Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides
|
|
|
- Kelley Magnus Cobb
- 5 years ago
- Views:
Transcription
1 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides Zaixing Yang 1,ǂ, Cuicui Ge 1,ǂ, Jiajia Liu 1, Yu Chong 1, Zonglin Gu 1, Camilo A. Jimenez- Cruz 2, Zhifang Chai 1, and Ruhong Zhou 1,2,3 * 1 School for Radiological and Interdisciplinary Sciences (RAD-X), Collaborative Innovation Center of Radiation Medicine of Jiangsu Higher Education Institutions, and Jiangsu Provincial Key Laboratory of Radiation Medicine and Protection, Soochow University, Suzhou , China 2 IBM Thomas J. Watson Research Center, Yorktown Heights, NY 10598, USA 3 Department of Chemistry, Columbia University, New York, NY 10027, USA ǂ These authors contribute equally *Corresponding author, ruhongz@us.ibm.com Context: PS1: Other representative trajectories for all the four types of simulated models with double or single graphene nanosheets. PS2: Instruction to the video. PS3: Two representative trajectories to show graphene sheet(s) spontaneously penetrating into the fibril. PS4: One representative trajectory to show a larger graphene nanosheet spontaneously penetrating into the fibril with subsequent peptide extraction. PS5: Role of interfacial water in assisting graphene s penetration into fibril. PS6: Time evolution of the interaction potential energy illustrating the step-wise decrease upon lipid extraction PS7: Potential of mean force for the estimate of binding free energy. PS8: AFM images of s-go and GO. PS9: Sketches of restraints in all four types of simulated systems.
2 Supporting Information PS1: Other representative trajectories for all the four types of simulated models with double or single graphene nanosheets. Fig. S1 A and B show similar trajectories as shown in Fig. 1 in the main text, but with additional runs (i.e., the two graphene nanosheets either attacking from the same side (Fig. S1A) or from both sides (Fig. S1B)). Again, both are shown to be capable of disrupting the preformed amyloid fibril through insertion/cutting and the direct extraction. Fig. S1C represents a typical trajectory of a single graphene nanosheet attacking from the edge, similar to that in the docking simulation in Fig. 2 of the main text. Again, the extraction of peptides is commonly seen. Fig. S1D shows a single graphene nanosheet attacking from the middle of fibril. The graphene can still cut into and extract peptides from the fibril. All these results indicate that the destruction mechanism with both insertion/cutting and direct extraction is very robust. Fig. S1. The additional representative trajectories of graphene nanosheet(s) interaction with mature fibrils for all the four simulated models, with (A) showing two graphene sheets attacking a preformed A amyloid fibril from the same side, (B) the two graphene sheets attacking from both sides, (C) the top-edge-restrained
3 graphene nanosheet docked at the edge of the A amyloid fibril extracting peptides, and (D) a graphene sheet attacking from the middle of fibril. The A peptides (total 24 monomers) are shown in sticks, with the two aromatic residues Phe shown in dark blue. The graphene sheet is shown as an orange-bonded sheet. Extracted peptides are highlighted with their Phe residues shown in larger van der Waals spheres. Color settings are the same as in Fig. 1, in the main text. PS2: Instruction to the video. The video shows how the two graphene nanosheets, attacking from both sides of the fibril, insert/cut into the fibril and extract large amount of peptides in atomic details (for simplicity the solvent and ions were not shown). PS3: Two representative trajectories to show graphene sheet spontaneously penetrating into the fibril. Two sets of additional simulations have been performed to investigate whether graphene can still dissociate the amyloid or not once the constraints on the graphene and peptides are removed. Both the cases with double graphene sheets or single sheet were studied. As shown in the new Fig. S3A, for the double graphene sheets case, once the constraints were removed, one graphene sheet quickly penetrated into the fibril and cut it into two halves (graphene staying inside the fibril), while the other sheet laid on the fibril surface till the end of the simulation. For the single graphene case (see Fig. S3B), the graphene also quickly penetrated into the fibril and cut it into two halves, and stayed inside the fibril. The peptide extraction was not observed though in these simulations, probably partly due to the too small size used (thus not enough graphene surface for additional extracted peptides), and partly due to the fact that at least one sheet was laying flat on fibril surface, which also leaves no room for peptide extraction.
4 Fig. S3. Two representative trajectories to show graphene sheet spontaneously penetrate into the fibril; (A) for double graphene sheets, and (B) for single sheet attacking an amyloid fibril from the same side. PS4: One representative trajectory to show a larger graphene nanosheet spontaneously penetrating into the fibril with subsequent peptide extraction. To further illustrate the lipid extraction process, we doubled the size of the attacking graphene nanosheet in the constraint-free simulations. As shown in Fig. S4, after the graphene nanosheet (the right one) fully penetrated into the fibril at t = ~50 ns, the peptide extraction process started. At ~200 ns all the hanging-out portion of the inserted graphene was fully covered by the extracted peptides from fibril. This is also consistent with another very recent study where both the inserted bare and serum protein BSA-coated graphene nanosheets can extract lipids from cell membranes to their hanging-out portions, but with the BSA-coated one extracting less and covering only those available surfaces aside from BSA (Nanoscale, DOI: /C5NR01839K, 2015).
