Niche specific amino acid features within the core genes of the genus Shewanella
|
|
|
- Camron Hill
- 5 years ago
- Views:
Transcription
1 Hypothesis Volume 8(19) Niche specific amino acid features within the core genes of the genus Shewanella Rachana Banerjee* & Subhasis Mukhopadhyay Department of Biophysics, Molecular Biology and Bioinformatics and Distributed Information Centre for Bioinformatics, University of Calcutta, 92, A.P. C. Road Calcutta ; Rachana Banerjee Phone: ; *Corresponding author Received September 15, 2012; Accepted September 19, 2012; Published October 01, 2012 Abstract: Shewanella species are found to dwell in various ecological niches. The widespread habitation where they live requires specific adaptations. Recent advances in genomic approaches, such as in sequencing technologies, generate huge amount of genomic data that lend support towards understanding the microbial evolution and diversity through comparative study. In this manuscript, we discuss a comparative analysis of core genes of phylogenetically related twelve members from the genus Shewanella. Phylogenetic analysis based on the core genes, differentiated two subgroups of the genus, one group comprises of species characterized as highpressure cold-adapted while the other group is characterized as mesophilic pressure-sensitive species. By analyzing the differences of amino acid composition of these two groups, we have identified the specific trend of amino acid usage that has been adopted by the psychro-peizo-tolerant Shewanella species. The functional categories have also been recognized which are responsible for rendering the particular amino acid compositional pattern in psychropeizophilic Shewanella species facilitating their niche adaptation. Keywords: Shewanella sp, core gene, amino acid composition, phylogenetic tree Background: The increasing number of complete genomic sequences has unwrapped loads of ways to interpret the adaptation of genomes in their respective niche in relation to their structure, function etc. [1]. An estimation of amino acid preferences of different organisms is now possible through comparative genomic study [2]. Comparative genomics helps to determine the genes that are present across bacterial genomes of the same species (or genus) known as conserved (core) genes and disparity within these core genes is an indication that can be used to unravel highly variable and conserved genes within the genomes of interest. The conserved genes usually evolve more slowly; therefore, they will play a role to infer molecular evolution of the genomes [3]. Moreover, molecular phylogenetic trees built using the sequences of core genes can also determine the relationship among the genomes. Shewanella is a genus from which adequate number of species had already been sequenced facilitating detection of core genes. Different species belonging to the genus Shewanella are capable of inhabiting many aquatic (from fresh water to deep sea) and sedimentary ecosystems under aerobic as well as anaerobic conditions [4]. Some of the Shewanella species, for example Shewanella violacea, had been isolated from cold environments, such as seawater in Antarctica or in the North Sea, implying that they are not only peizophilic (can breed better under high hydrostatic pressure conditions), but also psychrophilic (needs low temperatures, ranging from 15 C to +10 C for growth and breeding). They can be defined as psychro-peizo-tolerant Shewanella species. It is known that pressure influences protein structure and folding [5]. Functional properties of proteins might get affected by pressure range that is experienced by organism [6]. In this Bioinformation 8(19): (2012) Biomedical Informatics
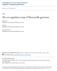
2 study, we discuss the comparative analysis of the core genes of phylogenetically related twelve members from the genus Shewanella and focus on the recognition of a common trend towards a certain pattern of amino acid usage of the psychropeizo-tolerant Shewanella species. Methodology: We have downloaded the fully annotated complete nucleotide sequences of all the twelve Shewanella genomes, considered in the present study, from NCBI FTP site ( Details of the 12 bacteria are listed in Table 2 (see supplementary material). core proteins (F Group2) at 90% or higher confidence level. If the frequency of residue or property group t-value is negative and less than , then the mean frequency of Group1 core proteins are (F Group1) is significantly less than that of Group2 core proteins (F Group2) at 90% or higher confidence level [8-10]. Functional classification The core proteins were functionally classified according to Clusters of Orthologous Groups (COGs) of proteins categories. BLAST homology study had been carried out against the COG database. Proteins that were classified in two COG categories were registered in both categories. Identification of core genes Highly similar paralogous sequences have been removed from each of the twelve set of genome sequences, based on a comparison of the gene list against itself (with identity >=90%) using Blastclust program ( ). From the rest of the genes, core genes were detected through putative one-to-one orthologous gene identification, using BLASTP (>= 85% identity and 0% gap). This search identified a total of 121 orthologous groups that had all twelve species represented, which were used for further analysis. Phylogenetic analysis All core genes from 12 Shewanella genomes were used for generating a neighbor joining tree. For each twelve species, the amino acid sequences of 121 core gene sets were concatenated in order to produce twelve continuous sequences of average length 34,746 amino acids. Multiple Alignments of these 12 sets of concatenated core amino acid sequences were performed using CLUSTALW ( A Neighbor-joining Phylogenetic tree based on these concatenated core amino acid sequences from the Shewanella species of interest, was generated using MEGA 5 [7].The bootstrap values are shown in the Phylogenetic tree. Estimation of amino acid composition We have computed the frequencies of amino acid residues in the protein sequences of two groups of Shewanella sp., as depicted in the Phylogenetic tree. To compare the means of two groups of data, t-test is essentially a good tool. The t-values are calculated as follows: t = (F Group1 - F Group2) / [(Var Group1/ n Group1) + (Var Group2/ n Group2)] where, Var Group1 and Var Group2 are the variance of amino acid residues. F Group1 and F Group2 are mean frequencies of Group1 and Group2 proteomes respectively. The n Group1 and n Group2 are the total number of Group1 and Group2 proteins investigated in this study. Based on student's t-distribution table of significance, critical values for such t-test at various