Supporting Information
|
|
|
- Ethan Watts
- 5 years ago
- Views:
Transcription
1 Supporting Information Development of a human vasopressin V 1a -receptor antagonist from an evolutionary-related insect neuropeptide Maria Giulia Di Giglio, Markus Muttenthaler, Kasper Harpsøe, Zita Liutkeviciute, Peter Keov, Thomas Eder, Thomas Rattei, Sarah Arrowsmith, Susan Wray, Ales Marek, Tomas Elbert, Paul F. Alewood, David E. Gloriam and Christian W. Gruber* *Correspondence and requests for materials should be addressed to C.W.G. ( christian.w.gruber@meduniwien.ac.at or c.gruber@uq.edu.au) -S1-
2 Supplementary Figures Supplementary Fig. 1. Phylogenetic tree of oxytocin/vasopressin (OT/AVP) receptors. Published invertebrate and vertebrate receptors were used together with the top three blast hits from the L. niger transcriptome (Lasius niger Assembly_contigs_1-3, shown in black) to reconstruct the phylogenetic relationship. Only the best blast hit of L. niger (Lasius niger Assembly_contig_1) clusters together with OT/AVP receptors in the same branch, while the other two hits cluster with invertebrate adipokinetic hormone receptors (AKHR). Vertebrate OT/AVP receptors are shown in violet and invertebrate OT/AVPR receptors are shown in pink. Putative OT/AVP-like ant receptors are shown in pink. Vertebrate neuropeptide S receptors (NPSR)/invertebrate crustacean cardioactive peptide receptors (CCAPR) are shown in green and vertebrate gonadotropin-releasing hormone receptors (GnRHR)/invertebrate AKHR are shown in blue. According to the study of Pitti and Manoj 1 the mouse gonadotropin-releasing hormone receptor (GnRHR) was used as outgroup. Numbers at nodes indicate confidence values and ExPASy/Genbank entry IDs of sequences are listed next to the receptor names. -S2-
3 Supplementary Fig. 2. Multiple sequence alignments between inotocin receptors (L. niger, T. castaneum) and human V 2 R, V 1a R, V 1b R and OTR. FASTA sequences were aligned through Clustal Omega and shown in Boxshade format. Colour coding is defined as follows: residues that are similar but non-identical are highlighted in grey; identical residues are highlighted in black. -S3-
4 Supplementary Fig. 3. Receptor pharmacology of [D-Arg8]-inotocin at inotocin and human oxytocin/vasopressin receptors. (A) Concentration-dependent displacement binding curves of inotocin at inotocin receptors (INTR) from Lasius niger ( ) (n = 2), Tribolium castaneum ( ) (n = 3). (B) Concentration-response curves of [D-Arg8]-inotocin at INTR from L. niger (n = 4) and T. castaneum (n = 3) through quantitation of increased intracellular IP 1. (C) Concentration-dependent displacement binding curves of inotocin at human OTR ( ), V 1a R ( ), V 1b R ( ) and V 2 R ( ) (n = 3). (D) Concentration-response curves of [D-Arg8]-inotocin at human OTR, V 1a R, V 1b R and V 2 R (n 3). Specific binding was calculated by subtraction of non-specific binding from total binding and normalized to the percentage (%) of maximal binding. Detailed descriptions of radioligand concentrations, membranes expressing receptors and dissociation constants are provided in the Methods section. Receptor activation was measured by IP 1 assays for the G q -coupled receptors (human OTR, V 1a R, V 1b R) and luciferase reporter assay with specific CRE response element for the G s -coupled human V 2 R, as described in Methods. Each data point was normalized to percentage of maximal activation, detected at the highest endogenous ligand concentration, being inotocin for INTR, vasopressin for V 1a R, V 1b R, V 2 R and oxytocin for OTR. Data is shown as mean ± SEM and fitted by nonlinear regression (sigmoidal, three-parameters, Hill slope of 1). -S4-
5 Supplementary Fig. 4. Structural representation of the human V 1a R homology model binding site. (A) 2-dimensional cartoon representation of receptor structure showing relative positions of residues identified from sequence and structural alignments to comprise the predicted binding pocket (circles). Key residues predicted to discriminate the binding and function of inotocin and [D-Arg8]-inotocin are highlighted (coloured circles). Potential binding residues of Arg8/D-Arg8 are shown in yellow; proposed Lys in V 2 R that impairs inotocin binding is shown in blue; proposed residues of the activation triad are shown in red; residues in human V 1b R, V 2 R, and OTR that potentially impair D-Arg8 binding are presented in green. (B) Van der Waals surface (grey transparent surface) of the binding site residues highlighted in panel A) is presented in the V 1a R homology model (cyan cartoon). One continuous cavity is observed within the upper part of the transmembrane domain comprised of 43 residue positions in TM1-7 and ECL1-2. -S5-
6 Supplementary Fig. 5. Human serum stability of the inotocin D-analogue. [D-Arg8]-inotocin ([D-Arg8]INT), inotocin (INT), vasopressin (AVP) and melittin (100 μm) were incubated in human serum and their stability monitored via HPLC over a time course of 2, 4, 6, 8, 10 and 24 h. Melittin, a haemolytic peptide from the bee venom was used as a positive control for peptide degradation. Area under the curves of samples was determined and correlated to the negative control analyte, which was dissolved in 0.1% TFA; thus they were not subject to degradation and were assumed therefore as 100%. Peptide stability was expressed in percentage of peptide remaining. -S6-