5 Fig. S4. One representative trajectory to show two graphene sheets spontaneously penetrate into the fibril, with the subsequent peptides extraction from the right graphene sheet. Other trajectories show the left graphene sheet extracts peptides first. If much longer simulations were performed, both graphene sheets will be covered by the extracted peptides eventually. PS5: Role of interfacial water in assisting graphene s penetration into fibril. The contributions of interfacial water between fibril and graphene to the processes of graphene s penetration can be described from the time evolution of the heavy atom contact number between graphene and fibril, as well as water solvation dynamics in the first solvation shell (FSS) for some representative residues, such as phenylalanine (aromatic) and lysine (basic) (Fig. S5). Initially, the sidechain of phenylalanine was buried in the hydrophobic core of fibril and remained dry. In this particular trajectory, as shown in Fig. S5, the -sheet structure started to get ruptured around t=100 ns, due to the asynchronous adsorption of peptides onto the surface of graphene, and water molecules quickly intruded into fibril s hydrophobic core. At = 105 ns, the target phenylalanine was already solvated by ~11 water molecules. From ~110 ns to ~180ns, as graphene further penetrating into the fibril hydrophobic core, the heavy atom contact number between the sidechain and graphene increased from 0 to ~65, while the number of water molecules in FSS of Phe side chain dramatically decreased from ~18 to ~0. In other words, a fascinating dewetting (drying) phenomenon occurred at the interface between graphene and the hydrophobic cluster near the target phenylalanine, which provided a strong driving force for graphene s further penetration. It is noteworthy that during this drying process, the aromatic ring of
6 target phenylalanine is actually perpendicular to the graphene. As the simulation time progresses (after ~250 ns), the sidechain of the target phenylalanine climbed onto the surface of graphene, accompanied by rotating its aromatic ring to be parallel to the graphene surface, interestingly. This results in more aromatic ring atoms in contact with graphene, with the heavy atom contact number increased from ~65 to ~200, and thus a significantly strengthened stacking interaction between graphene and phenylalanine. From then on, the target phenylalanine stayed in that conformation till to the end of the simulation, with both the heavy atom contact number and the number of water molecules in FSS nearly constant. For lysine, on the other hand, its long sidechain (and especially the positive charged -amino group, NH 3+ ) was fully solvated by ~18 water molecules (see the snapshot at t = 0 and 220 ns in Fig. S5) during the first ~250 ns of the simulation before it started to approach the graphene surface. Then, within a relatively short time interval from ~250 to ~300 ns, it was fully adsorbed onto the graphene surface. During this process, the heavy atom contact number between the side chain and graphene increased sharply from 0 to ~120, while the number of water molecules in its FSS only slightly decreased from ~18 to ~15 (see Fig. S5B), which is in contrast with the phenylalanine case. Taken together, we can conclude that water molecules near the interface have played a significant role during the graphene insertion and peptide adsorption process. These water molecules have played at least two roles in this case: (i) to provide a strong driving force through final drying of hydrophobic residues, particularly phenylalanine, upon binding onto graphene; and (ii) to facilitate as a lubricant for basic residues, such as lysine, to bind to graphene. Fig. S5. The number of water molecules in first solvation shell (FSS) of Phe (A) and Lys (B) residues (top panel in the first graph), and the heavy atom contact number
7 between the residue and graphene (bottom panel in the first graph). The representative snapshots showing the local solvation of Phe (A) and Lys (B) are also shown. The red spheres represent the water oxygen atoms in FSS of target residue sidechain. The A peptides (consisting of a total of 24 monomers) are shown in a cartoon representation, with the representative phenylalanine and lysine residues shown in blue and light blue surfaces. The graphene sheet is shown as an orange-flat-sheet. PS6: Time evolution of the interaction potential energy illustrating the step-wise decrease upon lipid extraction In addition to the PMF calculations, which clearly show the difference in binding energies (see Fig. S7), we also analyzed in detail the time evolution of the interaction potential energy changes to further illustrate this point. As shown in Fig. S6, two peptides with very similar positions with respect to the two graphene nanosheets were chosen to demonstrate their respective energy changes (step-wise decrease) once moving from the fibril to the graphene sheet. The interaction energies for peptide-1 and peptide-2 with fibril are about , and kcal/mol; while they are and kcal/mol with graphene once extracted (Fig. S6C). Thus, the net interaction potential energy will be significantly lower by and kcal/mol, respectively, for peptide-1 and peptide-2, displaying a step-wise decrease upon extraction. Fig. S6. (A) One representative trajectory showing the graphene nanosheet insertion and peptide extraction, featuring two graphene sheets attacking a pre-formed A amyloid fibril from the same side (the same with the trajectory shown in Fig.1A in the main text). (B) Time evolution of the interaction potential energy between the graphene nanosheet and the peptide amyloid fibril (in black), alongside the contact area between the graphene and peptides (in red) for the trajectory. (C) The interaction potential energy between the peptide-1 (top panel) and peptide-2 (bottom panel) with
8 the graphene nanosheet (black) and fibril (red) as the function of simulation time, displaying a clear step-wise decrease in interaction potential energy once the peptide is extracted onto the graphene sheet. PS7: Potential of mean force for the estimate of binding free energy. Three sets of umbrella sampling simulations were performed to estimate the PMF for the peptide-fibril (from both interior and terminal) and peptide-graphene binding systems. Fig. S7 shows the sketches of these binding systems: (A) a peptide being pulled from the interior of the fibril, (B) a peptide being pulled from the terminal of the fibril, and (C) a peptide being pulled from the graphene surface. The peptide in green is the target peptide for pulling and the arrow indicates the pulling direction (PMF reaction coordinate). The PMF curves were obtained by using 25, 31 and 43 sampling windows along the reaction coordinates of the peptide in the interior, at the terminal of the fibril, and the peptide-graphene complex, respectively. The binding free energy for a peptide in the interior and at the terminal are calculated to be kcal/mol and kcal/mol (Fig. S7A-B), respectively. While it is kcal/mol for the same peptide with graphene (Fig. S7C), which is about 3.12 and kcal/mol more favorable than the peptide in the interior or at the terminal. These results further confirm that extraction of peptides from fibril to the graphene surface is energetically favorable.
9 Fig. S7. The potential of mean force (PMF) calculations for the peptide KLVFFA being pulled from the interior (A) of the fibril, the terminal (B) of the fibril, and graphene surface (C). The left subfigure shows a sketch of the system, while the right subfigure shows the corresponding PMF. The peptide in green is the target peptide for pulling and the arrow indicates the pulling direction. PS8: AFM images of s-go and GO. Fig. S8. AFM images of small sized graphene oxide (s-go, with a lateral size <15 nm) (A), and normal sized GO with a lateral size of 0.5-3µm (B). PS9: Sketches of restraints in all four types of simulated systems. Fig. S9A-D show the sketches of restraints in all four types of simulated systems. All constrained atoms on graphene and peptide have been shown with large balls. Except for the graphene docking simulations, the positions of sixteen carbon atoms in four carbon rings at one of the corners were constrained by using a position restraint, while the other carbon atoms were left free to move. For the graphene docking simulations, the entire top row of 6-membered rings was restrained. The two C atoms at the two ends of the terminal four peptide chains in the -sheet were also restrained, while the remaining peptides were allowed to move freely.
