AS A SERVICE TO THE RESEARCH COMMUNITY, GENOME BIOLOGY PROVIDES A 'PREPRINT' DEPOSITORY
|
|
|
- Emmeline Montgomery
- 5 years ago
- Views:
Transcription
1 This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article MRD: a microsatellite repeats database for prokaryotic and eukaryotic genomes Subbaya Subramanian, Vamsi M Madgula #, Ranjan George #, Rakesh K Mishra, Madhusudhan W Pandit, Chandrashekar S Kumar # and Lalji Singh Addresses: Centre for Cellular and Molecular Biology, Uppal Road, Hyderabad , India. #Ingenovis, ilabs ltd., 97, Road No.3, Banjara Hills, Hyderabad, , India Correspondence: Lalji Singh. lalji@ccmb.res.in Posted: 13 November 2002 Genome Biology 2002, 3(12):preprint The electronic version of this article is the complete one and can be found online at BioMed Central Ltd (Print ISSN ; Online ISSN ).deposited research Received: 8 November 2002 This is the first version of this article to be made available publicly and no other version is available at present. AS A SERVICE TO THE RESEARCH COMMUNITY, GENOME BIOLOGY PROVIDES A 'PREPRINT' DEPOSITORY TO WHICH ANY ORIGINAL RESEARCH CAN BE SUBMITTED AND WHICH ALL INDIVIDUALS CAN ACCESS FREE OF CHARGE. ANY ARTICLE CAN BE SUBMITTED BY AUTHORS, WHO HAVE SOLE RESPONSIBILITY FOR THE ARTICLE'S CONTENT. THE ONLY SCREENING IS TO ENSURE RELEVANCE OF THE PREPRINT TO GENOME BIOLOGY'S SCOPE AND TO AVOID ABUSIVE, LIBELLOUS OR INDECENT ARTICLES. ARTICLES IN THIS SECTION OF THE JOURNAL HAVE NOT BEEN PEER-REVIEWED. EACH PREPRINT HAS A PERMANENT URL, BY WHICH IT CAN BE CITED. RESEARCH SUBMITTED TO THE PREPRINT DEPOSITORY MAY BE SIMULTANEOUSLY OR SUBSEQUENTLY SUBMITTED TO GENOME BIOLOGY OR ANY OTHER PUBLICATION FOR PEER REVIEW; THE ONLY REQUIREMENT IS AN EXPLICIT CITATION OF, AND LINK TO, THE PREPRINT IN ANY VERSION OF THE ARTICLE THAT IS EVENTUALLY PUBLISHED. IF POSSIBLE, GENOME BIOLOGY WILL PROVIDE A RECIPROCAL LINK FROM THE PREPRINT TO THE PUBLISHED ARTICLE. comment reviews reports deposited research refereed research interactions information
2 2 Genome Biology Deposited research (preprint) MRD: A Microsatellite Repeats Database for prokaryotic and eukaryotic Genomes Subbaya Subramanian, Vamsi M Madgula #, Ranjan George #, Rakesh K Mishra, Madhusudhan W Pandit, Chandrashekar S Kumar # and Lalji Singh Centre for Cellular and Molecular Biology, Uppal Road, Hyderabad , INDIA and # Ingenovis, ilabs ltd., 97, Road No.3, Banjara Hills, Hyderabad, , INDIA Running Title Database for microsatellite repeats Correspondence to Dr Lalji Singh Centre for Cellular and Molecular Biology Uppal Road, Hyderabad INDIA Tel: Fax: lalji@ccmb.res.in 2
3 ABSTRACT MRD is a database system to access the microsatellite repeats information of genomes such as archea, eubacteria, and other eukaryotic genomes whose sequence information is available in public domains. MRD stores information about simple tandemly repeated k-mer sequences where k= 1 to 6, i.e. monomer to hexamer. The web interface allows the users to search for the repeat of their interest and to know about the association of the repeat with genes and genomic regions in the specific organism. The data contains the abundance and distribution of microsatellites in the coding and non-coding regions of the genome. The exact location of repeats with respect to genomic regions of interest (such as UTR, exon, intron or intergenic regions) whichever is applicable to organism is highlighted. MRD is available on the World Wide Web at and/or The database is designed as an open-ended system to accommodate the microsatellite repeats information of other genomes whose complete sequences will be available in future through public domain. 3
4 4 Genome Biology Deposited research (preprint) INTRODUCTION Microsatellites are tandemly repeated sequence motifs of 1 to 6 base pairs [1] found in abundance in the genomes of prokaryotes [2] and eukaryotes [3]. These repeats are found in both coding and non-coding regions of the genome. The presence of microsatellites in the coding region and in the regulatory region of the genome can directly influence the gene expression. Studies have indicated that microsatellites are predominantly present in the non-coding part of the genome and play a significant role in the genome evolution and possibly in gene regulation [4]. While, there is no direct correlation of the microsatellite content with the genome size, it is generally believed that microsatellite content of a genome depends on the genome size [5]. Microsatellites show a high degree of length polymorphisms and are extremely useful in human genetic studies. Many markers have been developed from the known sequences containing these repeats available from databases as well as derived from screening genomic libraries. In spite of the recognition of simple repeats as markers, mechanisms underlying the microsatellite allelic diversity are still poorly understood. Strand slippage during replication has been suggested to be the most likely mechanism in generation of mutation and polymorphism in the microsatellites [6]. Questions such as why certain repeats motifs are common than others and why there exists a variation of such repeats among taxa are important from evolutionary point of view. Though there are extensive studies on the microsatellite repeats in the human and other genomes, a complete inventory of the microsatellite repeats in the human genome and other genomes, as a single resource is still not available. However with the completion of many prokaryotic and eukaryotic genomes this analysis has become possible. Realising the importance of 4