7 Supplementary Fig. 6. Concentration-dependent inhibitory effects of [D-Arg8]-inotocin and SR49059 on vasopressin-augmented (0.5 nm) ex vivo uterine contractions. Under [D-Arg8]- inotocin, contraction amplitude was significantly reduced at 10, 30 and 100 nm (A) whilst areaunder-the-curve (AUC) was significantly reduced at 100 nm (B). Following treatment with SR49059, amplitude of contraction and AUC were significantly reduced at 30 nm and 100 nm (C and D, respectively); *P<0.05, **P<0.01, ***P<0.001 (n = 5, one-way ANOVA, Tukey s post hoc analysis). -S7-
8 Supplementary Fig. 7. Desmopressin is inactive at the insect receptors. Functional second messenger quantification (IP 1 formation) on CHO cells transiently expressing inotocin receptors (INTR) of L. niger and T. castaneum, respectively, and HEK293 cells expressing human V 1a R. The effect of [D-Arg8]-inotocin was analysed in comparison to endogenous inotocin and desmopressin (1-desamino-8-D-arginine vasopressin, ddavp), carrying also a D-aa analogue in position 8 (n = 3). Cells were treated with 10 µm of each peptide. -S8-
9 Supplementary References 1 Pitti, T. & Manoj, N. Molecular evolution of the neuropeptide S receptor. PLoS One 7, e34046, (2012). -S9-
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
08/21/2017 BLAST. Multiple Sequence Alignments: Clustal Omega
 BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions
 Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions undergo endocytosis into clathrin-coated vesicles dependent
Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions undergo endocytosis into clathrin-coated vesicles dependent
Supporting information
 Supporting information Fluorescent derivatives of AC-42 to probe bitopic orthosteric/allosteric binding mechanisms on muscarinic M1 receptors Sandrine B. Daval, Céline Valant, Dominique Bonnet, Esther
Supporting information Fluorescent derivatives of AC-42 to probe bitopic orthosteric/allosteric binding mechanisms on muscarinic M1 receptors Sandrine B. Daval, Céline Valant, Dominique Bonnet, Esther
Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase Cwc27
 Acta Cryst. (2014). D70, doi:10.1107/s1399004714021695 Supporting information Volume 70 (2014) Supporting information for article: Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase
Acta Cryst. (2014). D70, doi:10.1107/s1399004714021695 Supporting information Volume 70 (2014) Supporting information for article: Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase
SUPPLEMENTARY MATERIAL
 10.101/CH140_AC CSIRO 201 Australian Journal of Chemistry 201, 0(2), 12-11 SUPPLEMENTARY MATERIAL Synthesis of Multivalent [lys ]-Oxytocin Dendrimers that Inhibit Visceral Nociceptive Responses Jingjing
10.101/CH140_AC CSIRO 201 Australian Journal of Chemistry 201, 0(2), 12-11 SUPPLEMENTARY MATERIAL Synthesis of Multivalent [lys ]-Oxytocin Dendrimers that Inhibit Visceral Nociceptive Responses Jingjing
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
MOLECULAR DRUG TARGETS
 MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises
 Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises six Orai subunits, each with identical amino acid sequences
Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises six Orai subunits, each with identical amino acid sequences
In order to confirm the contribution of diffusion to the FRAP recovery curves of
 Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
SIMPLE MODEL Direct Binding Analysis
 Neurochemistry, 56:120:575 Dr. Patrick J. McIlroy Supplementary Notes SIMPLE MODEL Direct Binding Analysis The interaction of a (radio)ligand, L, with its receptor, R, to form a non-covalent complex, RL,
Neurochemistry, 56:120:575 Dr. Patrick J. McIlroy Supplementary Notes SIMPLE MODEL Direct Binding Analysis The interaction of a (radio)ligand, L, with its receptor, R, to form a non-covalent complex, RL,
Molecular docking-based test for affinities of two ligands toward vasopressin and oxytocin receptors
 Vol. 48 No. 1/2001 131 135 QUARTERLY Communication Molecular docking-based test for affinities of two ligands toward vasopressin and oxytocin receptors Rafa³ Œlusarz, Rajmund KaŸmierkiewicz, Artur Gie³doñ,
Vol. 48 No. 1/2001 131 135 QUARTERLY Communication Molecular docking-based test for affinities of two ligands toward vasopressin and oxytocin receptors Rafa³ Œlusarz, Rajmund KaŸmierkiewicz, Artur Gie³doñ,
Supporting Information. Molecular Engineering of Acoustic Protein Nanostructures
 Supporting Information Molecular Engineering of Acoustic Protein Nanostructures Authors: Anupama Lakshmanan, Arash Farhadi, Suchita P. Nety, Audrey Lee-Gosselin, Raymond W. Bourdeau, David Maresca and
Supporting Information Molecular Engineering of Acoustic Protein Nanostructures Authors: Anupama Lakshmanan, Arash Farhadi, Suchita P. Nety, Audrey Lee-Gosselin, Raymond W. Bourdeau, David Maresca and
A High Sensitivity Dual Solid Phase Extraction LC-MS/MS Assay for the Determination of the Therapeutic Peptide Desmopressin in Human Plasma
 White Paper A High Sensitivity Dual Solid Phase Extraction LC-MS/MS Assay for the Determination of the Therapeutic Peptide Desmopressin in Human Plasma Lars Neudert, MSc, Senior Scientist Method Development
White Paper A High Sensitivity Dual Solid Phase Extraction LC-MS/MS Assay for the Determination of the Therapeutic Peptide Desmopressin in Human Plasma Lars Neudert, MSc, Senior Scientist Method Development
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Nature Neuroscience: doi: /nn Supplementary Figure 1. Amygdaloid complex and evoked synaptic currents recorded in CeM amygdala neurons.
