7 Multiple Genome Alignment
|
|
|
- Barry Carpenter
- 5 years ago
- Views:
Transcription
1 94 Bioinformatics I, WS /3, D. Huson, December 3, 0 7 Multiple Genome Alignment Assume we have a set of genomes G,..., G t that we want to align with each other. If they are short and very closely related, then one can use a multiple sequence alignment tool such as ClustalW to align them. Example: Generally, genomes will differ by: G A C C A A T T A C C T T - - A T C C C T A C - - T A A A C T subsitutions, insertions or deletions of bases, but also: replacements, insertions, deletions, duplications, translocations or inversions of whole segments of the sequence. If differences of the latter type occur, then simple multiple sequence alignment is not sufficient and a method is required that explicitly takes such large-scale rearrangments into account. The typical situation looks like this: Genome A B C D Genome A' C' B' D' Genome 3 C'' B'' E D'' 7. Multiple Alignment of Rearranged Sequences In the comparison of multiple genomes, we must solve the following computational problem: Multiple Alignment of Rearranged Genomic Sequences Problem: Input: Genome sequences G,..., G t. Output: Set of conserved segments and alignments between them. 7. Mauve Mauve is a widely used tool that addresses the stated problem. Here is an overview: A. Darling et al., Genome Research (004)
2 Bioinformatics I, WS /3, D. Huson, December 3, 0 95 (a) Quickly find long stretches of sequences that match two or more genomes, these are called anchors. Example: anchor G (b) Form a set of locally colinear blocks (LCBs) from the anchors. G LCB LCB LCB 3 3 LCB (c) Recursively try to improve LCBs. (d) Align the set of sequences in each LCB. 7.. MUMs and Multi MUMs Consider two genome sequences G and. Recall that a MUM (maximal unique match) is a subsequence that occurs exactly once in both G and and cannot be extended. A MUM can be written as M = (L, p, p ), where L is the length and p i is the left most position in G i. If the match is in the reverse complement of G i, then we use p i to denote its position in G i. MUMs can be computed using a suffix tree. Consider genomes G,..., G t. A multi-mum is a subsequence present in t genomes, in each contained exactly once and not extentable. It is written as M = (l, p, p,..., p t ), where l = L(M) is the length of the subsequence and p i = p i (M) is it s position in the i-th genome. Set p i = 0 to indicate that i-th genome is not involved in M. The multiplicity of M is d(m) = {i p i 0}, the number of genomes involved in M. Mauve uses multi-mums as anchors. 7.. Computation of multi-mums Multi-MUMs can be computed using a suffix tree (assignment). However, Mauve uses a simple seed-and-extend approach to find all multi-mums of length k, as follows:
3 96 Bioinformatics I, WS /3, D. Huson, December 3, 0 (a) Generate a hash table that maps each k-mer to a list of its occurrences in genomes G,..., G t. (b) For each k-mer that maps to two or more different genomes, extend all corresponding matches to the left and right until the first mismatch is found: k-mer match A Extend T A T C A The runtime of this approach is reported as: O(t n + tn log tn), where t is the number of genomes and n is their average length Anchors in Mauve The set of anchors used by Mauve is initially the set m of all multi-matches of full multiplicity t Determining LCBs Mauve uses breakpoint analysis to partition M into a set of L of LCBs, where each LCB consists of a set of matches (multi-mums) M,..., M f with p i (M j ) p i (M j+ ) i =,..., t and j =,..., f. G M M M 3 M M M 3 or To improve the set of LCBs, repeat the following until all remaining LCBs have a given minimum weight w: Let B be an LCB of smallest weight. Remove all matches in B from M. Recompute the set of LCBs from M. Here, the weight of LCB is the number of nucleotides that its matches cover. Example: G 3 3
4 Bioinformatics I, WS /3, D. Huson, December 3, 0 97 If the weight of LCB- is w then remove all matches of LCB- from M and then recalculate the set of LCBs: G G G3 Second illustration of block removal: 7..5 Multiple sequence alignment Once the set L of LCBs has been calculated, the last step is to compute a MSA for each LCB. For each LCB, Mauve computes a progressive alignment in three steps:. Pairwise alignment of all sequences to get a distance matrix D.. Compute a guide tree from D. 3. Progressively build a MSA by pairwise alignment of profiles along the guide tree. Step () is the most time-consuming operation in any progressive alignment approach. To avoid repeately performing steps () and (), Mauve computes a simple guide tree from the set of multimums by applying Neighbor-Joining to the following distances: d(gi, Gj ) = # bases shared by Gi and Gj Gi + Gj, based on the amount of identical sequence shared. This tree is used as guide tree for all LCB alignments.
