajectories in GROMACS David van der Spoel What does MD produce? Energy terms E(t) Coordinates x(t) Velocities v(t) Forces f(t) Managing your files trjcat - merging trajectories concatenating
Insights on Antibiotic Translocation : Molecular Dynamics Simulations
 I Insights on Antibiotic Translocation : Molecular Dynamics Simulations Amit Kumar, Eric Hajjar,Enrico Spiga, Francesa Collu, Paolo Ruggerone & Matteo Ceccarelli Marseille : 11 04 2008 Outline METHODS
I Insights on Antibiotic Translocation : Molecular Dynamics Simulations Amit Kumar, Eric Hajjar,Enrico Spiga, Francesa Collu, Paolo Ruggerone & Matteo Ceccarelli Marseille : 11 04 2008 Outline METHODS
Supplementary Materials for
 www.advances.sciencemag.org/cgi/content/full/1/7/e1500263/dc1 Supplementary Materials for Newton s cradle proton relay with amide imidic acid tautomerization in inverting cellulase visualized by neutron
www.advances.sciencemag.org/cgi/content/full/1/7/e1500263/dc1 Supplementary Materials for Newton s cradle proton relay with amide imidic acid tautomerization in inverting cellulase visualized by neutron
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
Unfolding CspB by means of biased molecular dynamics
 Chapter 4 Unfolding CspB by means of biased molecular dynamics 4.1 Introduction Understanding the mechanism of protein folding has been a major challenge for the last twenty years, as pointed out in the
Chapter 4 Unfolding CspB by means of biased molecular dynamics 4.1 Introduction Understanding the mechanism of protein folding has been a major challenge for the last twenty years, as pointed out in the
ψ j φ j+1 Motif position
 1.0 % Occupancy 0.8 0.6 0.4 0.2 0.0 ψ i-1 φ i ψ j φ j+1 ψ k-1 Motif position φ Supplementary Figure 1 Residue profiles for the positions in the β-sheet motif. Residue types as percentages for the six positions
1.0 % Occupancy 0.8 0.6 0.4 0.2 0.0 ψ i-1 φ i ψ j φ j+1 ψ k-1 Motif position φ Supplementary Figure 1 Residue profiles for the positions in the β-sheet motif. Residue types as percentages for the six positions
Energetics and Dynamics of the Non-Natural Fluorescent 4AP:DAP Base pair
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Energetics and Dynamics of the Non-Natural Fluorescent 4AP:DAP Base pair Mohit
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Energetics and Dynamics of the Non-Natural Fluorescent 4AP:DAP Base pair Mohit
A: Up regulated proteins B: Down regulated proteins. Susceptible Resistant Susceptible Resistant Resistant Susceptible
 Supplementary Materials: Identification of Biomarkers for Resistance to Fusarium oxysporum f. sp. cubense Infection and in Silico Studies in Musa paradisiaca Cultivar Puttabale through Proteomic Approach
Supplementary Materials: Identification of Biomarkers for Resistance to Fusarium oxysporum f. sp. cubense Infection and in Silico Studies in Musa paradisiaca Cultivar Puttabale through Proteomic Approach
Modeling for 3D structure prediction
 Modeling for 3D structure prediction What is a predicted structure? A structure that is constructed using as the sole source of information data obtained from computer based data-mining. However, mixing
Modeling for 3D structure prediction What is a predicted structure? A structure that is constructed using as the sole source of information data obtained from computer based data-mining. However, mixing
Table S1. Primers used for the constructions of recombinant GAL1 and λ5 mutants. GAL1-E74A ccgagcagcgggcggctgtctttcc ggaaagacagccgcccgctgctcgg
 SUPPLEMENTAL DATA Table S1. Primers used for the constructions of recombinant GAL1 and λ5 mutants Sense primer (5 to 3 ) Anti-sense primer (5 to 3 ) GAL1 mutants GAL1-E74A ccgagcagcgggcggctgtctttcc ggaaagacagccgcccgctgctcgg
SUPPLEMENTAL DATA Table S1. Primers used for the constructions of recombinant GAL1 and λ5 mutants Sense primer (5 to 3 ) Anti-sense primer (5 to 3 ) GAL1 mutants GAL1-E74A ccgagcagcgggcggctgtctttcc ggaaagacagccgcccgctgctcgg
Energetics of Ion Permeation in an Open-Activated TRPV1 Channel