10 Fig. S9. A schematic picture of restrained atoms in all the four types of simulated systems. (A) two graphene sheets attacking a preformed A amyloid fibril from the same side, (B) the two graphene sheets attacking from both sides, (C) the top-edgerestrained graphene nanosheet docked near the surface of the A amyloid fibril, and (D) a graphene sheet attacking from the middle of fibril. All restrained atoms are shown in cyan vdw balls, graphene in orange-colored sheets, and A peptides (consisting of a total of 24 monomers) in grey cartoon representations.
A Tunable, Strain-Controlled Nanoporous MoS 2 Filter for Water Desalination
 Supporting Information A Tunable, Strain-Controlled Nanoporous MoS 2 Filter for Water Desalination Weifeng Li 1, Yanmei Yang 1, Jeffrey K. Weber 2, Gang Zhang* 3, Ruhong Zhou* 1,2,4 1. School for Radiological
Supporting Information A Tunable, Strain-Controlled Nanoporous MoS 2 Filter for Water Desalination Weifeng Li 1, Yanmei Yang 1, Jeffrey K. Weber 2, Gang Zhang* 3, Ruhong Zhou* 1,2,4 1. School for Radiological
Supplementary Information
 Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
Supplementary Information
 1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
Supporting Information for: Mechanism of Reversible Peptide-Bilayer. Attachment: Combined Simulation and Experimental Single-Molecule Study
 Supporting Information for: Mechanism of Reversible Peptide-Bilayer Attachment: Combined Simulation and Experimental Single-Molecule Study Nadine Schwierz a,, Stefanie Krysiak c, Thorsten Hugel c,d, Martin
Supporting Information for: Mechanism of Reversible Peptide-Bilayer Attachment: Combined Simulation and Experimental Single-Molecule Study Nadine Schwierz a,, Stefanie Krysiak c, Thorsten Hugel c,d, Martin
Problem Set 1
 2006 7.012 Problem Set 1 Due before 5 PM on FRIDAY, September 15, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. For each of the following parts, pick
2006 7.012 Problem Set 1 Due before 5 PM on FRIDAY, September 15, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. For each of the following parts, pick
Structural insights into Aspergillus fumigatus lectin specificity - AFL binding sites are functionally non-equivalent
 Acta Cryst. (2015). D71, doi:10.1107/s1399004714026595 Supporting information Volume 71 (2015) Supporting information for article: Structural insights into Aspergillus fumigatus lectin specificity - AFL
Acta Cryst. (2015). D71, doi:10.1107/s1399004714026595 Supporting information Volume 71 (2015) Supporting information for article: Structural insights into Aspergillus fumigatus lectin specificity - AFL
Experimental and Theoretical Calculation Investigation on
 Electronic Supplementary Material (ESI) for Environmental Science: Nano. This journal is The Royal Society of Chemistry 2018 Experimental and Theoretical Calculation Investigation on Efficient Pb(II) Adsorption
Electronic Supplementary Material (ESI) for Environmental Science: Nano. This journal is The Royal Society of Chemistry 2018 Experimental and Theoretical Calculation Investigation on Efficient Pb(II) Adsorption
Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate
 Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate Shidi Zhao a, Laura Restrepo-Pérez b, Misha Soskine c, Giovanni Maglia c, Chirlmin Joo b, Cees Dekker b and Aleksei Aksimentiev
Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate Shidi Zhao a, Laura Restrepo-Pérez b, Misha Soskine c, Giovanni Maglia c, Chirlmin Joo b, Cees Dekker b and Aleksei Aksimentiev
From Amino Acids to Proteins - in 4 Easy Steps
 From Amino Acids to Proteins - in 4 Easy Steps Although protein structure appears to be overwhelmingly complex, you can provide your students with a basic understanding of how proteins fold by focusing
From Amino Acids to Proteins - in 4 Easy Steps Although protein structure appears to be overwhelmingly complex, you can provide your students with a basic understanding of how proteins fold by focusing
Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine. JAK2 Selective Mechanism Combined Molecular Dynamics Simulation
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Supporting Information Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Supporting Information Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine
What makes a good graphene-binding peptide? Adsorption of amino acids and peptides at aqueous graphene interfaces: Electronic Supplementary
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 21 What makes a good graphene-binding peptide? Adsorption of amino acids and
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 21 What makes a good graphene-binding peptide? Adsorption of amino acids and
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached
 Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached to HpUreI. Urea hydrolysis products 2NH 3 and 1CO 2
Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached to HpUreI. Urea hydrolysis products 2NH 3 and 1CO 2
Supporting Information
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supporting Information Lipid molecules can induce an opening of membrane-facing
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supporting Information Lipid molecules can induce an opening of membrane-facing
School of Nuclear Science and Engineering, East China University of Technology,
 Large-pore Layered Networks, Polycatenated Frameworks and Three-dimensional Frameworks of Uranyl Tri(biphenyl)amine/Tri(phenyl)amine Tricarboxylate: Solvent/Ligand-dependent Dual Regulation Shuai Wang,,,#
Large-pore Layered Networks, Polycatenated Frameworks and Three-dimensional Frameworks of Uranyl Tri(biphenyl)amine/Tri(phenyl)amine Tricarboxylate: Solvent/Ligand-dependent Dual Regulation Shuai Wang,,,#
Au-C Au-Au. g(r) r/a. Supplementary Figures
 g(r) Supplementary Figures 60 50 40 30 20 10 0 Au-C Au-Au 2 4 r/a 6 8 Supplementary Figure 1 Radial bond distributions for Au-C and Au-Au bond. The zero density regime between the first two peaks in g
g(r) Supplementary Figures 60 50 40 30 20 10 0 Au-C Au-Au 2 4 r/a 6 8 Supplementary Figure 1 Radial bond distributions for Au-C and Au-Au bond. The zero density regime between the first two peaks in g
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
PROTEIN STRUCTURE AMINO ACIDS H R. Zwitterion (dipolar ion) CO 2 H. PEPTIDES Formal reactions showing formation of peptide bond by dehydration:
 PTEI STUTUE ydrolysis of proteins with aqueous acid or base yields a mixture of free amino acids. Each type of protein yields a characteristic mixture of the ~ 20 amino acids. AMI AIDS Zwitterion (dipolar
PTEI STUTUE ydrolysis of proteins with aqueous acid or base yields a mixture of free amino acids. Each type of protein yields a characteristic mixture of the ~ 20 amino acids. AMI AIDS Zwitterion (dipolar
Viewing and Analyzing Proteins, Ligands and their Complexes 2
 2 Viewing and Analyzing Proteins, Ligands and their Complexes 2 Overview Viewing the accessible surface Analyzing the properties of proteins containing thousands of atoms is best accomplished by representing
2 Viewing and Analyzing Proteins, Ligands and their Complexes 2 Overview Viewing the accessible surface Analyzing the properties of proteins containing thousands of atoms is best accomplished by representing
Supplementary Figure 1 Experimental setup for crystal growth. Schematic drawing of the experimental setup for C 8 -BTBT crystal growth.
