The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome
|
|
|
- Holly Gilbert
- 5 years ago
- Views:
Transcription
1 Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG GCAATGGGAAGCTCTTCTGGAGAGTGAGAGAAGCTTCCAGTTAAGGTGACATTGAAGC AAGTCCTGAAAGATGAGGAAGAGTTGTATGAGAGTGGGGAGGGAAGGGGGAGGTGGA GGGATGGGGAATGGGCCGGGATGGGATAGCGCAAACTGCCCGGGAAGGGAAACCAGCA TGTACAGACCTGAACAACGAAGATGGCATATTTTGTTCAGGGAATGGTGAATTAAGTGT GGCAGGAATGCTTTGTAGACACAGTAATTTGCTTGTATGGAATTTTGCCTGAGAGACCTC CTTCATCCATCACTGTCCTTGTCAAATAGTTTGGAACAGGTATAATGATCACAATAACCC AAGCATAATATTTCGTTAATTCTCACAGAATCACATATAGGTGCCACAGTTATCCCCATT TATGAATGGAGTMinimalProkaryoticGenomeGATGAAAACCTTAGGAATAATGAA GATTTGCGCAGGCTCACCTGGATATTAAGACTGAGTCAAATGTTGGGTCTGGTCTGACT TAATGTTTGCTTTGTTCATGAGCACCACATATTGCCTCTCCTATGCAGTTAAGCAGGTAG GTGACAGAAAAGCCCATGTTTGTCTCTACTCACACACTTCCGACTGAATGTATGTATGGA GTTTCTACACCAGATTCTTCAGTGCTCTGGATATTAACTGGGTATCCCATGACTTTATTCT GACACTACCTGGACCTTGTCAAATAGTTTGGACCTTGTCAAATAGTTTGGAGTCCTTGTCA AATAGTTTGGGGTTAGCACAGACCCCACAAGTTAGGGGCTCAGTCCCACGAGGCCATCCT ACTTCAGATGACAATGGCAAGTCCTAAGTTGTCACCATACTTTTGACCAACCTGTTACC AATCGGGGGTTCCCGTAACTGTCTTCTTGGGTTTAATAATTTGCTAGAACAGTTTACGGA ACTCAGAAAAACAGTTTATTTTCTTTTTTTCTGAGAGAGAGGGTCTTATTTTGTTGCCCAG GCTGGTGTGCAATGGTGCAGTCATAGCTCATTGCAGCCTTGATTGTCTGGGTTCCAGTGG TCTCCCACCTCAGCCTCCCTAGTAGCTGAGACTACATGCCTGCACCACCACATCTGGCTA GTTTCTTTTATTTTTTGTATAGATGGGGTCTTGTTGTGTTGGCCAGGCTGGCCACAAATTC TGGTCTCAAGTGATCCTCCCACCTCAGCCTCTGAAAGTGCTGGGATTACAGATGTGAGC ACCACATCTGGCCAGTTCATTTCCTATTACTGGTTCATTGTGAAGGATACATCTCAGAAA AGTCAATGAAAGAGACGTGCATGCTGGATGCAGTGGCTCATGCCTGTAATCTCAGCACT The minimal prokaryotic genome The minimal prokaryotic genome Mycoplasma genitalium ~ genes not required for growth in laboratory ~ genes required for growth in laboratory The minimal prokaryotic genome Diagram of the genome of Mycoplasma genitalium proteins The minimal prokaryotic genome Haemophilus influenzae (1703) Using transposon mutagenesis (one gene disruptions): 130 genes not required for growth in laboratory 350 genes required for growth in laboratory 240 Mycoplasma genitalium (468) C.M. Fraser et al., Science
2 The minimal gene set A synthetic minimal genome The only way to better understand the minimal component of cellular life and understand the evolution of life Koonin, E.V. NRM 2003 CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG GCAATGGGAAGCTCTTCTGGAGAGTGAGAGAAGCTTCCAGTTAAGGTGACATTGAAGC AAGTCCTGAAAGATGAGGAAGAGTTGTATGAGAGTGGGGAGGGAAGGGGGAGGTGGA GGGATGGGGAATGGGCCGGGATGGGATAGCGCAAACTGCCCGGGAAGGGAAACCAGCA TGTACAGACCTGAACAACGAAGATGGCATATTTTGTTCAGGGAATGGTGAATTAAGTGT GGCAGGAATGCTTTGTAGACACAGTAATTTGCTTGTATGGAATTTTGCCTGAGAGACCTC CTTCATCCATCACTGTCCTTGTCAAATAGTTTGGAACAGGTATAATGATCACAATAACCC AAGCATAATATTTCGTTAATTCTCACAGAATCACATATAGGTGCCACAGTTATCCCCATT TATGAATGGAGTProkaryoticCoreGenomeGATGAAAACCTTAGGAATAATGAATGA TTGCGCAGGCTCACCTGGATATTAAGACTGAGTCAAATGTTGGGTCTGGTCTGACTTTA ATGTTTGCTTTGTTCATGAGCACCACATATTGCCTCTCCTATGCAGTTAAGCAGGTAGGTG ACAGAAAAGCCCATGTTTGTCTCTACTCACACACTTCCGACTGAATGTATGTATGGAGTT CTACACCAGATTCTTCAGTGCTCTGGATATTAACTGGGTATCCCATGACTTTATTCTGAC ACTACCTGGACCTTGTCAAATAGTTTGGACCTTGTCAAATAGTTTGGAGTCCTTGTCAAAT AGTTTGGGGTTAGCACAGACCCCACAAGTTAGGGGCTCAGTCCCACGAGGCCATCCTCAC TCAGATGACAATGGCAAGTCCTAAGTTGTCACCATACTTTTGACCAACCTGTTACCAAT GGGGGTTCCCGTAACTGTCTTCTTGGGTTTAATAATTTGCTAGAACAGTTTACGGAACTC AGAAAAACAGTTTATTTTCTTTTTTTCTGAGAGAGAGGGTCTTATTTTGTTGCCCAGGCTG GTGTGCAATGGTGCAGTCATAGCTCATTGCAGCCTTGATTGTCTGGGTTCCAGTGGTTCTC CACCTCAGCCTCCCTAGTAGCTGAGACTACATGCCTGCACCACCACATCTGGCTAGTTT TTTTATTTTTTGTATAGATGGGGTCTTGTTGTGTTGGCCAGGCTGGCCACAAATTCCTGG CTCAAGTGATCCTCCCACCTCAGCCTCTGAAAGTGCTGGGATTACAGATGTGAGCCACC ACATCTGGCCAGTTCATTTCCTATTACTGGTTCATTGTGAAGGATACATCTCAGAAACAGT AATGAAAGAGACGTGCATGCTGGATGCAGTGGCTCATGCCTGTAATCTCAGCACTTTGG The prokaryotic core genome How big? How stable? Common history? Core genome is NOT = minimal genome!!! Why so small? Non-orthologous gene displacement Maybe, at deep divergences, many true orthologs fall below the radar screen of BLAST Maybe, a few radically reduced parasite genomes are skewing the analysis But just maybe, there really are only about 100 genes, mostly translational/transcriptional in the true core - all else is mix and match So what?!? Doolittle, F. Genomes2005, Halifax 2
