Around the dynamics of bacterial genomes
|
|
|
- Pearl Jones
- 6 years ago
- Views:
Transcription
1 Around the dynamics of bacterial genomes Eduardo P. C. Rocha Unité Génétique des Génomes Bactériens, CNRS URA2171,Institut Pasteur Atelier de BioInformatique, Université Pierre et Marie Curie
2 L ADN
3
4
5 ORFs, CDS, etc. ORF : Open Reading Frame CDS : CoDing Sequence 5' -35 signal -10 signal RBS 3' ORF CDS ORF Start : Pattern (ATG,GTG,TTG) RBS : Pattern (AGGAGGT) Début de la transcription Stop : Pattern (TAA,TGA,TAG) Début de la traduction Fin de la traduction Fin de la transcription rho indépendante -35 signal : Pattern (TTGACA) -10 signal : Pattern (TATAAT) } Consensus "classique" : TTGACA (15 à 19 bp) TATAAT Terminateur de la transcription -> structure en tige-boucle suivie d'un poly T + gènes de trna et gènes de rrna
6
7 ATCTTTTTCGGCTTTTTTTAGTATCCACAGAGGTTATCGACAACATTTTCACATTACCAACCCCTGTGGACAAGGTTTTTTCAACAGGTT GTCCGCTTTGTGGATAAGATTGTGACAACCATTGCAAGCTCTCGTTTATTTTGGTATTATATTTGTGTTTTAACTCTTGATTACTAATCC TACCTTTCCTCTTTATCCACAAAGTGTGGATAAGTTGTGGATTGATTTCACACAGCTTGTGTAGAAGGTTGTCCACAAGTTGTGAAATTT GTCGAAAAGCTATTTATCTACTATATTATATGTTTTCAACATTTAATGTGTACGAATGGTAAGCGCCATTTGCTCTTTTTTTGTGTTCTA TAACAGAGAAAGACGCCATTTTCTAAGAAAAGGAGGGACGTGCCGGAAGATGGAAAATATATTAGACCTGTGGAACCAAGCCCTTGCTCA AATCGAAAAAAAGTTGAGCAAACCGAGTTTTGAGACTTGGATGAAGTCAACCAAAGCCCACTCACTGCAAGGCGATACATTAACAATCAC GGCTCCCAATGAATTTGCCAGAGACTGGCTGGAGTCCAGATACTTGCATCTGATTGCAGATACTATATATGAATTAACCGGGGAAGAATT GAGCATTAAGTTTGTCATTCCTCAAAATCAAGATGTTGAGGACTTTATGCCGAAACCGCAAGTCAAAAAAGCGGTCAAAGAAGATACATC TGATTTTCCTCAAAATATGCTCAATCCAAAATATACTTTTGATACTTTTGTCATCGGATCTGGAAACCGATTTGCACATGCTGCTTCCCT CGCAGTAGCGGAAGCGCCCGCGAAAGCTTACAACCCTTTATTTATCTATGGGGGCGTCGGCTTAGGGAAAACACACTTAATGCATGCGAT CGGCCATTATGTAATAGATCATAATCCTTCTGCCAAAGTGGTTTATCTGTCTTCTGAGAAATTTACAAACGAATTCATCAACTCTATCCG AGATAATAAAGCCGTCGACTTCCGCAATCGCTATCGAAATGTTGATGTGCTTTTGATAGATGATATTCAATTTTTAGCGGGGAAAGAACA AACCCAGGAAGAATTTTTCCATACATTTAACACATTACACGAAGAAAGCAAACAAATCGTCATTTCAAGTGACCGGCCGCCAAAGGAAAT TCCGACACTTGAAGACAGATTGCGCTCACGTTTTGAATGGGGACTTATTACAGATATCACACCGCCTGATCTAGAAACGAGAATTGCAAT TTTAAGAAAAAAGGCCAAAGCAGAGGGCCTCGATATTCCGAACGAGGTTATGCTTTACATCGCGAATCAAATCGACAGCAATATTCGGGA ACTCGAAGGAGCATTAATCAGAGTTGTCGCTTATTCATCTTTAATTAATAAAGATATTAATGCTGATCTGGCCGCTGAGGCGTTGAAAGA TATTATTCCTTCCTCAAAACCGAAAGTCATTACGATAAAAGAAATTCAGAGGGTAGTAGGCCAGCAATTTAATATTAAACTCGAGGATTT CAAAGCAAAAAAACGGACAAAGTCAGTAGCTTTTCCGCGTCAAATCGCCATGTACTTATCAAGGGAAATGACTGATTCCTCTCTTCCTAA AATCGGTGAAGAGTTTGGAGGACGTGATCATACGACCGTTATTCATGCGCATGAAAAAATTTCAAAACTGCTGGCAGATGATGAACAGCT TCAGCAGCATGTAAAAGAAATTAAAGAACAGCTTAAATAGCAGGACCGGGGATCAATCGGGGAAAGTGTGAATAACTTTTCGGAAGTCAT ACACAGTCTGTCCACATGTGGATAGGCTGTGTTTCCTGTCTTTTTCACAACTTATCCACAAATCCACAGGCCCTACTATTACTTCTACTA TTTTTTATAAATATATATATTAATACATTATCCGTTAGGAGGATAAAAATGAAATTCACGATTCAAAAAGATCGTCTTGTTGAAAGTGTC CAAGATGTATTAAAAGCAGTTTCATCCAGAACCACGATTCCCATTCTGACTGGTATTAAAATTGTTGCATCAGATGATGGAGTATCCTTT ACAGGGAGTGACTCAGATATTTCTATTGAATCCTTCATTCCAAAAGAAGAAGGAGATAAAGAAATCGTCACTATTGAACAGCCCGGAAGC ATCGTTTTACAGGCTCGCTTTTTTAGTGAAATTGTAAAAAAATTGCCGATGGCAACTGTA
