A New Approach to EM of Helical Polymers Yields New Insights
|
|
|
- Alexandrina Townsend
- 6 years ago
- Views:
Transcription
1 A New Approach to EM of Helical Polymers Yields New Insights Helical polymers are ubiquitous in biology Actin,, microtubules, intermediate filaments, thick filaments, viruses, bacteriophage,, flagella, pili, RecA,, Rad51 Helical objects were the first to be reconstructed in 3D in the EM DeRosier & Klug (1968), tail of bacteriophage T4 Many advances since then Structure of one natural helical polymer (bacterial flagellar filament) determined at near atomic resolution (Yonekura( et al.,., 2003) Membrane proteins have been induced to form in vitro helical tubes for high resolution structural studies
2 Some definitions What is helical symmetry? A screw operation involves a coupled rotation and axial translation. Some terminology: axial rise is Δz rotation is Δϕ helical repeat (c) is the translation along the axis needed to bring one subunit into exact superposition with another subunit for an integer number of subunits/turn, or units/turn (u/t( u/t), repeat is given by u * Δz z = c in a crystal, the only allowed helical symmetries involve 2, 3, 4 or 6 subunits/turn outside of a crystal, there is no reason for any helix to have an integer number of u/t! outside of a crystal, there is no space group maintaining long- range order. So cannot have true 1D crystal
3 Problems with definition of helical repeat Very small changes in symmetry can lead to very large changes in the repeat Example of actin: u/t = 13/6,, c=355 Å, Δϕ= = º but Δ(Δϕ)=0.128º, Δϕ= = º, u/t=1299/600, c=35,463 Å!
4 In simplest case: Each layer line contains a single Bessel function indexing pattern requires determining Bessel function order for only two layer lines 3D reconstruction can then be made by Fourier-Bessel inversion If a polymer is highly ordered, homogeneous, does not have Bessel overlap, Fourier-Bessel methods work very well
5 But Most Helical Polymers Have Been Refractory to High Resolution EM Studies! Disorder or variability Heterogeneity (it is much greater than has been assumed!) Weak Scattering Bessel Overlap
6 New method: Iterative Helical Real Space Reconstruction and ~ 20 other papers published or in press
7 Iterative Helical Real Space Reconstruction cycle Egelman (2000), Ultramicroscopy 85, impose helical symmetry Helically symmetric 3D volume rotate azimuthally by 4 increment, project onto 2D image Δφ, Δz (helical parameters) determine helical symmetry by least squares fit Asymmetric 3D volume 90 reference projections multi-reference alignment In-plane rotation, x-shift, y-shift parameters set of raw images (thousands of segments from helical filaments) back projection Aligned, rotated images; azimuthal angular assignments
8 Algorithm is robust in that it is independent of starting model
9 Disorder can be enormous Dynamin-lipid tubes Chen et al., Nature SMB (2004) RecA-DNA filaments VanLoock et al., JMB (2003)
10 Myosin thick filament has been almost intractable The helical symmetry is such that, on a given layer- line, Bessel function contributions of different orders start to overlap at fairly low resolution and must therefore be separated computationally by combining data from different views
11 Problem non-existent using IHRSR Woodhead et al.,, Nature 436, (2005)
12 Resolution sufficient to fit atomic model Woodhead et al.,, Nature 436, (2005)
13 EspA of Enteropathogenic E. coli
14 EspA of Enteropathogenic E. coli Daniell et al., Mol. Micro. (2003)
15 EspA of Enteropathogenic E. coli
16 EspA of Enteropathogenic E. coli Such behavior is a prima facie argument for heterogeneity!
17 EspA of Enteropathogenic E. coli Most variation determined to be due to variable axial rise but is this sorting valid? Is there a reality check?
18 EspA of Enteropathogenic E. coli Power spectra show change in axial rise, no change in twist
19 EspA: : heterogeneity in axial rise Grey: 5.3 Å Yellow: 4.2 Å Right-handed 6-start Left-handed 5-start
20 EspA of Enteropathogenic E. coli
21 EspA of Enteropathogenic E. coli What happens in the central bin?
22 EspA of Enteropathogenic E. coli central bin (~ 4.6 Å) shows no major variability in rise, twist, but can separate out two states of connectivity around lumen!