3 The phylogenetic problem resolved? Phylogenetic incongruence (HGT - hidden paralogies) Patchy distribution (different gene content within species) Loss of phylogenetic signal for deep branches More data = more phylogenetic signal? Resist both loss and HGT? Daubin et al., GR 2002 All the possible comparisons between gene phylogenies by using principle component analysis 120 genes with common phylogenetic history Phylogenetic artifact Lack of signal HGT Daubin et al., GR 2002 Among these, 205 contain exactly one gene per species. We consider these 205 genes to represent likely orthologs and, consequently, to be good candidates for use in inferring the organismal phylogeny and the extent of LGT the Shimodaira Hasegawa (SH) test Lerat et al., PLOS 2004 Lerat et al., PLOS
4 Failure of rejection is not the same as support Ford W. Doolittle Genes with little signal may fail to reject many or even all topologies, but they cannot be said to support a certain topology it is possible that a robust tree based on concatenated sequences is well supported because different constituent genes contribute strong support to different individual nodes of the tree, without any supporting that tree over all statistical test for each gene against many topologies more nuanced than rejection or failure of rejection visualize the compatibility of all genes with all trees simultaneously Susko et al., MBE 2006 Heat map = simultaneous display of all combinations of genes and test topologies together with simultaneous clustering of both genes and topologies according to p- values Clustering of genes identifies the core set of genes with a similar evol. history. Clustering of toplogoies identifies which trees are (nearly) equally supported (= # best trees) Susko et al., MBE 2006 Susko et al., MBE 2006 Can we prove Darwin s s theory of evolution? Suited evidence would be: a molecular phylogeny similar to organism phylogeny Our phylogenetic analyses do not support treethinking.... Representations other than a tree should be investigated because a non-critical concatenation of markers could be highly misleading. Bapteste et al., 2005 But for prokaryotes we don t have anything else besides the molecular phylogeny to determine the organism phylogeny = CIRCULAR REASONING to use concatenated genes as a the organism phylogeny We have to live with that!! We could believe in the concatenated phylogeny IF most genes would support the same phylogeny We do live in an era in which we have an enormous amount of data (> 350 genomes) 4
5 Problems with this study according to Ford W. Doolittle: concatenated without evaluating congruence among genes (failure of rejection is not suport) individual genes are compared with the concatenated genes => acception of rejection Even if they were congruent, still it is no prove of Darwin s theory as ONLY 31 genes were considered -> can t be representative of the phylogenetic history of the organisms HGT is shown to make a substantial contribution to genome evolution (up to 25-30%)... therefore no tree can, in principle, fully reflect the course of evolution of species. Eugene V. Koonin What does it mean... to speak of an organismal genealogy when nearly all of the genes in the cell - genes that give its general character - do not share a common history? C.R. Woese 2002 CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG GCAATGGGAAGCTCTTCTGGAGAGTGAGAGAAGCTTCCAGTTAAGGTGACATTGAAGC AAGTCCTGAAAGATGAGGAAGAGTTGTATGAGAGTGGGGAGGGAAGGGGGAGGTGGA GGGATGGGGAATGGGCCGGGATGGGATAGCGCAAACTGCCCGGGAAGGGAAACCAGCA TGTACAGACCTGAACAACGAAGATGGCATATTTTGTTCAGGGAATGGTGAATTAAGTGT GGCAGGAATGCTTTGTAGACACAGTAATTTGCTTGTATGGAATTTTGCCTGAGAGACCTC CTTCATCCATCACTGTCCTTGTCAAATAGTTTGGAACAGGTATAATGATCACAATAACCC AAGCATAATATTTCGTTAATTCTCACAGAATCACATATAGGTGCCACAGTTATCCCCATT TATGAATGGAGTMicrobialPanGenomeGATGAAAACCTTAGGAATAATGAATGATT GCGCAGGCTCACCTGGATATTAAGACTGAGTCAAATGTTGGGTCTGGTCTGACTTTAAT GTTTGCTTTGTTCATGAGCACCACATATTGCCTCTCCTATGCAGTTAAGCAGGTAGGTGAC AGAAAAGCCCATGTTTGTCTCTACTCACACACTTCCGACTGAATGTATGTATGGAGTTTCT ACACCAGATTCTTCAGTGCTCTGGATATTAACTGGGTATCCCATGACTTTATTCTGACACT ACCTGGACCTTGTCAAATAGTTTGGACCTTGTCAAATAGTTTGGAGTCCTTGTCAAATAGT TGGGGTTAGCACAGACCCCACAAGTTAGGGGCTCAGTCCCACGAGGCCATCCTCACTTC AGATGACAATGGCAAGTCCTAAGTTGTCACCATACTTTTGACCAACCTGTTACCAATCGG GGGTTCCCGTAACTGTCTTCTTGGGTTTAATAATTTGCTAGAACAGTTTACGGAACTCAGA AAAACAGTTTATTTTCTTTTTTTCTGAGAGAGAGGGTCTTATTTTGTTGCCCAGGCTGGTG GCAATGGTGCAGTCATAGCTCATTGCAGCCTTGATTGTCTGGGTTCCAGTGGTTCTCCCA CTCAGCCTCCCTAGTAGCTGAGACTACATGCCTGCACCACCACATCTGGCTAGTTTCTTT ATTTTTTGTATAGATGGGGTCTTGTTGTGTTGGCCAGGCTGGCCACAAATTCCTGGTCTC AAGTGATCCTCCCACCTCAGCCTCTGAAAGTGCTGGGATTACAGATGTGAGCCACCACAT TGGCCAGTTCATTTCCTATTACTGGTTCATTGTGAAGGATACATCTCAGAAACAGTCAAT GAAAGAGACGTGCATGCTGGATGCAGTGGCTCATGCCTGTAATCTCAGCACTTTGGGAGG Intra-species comparisons A strain is only a single representative of a species, the members of which can be genotypically and phenotypically much more diverse. How many genomes are needed to fully describe a bacterial species? Most analyses have revealed large differences in gene content between closely related strains. Lawrence COiM 2005 Medini et al., COiGD
