Comparison of simple sequence repeats in Staphylococcus strains using in-silico approach
|
|
|
- Adela Allen
- 5 years ago
- Views:
Transcription
1 Hypothesis Volume 8(24) Comparison of simple sequence repeats in Staphylococcus strains using in-silico approach Sunil Thorat 1 * & Prashant Thakare 2 1Institute of Bioresources and Sustainable Development, Imphal , Manipur, India; 2 Department of Biotechnology, S.G.B. Amravati University, Amravati , Maharashtra, India; Thorat Sunil sunilsthorat.ibsd@nic.in; *Corresponding author Received November 15, 2012; Accepted November 16, 2012; Published December 08, 2012 Abstract: Staphylococci are Gram-positive bacteria which play an important role in infectious disease and are major causes of communityacquired and hospital-acquired infections. Strains of Staphylococcus aureus are reported as genomically and phenotypically highly heterogeneous; hence in-silico based comparison of genomic data on simple sequence repeats may provide valuable information for understanding the pathogenicity and control measures. This study determined the distribution of a specific group of Simple Sequence Repeats (SSRs), in genome sequences of six Staphylococcus strains (Staphylococcus aureus COL, S.aureus MRSA252, S.aureus MSSA476, S.aureus Mu50, S.aureus MW2, S.aureus N315) and plasmid sequences of four Staphylococcus strains (Staphylococcus aureus COL pt181, Staphylococcus aureus MSSA psas, Staphylococcus aureus VRSAp, Staphylococcus aureus, Staphylococcus aureus pn315 DNA) downloaded from the GenBank database for identifying abundance, distribution and composition of SSRs. The data obtained in the present study shows that (i) a large number of tandem repeats are distributed throughout the genome and plasmid sequences. (ii) Number of mononucleotide SSRs decreased rapidly with increase in size of repeat unit. (iii) Total frequency of SSRs in plasmid regions is less than genomic regions. (iv) In all investigated strains, ratios of AT/TA repeats are dominating over GC/CG repeats in genomics as well as plasmid sequences, and (v) Dinucleotide combination of AT is dominated in all the six Staphylococcus genome sequences. Background: Staphylococcus is a genus of Gram-positive bacteria, a major human pathogen. The Staphylococcus genus includes at least forty species, of which; nine have two subspecies and one has three subspecies. Most are harmless and reside normally on the skin and mucous membranes of humans and other organisms. Found worldwide, they are a small component of soil microbial flora. The continuous emergence and spread of antibiotic resistant strains (e.g. MRSA, VRSA) leads to limited treatment options. Being a commensal organism living asymptomatically in the nasal cavities of a large number of the human population, S.aureus is also responsible for a raft of different infections that range in both anatomical site and severity. These infections are assisted by a vast array of different virulence factors like adhesins, invasins, toxins and modulins, which enables evasion of host immune responses and also contribute to colonization, dissemination, tissue damage and transmission. Though some responsible for serious invasive diseases. S.aureus is a leading cause of sepsis and infective endocarditis. Colonization of the heart and subsequent formation of vegetations involves a number of complex interactions [1]. The first bacterial species in which tandem repeats were identified was Mycobacterium tuberculosis, being described as mycobacterial interspersed repeat units [2, 3]. The sequences of six Staphylococcus aureus genomes; COL, MRSA252, MSSA476, Mu50, MW2 and N315 were studied along with four Staphylococcus aureus plasmids; COL pt181, MSSA psas, VRSAp, pn315 DNA which were downloaded from the GeneBank database. Five strains (COL, MRSA252, Mu50, MW2 and N315) are MRSA, while MSSA476 is methicillin sensitive. N315 and Mu50 are closely related strains that are hospitalacquired and vancomycin-intermediate resistance respectively, while MRSA252 is an epidemic hospital strain. MW2 is a hypervirulent community strain. The sequencing of these infections are superficial and self-limiting, S.aureus is also Bioinformation 8(24): (2012) Biomedical Informatics
2 genomes has allowed us to identify novel multiple tandem repeats that are similar to MIRUs in M.tuberculosis, and the present study describes the nature and distribution of these repeats, and their potential as a novel tool for understanding the micro-evolution of hospital MRSA. Microsatellites are also called simple sequence repeats (SSRs) or short tandem repeats (STRs) which are a group of tandem repeated sequences. SSR are comprised of mono-, di-, tri-, tetra-, penta-, or hexa-nucleotide units and are widely present in plant and animal genomes. These repeats can be either perfect tandem repeats or interrupted by several non-repeat nucleotides or compound repeats called compound SSRs [4]. These sequences experience frequent mutations that alter the number of repeats. The distribution of particular motif classes within a genome can vary substantially among different species [5, 6]. Repetitive DNA consists of simple homopolymeric tracts of a single nucleotide type [poly (A), poly (G), poly (T), or poly(c)] or of large or small numbers of several multimeric classes of repeats. These multimeric repeats are built from identical units (homogeneous repeats), mixed units (heterogeneous repeats), or degenerate repeat sequence motifs [7]. SSRs are highly polymorphic characterized by high rates of insertion and deletion (INDEL) mutations of their repeat units. INDELs of repeat units in a SSR arise due to slipped strand mispairing during the process of DNA replication. Slippage on the template strand tends to contraction of SSRs whereas slippage on the growing strand manifests into expansion of SSRs. The bias in SSRs either toward expansion or contraction is referred to as the directionality of SSR evolution. Ever since it became known that several hereditary diseases are associated with expansion of triplet repeats and colon cancer is associated with instability of certain mono and di-nucleotide repeats, there has been a lot of interest on the discovery of the mechanisms behind directionality of SSR mutations. Despite some investigations into the directional evolution of the SSRs, there is still an insufficient understanding of the factors influencing directionality of SSR mutations [8]. other species. Canine epilepsy is reported