 Supplementary Figure 1 Experimental setup for crystal growth. Schematic drawing of the experimental setup for C 8 -BTBT crystal growth. Supplementary Figure 2 AFM study of the C 8 -BTBT crystal growth
Supplementary Figure 1 Experimental setup for crystal growth. Schematic drawing of the experimental setup for C 8 -BTBT crystal growth. Supplementary Figure 2 AFM study of the C 8 -BTBT crystal growth
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Amino Acids and Proteins at ZnO-water Interfaces in Molecular Dynamics Simulations: Electronic Supplementary Information
 Amino Acids and Proteins at ZnO-water Interfaces in Molecular Dynamics Simulations: Electronic Supplementary Information Grzegorz Nawrocki and Marek Cieplak Institute of Physics, Polish Academy of Sciences,
Amino Acids and Proteins at ZnO-water Interfaces in Molecular Dynamics Simulations: Electronic Supplementary Information Grzegorz Nawrocki and Marek Cieplak Institute of Physics, Polish Academy of Sciences,
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2720 1 2 3 Tuning underwater adhesion with cation-π interactions Matthew A. Gebbie, Wei Wei, Alex M. Schrader,
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2720 1 2 3 Tuning underwater adhesion with cation-π interactions Matthew A. Gebbie, Wei Wei, Alex M. Schrader,
The protein folding problem consists of two parts:
 Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Molecular Modeling Lecture 11 side chain modeling rotamers rotamer explorer buried cavities.
 Molecular Modeling 218 Lecture 11 side chain modeling rotamers rotamer explorer buried cavities. Sidechain Rotamers Discrete approximation of the continuous space of backbone angles. Sidechain conformations
Molecular Modeling 218 Lecture 11 side chain modeling rotamers rotamer explorer buried cavities. Sidechain Rotamers Discrete approximation of the continuous space of backbone angles. Sidechain conformations
Fibril Elongation by Aβ : Kinetic Network Analysis of Hybrid- Resolution Molecular Dynamics Simulations
 pubs.acs.org/jacs Terms of Use Fibril Elongation by Aβ 17 42 : Kinetic Network Analysis of Hybrid- Resolution Molecular Dynamics Simulations Wei Han, and Klaus Schulten*,,, Beckman Institute, Center for
pubs.acs.org/jacs Terms of Use Fibril Elongation by Aβ 17 42 : Kinetic Network Analysis of Hybrid- Resolution Molecular Dynamics Simulations Wei Han, and Klaus Schulten*,,, Beckman Institute, Center for
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Molecular Wire of Urea in Carbon Nanotube: A Molecular. Dynamics Study
 Molecular Wire of Urea in Carbon Nanotube: A Molecular Dynamics Study Peng Xiu, 1,2 Yusong Tu, 3 Xingling Tian, 1 Haiping Fang, 4* and Ruhong Zhou 5* 1 Bio-X Lab, Department of Physics, and Soft Matter
Molecular Wire of Urea in Carbon Nanotube: A Molecular Dynamics Study Peng Xiu, 1,2 Yusong Tu, 3 Xingling Tian, 1 Haiping Fang, 4* and Ruhong Zhou 5* 1 Bio-X Lab, Department of Physics, and Soft Matter
NH 2. Biochemistry I, Fall Term Sept 9, Lecture 5: Amino Acids & Peptides Assigned reading in Campbell: Chapter
 Biochemistry I, Fall Term Sept 9, 2005 Lecture 5: Amino Acids & Peptides Assigned reading in Campbell: Chapter 3.1-3.4. Key Terms: ptical Activity, Chirality Peptide bond Condensation reaction ydrolysis
Biochemistry I, Fall Term Sept 9, 2005 Lecture 5: Amino Acids & Peptides Assigned reading in Campbell: Chapter 3.1-3.4. Key Terms: ptical Activity, Chirality Peptide bond Condensation reaction ydrolysis
Cationic carbosilane dendrimers and oligonucleotide binding: an energetic affair
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Cationic carbosilane dendrimers and oligonucleotide binding: an energetic
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Cationic carbosilane dendrimers and oligonucleotide binding: an energetic
Molecular Basis of K + Conduction and Selectivity
 The Structure of the Potassium Channel: Molecular Basis of K + Conduction and Selectivity -Doyle, DA, et al. The structure of the potassium channel: molecular basis of K + conduction and selectivity. Science
The Structure of the Potassium Channel: Molecular Basis of K + Conduction and Selectivity -Doyle, DA, et al. The structure of the potassium channel: molecular basis of K + conduction and selectivity. Science
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/5/e1601684/dc1 Supplementary Materials for Directional mechanical stability of Bacteriophage ϕ29 motor s 3WJ-pRNA: Extraordinary robustness along portal axis
advances.sciencemag.org/cgi/content/full/3/5/e1601684/dc1 Supplementary Materials for Directional mechanical stability of Bacteriophage ϕ29 motor s 3WJ-pRNA: Extraordinary robustness along portal axis
SUPPLEMENTARY INFORMATION
 Supplementary Table S1 Kinetic Analyses of the AMSH-LP mutants AMSH-LP K M (μm) k cat x 10-3 (s -1 ) WT 71.8 ± 6.3 860 ± 65.4 T353A 76.8 ± 11.7 46.3 ± 3.7 F355A 58.9 ± 10.4 5.33 ± 0.30 proximal S358A 75.1
Supplementary Table S1 Kinetic Analyses of the AMSH-LP mutants AMSH-LP K M (μm) k cat x 10-3 (s -1 ) WT 71.8 ± 6.3 860 ± 65.4 T353A 76.8 ± 11.7 46.3 ± 3.7 F355A 58.9 ± 10.4 5.33 ± 0.30 proximal S358A 75.1
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
 This article was published in an Elsevier journal. The attached copy is furnished to the author for non-commercial research and education use, including for instruction at the author s institution, sharing
This article was published in an Elsevier journal. The attached copy is furnished to the author for non-commercial research and education use, including for instruction at the author s institution, sharing
SUPPLEMENTARY INFORMATION
 Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
AP Biology Unit 1, Chapters 2, 3, 4, 5
 Name Date AP Biology Unit 1, Chapters 2, 3, 4, 5 Research Question How are chemical structures visualized? Background You can represent a molecule with either a molecular formula or a structural formula.