5 98 Bioinformatics I, WS /3, D. Huson, December 3, Bacterial Example The Mauve paper presents the analysis of nine enterobacterial genomes, namely four E.coli, two Shigella and three Salmonella genomes : Species Genome size Reference E. coli K MG655 4,639, Blattner et al. 997 E. coli O57:H7 EDL933 5,54,97 Perna et al. 00 E. coli O57:H7 VT- Saka 5,498,450 Hayashi et al. 00 E. coli CFT073 5,3,48 Welch et al. 00 S. flexneri A 457T 4,599,354 Wei et al. 003 S. flexneri A 4,607,03 Jin et al. 00 S. enterica Typhimurium LT 4,857,43 McClelland et al. 00 S. enterica Typhi CT8 4,809,037 Parkhill et al. 00 S. enterica Typhi Ty 4,79,96 Deng et al. 003 Processing by Mauve took 3 hours on a single core. This is the tree that Mauve estimated from the genome sequences and which is used to generate multiple alignments on each of the locally colinear blocks: Here is the resulting genome alignment: A. Darling et al., Genome Research (004)
6 Bioinformatics I, WS /3, D. Huson, December 3, 0 99 Question: How much sequence is conserved over all nine enterobacterial genomes? To address this, define a conserved backbone segment as a segment of a Mauve alignment that contains more than 50 gap-free columns and does not have a run of 50 or more consecutive gaps in any single genome sequence. Based on this, the nine enterobacteria have.86 Mb of conserved backbone sequence partitioned into 5 backbone segments. 7.4 Eukaryotes Example Although Mauve was originally designed for bacterial-sized genomes, it can also be applied to large genomes as well. The calculation of a comparison of human, mouse and rat (without calculating the alignments of reach of the LCB s) takes about hours. Depending on the parameters used, this alignment has between 000 and 000 LCBs:
7 00 Bioinformatics I, WS /3, D. Huson, December 3, Yersinia Example In 00, only three species of the enterobacterial genus Yersinia, ones that cause invasive human diseases (Yersinia pestis, Yersinia pseudotuberculosis, and Yersinia enterocolitica), had been sequenced. However, there were no genomic data on the Yersinia species with more limited virulence potential, such as are frequently found in soil and water environments. A paper published in 003 provided new genome sequences for other Yersinia species. The paper presents a number of different types of comparisons of the different genomes, including a Mauve analysis: Image source: P. Chen et al, Genome Biology (00) 3 P. Chen et al, Genome Biology (00),
I519 Introduction to Bioinformatics, Genome Comparison. Yuzhen Ye School of Informatics & Computing, IUB
 I519 Introduction to Bioinformatics, 2015 Genome Comparison Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Whole genome comparison/alignment Build better phylogenies Identify polymorphism
I519 Introduction to Bioinformatics, 2015 Genome Comparison Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Whole genome comparison/alignment Build better phylogenies Identify polymorphism
BMI/CS 776 Lecture #20 Alignment of whole genomes. Colin Dewey (with slides adapted from those by Mark Craven)
 BMI/CS 776 Lecture #20 Alignment of whole genomes Colin Dewey (with slides adapted from those by Mark Craven) 2007.03.29 1 Multiple whole genome alignment Input set of whole genome sequences genomes diverged
BMI/CS 776 Lecture #20 Alignment of whole genomes Colin Dewey (with slides adapted from those by Mark Craven) 2007.03.29 1 Multiple whole genome alignment Input set of whole genome sequences genomes diverged
17 Non-collinear alignment Motivation A B C A B C A B C A B C D A C. This exposition is based on:
 17 Non-collinear alignment This exposition is based on: 1. Darling, A.E., Mau, B., Perna, N.T. (2010) progressivemauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5(6):e11147.
17 Non-collinear alignment This exposition is based on: 1. Darling, A.E., Mau, B., Perna, N.T. (2010) progressivemauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5(6):e11147.
I519 Introduction to Bioinformatics, Genome Comparison. Yuzhen Ye School of Informatics & Computing, IUB
 I519 Introduction to Bioinformatics, 2011 Genome Comparison Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Whole genome comparison/alignment Build better phylogenies Identify polymorphism
I519 Introduction to Bioinformatics, 2011 Genome Comparison Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Whole genome comparison/alignment Build better phylogenies Identify polymorphism
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss
 Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
Genômica comparativa. João Carlos Setubal IQ-USP outubro /5/2012 J. C. Setubal
 Genômica comparativa João Carlos Setubal IQ-USP outubro 2012 11/5/2012 J. C. Setubal 1 Comparative genomics There are currently (out/2012) 2,230 completed sequenced microbial genomes publicly available
Genômica comparativa João Carlos Setubal IQ-USP outubro 2012 11/5/2012 J. C. Setubal 1 Comparative genomics There are currently (out/2012) 2,230 completed sequenced microbial genomes publicly available
Whole Genome Alignments and Synteny Maps
 Whole Genome Alignments and Synteny Maps IINTRODUCTION It was not until closely related organism genomes have been sequenced that people start to think about aligning genomes and chromosomes instead of
Whole Genome Alignments and Synteny Maps IINTRODUCTION It was not until closely related organism genomes have been sequenced that people start to think about aligning genomes and chromosomes instead of
Inferring positional homologs with common intervals of sequences
 Outline Introduction Our approach Results Conclusion Inferring positional homologs with common intervals of sequences Guillaume Blin, Annie Chateau, Cedric Chauve, Yannick Gingras CGL - Université du Québec
Outline Introduction Our approach Results Conclusion Inferring positional homologs with common intervals of sequences Guillaume Blin, Annie Chateau, Cedric Chauve, Yannick Gingras CGL - Université du Québec
Multiple Whole Genome Alignment
 Multiple Whole Genome Alignment BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 206 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Multiple Whole Genome Alignment BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 206 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
General context Anchor-based method Evaluation Discussion. CoCoGen meeting. Accuracy of the anchor-based strategy for genome alignment.