6 the microbial pan-genome the microbial pan-genome From this study: This question was addressed by sequencing the genomes of 8 Streptococcus agalactiae group B strains (GBS) A bacterial species can be described by its pan-genome (pan, Greek for whole ) Each strain on average 1806 genes present in every strain (core genome) plus 439 genes that are absent in one or more strains (dispensable genome) the present GBS pan-genome contains 2713 genes unique genes will continue to emerge even after 100s or 1000s genomes on average 33 new genes with every new strain sequenced core = essence of the species dispensable = diversity of the species (conserved + strain-specific) The bacterial species will never be fully described, i.e. open pan-genome Claire Fraser Claire Fraser, TIGR GBS pan-genome GBS core genome the microbial pan-genome the microbial pan-genome open closed Core genome = essence, basic aspects of the biology of a sp and its major phenotypic traits Dispensable genome = supplementary biochemical pathways and functions that are not essential for bacterial growth but confer selective advantages,such as adaptation to different niches, virulence, capsular serotype, antibiotic resistance, or colonization of a new host. (conserved / unique) Medini COiGD
7 Open pan-genome: typical for species that colonize multiple environments and have multiple ways of exchanging genetic material: e.g. Streptococci, Meningococci, H. pylori, Salmonellae and E. coli each new genome of Streptococci: + 50 genes (Streptococci is an ecological and phenotypical uniform species) Each new genome of E. coli: genes (E. coli might be too heterogeneous to be one species) Closed pan-genome: more conserved, live in isolated niches with limited access to the global microbial gene pool: e.g. B. Anthracis, Mycobacterium tuberculosis, Buchnera aphidicola and Chlamydia trachomatis CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG GCAATGGGAAGCTCTTCTGGAGAGTGAGAGAAGCTTCCAGTTAAGGTGACATTGAAGC AAGTCCTGAAAGATGAGGAAGAGTTGTATGAGAGTGGGGAGGGAAGGGGGAGGTGGA GGGATGGGGAATGGGCCGGGATGGGATAGCGCAAACTGCCCGGGAAGGGAAACCAGCA TGTACAGACCTGAACAACGAAGATGGCATATTTTGTTCAGGGAATGGTGAATTAAGTGT GGCAGGAATGCTTTGTAGACACAGTAATTTGCTTGTATGGAATTTTGCCTGAGAGACCTC CTTCATCCATCACTGTCCTTGTCAAATAGTTTGGAACAGGTATAATGATCACAATAACCC AAGCATAATATTTCGTTAATTCTCACAGAATCACATATAGGTGCCACAGTTATCCCCATT TATGAATGGAGTSpeciesGenomeConceptGATGAAAACCTTAGGAATAATGAATGA TTGCGCAGGCTCACCTGGATATTAAGACTGAGTCAAATGTTGGGTCTGGTCTGACTTTA ATGTTTGCTTTGTTCATGAGCACCACATATTGCCTCTCCTATGCAGTTAAGCAGGTAGGTG ACAGAAAAGCCCATGTTTGTCTCTACTCACACACTTCCGACTGAATGTATGTATGGAGTT CTACACCAGATTCTTCAGTGCTCTGGATATTAACTGGGTATCCCATGACTTTATTCTGAC ACTACCTGGACCTTGTCAAATAGTTTGGACCTTGTCAAATAGTTTGGAGTCCTTGTCAAAT AGTTTGGGGTTAGCACAGACCCCACAAGTTAGGGGCTCAGTCCCACGAGGCCATCCTCAC TCAGATGACAATGGCAAGTCCTAAGTTGTCACCATACTTTTGACCAACCTGTTACCAAT GGGGGTTCCCGTAACTGTCTTCTTGGGTTTAATAATTTGCTAGAACAGTTTACGGAACTC AGAAAAACAGTTTATTTTCTTTTTTTCTGAGAGAGAGGGTCTTATTTTGTTGCCCAGGCTG GTGTGCAATGGTGCAGTCATAGCTCATTGCAGCCTTGATTGTCTGGGTTCCAGTGGTTCTC CACCTCAGCCTCCCTAGTAGCTGAGACTACATGCCTGCACCACCACATCTGGCTAGTTT TTTTATTTTTTGTATAGATGGGGTCTTGTTGTGTTGGCCAGGCTGGCCACAAATTCCTGG CTCAAGTGATCCTCCCACCTCAGCCTCTGAAAGTGCTGGGATTACAGATGTGAGCCACC ACATCTGGCCAGTTCATTTCCTATTACTGGTTCATTGTGAAGGATACATCTCAGAAACAGT AATGAAAGAGACGTGCATGCTGGATGCAGTGGCTCATGCCTGTAATCTCAGCACTTTGG Comparison between DDH and genome similarity What have we learned from almost a decade of extensive genome sequencing with respect to currently named bacterial species? DDH Can we improve the species definition/concept? ANI Konstantinidis PNAS 2005 unpublished study of 28 sequenced strains: y = 0.785x R 2 = Comparison between 16S and genome similarity % DNA-DNA Hybridization S seq id % Conserved DNA % conserved DNA: blastn of 1020 nt frags with 90% seq id. cut off Goris IJSEM (in press) ANI Konstantinidis PNAS
