Figure S1. Pangenome plots of ten recombining bacterial species based on RAST annotated
|
|
|
- Millicent Neal
- 5 years ago
- Views:
Transcription
1 Figure S1
2 Figure S2
3 Supplementary Figure legends Figure S1. Pangenome plots of ten recombining bacterial species based on RAST annotated genomes. To generate these plots all strains within a species were annotated using the RAST annotation webserver and the pangenomes were generated as described above. Two plots are provided for each species: Left: HGT-artifact corrected pangenome plot the protein pangenome was compared to the full DNA sequences of the strains from which it was generated. Pangenes appearing in less than 50% of strains within a species at the annotated protein level, but in more than 75% of the strains at the whole DNA level, were removed from the plot; Right: Protein length corrected pangenome plot for each pangene within the HGT-artifact corrected pangenome lengths of all genes within the pangene group were compared to the length of the longest gene within the group. Genes, whose length was shorter than two thirds of the longest gene length, were removed from the pangene cluster and the pangene distribution was re-drawn. Figure S2. Pangenome plots of four clonal bacterial species based on RAST annotated genomes. To generate these plots all strains within a species were annotated using the RAST annotation webserver and the pangenomes were generated as described above. Two plots are provided for each species: Left: HGT-artifact corrected pangenome plot the protein pangenome was compared to the full DNA sequences of the strains from which it was generated. Pangenes appearing in less than 50% of strains within a species at the annotated protein level, but in more than 75% of the strains at the whole DNA level, were removed from the plot; Right: Protein length corrected pangenome plot for each pangene within the HGT-artifact corrected pangenome lengths of all genes within the pangene group were compared to the length of the longest gene within the group. Genes, whose length was shorter than two thirds of the longest gene length, were removed from the pangene cluster and the pangene distribution was re-drawn.
4 Table S1. Investigated strains and the percentage of near core pangenes absent from each strain Bacilllus cereus genes a lost genes b NC_ Bacillus cereus ATCC NC_ Bacillus cereus ATCC NC_ Bacillus cereus E33L NC_ Bacillus cereus AH NC_ Bacillus cereus B NC_ Bacillus cereus G NC_ Bacillus cereus AH NC_ Bacillus cereus Q NC_ Bacillus cereus 03BB NC_ Bacillus cereus biovar anthracis str. CI NC_ Bacillus cereus NC NC_ Bacillus cereus F837/ NC_ Bacillus cereus FRI Campylobacter jejuni Number of Percent of lost genes genes NC_ Campylobacter jejuni RM NC_ Campylobacter jejuni subsp. jejuni NC_ Campylobacter jejuni subsp. doylei NC_ Campylobacter jejuni subsp. jejuni NC_ Campylobacter jejuni subsp. jejuni ICDCCJ NC_ Campylobacter jejuni subsp
5 jejuni IA3902 NC_ Campylobacter jejuni subsp. jejuni M NC_ Campylobacter jejuni subsp. jejuni S NC_ Campylobacter jejuni subsp. jejuni NCTC BN148, NC_ Campylobacter jejuni subsp. jejuni PT NC_ Campylobacter jejuni subsp. jejuni NC_ Campylobacter jejuni subsp. jejuni NC_ Campylobacter jejuni Clostridium botulinum genes lost genes NC_ Clostridium botulinum A str. ATCC NC_ Clostridium botulinum A str. ATCC NC_ Clostridium botulinum A str. Hall NC_ Clostridium botulinum F str. Langeland NC_ Clostridium botulinum B1 str. Okra NC_ Clostridium botulinum A3 str. Loch Maree NC_ Clostridium botulinum A2 str. Kyoto NC_ Clostridium botulinum Ba4 str
6 NC_ NC_ Accession number NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ Clostridium botulinum F str Clostridium botulinum H diphtheriae Strain Name genes lost genes diphtheriae NCTC diphtheriae diphtheriae INCA diphtheriae CDCE diphtheriae HC diphtheriae HC diphtheriae HC diphtheriae PW diphtheriae VA diphtheriae 31A diphtheriae BH diphtheriae C7 (beta) diphtheriae HC
7 Accession number NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ pseudotuberculosis Strain Name genes lost genes pseudotuberculosis FRC pseudotuberculosis 3/ pseudotuberculosis pseudotuberculosis P54B pseudotuberculosis pseudotuberculosis C pseudotuberculosis I pseudotuberculosis PAT pseudotuberculosis 42/02-A pseudotuberculosis CIP pseudotuberculosis 1/06-A pseudotuberculosis pseudotuberculosis pseudotuberculosis pseudotuberculosis Cp
8 Escherichia coli-shigella Group genes lost genes NC_ Escherichia coli str. K-12 substr. MG NC_ Escherichia coli O157:H7 str. EDL NC_ Escherichia coli O157:H7 str. Sakai NC_ Escherichia coli CFT NC_ Escherichia coli str. K-12 substr. W NC_ Escherichia coli UTI NC_ Escherichia coli NC_ Escherichia coli APEC O NC_ Escherichia coli HS NC_ Escherichia coli E24377A NC_ Escherichia coli ATCC NC_ Escherichia coli SMS NC_ Escherichia coli O157:H7 str. EC NC_ Escherichia coli SE NC_ Escherichia coli O127:H6 str. E2348/ NC_ Escherichia coli IAI NC_ Escherichia coli S NC_ Escherichia coli ED1a NC_ Escherichia coli NC_ Escherichia coli IAI NC_ Escherichia coli UMN NC_ Escherichia coli LF NC_ Escherichia coli BW NC_ Escherichia coli B str. REL
9 NC_ Escherichia coli BL21(DE3) NC_ Escherichia coli O157:H7 str. TW NC_ Escherichia coli O103:H2 str NC_ Escherichia coli O26:H11 str NC_ Escherichia coli O111:H- str NC_ Escherichia coli SE NC_ Escherichia coli O55:H7 str. CB NC_ Escherichia coli KO11FL NC_ Escherichia coli DH NC_ Escherichia coli NC_ Escherichia coli IHE NC_ Escherichia coli ABU NC_ Escherichia coli ETEC H NC_ Escherichia coli W NC_ Escherichia coli UMNK NC_ Escherichia coli O7:K1 str. CE NC_ Escherichia coli str. 'clone D i14' NC_ Escherichia coli O55:H7 str. RM NC_ Escherichia coli P12b NC_ Escherichia coli Xuzhou NC_ Escherichia coli O104:H4 str. 2009EL NC_ Escherichia coli O104:H4 str. 2011C NC_ Escherichia coli O104:H4 str
10 2009EL-2071 NC_ Escherichia coli APEC O NC_ Escherichia coli LY NC_ Escherichia coli PMV-1 main NC_ Escherichia coli JJ NC_ Shigella flexneri 2a str NC_ Shigella flexneri 2a str. 2457T NC_ Shigella sonnei Ss NC_ Shigella dysenteriae Sd NC_ Shigella boydii Sb NC_ Shigella flexneri 5 str NC_ Shigella boydii CDC NC_ Shigella sonnei 53G NC_ Shigella flexneri Francisella tularensis genes lost genes NC_ Francisella tularensis subsp. tularensis SCHU S NC_ Francisella tularensis subsp. holarctica LVS NC_ Francisella tularensis subsp. tularensis FSC NC_ Francisella tularensis subsp. holarctica OSU NC_ Francisella tularensis subsp. tularensis WY NC_ Francisella tularensis subsp. holarctica FTNF NC_ Francisella tularensis subsp. mediasiatica FSC NC_ Francisella tularensis TIGB NC_ Francisella tularensis subsp
11 tularensis TI0902 NC_ Francisella tularensis subsp. tularensis NE NC_ Francisella tularensis subsp. holarctica F NC_ Francisella tularensis subsp. holarctica FSC Haemophilus influenzae genes lost genes NC_ Haemophilus influenzae Rd KW NC_ Haemophilus influenzae NP NC_ Haemophilus influenzae PittEE NC_ Haemophilus influenzae PittGG NC_ Haemophilus influenzae F NC_ Haemophilus influenzae F NC_ Haemophilus influenzae NC_ Haemophilus influenzae R NC_ Haemophilus influenzae R NC_ Haemophilus influenzae KR genes lost genes NC_ serotype 4b str. F NC_ EGD-e NC_ HCC NC_