probabilities are as follows: Probability 10% 5% 2.5% 2% 1% 0.5% Critical Value If t-value is positive and greater than critical value at 10% probability (1.372), then the mean frequency (F Group1) of Group1 core proteins are significantly greater than that of the Group2 Figure 1: Phylogenetic tree constructed using core genes of 12 Shewanella species.group1 consists of psychro-peizo-tolerant Shewanella species and Group2 contains mesophilic pressuresensitive Shewanella species. Bootstrap values for all the branches are mentioned in the figure. Results and Discussion: A final list containing a total of 121 core gene sets from all the twelve Shewanella species, were used for further analyses. Alignment of these 12 sets of concatenated core amino acid sequences using CLUSTALW ( reveals 17% non-conserved amino acid sites. Neighbor-joining phylogenetic tree based on the concatenated core amino acid sequences from the Shewanella species of interest, was constructed using MEGA 5, which separates 12 Shewanella species into two groups i.e., Group 1 and Group 2 (Figure 1). Literature search describes Group 1 members of Shewanella as cold adapted species that grow at high pressure, while Group2 members are mesophilic, pressure sensitive species [11]. Amino acid composition preferences We have analyzed the amino acid composition of 121 core gene sequence sets for each of the twelve Shewanella species. The frequencies of individual amino acids were further analyzed using student t-test, which shows that a few of the amino acid differed significantly in the core gene sets of Shewanella species present in Group1 when compared to the core gene sets of Shewanella species present in Group 2 Table 1 (see supplementary material). As specified by the t-value, amino acid residues A, D, S, N and C, are significantly preferred (marked in bold red in Table 1), while G, T, V, M, I are moderately preferred and residues R, Y, E, P and L are significantly avoided (marked in blue in Table 1) by Group 1 Shewanella species compared to Group 2. Residues S and D are Bioinformation 8(19): (2012) Biomedical Informatics
3 helix breakers and residue E favors the formation of helical structure. On the other hand, presence of residue L stabilizes the helical conformations [12]. It is known that amino acid substitutions that diminish protein flexibility and compressibility results in an increase of stability of the protein at high pressure [13]. In addition, helix destabilizing beta branch residues I, T and V are preferred by peizophilic proteins [14]. All the above results signify that increased structural homogeneity is perhaps favored by the high-pressure environments of the deep-sea, which is attained by favoring a decreased number of helix breaking and helix destabilizing residues. Considering the amino acid properties, Group1 Shewanella members most significantly, favor the amino acid residues S, D, and N, all of which are strongly polar as well as have very low molecular weight. Interestingly, the hydrostatic pressure asymmetry index is positively correlated with the polarity of amino acids and inversely correlated with molecular weight of amino acids [15]. Thus, the amino acid composition of the core genes of the twelve species of Shewanella considered in our study shows a particular trend, which points towards a strong favor of polar and small amino acids with adequate propensity of breaking and destabilizing the helical structure, for the Group 1 members of Shewanella (Figure 1) sustaining their psychropeizophilic adaptation for residing in deep sea environment, while an opposite amino acid compositional trend is featured by the Group 2 members of Shewanella (Figure 1) supporting their mesophilic, pressure sensitive characteristics. Figure 2: Distribution of COG classification of 121 core genes in twelve Shewanella species considered in this study. Functional Classification Core genes considered in our study can be divided into two categories depending on the average percent identity of the blast score. (a) Core gene showing low variation (54% of the total core genes with identity >95%); (b) Core gene that are highly variable (46% of the total core genes with identity <95%). Functional profiles of the core genes of the twelve Shewanella species considered in our study have been determined based on COG categories. It has been found that higher proportion of the core genes (44%) are involved in information storage and processing, 62% of which are involved in translation, ribosomal structure and biogenesis (Figure 2). Variable core genes as well as conserved core genes are classified according to COG functional categories and compared. The comparison points to the fact that functional categories like Translation, ribosomal structure and biogenesis, Carbohydrate transport and metabolism and Amino acid transport and metabolism are more common in conserved core genes (Figure 3). On the other hand, functional categories like Energy production and conversion, Cell envelope biogenesis and Cell motility and secretion are more common in variable core genes. Core genes with low variation experience selection against mutations that leads to amino acid changes. But in case of highly variable core genes, positive selection for amino acid changes takes place [16]. Consequently, three functional categories which are more common in variable core genes are accountable for the specific amino acid compositional trend in Bioinformation 8(19): (2012) Biomedical Informatics
4 psychropeizophilic Shewanella species (members of Group 1), favoring polar and small amino acids with helix breaking and destabilizing property. them involved in translation, ribosomal structure and biogenesis, signifying the importance of these two functional categories in maintaining the most important cellular processes of the genus Shewanella. We have also divided the core genes into two types: conserved core genes and variable core genes, which are evaluated on the basis of COG functional classification. Three functional categories (Cell envelope biogenesis, Cell motility and secretion, Energy production and conversion) which are quite more common in variable core genes are responsible for displaying the specific amino acid compositional trend in psychropeizophilic Shewanella species (members of Group 1). Thus, they seem to have played an important role for niche adaptation of the twelve Shewanella species considered in our study. Acknowledgement: We wish to thank Department of Biotechnology, Government of India, for providing financial support through two project grants, (BT/BI/04/026/93 and BT/BI/010/019/99). Figure 3: COG classification terms for conserved core genes (red bars) and highly variable core genes (blue bars). Functional classes like Translation, ribosomal structure and