8 Gene content diversity within species Gene content diversity within species % conserved genes ANI Species may differ up to 35% gene content or 20% (excl. hypothetical and mobile elements) 70% DDH = min 80% gene content shared 20% = on average 1000 genes! Konstantinidis PNAS 2005 ANI Konstantinidis PNAS 2005 Evolutionary relatedness should be coupled to ecological relatedness This will give better predictive species definition than just an evolutionary one (what is the genetic basis for ecological distinctiveness) clusters or continuum of diversity? Do bacteria exhibit a genetic continuum in nature or are there coherent sequence/genomic clusters on which a species definition/concept could be based? Freq. 333 Hsp60 sequences from vibrio strains isolated from coastal bacterioplankton Current datasets in taxonomy consists of only a few representative strains per species, consequently borders based on this dataset might not hold when more intraspecies diversity is included! % sequence identity 8
9 Unsolved questions: How is selection acting on bacterial populations? Recombination rate at the whole genome level within a bacterial population? Answers to these questions will advance our knowledge of the two basic processes on which our current species concepts are based. One thing is certain: reconciling eukaryotic and bacterial species under the same biological species concept is NOT possible because these organisms are too different in terms of evolutionary processes 9
Stepping stones towards a new electronic prokaryotic taxonomy. The ultimate goal in taxonomy. Pragmatic towards diagnostics
 Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
The Minimal-Gene-Set -Kapil PHY498BIO, HW 3
 The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
Microbial Taxonomy and the Evolution of Diversity
 19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder
 Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder nrpace@colorado.edu Outline What is the Tree of Life? -- Historical Conceptually
Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder nrpace@colorado.edu Outline What is the Tree of Life? -- Historical Conceptually
Introduction to polyphasic taxonomy
 Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Outline. I. Methods. II. Preliminary Results. A. Phylogeny Methods B. Whole Genome Methods C. Horizontal Gene Transfer
 Comparative Genomics Preliminary Results April 4, 2016 Juan Castro, Aroon Chande, Cheng Chen, Evan Clayton, Hector Espitia, Alli Gombolay, Walker Gussler, Ken Lee, Tyrone Lee, Hari Prasanna, Carlos Ruiz,
Comparative Genomics Preliminary Results April 4, 2016 Juan Castro, Aroon Chande, Cheng Chen, Evan Clayton, Hector Espitia, Alli Gombolay, Walker Gussler, Ken Lee, Tyrone Lee, Hari Prasanna, Carlos Ruiz,
2 Genome evolution: gene fusion versus gene fission
 2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy. Slowly evolving molecules (e.g., rrna) used for large-scale structure; "fast- clock" molecules for fine-structure.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Using phylogenetics to estimate species divergence times... Basics and basic issues for Bayesian inference of divergence times (plus some digression)
 Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Microbial Taxonomy. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
# shared OGs (spa, spb) Size of the smallest genome. dist (spa, spb) = 1. Neighbor joining. OG1 OG2 OG3 OG4 sp sp sp
 Bioinformatics and Evolutionary Genomics: Genome Evolution in terms of Gene Content 3/10/2014 1 Gene Content Evolution What about HGT / genome sizes? Genome trees based on gene content: shared genes Haemophilus
Bioinformatics and Evolutionary Genomics: Genome Evolution in terms of Gene Content 3/10/2014 1 Gene Content Evolution What about HGT / genome sizes? Genome trees based on gene content: shared genes Haemophilus
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Microbial Taxonomy. C. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy 1. Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eucaryote, is in a mess we are stuck with it for traditional
Microbial Taxonomy 1. Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eucaryote, is in a mess we are stuck with it for traditional
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
Microbiology Helmut Pospiech
 Microbiology http://researchmagazine.uga.edu/summer2002/bacteria.htm 05.04.2018 Helmut Pospiech The Species Concept in Microbiology No universally accepted concept of species for prokaryotes Current definition
Microbiology http://researchmagazine.uga.edu/summer2002/bacteria.htm 05.04.2018 Helmut Pospiech The Species Concept in Microbiology No universally accepted concept of species for prokaryotes Current definition
Chapter 19. Microbial Taxonomy
 Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Phylogeny and systematics. Why are these disciplines important in evolutionary biology and how are they related to each other?
 Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
Horizontal Gene Transfer and the Emergence of Darwinian Evolution
 Horizontal Gene Transfer and the Emergence of Darwinian Evolution Thomas C. Butler May 8, 2006 Abstract In this article I discuss the broader view of evolution that is growing out of our increased understanding
Horizontal Gene Transfer and the Emergence of Darwinian Evolution Thomas C. Butler May 8, 2006 Abstract In this article I discuss the broader view of evolution that is growing out of our increased understanding
Microbial Taxonomy and Phylogeny: Extending from rrnas to Genomes
 Microbial Taxonomy and Phylogeny: Extending from rrnas to Genomes Dr. Kostas Konstantinidis Department of Civil and Environmental Engineering & Department of Biology (Adjunct), Center for Bioinformatics
Microbial Taxonomy and Phylogeny: Extending from rrnas to Genomes Dr. Kostas Konstantinidis Department of Civil and Environmental Engineering & Department of Biology (Adjunct), Center for Bioinformatics
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Big Idea 1: The process of evolution drives the diversity and unity of life.
 Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Today's project. Test input data Six alignments (from six independent markers) of Curcuma species
 DNA sequences II Analyses of multiple sequence data datasets, incongruence tests, gene trees vs. species tree reconstruction, networks, detection of hybrid species DNA sequences II Test of congruence of
DNA sequences II Analyses of multiple sequence data datasets, incongruence tests, gene trees vs. species tree reconstruction, networks, detection of hybrid species DNA sequences II Test of congruence of
HORIZONTAL TRANSFER IN EUKARYOTES KIMBERLEY MC GRAIL FERNÁNDEZ GENOMICS
 HORIZONTAL TRANSFER IN EUKARYOTES KIMBERLEY MC GRAIL FERNÁNDEZ GENOMICS OVERVIEW INTRODUCTION MECHANISMS OF HGT IDENTIFICATION TECHNIQUES EXAMPLES - Wolbachia pipientis - Fungus - Plants - Drosophila ananassae
HORIZONTAL TRANSFER IN EUKARYOTES KIMBERLEY MC GRAIL FERNÁNDEZ GENOMICS OVERVIEW INTRODUCTION MECHANISMS OF HGT IDENTIFICATION TECHNIQUES EXAMPLES - Wolbachia pipientis - Fungus - Plants - Drosophila ananassae
AP Curriculum Framework with Learning Objectives
 Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Unsupervised Learning in Spectral Genome Analysis
 Unsupervised Learning in Spectral Genome Analysis Lutz Hamel 1, Neha Nahar 1, Maria S. Poptsova 2, Olga Zhaxybayeva 3, J. Peter Gogarten 2 1 Department of Computer Sciences and Statistics, University of
Unsupervised Learning in Spectral Genome Analysis Lutz Hamel 1, Neha Nahar 1, Maria S. Poptsova 2, Olga Zhaxybayeva 3, J. Peter Gogarten 2 1 Department of Computer Sciences and Statistics, University of
Map of AP-Aligned Bio-Rad Kits with Learning Objectives
 Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Microbiota: Its Evolution and Essence. Hsin-Jung Joyce Wu "Microbiota and man: the story about us
 Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
BIOL 1010 Introduction to Biology: The Evolution and Diversity of Life. Spring 2011 Sections A & B
 BIOL 1010 Introduction to Biology: The Evolution and Diversity of Life. Spring 2011 Sections A & B Steve Thompson: stthompson@valdosta.edu http://www.bioinfo4u.net 1 ʻTree of Life,ʼ ʻprimitive,ʼ ʻprogressʼ
BIOL 1010 Introduction to Biology: The Evolution and Diversity of Life. Spring 2011 Sections A & B Steve Thompson: stthompson@valdosta.edu http://www.bioinfo4u.net 1 ʻTree of Life,ʼ ʻprimitive,ʼ ʻprogressʼ
Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes.
 Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes. Enduring understanding 3.A: Heritable information provides for continuity of life. Essential
Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes. Enduring understanding 3.A: Heritable information provides for continuity of life. Essential
Valley Central School District 944 State Route 17K Montgomery, NY Telephone Number: (845) ext Fax Number: (845)
 Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Big Idea #1: The process of evolution drives the diversity and unity of life
 BIG IDEA! Big Idea #1: The process of evolution drives the diversity and unity of life Key Terms for this section: emigration phenotype adaptation evolution phylogenetic tree adaptive radiation fertility
BIG IDEA! Big Idea #1: The process of evolution drives the diversity and unity of life Key Terms for this section: emigration phenotype adaptation evolution phylogenetic tree adaptive radiation fertility
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Enduring understanding 1.A: Change in the genetic makeup of a population over time is evolution.