to be caused by expansion of 12-nucleotide repeats in the single exon of the Epm 2b gene [12]. The increasing availability of prokaryotic genome sequences has shown that SSRs are also widespread in prokaryotes and that there is extensive variation in their length, number and distribution [13-17]. The present study is an attempt to analyze distribution and composition of SSRs among whole genomes of six strains of Staphylococcus aureus. Methodology: Tools for analysis of SSRs A tool for Simple Sequence Repeats was developed to find out the short tandem repeats from the input sequences, the SSR tool is hosted on a local webserver i.e. wampserver which has a combination of Apache, MySQL and PHP to run the scripts for analysis. The front-end of SSR tool has Perl based links for dinucleotides, tri-nucleotides, tetra-nucleotides, penta-nucleotides and hexa-nucleotides. This tool is used to find out the number of nucleotide repeats of all the possible (mono, di, tri, tetra and pentanucleotide) combinations out of input nucleotide sequence. For combined analysis of all the five repeats; a separate script is developed where an input sequence is screened for repeats from di- to penta-nucleotides. A script for finding repeats for selective combinations for all the five SSR is also developed where a user can select a particular repeat (di-, tri-, etc.) for selective combinations (di-aa, tri-atc, etc.). This program can also analyse whole genome sequences. Apart from finding occurrences of all possible combinations of repeats, graphical presentation along with numerical data for research analysis can be viewed using SSR tool. Mononucleotide AAAA Dinucleotide CACACACA Trinucleotide ATGATGATGATG Tetranucleotide GTATGTATGTATGTAT Pentanucleotide CGTAGCGTAGCGTAGCGTAG SSRs have been extensively studied in eukaryote genomes and are well-established targets for pedigree analysis [9]; though there is insufficient information about microsatellites in simple organisms [10]. Bacterial SSR-type DNA can be divided into four main categories. First, dispersed repeat motifs that generally do not occur in tandem. These are repeats that occur throughout genomes of a mass of microorganisms, and are sometimes organized in tandem. A second class is formed by the homopolymeric tracts. Multimers of one of the four nucleotides that are frequently encountered in the genome, for instance the homogenous stretches of S.cerevisiae have occurred for as much as 42 nucleotides. The third category is short-motif SSRs with repeat units differing from 2 to 6 bases, this class of repeats are more likely to unit number variation at a given locus. Mostly, when these short-motif repeats are located within genes and are not 3 or 6 nucleotides long, they are likely to upset the coding potential of a given transcript. Fourth, the repeats of more than 8 nucleotides per unit, form a separate category. In addition, the longer repeat unit, the greater the chance that point mutations will be introduced [11]. The pathogenic phenotypes caused by unstable tandem repeats have been described in human beings. However, a few notable examples of diseases caused by repeats have been described in For the present study, we have used two tools, a webserver based Simple Sequence Repeats (SSR) tool as mentioned above and software developed by Gur-Arie (Ssr.exe) downloaded from (ftp://ftp.technion.ac.il/pub/supported/biotech) to screen the genomes and plasmids of the selected Staphylococcus strains for comparative analysis. Using web based SSR, possible combinations of repeats were searched from the input sequence and results were displayed with its frequency occurrences. On comparing the repeats of six genomes and four plasmids of Staphylococcus strains, the result showed frequency of particular repeat conserved in all the selected genomes and plasmids and also showed which particular combination occur the most. The offline tool Ssr.exe was used for extensive study by setting parameters with minimal number of repeats = 2, minimal motif length = 1, length of whole SSR array = (2*1) = 2. This software searches for all of the SSRs with motif lengths up to 10bp; records motif, repeat number and genomic location and reports the results in an output file. DNA sequences The whole genome sequences of Staphylococcus aureus COL (NC_002951), S.aureus MRSA252 (NC_002952), S.aureus Bioinformation 8(24): (2012) Biomedical Informatics
3 MSSA476 (NC_002953), S.aureus Mu50 (NC_002758), S.aureus MW2 (NC_003923) and S.aureus N315 (NC_002745) were downloaded from the GeneBank database. Similarly plasmid sequences of Staphylococcus aureus COL pt181 (NC_006629), Staphylococcus aureus MSSA psas (BX571858), Staphylococcus aureus VRSAp (NC_002774) and Staphylococcus aureus pn315 DNA (AP003139) were downloaded from NCBI GenBank database. Figure 1: Frequency of repeats in (A) genomic; (B) plasmid sequences. Results and Discussion: From the investigated six whole genomes and four plasmids of Staphylococcus aureus strains, the observations were as under: (1) Large numbers of Simple Sequence Repeats were found to be scattered in whole genome and plasmid sequences of Staphylococcus aureus. Shorter repeats were found to be more than longer repeats. High density of repeats was occurred in genomic sequences; (2) The strain Staphylococcus aureus MRSA252 and VRSAp shows high density and more percentage of SSR in genomic and plasmid sequences. The SSR mononucleotide repeats were found to be large in number followed by dinucleotide, trinucleotide and higher motif repeats; (3) The number and percentage of repeats obtained were comparatively more in genomic regions as compare to plasmid sequences of all the studied strains of Staphylococcus; (4) Number of mononucleotide SSRs decreased rapidly with increase in size of repeat unit in all six genomic and four plasmid sequences of Staphylococcus; (5) Total frequency of SSRs in plasmid regions is less than genomic regions; (6) In all investigated strains, ratios of AT/TA repeats were overrepresented over GC/CG repeats in genomics as well as plasmid sequences; (7) Dinucleotide combination of AT dominated all the six Staphylococcus genome sequences. For the comparative genomic analysis of Staphylococcus genomes and plasmids; we searched for SSRs with minimal number of repeats 2 and motif length from 1 to 10 base pairs. The length of whole SSR array was multiples of 2 ie. 2*1 =2. No significantly large SSR with mononucleotide motif was identified among the studied genomic and plasmid sequences. Bioinformation 8(24): (2012) Biomedical Informatics