Name Date AP Biology Unit 1, Chapters 2, 3, 4, 5 Research Question How are chemical structures visualized? Background You can represent a molecule with either a molecular formula or a structural formula.
Supplementary information
 Supplementary information doi: 10.1038/nchem.247 Amyloid!-Protein Oligomerization and the Importance of Tetramers and Dodecamers in the Aetiology of Alzheimer s Disease Summer L. Bernstein, Nicholas F.
Supplementary information doi: 10.1038/nchem.247 Amyloid!-Protein Oligomerization and the Importance of Tetramers and Dodecamers in the Aetiology of Alzheimer s Disease Summer L. Bernstein, Nicholas F.
schematic diagram; EGF binding, dimerization, phosphorylation, Grb2 binding, etc.
 Lecture 1: Noncovalent Biomolecular Interactions Bioengineering and Modeling of biological processes -e.g. tissue engineering, cancer, autoimmune disease Example: RTK signaling, e.g. EGFR Growth responses
Lecture 1: Noncovalent Biomolecular Interactions Bioengineering and Modeling of biological processes -e.g. tissue engineering, cancer, autoimmune disease Example: RTK signaling, e.g. EGFR Growth responses
Let s continue our discussion on the interaction between Fe(III) and 6,7-dihydroxynaphthalene-2- sulfonate.
 Chemistry 5995(133)-8990(013) Bioinorganic Chemistry: The Good, the Bad, and the Potential of Metals Assignment 2- Aqueous Speciation, Magnetism, Redox, UV-Vis Spectroscopy, and Pymol Let s continue our
Chemistry 5995(133)-8990(013) Bioinorganic Chemistry: The Good, the Bad, and the Potential of Metals Assignment 2- Aqueous Speciation, Magnetism, Redox, UV-Vis Spectroscopy, and Pymol Let s continue our
T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y. jgp
 S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
L718Q mutant EGFR escapes covalent inhibition by stabilizing. a non-reactive conformation of the lung cancer drug. osimertinib
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Electronic Supplementary Information (ESI) for L718Q mutant EGFR escapes covalent inhibition
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2018 Electronic Supplementary Information (ESI) for L718Q mutant EGFR escapes covalent inhibition
Supporting information for: Electron Transport. Properties of Atomic Carbon Nanowires between. Graphene Electrodes
 Supporting information for: Electron Transport Properties of Atomic Carbon Nanowires between Graphene Electrodes Lei Shen,,, Minggang Zeng,,, Shuo-Wang Yang,, Chun Zhang, Xuefeng Wang, and Yuanping Feng,
Supporting information for: Electron Transport Properties of Atomic Carbon Nanowires between Graphene Electrodes Lei Shen,,, Minggang Zeng,,, Shuo-Wang Yang,, Chun Zhang, Xuefeng Wang, and Yuanping Feng,
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08 Å resolution: comparison with the Azotobacter vinelandii MoFe protein
 Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Layered SiC Sheets: A Potential Catalyst for Oxygen Reduction Reaction. Materials Science and Engineering, Jilin University, Changchun , China,
 Supporting Information Layered SiC Sheets: A Potential Catalyst for Oxygen Reduction Reaction P. Zhang 1,2, B. B. Xiao 1, X. L. Hou 1,2, Y. F. Zhu 1,* Q. Jiang 1 1 Key Laboratory of Automobile Materials,
Supporting Information Layered SiC Sheets: A Potential Catalyst for Oxygen Reduction Reaction P. Zhang 1,2, B. B. Xiao 1, X. L. Hou 1,2, Y. F. Zhu 1,* Q. Jiang 1 1 Key Laboratory of Automobile Materials,
Why Proteins Fold. How Proteins Fold? e - ΔG/kT. Protein Folding, Nonbonding Forces, and Free Energy
 Why Proteins Fold Proteins are the action superheroes of the body. As enzymes, they make reactions go a million times faster. As versatile transport vehicles, they carry oxygen and antibodies to fight
Why Proteins Fold Proteins are the action superheroes of the body. As enzymes, they make reactions go a million times faster. As versatile transport vehicles, they carry oxygen and antibodies to fight
Water-methanol separation with carbon nanotubes and electric fields
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 215 Supplementary Information: Water-methanol separation with carbon nanotubes and electric fields
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 215 Supplementary Information: Water-methanol separation with carbon nanotubes and electric fields
SUPPLEMENTARY INFORMATION
 Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Table 1. Crystallographic data collection, phasing and refinement statistics. Native Hg soaked Mn soaked 1 Mn soaked 2
 Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Supporting Information
 Supporting Information Micelle-Triggered b-hairpin to a-helix Transition in a 14-Residue Peptide from a Choline-Binding Repeat of the Pneumococcal Autolysin LytA HØctor Zamora-Carreras, [a] Beatriz Maestro,
Supporting Information Micelle-Triggered b-hairpin to a-helix Transition in a 14-Residue Peptide from a Choline-Binding Repeat of the Pneumococcal Autolysin LytA HØctor Zamora-Carreras, [a] Beatriz Maestro,
Supplemental Information for: Characterizing the Membrane-Bound State of Cytochrome P450 3A4: Structure, Depth of Insertion and Orientation
 Supplemental Information for: Characterizing the Membrane-Bound State of Cytochrome P450 3A4: Structure, Depth of Insertion and Orientation Javier L. Baylon, Ivan L. Lenov, Stephen G. Sligar and Emad Tajkhorshid
Supplemental Information for: Characterizing the Membrane-Bound State of Cytochrome P450 3A4: Structure, Depth of Insertion and Orientation Javier L. Baylon, Ivan L. Lenov, Stephen G. Sligar and Emad Tajkhorshid
Potassium channel gating and structure!