23 Bacterial pili: : example of even more weakly scattering objects
24 Unambiguous fit of monomeric crystal subunit from Neisseria gonorrhoeae
25 Very weak scattering is no longer a problem!
26 Filamentous bacteriophage (M13/fd) Model systems in understanding: DNA packaging Assembly of a protein polymer from a small integral membrane protein Important in cloning, phage display, etc. Such polymorphism should not be surprising, given that 41/50 residues can be mutated to Ala and the subunit still co-assembles almost as efficiently as wt! (Roth et al., JMB 322,357-67, 2002)
27 Filamentous bacteriophage fd 1/(7 Å)
28 EM reconstruction cannot be reconciled with existing models Cyan: Solid State NMR Model, Zeri et al., PNAS (2003) - calculation of the threedimensional structure of the protein from orientational restraints with an accuracy equivalent to an rms deviation of approximately 1 Å Red: X-ray fiber diffraction (refined against NMR data), Marvin et al., JMB (2006) Here we show that reinterpreted NMR data are also consistent with the model derived from X-ray fibre diffraction studies
29 Conclusions I Outside of a crystal, there is nothing to maintain long-range order in a macromolecular assembly Many helical protein polymers are more polymorphic than assumed Originally showed this for F-actinF and RecA Can now demonstrate similar polymorphisms in bacteriophage fd,, T3SS EspA, dynamin tubes, Plasticity of protein-protein interface in these structures must reflect evolutionary selection
30 Conclusions II Polymorphism and heterogeneity not necessarily revealed by casual observation or global averaging Bessel overlap occurs in many structures where it cannot be ignored In every instance, we can do better with the IHRSR approach than Fourier- Bessel methods
31 Acknowledgments Vitold Galkin, Albina Orlova, Xiong Yu, Ying Wang, Olga Cherepanova Margaret VanLoock, Yen-Ju Chen, Natasha Lukoyanova Myosin: Raul Padron (Caracas), Roger Craig (U. Mass.) EspA: : Natalie Strynadka (UBC) GC Pili: : John Tainer (Scripps), Lisa Craig (Simon Fraser) Bacteriophage fd: : George Thomas (UMKC)
History of 3D Electron Microscopy and Helical Reconstruction
 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N History of 3D Electron Microscopy and Helical Reconstruction For students of HI
T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N History of 3D Electron Microscopy and Helical Reconstruction For students of HI
ParM filament images were extracted and from the electron micrographs and
 Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Nuclear import of DNA: genetic modification of plants
 Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
A New Internal Mode in F-Actin Helps Explain the Remarkable Evolutionary Conservation of Actin s Sequence and Structure
 Current Biology, Vol. 12, 570 575, April 2, 2002, 2002 Elsevier Science Ltd. All rights reserved. PII S0960-9822(02)00742-X A New Internal Mode in F-Actin Helps Explain the Remarkable Evolutionary Conservation
Current Biology, Vol. 12, 570 575, April 2, 2002, 2002 Elsevier Science Ltd. All rights reserved. PII S0960-9822(02)00742-X A New Internal Mode in F-Actin Helps Explain the Remarkable Evolutionary Conservation
Three-dimensional structure of a viral genome-delivery portal vertex
 Three-dimensional structure of a viral genome-delivery portal vertex Adam S. Olia 1, Peter E. Prevelige Jr. 2, John E. Johnson 3 and Gino Cingolani 4 1 Department of Biological Sciences, Purdue University,
Three-dimensional structure of a viral genome-delivery portal vertex Adam S. Olia 1, Peter E. Prevelige Jr. 2, John E. Johnson 3 and Gino Cingolani 4 1 Department of Biological Sciences, Purdue University,
NOTE: Questions are written on both sides of the sheets of paper making up this exam booklet!