 Supplementary Figure 1 Amygdaloid complex and evoked synaptic currents recorded in CeM amygdala neurons. (a) Left: Schematic representation (modified from: Allen Brain Atlas) of coronal sections containing
Supplementary Figure 1 Amygdaloid complex and evoked synaptic currents recorded in CeM amygdala neurons. (a) Left: Schematic representation (modified from: Allen Brain Atlas) of coronal sections containing
Using Bioinformatics to Study Evolutionary Relationships Instructions
 3 Using Bioinformatics to Study Evolutionary Relationships Instructions Student Researcher Background: Making and Using Multiple Sequence Alignments One of the primary tasks of genetic researchers is comparing
3 Using Bioinformatics to Study Evolutionary Relationships Instructions Student Researcher Background: Making and Using Multiple Sequence Alignments One of the primary tasks of genetic researchers is comparing
Supplementary Figures
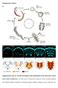 Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Analysis of Allosterism in Functional Assays a
 JPET This Fast article Forward. has not been Published copyedited and on formatted. July 26, The 2005 final as version DOI:10.1124/jpet.105.090886 may differ from this version. Analysis of Allosterism
JPET This Fast article Forward. has not been Published copyedited and on formatted. July 26, The 2005 final as version DOI:10.1124/jpet.105.090886 may differ from this version. Analysis of Allosterism
Nature Structural & Molecular Biology doi: /nsmb Supplementary Figure 1. CRBN binding assay with thalidomide enantiomers.
 Supplementary Figure 1 CRBN binding assay with thalidomide enantiomers. (a) Competitive elution assay using thalidomide-immobilized beads coupled with racemic thalidomide. Beads were washed three times
Supplementary Figure 1 CRBN binding assay with thalidomide enantiomers. (a) Competitive elution assay using thalidomide-immobilized beads coupled with racemic thalidomide. Beads were washed three times
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Discovery of a potent class I PRMT fragment inhibitor Renato Ferreira de Freitas 1,$, Mohammad S. Eram 1,$, Magdalena M. Szewczyk 1, Holger Steuber 2, David Smil 1, Hong Wu 1, Fengling
SUPPORTING INFORMATION Discovery of a potent class I PRMT fragment inhibitor Renato Ferreira de Freitas 1,$, Mohammad S. Eram 1,$, Magdalena M. Szewczyk 1, Holger Steuber 2, David Smil 1, Hong Wu 1, Fengling
Supplementary Figure 1 The number of differentially expressed genes for uniparental males (green), uniparental females (yellow), biparental males
 Supplementary Figure 1 The number of differentially expressed genes for males (green), females (yellow), males (red), and females (blue) in caring vs. control comparisons in the caring gene set and the
Supplementary Figure 1 The number of differentially expressed genes for males (green), females (yellow), males (red), and females (blue) in caring vs. control comparisons in the caring gene set and the
Supplementary Information
 Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
Supporting Text 1. Comparison of GRoSS sequence alignment to HMM-HMM and GPCRDB
 Structure-Based Sequence Alignment of the Transmembrane Domains of All Human GPCRs: Phylogenetic, Structural and Functional Implications, Cvicek et al. Supporting Text 1 Here we compare the GRoSS alignment
Structure-Based Sequence Alignment of the Transmembrane Domains of All Human GPCRs: Phylogenetic, Structural and Functional Implications, Cvicek et al. Supporting Text 1 Here we compare the GRoSS alignment
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Quantitation of the binding of pro53 peptide to sorla Vps10p measured by the AP reporter assay. The graph shows tracings of the typical chromogenic AP reaction observed with AP-pro53
Supplementary Figure 1 Quantitation of the binding of pro53 peptide to sorla Vps10p measured by the AP reporter assay. The graph shows tracings of the typical chromogenic AP reaction observed with AP-pro53
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Carbazole Derivatives Binding to c-kit G-quadruplex DNA
 Supplementary Materials Carbazole Derivatives Binding to c-kit G-quadruplex DNA Agata Głuszyńska 1, *, Bernard Juskowiak 1, Martyna Kuta-Siejkowska 2, Marcin Hoffmann 2 and Shozeb Haider 3 1 Laboratory
Supplementary Materials Carbazole Derivatives Binding to c-kit G-quadruplex DNA Agata Głuszyńska 1, *, Bernard Juskowiak 1, Martyna Kuta-Siejkowska 2, Marcin Hoffmann 2 and Shozeb Haider 3 1 Laboratory
TM-MOTIF: an alignment viewer to annotate predicted transmembrane helices and conserved motifs in aligned set of sequences
 www.bioinformation.net Software Volume 7(5) TM-MOTIF: an alignment viewer to annotate predicted transmembrane helices and conserved motifs in aligned set of sequences Balasubramanian Nagarathnam 1, Kannan
www.bioinformation.net Software Volume 7(5) TM-MOTIF: an alignment viewer to annotate predicted transmembrane helices and conserved motifs in aligned set of sequences Balasubramanian Nagarathnam 1, Kannan
Sequence Based Bioinformatics