 Reading: Potassium channel gating and structure Hille (3rd ed.) chapts 10, 13, 17 Doyle et al. The Structure of the Potassium Channel: Molecular Basis of K1 Conduction and Selectivity. Science 280:70-77
Reading: Potassium channel gating and structure Hille (3rd ed.) chapts 10, 13, 17 Doyle et al. The Structure of the Potassium Channel: Molecular Basis of K1 Conduction and Selectivity. Science 280:70-77
Bulk behaviour. Alanine. FIG. 1. Chemical structure of the RKLPDA peptide. Numbers on the left mark alpha carbons.
 Bulk behaviour To characterise the conformational behaviour of the peptide, first we looked at the statistics of alpha carbons and the torsion angles. Then they were correlated with positions of positively
Bulk behaviour To characterise the conformational behaviour of the peptide, first we looked at the statistics of alpha carbons and the torsion angles. Then they were correlated with positions of positively
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
CHAPTER 29 HW: AMINO ACIDS + PROTEINS
 CAPTER 29 W: AMI ACIDS + PRTEIS For all problems, consult the table of 20 Amino Acids provided in lecture if an amino acid structure is needed; these will be given on exams. Use natural amino acids (L)
CAPTER 29 W: AMI ACIDS + PRTEIS For all problems, consult the table of 20 Amino Acids provided in lecture if an amino acid structure is needed; these will be given on exams. Use natural amino acids (L)
Time-dependence of key H-bond/electrostatic interaction distances in the sirna5-hago2 complexes... Page S14
 Supporting Information Probing the Binding Interactions between Chemically Modified sirnas and Human Argonaute 2 Using Microsecond Molecular Dynamics Simulations S. Harikrishna* and P. I. Pradeepkumar*
Supporting Information Probing the Binding Interactions between Chemically Modified sirnas and Human Argonaute 2 Using Microsecond Molecular Dynamics Simulations S. Harikrishna* and P. I. Pradeepkumar*
Supplementary Figure 1 A schematic representation of the different reaction mechanisms
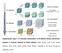 Supplementary Figure 1 A schematic representation of the different reaction mechanisms observed in electrode materials for lithium batteries. Black circles: voids in the crystal structure, blue circles:
Supplementary Figure 1 A schematic representation of the different reaction mechanisms observed in electrode materials for lithium batteries. Black circles: voids in the crystal structure, blue circles:
Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6
 Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Housekeeping. Housekeeping. Molecules of Life: Biopolymers
 Molecules of Life: Biopolymers Dr. Dale Hancock D.Hancock@mmb.usyd.edu.au Room 377 Biochemistry building Housekeeping Answers to the practise calculations and a narration are on WebT. Access these through
Molecules of Life: Biopolymers Dr. Dale Hancock D.Hancock@mmb.usyd.edu.au Room 377 Biochemistry building Housekeeping Answers to the practise calculations and a narration are on WebT. Access these through
TiC 2 : A New Two Dimensional Sheet beyond MXenes
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Supplementary Information (SI) TiC 2 : A New Two Dimensional Sheet beyond MXenes Tianshan Zhao,
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Supplementary Information (SI) TiC 2 : A New Two Dimensional Sheet beyond MXenes Tianshan Zhao,
BIOC : Homework 1 Due 10/10
 Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
Molecular Dynamics, Monte Carlo and Docking. Lecture 21. Introduction to Bioinformatics MNW2
 Molecular Dynamics, Monte Carlo and Docking Lecture 21 Introduction to Bioinformatics MNW2 Allowed phi-psi angles Red areas are preferred, yellow areas are allowed, and white is avoided 2.3a Hamiltonian
Molecular Dynamics, Monte Carlo and Docking Lecture 21 Introduction to Bioinformatics MNW2 Allowed phi-psi angles Red areas are preferred, yellow areas are allowed, and white is avoided 2.3a Hamiltonian
Rotamers in the CHARMM19 Force Field
 Appendix A Rotamers in the CHARMM19 Force Field The people may be made to follow a path of action, but they may not be made to understand it. Confucius (551 BC - 479 BC) ( ) V r 1 (j),r 2 (j),r 3 (j),...,r
Appendix A Rotamers in the CHARMM19 Force Field The people may be made to follow a path of action, but they may not be made to understand it. Confucius (551 BC - 479 BC) ( ) V r 1 (j),r 2 (j),r 3 (j),...,r
Universal Repulsive Contribution to the. Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information
 Universal Repulsive Contribution to the Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information B. Shadrack Jabes, Dusan Bratko, and Alenka Luzar Department of Chemistry,
Universal Repulsive Contribution to the Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information B. Shadrack Jabes, Dusan Bratko, and Alenka Luzar Department of Chemistry,
Exploratory Examination of Photosystem I in Arabidopsis thaliana
 Exploratory Examination of Photosystem I in Arabidopsis thaliana A. Role of Photosystem I Photosystem I (PSI) captures the sunlight and transfers the energy through a pigment network to the center of the
Exploratory Examination of Photosystem I in Arabidopsis thaliana A. Role of Photosystem I Photosystem I (PSI) captures the sunlight and transfers the energy through a pigment network to the center of the
Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets
 Supporting information Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets Wan-Na Chen, Christoph Nitsche, Kala Bharath Pilla, Bim Graham, Thomas
Supporting information Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets Wan-Na Chen, Christoph Nitsche, Kala Bharath Pilla, Bim Graham, Thomas
Section Week 3. Junaid Malek, M.D.