 The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Evolution AP Biology
 Darwin s Theory of Evolution How do biologists use evolutionary theory to develop better flu vaccines? Theory: Evolutionary Theory: Why do we need to understand the Theory of Evolution? Charles Darwin:
Darwin s Theory of Evolution How do biologists use evolutionary theory to develop better flu vaccines? Theory: Evolutionary Theory: Why do we need to understand the Theory of Evolution? Charles Darwin:
Quantitative Genetics & Evolutionary Genetics
 Quantitative Genetics & Evolutionary Genetics (CHAPTER 24 & 26- Brooker Text) May 14, 2007 BIO 184 Dr. Tom Peavy Quantitative genetics (the study of traits that can be described numerically) is important
Quantitative Genetics & Evolutionary Genetics (CHAPTER 24 & 26- Brooker Text) May 14, 2007 BIO 184 Dr. Tom Peavy Quantitative genetics (the study of traits that can be described numerically) is important
AP Biology Curriculum Framework
 AP Biology Curriculum Framework This chart correlates the College Board s Advanced Placement Biology Curriculum Framework to the corresponding chapters and Key Concept numbers in Campbell BIOLOGY IN FOCUS,
AP Biology Curriculum Framework This chart correlates the College Board s Advanced Placement Biology Curriculum Framework to the corresponding chapters and Key Concept numbers in Campbell BIOLOGY IN FOCUS,
Microbial Diversity and Assessment (II) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Visualizing and Assessing Phylogenetic Congruence of Core Gene Sets: A Case Study of the g-proteobacteria
 RESEARCH ARTICLES Visualizing and Assessing Phylogenetic Congruence of Core Gene Sets: A Case Study of the g-proteobacteria E. Susko,* 1 J. Leigh, W. F. Doolittle, and E. Bapteste 1 *Genome Atlantic, Department
RESEARCH ARTICLES Visualizing and Assessing Phylogenetic Congruence of Core Gene Sets: A Case Study of the g-proteobacteria E. Susko,* 1 J. Leigh, W. F. Doolittle, and E. Bapteste 1 *Genome Atlantic, Department
Microbial Diversity. Yuzhen Ye I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University
 Microbial Diversity Yuzhen Ye (yye@indiana.edu) I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University Contents Microbial diversity Morphological, structural,
Microbial Diversity Yuzhen Ye (yye@indiana.edu) I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University Contents Microbial diversity Morphological, structural,
Chapters AP Biology Objectives. Objectives: You should know...
 Objectives: You should know... Notes 1. Scientific evidence supports the idea that evolution has occurred in all species. 2. Scientific evidence supports the idea that evolution continues to occur. 3.
Objectives: You should know... Notes 1. Scientific evidence supports the idea that evolution has occurred in all species. 2. Scientific evidence supports the idea that evolution continues to occur. 3.
doi: / _25
 Boc, A., P. Legendre and V. Makarenkov. 2013. An efficient algorithm for the detection and classification of horizontal gene transfer events and identification of mosaic genes. Pp. 253-260 in: B. Lausen,
Boc, A., P. Legendre and V. Makarenkov. 2013. An efficient algorithm for the detection and classification of horizontal gene transfer events and identification of mosaic genes. Pp. 253-260 in: B. Lausen,
A. Incorrect! In the binomial naming convention the Kingdom is not part of the name.
 Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Phylogenetics. Applications of phylogenetics. Unrooted networks vs. rooted trees. Outline
 Phylogenetics Todd Vision iology 522 March 26, 2007 pplications of phylogenetics Studying organismal or biogeographic history Systematics ating events in the fossil record onservation biology Studying
Phylogenetics Todd Vision iology 522 March 26, 2007 pplications of phylogenetics Studying organismal or biogeographic history Systematics ating events in the fossil record onservation biology Studying
Real species are typically defined by the ability of their
 Colloquium Examining bacterial species under the specter of gene transfer and exchange Howard Ochman*, Emmanuelle Lerat, and Vincent Daubin* Departments of *Biochemistry and Molecular Biophysics and Ecology
Colloquium Examining bacterial species under the specter of gene transfer and exchange Howard Ochman*, Emmanuelle Lerat, and Vincent Daubin* Departments of *Biochemistry and Molecular Biophysics and Ecology
Evolutionary Genetics: Part 0.2 Introduction to Population genetics
 Evolutionary Genetics: Part 0.2 Introduction to Population genetics S. chilense S. peruvianum Winter Semester 2012-2013 Prof Aurélien Tellier FG Populationsgenetik Population genetics Evolution = changes
Evolutionary Genetics: Part 0.2 Introduction to Population genetics S. chilense S. peruvianum Winter Semester 2012-2013 Prof Aurélien Tellier FG Populationsgenetik Population genetics Evolution = changes
Evolutionary Genomics and Proteomics
 Evolutionary Genomics and Proteomics Mark Pagel Andrew Pomiankowski Editors Sinauer Associates, Inc. Publishers Sunderland, Massachusetts 01375 Table of Contents Preface xiii Contributors xv CHAPTER 1
Evolutionary Genomics and Proteomics Mark Pagel Andrew Pomiankowski Editors Sinauer Associates, Inc. Publishers Sunderland, Massachusetts 01375 Table of Contents Preface xiii Contributors xv CHAPTER 1