4 The longest repeat in genomic sequences was T 11 and A 10, which showed similarity with hypothetical proteins and transposase genes. The over-representation of nucleotides was tested among six genomic and four plasmid sequences and the results Table 1 (see supplementary material) showed that AT/TA repeats were dominating. Conclusion: SSR of many types are found in prokaryotic genomes as well. These are present in functional domains and play an important role in functional alterations and implications in mutation helping the organism to adapt to its surroundings. Higher nucleotide repeats have been observed in our study. The environmental changes cause because of stress reactions such as change in copy number of tandem repeats. With this high density of SSR stress response genes and virulent genes may undergo such change resulting in a change in activity of additionally relevant genes and further relaxation of stress by adapting to changed environment [18]. The overrepresentation of A and T mononucleotide SSR can be explained by different ways like slipped strand mispairing, which is more likely for poly A or poly T as strand separation is energetically more favorable compared to poly GC. Similarly the higher energy cost of synthesis of CG dntps by the cells [19]. Longest SSR in sequences was 11bp for T and 10bp for A, whereas plasmids of selected four strains showed 8bp for A, 8bp for T, 6bp for G, and 6bp for T. There is overrepresentation of A and T mononucleotide SSR in plasmids as well as genomic sequences. The frequency of AT/TA dinucleotide repeats was higher in genomic sequences as compared to plasmids (Figure 1). It has been proposed that GT, CA, CT, GA GC or AT repeats binding proteins could participate in recombination process by inducing Z conformation of DNA or other alternative secondary DNA structures. Presence of GC/CG dinucleotide in genome more frequently compared to AT/TA could be due to the fact that TA forms thermodynamically least stable DNA. RNases preferentially degrade UA dinucleotides in mrna. The motifs containing predominantly A and T are found to be over represented in genome. Whereas, the motifs containing predominantly G and C, are found to be over represented in plasmids except in plasmid of strain COL pt181. The over representation of poly (A) and poly (T) mononucleotide repeats in all the Staphylococcus strains could be explained by the fact that strand separation for these poly (A) and poly (T) tracts is considerably easier than for poly (G) or poly(c) tracts, increasing the possibility of slipped strand mispairing. The distribution of tri and hexa-nucleotide repeats reflects codon repetition and that of amino acids suggesting these repeats are strongly selected and shows its association with protein function. The mono nucleotide repeats were over represented in plasmids whereas penta-nucleotide repeats are slightly over represented in genome. The penta nucleotide repeats are present more in genome. Our study suggests that genomic distribution of SSR is non random and apart from nucleotide composition of repeats the characteristic DNA replication, repair and recombination machinery might have important role in the evolution of SSR. Our analyses performed on the genome and plasmid of genes of Staphylococcus aureus clearly indicates that, due to the presence of this large number of SSRs, the organism has an enormous potential for generating this genomic and phenotypic diversity. Though SSRs play an important role in the dynamics of eukaryotes, their presence in bacteria is rare, and mostly reduced to pathogenic organisms. The results found from the present study are difficult to analyse and predict the instability of such small SSRs from sequence alone. Nevertheless, these results can be useful for further experimental studies. References: [1] Edwards AM et al. PLoS Pathog : 6 [PMID: ] [2] Supply P et al. Mol Microbiol : 1003 [PMID: ] [3] Supply P et al. Mol Microbiol : 762 [PMID: ] [4] Xin D et al. Mol Biol Rep : 9047 [PMID: ] [5] Levinson G et al. Mol Biol Evol : 203 [PMID: ] [6] Fodon JW et al. Trends Neurosci : 328 [PMID: ] [7] Jeffreys AJ et al. Biotechnology : 467 [PMID: ] [8] Kumar P et al. J Mol Evol : 127 [PMID: ] [9] Jeffreys AJ et al. Am J Hum Genet : 11 [PMID: ] [10] Field D et al. Proc Biol Sci : 209 [PMID: ] [11] Belkum A et al. Microbiol Mol Biol Rev : 275 [PMID: PMC98915] [12] Gemayel R et al. Annu Rev Genet : 445 [PMID: ] [13] Cox R et al. Proc Natl Acad Sci USA : 5237 [PMID: ] [14] Field D et al. Proc Natl Acad Sci USA : 1647 [PMID: ] [15] Gur-Arie R et al. Genome Res : 62 [PMID: ] [16] Coenye T et al. BMC Genomics : 10 [PMID: ] [17] Yang J et al. Gene : 85 [PMID: ] [18] Trifonov EN, Ann N Y Acad Sci : 330 [PMID: ] [19] Rocha EP et al. Nucleic Acids Res : 1886 [PMID: ] Edited by P Kangueane Citation: Thorat & Thakare, Bioinformation 8(24): (2012) License statement: This is an open-access article, which permits unrestricted use, distribution, and reproduction in any medium, for non-commercial purposes, provided the original author and source are credited Bioinformation 8(24): (2012) Biomedical Informatics
5 Supplementary material: Table 1: Frequency of SSRs in genomic and plasmid of Staphylococcus strains. S. aureus strain Length (bp) Mononucleotide repeats A T G C AT/TA GC/CG COL MRSA MSSA Mu MW N COL pt MSSA psas VRSAp pn315 DNA Bioinformation 8(24): (2012) Biomedical Informatics
15.2 Prokaryotic Transcription *
 OpenStax-CNX module: m52697 1 15.2 Prokaryotic Transcription * Shannon McDermott Based on Prokaryotic Transcription by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
OpenStax-CNX module: m52697 1 15.2 Prokaryotic Transcription * Shannon McDermott Based on Prokaryotic Transcription by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
Horizontal transfer and pathogenicity
 Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
AS A SERVICE TO THE RESEARCH COMMUNITY, GENOME BIOLOGY PROVIDES A 'PREPRINT' DEPOSITORY
 http://genomebiology.com/2002/3/12/preprint/0011.1 This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article MRD: a microsatellite
http://genomebiology.com/2002/3/12/preprint/0011.1 This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article MRD: a microsatellite
(Lys), resulting in translation of a polypeptide without the Lys amino acid. resulting in translation of a polypeptide without the Lys amino acid.