 Section Week 3 Junaid Malek, M.D. Biological Polymers DA 4 monomers (building blocks), limited structure (double-helix) RA 4 monomers, greater flexibility, multiple structures Proteins 20 Amino Acids,
Section Week 3 Junaid Malek, M.D. Biological Polymers DA 4 monomers (building blocks), limited structure (double-helix) RA 4 monomers, greater flexibility, multiple structures Proteins 20 Amino Acids,
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 218 Supporting Information Multi-Functional Organosilane-Polymerized Carbon Dots Inverse Opals Junchao
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 218 Supporting Information Multi-Functional Organosilane-Polymerized Carbon Dots Inverse Opals Junchao
Protein Structure Bioinformatics Introduction
 1 Swiss Institute of Bioinformatics Protein Structure Bioinformatics Introduction Basel, 27. September 2004 Torsten Schwede Biozentrum - Universität Basel Swiss Institute of Bioinformatics Klingelbergstr
1 Swiss Institute of Bioinformatics Protein Structure Bioinformatics Introduction Basel, 27. September 2004 Torsten Schwede Biozentrum - Universität Basel Swiss Institute of Bioinformatics Klingelbergstr
Practice Midterm Exam 200 points total 75 minutes Multiple Choice (3 pts each 30 pts total) Mark your answers in the space to the left:
 MITES ame Practice Midterm Exam 200 points total 75 minutes Multiple hoice (3 pts each 30 pts total) Mark your answers in the space to the left: 1. Amphipathic molecules have regions that are: a) polar
MITES ame Practice Midterm Exam 200 points total 75 minutes Multiple hoice (3 pts each 30 pts total) Mark your answers in the space to the left: 1. Amphipathic molecules have regions that are: a) polar
Supporting Information
 Supporting Information Allosteric communication disrupted by small molecule binding to the Imidazole glycerol phosphate synthase protein-protein interface. Ivan Rivalta*,#, George P. Lisi #, Ning-Shiuan
Supporting Information Allosteric communication disrupted by small molecule binding to the Imidazole glycerol phosphate synthase protein-protein interface. Ivan Rivalta*,#, George P. Lisi #, Ning-Shiuan
Supporting Information
 Supporting Information Interface-Induced Affinity Sieving in Nanoporous Graphenes for Liquid-Phase Mixtures Yanan Hou, Zhijun Xu, Xiaoning Yang * State Key Laboratory of Material-Orientated Chemical Engineering,
Supporting Information Interface-Induced Affinity Sieving in Nanoporous Graphenes for Liquid-Phase Mixtures Yanan Hou, Zhijun Xu, Xiaoning Yang * State Key Laboratory of Material-Orientated Chemical Engineering,
Supplementary Figure 1 a) Scheme of microfluidic device fabrication by photo and soft lithography,
 a b 1 mm Supplementary Figure 1 a) Scheme of microfluidic device fabrication by photo and soft lithography, (a1, a2) 50nm Pd evaporated on Si wafer with 100 nm Si 2 insulating layer and 5nm Cr as an adhesion
a b 1 mm Supplementary Figure 1 a) Scheme of microfluidic device fabrication by photo and soft lithography, (a1, a2) 50nm Pd evaporated on Si wafer with 100 nm Si 2 insulating layer and 5nm Cr as an adhesion
Protein Structure Basics
 Protein Structure Basics Presented by Alison Fraser, Christine Lee, Pradhuman Jhala, Corban Rivera Importance of Proteins Muscle structure depends on protein-protein interactions Transport across membranes
Protein Structure Basics Presented by Alison Fraser, Christine Lee, Pradhuman Jhala, Corban Rivera Importance of Proteins Muscle structure depends on protein-protein interactions Transport across membranes
MARTINI simulation details
 S1 Appendix MARTINI simulation details MARTINI simulation initialization and equilibration In this section, we describe the initialization of simulations from Main Text section Residue-based coarsegrained
S1 Appendix MARTINI simulation details MARTINI simulation initialization and equilibration In this section, we describe the initialization of simulations from Main Text section Residue-based coarsegrained
1. What is an ångstrom unit, and why is it used to describe molecular structures?
 1. What is an ångstrom unit, and why is it used to describe molecular structures? The ångstrom unit is a unit of distance suitable for measuring atomic scale objects. 1 ångstrom (Å) = 1 10-10 m. The diameter
1. What is an ångstrom unit, and why is it used to describe molecular structures? The ångstrom unit is a unit of distance suitable for measuring atomic scale objects. 1 ångstrom (Å) = 1 10-10 m. The diameter
Protein Bioinformatics Computer lab #1 Friday, April 11, 2008 Sean Prigge and Ingo Ruczinski
 Protein Bioinformatics 260.655 Computer lab #1 Friday, April 11, 2008 Sean Prigge and Ingo Ruczinski Goals: Approx. Time [1] Use the Protein Data Bank PDB website. 10 minutes [2] Use the WebMol Viewer.
Protein Bioinformatics 260.655 Computer lab #1 Friday, April 11, 2008 Sean Prigge and Ingo Ruczinski Goals: Approx. Time [1] Use the Protein Data Bank PDB website. 10 minutes [2] Use the WebMol Viewer.
Exploring the Changes in the Structure of α-helical Peptides Adsorbed onto Carbon and Boron Nitride based Nanomaterials
 Exploring the Changes in the Structure of α-helical Peptides Adsorbed onto Carbon and Boron Nitride based Nanomaterials Dr. V. Subramanian Chemical Laboratory, IPC Division CSIR-Central Leather Research
Exploring the Changes in the Structure of α-helical Peptides Adsorbed onto Carbon and Boron Nitride based Nanomaterials Dr. V. Subramanian Chemical Laboratory, IPC Division CSIR-Central Leather Research
Nature Structural & Molecular Biology doi: /nsmb Supplementary Figure 1. CRBN binding assay with thalidomide enantiomers.
 Supplementary Figure 1 CRBN binding assay with thalidomide enantiomers. (a) Competitive elution assay using thalidomide-immobilized beads coupled with racemic thalidomide. Beads were washed three times
Supplementary Figure 1 CRBN binding assay with thalidomide enantiomers. (a) Competitive elution assay using thalidomide-immobilized beads coupled with racemic thalidomide. Beads were washed three times
Cks1 CDK1 CDK1 CDK1 CKS1. are ice- lobe. conserved. conserved
 Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. The asymmetric unit in para-iodio-phenylalanine crystal. The 50% probability ellipsoid representation was prepared using the Mercury Software. Colors are as
Supplementary Figures Supplementary Figure 1. The asymmetric unit in para-iodio-phenylalanine crystal. The 50% probability ellipsoid representation was prepared using the Mercury Software. Colors are as
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
Essential Forces in Protein Folding
 Essential Forces in Protein Folding Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Amino Acid sequence
Essential Forces in Protein Folding Dr. Mohammad Alsenaidy Department of Pharmaceutics College of Pharmacy King Saud University Office: AA 101 msenaidy@ksu.edu.sa Previously on PHT 426!! Amino Acid sequence
Design of Efficient Catalysts with Double Transition Metal. Atoms on C 2 N Layer
 Supporting Information Design of Efficient Catalysts with Double Transition Metal Atoms on C 2 N Layer Xiyu Li, 1, Wenhui Zhong, 2, Peng Cui, 1 Jun Li, 1 Jun Jiang 1, * 1 Hefei National Laboratory for
Supporting Information Design of Efficient Catalysts with Double Transition Metal Atoms on C 2 N Layer Xiyu Li, 1, Wenhui Zhong, 2, Peng Cui, 1 Jun Li, 1 Jun Jiang 1, * 1 Hefei National Laboratory for
HOMOLOGY MODELING. The sequence alignment and template structure are then used to produce a structural model of the target.