8 Des données à l information ORIGIN - origine de réplication; TCGA - restriction (défense?); TGTGGATAA - promoteur (transcription); souligné - DnaA Box (réplication); ATG/TAA/RBS - codon start/stop /RBS (traduction); séquence codante (traduction). ATCTTTTTCGGCTTTTTTTAGTATCCACAGAGGTTATCGACAACATTTTCACATTACCAACCCCTGTGGACAAGGTTTTTTCAACAGGTT GTCCGCTTTGTGGATAAGATTGTGACAACCATTGCAAGCTCTCGTTTATTTTGGTATTATATTTGTGTTTTAACTCTTGATTACTAATCC TACCTTTCCTCTTTATCCACAAAGTGTGGATAAGTTGTGGATTGATTTCACACAGCTTGTGTAGAAGGTTGTCCACAAGTTGTGAAATTT GTCGAAAAGCTATTTATCTACTATATTATATGTTTTCAACATTTAATGTGTACGAATGGTAAGCGCCATTTGCTCTTTTTTTGTGTTCTA TAACAGAGAAAGACGCCATTTTCTAAGAAAAGGAGGGACGTGCCGGAAGATGGAAAATATATTAGACCTGTGGAACCAAGCCCTTGCTCA AATCGAAAAAAAGTTGAGCAAACCGAGTTTTGAGACTTGGATGAAGTCAACCAAAGCCCACTCACTGCAAGGCGATACATTAACAATCAC GGCTCCCAATGAATTTGCCAGAGACTGGCTGGAGTCCAGATACTTGCATCTGATTGCAGATACTATATATGAATTAACCGGGGAAGAATT GAGCATTAAGTTTGTCATTCCTCAAAATCAAGATGTTGAGGACTTTATGCCGAAACCGCAAGTCAAAAAAGCGGTCAAAGAAGATACATC TGATTTTCCTCAAAATATGCTCAATCCAAAATATACTTTTGATACTTTTGTCATCGGATCTGGAAACCGATTTGCACATGCTGCTTCCCT CGCAGTAGCGGAAGCGCCCGCGAAAGCTTACAACCCTTTATTTATCTATGGGGGCGTCGGCTTAGGGAAAACACACTTAATGCATGCGAT CGGCCATTATGTAATAGATCATAATCCTTCTGCCAAAGTGGTTTATCTGTCTTCTGAGAAATTTACAAACGAATTCATCAACTCTATCCG AGATAATAAAGCCGTCGACTTCCGCAATCGCTATCGAAATGTTGATGTGCTTTTGATAGATGATATTCAATTTTTAGCGGGGAAAGAACA AACCCAGGAAGAATTTTTCCATACATTTAACACATTACACGAAGAAAGCAAACAAATCGTCATTTCAAGTGACCGGCCGCCAAAGGAAAT TCCGACACTTGAAGACAGATTGCGCTCACGTTTTGAATGGGGACTTATTACAGATATCACACCGCCTGATCTAGAAACGAGAATTGCAAT TTTAAGAAAAAAGGCCAAAGCAGAGGGCCTCGATATTCCGAACGAGGTTATGCTTTACATCGCGAATCAAATCGACAGCAATATTCGGGA ACTCGAAGGAGCATTAATCAGAGTTGTCGCTTATTCATCTTTAATTAATAAAGATATTAATGCTGATCTGGCCGCTGAGGCGTTGAAAGA TATTATTCCTTCCTCAAAACCGAAAGTCATTACGATAAAAGAAATTCAGAGGGTAGTAGGCCAGCAATTTAATATTAAACTCGAGGATTT CAAAGCAAAAAAACGGACAAAGTCAGTAGCTTTTCCGCGTCAAATCGCCATGTACTTATCAAGGGAAATGACTGATTCCTCTCTTCCTAA AATCGGTGAAGAGTTTGGAGGACGTGATCATACGACCGTTATTCATGCGCATGAAAAAATTTCAAAACTGCTGGCAGATGATGAACAGCT TCAGCAGCATGTAAAAGAAATTAAAGAACAGCTTAAATAGCAGGACCGGGGATCAATCGGGGAAAGTGTGAATAACTTTTCGGAAGTCAT ACACAGTCTGTCCACATGTGGATAGGCTGTGTTTCCTGTCTTTTTCACAACTTATCCACAAATCCACAGGCCCTACTATTACTTCTACTA TTTTTTATAAATATATATATTAATACATTATCCGTTAGGAGGATAAAAATGAAATTCACGATTCAAAAAGATCGTCTTGTTGAAAGTGTC CAAGATGTATTAAAAGCAGTTTCATCCAGAACCACGATTCCCATTCTGACTGGTATTAAAATTGTTGCATCAGATGATGGAGTATCCTTT ACAGGGAGTGACTCAGATATTTCTATTGAATCCTTCATTCCAAAAGAAGAAGGAGATAAAGAAATCGTCACTATTGAACAGCCCGGAAGC ATCGTTTTACAGGCTCGCTTTTTTAGTGAAATTGTAAAAAAATTGCCGATGGCAACTGTA
9 (Himmelreich et al, NAR, 96)
10 Mutation & Recombination
11 Fading homologies (Clarck, TREE, 06)
12 Bioinformatics beyond model organisms ripr chtr pyho aqae trpa mytu arfu bobu meth basu hepy esco mypn sysp meja myge hain ,190 14,357 11,082 Nombre de papiers citant chaque espèce 167, Nombre de gènes classés fonctionnellement
13
14 Production d hypothèses Données Théorie Tests d hypothèses
15 Summary i- Genome stability ii- Genome organization iii -Conflicts
16 Gene order conservation Data Model A α i β j B GOC is derived from the order of pairs of orthologues : α i α i+1 β j β j+k GOC i =1 iff k =1, GOC = i genes GOC i N (method adapted from Huynen, PNAS, 98) If all pairs are conserved with equal probability (p) per unit of time : GOC α p t If there are 2 classes of conservation : GOC = p fast t + p slow t
17 Gene order conservation (GOC) GOC = p t GOC= p fastt +p slow t A B Δ<k p - probability that two contiguous genes have contiguous orthologs p fast and p fast allow for pairs of genes with different rearrangement rates. Data: 126 genomes of 6 clades with more than 8 genomes, 16S rdna distances (HKY+Γ) (Rocha, Mol Biol Evol, 06)
18 Genome backbones are stable ~100 million years ~500 million years : ~50% of contiguity is conserved (Rocha, Mol Biol Evol, 06)
19 What is the E. coli genome? Core genome evolution 1 2 Pan genome evolution ~18000 number of genes number of genes ~ ~1800 ~4600 number of genomes number of genomes
20 Genome backbones are stable ~100 million years ~500 million years : ~50% of contiguity is conserved (Fischer, PlosG, 06)