 1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Computational Biology and Chemistry
 Computational Biology and Chemistry 33 (2009) 245 252 Contents lists available at ScienceDirect Computational Biology and Chemistry journal homepage: www.elsevier.com/locate/compbiolchem Research Article
Computational Biology and Chemistry 33 (2009) 245 252 Contents lists available at ScienceDirect Computational Biology and Chemistry journal homepage: www.elsevier.com/locate/compbiolchem Research Article
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss
 Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
International Journal of Scientific & Engineering Research, Volume 4, Issue 12, December-2013 ISSN Sunil S. Thorat and Prashant V.
 International Journal of Scientific & Engineering Research, Volume 4, Issue 12, December-2013 Analysis of Staphylococcus using comparative genomics. Sunil S. Thorat and Prashant V. Thakare Abstract These
International Journal of Scientific & Engineering Research, Volume 4, Issue 12, December-2013 Analysis of Staphylococcus using comparative genomics. Sunil S. Thorat and Prashant V. Thakare Abstract These
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Supplementary Figures.
 Supplementary Figures. Supplementary Figure 1 The extended compartment model. Sub-compartment C (blue) and 1-C (yellow) represent the fractions of allele carriers and non-carriers in the focal patch, respectively,
Supplementary Figures. Supplementary Figure 1 The extended compartment model. Sub-compartment C (blue) and 1-C (yellow) represent the fractions of allele carriers and non-carriers in the focal patch, respectively,
no.1 Raya Ayman Anas Abu-Humaidan
 no.1 Raya Ayman Anas Abu-Humaidan Introduction to microbiology Let's start! As you might have concluded, microbiology is the study of all organisms that are too small to be seen with the naked eye, Ex:
no.1 Raya Ayman Anas Abu-Humaidan Introduction to microbiology Let's start! As you might have concluded, microbiology is the study of all organisms that are too small to be seen with the naked eye, Ex:
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
2012 Univ Aguilera Lecture. Introduction to Molecular and Cell Biology
 2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
Bio 119 Bacterial Genomics 6/26/10
 BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
FINAL VERSION_ Secondary Preservice Teacher Standards -- Life Science AFK12SE/NGSS Strand Disciplinary Core Idea
 Secondary Preservice Teacher Standards -- Life Science AFK12SE/NGSS Strand Disciplinary Core Idea LS1: From Molecules to Organisms: Structures and Processes LS1.A: Structure and Function How do the structures
Secondary Preservice Teacher Standards -- Life Science AFK12SE/NGSS Strand Disciplinary Core Idea LS1: From Molecules to Organisms: Structures and Processes LS1.A: Structure and Function How do the structures
DNA Technology, Bacteria, Virus and Meiosis Test REVIEW
 Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
Ledyard Public Schools Science Curriculum. Biology. Level-2. Instructional Council Approval June 1, 2005
 Ledyard Public Schools Science Curriculum Biology Level-2 1422 Instructional Council Approval June 1, 2005 Suggested Time: Approximately 9 weeks Essential Question Cells & Cell Processes 1. What compounds
Ledyard Public Schools Science Curriculum Biology Level-2 1422 Instructional Council Approval June 1, 2005 Suggested Time: Approximately 9 weeks Essential Question Cells & Cell Processes 1. What compounds
TE content correlates positively with genome size
 TE content correlates positively with genome size Mb 3000 Genomic DNA 2500 2000 1500 1000 TE DNA Protein-coding DNA 500 0 Feschotte & Pritham 2006 Transposable elements. Variation in gene numbers cannot
TE content correlates positively with genome size Mb 3000 Genomic DNA 2500 2000 1500 1000 TE DNA Protein-coding DNA 500 0 Feschotte & Pritham 2006 Transposable elements. Variation in gene numbers cannot
Lecture Notes: BIOL2007 Molecular Evolution
 Lecture Notes: BIOL2007 Molecular Evolution Kanchon Dasmahapatra (k.dasmahapatra@ucl.ac.uk) Introduction By now we all are familiar and understand, or think we understand, how evolution works on traits
Lecture Notes: BIOL2007 Molecular Evolution Kanchon Dasmahapatra (k.dasmahapatra@ucl.ac.uk) Introduction By now we all are familiar and understand, or think we understand, how evolution works on traits
Bio 1B Lecture Outline (please print and bring along) Fall, 2007
 Bio 1B Lecture Outline (please print and bring along) Fall, 2007 B.D. Mishler, Dept. of Integrative Biology 2-6810, bmishler@berkeley.edu Evolution lecture #5 -- Molecular genetics and molecular evolution
Bio 1B Lecture Outline (please print and bring along) Fall, 2007 B.D. Mishler, Dept. of Integrative Biology 2-6810, bmishler@berkeley.edu Evolution lecture #5 -- Molecular genetics and molecular evolution
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Text of objective. Investigate and describe the structure and functions of cells including: Cell organelles
 This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
Reading Assignments. A. Genes and the Synthesis of Polypeptides. Lecture Series 7 From DNA to Protein: Genotype to Phenotype
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Nature Genetics: doi: /ng Supplementary Figure 1. Icm/Dot secretion system region I in 41 Legionella species.