biogenesis (J), Carbohydrate transport and metabolism (G), Amino acid transport and metabolism (E), are more common in conserved core genes. Variable core genes have a preference over functional categories like Energy production and conversion (C), Cell envelope biogenesis (M) and Cell motility and secretion (N). Conclusion: Phylogenetic study based on the concatenated core amino acid sequences of twelve Shewanella species separated two distinct groups of Shewanella. Group 1 comprises of psychropeizophilic Shewanella species, whereas Group2 members are mesophilic, pressure sensitive species. Our studies on the composition of individual amino acid residues within these two groups of bacteria revealed that the psychropeizophilic Shewanella species show a specific trend of amino acid usage that favors the increase in frequency of strongly polar, small and tiny amino acids having the potential of avoiding helical structures. Amino acid residues S, D and N are mostly preferred by them. Functional profiles of the core genes of the twelve Shewanella species show that information storage and processing represents higher proportion of the core genes, with 62% of References: [1] Medigue C et al. Genome Res : 1325 [PMID: ] [2] Saunders NF et al. Genome Res : 1580 [PMID: ] [3] Urwin R et al. Trends Microbiol : 479 [PMID: ] [4] Venkateswaran K et al. Int J Syst Bacteriol : 705 [PMID: ] [5] Gross M & Jaenicke R, Eur J Biochem : 617 [PMID: ] [6] Li H et al. J Agric Food Chem : [PMID: ] [7] Tamura K et al. Mol Biol Evol : 2731 [PMID: ] [8] Jahandideh S et al. J Theor Biol : 159 [PMID: ] [9] Pack SP & Yoo YJ, J Biotechnol : 269 [PMID: ] [10] Vogt G et al. J Mol Biol : 816 [PMID: ] [11] Kato C & Nogi Y, FEMS Microbiol Ecol : 223 [PMID: ] [12] Lewis PN et al. Proc Natl Acad Sci U S A : 810 [PMID: ] [13] Royer CA et al. Biochemistry : 5222 [PMID: ] [14] Kumar S et al. Protein Eng : 179 [PMID: ] [15] Simonato F et al. J Biotechnol : 11 [PMID: ] [16] Leekitcharoenphon P et al. BMC Genomics : 88 [PMID: ] Edited by P Kangueane Citation: Banerjee & Mukhopadhyay, Bioinformation 8(19): (2012) License statement: This is an open-access article, which permits unrestricted use, distribution, and reproduction in any medium, for non-commercial purposes, provided the original author and source are credited Bioinformation 8(19): (2012) Biomedical Informatics
5 Supplementary material: Table 1: The composition of individual amino acids in protein sequences of core genes of Group1 and Group2 Shewanella sp. Groups are referred as depicted by the phylogenetic tree in Figure 1. Significantly preferred and avoided amino acids by Group1 members of Shewanella, as indicated by the t-test parameters are marked in bold red and bold blue respectively. Group 1 Group 2 Amio acids Average Variance Average Variance t-test PHE [F] LEU [L] ILE [I] MET [M] VAL [V] SER [S] PRO [P] THR [T] ALA [A] TYR [Y] HIS [H] GLN [Q] ASN [N] LYS [K] ASP [D] GLU [E] CYS [C] TRP [W] ARG [R] GLY [G] Table 2: Information about Shewanella strains used in this study Genome Name NCBI Taxon ID RefSeq Project ID GenBank Project ID Genome Size CDS Count Shewanella amazonensis SB2B Shewanella baltica OS Shewanella frigidimarina NCIMB Shewanella halifaxensis HAW-EB Shewanella loihica PV Shewanella oneidensis MR Shewanella pealeana ANG-SQ1, ATCC Shewanella piezotolerans WP Shewanella putrefaciens CN Shewanella sediminis HAW-EB Shewanella violacea DSS Shewanella woodyi MS32, ATCC Bioinformation 8(19): (2012) Biomedical Informatics
Secondary Structure. Bioch/BIMS 503 Lecture 2. Structure and Function of Proteins. Further Reading. Φ, Ψ angles alone determine protein structure
 Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Protein structure. Protein structure. Amino acid residue. Cell communication channel. Bioinformatics Methods
 Cell communication channel Bioinformatics Methods Iosif Vaisman Email: ivaisman@gmu.edu SEQUENCE STRUCTURE DNA Sequence Protein Sequence Protein Structure Protein structure ATGAAATTTGGAAACTTCCTTCTCACTTATCAGCCACCT...
Cell communication channel Bioinformatics Methods Iosif Vaisman Email: ivaisman@gmu.edu SEQUENCE STRUCTURE DNA Sequence Protein Sequence Protein Structure Protein structure ATGAAATTTGGAAACTTCCTTCTCACTTATCAGCCACCT...
Packing of Secondary Structures
 7.88 Lecture Notes - 4 7.24/7.88J/5.48J The Protein Folding and Human Disease Professor Gossard Retrieving, Viewing Protein Structures from the Protein Data Base Helix helix packing Packing of Secondary
7.88 Lecture Notes - 4 7.24/7.88J/5.48J The Protein Folding and Human Disease Professor Gossard Retrieving, Viewing Protein Structures from the Protein Data Base Helix helix packing Packing of Secondary
Protein Structures: Experiments and Modeling. Patrice Koehl
 Protein Structures: Experiments and Modeling Patrice Koehl Structural Bioinformatics: Proteins Proteins: Sources of Structure Information Proteins: Homology Modeling Proteins: Ab initio prediction Proteins:
Protein Structures: Experiments and Modeling Patrice Koehl Structural Bioinformatics: Proteins Proteins: Sources of Structure Information Proteins: Homology Modeling Proteins: Ab initio prediction Proteins:
Physiochemical Properties of Residues
 Physiochemical Properties of Residues Various Sources C N Cα R Slide 1 Conformational Propensities Conformational Propensity is the frequency in which a residue adopts a given conformation (in a polypeptide)
Physiochemical Properties of Residues Various Sources C N Cα R Slide 1 Conformational Propensities Conformational Propensity is the frequency in which a residue adopts a given conformation (in a polypeptide)
Clustering and Model Integration under the Wasserstein Metric. Jia Li Department of Statistics Penn State University
 Clustering and Model Integration under the Wasserstein Metric Jia Li Department of Statistics Penn State University Clustering Data represented by vectors or pairwise distances. Methods Top- down approaches
Clustering and Model Integration under the Wasserstein Metric Jia Li Department of Statistics Penn State University Clustering Data represented by vectors or pairwise distances. Methods Top- down approaches
PROTEIN SECONDARY STRUCTURE PREDICTION: AN APPLICATION OF CHOU-FASMAN ALGORITHM IN A HYPOTHETICAL PROTEIN OF SARS VIRUS
 Int. J. LifeSc. Bt & Pharm. Res. 2012 Kaladhar, 2012 Research Paper ISSN 2250-3137 www.ijlbpr.com Vol.1, Issue. 1, January 2012 2012 IJLBPR. All Rights Reserved PROTEIN SECONDARY STRUCTURE PREDICTION:
Int. J. LifeSc. Bt & Pharm. Res. 2012 Kaladhar, 2012 Research Paper ISSN 2250-3137 www.ijlbpr.com Vol.1, Issue. 1, January 2012 2012 IJLBPR. All Rights Reserved PROTEIN SECONDARY STRUCTURE PREDICTION:
Introduction to Comparative Protein Modeling. Chapter 4 Part I
 Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Sequence Alignments. Dynamic programming approaches, scoring, and significance. Lucy Skrabanek ICB, WMC January 31, 2013