 CoCoGen meeting Accuracy of the anchor-based strategy for genome alignment Raluca Uricaru LIRMM, CNRS Université de Montpellier 2 3 octobre 2008 1 / 31 Summary 1 General context 2 Global alignment : anchor-based
CoCoGen meeting Accuracy of the anchor-based strategy for genome alignment Raluca Uricaru LIRMM, CNRS Université de Montpellier 2 3 octobre 2008 1 / 31 Summary 1 General context 2 Global alignment : anchor-based
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Genome Rearrangements In Man and Mouse. Abhinav Tiwari Department of Bioengineering
 Genome Rearrangements In Man and Mouse Abhinav Tiwari Department of Bioengineering Genome Rearrangement Scrambling of the order of the genome during evolution Operations on chromosomes Reversal Translocation
Genome Rearrangements In Man and Mouse Abhinav Tiwari Department of Bioengineering Genome Rearrangement Scrambling of the order of the genome during evolution Operations on chromosomes Reversal Translocation
An Integrative Method for Accurate Comparative Genome Mapping
 An Integrative Method for Accurate Comparative Genome Mapping Firas Swidan 1,2*, Eduardo P. C. Rocha 3,4, Michael Shmoish 1, Ron Y. Pinter 1 1 Department of Computer Science, Technion, Israel Institute
An Integrative Method for Accurate Comparative Genome Mapping Firas Swidan 1,2*, Eduardo P. C. Rocha 3,4, Michael Shmoish 1, Ron Y. Pinter 1 1 Department of Computer Science, Technion, Israel Institute
Design of an Enterobacteriaceae Pan-genome Microarray Chip
 Design of an Enterobacteriaceae Pan-genome Microarray Chip Oksana Lukjancenko and David W. Ussery DTU CBS 2010 2 Background Pan-genome complete collection of variuos genes located within populations at
Design of an Enterobacteriaceae Pan-genome Microarray Chip Oksana Lukjancenko and David W. Ussery DTU CBS 2010 2 Background Pan-genome complete collection of variuos genes located within populations at
Understanding microbes through the lens of comparative genomics... a biased perspective. Aaron Darling A/Prof. ithree institute UTS
 Understanding microbes through the lens of comparative genomics... a biased perspective Aaron Darling A/Prof. ithree institute UTS Molecular Evolution of Bacteria Bacteria reproduce clonally A brief history
Understanding microbes through the lens of comparative genomics... a biased perspective Aaron Darling A/Prof. ithree institute UTS Molecular Evolution of Bacteria Bacteria reproduce clonally A brief history
arxiv: v1 [q-bio.gn] 30 Oct 2009
![arxiv: v1 [q-bio.gn] 30 Oct 2009 arxiv: v1 [q-bio.gn] 30 Oct 2009](/thumbs/84/90800978.jpg) arxiv:0910.5780v1 [q-bio.gn] 30 Oct 2009 Progressive Mauve: Multiple alignment of genomes with gene flux and rearrangement Aaron E. Darling 1,2,3 Bob Mau 4 Nicole T. Perna 5 Running title: Multiple genome
arxiv:0910.5780v1 [q-bio.gn] 30 Oct 2009 Progressive Mauve: Multiple alignment of genomes with gene flux and rearrangement Aaron E. Darling 1,2,3 Bob Mau 4 Nicole T. Perna 5 Running title: Multiple genome
Course: Visual Analytics of largescale biological data. Kay Nieselt Center for Bioinformatics Tübingen University of Tübingen
 Course: Visual Analytics of largescale biological data Kay Nieselt Center for Bioinformatics Tübingen University of Tübingen THE SUPERGENOME AND GENOMERING Overview A revolution in genomics Flood of genomes:
Course: Visual Analytics of largescale biological data Kay Nieselt Center for Bioinformatics Tübingen University of Tübingen THE SUPERGENOME AND GENOMERING Overview A revolution in genomics Flood of genomes:
Comparison of 61 E. coli genomes
 Comparison of 61 E. coli genomes Center for Biological Sequence Analysis Department of Systems Biology Dave Ussery! DTU course 27105 - Comparative Genomics Oksana s 61 E. coli genomes paper! Monday, 23
Comparison of 61 E. coli genomes Center for Biological Sequence Analysis Department of Systems Biology Dave Ussery! DTU course 27105 - Comparative Genomics Oksana s 61 E. coli genomes paper! Monday, 23
Chromosomal rearrangements in mammalian genomes : characterising the breakpoints. Claire Lemaitre
 PhD defense Chromosomal rearrangements in mammalian genomes : characterising the breakpoints Claire Lemaitre Laboratoire de Biométrie et Biologie Évolutive Université Claude Bernard Lyon 1 6 novembre 2008
PhD defense Chromosomal rearrangements in mammalian genomes : characterising the breakpoints Claire Lemaitre Laboratoire de Biométrie et Biologie Évolutive Université Claude Bernard Lyon 1 6 novembre 2008
The Evolution of Infectious Disease
 The Evolution of Infectious Disease Why are some bacteria pathogenic to humans while other (closely-related) bacteria are not? This question can be approached from two directions: 1.From the point of view
The Evolution of Infectious Disease Why are some bacteria pathogenic to humans while other (closely-related) bacteria are not? This question can be approached from two directions: 1.From the point of view
Genomes and Their Evolution
 Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Effects of Gap Open and Gap Extension Penalties
 Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Overview of IslandPick pipeline and the generation of GI datasets
 Overview of IslandPick pipeline and the generation of GI datasets Predicting GIs using comparative genomics By using whole genome alignments we can identify regions that are present in one genome but not
Overview of IslandPick pipeline and the generation of GI datasets Predicting GIs using comparative genomics By using whole genome alignments we can identify regions that are present in one genome but not
Phylogenetics without multiple sequence alignment
 Phylogenetics without multiple sequence alignment Mark Ragan Institute for Molecular Bioscience and School of Information Technology & Electrical Engineering The University of Queensland, Brisbane, Australia
Phylogenetics without multiple sequence alignment Mark Ragan Institute for Molecular Bioscience and School of Information Technology & Electrical Engineering The University of Queensland, Brisbane, Australia
Database and Comparative Identification of Prophages
 Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy
Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy
Comparing whole genomes
 BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
Pyrobayes: an improved base caller for SNP discovery in pyrosequences
 Pyrobayes: an improved base caller for SNP discovery in pyrosequences Aaron R Quinlan, Donald A Stewart, Michael P Strömberg & Gábor T Marth Supplementary figures and text: Supplementary Figure 1. The
Pyrobayes: an improved base caller for SNP discovery in pyrosequences Aaron R Quinlan, Donald A Stewart, Michael P Strömberg & Gábor T Marth Supplementary figures and text: Supplementary Figure 1. The
Sequence comparison by compression
 Sequence comparison by compression Motivation similarity as a marker for homology. And homology is used to infer function. Sometimes, we are only interested in a numerical distance between two sequences.