 Biophysical Journal, Volume 111 Supplemental Information Energetics of Ion Permeation in an Open-Activated TRPV1 Channel Christian Jorgensen, Simone Furini, and Carmen Domene Supporting Material Energetics
Biophysical Journal, Volume 111 Supplemental Information Energetics of Ion Permeation in an Open-Activated TRPV1 Channel Christian Jorgensen, Simone Furini, and Carmen Domene Supporting Material Energetics
Supplementary Information
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Supplementary Information Dynamics of the thumb-finger regions in GH11 xylanase Bacillus circulans:
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Supplementary Information Dynamics of the thumb-finger regions in GH11 xylanase Bacillus circulans:
Table 1. Crystallographic data collection, phasing and refinement statistics. Native Hg soaked Mn soaked 1 Mn soaked 2
 Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Protein Structure Prediction, Engineering & Design CHEM 430
 Protein Structure Prediction, Engineering & Design CHEM 430 Eero Saarinen The free energy surface of a protein Protein Structure Prediction & Design Full Protein Structure from Sequence - High Alignment
Protein Structure Prediction, Engineering & Design CHEM 430 Eero Saarinen The free energy surface of a protein Protein Structure Prediction & Design Full Protein Structure from Sequence - High Alignment
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Supplementary information for:
 SUPPLEMETARY IFRMATI Supplementary information for: Structure of a β 1 -adrenergic G protein-coupled receptor Tony Warne, Maria J. Serrano-Vega, Jillian G. Baker#, Rouslan Moukhametzianov, Patricia C.
SUPPLEMETARY IFRMATI Supplementary information for: Structure of a β 1 -adrenergic G protein-coupled receptor Tony Warne, Maria J. Serrano-Vega, Jillian G. Baker#, Rouslan Moukhametzianov, Patricia C.
Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08 Å resolution: comparison with the Azotobacter vinelandii MoFe protein
 Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data
 Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN
 THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figures. Measuring Similarity Between Dynamic Ensembles of Biomolecules
 Supplementary Figures Measuring Similarity Between Dynamic Ensembles of Biomolecules Shan Yang, Loïc Salmon 2, and Hashim M. Al-Hashimi 3*. Department of Chemistry, University of Michigan, Ann Arbor, MI,
Supplementary Figures Measuring Similarity Between Dynamic Ensembles of Biomolecules Shan Yang, Loïc Salmon 2, and Hashim M. Al-Hashimi 3*. Department of Chemistry, University of Michigan, Ann Arbor, MI,
The Effect of Linker DNA on the Structure and Interaction of Nucleosome Core Particles
 Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 SUPPLEMENTARY INFORMATION The Effect of Linker DNA on the Structure and Interaction of Nucleosome
Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 SUPPLEMENTARY INFORMATION The Effect of Linker DNA on the Structure and Interaction of Nucleosome
of the Guanine Nucleotide Exchange Factor FARP2
 Structure, Volume 21 Supplemental Information Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2 Xiaojing He, Yi-Chun Kuo, Tyler J. Rosche, and Xuewu Zhang Inventory of
Structure, Volume 21 Supplemental Information Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2 Xiaojing He, Yi-Chun Kuo, Tyler J. Rosche, and Xuewu Zhang Inventory of
Time-dependence of key H-bond/electrostatic interaction distances in the sirna5-hago2 complexes... Page S14
 Supporting Information Probing the Binding Interactions between Chemically Modified sirnas and Human Argonaute 2 Using Microsecond Molecular Dynamics Simulations S. Harikrishna* and P. I. Pradeepkumar*
Supporting Information Probing the Binding Interactions between Chemically Modified sirnas and Human Argonaute 2 Using Microsecond Molecular Dynamics Simulations S. Harikrishna* and P. I. Pradeepkumar*
Structural characterization of NiV N 0 P in solution and in crystal.
 Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides Zaixing
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides Zaixing
Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits. AhpC and AhpF from Escherichia coli
 Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits AhpC and AhpF from Escherichia coli Phat Vinh Dip 1,#, Neelagandan Kamariah 2,#, Malathy Sony Subramanian Manimekalai
Structure, mechanism and ensemble formation of the Alkylhydroperoxide Reductase subunits AhpC and AhpF from Escherichia coli Phat Vinh Dip 1,#, Neelagandan Kamariah 2,#, Malathy Sony Subramanian Manimekalai
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
CONFORMATIONAL SEARCH OF PROTEINS AND PROTEIN LOOPS
 CONFORMATIONAL SEARCH OF PROTEINS AND PROTEIN LOOPS By Ranjitha Venkataramani Submitted to the Department of Chemistry and the Faculty of the Graduate School of the University of Kansas in partial fulfillment
CONFORMATIONAL SEARCH OF PROTEINS AND PROTEIN LOOPS By Ranjitha Venkataramani Submitted to the Department of Chemistry and the Faculty of the Graduate School of the University of Kansas in partial fulfillment
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R)
 Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Type II Kinase Inhibitors Show an Unexpected Inhibition Mode against Parkinson s Disease-Linked LRRK2 Mutant G2019S.
 Type II Kinase Inhibitors Show an Unexpected Inhibition Mode against Parkinson s Disease-Linked LRRK2 Mutant G219S. Min Liu@&*, Samantha A. Bender%*, Gregory D Cuny@, Woody Sherman, Marcie Glicksman@ Soumya
Type II Kinase Inhibitors Show an Unexpected Inhibition Mode against Parkinson s Disease-Linked LRRK2 Mutant G219S. Min Liu@&*, Samantha A. Bender%*, Gregory D Cuny@, Woody Sherman, Marcie Glicksman@ Soumya
SUPPLEMENTARY FIGURES. Figure S1
 SUPPLEMENTARY FIGURES Figure S1 The substrate for DH domain (2R,3R,4R,6R,7S,8S,9R)-3,7,9-trihydroxy-5-oxo-2,4,6,8 tetramethylundecanoate) was docked as two separate fragments shown in magenta and blue
SUPPLEMENTARY FIGURES Figure S1 The substrate for DH domain (2R,3R,4R,6R,7S,8S,9R)-3,7,9-trihydroxy-5-oxo-2,4,6,8 tetramethylundecanoate) was docked as two separate fragments shown in magenta and blue
This document is downloaded from DR-NTU, Nanyang Technological University Library, Singapore.
 This document is downloaded from DR-NTU, Nanyang Technological University Library, Singapore. Title Author(s) Citation Coarse-Grained Molecular Modeling of the Solution Structure Ensemble of Dengue Virus
This document is downloaded from DR-NTU, Nanyang Technological University Library, Singapore. Title Author(s) Citation Coarse-Grained Molecular Modeling of the Solution Structure Ensemble of Dengue Virus
Structure Investigation of Fam20C, a Golgi Casein Kinase
 Structure Investigation of Fam20C, a Golgi Casein Kinase Sharon Grubner National Taiwan University, Dr. Jung-Hsin Lin University of California San Diego, Dr. Rommie Amaro Abstract This research project
Structure Investigation of Fam20C, a Golgi Casein Kinase Sharon Grubner National Taiwan University, Dr. Jung-Hsin Lin University of California San Diego, Dr. Rommie Amaro Abstract This research project
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 28 Electronic Supplementary Information ph-dependent Cooperativity and Existence of
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 28 Electronic Supplementary Information ph-dependent Cooperativity and Existence of
Essential dynamics sampling of proteins. Tuorial 6 Neva Bešker
 Essential dynamics sampling of proteins Tuorial 6 Neva Bešker Relevant time scale Why we need enhanced sampling? Interconvertion between basins is infrequent at the roomtemperature: kinetics and thermodynamics
Essential dynamics sampling of proteins Tuorial 6 Neva Bešker Relevant time scale Why we need enhanced sampling? Interconvertion between basins is infrequent at the roomtemperature: kinetics and thermodynamics
Supplementary Information: Table S1. Potential energy and essential dynamics (ED) analysis of the MD simulations of the Bcl-2 complexes under study.
