Database and Comparative Identification of Prophages
|
|
|
- Sophia Short
- 5 years ago
- Views:
Transcription
1 Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy 1 1 Bioinformatics centre, School of Biotechnology,Madurai Kamaraj University,Madurai krishna@mrna.tn.nic.in 2 Biological Services,Weizmann Institute of Science Rehovot 76100, Israel 3 Department of Structural Biology, Weizmann Institute of Science Rehovot 76100, Israel Abstract. Prophages are integrated viral genomes in bacteria. Prophages are distinct from other genomic segments encoding virulence factors that have been acquired by horizontal gene transfer events. A database for prophages ( has been constructed with data available from literature reports. To date other than bacteriophage corner stone genes based iterative searches, no other exhaustive approach unique for identifying prophage elements is available. Here we report detection of prophages based on proteomic signature comparison using a prophage proteome as reference set. This method was tested with using the database and then extended over newly sequenced bacterial genomes with no reported prophages. The approach of using similarity of proteins over a given region helped identify twenty putative prophage regions in nine different bacterial genomes. 1 Introduction Bacteriophages are viruses infecting bacteria. Bacteriophages take up two life cycles, one being lytic infects, multiplies, and lyses host bacterium during progeny release [1] whereas in the other temperate mode the phage DNA integrates with the bacterial genome and is termed as prophage. Prophages range from fully viable to cryptic. Cryptic prophages harbor mutational decay and do not result in lytic growth. Prophages can constitute as much as 10-20% of a bacterium s genome (Escherichia coli O157:H7 strain Sakai contains 18 prophage elements constituting 16% of the genome) [2]. Prophage sequences contribute to interstrain variability [3]. At present 230 prophages have been reported in 82 bacterial genomes [4]. In addition, prophages are important vehicles for horizontal gene exchange between different bacterial species. Virulence factors in many pathogenic bacteria are observed to be located on prophage locus, indicating the possible role played by prophages in conferring pathogenicity to host bacterium [5][6][7]. The impact of prophages on bacterial evolution has been reviewed extensively [8]. Prophages do not seem to be a homogenous group and show mosaic nature. Their diverse nature is also reflected by the diversity of genome sizes ranging from 549 kb (Flex9 prophage of Shigella flexneri 2a301) to kb (Bh1 prophage from Bacillus halodurans).analysis of e14 prophage of E coli K-12 revealed the modular nature of the element [9] and provided the basis for the approach of using similarity of proteins over a given region [10]. D.-S. Huang, K. Li, and G.W. Irwin (Eds.): ICIC 2006, LNCIS 344, pp , Springer-Verlag Berlin Heidelberg 2006
2 864 K.V. Srividhya et al. Identifying and understanding prophage elements is medically important as some phage genes are known to increase the survival fitness of lysogens [3]. Unambiguous detection of cryptic prophages is difficult as these defective prophages may be devoid of any corner stone genes. A prophage database has been initiated with the information available from the integrated prophages of sequenced bacterial genomes. At present, the database contains 227 prophage sequences from 49 organisms and details on the twenty putative prophage regions identified in 9 bacterial genomes by protein similarity approach. 2 Methods The prophage database was constructed using PostgreSQL server at backend and PHP in front end. All genomes, proteomes and protein table files were downloaded from ftp://ftp.ncbi.nih.gov/genomes/bacteria. Currently the database covers prophages reported by Casjens [4]. In order to identify e14 homologs, similarity searches at the protein level were done taking the twenty-three e14 proteins as query and the bacterial proteomes as target. Similarity searches were done using blastp, (local version of the WWW-BLAST) [11] using an e-value of 0.01 and Blosum62 as the scoring matrix. 3 Results and Discussion 3.1 Database for Prophage Elements Prophage database covers genome data (GC content, integration site, location, genome size) and protein data (location, protein annotation, related COG information, PDB homologs and Unfoldability index by FoldIndex ). Fig 1 represents the screenshot of prophage database homepage. Fig. 1. Screenshot of Prophage Database home page
3 Database and Comparative Identification of Prophages Identifying Prophage Elements in Bacterial GenoMes The e14 element is a very well characterized prophage element [9], which contains all the highly conserved prophage genes like the integrase, excisionase, phage portal, cro type regulator, repressor and terminase genes elements. Genes encoding the BLAST hits for the different e14 proteins, which were within a particular distance (this distance varies from one organism to another; it is the size of the longest prophage in the organism s genome) were then clubbed together. Any regions with two or more genes in this cluster were considered as putative prophage elements and further analyzed [10]. Fig 2 summarizes the method. Fig. 2. Schematic representation of protein similarity method A set of 61 bacterial genomes was chosen for prophage detection. To confirm the sensitivity of the approach bacterial genomes with prophage incorporated in prophage database data was used for comparison (BLAST hits (e < = 0.01). Out of 174 reported prophages, 87 loci were identified by protein similarity approach. Among them include 27 from E coli (K-12 O157 VT-2 Sakai, CFT073, EDL933) out of 61 reported in literature. With M. tuberculosis 3 loci were detected among the reported 6 prophages.with S.aureus strains 5 could be identified among 7 reported, 10 amongst 19 in Salmonella, 4 out of 10 in Yersinia, 9 among 15 in S.pyogenes. Samples results reflecting 14 genome sequences is presented in Table 1. Hence the method was further extended on to genomes with no reports of prophage. Out of 9 genomes, 24 loci were identified among which 9 were found to be highly probable prophage locations. Table 2 details the prophages identified and respective organisms. For the former, prophage regions were delimited using data from the prophage database and from literature [4]. It was observed that most putative prophage regions encode hypothetical proteins suggesting that these regions need to be characterized further. Interestingly among the newly identified prophages, five are