5 microsatellites in the genome, we under took to analyse in detail the repeat distribution and genes associated with them. IMPLEMENTATION We have created a microsatellite repeats database of all repeat combinations from mono- to hexanucleotide repeats. For the analysis we have used the sequence information that is available on the Genbank genomes FTP site. The build number and the release date for the genomes are given in the database. All 501 theoretically possible non-overlapping repeat types were searched [7]. We have analysed the distribution of perfect repeats of >=12 base pairs. The rationale for choosing the small cutoff value was that the microsatellites are often disrupted by single base substitutions. We have also included an extensive analysis of distribution of these repeats and their association with coding and non-coding regions of the genome, such as exon, intron and intergenic regions wherever applicable. MRD provides a comprehensive resource for studying various aspects of microsatellite repeats in prokaryotic and eukaryotic genomes which may help in understanding their probable function and evolutionary significance. The FTP sites of the database from where various genome sequences are obtained are given in the MRD database. The entire data is stored in an Oracle Database. DATA STRUCTURE The database is presented as various views that tabulate the details about a particular repeat type and the repeat. In the current form we have analysed 77 prokaryotic genomes and 7 eukaryotic genomes such as human, mouse, Drosophila, Yeast (S cerevesiae and S pombe), C elegans, and Arabidopsis thaliana. For each organism the microsatellite repeats were analysed and the density and distribution has been tabulated. The repeat associations with specific genomic 5
6 6 Genome Biology Deposited research (preprint) regions are tabulated for each repeat. Both the strands in the sequence were searched for the microsatellite repeats. The microsatellites were searched for a min cut-off of 12bp in both prokaryotes and eukaryotes. The database is organized so as to provide summary as well as detailed views of the repeat regions and their associations with genomic regions, genes. The complete data is organized into six tables. Size of a chromosome refers to the cumulative size of those regions for which the sequence is known and analyzed. All densities mentioned in the tables are with respect to this size. All numbers mentioned are a sum of the occurrences of the particular pattern and its reverse complement. The first page of the database provides a brief description of the database and a link to the page that enables the user to select the genome and repeat class of interest. In its current shape, the database deals only with microsatellites. The database has been designed in an open-ended fashion so that it is possible to add other types of repeats in the future. Once the user has chosen the genome of interest, he/she may view information organized within five views. View 1: Abundance and distribution of monomer to hexamer repeat types This table presents the abundance and density of each of the six repeat types, i.e. monomer to hexamer, across each chromosome of the organism selected. The total for all repeats types across a particular chromosome and for all chromosomes for a particular repeat type are also presented. Density of each repeat type in terms of repeats per mb (mega base pairs) of chromosome is given. 6
7 View 2: Abundance and distribution of all 501 repeats across the genome selected This table gives the same information as above but for all repeats in a particular repeat type. One is thus able to view details of occurrence and density for each of the 501 repeats across all chromosomes or the whole genome in case of prokaryotes. View 3: Abundance and distribution of monomer to hexamer repeat types across genomic regions for a selected genome It is essential and interesting to know the distribution of microsatellites across genomic regions, i.e. exon, intron and intergenic regions. The sizes of exon, intron and intergenic regions (in terms of base pairs) for each chromosome have been calculated from data given in the annotation lists of Genbank entries. For each repeat found, the genomic region it belongs to is captured. Thus for a given chromosome, the density of repeat types on exon, intron and intergenic regions is presented. However, only those repeats that start and end in the same region have been considered for density calculations. Repeats which span regions, say, start in an exon and end in an intron, have not been considered; the occurrence of such repeats is, in any case, rare. In the case of prokaryotic genomes the repeat abundance and density is analysed based on the coding and non-coding regions of the genome. View 4: Detailed view of each repeat, association with proximal genes and STS markers This table gives complete details of each repeat found in a given genome. For a given repeat, the repeat number and the length of the total repeated sequence are given. The start position of the repeat, both with respect to the contig sequence and the original Genbank entry (denoted by its accession id) where the repeat is found, is given. If the repeat is found on a particular gene, the 7
8 8 Genome Biology Deposited research (preprint) name of the gene and the exact regions of the gene where the repeat is found (exon number or intron number) are displayed. (In the case of prokaryotic genomes, currently the database refers to the coding sequences as exons). Wherever applicable if a repeat is found to lie in the UTR region of an mrna, the region is mentioned as UTR1 or UTR2, as the case may be. UTR1 refers to the distance between the first exon on the mrna and its corresponding coding sequence. UTR2 refers to the distance between the stop codon of the mrna and the last base pair of the transcript. However if more than one mrna is present for a given gene, the mrna with the largest exon regions (sum of all exon lengths) and/or starting the earliest is considered. In the event that a repeat lies in an intergenic region, the nearest downstream gene and the distance between the repeat and the gene (in terms of base pairs) is given. A distinction is also made between intergenic regions, which are upstream and downstream of a gene. The terms upstream and downstream are used with reference to the sequence as given in the Genbank entries. In the case of the human genome, association of microsatellites with Sequence Tagged Sites (STS) is also said to be important and revealing. Thus, if a repeat lies on a STS marker, we have therefore included the Standard Name of the STS, as mentioned in the annotation. Otherwise, the nearest STS marker and the distance between the STS marker and the repeat are given. The start and stop positions of the repeat vis-à-vis the STS, i.e. whether they are upstream or downstream of the STS, are also given. This however, is not applicable in the case of other genomes. 8