Sequence comparison by compression Motivation similarity as a marker for homology. And homology is used to infer function. Sometimes, we are only interested in a numerical distance between two sequences.
Phylogenies Scores for Exhaustive Maximum Likelihood and Parsimony Scores Searches
 Int. J. Bioinformatics Research and Applications, Vol. x, No. x, xxxx Phylogenies Scores for Exhaustive Maximum Likelihood and s Searches Hyrum D. Carroll, Perry G. Ridge, Mark J. Clement, Quinn O. Snell
Int. J. Bioinformatics Research and Applications, Vol. x, No. x, xxxx Phylogenies Scores for Exhaustive Maximum Likelihood and s Searches Hyrum D. Carroll, Perry G. Ridge, Mark J. Clement, Quinn O. Snell
Genes order and phylogenetic reconstruction: application to γ-proteobacteria
 Genes order and phylogenetic reconstruction: application to γ-proteobacteria Guillaume Blin 1, Cedric Chauve 2 and Guillaume Fertin 1 1 LINA FRE CNRS 2729, Université de Nantes 2 rue de la Houssinière,
Genes order and phylogenetic reconstruction: application to γ-proteobacteria Guillaume Blin 1, Cedric Chauve 2 and Guillaume Fertin 1 1 LINA FRE CNRS 2729, Université de Nantes 2 rue de la Houssinière,
Genetic Basis of Variation in Bacteria
 Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Whole Genome Alignment. Adam Phillippy University of Maryland, Fall 2012
 Whole Genome Alignment Adam Phillippy University of Maryland, Fall 2012 Motivation cancergenome.nih.gov Breast cancer karyotypes www.path.cam.ac.uk Goal of whole-genome alignment } For two genomes, A and
Whole Genome Alignment Adam Phillippy University of Maryland, Fall 2012 Motivation cancergenome.nih.gov Breast cancer karyotypes www.path.cam.ac.uk Goal of whole-genome alignment } For two genomes, A and
A Phylogenetic Gibbs Recursive Sampler for Locating Transcription Factor Binding Sites
 A for Locating Transcription Factor Binding Sites Sean P. Conlan 1 Lee Ann McCue 2 1,3 Thomas M. Smith 3 William Thompson 4 Charles E. Lawrence 4 1 Wadsworth Center, New York State Department of Health
A for Locating Transcription Factor Binding Sites Sean P. Conlan 1 Lee Ann McCue 2 1,3 Thomas M. Smith 3 William Thompson 4 Charles E. Lawrence 4 1 Wadsworth Center, New York State Department of Health
Copyright 2000 N. AYDIN. All rights reserved. 1
 Introduction to Bioinformatics Prof. Dr. Nizamettin AYDIN naydin@yildiz.edu.tr Multiple Sequence Alignment Outline Multiple sequence alignment introduction to msa methods of msa progressive global alignment
Introduction to Bioinformatics Prof. Dr. Nizamettin AYDIN naydin@yildiz.edu.tr Multiple Sequence Alignment Outline Multiple sequence alignment introduction to msa methods of msa progressive global alignment
Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and
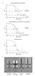 Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and comparisons to other published in situ T3SS structures. a, Resolution estimates after applying Fourier shell correlation
Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and comparisons to other published in situ T3SS structures. a, Resolution estimates after applying Fourier shell correlation
Sequence Database Search Techniques I: Blast and PatternHunter tools
 Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
Evidence That Mutation Is Universally Biased towards AT in Bacteria
 Evidence That Mutation Is Universally Biased towards AT in Bacteria Ruth Hershberg*, Dmitri A. Petrov Department of Biology, Stanford University, Stanford, California, United States of America Abstract
Evidence That Mutation Is Universally Biased towards AT in Bacteria Ruth Hershberg*, Dmitri A. Petrov Department of Biology, Stanford University, Stanford, California, United States of America Abstract
Network alignment and querying
 Network biology minicourse (part 4) Algorithmic challenges in genomics Network alignment and querying Roded Sharan School of Computer Science, Tel Aviv University Multiple Species PPI Data Rapid growth
Network biology minicourse (part 4) Algorithmic challenges in genomics Network alignment and querying Roded Sharan School of Computer Science, Tel Aviv University Multiple Species PPI Data Rapid growth
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Multiple sequence alignment
 Multiple sequence alignment Multiple sequence alignment: today s goals to define what a multiple sequence alignment is and how it is generated; to describe profile HMMs to introduce databases of multiple
Multiple sequence alignment Multiple sequence alignment: today s goals to define what a multiple sequence alignment is and how it is generated; to describe profile HMMs to introduce databases of multiple
Alignment Strategies for Large Scale Genome Alignments
 Alignment Strategies for Large Scale Genome Alignments CSHL Computational Genomics 9 November 2003 Algorithms for Biological Sequence Comparison algorithm value scoring gap time calculated matrix penalty
Alignment Strategies for Large Scale Genome Alignments CSHL Computational Genomics 9 November 2003 Algorithms for Biological Sequence Comparison algorithm value scoring gap time calculated matrix penalty
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT
 5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
Sequence Analysis '17- lecture 8. Multiple sequence alignment
 Sequence Analysis '17- lecture 8 Multiple sequence alignment Ex5 explanation How many random database search scores have e-values 10? (Answer: 10!) Why? e-value of x = m*p(s x), where m is the database