4 866 K.V. Srividhya et al. Table 1. Bacterial genomes and probable prophages identified using the Protein Similarity Approach method in comparison with literature reports Organism Literature PSA Prophages Detected reported detected B subtilis PBSX, SKIN C. tetani E Cpt2, Cpt3 E. coli K DLP12,QIN, Rac, KpLE1 E. coli O157 VT-2 Sakai Sp8, Sp9, Sp6, Sp4, Sp14, Sp3, Sp15, Sp1,Sp5, Sp12, Sp11 Sp7,Sp10,Sp18,Sp16, Sp17 H. influenzae Rd KW FluMu M. loti MAFF Meso2 M. tuberculosis H37Rv 3 1 phirv1 N. meningitidis Z Pnm2,Pnm1 S. aureus N phin315 S. enterica CT18 (serovar 12 5 Sti4b, Sti8, Sti3, Sti1,Sti7 Typhi) S. flexneri 2a Flex9, Flex5, Flex2 S. pyogenes M1 SF , X. fastidiosa 9a5c 9 1 XfP4 Y. pestis KIM 5 2 Yers3, Yers1 Table 2. Prophage regions detected using the PSA approach from six bacterial genomes. Indicated in # are genomes with no report of prophages. Organism Prophage Gene products S. enterica LT2 St1 Transposase, cyoplasmic (serovar proteins, phoqp Typhimurium) S. flexneri 2457T Sf1 Integrase, replication protein, helicase, mating formation S. pyogenes M18 Sp1 Efflux, phage portal protein, MGAS8232 transposase S. pyogenes M3 Spy1 Transposase, antiterminator MGAS315 P.luminescens subsp. laumondii TTO1 # Mycobacterium bovis AF2122/97 # Pl1,Pl2, P13 P14, 16,P17 4 putative prophages drug resistance protein DNA Invertase HIN, Mostly hypothetical proteins Integrase, transposase. Transcriptional regulatory
5 Database and Comparative Identification of Prophages 867 located near dehydrogenase genes. A priori there seems to be no attributable reason to this tendency for the putative lambdoid phages to get integrated near a dehydrogenase gene in the bacterial genome. However, it must be noted that the search template e14 is also integrated at the isocitrate dehydrogenase gene in the E. coli K12 genome. 3 Conclusion A prophage database has been constructed and used to devise a prophage identification approach using similarity of proteins over a given region. Prediction of prophage related areas in genomes is problematic due to low similarity between prophage genes and the mosaic nature. Five bacterial genomes for which no prophage has been reported in the literature were analyzed in detail. It was observed that most putative prophage regions encode hypothetical proteins suggesting that these regions need to be characterized further. The database provides information on prophages, cryptic prophages and phage remnants, providing effective and efficient way to access the prophage genomes. Prophage identification can be further extended over newly sequenced genomes and incorporated into the database. Acknowledgments We acknowledge the use of Bioinformatics centre facility funded by DBT, Govt of India, DBT and the Israel Ministry of Science and Technology (MOST) for INDO- ISRAEL project support, MOST s support for the Israel Structural Proteomics Center. References 1. Brussow, H., Hendrix, R.: Phage Genomics: Small Is Beautiful. Cell 108 (2002) Canchaya, C., Proux, C., Fournous,G., Bruttin, A., Brussow, H. : Prophage Genomics. Microbiol Mol Biol Rev. 67 (2003) Brussow, H., Canchaya, C., Hardt, W.D.: Phages And The Evolution Of Bacterial Pathogens: From Genomic Rearrangements To Lysogenic Conversion. Microbiol Mol Biol Rev. 68 (2004) Casjens, S.: Prophages And Bacterial Genomics: What Have We Learned So Far? Mol. Microbiol, 49 (2003) Waldor, M, K.: Bacteriophage Biology And Bacterial Virulence. Trends Microbiol, 6 (1998) Davis, B.M., Waldor, and M.K.: Filamentous Phages Linked To Virulence Of Vibrio cholerae. Curr Opin Microbiol, 36 (2000) Boyd, E.F., Brussow, H.: Common Themes Among Bacteriophage-Encoded Virulence Factors And Diversity Among The Bacteriophages Involved. Trends Microbiol, 10 (2002) Canchaya,C., Fournous, G., Brussow, H.: The Impact Of Prophages On Bacterial Chromosomes. Mol Microbiol. 53 (2004) 9-18
6 868 K.V. Srividhya et al. 9. Mehta, P., Casjens, S., Krishnaswamy, S.: Analysis Of The Lambdoid Prophage Element e14 In The E.Coli K12 Genome. BMC Microbiol, 4 (2004) Rao, G.V., Mehta, P.,. Srividhya, K.V., Krishnaswamy, S.: A Protein Similarity Approach For Detecting Prophage Regions In Bacterial Genomes. Genome Biology, 6, (2005) P11 ( 11. Altschul, S.F., Gish, W., Miller, W., Myers, E.W., Lipman, D.J.: Basic Local Alignment. Search Tool. J. Mol. Biol. 215 (1990)
7 Multiple Genome Alignment
 94 Bioinformatics I, WS /3, D. Huson, December 3, 0 7 Multiple Genome Alignment Assume we have a set of genomes G,..., G t that we want to align with each other. If they are short and very closely related,
94 Bioinformatics I, WS /3, D. Huson, December 3, 0 7 Multiple Genome Alignment Assume we have a set of genomes G,..., G t that we want to align with each other. If they are short and very closely related,
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
Genetic Basis of Variation in Bacteria
 Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Phages and the Evolution of Bacterial Pathogens: from Genomic Rearrangements to Lysogenic Conversion
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Sept. 2004, p. 560 602 Vol. 68, No. 3 1092-2172/04/$08.00 0 DOI: 10.1128/MMBR.68.3.560 602.2004 Copyright 2004, American Society for Microbiology. All Rights
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Sept. 2004, p. 560 602 Vol. 68, No. 3 1092-2172/04/$08.00 0 DOI: 10.1128/MMBR.68.3.560 602.2004 Copyright 2004, American Society for Microbiology. All Rights
SPECIES OF ARCHAEA ARE MORE CLOSELY RELATED TO EUKARYOTES THAN ARE SPECIES OF PROKARYOTES.