12 Clip81459 NC_ NC_ NC_ L NC_ M NC_ S NC_ J NC_ FSL R NC_ Finland NC_ ATCC NC_ SLCC NC_ SLCC NC_ SLCC NC_ SLCC NC_ SLCC NC_ SLCC NC_ serotype 7 str. SLCC NC_ SLCC NC_
13 SLCC7179 NC_ L NC_ serotype 4b str. LL NC_ J NC_ EGD Complex genes lost genes NC_ CDC NC_ Mycobacterium bovis AF2122/ NC_ Mycobacterium bovis BCG str. Pasteur 1173P NC_ H37Ra NC_ F NC_ Mycobacterium bovis BCG str. Tokyo NC_ KZN NC_ Mycobacterium africanum GM NC_ KZN NC_ Mycobacterium bovis BCG str. Mexico NC_ UT NC_ CCDC
14 NC_ NC_ NC_ NC_ NC_ NC_ NC_ NC_ Accession number CCDC CTRI KZN H37Rv complete genome Mycobacterium bovis BCG str. Korea 1168P str. Erdman = ATCC DNA str. Haarlem Neisseria meningitidis Strain Name genes lost genes NC_ Neisseria meningitidis MC NC_ Neisseria meningitidis Z NC_ Neisseria meningitidis FAM NC_ Neisseria meningitidis NC_ Neisseria meningitidis alpha NC_ Neisseria meningitidis NC_ Neisseria meningitidis alpha NC_ Neisseria meningitidis WUE NC_ Neisseria meningitidis G NC_ Neisseria meningitidis M
15 NC_ Neisseria meningitidis M NC_ Neisseria meningitidis H44/ NC_ Neisseria meningitidis M NC_ Neisseria meningitidis NZ- 05/ Pseudomonas aeruginosa genes lost genes NC_ Pseudomonas aeruginosa PAO NC_ Pseudomonas aeruginosa UCBPP-PA NC_ Pseudomonas aeruginosa PA7 chromosome NC_ Pseudomonas aeruginosa LESB NC_ Pseudomonas aeruginosa M NC_ Pseudomonas aeruginosa NCGM2.S NC_ Pseudomonas aeruginosa DK NC_ Pseudomonas aeruginosa B136-33, NC_ Pseudomonas aeruginosa RP NC_ Pseudomonas aeruginosa PA NC_ Pseudomonas aeruginosa MTB NC_ Pseudomonas aeruginosa LES NC_ Pseudomonas aeruginosa SCV
16 genes lost genes NC_ TIGR NC_ R NC_ D NC_ Hungary19A NC_ CGSP NC_ G NC_ ATCC NC_ JJA NC_ P NC_ NC_ Taiwan19F NC_ TCH8431/19A NC_ AP NC_ B NC_ INV NC_ OXC NC_ INV NC_
17 ST556 NC_ SPNA NC_ gampni NC_ A026 genome Yersinia pestis genes lost genes NC_ Yersinia pestis CO NC_ Yersinia pestis KIM NC_ Yersinia pestis biovar Microtus str NC_ Yersinia pestis Nepal NC_ Yersinia pestis Antiqua NC_ Yersinia pestis Pestoides F NC_ Yersinia pestis Angola NC_ Yersinia pestis Z NC_ Yersinia pestis D NC_ Yersinia pestis D NC_ Yersinia pestis A NC_ Yersinia pestis biovar Medievalis str. Harbin a Number of genes from each strain used for the construction of pangenomes (see Materials and Methods) b Percent of near core gene loss. Calculated as 100*L/T, where L is the number of near core core pangenes that are absent from the genome and T is the total number of genes present within that genome, that were used to construct the species pangenome.
18 Table S2. List of bacterial strains removed from our analysis Campylobacter jejuni Accession number Strain Name NC_ Campylobacter jejuni subsp. jejuni NCTC = ATCC NC_ Campylobacter jejuni subsp. jejuni NC_ Campylobacter jejuni subsp. jejuni NC_ Campylobacter jejuni Clostridium botulinum NC_ Clostridium botulinum B str. Eklund 17B NC_ Clostridium botulinum E3 str. Alaska E43 NC_ Clostridium botulinum BKT Escherichia coli-shigella Group NC_ Escherichia coli NA114 NC_ Escherichia coli UM146 NC_ Escherichia coli O83:H1 str. NRG 857C NC_ Shigella dysenteriae 1617 NC_ Escherichia coli str. K-12 substr. MDS42 DNA NC_ Escherichia coli str. K-12 substr. DH10B NC_ Escherichia coli str. 'clone D i2' NC_ PF0776 NC_ La111 NC_ N53-1 NC_ J1-220 Mycobacterium Tuberculosis Cluster NC_ RGTB327 NC_ RGTB423
19 NC_ NC_ NC_ NC_ Accession number NC_ NC_ NC_ NC_ NC_ str. Beijing/NITR203 str. Haarlem/NITR202 CAS/NITR204 EAI5/NITR206 Pseudomonas aeruginosa Strain Name Pseudomonas aeruginosa c7447m Pseudomonas aeruginosa PA1R Pseudomonas aeruginosa PAO1-VE13 Pseudomonas aeruginosa PAO1-VE2 Pseudomonas aeruginosa PAO581
20 Table S3. Summary of recombination analyses by PHIpack software for 14 analyzed species Species a Number of tested genes Recombinant genes (%) Nonrecombinant genes (%) Noninformative genes (%) B. cereus C. jejuni C. botulinum C. diphteriae C. pseudotuberculosis* E. coli-shigella F. tularensis* H. influenzae L. monocytogenes MTBC* N. meningitidis P. aeruginosa S. pneumoniae Y. pestis* a Clonal species are marked with an asterisk
21 Table S4. Summary of pangene frequencies within the protein pangenomes based on the NCBI annotations of the 14 studied species (prior to HGT correction) Species a Pangenome size b Rare pangenes c Near Core pangenes d Core pangenes e B.cereus C. jejuni C. botulinum C. diphtheriae C.pseudotuberculosis* E. coli-shigella F. tularensis* H. influenzae L. monocytogenes MTBC* N. meningitidis P. aeruginosa S. pneumoniae Y. pestis* a Clonal species are marked with an asterisk b Number of pangenes (orthologous gene clusters) within pangenome c % of pangenes that are found in less than 25% of strains of a species d % of pangenes that are found in over 75% of strains of a species, but are not found in all strains e % of pangenes found in all strains of a species
22 Table S5. Correlations between aaai a and Percentage of genes lost b in the different bacterial species Species c Number of genomes Spearman Rho P-value B.cereus C. jejuni C. botulinum C. diphtheriae C.pseudotuberculosis* E. coli-shigella F. tularensis* H. influenzae L. monocytogenes MTBC* N. meningitidis P. aeruginosa S. pneumoniae Y. pestis* a Denotes for each strain within each species the average AAI for its pairwise comparisons to all other strains within that species b For each strain we calculated the percentage of genes that were clearly lost from that genome as 100*L/T, where L is the number of near core core pangenes that are absent from the genome and T is the total number of genes present within that genome, that were used to construct the species pangenome c Clonal species are marked with an asterisk
23 Table S6. Summary of pangene frequencies within the RAST annotated, length-corrected protein pangenomes of the 14 studied species Species a Pangenome size b Rare pangenes c Near Core pangenes d Core pangenes e B.cereus C. jejuni C. botulinum C. diphtheriae C.pseudotuberculosis* E. coli-shigella F. tularensis* H. influenzae L. monocytogenes MTBC* N. meningitidis P. aeruginosa S. pneumoniae Y. pestis* a Clonal species are marked with an asterisk b Number of pangenes (orthologous gene clusters) within pangenome c % of pangenes that are found in less than 25% of strains of a species d % of pangenes that are found in over 75% of strains of a species, but are not found in all strains e % of pangenes found in all strains of a species
24
Comparison of 61 E. coli genomes
 Comparison of 61 E. coli genomes Center for Biological Sequence Analysis Department of Systems Biology Dave Ussery! DTU course 27105 - Comparative Genomics Oksana s 61 E. coli genomes paper! Monday, 23
Comparison of 61 E. coli genomes Center for Biological Sequence Analysis Department of Systems Biology Dave Ussery! DTU course 27105 - Comparative Genomics Oksana s 61 E. coli genomes paper! Monday, 23
Supplemental. Location Year Source. 1412G Dalidag, Georgia 1979 Flea
 Supplemental Table S1: pestis strains (n=59) a Strain designation / bei # Location Year Source -- 1391G 1392G 1393G Ninotsminda, 1979 Common vole 1412G Dalidag, 1979 Flea 1413G 1670G 1851G 1852G 1853G