 Structural and Functional Analysis of Inosine Monophosphate Dehydrogenase using Sequence-Based Bioinformatics Barry Sexton 1,2 and Troy Wymore 3 1 Bioengineering and Bioinformatics Summer Institute, Department
Structural and Functional Analysis of Inosine Monophosphate Dehydrogenase using Sequence-Based Bioinformatics Barry Sexton 1,2 and Troy Wymore 3 1 Bioengineering and Bioinformatics Summer Institute, Department
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Evolutionary divergence of the plant elicitor peptides (Peps)
 Evolutionary divergence of the plant elicitor peptides (Peps) and their receptors: interfamily incompatibility of perception but compatibility of downstream signalling Item Type Article Authors Lori, M.;
Evolutionary divergence of the plant elicitor peptides (Peps) and their receptors: interfamily incompatibility of perception but compatibility of downstream signalling Item Type Article Authors Lori, M.;
Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides Zaixing
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides Zaixing
RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES
 Molecular Biology-2018 1 Definitions: RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES Heterologues: Genes or proteins that possess different sequences and activities. Homologues: Genes or proteins that
Molecular Biology-2018 1 Definitions: RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES Heterologues: Genes or proteins that possess different sequences and activities. Homologues: Genes or proteins that
Supplementary Figures
 1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Using Phylogenomics to Predict Novel Fungal Pathogenicity Genes
 Using Phylogenomics to Predict Novel Fungal Pathogenicity Genes David DeCaprio, Ying Li, Hung Nguyen (sequenced Ascomycetes genomes courtesy of the Broad Institute) Phylogenomics Combining whole genome
Using Phylogenomics to Predict Novel Fungal Pathogenicity Genes David DeCaprio, Ying Li, Hung Nguyen (sequenced Ascomycetes genomes courtesy of the Broad Institute) Phylogenomics Combining whole genome
Tracking Protein Allostery in Evolution
 Tracking Protein Allostery in Evolution Glycogen phosphorylase frees sugars to provide energy GP orthologs diverged 600,000,000 years can respond to transcription controls, metabolite concentrations and
Tracking Protein Allostery in Evolution Glycogen phosphorylase frees sugars to provide energy GP orthologs diverged 600,000,000 years can respond to transcription controls, metabolite concentrations and
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Supporting Information
 Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Sarcomere length-dependence of total fluorescence intensity in a relaxed muscle fibre containing BSR-RLC. a) Fluorescence intensity (I) relative to the value
Supplementary Figures Supplementary Figure 1. Sarcomere length-dependence of total fluorescence intensity in a relaxed muscle fibre containing BSR-RLC. a) Fluorescence intensity (I) relative to the value
concentration ( mol l -1 )
 concentration ( mol l -1 ) 8 10 0 20 40 60 80 100 120 140 160 180 methane sulfide ammonium oxygen sulfate (/10) b depth (m) 12 14 Supplementary Figure 1. Water column parameters from August 2011. Chemical
concentration ( mol l -1 ) 8 10 0 20 40 60 80 100 120 140 160 180 methane sulfide ammonium oxygen sulfate (/10) b depth (m) 12 14 Supplementary Figure 1. Water column parameters from August 2011. Chemical
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Tree Building Activity
 Tree Building Activity Introduction In this activity, you will construct phylogenetic trees using a phenotypic similarity (cartoon microbe pictures) and genotypic similarity (real microbe sequences). For
Tree Building Activity Introduction In this activity, you will construct phylogenetic trees using a phenotypic similarity (cartoon microbe pictures) and genotypic similarity (real microbe sequences). For
NGF - twenty years a-growing
 NGF - twenty years a-growing A molecule vital to brain growth It is twenty years since the structure of nerve growth factor (NGF) was determined [ref. 1]. This molecule is more than 'quite interesting'
NGF - twenty years a-growing A molecule vital to brain growth It is twenty years since the structure of nerve growth factor (NGF) was determined [ref. 1]. This molecule is more than 'quite interesting'
Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate
 Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate Shidi Zhao a, Laura Restrepo-Pérez b, Misha Soskine c, Giovanni Maglia c, Chirlmin Joo b, Cees Dekker b and Aleksei Aksimentiev
Electro-Mechanical Conductance Modulation of a Nanopore Using a Removable Gate Shidi Zhao a, Laura Restrepo-Pérez b, Misha Soskine c, Giovanni Maglia c, Chirlmin Joo b, Cees Dekker b and Aleksei Aksimentiev
of the Guanine Nucleotide Exchange Factor FARP2
 Structure, Volume 21 Supplemental Information Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2 Xiaojing He, Yi-Chun Kuo, Tyler J. Rosche, and Xuewu Zhang Inventory of
Structure, Volume 21 Supplemental Information Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2 Xiaojing He, Yi-Chun Kuo, Tyler J. Rosche, and Xuewu Zhang Inventory of