9 Detailed view of repeats contained within genes A separate option is provided in View 4 which facilitates the user to view details of only those repeats that are contained within genes or which span a gene and the nearing intergenic region. View 5: Details of each repeat for a specific gene This table allows the user to specify the gene of interest and view details of all repeats associated with the gene, i.e. those that are either contained within the gene or are proximal to it. The names of the genes have been taken from the contig files of Genbank. The database therefore currently accepts gene names written in Genbank nomenclature only. AVAILABILITY MRD is available on the World Wide Web at and A user-friendly interactive interface is in place that provides researchers the facility to view details of microsatellites of their interest. The database also includes detailed instructions on how to access and utilize the resource. Technical concerns and queries may be directed to subree@ccmb.res.in or vamsi.madhav@ingenovis.com CONCLUSION The high lights of MRD database are as follows 1. The database includes more than eighty different genome information for the microsatellite repeats and abundance, density and distribution. 9
10 10 Genome Biology Deposited research (preprint) 2. Comprehensive tables which details about the repeat association with specific genomic region. In case of human and mouse the STS marker association is also included which will help in researches to identify more STRs. 3. Using MRD it is possible to compare different repeats and investigate the evolution of associated genomic regions. For example, looking at the triplet repeat containing genes in different organisms can reveal its potential for repeat expansion associated disease. 4. The database is expected provide an excellent platform for the analysis of microsatellite evolution and their possible role in genome organization. 5. One of the possible roles of such repeat is suggested to be in gene regulation. Study of the association of these repeats in the flanking sequences of the gene may be the first step to understand role of microsatellites in gene regulation. We have provided an option of searching repeats in the flanking sequence of specific gene. 6. Using this database it is possible to list out most abundant or rare repeats in different organisms. We hope that this database will be a comprehensive resource for studying various aspects of microsatellite repeats in the human and other genomes and that it will be helpful in identifying their probable function and evolutionary significance. Information on the abundance of microsatellites, coupled with the distribution patterns in the coding as well as non-coding regions of the genome and their associations with genes and STS markers will shed more light on the function of microsatellites in gene regulation. As and when genome sequences of new species become available through the public domain the list of genomes in the MRD database will be expanded. 10
11 Acknowledgements The authors are thankful to Sreedhar, Siva Prasad, Saritha and Kavitha for their support in developing the MRD database. We are grateful to Thangaraj and Ramesh Aggarwal for providing helpful discussions. Financial support from CSIR and DBT is duly acknowledged. 11
12 12 Genome Biology Deposited research (preprint) REFERENCES 1. Vogt P: Potential genetic functions of tandem repeated DNA sequence blocks in the human genome are based on a highly conserved "chromatin folding code. Hum. Genet 1990, 84: Gur-Arie R, Cohen C, Eitan Y, Shelef L, Hallerman EM, Kashi Y: Simple sequence repeats in Escherichia coli: abundance, distribution, composition, and polymorphism. Genome Res 2000, 10: Toth G, Gaspari Z, Jurka J: Microsatellites in different eukaryotic genomes: survey and analysis. Genome Res 2000, 10: Kashi Y, King D, Soller M: () Simple sequence repeats as a source of quantitative genetic variation. Trends Genet 1997, 13: Primmer CR, Raudsepp T, Chowdhary BP, Moller AP, Ellegren H: Low frequency of microsatellites in the avian genome. Genome Res 1997, 7: Pearson CE, Sinden RR: Trinucleotide repeat DNA structures: dynamic mutations from dynanic DNA Curr Opin Struct Biol 1998, 8: Jurka, J., and Pethiyagoda, C. Simple repetitive DNA sequences from primates: compilation and analysis. J Mol Evol 1995, 40:
13 Figure 1 11
Biology. Biology. Slide 1 of 26. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
How much non-coding DNA do eukaryotes require?
 How much non-coding DNA do eukaryotes require? Andrei Zinovyev UMR U900 Computational Systems Biology of Cancer Institute Curie/INSERM/Ecole de Mine Paritech Dr. Sebastian Ahnert Dr. Thomas Fink Bioinformatics
How much non-coding DNA do eukaryotes require? Andrei Zinovyev UMR U900 Computational Systems Biology of Cancer Institute Curie/INSERM/Ecole de Mine Paritech Dr. Sebastian Ahnert Dr. Thomas Fink Bioinformatics
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
The Gene The gene; Genes Genes Allele;
 Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
TE content correlates positively with genome size
 TE content correlates positively with genome size Mb 3000 Genomic DNA 2500 2000 1500 1000 TE DNA Protein-coding DNA 500 0 Feschotte & Pritham 2006 Transposable elements. Variation in gene numbers cannot
TE content correlates positively with genome size Mb 3000 Genomic DNA 2500 2000 1500 1000 TE DNA Protein-coding DNA 500 0 Feschotte & Pritham 2006 Transposable elements. Variation in gene numbers cannot
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
AS A SERVICE TO THE RESEARCH COMMUNITY, GENOME BIOLOGY PROVIDES A 'PREPRINT' DEPOSITORY
 This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article Universality in large-scale structure of complete genomes Li-Ching
This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article Universality in large-scale structure of complete genomes Li-Ching
Genomes and Their Evolution
 Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
(Lys), resulting in translation of a polypeptide without the Lys amino acid. resulting in translation of a polypeptide without the Lys amino acid.