 THE TERMS RUN AND TUMBLE ARE GENERALLY ASSOCIATED WITH A) cell wall fluidity. B) cell membrane structures. C) taxic movements of the cell. D) clustering properties of certain rod-shaped bacteria. A MAJOR
THE TERMS RUN AND TUMBLE ARE GENERALLY ASSOCIATED WITH A) cell wall fluidity. B) cell membrane structures. C) taxic movements of the cell. D) clustering properties of certain rod-shaped bacteria. A MAJOR
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
The Evolution of Infectious Disease
 The Evolution of Infectious Disease Why are some bacteria pathogenic to humans while other (closely-related) bacteria are not? This question can be approached from two directions: 1.From the point of view
The Evolution of Infectious Disease Why are some bacteria pathogenic to humans while other (closely-related) bacteria are not? This question can be approached from two directions: 1.From the point of view
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
Bacterial Genetics & Operons
 Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss
 Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
In-Depth Assessment of Local Sequence Alignment
 2012 International Conference on Environment Science and Engieering IPCBEE vol.3 2(2012) (2012)IACSIT Press, Singapoore In-Depth Assessment of Local Sequence Alignment Atoosa Ghahremani and Mahmood A.
2012 International Conference on Environment Science and Engieering IPCBEE vol.3 2(2012) (2012)IACSIT Press, Singapoore In-Depth Assessment of Local Sequence Alignment Atoosa Ghahremani and Mahmood A.
BLAST. Varieties of BLAST
 BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
A pathogen is an agent or microrganism that causes a disease in its host. Pathogens can be viruses, bacteria, fungi or protozoa.
 1 A pathogen is an agent or microrganism that causes a disease in its host. Pathogens can be viruses, bacteria, fungi or protozoa. Protozoa are single celled eukaryotic organisms. Some protozoa are pathogens.
1 A pathogen is an agent or microrganism that causes a disease in its host. Pathogens can be viruses, bacteria, fungi or protozoa. Protozoa are single celled eukaryotic organisms. Some protozoa are pathogens.
An Introduction to Sequence Similarity ( Homology ) Searching
 An Introduction to Sequence Similarity ( Homology ) Searching Gary D. Stormo 1 UNIT 3.1 1 Washington University, School of Medicine, St. Louis, Missouri ABSTRACT Homologous sequences usually have the same,
An Introduction to Sequence Similarity ( Homology ) Searching Gary D. Stormo 1 UNIT 3.1 1 Washington University, School of Medicine, St. Louis, Missouri ABSTRACT Homologous sequences usually have the same,
Supplemental Materials
 JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
Obligate anaerobes - cannot grow in the presence of oxygen Facultative anaerobes - can grow with or without oxygen Aerobic - require oxygen
 PROKARYOTES *include bacteria and archaea *singular: bacterium / plural: bacteria PROPERTIES 1. Bacteria are classified into two kingdoms: Eubacteria (true bacteria) and Archaebacteria (Ancient Bacteria).
PROKARYOTES *include bacteria and archaea *singular: bacterium / plural: bacteria PROPERTIES 1. Bacteria are classified into two kingdoms: Eubacteria (true bacteria) and Archaebacteria (Ancient Bacteria).