21 Different constraints GOC = p fastt +p slow t pairs slowly separating pairs quickly separating
22 Détection d opérons
23 Détection d opérons 1. Co-orientation : un opéron est une séquence de gènes coorientés contigus.
24 Détection d opérons 1. Co-orientation : un opéron est une séquence de gènes coorientés contigus. 2. Dans la plupart des génomes la majorité des opérons a un signal de terminaison de transcription rho-indépendant.
25 Détection d opérons 1. Co-orientation : un opéron est une séquence de gènes coorientés contigus. 2. Dans la plupart des génomes la majorité des opérons a un signal de terminaison de transcription rho-indépendant. 3. Si la distance entre deux gènes est très grande (>~150 nt) ils ont une forte probabilité d être dans des opérons diférents
26 Détection d opérons 1. Co-orientation : un opéron est une séquence de gènes co-orientés contigus. 2. Dans la plupart des génomes la majorité des opérons a un signal de terminaison de transcription rho-indépendant. 3. Si la distance entre deux gènes est très grande (>~150 nt) ils ont une forte probabilité d être dans des opérons différents. 4. Naturellement d autres données peuvent améliorer les prédictions
27 Operons organize genomes locally observed fit vs expected fit operons inter-operons Genes contiguous in the same operon Genes contiguous in different operons (Rocha, Mol Biol Evol, 06)
28 Defining genome stability + stable Obs>Expected - stable Obs<Expected Genome stability can be defined as the average of the residuals for the comparisons in which the genome participates.
29 Repeats decrease stability Stability density of repeats (Rocha, Trends Genetics, 03)
30 Why repeats? Creation mechanisms : Molecular parasites (e.g. transposases) Intra-chromosomal recombination Horizontal transfer Fixation in populations: For dosage effects (e.g. ARNr) To generate variability by recombination (e.g. antigenic variation in pathogens) In the way for sub- or neo-functionalization
31 Summary i- Genome stability Bacterial genomes are stable Some genomes are more stable than others depending on mutational events At a local level, transcription constrains genome rearrangements. ii- Genome organization iii -Conflicts
32 Replication-associated gene dosage Time replication round Ori R = replication time/doubling time # replication forks = 2x(2 R -1) # origins / # termini = 2 R R can be up to 4! -> 30 replication forks and 16 ori/ter. Θ Ter (Couturier, Mol Microbiol, 06)
33 Replication-associated gene dosage fast growing slow growing Ori Θ Highly expressed genes group near the origin of replication in fast growing bacteria The regulatory cascade is imprinted in the organization of the genome: RNAP>RDNA>RP Ter (Couturier, Mol Microbiol, 06)
34 In short
35 Not all rearrangements are alike ter A ter B before before A C before B no change ter before C after ter Op- operons Gi- genomic islands Sb- strand bias Gd- gene dosage Rs- segregation Cs- symmetry after after
36 . Keeping order at a global scale Buchnera Bp Chlamydia muridarum 16S:.076 nt-1 Buchnera Ap Chlamydia pneumoniae 16S:.071 nt -1 Escherichia coli origin terminus Bacillus subtilis 16S:.063 nt -1 Yersinia pestis Bacillus halodurans 16S:.051 nt -1
37 Summary i- Genome stability Bacterial genomes are stable Some genomes are more stable than others depending on mutational events At a local level, transcription constrains genome rearrangements. ii- Genome organization Genomes are organized at local and global scales Replication structures the chromosome at a large-scale The organization depends on cell machinery and cell structure iii -Conflicts
38 Taming disorder I regionalization Unstable Streptomyces coelicolor Stable (Bentley, Nature, 02)
39 (Himmelreich et al, NAR, 96)
40 Rearrangements in Mycoplasma genome M. genitalium M. pneumoniae D/I M. pneumoniae M. genitalium (Rocha, NAR, 02)
41 Taming disorder III delocalizing repeats Borrelia burgdorferi Plasmids (600 kb) : 1256 repeats Chromosome (900 kb) : 21 repeats (Fraser, Nature, 97)
42 Genome organization Genome organization is driven by the cellular processes interacting with the chromosome. But is constantly challenged by the fluidity of the chromosome. Thus, only organizational elements under strong selection endure. The genome allows inferring cellular mechanisms and phenotypic characteristics underlying its organisation. (Rocha, Curr Op Microb, 04)
43 All things have varying capacities. Consequently he who wants to have right without wrong, order without disorder, does not understand [ ] how things hang together. Chuang-Tzu (300 BC) «Forgetting about preferences»
Essentiality in B. subtilis
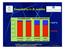 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Fitness constraints on horizontal gene transfer
 Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
Bio 119 Bacterial Genomics 6/26/10
 BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
Vital Statistics Derived from Complete Genome Sequencing (for E. coli MG1655)
 We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
2 Genome evolution: gene fusion versus gene fission
 2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
Topology. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 Topology 1 Introduction 2 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 G+C content 7 Codon usage 27 marc.bailly-bechet@univ-lyon1.fr The big picture Eukaryota Bacteria Many linear chromosomes