 1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
L3.1: Circuits: Introduction to Transcription Networks. Cellular Design Principles Prof. Jenna Rickus
 L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
Research Article Expression Divergence of Tandemly Arrayed Genes in Human and Mouse
 Hindawi Publishing Corporation Comparative and Functional Genomics Volume 27, Article ID 6964, 8 pages doi:1.1155/27/6964 Research Article Expression Divergence of Tandemly Arrayed Genes in Human and Mouse
Hindawi Publishing Corporation Comparative and Functional Genomics Volume 27, Article ID 6964, 8 pages doi:1.1155/27/6964 Research Article Expression Divergence of Tandemly Arrayed Genes in Human and Mouse
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Related Courses He who asks is a fool for five minutes, but he who does not ask remains a fool forever.
 CSE 527 Computational Biology http://www.cs.washington.edu/527 Lecture 1: Overview & Bio Review Autumn 2004 Larry Ruzzo Related Courses He who asks is a fool for five minutes, but he who does not ask remains
CSE 527 Computational Biology http://www.cs.washington.edu/527 Lecture 1: Overview & Bio Review Autumn 2004 Larry Ruzzo Related Courses He who asks is a fool for five minutes, but he who does not ask remains
A DNA Sequence 2017/12/6 1
 A DNA Sequence ccgtacgtacgtagagtgctagtctagtcgtagcgccgtagtcgatcgtgtgg gtagtagctgatatgatgcgaggtaggggataggatagcaacagatgagc ggatgctgagtgcagtggcatgcgatgtcgatgatagcggtaggtagacttc gcgcataaagctgcgcgagatgattgcaaagragttagatgagctgatgcta
A DNA Sequence ccgtacgtacgtagagtgctagtctagtcgtagcgccgtagtcgatcgtgtgg gtagtagctgatatgatgcgaggtaggggataggatagcaacagatgagc ggatgctgagtgcagtggcatgcgatgtcgatgatagcggtaggtagacttc gcgcataaagctgcgcgagatgattgcaaagragttagatgagctgatgcta
12-5 Gene Regulation
 12-5 Gene Regulation Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 1 of 26 12-5 Gene Regulation Gene Regulation: An Example Gene
12-5 Gene Regulation Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 1 of 26 12-5 Gene Regulation Gene Regulation: An Example Gene
Flow of Genetic Information
 presents Flow of Genetic Information A Montagud E Navarro P Fernández de Córdoba JF Urchueguía Elements Nucleic acid DNA RNA building block structure & organization genome building block types Amino acid
presents Flow of Genetic Information A Montagud E Navarro P Fernández de Córdoba JF Urchueguía Elements Nucleic acid DNA RNA building block structure & organization genome building block types Amino acid
Biology 105/Summer Bacterial Genetics 8/12/ Bacterial Genomes p Gene Transfer Mechanisms in Bacteria p.
 READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Algorithms in Computational Biology (236522) spring 2008 Lecture #1
 Algorithms in Computational Biology (236522) spring 2008 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: 15:30-16:30/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office hours:??
Algorithms in Computational Biology (236522) spring 2008 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: 15:30-16:30/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office hours:??