Supplemental Table S1: pestis strains (n=59) a Strain designation / bei # Location Year Source -- 1391G 1392G 1393G Ninotsminda, 1979 Common vole 1412G Dalidag, 1979 Flea 1413G 1670G 1851G 1852G 1853G
Microbial Typing by Machine Learned DNA Melt Signatures
 Microbial Typing by Machine Learned DNA Melt Signatures Nadya Andini 1, Bo Wang 2, Pornpat Athamanolap 3, Justin Hardick 4, Billie J. Masek 5, Simone Thair 1, Annie Hu 1, Gideon Avornu 5, Stephen Peterson
Microbial Typing by Machine Learned DNA Melt Signatures Nadya Andini 1, Bo Wang 2, Pornpat Athamanolap 3, Justin Hardick 4, Billie J. Masek 5, Simone Thair 1, Annie Hu 1, Gideon Avornu 5, Stephen Peterson
10ml. Set (4 poly and 17 monovalent, 2ml each)
 120314TR Bordetella pertussis Antigen 10ml Contents 1 Bordetella pertussis Antigen 1 Clostridium perfringens Type A 2 Escherichia coli 5 Legionella pneumophila 5 Listeria monocytogenes 6 Pseudomonas aeruginosa
120314TR Bordetella pertussis Antigen 10ml Contents 1 Bordetella pertussis Antigen 1 Clostridium perfringens Type A 2 Escherichia coli 5 Legionella pneumophila 5 Listeria monocytogenes 6 Pseudomonas aeruginosa
Product Catalogue 2015 Clinical and Industrial Microbiology
 Acinetobacter baumannii ATCC BAA-747 * 89141 Actinomyces odontolyticus ATCC 17929 * 89114 Aeromonas hydrophila ATCC 7966 * 89119 Aggregatibacter aphrophilus ATCC 7901 * 89091 Aspergillus brasiliensis ATCC
Acinetobacter baumannii ATCC BAA-747 * 89141 Actinomyces odontolyticus ATCC 17929 * 89114 Aeromonas hydrophila ATCC 7966 * 89119 Aggregatibacter aphrophilus ATCC 7901 * 89091 Aspergillus brasiliensis ATCC
INTERPRETATION OF THE GRAM STAIN
 INTERPRETATION OF THE GRAM STAIN DISCLOSURE Relevant relationships with commercial entities none Potential for conflicts of interest within this presentation none Steps taken to review and mitigate potential
INTERPRETATION OF THE GRAM STAIN DISCLOSURE Relevant relationships with commercial entities none Potential for conflicts of interest within this presentation none Steps taken to review and mitigate potential
Product Catalogue 2016 Clinical and Industrial Microbiology
 Acinetobacter baumannii ATCC BAA-747 * 89141 Acinetobacter baumannii ATCC 19606 * 89174 Actinomyces odontolyticus ATCC 17929 * 89114 Aeromonas hydrophila ATCC 7966 * 89119 Aeromonas hydrophila ATCC 35654
Acinetobacter baumannii ATCC BAA-747 * 89141 Acinetobacter baumannii ATCC 19606 * 89174 Actinomyces odontolyticus ATCC 17929 * 89114 Aeromonas hydrophila ATCC 7966 * 89119 Aeromonas hydrophila ATCC 35654
Saturday (18 Aug 2007)
 E rrl Center for Biological Sequence analysis Comparative icrobial Genomics group BLAST atlases O r i g i 0 rrlc 4 0.5 rrla 3.5 4,641,433 bp rrld rrlb 1.5 3 2 2.5 Te r mi Dave Ussery nu 4th annual workshop
E rrl Center for Biological Sequence analysis Comparative icrobial Genomics group BLAST atlases O r i g i 0 rrlc 4 0.5 rrla 3.5 4,641,433 bp rrld rrlb 1.5 3 2 2.5 Te r mi Dave Ussery nu 4th annual workshop
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss
 Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
D SWATI. In silico comparison of bacterial strains using mutual information 1185
 In silico comparison of bacterial strains using mutual information In silico comparison of bacterial strains using mutual information 1185 D SWATI, 1169 1184 Indian Academy of Sciences Supplementary Material
In silico comparison of bacterial strains using mutual information In silico comparison of bacterial strains using mutual information 1185 D SWATI, 1169 1184 Indian Academy of Sciences Supplementary Material
Whole Genome based Phylogeny
 Whole Genome based Phylogeny Johanne Ahrenfeldt PhD student DTU Bioinformatics Short about me Johanne Ahrenfeldt johah@dtu.dk PhD student at DTU Bioinformatics Whole Genome based Phylogeny Graduate Engineer
Whole Genome based Phylogeny Johanne Ahrenfeldt PhD student DTU Bioinformatics Short about me Johanne Ahrenfeldt johah@dtu.dk PhD student at DTU Bioinformatics Whole Genome based Phylogeny Graduate Engineer
EZ-COMP EZ-COMP For Training and Proficiency Testing Product Details
 EZ-COMP For Training and Proficiency Testing Mixed microorganism populations Identified by codes rather than descriptions Refrigerated storage Traceable to reference culture Product warranty Product Details
EZ-COMP For Training and Proficiency Testing Mixed microorganism populations Identified by codes rather than descriptions Refrigerated storage Traceable to reference culture Product warranty Product Details
Inferring positional homologs with common intervals of sequences
 Outline Introduction Our approach Results Conclusion Inferring positional homologs with common intervals of sequences Guillaume Blin, Annie Chateau, Cedric Chauve, Yannick Gingras CGL - Université du Québec
Outline Introduction Our approach Results Conclusion Inferring positional homologs with common intervals of sequences Guillaume Blin, Annie Chateau, Cedric Chauve, Yannick Gingras CGL - Université du Québec
Essentiality in B. subtilis
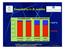 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Pan- and core- Genomics
 Pan- and core- Genomics Dave Ussery Workshop on Comparative Genomics King Mongkut's University of Technology Thonburi Bangkok, Thailand First Talk for Wednesday 22 June, 2011 Center for Biological Sequence
Pan- and core- Genomics Dave Ussery Workshop on Comparative Genomics King Mongkut's University of Technology Thonburi Bangkok, Thailand First Talk for Wednesday 22 June, 2011 Center for Biological Sequence
Database and Comparative Identification of Prophages
 Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy
Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy
Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana
 Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana Table S1 The species marked with belong to the Proteobacteria subset and those marked
Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana Table S1 The species marked with belong to the Proteobacteria subset and those marked
SENCA: A codon substitution model to better estimate evolutionary processes
 SENCA: A codon substitution model to better estimate evolutionary processes Fanny Pouyet Marc Bailly-Bechet, Dominique Mouchiroud and Laurent Guéguen LBBE - UCB Lyon - UMR 5558 June 2015, Porquerolles,
SENCA: A codon substitution model to better estimate evolutionary processes Fanny Pouyet Marc Bailly-Bechet, Dominique Mouchiroud and Laurent Guéguen LBBE - UCB Lyon - UMR 5558 June 2015, Porquerolles,
Pan-genome analysis provides much higher strain typing resolution than multi-locus sequence typing
 Microbiology (2010), 156, 1060 1068 DOI 10.1099/mic.0.035188-0 Pan-genome analysis provides much higher strain typing resolution than multi-locus sequence typing Barry G. Hall, 1,2 Garth D. Ehrlich 2,3
Microbiology (2010), 156, 1060 1068 DOI 10.1099/mic.0.035188-0 Pan-genome analysis provides much higher strain typing resolution than multi-locus sequence typing Barry G. Hall, 1,2 Garth D. Ehrlich 2,3