Topology 1 Introduction 2 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 G+C content 7 Codon usage 27 marc.bailly-bechet@univ-lyon1.fr The big picture Eukaryota Bacteria Many linear chromosomes
Flow of Genetic Information
 presents Flow of Genetic Information A Montagud E Navarro P Fernández de Córdoba JF Urchueguía Elements Nucleic acid DNA RNA building block structure & organization genome building block types Amino acid
presents Flow of Genetic Information A Montagud E Navarro P Fernández de Córdoba JF Urchueguía Elements Nucleic acid DNA RNA building block structure & organization genome building block types Amino acid
the noisy gene Biology of the Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB)
 Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Chromosomal rearrangements in mammalian genomes : characterising the breakpoints. Claire Lemaitre
 PhD defense Chromosomal rearrangements in mammalian genomes : characterising the breakpoints Claire Lemaitre Laboratoire de Biométrie et Biologie Évolutive Université Claude Bernard Lyon 1 6 novembre 2008
PhD defense Chromosomal rearrangements in mammalian genomes : characterising the breakpoints Claire Lemaitre Laboratoire de Biométrie et Biologie Évolutive Université Claude Bernard Lyon 1 6 novembre 2008
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
THE advances of the last decade on genome sequenc- very different compositional bias, as well as different gene
 Copyright 2003 by the Genetics Society of America Associations Between Inverted Repeats and the Structural Evolution of Bacterial Genomes Guillaume Achaz,*,1 Eric Coissac,*,2 Pierre Netter* and Eduardo
Copyright 2003 by the Genetics Society of America Associations Between Inverted Repeats and the Structural Evolution of Bacterial Genomes Guillaume Achaz,*,1 Eric Coissac,*,2 Pierre Netter* and Eduardo
# shared OGs (spa, spb) Size of the smallest genome. dist (spa, spb) = 1. Neighbor joining. OG1 OG2 OG3 OG4 sp sp sp
 Bioinformatics and Evolutionary Genomics: Genome Evolution in terms of Gene Content 3/10/2014 1 Gene Content Evolution What about HGT / genome sizes? Genome trees based on gene content: shared genes Haemophilus
Bioinformatics and Evolutionary Genomics: Genome Evolution in terms of Gene Content 3/10/2014 1 Gene Content Evolution What about HGT / genome sizes? Genome trees based on gene content: shared genes Haemophilus
The Minimal-Gene-Set -Kapil PHY498BIO, HW 3
 The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome
 Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
TE content correlates positively with genome size
 TE content correlates positively with genome size Mb 3000 Genomic DNA 2500 2000 1500 1000 TE DNA Protein-coding DNA 500 0 Feschotte & Pritham 2006 Transposable elements. Variation in gene numbers cannot
TE content correlates positively with genome size Mb 3000 Genomic DNA 2500 2000 1500 1000 TE DNA Protein-coding DNA 500 0 Feschotte & Pritham 2006 Transposable elements. Variation in gene numbers cannot
Biology 105/Summer Bacterial Genetics 8/12/ Bacterial Genomes p Gene Transfer Mechanisms in Bacteria p.
 READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
Inferring positional homologs with common intervals of sequences
 Outline Introduction Our approach Results Conclusion Inferring positional homologs with common intervals of sequences Guillaume Blin, Annie Chateau, Cedric Chauve, Yannick Gingras CGL - Université du Québec
Outline Introduction Our approach Results Conclusion Inferring positional homologs with common intervals of sequences Guillaume Blin, Annie Chateau, Cedric Chauve, Yannick Gingras CGL - Université du Québec
Supplementary Information
 Supplementary Information 1 List of Figures 1 Models of circular chromosomes. 2 Distribution of distances between core genes in Escherichia coli K12, arc based model. 3 Distribution of distances between
Supplementary Information 1 List of Figures 1 Models of circular chromosomes. 2 Distribution of distances between core genes in Escherichia coli K12, arc based model. 3 Distribution of distances between
AP Bio Module 16: Bacterial Genetics and Operons, Student Learning Guide
 Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Bacterial Genetics & Operons
 Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
The replication-related organization of bacterial genomes
 Microbiology (2004), 150, 1609 1627 DOI 10.1099/mic.0.26974-0 Review The replication-related organization of bacterial genomes Eduardo P. C. Rocha Correspondence erocha@abi.snv.jussieu.fr Atelier de Bioinformatique,
Microbiology (2004), 150, 1609 1627 DOI 10.1099/mic.0.26974-0 Review The replication-related organization of bacterial genomes Eduardo P. C. Rocha Correspondence erocha@abi.snv.jussieu.fr Atelier de Bioinformatique,
Base Composition Skews, Replication Orientation, and Gene Orientation in 12 Prokaryote Genomes
 J Mol Evol (1998) 47:691 696 Springer-Verlag New York Inc. 1998 Base Composition Skews, Replication Orientation, and Gene Orientation in 12 Prokaryote Genomes Michael J. McLean, Kenneth H. Wolfe, Kevin
J Mol Evol (1998) 47:691 696 Springer-Verlag New York Inc. 1998 Base Composition Skews, Replication Orientation, and Gene Orientation in 12 Prokaryote Genomes Michael J. McLean, Kenneth H. Wolfe, Kevin
Genomic Arrangement of Regulons in Bacterial Genomes
 l Genomes Han Zhang 1,2., Yanbin Yin 1,3., Victor Olman 1, Ying Xu 1,3,4 * 1 Computational Systems Biology Laboratory, Department of Biochemistry and Molecular Biology and Institute of Bioinformatics,