GENOME-WIDE ANALYSIS OF CORE PROMOTER REGIONS IN EMILIANIA HUXLEYI
 1 GENOME-WIDE ANALYSIS OF CORE PROMOTER REGIONS IN EMILIANIA HUXLEYI Justin Dailey and Xiaoyu Zhang Department of Computer Science, California State University San Marcos San Marcos, CA 92096 Email: daile005@csusm.edu,
1 GENOME-WIDE ANALYSIS OF CORE PROMOTER REGIONS IN EMILIANIA HUXLEYI Justin Dailey and Xiaoyu Zhang Department of Computer Science, California State University San Marcos San Marcos, CA 92096 Email: daile005@csusm.edu,
Comparison of simple sequence repeats in Staphylococcus strains using in-silico approach
 www.bioinformation.net Hypothesis Volume 8(24) Comparison of simple sequence repeats in Staphylococcus strains using in-silico approach Sunil Thorat 1 * & Prashant Thakare 2 1Institute of Bioresources
www.bioinformation.net Hypothesis Volume 8(24) Comparison of simple sequence repeats in Staphylococcus strains using in-silico approach Sunil Thorat 1 * & Prashant Thakare 2 1Institute of Bioresources
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST
 Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Bio 119 Bacterial Genomics 6/26/10
 BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Simulation of Gene Regulatory Networks
 Simulation of Gene Regulatory Networks Overview I have been assisting Professor Jacques Cohen at Brandeis University to explore and compare the the many available representations and interpretations of
Simulation of Gene Regulatory Networks Overview I have been assisting Professor Jacques Cohen at Brandeis University to explore and compare the the many available representations and interpretations of
Supplemental Materials
 JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
Tandem repeat 16,225 20,284. 0kb 5kb 10kb 15kb 20kb 25kb 30kb 35kb
 Overview Fosmid XAAA112 consists of 34,783 nucleotides. Blat results indicate that this fosmid has significant identity to the 2R chromosome of D.melanogaster. Evidence suggests that fosmid XAAA112 contains
Overview Fosmid XAAA112 consists of 34,783 nucleotides. Blat results indicate that this fosmid has significant identity to the 2R chromosome of D.melanogaster. Evidence suggests that fosmid XAAA112 contains
Small RNA in rice genome
 Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Lecture Notes: BIOL2007 Molecular Evolution
 Lecture Notes: BIOL2007 Molecular Evolution Kanchon Dasmahapatra (k.dasmahapatra@ucl.ac.uk) Introduction By now we all are familiar and understand, or think we understand, how evolution works on traits
Lecture Notes: BIOL2007 Molecular Evolution Kanchon Dasmahapatra (k.dasmahapatra@ucl.ac.uk) Introduction By now we all are familiar and understand, or think we understand, how evolution works on traits
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Eukaryotic vs. Prokaryotic genes
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
The nature of genomes. Viral genomes. Prokaryotic genome. Nonliving particle. DNA or RNA. Compact genomes with little spacer DNA
 The nature of genomes Genomics: study of structure and function of genomes Genome size variable, by orders of magnitude number of genes roughly proportional to genome size Plasmids symbiotic DNA molecules,
The nature of genomes Genomics: study of structure and function of genomes Genome size variable, by orders of magnitude number of genes roughly proportional to genome size Plasmids symbiotic DNA molecules,
Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites
 Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites Paper by: James P. Balhoff and Gregory A. Wray Presentation by: Stephanie Lucas Reviewed
Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites Paper by: James P. Balhoff and Gregory A. Wray Presentation by: Stephanie Lucas Reviewed
Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are:
 Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
MOLECULAR BIOLOGY BIOL 021 SEMESTER 2 (2015) COURSE OUTLINE
 COURSE OUTLINE 1 COURSE GENERAL INFORMATION 1 Course Title & Course Code Molecular Biology: 2 Credit (Contact hour) 3 (2+1+0) 3 Title(s) of program(s) within which the subject is taught. Preparatory Program
COURSE OUTLINE 1 COURSE GENERAL INFORMATION 1 Course Title & Course Code Molecular Biology: 2 Credit (Contact hour) 3 (2+1+0) 3 Title(s) of program(s) within which the subject is taught. Preparatory Program
Предсказание и анализ промотерных последовательностей. Татьяна Татаринова
 Предсказание и анализ промотерных последовательностей Татьяна Татаринова Eukaryotic Transcription 2 Initiation Promoter: the DNA sequence that initially binds the RNA polymerase The structure of promoter-polymerase
Предсказание и анализ промотерных последовательностей Татьяна Татаринова Eukaryotic Transcription 2 Initiation Promoter: the DNA sequence that initially binds the RNA polymerase The structure of promoter-polymerase
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Frequently Asked Questions (FAQs)
 Frequently Asked Questions (FAQs) Q1. What is meant by Satellite and Repetitive DNA? Ans: Satellite and repetitive DNA generally refers to DNA whose base sequence is repeated many times throughout the
Frequently Asked Questions (FAQs) Q1. What is meant by Satellite and Repetitive DNA? Ans: Satellite and repetitive DNA generally refers to DNA whose base sequence is repeated many times throughout the
2012 Univ Aguilera Lecture. Introduction to Molecular and Cell Biology
 2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
Computational Biology and Chemistry
 Computational Biology and Chemistry 33 (2009) 245 252 Contents lists available at ScienceDirect Computational Biology and Chemistry journal homepage: www.elsevier.com/locate/compbiolchem Research Article
Computational Biology and Chemistry 33 (2009) 245 252 Contents lists available at ScienceDirect Computational Biology and Chemistry journal homepage: www.elsevier.com/locate/compbiolchem Research Article
Grundlagen der Bioinformatik Summer semester Lecturer: Prof. Daniel Huson
 Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Universal power law behaviors in genomic sequences and evolutionary models
 Universal power law behaviors in genomic sequences and evolutionary models Loredana Martignetti* and Michele Caselle Dipartimento di Fisica Teoric, Università di Torino and INFN, Via Pietro Giuria 1, I-10125
Universal power law behaviors in genomic sequences and evolutionary models Loredana Martignetti* and Michele Caselle Dipartimento di Fisica Teoric, Università di Torino and INFN, Via Pietro Giuria 1, I-10125
GENE REGULATION AND PROBLEMS OF DEVELOPMENT
 GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
Molecular evolution - Part 1. Pawan Dhar BII
 Molecular evolution - Part 1 Pawan Dhar BII Theodosius Dobzhansky Nothing in biology makes sense except in the light of evolution Age of life on earth: 3.85 billion years Formation of planet: 4.5 billion
Molecular evolution - Part 1 Pawan Dhar BII Theodosius Dobzhansky Nothing in biology makes sense except in the light of evolution Age of life on earth: 3.85 billion years Formation of planet: 4.5 billion
AP Biology. Read college-level text for understanding and be able to summarize main concepts
 St. Mary's College AP Biology Continuity and Change Consider how specific changes to an ecosystem (geological, climatic, introduction of new organisms, etc.) can affect the organisms that live within it.