 Supplementary Figure 1 Icm/Dot secretion system region I in 41 Legionella species. Homologs of the effector-coding gene lega15 (orange) were found within Icm/Dot region I in 13 Legionella species. In four
Supplementary Figure 1 Icm/Dot secretion system region I in 41 Legionella species. Homologs of the effector-coding gene lega15 (orange) were found within Icm/Dot region I in 13 Legionella species. In four
Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1
 Name I. Multiple Choice (1 point each) Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1 B 1. Which is possessed by eukaryotes but not by prokaryotes? A. Cell wall B. Distinct nucleus
Name I. Multiple Choice (1 point each) Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1 B 1. Which is possessed by eukaryotes but not by prokaryotes? A. Cell wall B. Distinct nucleus
BIOLOGY STANDARDS BASED RUBRIC
 BIOLOGY STANDARDS BASED RUBRIC STUDENTS WILL UNDERSTAND THAT THE FUNDAMENTAL PROCESSES OF ALL LIVING THINGS DEPEND ON A VARIETY OF SPECIALIZED CELL STRUCTURES AND CHEMICAL PROCESSES. First Semester Benchmarks:
BIOLOGY STANDARDS BASED RUBRIC STUDENTS WILL UNDERSTAND THAT THE FUNDAMENTAL PROCESSES OF ALL LIVING THINGS DEPEND ON A VARIETY OF SPECIALIZED CELL STRUCTURES AND CHEMICAL PROCESSES. First Semester Benchmarks:
THINGS I NEED TO KNOW:
 THINGS I NEED TO KNOW: 1. Prokaryotic and Eukaryotic Cells Prokaryotic cells do not have a true nucleus. In eukaryotic cells, the DNA is surrounded by a membrane. Both types of cells have ribosomes. Some
THINGS I NEED TO KNOW: 1. Prokaryotic and Eukaryotic Cells Prokaryotic cells do not have a true nucleus. In eukaryotic cells, the DNA is surrounded by a membrane. Both types of cells have ribosomes. Some
Frequently Asked Questions (FAQs)
 Frequently Asked Questions (FAQs) Q1. What is meant by Satellite and Repetitive DNA? Ans: Satellite and repetitive DNA generally refers to DNA whose base sequence is repeated many times throughout the
Frequently Asked Questions (FAQs) Q1. What is meant by Satellite and Repetitive DNA? Ans: Satellite and repetitive DNA generally refers to DNA whose base sequence is repeated many times throughout the
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/8/16
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Variation of Traits. genetic variation: the measure of the differences among individuals within a population
 Genetic variability is the measure of the differences among individuals within a population. Because some traits are more suited to certain environments, creating particular niches and fits, we know that
Genetic variability is the measure of the differences among individuals within a population. Because some traits are more suited to certain environments, creating particular niches and fits, we know that
REQUIREMENTS FOR THE BIOCHEMISTRY MAJOR
 REQUIREMENTS FOR THE BIOCHEMISTRY MAJOR Grade Requirement: All courses required for the Biochemistry major (CH, MATH, PHYS, BI courses) must be graded and passed with a grade of C- or better. Core Chemistry
REQUIREMENTS FOR THE BIOCHEMISTRY MAJOR Grade Requirement: All courses required for the Biochemistry major (CH, MATH, PHYS, BI courses) must be graded and passed with a grade of C- or better. Core Chemistry
9/11/18. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Bacterial Genetics & Operons
 Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Drosophila melanogaster and D. simulans, two fruit fly species that are nearly
 Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Department of Mathematics. Mathematical study of competition between Staphylococcus strains within the host and at the host population level
 Department of Mathematics Mathematical study of competition between Staphylococcus strains within the host and at the host population level MATH554: Main Dissertation Written by Nouf Saleh Alghamdi ID
Department of Mathematics Mathematical study of competition between Staphylococcus strains within the host and at the host population level MATH554: Main Dissertation Written by Nouf Saleh Alghamdi ID
Vital Statistics Derived from Complete Genome Sequencing (for E. coli MG1655)
 We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
Hiromi Nishida. 1. Introduction. 2. Materials and Methods
 Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Bio/Life: Cell Biology
 Bio/Life: Cell Biology 1a The fundamental life processes of plants and animals depend on a variety of chemical reactions that occur in specialized areas of the organism's cells. As a basis for understanding
Bio/Life: Cell Biology 1a The fundamental life processes of plants and animals depend on a variety of chemical reactions that occur in specialized areas of the organism's cells. As a basis for understanding
2. What was the Avery-MacLeod-McCarty experiment and why was it significant? 3. What was the Hershey-Chase experiment and why was it significant?