l Genomes Han Zhang 1,2., Yanbin Yin 1,3., Victor Olman 1, Ying Xu 1,3,4 * 1 Computational Systems Biology Laboratory, Department of Biochemistry and Molecular Biology and Institute of Bioinformatics,
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
Supplemental Materials
 JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
Analyzing DNA Strand Compositional Asymmetry to Identify Candidate Replication Origins of Borrelia burgdorferi Linear and Circular Plasmids
 Letter Analyzing DNA Strand Compositional Asymmetry to Identify Candidate Replication Origins of Borrelia burgdorferi Linear and Circular Plasmids Mathieu Picardeau, 1,3 Jean R. Lobry, 2 and B. Joseph
Letter Analyzing DNA Strand Compositional Asymmetry to Identify Candidate Replication Origins of Borrelia burgdorferi Linear and Circular Plasmids Mathieu Picardeau, 1,3 Jean R. Lobry, 2 and B. Joseph
Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder
 Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder nrpace@colorado.edu Outline What is the Tree of Life? -- Historical Conceptually
Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder nrpace@colorado.edu Outline What is the Tree of Life? -- Historical Conceptually
Genomics Mycoplasma genitalium
 BIMM 122 Lecture Notes #4 Genomics Dr. Milton Saier Mycoplasma genitalium (C.M. Fraser et al., Science, 270, 397-403, 1995) Genome: 580,070 bp 470 genes* *(counting ones that code for >100 residue proteins)
BIMM 122 Lecture Notes #4 Genomics Dr. Milton Saier Mycoplasma genitalium (C.M. Fraser et al., Science, 270, 397-403, 1995) Genome: 580,070 bp 470 genes* *(counting ones that code for >100 residue proteins)
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss
 Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
Genetic transcription and regulation
 Genetic transcription and regulation Central dogma of biology DNA codes for DNA DNA codes for RNA RNA codes for proteins not surprisingly, many points for regulation of the process https://www.youtube.com/
Genetic transcription and regulation Central dogma of biology DNA codes for DNA DNA codes for RNA RNA codes for proteins not surprisingly, many points for regulation of the process https://www.youtube.com/
Genetic transcription and regulation
 Genetic transcription and regulation Central dogma of biology DNA codes for DNA DNA codes for RNA RNA codes for proteins not surprisingly, many points for regulation of the process DNA codes for DNA replication
Genetic transcription and regulation Central dogma of biology DNA codes for DNA DNA codes for RNA RNA codes for proteins not surprisingly, many points for regulation of the process DNA codes for DNA replication
Genetically Engineering Yeast to Understand Molecular Modes of Speciation
 Genetically Engineering Yeast to Understand Molecular Modes of Speciation Mark Umbarger Biophysics 242 May 6, 2004 Abstract: An understanding of the molecular mechanisms of speciation (reproductive isolation)
Genetically Engineering Yeast to Understand Molecular Modes of Speciation Mark Umbarger Biophysics 242 May 6, 2004 Abstract: An understanding of the molecular mechanisms of speciation (reproductive isolation)
Genetic Basis of Variation in Bacteria
 Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Introduction to polyphasic taxonomy
 Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
SUPPLEMENTARY INFORMATION
 VOLUME: 1 ARTICLE NUMBER: 0010 In the format provided by the authors and unedited. Multicopy plasmids potentiate the evolution of antibiotic resistance in bacteria Alvaro San Millan 1,2,*, Jose Antonio
VOLUME: 1 ARTICLE NUMBER: 0010 In the format provided by the authors and unedited. Multicopy plasmids potentiate the evolution of antibiotic resistance in bacteria Alvaro San Millan 1,2,*, Jose Antonio
The percentage of bacterial genes on leading versus lagging strands is influenced by multiple balancing forces
 The percentage of bacterial genes on leading versus lagging strands is influenced by multiple balancing forces Xizeng Mao 1, Han Zhang 1,4, Yanbin Yin 1, 2 1, 2, 3, Ying Xu 1 Computational Systems Biology
The percentage of bacterial genes on leading versus lagging strands is influenced by multiple balancing forces Xizeng Mao 1, Han Zhang 1,4, Yanbin Yin 1, 2 1, 2, 3, Ying Xu 1 Computational Systems Biology
Genetics 304 Lecture 6
 Genetics 304 Lecture 6 00/01/27 Assigned Readings Busby, S. and R.H. Ebright (1994). Promoter structure, promoter recognition, and transcription activation in prokaryotes. Cell 79:743-746. Reed, W.L. and
Genetics 304 Lecture 6 00/01/27 Assigned Readings Busby, S. and R.H. Ebright (1994). Promoter structure, promoter recognition, and transcription activation in prokaryotes. Cell 79:743-746. Reed, W.L. and
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Repeated Sequences SOPHIE BACHELLIER, ERIC GILSON, MAURICE HOFNUNG, AND CHARLES W. HILL
 Repeated Sequences SOPHIE BACHELLIER, ERIC GILSON, MAURICE HOFNUNG, AND CHARLES W. HILL 112 INTRODUCTION Sequence repetition in Escherichia coli and Salmonella typhimurium (official designation, Salmonella
Repeated Sequences SOPHIE BACHELLIER, ERIC GILSON, MAURICE HOFNUNG, AND CHARLES W. HILL 112 INTRODUCTION Sequence repetition in Escherichia coli and Salmonella typhimurium (official designation, Salmonella
This document describes the process by which operons are predicted for genes within the BioHealthBase database.