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Chemical structure of LPS and LPS biogenesis in Gram-negative bacteria. a. Chemical structure of LPS. LPS molecule consists of Lipid A, core oligosaccharide and O-antigen. The polar
Supplementary Figure 1 Chemical structure of LPS and LPS biogenesis in Gram-negative bacteria. a. Chemical structure of LPS. LPS molecule consists of Lipid A, core oligosaccharide and O-antigen. The polar
Bioinformatics Exercises
 Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Supplementary Information for
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2015 Supplementary Information for The use of Ion Mobility Mass Spectrometry to assist Protein Design:
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2015 Supplementary Information for The use of Ion Mobility Mass Spectrometry to assist Protein Design:
Supplementary Information
 Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Patrick: An Introduction to Medicinal Chemistry 5e Chapter 04
 01) Which of the following statements is not true about receptors? a. Most receptors are proteins situated inside the cell. b. Receptors contain a hollow or cleft on their surface which is known as a binding
01) Which of the following statements is not true about receptors? a. Most receptors are proteins situated inside the cell. b. Receptors contain a hollow or cleft on their surface which is known as a binding
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Disaggregation of Amylin Aggregate by Novel Conformationally. Restricted Aminobenzoic Acid containing α/β and α/γ Hybrid.
 Supporting Information Disaggregation of Amylin Aggregate by Novel Conformationally Restricted Aminobenzoic Acid containing α/β and α/γ Hybrid Peptidomimetics Ashim Paul 1, Sourav Kalita 1, Sujan Kalita
Supporting Information Disaggregation of Amylin Aggregate by Novel Conformationally Restricted Aminobenzoic Acid containing α/β and α/γ Hybrid Peptidomimetics Ashim Paul 1, Sourav Kalita 1, Sujan Kalita
Retrieving hits through in silico screening and expert assessment M. N. Drwal a,b and R. Griffith a
 Retrieving hits through in silico screening and expert assessment M.. Drwal a,b and R. Griffith a a: School of Medical Sciences/Pharmacology, USW, Sydney, Australia b: Charité Berlin, Germany Abstract:
Retrieving hits through in silico screening and expert assessment M.. Drwal a,b and R. Griffith a a: School of Medical Sciences/Pharmacology, USW, Sydney, Australia b: Charité Berlin, Germany Abstract:
Electronic Supplementary Information. A biologically relevant fluorescent probe to assess the binding of ceramide to the CERT transfer protein
 This journal is The Royal Society of Chemistry 20 Electronic Supplementary Information A biologically relevant fluorescent probe to assess the binding of ceramide to the CERT transfer protein Stéphanie
This journal is The Royal Society of Chemistry 20 Electronic Supplementary Information A biologically relevant fluorescent probe to assess the binding of ceramide to the CERT transfer protein Stéphanie
Structural insights into Aspergillus fumigatus lectin specificity - AFL binding sites are functionally non-equivalent
 Acta Cryst. (2015). D71, doi:10.1107/s1399004714026595 Supporting information Volume 71 (2015) Supporting information for article: Structural insights into Aspergillus fumigatus lectin specificity - AFL
Acta Cryst. (2015). D71, doi:10.1107/s1399004714026595 Supporting information Volume 71 (2015) Supporting information for article: Structural insights into Aspergillus fumigatus lectin specificity - AFL
Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae
 Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae Frederik Schulz, Ilias Lagkouvardos, Florian Wascher, Karin Aistleitner, Rok Kostanjšek, Matthias Horn Supplementary
Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae Frederik Schulz, Ilias Lagkouvardos, Florian Wascher, Karin Aistleitner, Rok Kostanjšek, Matthias Horn Supplementary
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/243/ra68/dc1 Supplementary Materials for Superbinder SH2 Domains Act as Antagonists of Cell Signaling Tomonori Kaneko, Haiming Huang, Xuan Cao, Xing Li, Chengjun
www.sciencesignaling.org/cgi/content/full/5/243/ra68/dc1 Supplementary Materials for Superbinder SH2 Domains Act as Antagonists of Cell Signaling Tomonori Kaneko, Haiming Huang, Xuan Cao, Xing Li, Chengjun
FW 1 CDR 1 FW 2 CDR 2
 Supplementary Figure 1 Supplementary Figure 1: Interface of the E9:Fas structure. The two interfaces formed by V H and V L of E9 with Fas are shown in stereo. The Fas receptor is represented as a surface
Supplementary Figure 1 Supplementary Figure 1: Interface of the E9:Fas structure. The two interfaces formed by V H and V L of E9 with Fas are shown in stereo. The Fas receptor is represented as a surface
Computational modeling of G-Protein Coupled Receptors (GPCRs) has recently become
 Homology Modeling and Docking of Melatonin Receptors Andrew Kohlway, UMBC Jeffry D. Madura, Duquesne University 6/18/04 INTRODUCTION Computational modeling of G-Protein Coupled Receptors (GPCRs) has recently