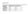 Supplementary Information: Table S1. Potential energy and essential dynamics (ED) analysis of the MD simulations of the Bcl-2 complexes under study. Bcl2 and complex Potential energy (kj/mol) ED 2D projection
Supplementary Information: Table S1. Potential energy and essential dynamics (ED) analysis of the MD simulations of the Bcl-2 complexes under study. Bcl2 and complex Potential energy (kj/mol) ED 2D projection
Improving De novo Protein Structure Prediction using Contact Maps Information
 CIBCB 2017 IEEE Conference on Computational Intelligence in Bioinformatics and Computational Biology Improving De novo Protein Structure Prediction using Contact Maps Information Karina Baptista dos Santos
CIBCB 2017 IEEE Conference on Computational Intelligence in Bioinformatics and Computational Biology Improving De novo Protein Structure Prediction using Contact Maps Information Karina Baptista dos Santos
An engineered scorpion toxin analogue with improved Kv1.3 selectivity displays reduced conformational flexibility
 An engineered scorpion toxin analogue with improved Kv1.3 selectivity displays reduced conformational flexibility Adam Bartok a,#, Krisztina Fehér b,c,#,, Andrea Bodor d, Kinga Rákosi e, Gábor K. Tóth
An engineered scorpion toxin analogue with improved Kv1.3 selectivity displays reduced conformational flexibility Adam Bartok a,#, Krisztina Fehér b,c,#,, Andrea Bodor d, Kinga Rákosi e, Gábor K. Tóth
What makes a good graphene-binding peptide? Adsorption of amino acids and peptides at aqueous graphene interfaces: Electronic Supplementary
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 21 What makes a good graphene-binding peptide? Adsorption of amino acids and
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 21 What makes a good graphene-binding peptide? Adsorption of amino acids and
Supplemental Data SUPPLEMENTAL FIGURES
 Supplemental Data CRYSTAL STRUCTURE OF THE MG.ADP-INHIBITED STATE OF THE YEAST F 1 C 10 ATP SYNTHASE Alain Dautant*, Jean Velours and Marie-France Giraud* From Université Bordeaux 2, CNRS; Institut de
Supplemental Data CRYSTAL STRUCTURE OF THE MG.ADP-INHIBITED STATE OF THE YEAST F 1 C 10 ATP SYNTHASE Alain Dautant*, Jean Velours and Marie-France Giraud* From Université Bordeaux 2, CNRS; Institut de
Improving Protein Function Prediction with Molecular Dynamics Simulations. Dariya Glazer Russ Altman
 Improving Protein Function Prediction with Molecular Dynamics Simulations Dariya Glazer Russ Altman Motivation Sometimes the 3D structure doesn t score well for a known function. The experimental structure
Improving Protein Function Prediction with Molecular Dynamics Simulations Dariya Glazer Russ Altman Motivation Sometimes the 3D structure doesn t score well for a known function. The experimental structure
Full wwpdb X-ray Structure Validation Report i
 Full wwpdb X-ray Structure Validation Report i Mar 14, 2018 02:00 pm GMT PDB ID : 3RRQ Title : Crystal structure of the extracellular domain of human PD-1 Authors : Lazar-Molnar, E.; Ramagopal, U.A.; Nathenson,
Full wwpdb X-ray Structure Validation Report i Mar 14, 2018 02:00 pm GMT PDB ID : 3RRQ Title : Crystal structure of the extracellular domain of human PD-1 Authors : Lazar-Molnar, E.; Ramagopal, U.A.; Nathenson,
Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6
 Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Cks1 CDK1 CDK1 CDK1 CKS1. are ice- lobe. conserved. conserved
 Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Stabilizing the CH2 domain of an Antibody by Engineering in an Enhanced Aromatic Sequon
 Stabilizing the CH2 domain of an Antibody by Engineering in an Enhanced Aromatic Sequon Wentao Chen,, Leopold Kong, Stephen Connelly, Julia M. Dendle,, Yu Liu,, Ian A. Wilson,#, Evan T. Powers, *, Jeffery
Stabilizing the CH2 domain of an Antibody by Engineering in an Enhanced Aromatic Sequon Wentao Chen,, Leopold Kong, Stephen Connelly, Julia M. Dendle,, Yu Liu,, Ian A. Wilson,#, Evan T. Powers, *, Jeffery
Structurale, Université Grenoble Alpes, CNRS, CEA, Grenoble, France
 Supplementary Information to Lysine relay mechanism coordinates intermediate transfer in vitamin B6 biosynthesis Matthew J. Rodrigues 1,2, Volker Windeisen 1,3, Yang Zhang 4, Gabriela Guédez 3, Stefan
Supplementary Information to Lysine relay mechanism coordinates intermediate transfer in vitamin B6 biosynthesis Matthew J. Rodrigues 1,2, Volker Windeisen 1,3, Yang Zhang 4, Gabriela Guédez 3, Stefan
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Temperature dependence of dynamic, tunnelling
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Temperature dependence of dynamic, tunnelling
PROTEIN STRUCTURE PREDICTION USING GAS PHASE MOLECULAR DYNAMICS SIMULATION: EOTAXIN-3 CYTOKINE AS A CASE STUDY
 International Conference Mathematical and Computational Biology 2011 International Journal of Modern Physics: Conference Series Vol. 9 (2012) 193 198 World Scientific Publishing Company DOI: 10.1142/S2010194512005259
International Conference Mathematical and Computational Biology 2011 International Journal of Modern Physics: Conference Series Vol. 9 (2012) 193 198 World Scientific Publishing Company DOI: 10.1142/S2010194512005259
Supporting Information
 Supporting Information The Predicted Ensemble of Low-Energy Conformations of Human Somatostatin Receptor Subtype 5 and the Binding of Antagonists Sijia S. Dong, [a] Ravinder Abrol, [a, b] and William A.