Essentiality in B. subtilis
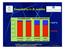 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Fitness constraints on horizontal gene transfer
 Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Introduction to characters and parsimony analysis
 Introduction to characters and parsimony analysis Genetic Relationships Genetic relationships exist between individuals within populations These include ancestordescendent relationships and more indirect
Introduction to characters and parsimony analysis Genetic Relationships Genetic relationships exist between individuals within populations These include ancestordescendent relationships and more indirect
Essential knowledge 1.A.2: Natural selection
 Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Biology. Revisiting Booklet. 6. Inheritance, Variation and Evolution. Name:
 Biology 6. Inheritance, Variation and Evolution Revisiting Booklet Name: Reproduction Name the process by which body cells divide:... What kind of cells are produced this way? Name the process by which
Biology 6. Inheritance, Variation and Evolution Revisiting Booklet Name: Reproduction Name the process by which body cells divide:... What kind of cells are produced this way? Name the process by which
The Prokaryotic World
 The Prokaryotic World A. An overview of prokaryotic life There is no doubt that prokaryotes are everywhere. By everywhere, I mean living in every geographic region, in extremes of environmental conditions,
The Prokaryotic World A. An overview of prokaryotic life There is no doubt that prokaryotes are everywhere. By everywhere, I mean living in every geographic region, in extremes of environmental conditions,
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Quantitative Exploration of the Occurrence of Lateral Gene Transfer Using Nitrogen Fixation Genes as a Case Study
 Lin 1 Quantitative Exploration of the Occurrence of Lateral Gene Transfer Using Nitrogen Fixation Genes as a Case Study by Jason Lin Advisor: Professor Peter Bickel Introduction Under the concept of evolution,
Lin 1 Quantitative Exploration of the Occurrence of Lateral Gene Transfer Using Nitrogen Fixation Genes as a Case Study by Jason Lin Advisor: Professor Peter Bickel Introduction Under the concept of evolution,
OGtree: a tool for creating genome trees of prokaryotes based on overlapping genes
 Published online 2 May 2008 Nucleic Acids Research, 2008, Vol. 36, Web Server issue W475 W480 doi:10.1093/nar/gkn240 OGtree: a tool for creating genome trees of prokaryotes based on overlapping genes Li-Wei
Published online 2 May 2008 Nucleic Acids Research, 2008, Vol. 36, Web Server issue W475 W480 doi:10.1093/nar/gkn240 OGtree: a tool for creating genome trees of prokaryotes based on overlapping genes Li-Wei
A DNA Sequence 2017/12/6 1
 A DNA Sequence ccgtacgtacgtagagtgctagtctagtcgtagcgccgtagtcgatcgtgtgg gtagtagctgatatgatgcgaggtaggggataggatagcaacagatgagc ggatgctgagtgcagtggcatgcgatgtcgatgatagcggtaggtagacttc gcgcataaagctgcgcgagatgattgcaaagragttagatgagctgatgcta
A DNA Sequence ccgtacgtacgtagagtgctagtctagtcgtagcgccgtagtcgatcgtgtgg gtagtagctgatatgatgcgaggtaggggataggatagcaacagatgagc ggatgctgagtgcagtggcatgcgatgtcgatgatagcggtaggtagacttc gcgcataaagctgcgcgagatgattgcaaagragttagatgagctgatgcta
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
Classification and Phylogeny
 Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Origin and Evolution of Life
 Origin and Evolution of Life OCN 201 Science of the Sea Biology Lecture 2 The Handfish -BBC Blue Planet!1!1 Evolution Nothing in biology makes sense except in the light of evolution I am a creationist
Origin and Evolution of Life OCN 201 Science of the Sea Biology Lecture 2 The Handfish -BBC Blue Planet!1!1 Evolution Nothing in biology makes sense except in the light of evolution I am a creationist
Properties of Life. Levels of Organization. Levels of Organization. Levels of Organization. Levels of Organization. The Science of Biology.
 The Science of Biology Chapter 1 Properties of Life Living organisms: are composed of cells are complex and ordered respond to their environment can grow and reproduce obtain and use energy maintain internal
The Science of Biology Chapter 1 Properties of Life Living organisms: are composed of cells are complex and ordered respond to their environment can grow and reproduce obtain and use energy maintain internal
Chapter 17. Organizing Life's Diversity
 Chapter 17 Organizing Life's Diversity Key Concepts: Chapter 17 1. List the 3 domains and the 6 kingdoms. 2. Our current system of classification was originally based on structures; scientists now base
Chapter 17 Organizing Life's Diversity Key Concepts: Chapter 17 1. List the 3 domains and the 6 kingdoms. 2. Our current system of classification was originally based on structures; scientists now base
Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3]
![Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3] Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3]](/thumbs/75/72965665.jpg) Learning Objectives Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3] Refine evidence based on data from many scientific disciplines
Learning Objectives Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3] Refine evidence based on data from many scientific disciplines
Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics.
 Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics. Tree topologies Two topologies were examined, one favoring
Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics. Tree topologies Two topologies were examined, one favoring
AP Biology Review Packet 5- Natural Selection and Evolution & Speciation and Phylogeny
 AP Biology Review Packet 5- Natural Selection and Evolution & Speciation and Phylogeny 1A1- Natural selection is a major mechanism of evolution. 1A2: Natural selection acts on phenotypic variations in
AP Biology Review Packet 5- Natural Selection and Evolution & Speciation and Phylogeny 1A1- Natural selection is a major mechanism of evolution. 1A2: Natural selection acts on phenotypic variations in
 1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
Consensus Methods. * You are only responsible for the first two
 Consensus Trees * consensus trees reconcile clades from different trees * consensus is a conservative estimate of phylogeny that emphasizes points of agreement * philosophy: agreement among data sets is
Consensus Trees * consensus trees reconcile clades from different trees * consensus is a conservative estimate of phylogeny that emphasizes points of agreement * philosophy: agreement among data sets is
BLAST. Varieties of BLAST
 BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
BIOINFORMATICS: An Introduction
 BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
Classification and Phylogeny
 Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
West Windsor-Plainsboro Regional School District AP Biology Grades 11-12
 West Windsor-Plainsboro Regional School District AP Biology Grades 11-12 Unit 1: Chemistry of Life Content Area: Science Course & Grade Level: AP Biology, 11 12 Summary and Rationale The structural levels
West Windsor-Plainsboro Regional School District AP Biology Grades 11-12 Unit 1: Chemistry of Life Content Area: Science Course & Grade Level: AP Biology, 11 12 Summary and Rationale The structural levels
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Chapter 1: Introduction: Themes in the Study of Life
 Name Period Begin your study of biology this year by reading Chapter 1. It will serve as a reminder about biological concepts that you may have learned in an earlier course and give you an overview of
Name Period Begin your study of biology this year by reading Chapter 1. It will serve as a reminder about biological concepts that you may have learned in an earlier course and give you an overview of
Horizontal transfer and pathogenicity
 Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
The Science of Biology. Chapter 1
 The Science of Biology Chapter 1 Properties of Life Living organisms: are composed of cells are complex and ordered respond to their environment can grow and reproduce obtain and use energy maintain internal
The Science of Biology Chapter 1 Properties of Life Living organisms: are composed of cells are complex and ordered respond to their environment can grow and reproduce obtain and use energy maintain internal
Identify stages of plant life cycle Botany Oral/written pres, exams
 DPI Standards Biology Education (for students) 1. Characteristics of organisms Know Properties of living organisms, including: Acquire and use energy and materials Sense and respond to stimuli Reproduce
DPI Standards Biology Education (for students) 1. Characteristics of organisms Know Properties of living organisms, including: Acquire and use energy and materials Sense and respond to stimuli Reproduce
8/23/2014. Phylogeny and the Tree of Life
 Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Molecular phylogeny How to infer phylogenetic trees using molecular sequences
 Molecular phylogeny How to infer phylogenetic trees using molecular sequences ore Samuelsson Nov 200 Applications of phylogenetic methods Reconstruction of evolutionary history / Resolving taxonomy issues
Molecular phylogeny How to infer phylogenetic trees using molecular sequences ore Samuelsson Nov 200 Applications of phylogenetic methods Reconstruction of evolutionary history / Resolving taxonomy issues
Microbes and you ON THE LATEST HUMAN MICROBIOME DISCOVERIES, COMPUTATIONAL QUESTIONS AND SOME SOLUTIONS. Elizabeth Tseng
 Microbes and you ON THE LATEST HUMAN MICROBIOME DISCOVERIES, COMPUTATIONAL QUESTIONS AND SOME SOLUTIONS Elizabeth Tseng Dept. of CSE, University of Washington Johanna Lampe Lab, Fred Hutchinson Cancer
Microbes and you ON THE LATEST HUMAN MICROBIOME DISCOVERIES, COMPUTATIONAL QUESTIONS AND SOME SOLUTIONS Elizabeth Tseng Dept. of CSE, University of Washington Johanna Lampe Lab, Fred Hutchinson Cancer
Chapter 15: Darwin and Evolution
 Chapter 15: Darwin and Evolution AP Curriculum Alignment Big Idea 1 is about evolution. Charles Darwin is called the father of evolution because his theory of natural selection explains how evolution occurs.
Chapter 15: Darwin and Evolution AP Curriculum Alignment Big Idea 1 is about evolution. Charles Darwin is called the father of evolution because his theory of natural selection explains how evolution occurs.
Multiple Sequence Alignment. Sequences
 Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
Molecular phylogeny How to infer phylogenetic trees using molecular sequences
 Molecular phylogeny How to infer phylogenetic trees using molecular sequences ore Samuelsson Nov 2009 Applications of phylogenetic methods Reconstruction of evolutionary history / Resolving taxonomy issues
Molecular phylogeny How to infer phylogenetic trees using molecular sequences ore Samuelsson Nov 2009 Applications of phylogenetic methods Reconstruction of evolutionary history / Resolving taxonomy issues
SPRING GROVE AREA SCHOOL DISTRICT. Course Description. Instructional Strategies, Learning Practices, Activities, and Experiences.
 SPRING GROVE AREA SCHOOL DISTRICT PLANNED COURSE OVERVIEW Course Title: Advanced Placement Biology Grade Level(s): 12 Units of Credit: 1.50 Classification: Elective Length of Course: 30 cycles Periods
SPRING GROVE AREA SCHOOL DISTRICT PLANNED COURSE OVERVIEW Course Title: Advanced Placement Biology Grade Level(s): 12 Units of Credit: 1.50 Classification: Elective Length of Course: 30 cycles Periods
Microbiome: 16S rrna Sequencing 3/30/2018
 Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Microbial Genetics, Mutation and Repair. 2. State the function of Rec A proteins in homologous genetic recombination.
 Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Biology 105/Summer Bacterial Genetics 8/12/ Bacterial Genomes p Gene Transfer Mechanisms in Bacteria p.
 READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis
 Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
CSCI 4181 / CSCI 6802 Algorithms in Bioinformatics
 CSCI 4181 / CSCI 6802 Algorithms in Bioinformatics 1 "In science there is only physics; all the rest is stamp collecting." -Ernest Rutherford 2 Was I a stamp collector? Alan Turing 3 Inte skert, men vad
CSCI 4181 / CSCI 6802 Algorithms in Bioinformatics 1 "In science there is only physics; all the rest is stamp collecting." -Ernest Rutherford 2 Was I a stamp collector? Alan Turing 3 Inte skert, men vad
PACING GUIDE ADVANCED PLACEMENT BIOLOGY
 PACING GUIDE ADVANCED PLACEMENT BIOLOGY BIG IDEAS: 1: The process of evolution drives the diversity and unity of life. 2: Biological systems utilize free energy and molecular building blocks to grow, to
PACING GUIDE ADVANCED PLACEMENT BIOLOGY BIG IDEAS: 1: The process of evolution drives the diversity and unity of life. 2: Biological systems utilize free energy and molecular building blocks to grow, to