RGP finder: prediction of Genomic Islands
 Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
AP Bio Module 16: Bacterial Genetics and Operons, Student Learning Guide
 Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Figure S1. Pangenome plots of ten recombining bacterial species based on RAST annotated
 Figure S1 Figure S2 Supplementary Figure legends Figure S1. Pangenome plots of ten recombining bacterial species based on RAST annotated genomes. To generate these plots all strains within a species were
Figure S1 Figure S2 Supplementary Figure legends Figure S1. Pangenome plots of ten recombining bacterial species based on RAST annotated genomes. To generate these plots all strains within a species were
Genome reduction in prokaryotic obligatory intracellular parasites of humans: a comparative analysis
 International Journal of Systematic and Evolutionary Microbiology (2004), 54, 1937 1941 DOI 10.1099/ijs.0.63090-0 Genome reduction in prokaryotic obligatory intracellular parasites of humans: a comparative
International Journal of Systematic and Evolutionary Microbiology (2004), 54, 1937 1941 DOI 10.1099/ijs.0.63090-0 Genome reduction in prokaryotic obligatory intracellular parasites of humans: a comparative
Vital Statistics Derived from Complete Genome Sequencing (for E. coli MG1655)
 We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
Comparison of 61 E. coli genomes
 Comparison of 61 E. coli genomes Center for Biological Sequence Analysis Department of Systems Biology Dave Ussery! DTU course 27105 - Comparative Genomics Oksana s 61 E. coli genomes paper! Monday, 23
Comparison of 61 E. coli genomes Center for Biological Sequence Analysis Department of Systems Biology Dave Ussery! DTU course 27105 - Comparative Genomics Oksana s 61 E. coli genomes paper! Monday, 23
Sequence Alignment Techniques and Their Uses
 Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Horizontal transfer and pathogenicity
 Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
Bioinformatics Study On Mirror DNA In Mycobacterium Tuberculosis Using Fast Parallel Complement Blast Strategy.
 INTERNATIONAL JOURNAL OF SCIENTIFIC & TECHNOLOGY RESEARCH VOLUME 4, ISSUE 2, DECEMBER 205 ISSN 2277-866 Bioinformatics Study On Mirror DNA In Mycobacterium Tuberculosis Using Fast Parallel Complement Blast
INTERNATIONAL JOURNAL OF SCIENTIFIC & TECHNOLOGY RESEARCH VOLUME 4, ISSUE 2, DECEMBER 205 ISSN 2277-866 Bioinformatics Study On Mirror DNA In Mycobacterium Tuberculosis Using Fast Parallel Complement Blast
UNIVERSITY OF YORK. BA, BSc, and MSc Degree Examinations Department : BIOLOGY. Title of Exam: Molecular microbiology
 Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Tiffany Samaroo MB&B 452a December 8, Take Home Final. Topic 1
 Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Bio 119 Bacterial Genomics 6/26/10
 BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids. Prepared by Woojoo Choi
 Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids Prepared by Woojoo Choi Dead or alive? 1) In this chapter we will explore the twilight zone of biology and the gene creature who live there.
Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids Prepared by Woojoo Choi Dead or alive? 1) In this chapter we will explore the twilight zone of biology and the gene creature who live there.
Grundlagen der Bioinformatik, SS 08, D. Huson, May 2,
 Grundlagen der Bioinformatik, SS 08, D. Huson, May 2, 2008 39 5 Blast This lecture is based on the following, which are all recommended reading: R. Merkl, S. Waack: Bioinformatik Interaktiv. Chapter 11.4-11.7
Grundlagen der Bioinformatik, SS 08, D. Huson, May 2, 2008 39 5 Blast This lecture is based on the following, which are all recommended reading: R. Merkl, S. Waack: Bioinformatik Interaktiv. Chapter 11.4-11.7
3.B.1 Gene Regulation. Gene regulation results in differential gene expression, leading to cell specialization.
 3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
CISC 889 Bioinformatics (Spring 2004) Sequence pairwise alignment (I)
 CISC 889 Bioinformatics (Spring 2004) Sequence pairwise alignment (I) Contents Alignment algorithms Needleman-Wunsch (global alignment) Smith-Waterman (local alignment) Heuristic algorithms FASTA BLAST
CISC 889 Bioinformatics (Spring 2004) Sequence pairwise alignment (I) Contents Alignment algorithms Needleman-Wunsch (global alignment) Smith-Waterman (local alignment) Heuristic algorithms FASTA BLAST
Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1
 Name I. Multiple Choice (1 point each) Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1 B 1. Which is possessed by eukaryotes but not by prokaryotes? A. Cell wall B. Distinct nucleus
Name I. Multiple Choice (1 point each) Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1 B 1. Which is possessed by eukaryotes but not by prokaryotes? A. Cell wall B. Distinct nucleus
Regulation of Gene Expression in Bacteria and Their Viruses
 11 Regulation of Gene Expression in Bacteria and Their Viruses WORKING WITH THE FIGURES 1. Compare the structure of IPTG shown in Figure 11-7 with the structure of galactose shown in Figure 11-5. Why is