 Sequence Alignments Dynamic programming approaches, scoring, and significance Lucy Skrabanek ICB, WMC January 31, 213 Sequence alignment Compare two (or more) sequences to: Find regions of conservation
Sequence Alignments Dynamic programming approaches, scoring, and significance Lucy Skrabanek ICB, WMC January 31, 213 Sequence alignment Compare two (or more) sequences to: Find regions of conservation
Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase Cwc27
 Acta Cryst. (2014). D70, doi:10.1107/s1399004714021695 Supporting information Volume 70 (2014) Supporting information for article: Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase
Acta Cryst. (2014). D70, doi:10.1107/s1399004714021695 Supporting information Volume 70 (2014) Supporting information for article: Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
What makes a good graphene-binding peptide? Adsorption of amino acids and peptides at aqueous graphene interfaces: Electronic Supplementary
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 21 What makes a good graphene-binding peptide? Adsorption of amino acids and
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 21 What makes a good graphene-binding peptide? Adsorption of amino acids and
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
Supplemental Materials for. Structural Diversity of Protein Segments Follows a Power-law Distribution
 Supplemental Materials for Structural Diversity of Protein Segments Follows a Power-law Distribution Yoshito SAWADA and Shinya HONDA* National Institute of Advanced Industrial Science and Technology (AIST),
Supplemental Materials for Structural Diversity of Protein Segments Follows a Power-law Distribution Yoshito SAWADA and Shinya HONDA* National Institute of Advanced Industrial Science and Technology (AIST),
Phylogenetic and Comparative Sequence Analysis of Thermostable Alpha Amylases of kingdom Archea, Prokaryotes and Eukaryotes
 www.bioinformation.net Hypothesis Volume 10(7) Phylogenetic and Comparative Sequence Analysis of Thermostable Alpha Amylases of kingdom Archea, Prokaryotes and Eukaryotes Tayyaba Huma 1 *, Arooma Maryam
www.bioinformation.net Hypothesis Volume 10(7) Phylogenetic and Comparative Sequence Analysis of Thermostable Alpha Amylases of kingdom Archea, Prokaryotes and Eukaryotes Tayyaba Huma 1 *, Arooma Maryam
Protein Fragment Search Program ver Overview: Contents:
 Protein Fragment Search Program ver 1.1.1 Developed by: BioPhysics Laboratory, Faculty of Life and Environmental Science, Shimane University 1060 Nishikawatsu-cho, Matsue-shi, Shimane, 690-8504, Japan
Protein Fragment Search Program ver 1.1.1 Developed by: BioPhysics Laboratory, Faculty of Life and Environmental Science, Shimane University 1060 Nishikawatsu-cho, Matsue-shi, Shimane, 690-8504, Japan
Lecture 15: Realities of Genome Assembly Protein Sequencing
 Lecture 15: Realities of Genome Assembly Protein Sequencing Study Chapter 8.10-8.15 1 Euler s Theorems A graph is balanced if for every vertex the number of incoming edges equals to the number of outgoing
Lecture 15: Realities of Genome Assembly Protein Sequencing Study Chapter 8.10-8.15 1 Euler s Theorems A graph is balanced if for every vertex the number of incoming edges equals to the number of outgoing
Proteins: Characteristics and Properties of Amino Acids
 SBI4U:Biochemistry Macromolecules Eachaminoacidhasatleastoneamineandoneacidfunctionalgroupasthe nameimplies.thedifferentpropertiesresultfromvariationsinthestructuresof differentrgroups.thergroupisoftenreferredtoastheaminoacidsidechain.
SBI4U:Biochemistry Macromolecules Eachaminoacidhasatleastoneamineandoneacidfunctionalgroupasthe nameimplies.thedifferentpropertiesresultfromvariationsinthestructuresof differentrgroups.thergroupisoftenreferredtoastheaminoacidsidechain.
Research Proposal. Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family.
 Research Proposal Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family. Name: Minjal Pancholi Howard University Washington, DC. June 19, 2009 Research
Research Proposal Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family. Name: Minjal Pancholi Howard University Washington, DC. June 19, 2009 Research
CSE 549: Computational Biology. Substitution Matrices
 CSE 9: Computational Biology Substitution Matrices How should we score alignments So far, we ve looked at arbitrary schemes for scoring mutations. How can we assign scores in a more meaningful way? Are
CSE 9: Computational Biology Substitution Matrices How should we score alignments So far, we ve looked at arbitrary schemes for scoring mutations. How can we assign scores in a more meaningful way? Are
Sequences, Structures, and Gene Regulatory Networks
 Sequences, Structures, and Gene Regulatory Networks Learning Outcomes After this class, you will Understand gene expression and protein structure in more detail Appreciate why biologists like to align
Sequences, Structures, and Gene Regulatory Networks Learning Outcomes After this class, you will Understand gene expression and protein structure in more detail Appreciate why biologists like to align
Heteropolymer. Mostly in regular secondary structure
 Heteropolymer - + + - Mostly in regular secondary structure 1 2 3 4 C >N trace how you go around the helix C >N C2 >N6 C1 >N5 What s the pattern? Ci>Ni+? 5 6 move around not quite 120 "#$%&'!()*(+2!3/'!4#5'!1/,#64!#6!,6!
Heteropolymer - + + - Mostly in regular secondary structure 1 2 3 4 C >N trace how you go around the helix C >N C2 >N6 C1 >N5 What s the pattern? Ci>Ni+? 5 6 move around not quite 120 "#$%&'!()*(+2!3/'!4#5'!1/,#64!#6!,6!
Similarity or Identity? When are molecules similar?
 Similarity or Identity? When are molecules similar? Mapping Identity A -> A T -> T G -> G C -> C or Leu -> Leu Pro -> Pro Arg -> Arg Phe -> Phe etc If we map similarity using identity, how similar are
Similarity or Identity? When are molecules similar? Mapping Identity A -> A T -> T G -> G C -> C or Leu -> Leu Pro -> Pro Arg -> Arg Phe -> Phe etc If we map similarity using identity, how similar are
The cis-regulatory map of Shewanella genomes
 Washington University School of Medicine Digital Commons@Becker Open Access Publications 2008 The cis-regulatory map of Shewanella genomes Jiajian Liu Washington University School of Medicine in St. Louis
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2008 The cis-regulatory map of Shewanella genomes Jiajian Liu Washington University School of Medicine in St. Louis
Course Notes: Topics in Computational. Structural Biology.
 Course Notes: Topics in Computational Structural Biology. Bruce R. Donald June, 2010 Copyright c 2012 Contents 11 Computational Protein Design 1 11.1 Introduction.........................................