 Name Date Period AP Exam Review Part 6: Molecular Genetics I. DNA and RNA Basics A. History of finding out what DNA really is 1. What was Griffith s experiment and why was it significant? 1 2. What was
Name Date Period AP Exam Review Part 6: Molecular Genetics I. DNA and RNA Basics A. History of finding out what DNA really is 1. What was Griffith s experiment and why was it significant? 1 2. What was
Full file at CHAPTER 2 Genetics
 CHAPTER 2 Genetics MULTIPLE CHOICE 1. Chromosomes are a. small linear bodies. b. contained in cells. c. replicated during cell division. 2. A cross between true-breeding plants bearing yellow seeds produces
CHAPTER 2 Genetics MULTIPLE CHOICE 1. Chromosomes are a. small linear bodies. b. contained in cells. c. replicated during cell division. 2. A cross between true-breeding plants bearing yellow seeds produces
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/1/18
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Molecular Evolution & the Origin of Variation
 Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation
 Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
METHODS FOR DETERMINING PHYLOGENY. In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task.
 Chapter 12 (Strikberger) Molecular Phylogenies and Evolution METHODS FOR DETERMINING PHYLOGENY In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task. Modern
Chapter 12 (Strikberger) Molecular Phylogenies and Evolution METHODS FOR DETERMINING PHYLOGENY In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task. Modern
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
REQUIREMENTS FOR THE BIOCHEMISTRY MAJOR
 REQUIREMENTS FOR THE BIOCHEMISTRY MAJOR Grade Requirement: All courses required for the Biochemistry major (CH, MATH, PHYS, BI courses) must be graded and passed with a grade of C- or better. Core Chemistry
REQUIREMENTS FOR THE BIOCHEMISTRY MAJOR Grade Requirement: All courses required for the Biochemistry major (CH, MATH, PHYS, BI courses) must be graded and passed with a grade of C- or better. Core Chemistry
The Prokaryotic World
 The Prokaryotic World A. An overview of prokaryotic life There is no doubt that prokaryotes are everywhere. By everywhere, I mean living in every geographic region, in extremes of environmental conditions,
The Prokaryotic World A. An overview of prokaryotic life There is no doubt that prokaryotes are everywhere. By everywhere, I mean living in every geographic region, in extremes of environmental conditions,
Flow of Genetic Information
 presents Flow of Genetic Information A Montagud E Navarro P Fernández de Córdoba JF Urchueguía Elements Nucleic acid DNA RNA building block structure & organization genome building block types Amino acid
presents Flow of Genetic Information A Montagud E Navarro P Fernández de Córdoba JF Urchueguía Elements Nucleic acid DNA RNA building block structure & organization genome building block types Amino acid
Genetic Basis of Variation in Bacteria
 Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Niche specific amino acid features within the core genes of the genus Shewanella
 www.bioinformation.net Hypothesis Volume 8(19) Niche specific amino acid features within the core genes of the genus Shewanella Rachana Banerjee* & Subhasis Mukhopadhyay Department of Biophysics, Molecular
www.bioinformation.net Hypothesis Volume 8(19) Niche specific amino acid features within the core genes of the genus Shewanella Rachana Banerjee* & Subhasis Mukhopadhyay Department of Biophysics, Molecular
Multivariate analysis of genetic data: an introduction
 Multivariate analysis of genetic data: an introduction Thibaut Jombart MRC Centre for Outbreak Analysis and Modelling Imperial College London XXIV Simposio Internacional De Estadística Bogotá, 25th July
Multivariate analysis of genetic data: an introduction Thibaut Jombart MRC Centre for Outbreak Analysis and Modelling Imperial College London XXIV Simposio Internacional De Estadística Bogotá, 25th July
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Designer Genes C Test
 Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Chapters AP Biology Objectives. Objectives: You should know...
 Objectives: You should know... Notes 1. Scientific evidence supports the idea that evolution has occurred in all species. 2. Scientific evidence supports the idea that evolution continues to occur. 3.
Objectives: You should know... Notes 1. Scientific evidence supports the idea that evolution has occurred in all species. 2. Scientific evidence supports the idea that evolution continues to occur. 3.
Biology 10 th Grade. Textbook: Biology, Miller and Levine, Pearson (2010) Prerequisite: None
 Biology 10 th Grade SCI 401, 402 Biology 1 credit 5 days a week; 2 semesters Taught in English Biology - The Study of Life! This is a required course for all 10 th grade students in both the Mexican and/or
Biology 10 th Grade SCI 401, 402 Biology 1 credit 5 days a week; 2 semesters Taught in English Biology - The Study of Life! This is a required course for all 10 th grade students in both the Mexican and/or
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
BIOLOGY FINAL EXAM REVIEW SHEET Chapters 10-15, 17-30
 Name Hour Due Date: BIOLOGY FINAL EXAM REVIEW SHEET Chapters 10-15, 17-30 The exam was prepared by the Biology teachers in the science departments of CVHS and DHS. 1. What is a Punnett Square? 2. Cross
Name Hour Due Date: BIOLOGY FINAL EXAM REVIEW SHEET Chapters 10-15, 17-30 The exam was prepared by the Biology teachers in the science departments of CVHS and DHS. 1. What is a Punnett Square? 2. Cross
Grundlagen der Bioinformatik Summer semester Lecturer: Prof. Daniel Huson
 Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
Prokaryotes & Viruses. Practice Questions. Slide 1 / 71. Slide 2 / 71. Slide 3 / 71. Slide 4 / 71. Slide 6 / 71. Slide 5 / 71
 Slide 1 / 71 Slide 2 / 71 New Jersey Center for Teaching and Learning Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of
Slide 1 / 71 Slide 2 / 71 New Jersey Center for Teaching and Learning Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of
OCR Biology Checklist
 Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
20 Viruses and Prokaryotes Bacteria
 20 Viruses and Prokaryotes 20.2 - Bacteria Classifying Prokaryotes Prokaryote unicellular organisms that lacks a nucleus Most abundant and widespread organisms on Earth Divided into two groups Bacteria
20 Viruses and Prokaryotes 20.2 - Bacteria Classifying Prokaryotes Prokaryote unicellular organisms that lacks a nucleus Most abundant and widespread organisms on Earth Divided into two groups Bacteria
Curriculum Map. Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1)
 1 Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1) Focus Standards BIO1.LS1.2 Evaluate comparative models of various cell types with a focus on organic molecules
1 Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1) Focus Standards BIO1.LS1.2 Evaluate comparative models of various cell types with a focus on organic molecules
OCR Biology Checklist
 Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Basic Biology. Content Skills Learning Targets Assessment Resources & Technology
 Teacher: Lynn Dahring Basic Biology August 2014 Basic Biology CEQ (tri 1) 1. What are the parts of the biological scientific process? 2. What are the essential molecules and elements in living organisms?