Homology Modeling and Docking of Melatonin Receptors Andrew Kohlway, UMBC Jeffry D. Madura, Duquesne University 6/18/04 INTRODUCTION Computational modeling of G-Protein Coupled Receptors (GPCRs) has recently
SUPPLEMENTARY INFORMATION
 UPPEER ORO doi:10.1038/nature10753 D D D D P E ntracellular C1 W P P C EC1 D Q R H C D W D R C C2 D E D E C R Q Q W P W W R P P EC2 EC3 P C C P W P W W P C W H R C R E C3 P R R P P P C Extracellular embrane
UPPEER ORO doi:10.1038/nature10753 D D D D P E ntracellular C1 W P P C EC1 D Q R H C D W D R C C2 D E D E C R Q Q W P W W R P P EC2 EC3 P C C P W P W W P C W H R C R E C3 P R R P P P C Extracellular embrane
Rho1 binding site PtdIns(4,5)P2 binding site Both sites
 localization Mutation site DMSO LatB WT F77A I115A I131A K134A Rho1 binding site PtdIns(4,5)P2 binding site Both sites E186A E199A N201A R84A-E186A-E199A L131A-K136A-E186A L131A-E186A-E199A K136A-E186A-E199A
localization Mutation site DMSO LatB WT F77A I115A I131A K134A Rho1 binding site PtdIns(4,5)P2 binding site Both sites E186A E199A N201A R84A-E186A-E199A L131A-K136A-E186A L131A-E186A-E199A K136A-E186A-E199A
Supplementary information
 Supplementary information The structural basis of modularity in ECF-type ABC transporters Guus B. Erkens 1,2, Ronnie P-A. Berntsson 1,2, Faizah Fulyani 1,2, Maria Majsnerowska 1,2, Andreja Vujičić-Žagar
Supplementary information The structural basis of modularity in ECF-type ABC transporters Guus B. Erkens 1,2, Ronnie P-A. Berntsson 1,2, Faizah Fulyani 1,2, Maria Majsnerowska 1,2, Andreja Vujičić-Žagar
Structural characterization of NiV N 0 P in solution and in crystal.
 Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Waithe et al Supplementary Figures
 Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Data collection, phasing and refinement statistics ChbC/Ta 6 Br 12 Native ChbC Data collection Space group P4 3 2 1 2 P4 3 2 1 2 Cell dimensions a, c (Å) 132.75, 453.57 132.81, 452.95
Supplementary Table 1: Data collection, phasing and refinement statistics ChbC/Ta 6 Br 12 Native ChbC Data collection Space group P4 3 2 1 2 P4 3 2 1 2 Cell dimensions a, c (Å) 132.75, 453.57 132.81, 452.95
Neurons, Synapses, and Signaling
 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 48 Neurons, Synapses, and Signaling
LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 48 Neurons, Synapses, and Signaling
SUPPLEMENTARY INFORMATION
 Data collection Supplementary Table 1 Statistics of data collection, phasing and refinement Native Se-MAD Space group P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 50.4, 94.2, 115.4 49.8, 94.2,
Data collection Supplementary Table 1 Statistics of data collection, phasing and refinement Native Se-MAD Space group P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 50.4, 94.2, 115.4 49.8, 94.2,
Regression. Estimation of the linear function (straight line) describing the linear component of the joint relationship between two variables X and Y.
 Regression Bivariate i linear regression: Estimation of the linear function (straight line) describing the linear component of the joint relationship between two variables and. Generally describe as a
Regression Bivariate i linear regression: Estimation of the linear function (straight line) describing the linear component of the joint relationship between two variables and. Generally describe as a
Bacterial protease uses distinct thermodynamic signatures for substrate recognition
 Bacterial protease uses distinct thermodynamic signatures for substrate recognition Gustavo Arruda Bezerra, Yuko Ohara-Nemoto, Irina Cornaciu, Sofiya Fedosyuk, Guillaume Hoffmann, Adam Round, José A. Márquez,
Bacterial protease uses distinct thermodynamic signatures for substrate recognition Gustavo Arruda Bezerra, Yuko Ohara-Nemoto, Irina Cornaciu, Sofiya Fedosyuk, Guillaume Hoffmann, Adam Round, José A. Márquez,
SUPPLEMENTARY INFORMATION
 Supplementary Table S1 Kinetic Analyses of the AMSH-LP mutants AMSH-LP K M (μm) k cat x 10-3 (s -1 ) WT 71.8 ± 6.3 860 ± 65.4 T353A 76.8 ± 11.7 46.3 ± 3.7 F355A 58.9 ± 10.4 5.33 ± 0.30 proximal S358A 75.1
Supplementary Table S1 Kinetic Analyses of the AMSH-LP mutants AMSH-LP K M (μm) k cat x 10-3 (s -1 ) WT 71.8 ± 6.3 860 ± 65.4 T353A 76.8 ± 11.7 46.3 ± 3.7 F355A 58.9 ± 10.4 5.33 ± 0.30 proximal S358A 75.1
The circle and the basics of signal transduction. Course Outline. Topic #! Topic lecture! Silverthorn! Membranes (pre-requisite material)" "
 Homeostasis 03 The goal of this lecture is to discuss the concept of homeostasis and to introduce basic signal transduction mechanisms involved in homeostatic regulation The sections for this lecture are:
Homeostasis 03 The goal of this lecture is to discuss the concept of homeostasis and to introduce basic signal transduction mechanisms involved in homeostatic regulation The sections for this lecture are:
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Different crystal forms obtained for Sky
 Supplementary Figure 1 Different crystal forms obtained for Sky 1 353. (a) Crystal form 1 obtained in the presence of 20% PEG 3350 and 0.2 M ammonium citrate tribasic ph 7.0. (b) Crystal form 1 of the