Course Notes: Topics in Computational Structural Biology. Bruce R. Donald June, 2010 Copyright c 2012 Contents 11 Computational Protein Design 1 11.1 Introduction.........................................
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss
 Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
Supplementary Information for Hurst et al.: Causes of trends of amino acid gain and loss Methods Identification of orthologues, alignment and evolutionary distances A preliminary set of orthologues was
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Major Types of Association of Proteins with Cell Membranes. From Alberts et al
 Major Types of Association of Proteins with Cell Membranes From Alberts et al Proteins Are Polymers of Amino Acids Peptide Bond Formation Amino Acid central carbon atom to which are attached amino group
Major Types of Association of Proteins with Cell Membranes From Alberts et al Proteins Are Polymers of Amino Acids Peptide Bond Formation Amino Acid central carbon atom to which are attached amino group
Bioinformatics Exercises
 Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Sequence Based Bioinformatics
 Structural and Functional Analysis of Inosine Monophosphate Dehydrogenase using Sequence-Based Bioinformatics Barry Sexton 1,2 and Troy Wymore 3 1 Bioengineering and Bioinformatics Summer Institute, Department
Structural and Functional Analysis of Inosine Monophosphate Dehydrogenase using Sequence-Based Bioinformatics Barry Sexton 1,2 and Troy Wymore 3 1 Bioengineering and Bioinformatics Summer Institute, Department
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Supplementary Information. Broad Spectrum Anti-Influenza Agents by Inhibiting Self- Association of Matrix Protein 1
 Supplementary Information Broad Spectrum Anti-Influenza Agents by Inhibiting Self- Association of Matrix Protein 1 Philip D. Mosier 1, Meng-Jung Chiang 2, Zhengshi Lin 2, Yamei Gao 2, Bashayer Althufairi
Supplementary Information Broad Spectrum Anti-Influenza Agents by Inhibiting Self- Association of Matrix Protein 1 Philip D. Mosier 1, Meng-Jung Chiang 2, Zhengshi Lin 2, Yamei Gao 2, Bashayer Althufairi
Supplemental Materials
 JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but not the only one!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids repeated
Properties of amino acids in proteins one of the primary roles of DNA (but not the only one!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids repeated
Protein Structure Prediction and Display
 Protein Structure Prediction and Display Goal Take primary structure (sequence) and, using rules derived from known structures, predict the secondary structure that is most likely to be adopted by each
Protein Structure Prediction and Display Goal Take primary structure (sequence) and, using rules derived from known structures, predict the secondary structure that is most likely to be adopted by each
Analysis of correlated mutations in Ras G-domain
 www.bioinformation.net Volume 13(6) Hypothesis Analysis of correlated mutations in Ras G-domain Ekta Pathak * Bioinformatics Department, MMV, Banaras Hindu University. Ekta Pathak - E-mail: ektavpathak@gmail.com;
www.bioinformation.net Volume 13(6) Hypothesis Analysis of correlated mutations in Ras G-domain Ekta Pathak * Bioinformatics Department, MMV, Banaras Hindu University. Ekta Pathak - E-mail: ektavpathak@gmail.com;
Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India. 1 st November, 2013
 Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Using Higher Calculus to Study Biologically Important Molecules Julie C. Mitchell
 Using Higher Calculus to Study Biologically Important Molecules Julie C. Mitchell Mathematics and Biochemistry University of Wisconsin - Madison 0 There Are Many Kinds Of Proteins The word protein comes
Using Higher Calculus to Study Biologically Important Molecules Julie C. Mitchell Mathematics and Biochemistry University of Wisconsin - Madison 0 There Are Many Kinds Of Proteins The word protein comes
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Chapter 26: Phylogeny and the Tree of Life Phylogenies Show Evolutionary Relationships
 Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination
 Lecture 9 M230 Feigon Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination Reading resources v Roberts NMR of Macromolecules, Chap 4 by Christina Redfield
Lecture 9 M230 Feigon Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination Reading resources v Roberts NMR of Macromolecules, Chap 4 by Christina Redfield
UNIT TWELVE. a, I _,o "' I I I. I I.P. l'o. H-c-c. I ~o I ~ I / H HI oh H...- I II I II 'oh. HO\HO~ I "-oh
 UNT TWELVE PROTENS : PEPTDE BONDNG AND POLYPEPTDES 12 CONCEPTS Many proteins are important in biological structure-for example, the keratin of hair, collagen of skin and leather, and fibroin of silk. Other
UNT TWELVE PROTENS : PEPTDE BONDNG AND POLYPEPTDES 12 CONCEPTS Many proteins are important in biological structure-for example, the keratin of hair, collagen of skin and leather, and fibroin of silk. Other
Protein Structure. W. M. Grogan, Ph.D. OBJECTIVES
 Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
BLAST. Varieties of BLAST
 BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
April, The energy functions include:
 REDUX A collection of Python scripts for torsion angle Monte Carlo protein molecular simulations and analysis The program is based on unified residue peptide model and is designed for more efficient exploration
REDUX A collection of Python scripts for torsion angle Monte Carlo protein molecular simulations and analysis The program is based on unified residue peptide model and is designed for more efficient exploration
Introduction to Evolutionary Concepts
 Introduction to Evolutionary Concepts and VMD/MultiSeq - Part I Zaida (Zan) Luthey-Schulten Dept. Chemistry, Beckman Institute, Biophysics, Institute of Genomics Biology, & Physics NIH Workshop 2009 VMD/MultiSeq
Introduction to Evolutionary Concepts and VMD/MultiSeq - Part I Zaida (Zan) Luthey-Schulten Dept. Chemistry, Beckman Institute, Biophysics, Institute of Genomics Biology, & Physics NIH Workshop 2009 VMD/MultiSeq
Supplementary Information
 Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Gene regulation II Biochemistry 302. February 27, 2006
 Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Bioinformatics. Dept. of Computational Biology & Bioinformatics
 Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Massachusetts Institute of Technology Computational Evolutionary Biology, Fall, 2005 Notes for November 7: Molecular evolution
 Massachusetts Institute of Technology 6.877 Computational Evolutionary Biology, Fall, 2005 Notes for November 7: Molecular evolution 1. Rates of amino acid replacement The initial motivation for the neutral
Massachusetts Institute of Technology 6.877 Computational Evolutionary Biology, Fall, 2005 Notes for November 7: Molecular evolution 1. Rates of amino acid replacement The initial motivation for the neutral
Supplementary Figure 3 a. Structural comparison between the two determined structures for the IL 23:MA12 complex. The overall RMSD between the two
 Supplementary Figure 1. Biopanningg and clone enrichment of Alphabody binders against human IL 23. Positive clones in i phage ELISA with optical density (OD) 3 times higher than background are shown for
Supplementary Figure 1. Biopanningg and clone enrichment of Alphabody binders against human IL 23. Positive clones in i phage ELISA with optical density (OD) 3 times higher than background are shown for
Protein Data Bank Contents Guide: Atomic Coordinate Entry Format Description. Version Document Published by the wwpdb
 Protein Data Bank Contents Guide: Atomic Coordinate Entry Format Description Version 3.30 Document Published by the wwpdb This format complies with the PDB Exchange Dictionary (PDBx) http://mmcif.pdb.org/dictionaries/mmcif_pdbx.dic/index/index.html.