St. Mary's College AP Biology Continuity and Change Consider how specific changes to an ecosystem (geological, climatic, introduction of new organisms, etc.) can affect the organisms that live within it.
Annotation of Plant Genomes using RNA-seq. Matteo Pellegrini (UCLA) In collaboration with Sabeeha Merchant (UCLA)
 Annotation of Plant Genomes using RNA-seq Matteo Pellegrini (UCLA) In collaboration with Sabeeha Merchant (UCLA) inuscu1-35bp 5 _ 0 _ 5 _ What is Annotation inuscu2-75bp luscu1-75bp 0 _ 5 _ Reconstruction
Annotation of Plant Genomes using RNA-seq Matteo Pellegrini (UCLA) In collaboration with Sabeeha Merchant (UCLA) inuscu1-35bp 5 _ 0 _ 5 _ What is Annotation inuscu2-75bp luscu1-75bp 0 _ 5 _ Reconstruction
Comparing whole genomes
 BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
GSBHSRSBRSRRk IZTI/^Q. LlML. I Iv^O IV I I I FROM GENES TO GENOMES ^^^H*" ^^^^J*^ ill! BQPIP. illt. goidbkc. itip31. li4»twlil FIFTH EDITION
 FIFTH EDITION IV I ^HHk ^ttm IZTI/^Q i I II MPHBBMWBBIHB '-llwmpbi^hbwm^^pfc ' GSBHSRSBRSRRk LlML I I \l 1MB ^HP'^^MMMP" jflp^^^^^^^^st I Iv^O FROM GENES TO GENOMES %^MiM^PM^^MWi99Mi$9i0^^ ^^^^^^^^^^^^^V^^^fii^^t^i^^^^^
FIFTH EDITION IV I ^HHk ^ttm IZTI/^Q i I II MPHBBMWBBIHB '-llwmpbi^hbwm^^pfc ' GSBHSRSBRSRRk LlML I I \l 1MB ^HP'^^MMMP" jflp^^^^^^^^st I Iv^O FROM GENES TO GENOMES %^MiM^PM^^MWi99Mi$9i0^^ ^^^^^^^^^^^^^V^^^fii^^t^i^^^^^
REVIEW SESSION. Wednesday, September 15 5:30 PM SHANTZ 242 E
 REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA
 INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA XIUFENG WAN xw6@cs.msstate.edu Department of Computer Science Box 9637 JOHN A. BOYLE jab@ra.msstate.edu Department of Biochemistry and Molecular Biology
INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA XIUFENG WAN xw6@cs.msstate.edu Department of Computer Science Box 9637 JOHN A. BOYLE jab@ra.msstate.edu Department of Biochemistry and Molecular Biology
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/8/16
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00.
 Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
Molecular Evolution & the Origin of Variation
 Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation
 Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Designer Genes C Test
 Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
UNIT 5. Protein Synthesis 11/22/16
 UNIT 5 Protein Synthesis IV. Transcription (8.4) A. RNA carries DNA s instruction 1. Francis Crick defined the central dogma of molecular biology a. Replication copies DNA b. Transcription converts DNA
UNIT 5 Protein Synthesis IV. Transcription (8.4) A. RNA carries DNA s instruction 1. Francis Crick defined the central dogma of molecular biology a. Replication copies DNA b. Transcription converts DNA
Identification and annotation of promoter regions in microbial genome sequences on the basis of DNA stability
 Annotation of promoter regions in microbial genomes 851 Identification and annotation of promoter regions in microbial genome sequences on the basis of DNA stability VETRISELVI RANGANNAN and MANJU BANSAL*
Annotation of promoter regions in microbial genomes 851 Identification and annotation of promoter regions in microbial genome sequences on the basis of DNA stability VETRISELVI RANGANNAN and MANJU BANSAL*
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
GEP Annotation Report
 GEP Annotation Report Note: For each gene described in this annotation report, you should also prepare the corresponding GFF, transcript and peptide sequence files as part of your submission. Student name:
GEP Annotation Report Note: For each gene described in this annotation report, you should also prepare the corresponding GFF, transcript and peptide sequence files as part of your submission. Student name:
Reading Assignments. A. Genes and the Synthesis of Polypeptides. Lecture Series 7 From DNA to Protein: Genotype to Phenotype
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Hiromi Nishida. 1. Introduction. 2. Materials and Methods
 Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Introduction to Hidden Markov Models for Gene Prediction ECE-S690
 Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Graduate Funding Information Center
 Graduate Funding Information Center UNC-Chapel Hill, The Graduate School Graduate Student Proposal Sponsor: Program Title: NESCent Graduate Fellowship Department: Biology Funding Type: Fellowship Year:
Graduate Funding Information Center UNC-Chapel Hill, The Graduate School Graduate Student Proposal Sponsor: Program Title: NESCent Graduate Fellowship Department: Biology Funding Type: Fellowship Year:
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p
 Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
BioControl - Week 6, Lecture 1
 BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
Chapter 18 Active Reading Guide Genomes and Their Evolution
 Name: AP Biology Mr. Croft Chapter 18 Active Reading Guide Genomes and Their Evolution Most AP Biology teachers think this chapter involves an advanced topic. The questions posed here will help you understand
Name: AP Biology Mr. Croft Chapter 18 Active Reading Guide Genomes and Their Evolution Most AP Biology teachers think this chapter involves an advanced topic. The questions posed here will help you understand
MCB 110. "Molecular Biology: Macromolecular Synthesis and Cellular Function" Spring, 2018
 MCB 110 "Molecular Biology: Macromolecular Synthesis and Cellular Function" Spring, 2018 Faculty Instructors: Prof. Jeremy Thorner Prof. Qiang Zhou Prof. Eva Nogales GSIs:!!!! Ms. Samantha Fernandez Mr.