11 Regulation of Gene Expression in Bacteria and Their Viruses WORKING WITH THE FIGURES 1. Compare the structure of IPTG shown in Figure 11-7 with the structure of galactose shown in Figure 11-5. Why is
Fitness constraints on horizontal gene transfer
 Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
BLAST Database Searching. BME 110: CompBio Tools Todd Lowe April 8, 2010
 BLAST Database Searching BME 110: CompBio Tools Todd Lowe April 8, 2010 Admin Reading: Read chapter 7, and the NCBI Blast Guide and tutorial http://www.ncbi.nlm.nih.gov/blast/why.shtml Read Chapter 8 for
BLAST Database Searching BME 110: CompBio Tools Todd Lowe April 8, 2010 Admin Reading: Read chapter 7, and the NCBI Blast Guide and tutorial http://www.ncbi.nlm.nih.gov/blast/why.shtml Read Chapter 8 for
Comparative Genomics Background & Strategy. Faction 2
 Comparative Genomics Background & Strategy Faction 2 Overview Introduction to comparative genomics Salmonella enterica subsp. enterica serovar Heidelberg Comparative Genomics Faction 2 Objectives Genomic
Comparative Genomics Background & Strategy Faction 2 Overview Introduction to comparative genomics Salmonella enterica subsp. enterica serovar Heidelberg Comparative Genomics Faction 2 Objectives Genomic
Homology Modeling. Roberto Lins EPFL - summer semester 2005
 Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Introduction to Bioinformatics
 Introduction to Bioinformatics Jianlin Cheng, PhD Department of Computer Science Informatics Institute 2011 Topics Introduction Biological Sequence Alignment and Database Search Analysis of gene expression
Introduction to Bioinformatics Jianlin Cheng, PhD Department of Computer Science Informatics Institute 2011 Topics Introduction Biological Sequence Alignment and Database Search Analysis of gene expression
Prokaryotes & Viruses. Practice Questions. Slide 1 / 71. Slide 2 / 71. Slide 3 / 71. Slide 4 / 71. Slide 6 / 71. Slide 5 / 71
 Slide 1 / 71 Slide 2 / 71 New Jersey Center for Teaching and Learning Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of
Slide 1 / 71 Slide 2 / 71 New Jersey Center for Teaching and Learning Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Design of an Enterobacteriaceae Pan-genome Microarray Chip
 Design of an Enterobacteriaceae Pan-genome Microarray Chip Oksana Lukjancenko and David W. Ussery DTU CBS 2010 2 Background Pan-genome complete collection of variuos genes located within populations at
Design of an Enterobacteriaceae Pan-genome Microarray Chip Oksana Lukjancenko and David W. Ussery DTU CBS 2010 2 Background Pan-genome complete collection of variuos genes located within populations at
Biology 105/Summer Bacterial Genetics 8/12/ Bacterial Genomes p Gene Transfer Mechanisms in Bacteria p.
 READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
COMPARATIVE PATHWAY ANNOTATION WITH PROTEIN-DNA INTERACTION AND OPERON INFORMATION VIA GRAPH TREE DECOMPOSITION
 COMPARATIVE PATHWAY ANNOTATION WITH PROTEIN-DNA INTERACTION AND OPERON INFORMATION VIA GRAPH TREE DECOMPOSITION JIZHEN ZHAO, DONGSHENG CHE AND LIMING CAI Department of Computer Science, University of Georgia,
COMPARATIVE PATHWAY ANNOTATION WITH PROTEIN-DNA INTERACTION AND OPERON INFORMATION VIA GRAPH TREE DECOMPOSITION JIZHEN ZHAO, DONGSHENG CHE AND LIMING CAI Department of Computer Science, University of Georgia,
Tutorial 4 Substitution matrices and PSI-BLAST
 Tutorial 4 Substitution matrices and PSI-BLAST 1 Agenda Substitution Matrices PAM - Point Accepted Mutations BLOSUM - Blocks Substitution Matrix PSI-BLAST Cool story of the day: Why should we care about
Tutorial 4 Substitution matrices and PSI-BLAST 1 Agenda Substitution Matrices PAM - Point Accepted Mutations BLOSUM - Blocks Substitution Matrix PSI-BLAST Cool story of the day: Why should we care about
Molecular archaeology of the Escherichia coli genome
 Proc. Natl. Acad. Sci. USA Vol. 95, pp. 9413 9417, August 1998 Evolution Molecular archaeology of the Escherichia coli genome JEFFREY G. LAWRENCE* AND HOWARD OCHMAN *Department of Biological Sciences,
Proc. Natl. Acad. Sci. USA Vol. 95, pp. 9413 9417, August 1998 Evolution Molecular archaeology of the Escherichia coli genome JEFFREY G. LAWRENCE* AND HOWARD OCHMAN *Department of Biological Sciences,
Prokaryotes & Viruses. Multiple Choice Review. Slide 1 / 47. Slide 2 / 47. Slide 3 / 47
 New Jersey enter for Teaching and Learning Slide 1 / 47 Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of students and
New Jersey enter for Teaching and Learning Slide 1 / 47 Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of students and
Bioinformatics Chapter 1. Introduction
 Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Chapters AP Biology Objectives. Objectives: You should know...
 Objectives: You should know... Notes 1. Scientific evidence supports the idea that evolution has occurred in all species. 2. Scientific evidence supports the idea that evolution continues to occur. 3.
Objectives: You should know... Notes 1. Scientific evidence supports the idea that evolution has occurred in all species. 2. Scientific evidence supports the idea that evolution continues to occur. 3.