Teacher: Lynn Dahring Basic Biology August 2014 Basic Biology CEQ (tri 1) 1. What are the parts of the biological scientific process? 2. What are the essential molecules and elements in living organisms?
Genetics 275 Notes Week 7
 Cytoplasmic Inheritance Genetics 275 Notes Week 7 Criteriafor recognition of cytoplasmic inheritance: 1. Reciprocal crosses give different results -mainly due to the fact that the female parent contributes
Cytoplasmic Inheritance Genetics 275 Notes Week 7 Criteriafor recognition of cytoplasmic inheritance: 1. Reciprocal crosses give different results -mainly due to the fact that the female parent contributes
Ap Biology Chapter 17 Packet Answers
 AP BIOLOGY CHAPTER 17 PACKET ANSWERS PDF - Are you looking for ap biology chapter 17 packet answers Books? Now, you will be happy that at this time ap biology chapter 17 packet answers PDF is available
AP BIOLOGY CHAPTER 17 PACKET ANSWERS PDF - Are you looking for ap biology chapter 17 packet answers Books? Now, you will be happy that at this time ap biology chapter 17 packet answers PDF is available
Analysis of correlated mutations in Ras G-domain
 www.bioinformation.net Volume 13(6) Hypothesis Analysis of correlated mutations in Ras G-domain Ekta Pathak * Bioinformatics Department, MMV, Banaras Hindu University. Ekta Pathak - E-mail: ektavpathak@gmail.com;
www.bioinformation.net Volume 13(6) Hypothesis Analysis of correlated mutations in Ras G-domain Ekta Pathak * Bioinformatics Department, MMV, Banaras Hindu University. Ekta Pathak - E-mail: ektavpathak@gmail.com;
Microbiology / Active Lecture Questions Chapter 10 Classification of Microorganisms 1 Chapter 10 Classification of Microorganisms
 1 2 Bergey s Manual of Systematic Bacteriology differs from Bergey s Manual of Determinative Bacteriology in that the former a. groups bacteria into species. b. groups bacteria according to phylogenetic
1 2 Bergey s Manual of Systematic Bacteriology differs from Bergey s Manual of Determinative Bacteriology in that the former a. groups bacteria into species. b. groups bacteria according to phylogenetic
RGP finder: prediction of Genomic Islands
 Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
Introduction to Hidden Markov Models for Gene Prediction ECE-S690
 Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Prokaryotes & Viruses. Multiple Choice Review. Slide 1 / 47. Slide 2 / 47. Slide 3 / 47
 New Jersey enter for Teaching and Learning Slide 1 / 47 Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of students and
New Jersey enter for Teaching and Learning Slide 1 / 47 Progressive Science Initiative This material is made freely available at www.njctl.org and is intended for the non-commercial use of students and
Curriculum Links. AQA GCE Biology. AS level
 Curriculum Links AQA GCE Biology Unit 2 BIOL2 The variety of living organisms 3.2.1 Living organisms vary and this variation is influenced by genetic and environmental factors Causes of variation 3.2.2
Curriculum Links AQA GCE Biology Unit 2 BIOL2 The variety of living organisms 3.2.1 Living organisms vary and this variation is influenced by genetic and environmental factors Causes of variation 3.2.2
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p
 Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids. Prepared by Woojoo Choi
 Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids Prepared by Woojoo Choi Dead or alive? 1) In this chapter we will explore the twilight zone of biology and the gene creature who live there.
Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids Prepared by Woojoo Choi Dead or alive? 1) In this chapter we will explore the twilight zone of biology and the gene creature who live there.
EVOLUTION ALGEBRA Hartl-Clark and Ayala-Kiger
 EVOLUTION ALGEBRA Hartl-Clark and Ayala-Kiger Freshman Seminar University of California, Irvine Bernard Russo University of California, Irvine Winter 2015 Bernard Russo (UCI) EVOLUTION ALGEBRA 1 / 10 Hartl
EVOLUTION ALGEBRA Hartl-Clark and Ayala-Kiger Freshman Seminar University of California, Irvine Bernard Russo University of California, Irvine Winter 2015 Bernard Russo (UCI) EVOLUTION ALGEBRA 1 / 10 Hartl
9/2/17. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
Outline. Collective behavior in bacteria. Know your horsemen. Importance. Cooperation and disease. Medical applications?
 Collective behavior in bacteria Will Driscoll April 30 th, 2008 Outline Importance Our macrobial bias Quorum sensing Biofilms Physiology Development Prokaryotic stab at multicellularity? Discussion But
Collective behavior in bacteria Will Driscoll April 30 th, 2008 Outline Importance Our macrobial bias Quorum sensing Biofilms Physiology Development Prokaryotic stab at multicellularity? Discussion But
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
The two daughter cells are genetically identical to each other and the parent cell.