Bacterial phylogenetic tree construction based on genomic translation stop signals
 Xu et al. Microbial Informatics and Experimentation 2012, 2:6 RESEARCH Open Access Bacterial phylogenetic tree construction based on genomic translation stop signals Lijing Xu 1, Jimmy Kuo 2, Jong-Kang
Xu et al. Microbial Informatics and Experimentation 2012, 2:6 RESEARCH Open Access Bacterial phylogenetic tree construction based on genomic translation stop signals Lijing Xu 1, Jimmy Kuo 2, Jong-Kang
Opening the pan-genomics box for E. coli
 Opening the pan-genomics box for E. coli Comparison of 20 sequenced genomes Dave Ussery Workshop on Comparative Microbial Genomics Oslo, Norway 19 October, 2006 Dave Ussery Genomics of pathogenic : European
Opening the pan-genomics box for E. coli Comparison of 20 sequenced genomes Dave Ussery Workshop on Comparative Microbial Genomics Oslo, Norway 19 October, 2006 Dave Ussery Genomics of pathogenic : European
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Genetic Basis of Variation in Bacteria
 Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
The impact of the neisserial DNA uptake sequence on genome evolution and stability
 The impact of the neisserial DNA uptake sequence on genome evolution and stability Ole Herman Ambur The Tønjum Group Transformation Neisserial transformation requires: DNA uptake sequence (DUS) Transformation
The impact of the neisserial DNA uptake sequence on genome evolution and stability Ole Herman Ambur The Tønjum Group Transformation Neisserial transformation requires: DNA uptake sequence (DUS) Transformation
Design of an Enterobacteriaceae Pan-genome Microarray Chip
 Design of an Enterobacteriaceae Pan-genome Microarray Chip Oksana Lukjancenko and David W. Ussery DTU CBS 2010 2 Background Pan-genome complete collection of variuos genes located within populations at
Design of an Enterobacteriaceae Pan-genome Microarray Chip Oksana Lukjancenko and David W. Ussery DTU CBS 2010 2 Background Pan-genome complete collection of variuos genes located within populations at
Bacterial pan-genomics
 Bacterial pan-genomics Dave Ussery 4 th annual workshop on Comparative Microbial Genomics and Taxonomy Petropolois, Brazil Lecture #3 Wednesday, 5 August, 2009 What we DO know What we DON'T know Outline
Bacterial pan-genomics Dave Ussery 4 th annual workshop on Comparative Microbial Genomics and Taxonomy Petropolois, Brazil Lecture #3 Wednesday, 5 August, 2009 What we DO know What we DON'T know Outline
Understanding microbes through the lens of comparative genomics... a biased perspective. Aaron Darling A/Prof. ithree institute UTS
 Understanding microbes through the lens of comparative genomics... a biased perspective Aaron Darling A/Prof. ithree institute UTS Molecular Evolution of Bacteria Bacteria reproduce clonally A brief history
Understanding microbes through the lens of comparative genomics... a biased perspective Aaron Darling A/Prof. ithree institute UTS Molecular Evolution of Bacteria Bacteria reproduce clonally A brief history
MICROBIAL BIOCHEMISTRY BIOT 309. Dr. Leslye Johnson Sept. 30, 2012
 MICROBIAL BIOCHEMISTRY BIOT 309 Dr. Leslye Johnson Sept. 30, 2012 Phylogeny study of evoluhonary relatedness among groups of organisms (e.g. species, populahons), which is discovered through molecular
MICROBIAL BIOCHEMISTRY BIOT 309 Dr. Leslye Johnson Sept. 30, 2012 Phylogeny study of evoluhonary relatedness among groups of organisms (e.g. species, populahons), which is discovered through molecular
Product List. - Kits & Reagents
 Product List - Kits & Reagents Bacterial Pathogens Viral Pathogens Beverage Spoilage Organisms Yeast and Mold GMOs Allergens Animal Identification Veterinary Environmental Pharma DNA/RNA Extraction Additional
Product List - Kits & Reagents Bacterial Pathogens Viral Pathogens Beverage Spoilage Organisms Yeast and Mold GMOs Allergens Animal Identification Veterinary Environmental Pharma DNA/RNA Extraction Additional
Structural insights into bacterial flagellar hooks similarities and specificities
 Supplementary information Structural insights into bacterial flagellar hooks similarities and specificities Young-Ho Yoon, Clive S. Barker, Paula V. Bulieris, Hideyuki Matsunami, Fadel A. Samatey* Affiliation:
Supplementary information Structural insights into bacterial flagellar hooks similarities and specificities Young-Ho Yoon, Clive S. Barker, Paula V. Bulieris, Hideyuki Matsunami, Fadel A. Samatey* Affiliation:
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett. Introduction
 The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
Evolutionary Analysis by Whole-Genome Comparisons
 JOURNAL OF BACTERIOLOGY, Apr. 2002, p. 2260 2272 Vol. 184, No. 8 0021-9193/02/$04.00 0 DOI: 184.8.2260 2272.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Evolutionary Analysis
JOURNAL OF BACTERIOLOGY, Apr. 2002, p. 2260 2272 Vol. 184, No. 8 0021-9193/02/$04.00 0 DOI: 184.8.2260 2272.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Evolutionary Analysis
Evidence That Mutation Is Universally Biased towards AT in Bacteria
 Evidence That Mutation Is Universally Biased towards AT in Bacteria Ruth Hershberg*, Dmitri A. Petrov Department of Biology, Stanford University, Stanford, California, United States of America Abstract
Evidence That Mutation Is Universally Biased towards AT in Bacteria Ruth Hershberg*, Dmitri A. Petrov Department of Biology, Stanford University, Stanford, California, United States of America Abstract
Species Identification and Strain Attribution with Unassembled Sequencing Data
 Brigham Young University BYU ScholarsArchive All Theses and Dissertations 2012-04-18 Species Identification and Strain Attribution with Unassembled Sequencing Data Owen Eric Francis Brigham Young University
Brigham Young University BYU ScholarsArchive All Theses and Dissertations 2012-04-18 Species Identification and Strain Attribution with Unassembled Sequencing Data Owen Eric Francis Brigham Young University
Genomic insights into the taxonomic status of the Bacillus cereus group. Laboratory of Marine Genetic Resources, Third Institute of Oceanography,
 1 2 3 Genomic insights into the taxonomic status of the Bacillus cereus group Yang Liu 1, Qiliang Lai 1, Markus Göker 2, Jan P. Meier-Kolthoff 2, Meng Wang 3, Yamin Sun 3, Lei Wang 3 and Zongze Shao 1*
1 2 3 Genomic insights into the taxonomic status of the Bacillus cereus group Yang Liu 1, Qiliang Lai 1, Markus Göker 2, Jan P. Meier-Kolthoff 2, Meng Wang 3, Yamin Sun 3, Lei Wang 3 and Zongze Shao 1*
colony size color morphology haemolysis S. aureus S. epidermidis
 practical 2.: STAPHYLOCOCCUS 1. Prepare a heat fixed smear of the culture of S.aureus. (Gram staining, microscopy). 2. Prepare a heat fixed smear of the culture of S.aureus. and S.epidermidis (mixed smear),
practical 2.: STAPHYLOCOCCUS 1. Prepare a heat fixed smear of the culture of S.aureus. (Gram staining, microscopy). 2. Prepare a heat fixed smear of the culture of S.aureus. and S.epidermidis (mixed smear),
Identification of Bacteria Using Phylogenetic Relationships Revealed by MS/MS Sequencing of Tryptic Peptides Derived from Cellular Proteins