Sequence Analysis '17- lecture 8 Multiple sequence alignment Ex5 explanation How many random database search scores have e-values 10? (Answer: 10!) Why? e-value of x = m*p(s x), where m is the database
Improved Sensitivity And Reliability Of Anchor Based Genome Alignment
 Improved Sensitivity And Reliability Of Anchor Based Genome Alignment Raluca Uricaru, Célia Michotey, Laurent Noé, Hélène Chiapello, Eric Rivals To cite this version: Raluca Uricaru, Célia Michotey, Laurent
Improved Sensitivity And Reliability Of Anchor Based Genome Alignment Raluca Uricaru, Célia Michotey, Laurent Noé, Hélène Chiapello, Eric Rivals To cite this version: Raluca Uricaru, Célia Michotey, Laurent
Copyright Warning & Restrictions
 Copyright Warning & Restrictions The copyright law of the United States (Title 17, United States Code) governs the making of photocopies or other reproductions of copyrighted material. Under certain conditions
Copyright Warning & Restrictions The copyright law of the United States (Title 17, United States Code) governs the making of photocopies or other reproductions of copyrighted material. Under certain conditions
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 05: Index-based alignment algorithms Slides adapted from Dr. Shaojie Zhang (University of Central Florida) Real applications of alignment Database search
EECS730: Introduction to Bioinformatics Lecture 05: Index-based alignment algorithms Slides adapted from Dr. Shaojie Zhang (University of Central Florida) Real applications of alignment Database search
The breakpoint distance for signed sequences
 The breakpoint distance for signed sequences Guillaume Blin 1, Cedric Chauve 2 Guillaume Fertin 1 and 1 LINA, FRE CNRS 2729 2 LACIM et Département d'informatique, Université de Nantes, Université du Québec
The breakpoint distance for signed sequences Guillaume Blin 1, Cedric Chauve 2 Guillaume Fertin 1 and 1 LINA, FRE CNRS 2729 2 LACIM et Département d'informatique, Université de Nantes, Université du Québec
Pairwise & Multiple sequence alignments
 Pairwise & Multiple sequence alignments Urmila Kulkarni-Kale Bioinformatics Centre 411 007 urmila@bioinfo.ernet.in Basis for Sequence comparison Theory of evolution: gene sequences have evolved/derived
Pairwise & Multiple sequence alignments Urmila Kulkarni-Kale Bioinformatics Centre 411 007 urmila@bioinfo.ernet.in Basis for Sequence comparison Theory of evolution: gene sequences have evolved/derived
Reversing Gene Erosion Reconstructing Ancestral Bacterial Genomes from Gene-Content and Order Data
 Reversing Gene Erosion Reconstructing Ancestral Bacterial Genomes from Gene-Content and Order Data Joel V. Earnest-DeYoung 1, Emmanuelle Lerat 2, and Bernard M.E. Moret 1,3 Abstract In the last few years,
Reversing Gene Erosion Reconstructing Ancestral Bacterial Genomes from Gene-Content and Order Data Joel V. Earnest-DeYoung 1, Emmanuelle Lerat 2, and Bernard M.E. Moret 1,3 Abstract In the last few years,
Supplemental. Location Year Source. 1412G Dalidag, Georgia 1979 Flea
 Supplemental Table S1: pestis strains (n=59) a Strain designation / bei # Location Year Source -- 1391G 1392G 1393G Ninotsminda, 1979 Common vole 1412G Dalidag, 1979 Flea 1413G 1670G 1851G 1852G 1853G
Supplemental Table S1: pestis strains (n=59) a Strain designation / bei # Location Year Source -- 1391G 1392G 1393G Ninotsminda, 1979 Common vole 1412G Dalidag, 1979 Flea 1413G 1670G 1851G 1852G 1853G
Sequence Alignment (chapter 6)
 Sequence lignment (chapter 6) he biological problem lobal alignment Local alignment Multiple alignment Introduction to bioinformatics, utumn 6 Background: comparative genomics Basic question in biology:
Sequence lignment (chapter 6) he biological problem lobal alignment Local alignment Multiple alignment Introduction to bioinformatics, utumn 6 Background: comparative genomics Basic question in biology:
Pairwise Alignment. Guan-Shieng Huang. Dept. of CSIE, NCNU. Pairwise Alignment p.1/55
 Pairwise Alignment Guan-Shieng Huang shieng@ncnu.edu.tw Dept. of CSIE, NCNU Pairwise Alignment p.1/55 Approach 1. Problem definition 2. Computational method (algorithms) 3. Complexity and performance Pairwise
Pairwise Alignment Guan-Shieng Huang shieng@ncnu.edu.tw Dept. of CSIE, NCNU Pairwise Alignment p.1/55 Approach 1. Problem definition 2. Computational method (algorithms) 3. Complexity and performance Pairwise
Algorithms in Bioinformatics FOUR Pairwise Sequence Alignment. Pairwise Sequence Alignment. Convention: DNA Sequences 5. Sequence Alignment
 Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
BIOINFORMATICS ORIGINAL PAPER
 BIOINFORMATICS ORIGINAL PAPER Vol. 25 no. 24 2009, pages 3221 3227 doi:10.1093/bioinformatics/btp590 Genome analysis Efficient estimation of pairwise distances between genomes Mirjana Domazet-Lošo 1,2
BIOINFORMATICS ORIGINAL PAPER Vol. 25 no. 24 2009, pages 3221 3227 doi:10.1093/bioinformatics/btp590 Genome analysis Efficient estimation of pairwise distances between genomes Mirjana Domazet-Lošo 1,2
Handling Rearrangements in DNA Sequence Alignment
 Handling Rearrangements in DNA Sequence Alignment Maneesh Bhand 12/5/10 1 Introduction Sequence alignment is one of the core problems of bioinformatics, with a broad range of applications such as genome
Handling Rearrangements in DNA Sequence Alignment Maneesh Bhand 12/5/10 1 Introduction Sequence alignment is one of the core problems of bioinformatics, with a broad range of applications such as genome
Bioinformatics Exercises
 Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Multiple Alignment of Genomic Sequences
 Ross Metzger June 4, 2004 Biochemistry 218 Multiple Alignment of Genomic Sequences Genomic sequence is currently available from ENTREZ for more than 40 eukaryotic and 157 prokaryotic organisms. As part
Ross Metzger June 4, 2004 Biochemistry 218 Multiple Alignment of Genomic Sequences Genomic sequence is currently available from ENTREZ for more than 40 eukaryotic and 157 prokaryotic organisms. As part
Bioinformatics and BLAST
 Bioinformatics and BLAST Overview Recap of last time Similarity discussion Algorithms: Needleman-Wunsch Smith-Waterman BLAST Implementation issues and current research Recap from Last Time Genome consists
Bioinformatics and BLAST Overview Recap of last time Similarity discussion Algorithms: Needleman-Wunsch Smith-Waterman BLAST Implementation issues and current research Recap from Last Time Genome consists
Reducing storage requirements for biological sequence comparison
 Bioinformatics Advance Access published July 15, 2004 Bioinfor matics Oxford University Press 2004; all rights reserved. Reducing storage requirements for biological sequence comparison Michael Roberts,
Bioinformatics Advance Access published July 15, 2004 Bioinfor matics Oxford University Press 2004; all rights reserved. Reducing storage requirements for biological sequence comparison Michael Roberts,
InDel 3-5. InDel 8-9. InDel 3-5. InDel 8-9. InDel InDel 8-9
 Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family
 A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family Jieming Shen 1,2 and Hugh B. Nicholas, Jr. 3 1 Bioengineering and Bioinformatics Summer
A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family Jieming Shen 1,2 and Hugh B. Nicholas, Jr. 3 1 Bioengineering and Bioinformatics Summer
Sequence Alignment: Scoring Schemes. COMP 571 Luay Nakhleh, Rice University
 Sequence Alignment: Scoring Schemes COMP 571 Luay Nakhleh, Rice University Scoring Schemes Recall that an alignment score is aimed at providing a scale to measure the degree of similarity (or difference)
Sequence Alignment: Scoring Schemes COMP 571 Luay Nakhleh, Rice University Scoring Schemes Recall that an alignment score is aimed at providing a scale to measure the degree of similarity (or difference)
Sequence analysis and comparison
 The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
Phylogenetic Gibbs Recursive Sampler. Locating Transcription Factor Binding Sites
 A for Locating Transcription Factor Binding Sites Sean P. Conlan 1 Lee Ann McCue 2 1,3 Thomas M. Smith 3 William Thompson 4 Charles E. Lawrence 4 1 Wadsworth Center, New York State Department of Health
A for Locating Transcription Factor Binding Sites Sean P. Conlan 1 Lee Ann McCue 2 1,3 Thomas M. Smith 3 William Thompson 4 Charles E. Lawrence 4 1 Wadsworth Center, New York State Department of Health
Algorithms for Bioinformatics
 Adapted from slides by Alexandru Tomescu, Leena Salmela, Veli Mäkinen, Esa Pitkänen 582670 Algorithms for Bioinformatics Lecture 5: Combinatorial Algorithms and Genomic Rearrangements 1.10.2015 Background
Adapted from slides by Alexandru Tomescu, Leena Salmela, Veli Mäkinen, Esa Pitkänen 582670 Algorithms for Bioinformatics Lecture 5: Combinatorial Algorithms and Genomic Rearrangements 1.10.2015 Background
Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:
 Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:50 5001 5 Multiple Sequence Alignment The first part of this exposition is based on the following sources, which are recommended reading:
Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:50 5001 5 Multiple Sequence Alignment The first part of this exposition is based on the following sources, which are recommended reading:
Discovery of Genomic Structural Variations with Next-Generation Sequencing Data
 Discovery of Genomic Structural Variations with Next-Generation Sequencing Data Advanced Topics in Computational Genomics Slides from Marcel H. Schulz, Tobias Rausch (EMBL), and Kai Ye (Leiden University)
Discovery of Genomic Structural Variations with Next-Generation Sequencing Data Advanced Topics in Computational Genomics Slides from Marcel H. Schulz, Tobias Rausch (EMBL), and Kai Ye (Leiden University)
Gram negative bacilli
 Gram negative bacilli 1-Enterobacteriaceae Gram negative bacilli-rods Enterobacteriaceae Are everywhere Part of normal flora of humans and most animals They are cause of -30-35% septisemia -more than 70%
Gram negative bacilli 1-Enterobacteriaceae Gram negative bacilli-rods Enterobacteriaceae Are everywhere Part of normal flora of humans and most animals They are cause of -30-35% septisemia -more than 70%
Greedy Algorithms. CS 498 SS Saurabh Sinha
 Greedy Algorithms CS 498 SS Saurabh Sinha Chapter 5.5 A greedy approach to the motif finding problem Given t sequences of length n each, to find a profile matrix of length l. Enumerative approach O(l n
Greedy Algorithms CS 498 SS Saurabh Sinha Chapter 5.5 A greedy approach to the motif finding problem Given t sequences of length n each, to find a profile matrix of length l. Enumerative approach O(l n
Background: comparative genomics. Sequence similarity. Homologs. Similarity vs homology (2) Similarity vs homology. Sequence Alignment (chapter 6)
 Sequence lignment (chapter ) he biological problem lobal alignment Local alignment Multiple alignment Background: comparative genomics Basic question in biology: what properties are shared among organisms?