Supplementary Figure 1 Different crystal forms obtained for Sky 1 353. (a) Crystal form 1 obtained in the presence of 20% PEG 3350 and 0.2 M ammonium citrate tribasic ph 7.0. (b) Crystal form 1 of the
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/6/e1700147/dc1 Supplementary Materials for Ribosome rearrangements at the onset of translational bypassing Xabier Agirrezabala, Ekaterina Samatova, Mariia Klimova,
advances.sciencemag.org/cgi/content/full/3/6/e1700147/dc1 Supplementary Materials for Ribosome rearrangements at the onset of translational bypassing Xabier Agirrezabala, Ekaterina Samatova, Mariia Klimova,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11744 Supplementary Table 1. Crystallographic data collection and refinement statistics. Wild-type Se-Met-BcsA-B SmCl 3 -soaked EMTS-soaked Data collection Space
SUPPLEMENTARY INFORMATION doi:10.1038/nature11744 Supplementary Table 1. Crystallographic data collection and refinement statistics. Wild-type Se-Met-BcsA-B SmCl 3 -soaked EMTS-soaked Data collection Space
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R)
 Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Drug binding and subtype selectivity in G-protein-coupled receptors
 Drug binding and subtype selectivity in G-protein-coupled receptors Albert C. Pan The Aspect of Time in Drug Design Schloss Rauischholzhausen Marburg, Germany Thursday, March 27 th, 2014 D. E. Shaw Research
Drug binding and subtype selectivity in G-protein-coupled receptors Albert C. Pan The Aspect of Time in Drug Design Schloss Rauischholzhausen Marburg, Germany Thursday, March 27 th, 2014 D. E. Shaw Research
Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08 Å resolution: comparison with the Azotobacter vinelandii MoFe protein
 Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
From the Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences,16/10 Miklukho- Maklaya Street, Moscow, Russia;
 Supplementary information High-Affinity α-conotoxin PnIA Analogs Designed on the Basis of Protein Surface Topography Method Igor E. Kasheverov 1*, Anton O. Chugunov 1*, Denis S. Kudryavtsev 1, Igor A.
Supplementary information High-Affinity α-conotoxin PnIA Analogs Designed on the Basis of Protein Surface Topography Method Igor E. Kasheverov 1*, Anton O. Chugunov 1*, Denis S. Kudryavtsev 1, Igor A.
Nervous Tissue. Neurons Electrochemical Gradient Propagation & Transduction Neurotransmitters Temporal & Spatial Summation
 Nervous Tissue Neurons Electrochemical Gradient Propagation & Transduction Neurotransmitters Temporal & Spatial Summation What is the function of nervous tissue? Maintain homeostasis & respond to stimuli
Nervous Tissue Neurons Electrochemical Gradient Propagation & Transduction Neurotransmitters Temporal & Spatial Summation What is the function of nervous tissue? Maintain homeostasis & respond to stimuli
5- Semaphorin-Plexin-Neuropilin
 5- Semaphorin-Plexin-Neuropilin 1 SEMAPHORINS-PLEXINS-NEUROPILINS ligands receptors co-receptors semaphorins and their receptors are known signals for: -axon guidance -cell migration -morphogenesis -immune
5- Semaphorin-Plexin-Neuropilin 1 SEMAPHORINS-PLEXINS-NEUROPILINS ligands receptors co-receptors semaphorins and their receptors are known signals for: -axon guidance -cell migration -morphogenesis -immune
Analyzing Radioligand Binding Data
 Analyzing Radioligand Binding Data APPENDIX 3H A radioligand is a radioactively labeled drug that can associate with a receptor, transporter, enzyme, or any protein of interest. The term ligand derives
Analyzing Radioligand Binding Data APPENDIX 3H A radioligand is a radioactively labeled drug that can associate with a receptor, transporter, enzyme, or any protein of interest. The term ligand derives
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11510 Supplementary Table 1. Indel Index Removal Gene Number of Starting Sequences Number of Final Sequences Percentage of Sequences Removed based on the Indel
SUPPLEMENTARY INFORMATION doi:10.1038/nature11510 Supplementary Table 1. Indel Index Removal Gene Number of Starting Sequences Number of Final Sequences Percentage of Sequences Removed based on the Indel
Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets
 Supporting information Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets Wan-Na Chen, Christoph Nitsche, Kala Bharath Pilla, Bim Graham, Thomas
Supporting information Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets Wan-Na Chen, Christoph Nitsche, Kala Bharath Pilla, Bim Graham, Thomas
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.1821 Cloud-based simulations on Google Exacycle reveal ligand-modulation of GPCR activation pathways Kai J. Kohlhoff 1,4 *, Diwakar Shukla 1,2, Morgan Lawrenz 2, Gregory R. Bowman 2,
DOI: 10.1038/NCHEM.1821 Cloud-based simulations on Google Exacycle reveal ligand-modulation of GPCR activation pathways Kai J. Kohlhoff 1,4 *, Diwakar Shukla 1,2, Morgan Lawrenz 2, Gregory R. Bowman 2,
Procheck output. Bond angles (Procheck) Structure verification and validation Bond lengths (Procheck) Introduction to Bioinformatics.