 Identification of Bacteria Using Phylogenetic Relationships Revealed by MS/MS Sequencing of Tryptic Peptides Derived from Cellular Proteins Jacek P. Dworzanski Geo-Centers, Inc., Aberdeen Proving Ground,
Identification of Bacteria Using Phylogenetic Relationships Revealed by MS/MS Sequencing of Tryptic Peptides Derived from Cellular Proteins Jacek P. Dworzanski Geo-Centers, Inc., Aberdeen Proving Ground,
The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome
 Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
7 Multiple Genome Alignment
 94 Bioinformatics I, WS /3, D. Huson, December 3, 0 7 Multiple Genome Alignment Assume we have a set of genomes G,..., G t that we want to align with each other. If they are short and very closely related,
94 Bioinformatics I, WS /3, D. Huson, December 3, 0 7 Multiple Genome Alignment Assume we have a set of genomes G,..., G t that we want to align with each other. If they are short and very closely related,
Product List. - Kits & Reagents
 Product List - Kits & Reagents Bacterial Pathogens Viral Pathogens Beverage Spoilage Organisms Yeast and Mold GMOs Allergens Animal Identification Veterinary Environmental Pharma DNA/RNA Extraction Additional
Product List - Kits & Reagents Bacterial Pathogens Viral Pathogens Beverage Spoilage Organisms Yeast and Mold GMOs Allergens Animal Identification Veterinary Environmental Pharma DNA/RNA Extraction Additional
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
A L A BA M A L A W R E V IE W
 A L A BA M A L A W R E V IE W Volume 52 Fall 2000 Number 1 B E F O R E D I S A B I L I T Y C I V I L R I G HT S : C I V I L W A R P E N S I O N S A N D TH E P O L I T I C S O F D I S A B I L I T Y I N
A L A BA M A L A W R E V IE W Volume 52 Fall 2000 Number 1 B E F O R E D I S A B I L I T Y C I V I L R I G HT S : C I V I L W A R P E N S I O N S A N D TH E P O L I T I C S O F D I S A B I L I T Y I N
Our product offering. The ATCC Licensed Derivative Program
 Our product offering Currently MECCONTI offers three different product lines of lyophilized micro-organisms: MicroSwabs: A MicroSwab consists of a lyophilized pellet of a single micro-organism strain inside
Our product offering Currently MECCONTI offers three different product lines of lyophilized micro-organisms: MicroSwabs: A MicroSwab consists of a lyophilized pellet of a single micro-organism strain inside
AUTOREGRESSIVE MODELING OF CODING SEQUENCE LENGTHS IN BACTERIAL GENOME
 Fluctuation and oise Letters Vol. 0, o. 0 (0000) 000 000 World Scientific Publishing Company AUTOREGRESSIVE MODELIG OF CODIG SEQUECE LEGTHS I BACTERIAL GEOME VASILE V. MORARIU 1,, LUIZA BUIMAGA-IARICA
Fluctuation and oise Letters Vol. 0, o. 0 (0000) 000 000 World Scientific Publishing Company AUTOREGRESSIVE MODELIG OF CODIG SEQUECE LEGTHS I BACTERIAL GEOME VASILE V. MORARIU 1,, LUIZA BUIMAGA-IARICA
CATALOGUE OF TEST, CONTROL OR BIOASSAY STRAINS FROM BCCM CULTURE COLLECTIONS
 CATALOGUE OF TEST, CONTROL OR BIOASSAY STRAINS FROM BCCM CULTURE COLLECTIONS LIST OF STRAINS INVOLVED IN NORMS AND STANDARDISED MICROBIOLOGICAL METHODS The Belgian Co-ordinated Collections of Micro-organisms
CATALOGUE OF TEST, CONTROL OR BIOASSAY STRAINS FROM BCCM CULTURE COLLECTIONS LIST OF STRAINS INVOLVED IN NORMS AND STANDARDISED MICROBIOLOGICAL METHODS The Belgian Co-ordinated Collections of Micro-organisms
The use of gene clusters to infer functional coupling
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2896 2901, March 1999 Genetics The use of gene clusters to infer functional coupling ROSS OVERBEEK*, MICHAEL FONSTEIN, MARK D SOUZA*, GORDON D. PUSCH*, AND NATALIA
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2896 2901, March 1999 Genetics The use of gene clusters to infer functional coupling ROSS OVERBEEK*, MICHAEL FONSTEIN, MARK D SOUZA*, GORDON D. PUSCH*, AND NATALIA
Outline. I. Methods. II. Preliminary Results. A. Phylogeny Methods B. Whole Genome Methods C. Horizontal Gene Transfer
 Comparative Genomics Preliminary Results April 4, 2016 Juan Castro, Aroon Chande, Cheng Chen, Evan Clayton, Hector Espitia, Alli Gombolay, Walker Gussler, Ken Lee, Tyrone Lee, Hari Prasanna, Carlos Ruiz,
Comparative Genomics Preliminary Results April 4, 2016 Juan Castro, Aroon Chande, Cheng Chen, Evan Clayton, Hector Espitia, Alli Gombolay, Walker Gussler, Ken Lee, Tyrone Lee, Hari Prasanna, Carlos Ruiz,
T h e C S E T I P r o j e c t
 T h e P r o j e c t T H E P R O J E C T T A B L E O F C O N T E N T S A r t i c l e P a g e C o m p r e h e n s i v e A s s es s m e n t o f t h e U F O / E T I P h e n o m e n o n M a y 1 9 9 1 1 E T
T h e P r o j e c t T H E P R O J E C T T A B L E O F C O N T E N T S A r t i c l e P a g e C o m p r e h e n s i v e A s s es s m e n t o f t h e U F O / E T I P h e n o m e n o n M a y 1 9 9 1 1 E T
Comprehensive Phylogenetic Analysis of Mycobacteria
 Preprints of the 12th IFAC Symposium on Computer Applications in Biotechnology The International Federation of Automatic Control 16-18, 2013, December. Mumbai, India Comprehensive Phylogenetic Analysis
Preprints of the 12th IFAC Symposium on Computer Applications in Biotechnology The International Federation of Automatic Control 16-18, 2013, December. Mumbai, India Comprehensive Phylogenetic Analysis
The Prokaryotes: Domains Bacteria and Archaea
 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 11 The Prokaryotes: Domains Bacteria and Archaea Table 11.1 Classification of Selected Prokaryotes*
PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 11 The Prokaryotes: Domains Bacteria and Archaea Table 11.1 Classification of Selected Prokaryotes*
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
GENE content variation as a key feature of bacterial
 Copyright Ó 2 by the Genetics Society of America DOI:.534/genetics..8448 Inferring Bacterial Genome Flux While Considering Truncated Genes Weilong Hao*, and G. Brian Golding *Department of Biology, Indiana
Copyright Ó 2 by the Genetics Society of America DOI:.534/genetics..8448 Inferring Bacterial Genome Flux While Considering Truncated Genes Weilong Hao*, and G. Brian Golding *Department of Biology, Indiana
176 5 t h Fl oo r. 337 P o ly me r Ma te ri al s
 A g la di ou s F. L. 462 E l ec tr on ic D ev el op me nt A i ng er A.W.S. 371 C. A. M. A l ex an de r 236 A d mi ni st ra ti on R. H. (M rs ) A n dr ew s P. V. 326 O p ti ca l Tr an sm is si on A p ps
A g la di ou s F. L. 462 E l ec tr on ic D ev el op me nt A i ng er A.W.S. 371 C. A. M. A l ex an de r 236 A d mi ni st ra ti on R. H. (M rs ) A n dr ew s P. V. 326 O p ti ca l Tr an sm is si on A p ps
Correlations between Shine-Dalgarno Sequences and Gene Features Such as Predicted Expression Levels and Operon Structures
 JOURNAL OF BACTERIOLOGY, Oct. 2002, p. 5733 5745 Vol. 184, No. 20 0021-9193/02/$04.00 0 DOI: 10.1128/JB.184.20.5733 5745.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Correlations
JOURNAL OF BACTERIOLOGY, Oct. 2002, p. 5733 5745 Vol. 184, No. 20 0021-9193/02/$04.00 0 DOI: 10.1128/JB.184.20.5733 5745.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Correlations
By signing below, you acknowledge that you have ensured that you are complying with the above statement.