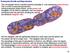 Prokaryote Growth and Reproduction This micrograph shows a bacillus bacteria (probably E. coli) undergoing binary fission. This is a form of asexual reproduction. During prokaryotic binary fission, as
Prokaryote Growth and Reproduction This micrograph shows a bacillus bacteria (probably E. coli) undergoing binary fission. This is a form of asexual reproduction. During prokaryotic binary fission, as
GENETICS - CLUTCH CH.1 INTRODUCTION TO GENETICS.
 !! www.clutchprep.com CONCEPT: HISTORY OF GENETICS The earliest use of genetics was through of plants and animals (8000-1000 B.C.) Selective breeding (artificial selection) is the process of breeding organisms
!! www.clutchprep.com CONCEPT: HISTORY OF GENETICS The earliest use of genetics was through of plants and animals (8000-1000 B.C.) Selective breeding (artificial selection) is the process of breeding organisms
9/8/2017. Bacteria and Archaea. Three domain system: The present tree of life. Structural and functional adaptations contribute to prokaryotic success
 5 m 2 m 9/8/2017 Three domain system: The present tree of life Bacteria and Archaea Chapter 27 Structural and functional adaptations contribute to prokaryotic success Unicellular Small Variety of shapes
5 m 2 m 9/8/2017 Three domain system: The present tree of life Bacteria and Archaea Chapter 27 Structural and functional adaptations contribute to prokaryotic success Unicellular Small Variety of shapes
BACTERIAL PHYSIOLOGY SMALL GROUP. Monday, August 25, :00pm. Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D.
 BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
What Kind Of Molecules Carry Protein Assembly Instructions From The Nucleus To The Cytoplasm
 What Kind Of Molecules Carry Protein Assembly Instructions From The Nucleus To The Cytoplasm What kind of reaction produces large molecules by linking small molecules? molecules carry protein assembly
What Kind Of Molecules Carry Protein Assembly Instructions From The Nucleus To The Cytoplasm What kind of reaction produces large molecules by linking small molecules? molecules carry protein assembly
The Gene The gene; Genes Genes Allele;
 Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Section 19 1 Bacteria (pages )
 Chapter 19 Bacteria and Viruses Section 19 1 Bacteria (pages 471 477) How do the two groups of prokaryotes differ? What factors are used to identify prokaryotes? What is the importance of bacteria? 13.
Chapter 19 Bacteria and Viruses Section 19 1 Bacteria (pages 471 477) How do the two groups of prokaryotes differ? What factors are used to identify prokaryotes? What is the importance of bacteria? 13.
A pathogen is an agent or microrganism that causes a disease in its host. Pathogens can be viruses, bacteria, fungi or protozoa.
 1 A pathogen is an agent or microrganism that causes a disease in its host. Pathogens can be viruses, bacteria, fungi or protozoa. Protozoa are single celled eukaryotic organisms. Some protozoa are pathogens.
1 A pathogen is an agent or microrganism that causes a disease in its host. Pathogens can be viruses, bacteria, fungi or protozoa. Protozoa are single celled eukaryotic organisms. Some protozoa are pathogens.
Bioinformatics Exercises
 Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Chapter 17. From Gene to Protein. Biology Kevin Dees
 Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
Guided Notes: Evolution. is the change in traits through generations over! Occurs in, NOT individual organisms
 Guided Notes: Evolution The Theory of Evolution is the change in traits through generations over! Occurs in, NOT individual organisms How Have Organisms Changed? At the time life emerged, the Earth was
Guided Notes: Evolution The Theory of Evolution is the change in traits through generations over! Occurs in, NOT individual organisms How Have Organisms Changed? At the time life emerged, the Earth was
Molecular evolution - Part 1. Pawan Dhar BII
 Molecular evolution - Part 1 Pawan Dhar BII Theodosius Dobzhansky Nothing in biology makes sense except in the light of evolution Age of life on earth: 3.85 billion years Formation of planet: 4.5 billion
Molecular evolution - Part 1 Pawan Dhar BII Theodosius Dobzhansky Nothing in biology makes sense except in the light of evolution Age of life on earth: 3.85 billion years Formation of planet: 4.5 billion
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Seminar 2 : Good Bugs
 Seminar 2 : Good Bugs Part 2 Viruses What is a virus? Microscopic particles that infect other organisms and can only replicate within a host cell Contain either contain DNA or RNA surrounded by a protective
Seminar 2 : Good Bugs Part 2 Viruses What is a virus? Microscopic particles that infect other organisms and can only replicate within a host cell Contain either contain DNA or RNA surrounded by a protective
BIOINFORMATICS: An Introduction
 BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
From gene to protein. Premedical biology
 From gene to protein Premedical biology Central dogma of Biology, Molecular Biology, Genetics transcription replication reverse transcription translation DNA RNA Protein RNA chemically similar to DNA,
From gene to protein Premedical biology Central dogma of Biology, Molecular Biology, Genetics transcription replication reverse transcription translation DNA RNA Protein RNA chemically similar to DNA,
Classifying Prokaryotes: Eubacteria Plasma Membrane. Ribosomes. Plasmid (DNA) Capsule. Cytoplasm. Outer Membrane DNA. Flagellum.
 Bacteria The yellow band surrounding this hot spring is sulfur, a waste product of extremophilic prokaryotes, probably of the Domain Archaea, Kingdom Archaebacteria. Bacteria are prokaryotic cells (no
Bacteria The yellow band surrounding this hot spring is sulfur, a waste product of extremophilic prokaryotes, probably of the Domain Archaea, Kingdom Archaebacteria. Bacteria are prokaryotic cells (no
ADVANCED PLACEMENT BIOLOGY
 ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