 1. Purpose This document describes the process by which operons are predicted for genes within the BioHealthBase database. 2. Methods Description An operon is a coexpressed set of genes, transcribed onto
1. Purpose This document describes the process by which operons are predicted for genes within the BioHealthBase database. 2. Methods Description An operon is a coexpressed set of genes, transcribed onto
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
METHODS FOR DETERMINING PHYLOGENY. In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task.
 Chapter 12 (Strikberger) Molecular Phylogenies and Evolution METHODS FOR DETERMINING PHYLOGENY In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task. Modern
Chapter 12 (Strikberger) Molecular Phylogenies and Evolution METHODS FOR DETERMINING PHYLOGENY In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task. Modern
The Gene The gene; Genes Genes Allele;
 Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
An Analysis of Determinants of Amino Acids Substitution Rates in Bacterial Proteins
 An Analysis of Determinants of Amino Acids Substitution Rates in Bacterial Proteins Eduardo P. C. Rocha* and Antoine Danchin*à *Unité GGB, Institut Pasteur, Paris, France; Atelier de BioInformatique, Université
An Analysis of Determinants of Amino Acids Substitution Rates in Bacterial Proteins Eduardo P. C. Rocha* and Antoine Danchin*à *Unité GGB, Institut Pasteur, Paris, France; Atelier de BioInformatique, Université
RGP finder: prediction of Genomic Islands
 Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
L épidémiologie moléculaire en laboratoire de recherche en bactériologie : l exemple d Escherichia coli
 L épidémiologie moléculaire en laboratoire de recherche en bactériologie : l exemple d Escherichia coli Académie Nationale de Pharmacie Paris 15 Mars 2017 The various E. coli lifestyles Primary habitat
L épidémiologie moléculaire en laboratoire de recherche en bactériologie : l exemple d Escherichia coli Académie Nationale de Pharmacie Paris 15 Mars 2017 The various E. coli lifestyles Primary habitat
Rise and Fall of Mutator Bacteria
 Rise and Fall of Mutator Bacteria The Evolution of Mutation Rates in Bacteria Yoav Ram Hadany Evolutionary Theory Lab 29 May 2011 Bacteria Bacteria are unicellular, haploid and asexual Reproduce by binary
Rise and Fall of Mutator Bacteria The Evolution of Mutation Rates in Bacteria Yoav Ram Hadany Evolutionary Theory Lab 29 May 2011 Bacteria Bacteria are unicellular, haploid and asexual Reproduce by binary
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
The two daughter cells are genetically identical to each other and the parent cell.
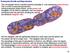 Prokaryote Growth and Reproduction This micrograph shows a bacillus bacteria (probably E. coli) undergoing binary fission. This is a form of asexual reproduction. During prokaryotic binary fission, as
Prokaryote Growth and Reproduction This micrograph shows a bacillus bacteria (probably E. coli) undergoing binary fission. This is a form of asexual reproduction. During prokaryotic binary fission, as
Microbial Taxonomy and the Evolution of Diversity
 19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
PROTEIN SYNTHESIS INTRO
 MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
The Small, Slow and Specialized CRISPR and Anti-CRISPR of Escherichia and Salmonella
 The Small, Slow and Specialized CRISPR and Anti-CRISPR of Escherichia and Salmonella Marie Touchon, Eduardo P. C. Rocha To cite this version: Marie Touchon, Eduardo P. C. Rocha. The Small, Slow and Specialized
The Small, Slow and Specialized CRISPR and Anti-CRISPR of Escherichia and Salmonella Marie Touchon, Eduardo P. C. Rocha To cite this version: Marie Touchon, Eduardo P. C. Rocha. The Small, Slow and Specialized
Horizontal transfer and pathogenicity
 Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
Designer Genes C Test
 Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
The use of gene clusters to infer functional coupling
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2896 2901, March 1999 Genetics The use of gene clusters to infer functional coupling ROSS OVERBEEK*, MICHAEL FONSTEIN, MARK D SOUZA*, GORDON D. PUSCH*, AND NATALIA
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2896 2901, March 1999 Genetics The use of gene clusters to infer functional coupling ROSS OVERBEEK*, MICHAEL FONSTEIN, MARK D SOUZA*, GORDON D. PUSCH*, AND NATALIA
Gene Network Science Diagrammatic Cell Language and Visual Cell
 Gene Network Science Diagrammatic Cell Language and Visual Cell Mr. Tan Chee Meng Scientific Programmer, System Biology Group, Bioinformatics Institute Overview Introduction Why? Challenges Diagrammatic
Gene Network Science Diagrammatic Cell Language and Visual Cell Mr. Tan Chee Meng Scientific Programmer, System Biology Group, Bioinformatics Institute Overview Introduction Why? Challenges Diagrammatic
Comparative Bioinformatics Midterm II Fall 2004
 Comparative Bioinformatics Midterm II Fall 2004 Objective Answer, part I: For each of the following, select the single best answer or completion of the phrase. (3 points each) 1. Deinococcus radiodurans
Comparative Bioinformatics Midterm II Fall 2004 Objective Answer, part I: For each of the following, select the single best answer or completion of the phrase. (3 points each) 1. Deinococcus radiodurans
The Evolution of Infectious Disease
 The Evolution of Infectious Disease Why are some bacteria pathogenic to humans while other (closely-related) bacteria are not? This question can be approached from two directions: 1.From the point of view
The Evolution of Infectious Disease Why are some bacteria pathogenic to humans while other (closely-related) bacteria are not? This question can be approached from two directions: 1.From the point of view
Biology. Biology. Slide 1 of 26. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Quantitative Molecular Biology
 Quantitative Molecular Biology PHYS 176/276 Instructor: Terry Hwa Winter 2018 What is quantitative biology? èquantitative biology biology + numbers/equations biology-inspired physics application of existing
Quantitative Molecular Biology PHYS 176/276 Instructor: Terry Hwa Winter 2018 What is quantitative biology? èquantitative biology biology + numbers/equations biology-inspired physics application of existing
Cladistics and Bioinformatics Questions 2013
 AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
Genômica comparativa. João Carlos Setubal IQ-USP outubro /5/2012 J. C. Setubal
 Genômica comparativa João Carlos Setubal IQ-USP outubro 2012 11/5/2012 J. C. Setubal 1 Comparative genomics There are currently (out/2012) 2,230 completed sequenced microbial genomes publicly available
Genômica comparativa João Carlos Setubal IQ-USP outubro 2012 11/5/2012 J. C. Setubal 1 Comparative genomics There are currently (out/2012) 2,230 completed sequenced microbial genomes publicly available
2. What was the Avery-MacLeod-McCarty experiment and why was it significant? 3. What was the Hershey-Chase experiment and why was it significant?