 Instructions: This exam consists of 31 multiple choice questions on 8 pages, including this one. Please submit your answers on the scantron sheet provided and on this copy of the exam. This exam is closed
Instructions: This exam consists of 31 multiple choice questions on 8 pages, including this one. Please submit your answers on the scantron sheet provided and on this copy of the exam. This exam is closed
Microbial Bioinformatics
 1 Microbial Bioinformatics Introduction These exercises are for you to learn how to use bioinformatics tools to explore bacterial genomes, and complements the Microbiology of Safe Food 2 nd Edition (Wiley-
1 Microbial Bioinformatics Introduction These exercises are for you to learn how to use bioinformatics tools to explore bacterial genomes, and complements the Microbiology of Safe Food 2 nd Edition (Wiley-
Tetracycline Rationale for the EUCAST clinical breakpoints, version th November 2009
 Tetracycline Rationale for the EUCAST clinical breakpoints, version 1.0 20 th November 2009 Introduction The natural tetracyclines, including tetracycline, chlortetracycline, oxytetracycline and demethylchlortetracycline
Tetracycline Rationale for the EUCAST clinical breakpoints, version 1.0 20 th November 2009 Introduction The natural tetracyclines, including tetracycline, chlortetracycline, oxytetracycline and demethylchlortetracycline
Retail Catalog 39th Edition Supplemental Information Individual Microorganisms Price Microorganism Description
 LYFO DISK KWIK-STIK Vial 6 Pk Pk Individual Microorganisms Price Microorganism Description ode BSL omment 049L 049K 049P 0645L 0645K 0645P Aggregatibacter aphrophilus AT 4997 D formerly Haemophilus paraphrophilus
LYFO DISK KWIK-STIK Vial 6 Pk Pk Individual Microorganisms Price Microorganism Description ode BSL omment 049L 049K 049P 0645L 0645K 0645P Aggregatibacter aphrophilus AT 4997 D formerly Haemophilus paraphrophilus
Introduction to polyphasic taxonomy
 Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Toronto General Hospital ANTIBIOGRAM Emergency Department January 1, December 31, 2016
 IV (meningitis) IV (non-meningitis) (meningitis) (non-meningitis) Blood Isolates % Susceptible 644 18 36 70 78 74 59 69 75 262 100 19 64 75 100 92 54 72 78 76 68 89 86 99 Escherichia coli 153 58 30 67
IV (meningitis) IV (non-meningitis) (meningitis) (non-meningitis) Blood Isolates % Susceptible 644 18 36 70 78 74 59 69 75 262 100 19 64 75 100 92 54 72 78 76 68 89 86 99 Escherichia coli 153 58 30 67
INTRODUCTION MATERIALS & METHODS
 Evaluation of Three Bacterial Transport Systems, The New Copan M40 Transystem, Remel Bactiswab And Medical Wire & Equipment Transwab, for Maintenance of Aerobic Fastidious and Non-Fastidious Organisms
Evaluation of Three Bacterial Transport Systems, The New Copan M40 Transystem, Remel Bactiswab And Medical Wire & Equipment Transwab, for Maintenance of Aerobic Fastidious and Non-Fastidious Organisms
11/5/2018. Update on Modern Bacterial Taxonomy for Bench Microbiologists. Why is Taxonomy Important? Bacterial Taxonomy for Clinical Microbiologists
 Update on Modern Bacterial Taxonomy for Bench Microbiologists J. Michael Janda Kern County Public Health Laboratory Bakersfield CA The Name Game Which Ones Different? Why is Taxonomy Important? Bacterial
Update on Modern Bacterial Taxonomy for Bench Microbiologists J. Michael Janda Kern County Public Health Laboratory Bakersfield CA The Name Game Which Ones Different? Why is Taxonomy Important? Bacterial
Fitness constraints on horizontal gene transfer
 Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Overview of the major bacterial pathogens The major bacterial pathogens are presented in this table:
 Practical Microbiology 30/11/2018 University of Sulaimani college of Pharmacy Year2 Lab. 5: Overview of the major bacterial pathogens The major bacterial pathogens are presented in this table: Major Bacterial
Practical Microbiology 30/11/2018 University of Sulaimani college of Pharmacy Year2 Lab. 5: Overview of the major bacterial pathogens The major bacterial pathogens are presented in this table: Major Bacterial
Gram negative bacilli
 Gram negative bacilli 1-Enterobacteriaceae Gram negative bacilli-rods Enterobacteriaceae Are everywhere Part of normal flora of humans and most animals They are cause of -30-35% septisemia -more than 70%
Gram negative bacilli 1-Enterobacteriaceae Gram negative bacilli-rods Enterobacteriaceae Are everywhere Part of normal flora of humans and most animals They are cause of -30-35% septisemia -more than 70%
WALTER REED ARMY INSTITUTE OF RESEARCH
 WALTER REED ARMY INSTITUTE OF RESEARCH BASIC CLINICAL MICROBIOLOGY MAJ Tom Palys Ph.D Walter Reed Army Institute of Research OUTLINE Introduction to Basic Clinical Microbiology Didactic Plate Rounds Practical
WALTER REED ARMY INSTITUTE OF RESEARCH BASIC CLINICAL MICROBIOLOGY MAJ Tom Palys Ph.D Walter Reed Army Institute of Research OUTLINE Introduction to Basic Clinical Microbiology Didactic Plate Rounds Practical
2 Genome evolution: gene fusion versus gene fission
 2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria)
 Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria) (don t study Concept 27.2) Some phyla Remember: Bacterial cell structure and shapes 1 Usually very small but some are unusually
Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria) (don t study Concept 27.2) Some phyla Remember: Bacterial cell structure and shapes 1 Usually very small but some are unusually
Microbiological Testing Summary
 BACTERICIDAL ACTIVITY EN 1276 - Chemical disinfectants and antiseptics - Quantitative suspension test for the evaluation of bactericidal activity of chemical disinfectants and antiseptics used in food,
BACTERICIDAL ACTIVITY EN 1276 - Chemical disinfectants and antiseptics - Quantitative suspension test for the evaluation of bactericidal activity of chemical disinfectants and antiseptics used in food,
Quality Control Solutions for Food Testing
 Quality Control Solutions for Food Testing QUALITY CONTROL SOLUTIONS FOR FOOD TESTING ATCC offers an expanding portfolio of quality control products to ensure the accuracy and reliability of your food
Quality Control Solutions for Food Testing QUALITY CONTROL SOLUTIONS FOR FOOD TESTING ATCC offers an expanding portfolio of quality control products to ensure the accuracy and reliability of your food
Most common dose (mg) 1g x 1 1g x 1 1g x 1 1g x 1 1g x 1 1g x 1. Maximum dose schedule (mg) 1g x 1 1g x 1 1g x 1 1g x 1 1g x 1 1g x 1
 Ertapenem Rationale for the EUCAST clinical breakpoints, version 1.3 1 st June 2009 Introduction Ertapenem is a carbapenem, available only for parenteral use. Ertapenem is relevant for therapy of septicaemia,
Ertapenem Rationale for the EUCAST clinical breakpoints, version 1.3 1 st June 2009 Introduction Ertapenem is a carbapenem, available only for parenteral use. Ertapenem is relevant for therapy of septicaemia,
What is Microbiology?