MCB 110 "Molecular Biology: Macromolecular Synthesis and Cellular Function" Spring, 2018 Faculty Instructors: Prof. Jeremy Thorner Prof. Qiang Zhou Prof. Eva Nogales GSIs:!!!! Ms. Samantha Fernandez Mr.
Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors
 Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors Arli Parikesit, Peter F. Stadler, Sonja J. Prohaska Bioinformatics Group Institute of Computer Science University
Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors Arli Parikesit, Peter F. Stadler, Sonja J. Prohaska Bioinformatics Group Institute of Computer Science University
Translation - Prokaryotes
 1 Translation - Prokaryotes Shine-Dalgarno (SD) Sequence rrna 3 -GAUACCAUCCUCCUUA-5 mrna...ggagg..(5-7bp)...aug Influences: Secondary structure!! SD and AUG in unstructured region Start AUG 91% GUG 8 UUG
1 Translation - Prokaryotes Shine-Dalgarno (SD) Sequence rrna 3 -GAUACCAUCCUCCUUA-5 mrna...ggagg..(5-7bp)...aug Influences: Secondary structure!! SD and AUG in unstructured region Start AUG 91% GUG 8 UUG
Model plants and their Role in genetic manipulation. Mitesh Shrestha
 Model plants and their Role in genetic manipulation Mitesh Shrestha Definition of Model Organism Specific species or organism Extensively studied in research laboratories Advance our understanding of Cellular
Model plants and their Role in genetic manipulation Mitesh Shrestha Definition of Model Organism Specific species or organism Extensively studied in research laboratories Advance our understanding of Cellular
Molecular Biology Of The Cell 6th Edition Alberts
 We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on your computer, you have convenient answers with molecular biology of
We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on your computer, you have convenient answers with molecular biology of
Hairpin Database: Why and How?
 Hairpin Database: Why and How? Clark Jeffries Research Professor Renaissance Computing Institute and School of Pharmacy University of North Carolina at Chapel Hill, United States Why should a database
Hairpin Database: Why and How? Clark Jeffries Research Professor Renaissance Computing Institute and School of Pharmacy University of North Carolina at Chapel Hill, United States Why should a database
G C T G U A. template strand
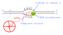 A 5 w 3 3 c 5 mrna RNA polymerase T G C A G C C T G U A coding or sense st AUG?? template strand The triplet code 3 bases = 1 amino acid Punctuation: sta rt: sto p: AUGA CUU CAGUAAC AU U AA C C AUG = methionine,
A 5 w 3 3 c 5 mrna RNA polymerase T G C A G C C T G U A coding or sense st AUG?? template strand The triplet code 3 bases = 1 amino acid Punctuation: sta rt: sto p: AUGA CUU CAGUAAC AU U AA C C AUG = methionine,
Biology I Fall Semester Exam Review 2014
 Biology I Fall Semester Exam Review 2014 Biomolecules and Enzymes (Chapter 2) 8 questions Macromolecules, Biomolecules, Organic Compunds Elements *From the Periodic Table of Elements Subunits Monomers,
Biology I Fall Semester Exam Review 2014 Biomolecules and Enzymes (Chapter 2) 8 questions Macromolecules, Biomolecules, Organic Compunds Elements *From the Periodic Table of Elements Subunits Monomers,
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Expanded View Figures
 Molecular Systems iology Evolutionary divergence of mouse P Mei-Sheng Xiao et al Expanded View Figures Nucleotide percentage (%) 0 20 40 60 80 100 T G Nucleotide percentage (%) 0 20 40 60 80 100 T G Frequency
Molecular Systems iology Evolutionary divergence of mouse P Mei-Sheng Xiao et al Expanded View Figures Nucleotide percentage (%) 0 20 40 60 80 100 T G Nucleotide percentage (%) 0 20 40 60 80 100 T G Frequency
Number of questions TEK (Learning Target) Biomolecules & Enzymes
 Unit Biomolecules & Enzymes Number of questions TEK (Learning Target) on Exam 8 questions 9A I can compare and contrast the structure and function of biomolecules. 9C I know the role of enzymes and how
Unit Biomolecules & Enzymes Number of questions TEK (Learning Target) on Exam 8 questions 9A I can compare and contrast the structure and function of biomolecules. 9C I know the role of enzymes and how
BIOLOGY STANDARDS BASED RUBRIC
 BIOLOGY STANDARDS BASED RUBRIC STUDENTS WILL UNDERSTAND THAT THE FUNDAMENTAL PROCESSES OF ALL LIVING THINGS DEPEND ON A VARIETY OF SPECIALIZED CELL STRUCTURES AND CHEMICAL PROCESSES. First Semester Benchmarks:
BIOLOGY STANDARDS BASED RUBRIC STUDENTS WILL UNDERSTAND THAT THE FUNDAMENTAL PROCESSES OF ALL LIVING THINGS DEPEND ON A VARIETY OF SPECIALIZED CELL STRUCTURES AND CHEMICAL PROCESSES. First Semester Benchmarks:
A Browser for Pig Genome Data
 A Browser for Pig Genome Data Thomas Mailund January 2, 2004 This report briefly describe the blast and alignment data available at http://www.daimi.au.dk/ mailund/pig-genome/ hits.html. The report describes
A Browser for Pig Genome Data Thomas Mailund January 2, 2004 This report briefly describe the blast and alignment data available at http://www.daimi.au.dk/ mailund/pig-genome/ hits.html. The report describes
From Gene to Protein
 From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
Full file at CHAPTER 2 Genetics
 CHAPTER 2 Genetics MULTIPLE CHOICE 1. Chromosomes are a. small linear bodies. b. contained in cells. c. replicated during cell division. 2. A cross between true-breeding plants bearing yellow seeds produces
CHAPTER 2 Genetics MULTIPLE CHOICE 1. Chromosomes are a. small linear bodies. b. contained in cells. c. replicated during cell division. 2. A cross between true-breeding plants bearing yellow seeds produces
Lecture 5: Processes and Timescales: Rates for the fundamental processes 5.1
 Lecture 5: Processes and Timescales: Rates for the fundamental processes 5.1 Reading Assignment for Lectures 5-6: Phillips, Kondev, Theriot (PKT), Chapter 3 Life is not static. Organisms as a whole are
Lecture 5: Processes and Timescales: Rates for the fundamental processes 5.1 Reading Assignment for Lectures 5-6: Phillips, Kondev, Theriot (PKT), Chapter 3 Life is not static. Organisms as a whole are
Lesson Overview. Gene Regulation and Expression. Lesson Overview Gene Regulation and Expression
 13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/1/18
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Differential Distribution of Simple Sequence Repeats in Eukaryotic Genome Sequences
 Differential Distribution of Simple Sequence in Eukaryotic Genome Sequences Mukund V. Katti, Prabhakar K. Ranjekar, and Vidya S. Gupta Plant Molecular Biology Unit, Division of Biochemical Sciences, National
Differential Distribution of Simple Sequence in Eukaryotic Genome Sequences Mukund V. Katti, Prabhakar K. Ranjekar, and Vidya S. Gupta Plant Molecular Biology Unit, Division of Biochemical Sciences, National
TRANSCRIPTION VS TRANSLATION FILE
 23 April, 2018 TRANSCRIPTION VS TRANSLATION FILE Document Filetype: PDF 352.85 KB 0 TRANSCRIPTION VS TRANSLATION FILE Get an answer for 'Compare and contrast transcription and translation in Prokaryotes
23 April, 2018 TRANSCRIPTION VS TRANSLATION FILE Document Filetype: PDF 352.85 KB 0 TRANSCRIPTION VS TRANSLATION FILE Get an answer for 'Compare and contrast transcription and translation in Prokaryotes
Whole-genome analysis of GCN4 binding in S.cerevisiae
 Whole-genome analysis of GCN4 binding in S.cerevisiae Lillian Dai Alex Mallet Gcn4/DNA diagram (CREB symmetric site and AP-1 asymmetric site: Song Tan, 1999) removed for copyright reasons. What is GCN4?
Whole-genome analysis of GCN4 binding in S.cerevisiae Lillian Dai Alex Mallet Gcn4/DNA diagram (CREB symmetric site and AP-1 asymmetric site: Song Tan, 1999) removed for copyright reasons. What is GCN4?
6.096 Algorithms for Computational Biology. Prof. Manolis Kellis
 6.096 Algorithms for Computational Biology Prof. Manolis Kellis Today s Goals Introduction Class introduction Challenges in Computational Biology Gene Regulation: Regulatory Motif Discovery Exhaustive
6.096 Algorithms for Computational Biology Prof. Manolis Kellis Today s Goals Introduction Class introduction Challenges in Computational Biology Gene Regulation: Regulatory Motif Discovery Exhaustive
ECOL/MCB 320 and 320H Genetics
 ECOL/MCB 320 and 320H Genetics Instructors Dr. C. William Birky, Jr. Dept. of Ecology and Evolutionary Biology Lecturing on Molecular genetics Transmission genetics Population and evolutionary genetics
ECOL/MCB 320 and 320H Genetics Instructors Dr. C. William Birky, Jr. Dept. of Ecology and Evolutionary Biology Lecturing on Molecular genetics Transmission genetics Population and evolutionary genetics
Microsatellites, Transposable Elements and the X Chromosome
 Microsatellites, Transposable Elements and the X Chromosome Philippe Jarne,* Patrice David,* and Frédérique Viard* *Génétique & Environnement CC065, Institut des Sciences de l Evolution, Université Montpellier
Microsatellites, Transposable Elements and the X Chromosome Philippe Jarne,* Patrice David,* and Frédérique Viard* *Génétique & Environnement CC065, Institut des Sciences de l Evolution, Université Montpellier