Prokaryotes & Viruses. Multiple Choice Review. Slide 2 / 47. Slide 1 / 47. Slide 3 (Answer) / 47. Slide 3 / 47. Slide 4 / 47. Slide 4 (Answer) / 47
 Slide 1 / 47 Slide 2 / 47 New Jersey enter for Teaching and Learning Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of
Slide 1 / 47 Slide 2 / 47 New Jersey enter for Teaching and Learning Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of
The invention of the microscope has opened to us a world of extraordinary numbers. A singular drop of pond water reveals countless life forms
 Biology Chapter 19 Notes - Bacteria and Viruses The invention of the microscope has opened to us a world of extraordinary numbers. A singular drop of pond water reveals countless life forms I. Classifying
Biology Chapter 19 Notes - Bacteria and Viruses The invention of the microscope has opened to us a world of extraordinary numbers. A singular drop of pond water reveals countless life forms I. Classifying
Chronology and pattern of integration of tandem genomic islands associated with the tmrna gene in Escherichia coli and Salmonella enterica
 Article Bioinformatics December 2011 Vol.56 No.35: 3836 3844 doi: 10.1007/s11434-011-4749-8 Chronology and pattern of integration of tandem genomic islands associated with the tmrna gene in Escherichia
Article Bioinformatics December 2011 Vol.56 No.35: 3836 3844 doi: 10.1007/s11434-011-4749-8 Chronology and pattern of integration of tandem genomic islands associated with the tmrna gene in Escherichia
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
PDB : 1ZTB Ligand : ZINC
 PDB : 1ZTB Ligand : ZINC00367031 Pantothenate biosynthesis Enzymes with invariant peptides : Pantothenate kinase (CoaA), Pantothenate Synthetase (PanC) Methyltransferase (PanB) Pathway3 Pathway2 E4 X X
PDB : 1ZTB Ligand : ZINC00367031 Pantothenate biosynthesis Enzymes with invariant peptides : Pantothenate kinase (CoaA), Pantothenate Synthetase (PanC) Methyltransferase (PanB) Pathway3 Pathway2 E4 X X
Prokaryotes & Viruses. Multiple Choice Review. Slide 1 / 47. Slide 2 / 47. Slide 3 / 47
 New Jersey enter for Teaching and Learning Slide 1 / 47 Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of students and
New Jersey enter for Teaching and Learning Slide 1 / 47 Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of students and
The Minimal-Gene-Set -Kapil PHY498BIO, HW 3
 The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
Small RNA in rice genome
 Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
T3DB: an integrated database for bacterial type III secretion system
 Wang et al. BMC Bioinformatics 2012, 13:66 DATABASE Open Access T3DB: an integrated database for bacterial type III secretion system Yejun Wang 1, He Huang 2, Ming an Sun 1, Qing Zhang 1 and Dianjing Guo
Wang et al. BMC Bioinformatics 2012, 13:66 DATABASE Open Access T3DB: an integrated database for bacterial type III secretion system Yejun Wang 1, He Huang 2, Ming an Sun 1, Qing Zhang 1 and Dianjing Guo
Part 2. The Basics of Biology:
 Part 2 The Basics of Biology: An Engineer s Perspective Chapter 2 An Overview of Biological Basics 21 2.1 Cells 2.2 Cell Construction 2.3 Cell Nutrient 2.1 Are all cells the same? Cells Basic unit of living
Part 2 The Basics of Biology: An Engineer s Perspective Chapter 2 An Overview of Biological Basics 21 2.1 Cells 2.2 Cell Construction 2.3 Cell Nutrient 2.1 Are all cells the same? Cells Basic unit of living
the noisy gene Biology of the Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB)
 Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Effective Cluster-Based Seed Design for Cross-Species Sequence Comparisons
 Effective Cluster-Based Seed Design for Cross-Species Sequence Comparisons Leming Zhou and Liliana Florea 1 Methods Supplementary Materials 1.1 Cluster-based seed design 1. Determine Homologous Genes.
Effective Cluster-Based Seed Design for Cross-Species Sequence Comparisons Leming Zhou and Liliana Florea 1 Methods Supplementary Materials 1.1 Cluster-based seed design 1. Determine Homologous Genes.
What are viruses? Marine Viruses I & II. OCN 626 Marine Microplankton Ecology. Characteristics, Abundance, and Diversity
 OCN 626 Marine Microplankton Ecology Marine Viruses I & II Characteristics, Abundance, and Diversity What Are Viruses? What are they made of? How do they replicate? Are they alive? What are viruses? Infectious
OCN 626 Marine Microplankton Ecology Marine Viruses I & II Characteristics, Abundance, and Diversity What Are Viruses? What are they made of? How do they replicate? Are they alive? What are viruses? Infectious
MATHEMATICAL MODELS - Vol. III - Mathematical Modeling and the Human Genome - Hilary S. Booth MATHEMATICAL MODELING AND THE HUMAN GENOME
 MATHEMATICAL MODELING AND THE HUMAN GENOME Hilary S. Booth Australian National University, Australia Keywords: Human genome, DNA, bioinformatics, sequence analysis, evolution. Contents 1. Introduction:
MATHEMATICAL MODELING AND THE HUMAN GENOME Hilary S. Booth Australian National University, Australia Keywords: Human genome, DNA, bioinformatics, sequence analysis, evolution. Contents 1. Introduction:
Tools and Algorithms in Bioinformatics
 Tools and Algorithms in Bioinformatics GCBA815, Fall 2015 Week-4 BLAST Algorithm Continued Multiple Sequence Alignment Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and
Tools and Algorithms in Bioinformatics GCBA815, Fall 2015 Week-4 BLAST Algorithm Continued Multiple Sequence Alignment Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
This document describes the process by which operons are predicted for genes within the BioHealthBase database.