 HOMOLOGY MODELING Homology modeling, also known as comparative modeling of protein refers to constructing an atomic-resolution model of the "target" protein from its amino acid sequence and an experimental
HOMOLOGY MODELING Homology modeling, also known as comparative modeling of protein refers to constructing an atomic-resolution model of the "target" protein from its amino acid sequence and an experimental
Supplementary information
 1 2 Supplementary information 3 4 5 6 Supplementary Figure 1 7 8 Supplementary Figure 1 ǀ Characterization of the lysozyme fibrils by atomic force microscopy 9 (AFM) and scanning electron microscopy (SEM).
1 2 Supplementary information 3 4 5 6 Supplementary Figure 1 7 8 Supplementary Figure 1 ǀ Characterization of the lysozyme fibrils by atomic force microscopy 9 (AFM) and scanning electron microscopy (SEM).
Supplementary Information for: Controlling Cellular Uptake of Nanoparticles with ph-sensitive Polymers
 Supplementary Information for: Controlling Cellular Uptake of Nanoparticles with ph-sensitive Polymers Hong-ming Ding 1 & Yu-qiang Ma 1,2, 1 National Laboratory of Solid State Microstructures and Department
Supplementary Information for: Controlling Cellular Uptake of Nanoparticles with ph-sensitive Polymers Hong-ming Ding 1 & Yu-qiang Ma 1,2, 1 National Laboratory of Solid State Microstructures and Department
Supporting Information Available:
 Supporting Information Available: Photoresponsive and Gas Sensing Field-Effect Transistors based on Multilayer WS 2 Nanoflakes Nengjie Huo 1, Shengxue Yang 1, Zhongming Wei 2, Shu-Shen Li 1, Jian-Bai Xia
Supporting Information Available: Photoresponsive and Gas Sensing Field-Effect Transistors based on Multilayer WS 2 Nanoflakes Nengjie Huo 1, Shengxue Yang 1, Zhongming Wei 2, Shu-Shen Li 1, Jian-Bai Xia
DISCRETE TUTORIAL. Agustí Emperador. Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING:
 DISCRETE TUTORIAL Agustí Emperador Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING: STRUCTURAL REFINEMENT OF DOCKING CONFORMATIONS Emperador
DISCRETE TUTORIAL Agustí Emperador Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING: STRUCTURAL REFINEMENT OF DOCKING CONFORMATIONS Emperador
Weyl semimetal phase in the non-centrosymmetric compound TaAs
 Weyl semimetal phase in the non-centrosymmetric compound TaAs L. X. Yang 1,2,3, Z. K. Liu 4,5, Y. Sun 6, H. Peng 2, H. F. Yang 2,7, T. Zhang 1,2, B. Zhou 2,3, Y. Zhang 3, Y. F. Guo 2, M. Rahn 2, P. Dharmalingam
Weyl semimetal phase in the non-centrosymmetric compound TaAs L. X. Yang 1,2,3, Z. K. Liu 4,5, Y. Sun 6, H. Peng 2, H. F. Yang 2,7, T. Zhang 1,2, B. Zhou 2,3, Y. Zhang 3, Y. F. Guo 2, M. Rahn 2, P. Dharmalingam
Journal of Pharmacology and Experimental Therapy-JPET#172536
 A NEW NON-PEPTIDIC INHIBITOR OF THE 14-3-3 DOCKING SITE INDUCES APOPTOTIC CELL DEATH IN CHRONIC MYELOID LEUKEMIA SENSITIVE OR RESISTANT TO IMATINIB Manuela Mancini, Valentina Corradi, Sara Petta, Enza
A NEW NON-PEPTIDIC INHIBITOR OF THE 14-3-3 DOCKING SITE INDUCES APOPTOTIC CELL DEATH IN CHRONIC MYELOID LEUKEMIA SENSITIVE OR RESISTANT TO IMATINIB Manuela Mancini, Valentina Corradi, Sara Petta, Enza
Xiang-Kui Gu,, Botao Qiao,,, Chuan-Qi Huang, Wu-Chen Ding, Keju Sun, Ensheng Zhan,, Tao Zhang, Jingyue Liu*,,, and Wei-Xue Li*,
 Supported Single Pt 1 /Au 1 Atoms for Methanol Steam Reforming Xiang-Kui Gu,, Botao Qiao,,, Chuan-Qi Huang, Wu-Chen Ding, Keju Sun, Ensheng Zhan,, Tao Zhang, Jingyue Liu*,,, and Wei-Xue Li*, State Key
Supported Single Pt 1 /Au 1 Atoms for Methanol Steam Reforming Xiang-Kui Gu,, Botao Qiao,,, Chuan-Qi Huang, Wu-Chen Ding, Keju Sun, Ensheng Zhan,, Tao Zhang, Jingyue Liu*,,, and Wei-Xue Li*, State Key
Supporting information for: Mechanism of lignin inhibition of enzymatic. biomass deconstruction
 Supporting information for: Mechanism of lignin inhibition of enzymatic biomass deconstruction Josh V. Vermaas,, Loukas Petridis, Xianghong Qi,, Roland Schulz,, Benjamin Lindner, and Jeremy C. Smith,,
Supporting information for: Mechanism of lignin inhibition of enzymatic biomass deconstruction Josh V. Vermaas,, Loukas Petridis, Xianghong Qi,, Roland Schulz,, Benjamin Lindner, and Jeremy C. Smith,,
Molecular dynamics simulations of anti-aggregation effect of ibuprofen. Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov
 Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Structuring of hydrophobic and hydrophilic polymers at interfaces Stephen Donaldson ChE 210D Final Project Abstract
 Structuring of hydrophobic and hydrophilic polymers at interfaces Stephen Donaldson ChE 210D Final Project Abstract In this work, a simplified Lennard-Jones (LJ) sphere model is used to simulate the aggregation,
Structuring of hydrophobic and hydrophilic polymers at interfaces Stephen Donaldson ChE 210D Final Project Abstract In this work, a simplified Lennard-Jones (LJ) sphere model is used to simulate the aggregation,