Protein Data Bank Contents Guide: Atomic Coordinate Entry Format Description Version 3.30 Document Published by the wwpdb This format complies with the PDB Exchange Dictionary (PDBx) http://mmcif.pdb.org/dictionaries/mmcif_pdbx.dic/index/index.html.
Microbial Taxonomy and the Evolution of Diversity
 19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
Homology Modeling. Roberto Lins EPFL - summer semester 2005
 Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Bahnson Biochemistry Cume, April 8, 2006 The Structural Biology of Signal Transduction
 Name page 1 of 6 Bahnson Biochemistry Cume, April 8, 2006 The Structural Biology of Signal Transduction Part I. The ion Ca 2+ can function as a 2 nd messenger. Pick a specific signal transduction pathway
Name page 1 of 6 Bahnson Biochemistry Cume, April 8, 2006 The Structural Biology of Signal Transduction Part I. The ion Ca 2+ can function as a 2 nd messenger. Pick a specific signal transduction pathway
BIOINFORMATICS: An Introduction
 BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
DATE A DAtabase of TIM Barrel Enzymes
 DATE A DAtabase of TIM Barrel Enzymes 2 2.1 Introduction.. 2.2 Objective and salient features of the database 2.2.1 Choice of the dataset.. 2.3 Statistical information on the database.. 2.4 Features....
DATE A DAtabase of TIM Barrel Enzymes 2 2.1 Introduction.. 2.2 Objective and salient features of the database 2.2.1 Choice of the dataset.. 2.3 Statistical information on the database.. 2.4 Features....
Journal of Proteomics & Bioinformatics - Open Access
 Abstract Methodology for Phylogenetic Tree Construction Kudipudi Srinivas 2, Allam Appa Rao 1, GR Sridhar 3, Srinubabu Gedela 1* 1 International Center for Bioinformatics & Center for Biotechnology, Andhra
Abstract Methodology for Phylogenetic Tree Construction Kudipudi Srinivas 2, Allam Appa Rao 1, GR Sridhar 3, Srinubabu Gedela 1* 1 International Center for Bioinformatics & Center for Biotechnology, Andhra
Amino Acid Side Chain Induced Selectivity in the Hydrolysis of Peptides Catalyzed by a Zr(IV)-Substituted Wells-Dawson Type Polyoxometalate
 Amino Acid Side Chain Induced Selectivity in the Hydrolysis of Peptides Catalyzed by a Zr(IV)-Substituted Wells-Dawson Type Polyoxometalate Stef Vanhaecht, Gregory Absillis, Tatjana N. Parac-Vogt* Department
Amino Acid Side Chain Induced Selectivity in the Hydrolysis of Peptides Catalyzed by a Zr(IV)-Substituted Wells-Dawson Type Polyoxometalate Stef Vanhaecht, Gregory Absillis, Tatjana N. Parac-Vogt* Department
Protein Secondary Structure Prediction using Feed-Forward Neural Network
 COPYRIGHT 2010 JCIT, ISSN 2078-5828 (PRINT), ISSN 2218-5224 (ONLINE), VOLUME 01, ISSUE 01, MANUSCRIPT CODE: 100713 Protein Secondary Structure Prediction using Feed-Forward Neural Network M. A. Mottalib,
COPYRIGHT 2010 JCIT, ISSN 2078-5828 (PRINT), ISSN 2218-5224 (ONLINE), VOLUME 01, ISSUE 01, MANUSCRIPT CODE: 100713 Protein Secondary Structure Prediction using Feed-Forward Neural Network M. A. Mottalib,
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT
 5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Advanced Topics in RNA and DNA. DNA Microarrays Aptamers
 Quiz 1 Advanced Topics in RNA and DNA DNA Microarrays Aptamers 2 Quantifying mrna levels to asses protein expression 3 The DNA Microarray Experiment 4 Application of DNA Microarrays 5 Some applications
Quiz 1 Advanced Topics in RNA and DNA DNA Microarrays Aptamers 2 Quantifying mrna levels to asses protein expression 3 The DNA Microarray Experiment 4 Application of DNA Microarrays 5 Some applications
Computer simulations of protein folding with a small number of distance restraints
 Vol. 49 No. 3/2002 683 692 QUARTERLY Computer simulations of protein folding with a small number of distance restraints Andrzej Sikorski 1, Andrzej Kolinski 1,2 and Jeffrey Skolnick 2 1 Department of Chemistry,
Vol. 49 No. 3/2002 683 692 QUARTERLY Computer simulations of protein folding with a small number of distance restraints Andrzej Sikorski 1, Andrzej Kolinski 1,2 and Jeffrey Skolnick 2 1 Department of Chemistry,
BIOINF 4120 Bioinformatics 2 - Structures and Systems - Oliver Kohlbacher Summer Protein Structure Prediction I
 BIOINF 4120 Bioinformatics 2 - Structures and Systems - Oliver Kohlbacher Summer 2013 9. Protein Structure Prediction I Structure Prediction Overview Overview of problem variants Secondary structure prediction
BIOINF 4120 Bioinformatics 2 - Structures and Systems - Oliver Kohlbacher Summer 2013 9. Protein Structure Prediction I Structure Prediction Overview Overview of problem variants Secondary structure prediction
Computational Protein Design
 11 Computational Protein Design This chapter introduces the automated protein design and experimental validation of a novel designed sequence, as described in Dahiyat and Mayo [1]. 11.1 Introduction Given
11 Computational Protein Design This chapter introduces the automated protein design and experimental validation of a novel designed sequence, as described in Dahiyat and Mayo [1]. 11.1 Introduction Given
Supplementary figure 1. Comparison of unbound ogm-csf and ogm-csf as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine
 Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett. Introduction
 The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
Translation. A ribosome, mrna, and trna.