 1. Purpose This document describes the process by which operons are predicted for genes within the BioHealthBase database. 2. Methods Description An operon is a coexpressed set of genes, transcribed onto
1. Purpose This document describes the process by which operons are predicted for genes within the BioHealthBase database. 2. Methods Description An operon is a coexpressed set of genes, transcribed onto
Assist. Prof. Martina Šeruga Musić acad. year 2016/17
 Assist. Prof. Martina Šeruga Musić acad. year 2016/17 PHYTOPATHOGENIC BACTERIA there are more than 100 species of known phytopathogenic bacteria genera Agrobacterium, Erwinia, Ralstonia, Pseudomonas, Xanthomonas,
Assist. Prof. Martina Šeruga Musić acad. year 2016/17 PHYTOPATHOGENIC BACTERIA there are more than 100 species of known phytopathogenic bacteria genera Agrobacterium, Erwinia, Ralstonia, Pseudomonas, Xanthomonas,
BACTERIAL PHYSIOLOGY SMALL GROUP. Monday, August 25, :00pm. Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D.
 BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
2012 Univ Aguilera Lecture. Introduction to Molecular and Cell Biology
 2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
Essentiality in B. subtilis
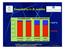 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Topology. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 Topology 1 Introduction 2 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 G+C content 7 Codon usage 27 marc.bailly-bechet@univ-lyon1.fr The big picture Eukaryota Bacteria Many linear chromosomes
Topology 1 Introduction 2 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 G+C content 7 Codon usage 27 marc.bailly-bechet@univ-lyon1.fr The big picture Eukaryota Bacteria Many linear chromosomes
Bioinformatics and BLAST
 Bioinformatics and BLAST Overview Recap of last time Similarity discussion Algorithms: Needleman-Wunsch Smith-Waterman BLAST Implementation issues and current research Recap from Last Time Genome consists
Bioinformatics and BLAST Overview Recap of last time Similarity discussion Algorithms: Needleman-Wunsch Smith-Waterman BLAST Implementation issues and current research Recap from Last Time Genome consists
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment
 Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
2. What was the Avery-MacLeod-McCarty experiment and why was it significant? 3. What was the Hershey-Chase experiment and why was it significant?
 Name Date Period AP Exam Review Part 6: Molecular Genetics I. DNA and RNA Basics A. History of finding out what DNA really is 1. What was Griffith s experiment and why was it significant? 1 2. What was
Name Date Period AP Exam Review Part 6: Molecular Genetics I. DNA and RNA Basics A. History of finding out what DNA really is 1. What was Griffith s experiment and why was it significant? 1 2. What was
ANTIMICROBIAL TESTING. E-Coli K-12 - E-Coli 0157:H7. Salmonella Enterica Servoar Typhimurium LT2 Enterococcus Faecalis
 ANTIMICROBIAL TESTING E-Coli K-12 - E-Coli 0157:H7 Salmonella Enterica Servoar Typhimurium LT2 Enterococcus Faecalis Staphylococcus Aureus (Staph Infection MRSA) Streptococcus Pyrogenes Anti Bacteria effect
ANTIMICROBIAL TESTING E-Coli K-12 - E-Coli 0157:H7 Salmonella Enterica Servoar Typhimurium LT2 Enterococcus Faecalis Staphylococcus Aureus (Staph Infection MRSA) Streptococcus Pyrogenes Anti Bacteria effect
Virus Classifications Based on the Haar Wavelet Transform of Signal Representation of DNA Sequences
 Virus Classifications Based on the Haar Wavelet Transform of Signal Representation of DNA Sequences MOHAMED EL-ZANATY 1, MAGDY SAEB 1, A. BAITH MOHAMED 1, SHAWKAT K. GUIRGUIS 2, EMAN EL-ABD 3 1. School
Virus Classifications Based on the Haar Wavelet Transform of Signal Representation of DNA Sequences MOHAMED EL-ZANATY 1, MAGDY SAEB 1, A. BAITH MOHAMED 1, SHAWKAT K. GUIRGUIS 2, EMAN EL-ABD 3 1. School
CRISPR-SeroSeq: A Developing Technique for Salmonella Subtyping
 Department of Biological Sciences Seminar Blog Seminar Date: 3/23/18 Speaker: Dr. Nikki Shariat, Gettysburg College Title: Probing Salmonella population diversity using CRISPRs CRISPR-SeroSeq: A Developing
Department of Biological Sciences Seminar Blog Seminar Date: 3/23/18 Speaker: Dr. Nikki Shariat, Gettysburg College Title: Probing Salmonella population diversity using CRISPRs CRISPR-SeroSeq: A Developing
chapter 5 the mammalian cell entry 1 (mce1) operon of Mycobacterium Ieprae and Mycobacterium tuberculosis
 chapter 5 the mammalian cell entry 1 (mce1) operon of Mycobacterium Ieprae and Mycobacterium tuberculosis chapter 5 Harald G. Wiker, Eric Spierings, Marc A. B. Kolkman, Tom H. M. Ottenhoff, and Morten
chapter 5 the mammalian cell entry 1 (mce1) operon of Mycobacterium Ieprae and Mycobacterium tuberculosis chapter 5 Harald G. Wiker, Eric Spierings, Marc A. B. Kolkman, Tom H. M. Ottenhoff, and Morten
The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome
 Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Section 19 1 Bacteria (pages )
 Chapter 19 Bacteria and Viruses Section 19 1 Bacteria (pages 471 477) How do the two groups of prokaryotes differ? What factors are used to identify prokaryotes? What is the importance of bacteria? 13.
Chapter 19 Bacteria and Viruses Section 19 1 Bacteria (pages 471 477) How do the two groups of prokaryotes differ? What factors are used to identify prokaryotes? What is the importance of bacteria? 13.