 Microbiology What is Microbiology? Microbiology is the Science that studies Microorganisms. Microorganisms, roughly, are those living things that are too small to be seen with the naked eye. Microorganisms
Microbiology What is Microbiology? Microbiology is the Science that studies Microorganisms. Microorganisms, roughly, are those living things that are too small to be seen with the naked eye. Microorganisms
D-10 VALUES OF MICRO-ORGANISMS
 INFORMATION SHEET SYNERGY HEALTH D-10 VALUES OF MICRO-ORGANISMS Synergy Health Applied Sterilisation Technologies (AST) operates as a global supplier of sterilisation, specialist applications and contamination
INFORMATION SHEET SYNERGY HEALTH D-10 VALUES OF MICRO-ORGANISMS Synergy Health Applied Sterilisation Technologies (AST) operates as a global supplier of sterilisation, specialist applications and contamination
The Evolution of Infectious Disease
 The Evolution of Infectious Disease Why are some bacteria pathogenic to humans while other (closely-related) bacteria are not? This question can be approached from two directions: 1.From the point of view
The Evolution of Infectious Disease Why are some bacteria pathogenic to humans while other (closely-related) bacteria are not? This question can be approached from two directions: 1.From the point of view
Survey of plasmid profiles of Shigella species isolated in Malaysia during
 World Journal of Microbiology & Biotechnology (2005) 21: 271 278 Ó Springer 2005 DOI 10.1007/s11274-004-3631-0 Survey of plasmid profiles of Shigella species isolated in Malaysia during 1994 2000 C.H.
World Journal of Microbiology & Biotechnology (2005) 21: 271 278 Ó Springer 2005 DOI 10.1007/s11274-004-3631-0 Survey of plasmid profiles of Shigella species isolated in Malaysia during 1994 2000 C.H.
P a g e 3 6 of R e p o r t P B 4 / 0 9
 P a g e 3 6 of R e p o r t P B 4 / 0 9 p r o t e c t h um a n h e a l t h a n d p r o p e r t y fr om t h e d a n g e rs i n h e r e n t i n m i n i n g o p e r a t i o n s s u c h a s a q u a r r y. J
P a g e 3 6 of R e p o r t P B 4 / 0 9 p r o t e c t h um a n h e a l t h a n d p r o p e r t y fr om t h e d a n g e rs i n h e r e n t i n m i n i n g o p e r a t i o n s s u c h a s a q u a r r y. J
CHEM 10113, Quiz 5 October 26, 2011
 CHEM 10113, Quiz 5 October 26, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
CHEM 10113, Quiz 5 October 26, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
Microbial Taxonomy. Classification of living organisms into groups. A group or level of classification
 Lec 2 Oral Microbiology Dr. Chatin Purpose Microbial Taxonomy Classification Systems provide an easy way grouping of diverse and huge numbers of microbes To provide an overview of how physicians think
Lec 2 Oral Microbiology Dr. Chatin Purpose Microbial Taxonomy Classification Systems provide an easy way grouping of diverse and huge numbers of microbes To provide an overview of how physicians think
Dr. habil. Anna Salek. Mikrobiologist Biotechnologist Research Associate
 Dr. habil. Anna Salek Mikrobiologist Biotechnologist Research Associate BIOTECHNOLOGY of Food Science Cell Biology of Microorganisms Physiology of Microorganisms Biochemistry of Microorganisms Molecularbiology
Dr. habil. Anna Salek Mikrobiologist Biotechnologist Research Associate BIOTECHNOLOGY of Food Science Cell Biology of Microorganisms Physiology of Microorganisms Biochemistry of Microorganisms Molecularbiology
8. Relax and do well.
 CHEM 1225 Exam III John III. Gelder April 8, 1999 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last two pages includes a periodic table and
CHEM 1225 Exam III John III. Gelder April 8, 1999 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last two pages includes a periodic table and
CONTROL ORGANISMS ss
 CONTROL ORGANISMS 032918ss CONTENTS CONTROL ORGANISMS 1 Lyophilized KWIK-STIK & LYFO DISK 29 Epower 32 Epower CRM 34 EZ-Accu Shot 36 EZ-Accu Shot Select 36 EZ-CFU 38 EZ-CFU One Step 41 EZ-Hydro Shot 40
CONTROL ORGANISMS 032918ss CONTENTS CONTROL ORGANISMS 1 Lyophilized KWIK-STIK & LYFO DISK 29 Epower 32 Epower CRM 34 EZ-Accu Shot 36 EZ-Accu Shot Select 36 EZ-CFU 38 EZ-CFU One Step 41 EZ-Hydro Shot 40
CHN62: REPORTING OF MICROBIOLOGY RESULTS
 CHN62: 1.1 Introduction This SOP provides guidance on reporting of all the microbiological results in the Microbiology laboratory for CHAIN study. 1.2 Purpose This SOP will aid in a standard reporting
CHN62: 1.1 Introduction This SOP provides guidance on reporting of all the microbiological results in the Microbiology laboratory for CHAIN study. 1.2 Purpose This SOP will aid in a standard reporting
HANDOUT SET GENERAL CHEMISTRY II
 HANDOUT SET GENERAL CHEMISTRY II Periodic Table of the Elements 1 2 3 4 5 6 7 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 IA VIIIA 1 2 H He 1.00794 IIA IIIA IVA VA VIA VIIA 4.00262 3 Li 6.941 11 Na 22.9898
HANDOUT SET GENERAL CHEMISTRY II Periodic Table of the Elements 1 2 3 4 5 6 7 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 IA VIIIA 1 2 H He 1.00794 IIA IIIA IVA VA VIA VIIA 4.00262 3 Li 6.941 11 Na 22.9898
arxiv: v1 [q-bio.qm] 18 Nov 2013
![arxiv: v1 [q-bio.qm] 18 Nov 2013 arxiv: v1 [q-bio.qm] 18 Nov 2013](/thumbs/72/66745314.jpg) Markovianness and Conditional Independence in Annotated Bacterial DNA Andrew Hart Servet Martínez arxiv:1311.4411v1 [q-bio.qm] 18 Nov 2013 November 19, 2013 Abstract We explore the probabilistic structure
Markovianness and Conditional Independence in Annotated Bacterial DNA Andrew Hart Servet Martínez arxiv:1311.4411v1 [q-bio.qm] 18 Nov 2013 November 19, 2013 Abstract We explore the probabilistic structure
An Inclusivity and Exclusivity Evaluation of the Invisible Sentinel Veriflow Salmonella Species Method for the Detection of Salmonella species
 An Inclusivity and Exclusivity Evaluation of the Invisible Sentinel Veriflow Salmonella Species for the Detection of Salmonella species AOAC PTM Evaluation September 20, 2013 Erin S. Crowley, Patrick M.