 Name Date Period AP Exam Review Part 6: Molecular Genetics I. DNA and RNA Basics A. History of finding out what DNA really is 1. What was Griffith s experiment and why was it significant? 1 2. What was
Name Date Period AP Exam Review Part 6: Molecular Genetics I. DNA and RNA Basics A. History of finding out what DNA really is 1. What was Griffith s experiment and why was it significant? 1 2. What was
Lecture 5: Processes and Timescales: Rates for the fundamental processes 5.1
 Lecture 5: Processes and Timescales: Rates for the fundamental processes 5.1 Reading Assignment for Lectures 5-6: Phillips, Kondev, Theriot (PKT), Chapter 3 Life is not static. Organisms as a whole are
Lecture 5: Processes and Timescales: Rates for the fundamental processes 5.1 Reading Assignment for Lectures 5-6: Phillips, Kondev, Theriot (PKT), Chapter 3 Life is not static. Organisms as a whole are
Eukaryotic vs. Prokaryotic genes
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome reduction in prokaryotic obligatory intracellular parasites of humans: a comparative analysis
 International Journal of Systematic and Evolutionary Microbiology (2004), 54, 1937 1941 DOI 10.1099/ijs.0.63090-0 Genome reduction in prokaryotic obligatory intracellular parasites of humans: a comparative
International Journal of Systematic and Evolutionary Microbiology (2004), 54, 1937 1941 DOI 10.1099/ijs.0.63090-0 Genome reduction in prokaryotic obligatory intracellular parasites of humans: a comparative
12-5 Gene Regulation
 12-5 Gene Regulation Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 1 of 26 12-5 Gene Regulation Gene Regulation: An Example Gene
12-5 Gene Regulation Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 1 of 26 12-5 Gene Regulation Gene Regulation: An Example Gene
UNIVERSITY OF YORK. BA, BSc, and MSc Degree Examinations Department : BIOLOGY. Title of Exam: Molecular microbiology
 Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Homology Modeling. Roberto Lins EPFL - summer semester 2005
 Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Database and Comparative Identification of Prophages
 Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy
Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy
Initiation of DNA Replication Lecture 3! Linda Bloom! Office: ARB R3-165! phone: !
 Initiation of DNA Replication Lecture 3! Linda Bloom! Office: ARB R3-165! email: lbloom@ufl.edu! phone: 392-8708! 1! Lecture 3 Outline! Students will learn! Basic techniques for visualizing replicating
Initiation of DNA Replication Lecture 3! Linda Bloom! Office: ARB R3-165! email: lbloom@ufl.edu! phone: 392-8708! 1! Lecture 3 Outline! Students will learn! Basic techniques for visualizing replicating
Genomes and Their Evolution
 Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Prediction of response to selection within families
 Note Prediction of response to selection within families WG Hill A Caballero L Dempfle 2 1 Institzite of Cell, Animal and Population Biology, University of Edinburgh, West Mains Road, Edinburgh, EH9 3JT,
Note Prediction of response to selection within families WG Hill A Caballero L Dempfle 2 1 Institzite of Cell, Animal and Population Biology, University of Edinburgh, West Mains Road, Edinburgh, EH9 3JT,
Special Topics on Genetics
 ARISTOTLE UNIVERSITY OF THESSALONIKI OPEN COURSES Section 9: Transposable elements Drosopoulou E License The offered educational material is subject to Creative Commons licensing. For educational material,
ARISTOTLE UNIVERSITY OF THESSALONIKI OPEN COURSES Section 9: Transposable elements Drosopoulou E License The offered educational material is subject to Creative Commons licensing. For educational material,
Phylogenetic relationship among S. castellii, S. cerevisiae and C. glabrata.
 Supplementary Note S2 Phylogenetic relationship among S. castellii, S. cerevisiae and C. glabrata. Phylogenetic trees reconstructed by a variety of methods from either single-copy orthologous loci (Class
Supplementary Note S2 Phylogenetic relationship among S. castellii, S. cerevisiae and C. glabrata. Phylogenetic trees reconstructed by a variety of methods from either single-copy orthologous loci (Class
Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy. Slowly evolving molecules (e.g., rrna) used for large-scale structure; "fast- clock" molecules for fine-structure.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools. Evaluation. Course Homepage.
 CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p
 Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Mining Infrequent Patterns of Two Frequent Substrings from a Single Set of Biological Sequences
 Mining Infrequent Patterns of Two Frequent Substrings from a Single Set of Biological Sequences Daisuke Ikeda Department of Informatics, Kyushu University 744 Moto-oka, Fukuoka 819-0395, Japan. daisuke@inf.kyushu-u.ac.jp
Mining Infrequent Patterns of Two Frequent Substrings from a Single Set of Biological Sequences Daisuke Ikeda Department of Informatics, Kyushu University 744 Moto-oka, Fukuoka 819-0395, Japan. daisuke@inf.kyushu-u.ac.jp
Streptomyces Linear Plasmids: Their Discovery, Functions, Interactions with Other Replicons, and Evolutionary Significance
 Microbiol Monogr (7) F. Meinhardt R. Klassen: Microbial Linear Plasmids DOI 10.1007/7171_2007_097/Published online: 27 June 2007 Springer-Verlag Berlin Heidelberg 2007 Streptomyces Linear Plasmids: Their
Microbiol Monogr (7) F. Meinhardt R. Klassen: Microbial Linear Plasmids DOI 10.1007/7171_2007_097/Published online: 27 June 2007 Springer-Verlag Berlin Heidelberg 2007 Streptomyces Linear Plasmids: Their
ABSTRACT. As a result of recent successes in genome scale studies, especially genome
 ABSTRACT Title of Dissertation / Thesis: COMPUTATIONAL ANALYSES OF MICROBIAL GENOMES OPERONS, PROTEIN FAMILIES AND LATERAL GENE TRANSFER. Yongpan Yan, Doctor of Philosophy, 2005 Dissertation / Thesis Directed
ABSTRACT Title of Dissertation / Thesis: COMPUTATIONAL ANALYSES OF MICROBIAL GENOMES OPERONS, PROTEIN FAMILIES AND LATERAL GENE TRANSFER. Yongpan Yan, Doctor of Philosophy, 2005 Dissertation / Thesis Directed
GENE REGULATION AND PROBLEMS OF DEVELOPMENT
 GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
Caulobacter crescentus as a model for the study of bacterial cell cycle regulation.
 Caulobacter crescentus as a model for the study of bacterial cell cycle regulation. Leticia Britos Cavagnaro Shapiro Lab Developmental Biology Department Stanford University Note:This is a modified version
Caulobacter crescentus as a model for the study of bacterial cell cycle regulation. Leticia Britos Cavagnaro Shapiro Lab Developmental Biology Department Stanford University Note:This is a modified version
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
GENE ACTIVITY Gene structure Transcription Transcript processing mrna transport mrna stability Translation Posttranslational modifications
 1 GENE ACTIVITY Gene structure Transcription Transcript processing mrna transport mrna stability Translation Posttranslational modifications 2 DNA Promoter Gene A Gene B Termination Signal Transcription
1 GENE ACTIVITY Gene structure Transcription Transcript processing mrna transport mrna stability Translation Posttranslational modifications 2 DNA Promoter Gene A Gene B Termination Signal Transcription
Microbial Genetics, Mutation and Repair. 2. State the function of Rec A proteins in homologous genetic recombination.
 Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Hiromi Nishida. 1. Introduction. 2. Materials and Methods
 Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
A genomic-scale search for regulatory binding sites in the integration host factor regulon of Escherichia coli K12
 The integration host factor regulon of E. coli K12 genome 783 A genomic-scale search for regulatory binding sites in the integration host factor regulon of Escherichia coli K12 M. Trindade dos Santos and
The integration host factor regulon of E. coli K12 genome 783 A genomic-scale search for regulatory binding sites in the integration host factor regulon of Escherichia coli K12 M. Trindade dos Santos and
1. In most cases, genes code for and it is that
 Name Chapter 10 Reading Guide From DNA to Protein: Gene Expression Concept 10.1 Genetics Shows That Genes Code for Proteins 1. In most cases, genes code for and it is that determine. 2. Describe what Garrod
Name Chapter 10 Reading Guide From DNA to Protein: Gene Expression Concept 10.1 Genetics Shows That Genes Code for Proteins 1. In most cases, genes code for and it is that determine. 2. Describe what Garrod
Phylogeny and systematics. Why are these disciplines important in evolutionary biology and how are they related to each other?
 Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
WHERE DOES THE VARIATION COME FROM IN THE FIRST PLACE?
 What factors contribute to phenotypic variation? The world s tallest man, Sultan Kosen (8 feet 1 inch) towers over the world s smallest, He Ping (2 feet 5 inches). WHERE DOES THE VARIATION COME FROM IN
What factors contribute to phenotypic variation? The world s tallest man, Sultan Kosen (8 feet 1 inch) towers over the world s smallest, He Ping (2 feet 5 inches). WHERE DOES THE VARIATION COME FROM IN
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
Revisiting the Central Dogma The role of Small RNA in Bacteria
 Graduate Student Seminar Revisiting the Central Dogma The role of Small RNA in Bacteria The Chinese University of Hong Kong Supervisor : Prof. Margaret Ip Faculty of Medicine Student : Helen Ma (PhD student)
Graduate Student Seminar Revisiting the Central Dogma The role of Small RNA in Bacteria The Chinese University of Hong Kong Supervisor : Prof. Margaret Ip Faculty of Medicine Student : Helen Ma (PhD student)
Translation - Prokaryotes
 1 Translation - Prokaryotes Shine-Dalgarno (SD) Sequence rrna 3 -GAUACCAUCCUCCUUA-5 mrna...ggagg..(5-7bp)...aug Influences: Secondary structure!! SD and AUG in unstructured region Start AUG 91% GUG 8 UUG
1 Translation - Prokaryotes Shine-Dalgarno (SD) Sequence rrna 3 -GAUACCAUCCUCCUUA-5 mrna...ggagg..(5-7bp)...aug Influences: Secondary structure!! SD and AUG in unstructured region Start AUG 91% GUG 8 UUG