 Structure verification and validation Bond lengths (Procheck) Introduction to Bioinformatics Iosif Vaisman Email: ivaisman@gmu.edu ----------------------------------------------------------------- Bond
Structure verification and validation Bond lengths (Procheck) Introduction to Bioinformatics Iosif Vaisman Email: ivaisman@gmu.edu ----------------------------------------------------------------- Bond
Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction. Lesser Tenrec (Echinops telfairi)
 Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction Lesser Tenrec (Echinops telfairi) Goals: 1. Use phylogenetic experimental design theory to select optimal taxa to
Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction Lesser Tenrec (Echinops telfairi) Goals: 1. Use phylogenetic experimental design theory to select optimal taxa to
SUPPLEMENTARY INFORMATION
 Figure S1 Multiplicative scaling of the granule cell input-output relation is not dependent on input rate. Input-output relations with short-term depression (STD) from Fig. 1d after normalizing by the
Figure S1 Multiplicative scaling of the granule cell input-output relation is not dependent on input rate. Input-output relations with short-term depression (STD) from Fig. 1d after normalizing by the
SUPPLEMENTARY INFORMATION
 Supplementary Results DNA binding property of the SRA domain was examined by an electrophoresis mobility shift assay (EMSA) using synthesized 12-bp oligonucleotide duplexes containing unmodified, hemi-methylated,
Supplementary Results DNA binding property of the SRA domain was examined by an electrophoresis mobility shift assay (EMSA) using synthesized 12-bp oligonucleotide duplexes containing unmodified, hemi-methylated,
Nature Methods: doi: /nmeth Supplementary Figure 1. In vitro screening of recombinant R-CaMP2 variants.
 Supplementary Figure 1 In vitro screening of recombinant R-CaMP2 variants. Baseline fluorescence compared to R-CaMP1.07 at nominally zero calcium plotted versus dynamic range ( F/F) for 150 recombinant
Supplementary Figure 1 In vitro screening of recombinant R-CaMP2 variants. Baseline fluorescence compared to R-CaMP1.07 at nominally zero calcium plotted versus dynamic range ( F/F) for 150 recombinant
A Review of camp-dependent Protein Kinase A Catalytic Subunit Structure, Function and Regulation
 A Review of camp-dependent Protein Kinase A Catalytic Subunit Structure, Function and Regulation Kaitlyn McLeod* Department of Chemistry and Biochemistry, University of Arizona, Tucson, AZ 85721, USA Abstract
A Review of camp-dependent Protein Kinase A Catalytic Subunit Structure, Function and Regulation Kaitlyn McLeod* Department of Chemistry and Biochemistry, University of Arizona, Tucson, AZ 85721, USA Abstract
Supplementary information for:
 SUPPLEMETARY IFRMATI Supplementary information for: Structure of a β 1 -adrenergic G protein-coupled receptor Tony Warne, Maria J. Serrano-Vega, Jillian G. Baker#, Rouslan Moukhametzianov, Patricia C.
SUPPLEMETARY IFRMATI Supplementary information for: Structure of a β 1 -adrenergic G protein-coupled receptor Tony Warne, Maria J. Serrano-Vega, Jillian G. Baker#, Rouslan Moukhametzianov, Patricia C.
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/11/eaao4709/dc1 Supplementary Materials for Pushing the limits of photoreception in twilight conditions: The rod-like cone retina of the deep-sea pearlsides Fanny
advances.sciencemag.org/cgi/content/full/3/11/eaao4709/dc1 Supplementary Materials for Pushing the limits of photoreception in twilight conditions: The rod-like cone retina of the deep-sea pearlsides Fanny
SUPPLEMENTARY INFORMATION
 Table of Contents Page Supplementary Table 1. Diffraction data collection statistics 2 Supplementary Table 2. Crystallographic refinement statistics 3 Supplementary Fig. 1. casic1mfc packing in the R3
Table of Contents Page Supplementary Table 1. Diffraction data collection statistics 2 Supplementary Table 2. Crystallographic refinement statistics 3 Supplementary Fig. 1. casic1mfc packing in the R3
Structural basis of PROTAC cooperative recognition for selective protein degradation
 SUPPLEMENTARY INFORMATION Structural basis of PROTAC cooperative recognition for selective protein degradation Morgan S. Gadd 1, Andrea Testa 1, Xavier Lucas 1, Kwok-Ho Chan, Wenzhang Chen, Douglas J.
SUPPLEMENTARY INFORMATION Structural basis of PROTAC cooperative recognition for selective protein degradation Morgan S. Gadd 1, Andrea Testa 1, Xavier Lucas 1, Kwok-Ho Chan, Wenzhang Chen, Douglas J.