 Translation The basic processes of translation are conserved among prokaryotes and eukaryotes. Prokaryotic Translation A ribosome, mrna, and trna. In the initiation of translation in prokaryotes, the Shine-Dalgarno
Translation The basic processes of translation are conserved among prokaryotes and eukaryotes. Prokaryotic Translation A ribosome, mrna, and trna. In the initiation of translation in prokaryotes, the Shine-Dalgarno
Sequence analysis and comparison
 The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
Ramachandran Plot. 4ysz Phi (degrees) Plot statistics
 B Ramachandran Plot ~b b 135 b ~b ~l l Psi (degrees) 5-5 a A ~a L - -135 SER HIS (F) 59 (G) SER (B) ~b b LYS ASP ASP 315 13 13 (A) (F) (B) LYS ALA ALA 315 173 (E) 173 (E)(A) ~p p ~b - -135 - -5 5 135 (degrees)
B Ramachandran Plot ~b b 135 b ~b ~l l Psi (degrees) 5-5 a A ~a L - -135 SER HIS (F) 59 (G) SER (B) ~b b LYS ASP ASP 315 13 13 (A) (F) (B) LYS ALA ALA 315 173 (E) 173 (E)(A) ~p p ~b - -135 - -5 5 135 (degrees)
Comparative analysis of Wolbachia surface protein in D. melanoagster, A. tabida and B. malayi
 www.bioinformation.net Hypothesis Volume 8(15) Comparative analysis of Wolbachia surface protein in D. melanoagster, A. tabida and B. malayi Jayaramaiah Uday & Hosagavi Puttegowda Puttaraju* Division of
www.bioinformation.net Hypothesis Volume 8(15) Comparative analysis of Wolbachia surface protein in D. melanoagster, A. tabida and B. malayi Jayaramaiah Uday & Hosagavi Puttegowda Puttaraju* Division of
Biotechnology of Proteins. The Source of Stability in Proteins (III) Fall 2015
 Biotechnology of Proteins The Source of Stability in Proteins (III) Fall 2015 Conformational Entropy of Unfolding It is The factor that makes the greatest contribution to stabilization of the unfolded
Biotechnology of Proteins The Source of Stability in Proteins (III) Fall 2015 Conformational Entropy of Unfolding It is The factor that makes the greatest contribution to stabilization of the unfolded
Any protein that can be labelled by both procedures must be a transmembrane protein.
 1. What kind of experimental evidence would indicate that a protein crosses from one side of the membrane to the other? Regions of polypeptide part exposed on the outside of the membrane can be probed
1. What kind of experimental evidence would indicate that a protein crosses from one side of the membrane to the other? Regions of polypeptide part exposed on the outside of the membrane can be probed
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Basic Local Alignment Search Tool
 Basic Local Alignment Search Tool Alignments used to uncover homologies between sequences combined with phylogenetic studies o can determine orthologous and paralogous relationships Local Alignment uses
Basic Local Alignment Search Tool Alignments used to uncover homologies between sequences combined with phylogenetic studies o can determine orthologous and paralogous relationships Local Alignment uses
Aoife McLysaght Dept. of Genetics Trinity College Dublin
 Aoife McLysaght Dept. of Genetics Trinity College Dublin Evolution of genome arrangement Evolution of genome content. Evolution of genome arrangement Gene order changes Inversions, translocations Evolution
Aoife McLysaght Dept. of Genetics Trinity College Dublin Evolution of genome arrangement Evolution of genome content. Evolution of genome arrangement Gene order changes Inversions, translocations Evolution
Supporting information to: Time-resolved observation of protein allosteric communication. Sebastian Buchenberg, Florian Sittel and Gerhard Stock 1
 Supporting information to: Time-resolved observation of protein allosteric communication Sebastian Buchenberg, Florian Sittel and Gerhard Stock Biomolecular Dynamics, Institute of Physics, Albert Ludwigs
Supporting information to: Time-resolved observation of protein allosteric communication Sebastian Buchenberg, Florian Sittel and Gerhard Stock Biomolecular Dynamics, Institute of Physics, Albert Ludwigs
Geometrical Concept-reduction in conformational space.and his Φ-ψ Map. G. N. Ramachandran
 Geometrical Concept-reduction in conformational space.and his Φ-ψ Map G. N. Ramachandran Communication paths in trna-synthetase: Insights from protein structure networks and MD simulations Saraswathi Vishveshwara
Geometrical Concept-reduction in conformational space.and his Φ-ψ Map G. N. Ramachandran Communication paths in trna-synthetase: Insights from protein structure networks and MD simulations Saraswathi Vishveshwara
Effects of Gap Open and Gap Extension Penalties
 Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Unraveling the degradation of artificial amide bonds in Nylon oligomer hydrolase: From induced-fit to acylation processes
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting Information for Unraveling the degradation of artificial amide bonds
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting Information for Unraveling the degradation of artificial amide bonds
Peptides And Proteins
 Kevin Burgess, May 3, 2017 1 Peptides And Proteins from chapter(s) in the recommended text A. Introduction B. omenclature And Conventions by amide bonds. on the left, right. 2 -terminal C-terminal triglycine
Kevin Burgess, May 3, 2017 1 Peptides And Proteins from chapter(s) in the recommended text A. Introduction B. omenclature And Conventions by amide bonds. on the left, right. 2 -terminal C-terminal triglycine
B O C 4 H 2 O O. NOTE: The reaction proceeds with a carbonium ion stabilized on the C 1 of sugar A.