Sequence lignment (chapter ) he biological problem lobal alignment Local alignment Multiple alignment Background: comparative genomics Basic question in biology: what properties are shared among organisms?
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Sara C. Madeira. Universidade da Beira Interior. (Thanks to Ana Teresa Freitas, IST for useful resources on this subject)
 Bioinformática Sequence Alignment Pairwise Sequence Alignment Universidade da Beira Interior (Thanks to Ana Teresa Freitas, IST for useful resources on this subject) 1 16/3/29 & 23/3/29 27/4/29 Outline
Bioinformática Sequence Alignment Pairwise Sequence Alignment Universidade da Beira Interior (Thanks to Ana Teresa Freitas, IST for useful resources on this subject) 1 16/3/29 & 23/3/29 27/4/29 Outline
1 Introduction. Abstract
 CBS 530 Assignment No 2 SHUBHRA GUPTA shubhg@asu.edu 993755974 Review of the papers: Construction and Analysis of a Human-Chimpanzee Comparative Clone Map and Intra- and Interspecific Variation in Primate
CBS 530 Assignment No 2 SHUBHRA GUPTA shubhg@asu.edu 993755974 Review of the papers: Construction and Analysis of a Human-Chimpanzee Comparative Clone Map and Intra- and Interspecific Variation in Primate
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Bioinformatics 1. Sepp Hochreiter. Biology, Sequences, Phylogenetics Part 4. Bioinformatics 1: Biology, Sequences, Phylogenetics
 Bioinformatics 1 Biology, Sequences, Phylogenetics Part 4 Sepp Hochreiter Klausur Mo. 30.01.2011 Zeit: 15:30 17:00 Raum: HS14 Anmeldung Kusss Contents Methods and Bootstrapping of Maximum Methods Methods
Bioinformatics 1 Biology, Sequences, Phylogenetics Part 4 Sepp Hochreiter Klausur Mo. 30.01.2011 Zeit: 15:30 17:00 Raum: HS14 Anmeldung Kusss Contents Methods and Bootstrapping of Maximum Methods Methods
20 Grundlagen der Bioinformatik, SS 08, D. Huson, May 27, Global and local alignment of two sequences using dynamic programming
 20 Grundlagen der Bioinformatik, SS 08, D. Huson, May 27, 2008 4 Pairwise alignment We will discuss: 1. Strings 2. Dot matrix method for comparing sequences 3. Edit distance 4. Global and local alignment
20 Grundlagen der Bioinformatik, SS 08, D. Huson, May 27, 2008 4 Pairwise alignment We will discuss: 1. Strings 2. Dot matrix method for comparing sequences 3. Edit distance 4. Global and local alignment
Linear-Space Alignment
 Linear-Space Alignment Subsequences and Substrings Definition A string x is a substring of a string x, if x = ux v for some prefix string u and suffix string v (similarly, x = x i x j, for some 1 i j x
Linear-Space Alignment Subsequences and Substrings Definition A string x is a substring of a string x, if x = ux v for some prefix string u and suffix string v (similarly, x = x i x j, for some 1 i j x
Grundlagen der Bioinformatik, SS 08, D. Huson, May 2,
 Grundlagen der Bioinformatik, SS 08, D. Huson, May 2, 2008 39 5 Blast This lecture is based on the following, which are all recommended reading: R. Merkl, S. Waack: Bioinformatik Interaktiv. Chapter 11.4-11.7
Grundlagen der Bioinformatik, SS 08, D. Huson, May 2, 2008 39 5 Blast This lecture is based on the following, which are all recommended reading: R. Merkl, S. Waack: Bioinformatik Interaktiv. Chapter 11.4-11.7
Comparative Genomics Background and Strategies. Nitya Sharma, Emily Rogers, Kanika Arora, Zhiming Zhao, Yun Gyeong Lee
 Comparative Genomics Background and Strategies Nitya Sharma, Emily Rogers, Kanika Arora, Zhiming Zhao, Yun Gyeong Lee Introduction Why comparative genomes? h"p://www.ensembl.org/info/about/species.html
Comparative Genomics Background and Strategies Nitya Sharma, Emily Rogers, Kanika Arora, Zhiming Zhao, Yun Gyeong Lee Introduction Why comparative genomes? h"p://www.ensembl.org/info/about/species.html
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
MULTIPLE SEQUENCE ALIGNMENT FOR CONSTRUCTION OF PHYLOGENETIC TREE
 MULTIPLE SEQUENCE ALIGNMENT FOR CONSTRUCTION OF PHYLOGENETIC TREE Manmeet Kaur 1, Navneet Kaur Bawa 2 1 M-tech research scholar (CSE Dept) ACET, Manawala,Asr 2 Associate Professor (CSE Dept) ACET, Manawala,Asr
MULTIPLE SEQUENCE ALIGNMENT FOR CONSTRUCTION OF PHYLOGENETIC TREE Manmeet Kaur 1, Navneet Kaur Bawa 2 1 M-tech research scholar (CSE Dept) ACET, Manawala,Asr 2 Associate Professor (CSE Dept) ACET, Manawala,Asr
Sequence Bioinformatics. Multiple Sequence Alignment Waqas Nasir
 Sequence Bioinformatics Multiple Sequence Alignment Waqas Nasir 2010-11-12 Multiple Sequence Alignment One amino acid plays coy; a pair of homologous sequences whisper; many aligned sequences shout out
Sequence Bioinformatics Multiple Sequence Alignment Waqas Nasir 2010-11-12 Multiple Sequence Alignment One amino acid plays coy; a pair of homologous sequences whisper; many aligned sequences shout out
Introduction to Bioinformatics
 Introduction to Bioinformatics Lecture : p he biological problem p lobal alignment p Local alignment p Multiple alignment 6 Background: comparative genomics p Basic question in biology: what properties
Introduction to Bioinformatics Lecture : p he biological problem p lobal alignment p Local alignment p Multiple alignment 6 Background: comparative genomics p Basic question in biology: what properties
98 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 6, 2006
 98 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 6, 2006 8.3.1 Simple energy minimization Maximizing the number of base pairs as described above does not lead to good structure predictions.