An Inclusivity and Exclusivity Evaluation of the Invisible Sentinel Veriflow Salmonella Species for the Detection of Salmonella species AOAC PTM Evaluation September 20, 2013 Erin S. Crowley, Patrick M.
8. Relax and do well.
 CHEM 1314.03 Exam I John I. Gelder September 25, 1997 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
CHEM 1314.03 Exam I John I. Gelder September 25, 1997 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
Perspective. Abundant Microbial Inorganic Polyphosphate, Poly P Kinase Are Underappreciated
 Abundant Microbial Inorganic Polyphosphate, Poly P Kinase Are Underappreciated Although some view poly P as a mere molecular fossil, the author thought that it plays a more prominent role in microorganisms
Abundant Microbial Inorganic Polyphosphate, Poly P Kinase Are Underappreciated Although some view poly P as a mere molecular fossil, the author thought that it plays a more prominent role in microorganisms
AP Bio Module 16: Bacterial Genetics and Operons, Student Learning Guide
 Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
UNCORRECTED PROOF. AUTHOR'S PROOF Microb Ecol DOI /s
 Microb Ecol DOI 10.1007/s00248-011-9880-1 1 3 MINIREVIEWS 2 4 The Salmonella enterica Pan-genome 5 Annika Jacobsen & Rene S. Hendriksen & 6 Frank M. Aaresturp & David W. Ussery & Carsten Friis Q2 Q3 7
Microb Ecol DOI 10.1007/s00248-011-9880-1 1 3 MINIREVIEWS 2 4 The Salmonella enterica Pan-genome 5 Annika Jacobsen & Rene S. Hendriksen & 6 Frank M. Aaresturp & David W. Ussery & Carsten Friis Q2 Q3 7
8. Relax and do well.
 CHEM 1215 Exam III John III. Gelder November 11, 1998 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last page includes a periodic table and
CHEM 1215 Exam III John III. Gelder November 11, 1998 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last page includes a periodic table and
HACCP: INTRODUCTION AND HAZARD ANALYSIS
 Food Hygiene HACCP: INTRODUCTION AND HAZARD ANALYSIS State: December 15, 2004 WPF 5/0 Important milestones in the development of food safety systems Time Activity Distant past Use of prohibition principle
Food Hygiene HACCP: INTRODUCTION AND HAZARD ANALYSIS State: December 15, 2004 WPF 5/0 Important milestones in the development of food safety systems Time Activity Distant past Use of prohibition principle
Practical examination
 Practical examination I. Sterile media 1. Bouillon, 2. Slant agar, tube agar 4. Enrichment media: meat bouillon 3., 5., 6.: Agar, blood agar and chocolate agar plates 7. Selective and differentiating media
Practical examination I. Sterile media 1. Bouillon, 2. Slant agar, tube agar 4. Enrichment media: meat bouillon 3., 5., 6.: Agar, blood agar and chocolate agar plates 7. Selective and differentiating media
Comparative Studies of Transcriptional Regulation Mechanisms in a Group of Eight Gamma-proteobacterial Genomes
 doi:10.1016/j.jmb.2005.09.037 J. Mol. Biol. (2005) 354, 184 199 Comparative Studies of Transcriptional Regulation Mechanisms in a Group of Eight Gamma-proteobacterial Genomes Vladimir Espinosa 1, Abel
doi:10.1016/j.jmb.2005.09.037 J. Mol. Biol. (2005) 354, 184 199 Comparative Studies of Transcriptional Regulation Mechanisms in a Group of Eight Gamma-proteobacterial Genomes Vladimir Espinosa 1, Abel
genomes The minimal set of conserved proteins across a thousand bacterial Center for Biological Sequence Analysis Department of Systems Biology
 The minimal set of conserved proteins across a thousand bacterial genomes Dave Ussery Comparative Bacterial Genomics Workshop Centers for Disease Control Atlanta, Georgia, USA Thursday, 3 August, 212 Center
The minimal set of conserved proteins across a thousand bacterial genomes Dave Ussery Comparative Bacterial Genomics Workshop Centers for Disease Control Atlanta, Georgia, USA Thursday, 3 August, 212 Center
Chem Exam 1. September 26, Dr. Susan E. Bates. Name 9:00 OR 10:00
 Chem 1711 Exam 1 September 26, 2013 Dr. Susan E. Bates Name 9:00 OR 10:00 N A = 6.022 x 10 23 mol 1 I A II A III B IV B V B VI B VII B VIII I B II B III A IV A V A VI A VII A inert gases 1 H 1.008 3 Li
Chem 1711 Exam 1 September 26, 2013 Dr. Susan E. Bates Name 9:00 OR 10:00 N A = 6.022 x 10 23 mol 1 I A II A III B IV B V B VI B VII B VIII I B II B III A IV A V A VI A VII A inert gases 1 H 1.008 3 Li
The bacterial pangenome as a new tool for analysing pathogenic bacteria
 ORIGINAL ARTICLE The bacterial pangenome as a new tool for analysing pathogenic bacteria L. Rouli, V. Merhej, P.-E. Fournier and D. Raoult Aix Marseille Université, URMITE, UM63, CNRS 7278, IRD 198, Inserm
ORIGINAL ARTICLE The bacterial pangenome as a new tool for analysing pathogenic bacteria L. Rouli, V. Merhej, P.-E. Fournier and D. Raoult Aix Marseille Université, URMITE, UM63, CNRS 7278, IRD 198, Inserm
World Journal of Pharmaceutical Research SJIF Impact Factor 8.074
 SJIF Impact Factor 8.074 Volume 7, Issue 5, 966-973. Research Article ISSN 2277 7105 MOLECULAR DETECTION OF ENTEROTOXIGENIC ISOLATES OF SALMONELLA TYPHIMURIUM, SHIGELLA FLEXNERI AND STAPHYLOCOCCUS AUREUS
SJIF Impact Factor 8.074 Volume 7, Issue 5, 966-973. Research Article ISSN 2277 7105 MOLECULAR DETECTION OF ENTEROTOXIGENIC ISOLATES OF SALMONELLA TYPHIMURIUM, SHIGELLA FLEXNERI AND STAPHYLOCOCCUS AUREUS
Software Process Models there are many process model s in th e li t e ra t u re, s om e a r e prescriptions and some are descriptions you need to mode
 Unit 2 : Software Process O b j ec t i ve This unit introduces software systems engineering through a discussion of software processes and their principal characteristics. In order to achieve the desireable
Unit 2 : Software Process O b j ec t i ve This unit introduces software systems engineering through a discussion of software processes and their principal characteristics. In order to achieve the desireable
Physical Chemistry I CHEM 4641 Final Exam 13 questions, 30 points
 Physical Chemistry I CHEM 4641 Final Exam 13 questions, 30 points Name: KEY Gas constant: R = 8.314 J mol -1 K -1 = 0.008314 kj mol -1 K -1. Boltzmann constant k = 1.381 10-23 J/K = 0.6950 cm -1 /K h =
Physical Chemistry I CHEM 4641 Final Exam 13 questions, 30 points Name: KEY Gas constant: R = 8.314 J mol -1 K -1 = 0.008314 kj mol -1 K -1. Boltzmann constant k = 1.381 10-23 J/K = 0.6950 cm -1 /K h =