Microbial Genetics, Mutation and Repair. 2. State the function of Rec A proteins in homologous genetic recombination.
 Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Boolean models of gene regulatory networks. Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016
 Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food Supervised by Henrik Hasman, PhD
 By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food elibi@food.dtu.dk Supervised by Henrik Hasman, PhD 1. Introduction to plasmid biology 2. Plasmid encoded resistance to β- lactams (basic theories)
By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food elibi@food.dtu.dk Supervised by Henrik Hasman, PhD 1. Introduction to plasmid biology 2. Plasmid encoded resistance to β- lactams (basic theories)
Sequence Database Search Techniques I: Blast and PatternHunter tools
 Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
Biology Tutorial. Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan
 Biology Tutorial Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan Viruses A T4 bacteriophage injecting DNA into a cell. Influenza A virus Electron micrograph of HIV. Cone-shaped cores are
Biology Tutorial Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan Viruses A T4 bacteriophage injecting DNA into a cell. Influenza A virus Electron micrograph of HIV. Cone-shaped cores are
Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes.
 Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes. Enduring understanding 3.A: Heritable information provides for continuity of life. Essential
Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes. Enduring understanding 3.A: Heritable information provides for continuity of life. Essential
- conserved in Eukaryotes. - proteins in the cluster have identifiable conserved domains. - human gene should be included in the cluster.
 NCBI BLAST Services DELTA-BLAST BLAST (http://blast.ncbi.nlm.nih.gov/), Basic Local Alignment Search tool, is a suite of programs for finding similarities between biological sequences. DELTA-BLAST is a
NCBI BLAST Services DELTA-BLAST BLAST (http://blast.ncbi.nlm.nih.gov/), Basic Local Alignment Search tool, is a suite of programs for finding similarities between biological sequences. DELTA-BLAST is a
Ti plasmid derived plant vector systems: binary and co - integrative vectors transformation process; regeneration of the transformed lines
 Ti plasmid derived plant vector systems: binary and co - integrative vectors transformation process; regeneration of the transformed lines Mitesh Shrestha Constraints of Wild type Ti/Ri-plasmid Very large
Ti plasmid derived plant vector systems: binary and co - integrative vectors transformation process; regeneration of the transformed lines Mitesh Shrestha Constraints of Wild type Ti/Ri-plasmid Very large
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
CSCE555 Bioinformatics. Protein Function Annotation
 CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: A new family of genes responsible for autoinducer production
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 1639 1644, February 1999 Microbiology Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: A new family of genes responsible for autoinducer
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 1639 1644, February 1999 Microbiology Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: A new family of genes responsible for autoinducer
Comparative Studies of Transcriptional Regulation Mechanisms in a Group of Eight Gamma-proteobacterial Genomes
 doi:10.1016/j.jmb.2005.09.037 J. Mol. Biol. (2005) 354, 184 199 Comparative Studies of Transcriptional Regulation Mechanisms in a Group of Eight Gamma-proteobacterial Genomes Vladimir Espinosa 1, Abel
doi:10.1016/j.jmb.2005.09.037 J. Mol. Biol. (2005) 354, 184 199 Comparative Studies of Transcriptional Regulation Mechanisms in a Group of Eight Gamma-proteobacterial Genomes Vladimir Espinosa 1, Abel
Single alignment: Substitution Matrix. 16 march 2017
 Single alignment: Substitution Matrix 16 march 2017 BLOSUM Matrix BLOSUM Matrix [2] (Blocks Amino Acid Substitution Matrices ) It is based on the amino acids substitutions observed in ~2000 conserved block
Single alignment: Substitution Matrix 16 march 2017 BLOSUM Matrix BLOSUM Matrix [2] (Blocks Amino Acid Substitution Matrices ) It is based on the amino acids substitutions observed in ~2000 conserved block
Supplementary Information
 Supplementary Information 1 List of Figures 1 Models of circular chromosomes. 2 Distribution of distances between core genes in Escherichia coli K12, arc based model. 3 Distribution of distances between
Supplementary Information 1 List of Figures 1 Models of circular chromosomes. 2 Distribution of distances between core genes in Escherichia coli K12, arc based model. 3 Distribution of distances between
Genomes and Their Evolution
 Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
PDF // IS BACTERIA A PROKARYOTE OR EUKARYOTE
 19 January, 2018 PDF // IS BACTERIA A PROKARYOTE OR EUKARYOTE Document Filetype: PDF 222.61 KB 0 PDF // IS BACTERIA A PROKARYOTE OR EUKARYOTE How to Tell the Difference Between Prokaryotes and Eukaryotes.
19 January, 2018 PDF // IS BACTERIA A PROKARYOTE OR EUKARYOTE Document Filetype: PDF 222.61 KB 0 PDF // IS BACTERIA A PROKARYOTE OR EUKARYOTE How to Tell the Difference Between Prokaryotes and Eukaryotes.
Comparison of simple sequence repeats in Staphylococcus strains using in-silico approach
 www.bioinformation.net Hypothesis Volume 8(24) Comparison of simple sequence repeats in Staphylococcus strains using in-silico approach Sunil Thorat 1 * & Prashant Thakare 2 1Institute of Bioresources
www.bioinformation.net Hypothesis Volume 8(24) Comparison of simple sequence repeats in Staphylococcus strains using in-silico approach Sunil Thorat 1 * & Prashant Thakare 2 1Institute of Bioresources