98 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 6, 2006 8.3.1 Simple energy minimization Maximizing the number of base pairs as described above does not lead to good structure predictions.
Bioinformatics. Part 8. Sequence Analysis An introduction. Mahdi Vasighi
 Bioinformatics Sequence Analysis An introduction Part 8 Mahdi Vasighi Sequence analysis Some of the earliest problems in genomics concerned how to measure similarity of DNA and protein sequences, either
Bioinformatics Sequence Analysis An introduction Part 8 Mahdi Vasighi Sequence analysis Some of the earliest problems in genomics concerned how to measure similarity of DNA and protein sequences, either
Multiple Sequence Alignment
 Multiple Sequence Alignment Multiple Alignment versus Pairwise Alignment Up until now we have only tried to align two sequences. What about more than two? And what for? A faint similarity between two sequences
Multiple Sequence Alignment Multiple Alignment versus Pairwise Alignment Up until now we have only tried to align two sequences. What about more than two? And what for? A faint similarity between two sequences
Lecture 2: Pairwise Alignment. CG Ron Shamir
 Lecture 2: Pairwise Alignment 1 Main source 2 Why compare sequences? Human hexosaminidase A vs Mouse hexosaminidase A 3 www.mathworks.com/.../jan04/bio_genome.html Sequence Alignment עימוד רצפים The problem:
Lecture 2: Pairwise Alignment 1 Main source 2 Why compare sequences? Human hexosaminidase A vs Mouse hexosaminidase A 3 www.mathworks.com/.../jan04/bio_genome.html Sequence Alignment עימוד רצפים The problem:
A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS
 A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS CRYSTAL L. KAHN and BENJAMIN J. RAPHAEL Box 1910, Brown University Department of Computer Science & Center for Computational Molecular Biology
A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS CRYSTAL L. KAHN and BENJAMIN J. RAPHAEL Box 1910, Brown University Department of Computer Science & Center for Computational Molecular Biology
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Plant transformation
 Plant transformation Objectives: 1. What is plant transformation? 2. What is Agrobacterium? How and why does it transform plant cells? 3. How is Agrobacterium used as a tool in molecular genetics? References:
Plant transformation Objectives: 1. What is plant transformation? 2. What is Agrobacterium? How and why does it transform plant cells? 3. How is Agrobacterium used as a tool in molecular genetics? References:
Multiple Sequence Alignment. Sequences
 Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools. Evaluation. Course Homepage.
 CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Sequence Comparison. mouse human
 Sequence Comparison Sequence Comparison mouse human Why Compare Sequences? The first fact of biological sequence analysis In biomolecular sequences (DNA, RNA, or amino acid sequences), high sequence similarity
Sequence Comparison Sequence Comparison mouse human Why Compare Sequences? The first fact of biological sequence analysis In biomolecular sequences (DNA, RNA, or amino acid sequences), high sequence similarity
Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment
 Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment Introduction to Bioinformatics online course : IBT Jonathan Kayondo Learning Objectives Understand
Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment Introduction to Bioinformatics online course : IBT Jonathan Kayondo Learning Objectives Understand
Genomes Comparision via de Bruijn graphs
 Genomes Comparision via de Bruijn graphs Student: Ilya Minkin Advisor: Son Pham St. Petersburg Academic University June 4, 2012 1 / 19 Synteny Blocks: Algorithmic challenge Suppose that we are given two
Genomes Comparision via de Bruijn graphs Student: Ilya Minkin Advisor: Son Pham St. Petersburg Academic University June 4, 2012 1 / 19 Synteny Blocks: Algorithmic challenge Suppose that we are given two
Tools and Algorithms in Bioinformatics
 Tools and Algorithms in Bioinformatics GCBA815, Fall 2015 Week-4 BLAST Algorithm Continued Multiple Sequence Alignment Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and
Tools and Algorithms in Bioinformatics GCBA815, Fall 2015 Week-4 BLAST Algorithm Continued Multiple Sequence Alignment Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and
Biochemistry 324 Bioinformatics. Pairwise sequence alignment
 Biochemistry 324 Bioinformatics Pairwise sequence alignment How do we compare genes/proteins? When we have sequenced a genome, we try and identify the function of unknown genes by finding a similar gene
Biochemistry 324 Bioinformatics Pairwise sequence alignment How do we compare genes/proteins? When we have sequenced a genome, we try and identify the function of unknown genes by finding a similar gene
Introduction to Bioinformatics Introduction to Bioinformatics
 Dr. rer. nat. Gong Jing Cancer Research Center Medicine School of Shandong University 2012.11.09 1 Chapter 4 Phylogenetic Tree 2 Phylogeny Evidence from morphological ( 形态学的 ), biochemical, and gene sequence
Dr. rer. nat. Gong Jing Cancer Research Center Medicine School of Shandong University 2012.11.09 1 Chapter 4 Phylogenetic Tree 2 Phylogeny Evidence from morphological ( 形态学的 ), biochemical, and gene sequence