Supporting Information The Predicted Ensemble of Low-Energy Conformations of Human Somatostatin Receptor Subtype 5 and the Binding of Antagonists Sijia S. Dong, [a] Ravinder Abrol, [a, b] and William A.
Docking. GBCB 5874: Problem Solving in GBCB
 Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Summary of Experimental Protein Structure Determination. Key Elements
 Programme 8.00-8.20 Summary of last week s lecture and quiz 8.20-9.00 Structure validation 9.00-9.15 Break 9.15-11.00 Exercise: Structure validation tutorial 11.00-11.10 Break 11.10-11.40 Summary & discussion
Programme 8.00-8.20 Summary of last week s lecture and quiz 8.20-9.00 Structure validation 9.00-9.15 Break 9.15-11.00 Exercise: Structure validation tutorial 11.00-11.10 Break 11.10-11.40 Summary & discussion
Molecular Mechanism for Conformational Dynamics of Ras GTP Elucidated from In-Situ Structural Transition in Crystal
 Molecular Mechanism for Conformational Dynamics of Ras GTP Elucidated from In-Situ Structural Transition in Crystal Shigeyuki Matsumoto, Nao Miyano, Seiki Baba, Jingling Liao, Takashi Kawamura, Chiemi
Molecular Mechanism for Conformational Dynamics of Ras GTP Elucidated from In-Situ Structural Transition in Crystal Shigeyuki Matsumoto, Nao Miyano, Seiki Baba, Jingling Liao, Takashi Kawamura, Chiemi
Goals. Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions
 Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions Jamie Duke 1,2 and Carlos Camacho 3 1 Bioengineering and Bioinformatics Summer Institute,
Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions Jamie Duke 1,2 and Carlos Camacho 3 1 Bioengineering and Bioinformatics Summer Institute,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Cationic carbosilane dendrimers and oligonucleotide binding: an energetic affair
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Cationic carbosilane dendrimers and oligonucleotide binding: an energetic
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Cationic carbosilane dendrimers and oligonucleotide binding: an energetic
Probing Molecular Mechanisms Of the Hsp90 Chaperone: Biophysical Modeling Identifies Key Regulators Of Functional Dynamics
 Chapman University Chapman University Digital Commons Mathematics, Physics, and Computer Science Faculty Articles and Research Science and Technology Faculty Articles and Research 2012 Probing Molecular
Chapman University Chapman University Digital Commons Mathematics, Physics, and Computer Science Faculty Articles and Research Science and Technology Faculty Articles and Research 2012 Probing Molecular
Table S1 Crystallographic data and structure refinement for complexes 1 and 2. Complex H 2 O
 Electronic Supplementary Information: Ternary oxovanadium(iv) complexes of ONO-donor Schiff base and polypyridyl derivatives as protein tyrosine phosphatase inhibitors: synthesis, characterization and
Electronic Supplementary Information: Ternary oxovanadium(iv) complexes of ONO-donor Schiff base and polypyridyl derivatives as protein tyrosine phosphatase inhibitors: synthesis, characterization and