 hbcse 33 rd International Page 101 hemistry lympiad Preparatory 05/02/01 Problems d. In the hydrolysis of the glycosidic bond, the glycosidic bridge oxygen goes with 4 of the sugar B. n cleavage, 18 from
hbcse 33 rd International Page 101 hemistry lympiad Preparatory 05/02/01 Problems d. In the hydrolysis of the glycosidic bond, the glycosidic bridge oxygen goes with 4 of the sugar B. n cleavage, 18 from
Structural insights into energy regulation of light-harvesting complex from spinach CP29
 SUPPLEMENTARY INFORMATION Structural insights into energy regulation of light-harvesting complex from spinach CP29 Xiaowei Pan 1, Mei Li 1, Tao Wan 1,2, Longfei Wang 1,2, Chenjun Jia 1,2, Zhiqiang Hou
SUPPLEMENTARY INFORMATION Structural insights into energy regulation of light-harvesting complex from spinach CP29 Xiaowei Pan 1, Mei Li 1, Tao Wan 1,2, Longfei Wang 1,2, Chenjun Jia 1,2, Zhiqiang Hou
Example of Function Prediction
 Find similar genes Example of Function Prediction Suggesting functions of newly identified genes It was known that mutations of NF1 are associated with inherited disease neurofibromatosis 1; but little
Find similar genes Example of Function Prediction Suggesting functions of newly identified genes It was known that mutations of NF1 are associated with inherited disease neurofibromatosis 1; but little
08/21/2017 BLAST. Multiple Sequence Alignments: Clustal Omega
 BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
Folding of Polypeptide Chains in Proteins: A Proposed Mechanism for Folding
 Proc. Nat. Acad. Sci. USA Vol. 68, No. 9, pp. 2293-2297, September 1971 Folding of Polypeptide Chains in Proteins: A Proposed Mechanism for Folding PETER N. LEWS, FRANK A. MOMANY, AND HAROLD A. SCHERAGA*
Proc. Nat. Acad. Sci. USA Vol. 68, No. 9, pp. 2293-2297, September 1971 Folding of Polypeptide Chains in Proteins: A Proposed Mechanism for Folding PETER N. LEWS, FRANK A. MOMANY, AND HAROLD A. SCHERAGA*
Electronic Supplementary Information
 Electronic Supplementary Information A Sensitive Phosphorescent Thiol Chemosensor Based on an Iridium(III) Complex with α,β-unsaturated Ketone Functionalized 2,2 -Bipyridyl Ligand Na Zhao, a Yu-Hui Wu,
Electronic Supplementary Information A Sensitive Phosphorescent Thiol Chemosensor Based on an Iridium(III) Complex with α,β-unsaturated Ketone Functionalized 2,2 -Bipyridyl Ligand Na Zhao, a Yu-Hui Wu,
Volume 12(2)
 Open access www.bioinformation.net Volume 12(2) Hypothesis Insights from molecular modeling, docking and simulation of imidazole nucleus containing chalcones with EGFR kinase domain for improved binding
Open access www.bioinformation.net Volume 12(2) Hypothesis Insights from molecular modeling, docking and simulation of imidazole nucleus containing chalcones with EGFR kinase domain for improved binding
Hidden symmetries in primary sequences of small α proteins
 Hidden symmetries in primary sequences of small α proteins Ruizhen Xu, Yanzhao Huang, Mingfen Li, Hanlin Chen, and Yi Xiao * Biomolecular Physics and Modeling Group, Department of Physics, Huazhong University
Hidden symmetries in primary sequences of small α proteins Ruizhen Xu, Yanzhao Huang, Mingfen Li, Hanlin Chen, and Yi Xiao * Biomolecular Physics and Modeling Group, Department of Physics, Huazhong University
Statistical Distributions of Optimal Global Alignment Scores of Random Protein Sequences
 BMC Bioinformatics This Provisional PDF corresponds to the article as it appeared upon acceptance. The fully-formatted PDF version will become available shortly after the date of publication, from the
BMC Bioinformatics This Provisional PDF corresponds to the article as it appeared upon acceptance. The fully-formatted PDF version will become available shortly after the date of publication, from the
Introduction to the Ribosome Overview of protein synthesis on the ribosome Prof. Anders Liljas
 Introduction to the Ribosome Molecular Biophysics Lund University 1 A B C D E F G H I J Genome Protein aa1 aa2 aa3 aa4 aa5 aa6 aa7 aa10 aa9 aa8 aa11 aa12 aa13 a a 14 How is a polypeptide synthesized? 2
Introduction to the Ribosome Molecular Biophysics Lund University 1 A B C D E F G H I J Genome Protein aa1 aa2 aa3 aa4 aa5 aa6 aa7 aa10 aa9 aa8 aa11 aa12 aa13 a a 14 How is a polypeptide synthesized? 2
Viewing and Analyzing Proteins, Ligands and their Complexes 2
 2 Viewing and Analyzing Proteins, Ligands and their Complexes 2 Overview Viewing the accessible surface Analyzing the properties of proteins containing thousands of atoms is best accomplished by representing
2 Viewing and Analyzing Proteins, Ligands and their Complexes 2 Overview Viewing the accessible surface Analyzing the properties of proteins containing thousands of atoms is best accomplished by representing
Resonance assignments in proteins. Christina Redfield
 Resonance assignments in proteins Christina Redfield 1. Introduction The assignment of resonances in the complex NMR spectrum of a protein is the first step in any study of protein structure, function
Resonance assignments in proteins Christina Redfield 1. Introduction The assignment of resonances in the complex NMR spectrum of a protein is the first step in any study of protein structure, function
Objective: You will be able to justify the claim that organisms share many conserved core processes and features.
 Objective: You will be able to justify the claim that organisms share many conserved core processes and features. Do Now: Read Enduring Understanding B Essential knowledge: Organisms share many conserved
Objective: You will be able to justify the claim that organisms share many conserved core processes and features. Do Now: Read Enduring Understanding B Essential knowledge: Organisms share many conserved
BIOINFORMATICS LAB AP BIOLOGY
 BIOINFORMATICS LAB AP BIOLOGY Bioinformatics is the science of collecting and analyzing complex biological data. Bioinformatics combines computer science, statistics and biology to allow scientists to
BIOINFORMATICS LAB AP BIOLOGY Bioinformatics is the science of collecting and analyzing complex biological data. Bioinformatics combines computer science, statistics and biology to allow scientists to
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Energy and Cellular Metabolism
 1 Chapter 4 About This Chapter Energy and Cellular Metabolism 2 Energy in biological systems Chemical reactions Enzymes Metabolism Figure 4.1 Energy transfer in the environment Table 4.1 Properties of
1 Chapter 4 About This Chapter Energy and Cellular Metabolism 2 Energy in biological systems Chemical reactions Enzymes Metabolism Figure 4.1 Energy transfer in the environment Table 4.1 Properties of