 Biology 1010 Section A Midterm 1 January 30, 2008 (print): ANSWER KEY Name (signature): Student I.D. #: Time: 50 minutes Read the following instructions: 1. Do not open the examination until you are instructed
Biology 1010 Section A Midterm 1 January 30, 2008 (print): ANSWER KEY Name (signature): Student I.D. #: Time: 50 minutes Read the following instructions: 1. Do not open the examination until you are instructed
R-factor-fit R-factor-non-fit
 Supplementary Figures and Legends R-factor-fit R-factor-non-fit R-factor-non-fit (overall) Supplementary figure 1 The areas where the R-factor-fit and the R-factor-non-fit were calculated. The edges of
Supplementary Figures and Legends R-factor-fit R-factor-non-fit R-factor-non-fit (overall) Supplementary figure 1 The areas where the R-factor-fit and the R-factor-non-fit were calculated. The edges of
NMR, X-ray Diffraction, Protein Structure, and RasMol
 NMR, X-ray Diffraction, Protein Structure, and RasMol Introduction So far we have been mostly concerned with the proteins themselves. The techniques (NMR or X-ray diffraction) used to determine a structure
NMR, X-ray Diffraction, Protein Structure, and RasMol Introduction So far we have been mostly concerned with the proteins themselves. The techniques (NMR or X-ray diffraction) used to determine a structure
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.1290 Metal-directed, chemically tunable assembly of one-, two- and threedimensional crystalline protein arrays 1 2 1 1 1,3 Jeffrey D. Brodin 1, X. I. Ambroggio 2, Chunyan Tang 1, Kristin
DOI: 10.1038/NCHEM.1290 Metal-directed, chemically tunable assembly of one-, two- and threedimensional crystalline protein arrays 1 2 1 1 1,3 Jeffrey D. Brodin 1, X. I. Ambroggio 2, Chunyan Tang 1, Kristin
T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y. jgp
 S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
Application examples of single particle 3D reconstruction. Ning Gao Tsinghua University
 Application examples of single particle 3D reconstruction Ning Gao Tsinghua University ninggao@tsinghua.edu.cn Electron Microscopes First electron microscope constructed by Ernst Ruska in 1930 s (1986
Application examples of single particle 3D reconstruction Ning Gao Tsinghua University ninggao@tsinghua.edu.cn Electron Microscopes First electron microscope constructed by Ernst Ruska in 1930 s (1986
Three-Dimensional Electron Microscopy of Macromolecular Assemblies
 Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Supramolecular self-assembly: molecular dynamics modeling of polyhedral shell formation
 Computer Physics Communications 121 122 (1999) 231 235 www.elsevier.nl/locate/cpc Supramolecular self-assembly: molecular dynamics modeling of polyhedral shell formation D.C. Rapaport a,1, J.E. Johnson
Computer Physics Communications 121 122 (1999) 231 235 www.elsevier.nl/locate/cpc Supramolecular self-assembly: molecular dynamics modeling of polyhedral shell formation D.C. Rapaport a,1, J.E. Johnson
sin" =1.22 # D "l =1.22 f# D I: In introduction to molecular electron microscopy - Imaging macromolecular assemblies
 I: In introduction to molecular electron microscopy - Imaging macromolecular assemblies Yifan Cheng Department of Biochemistry & Biophysics office: GH-S472D; email: ycheng@ucsf.edu 2/20/2015 - Introduction
I: In introduction to molecular electron microscopy - Imaging macromolecular assemblies Yifan Cheng Department of Biochemistry & Biophysics office: GH-S472D; email: ycheng@ucsf.edu 2/20/2015 - Introduction
Fig. 1. Stereo images showing (A) the best fit of the atomic model for F actin and the F actin map obtained by cryo-em and image analysis, and (B) goo
 Fig. 1. Stereo images showing (A) the best fit of the atomic model for F actin and the F actin map obtained by cryo-em and image analysis, and (B) good correspondence between the location of Cys374 and
Fig. 1. Stereo images showing (A) the best fit of the atomic model for F actin and the F actin map obtained by cryo-em and image analysis, and (B) good correspondence between the location of Cys374 and
Supplementary Information. Cryo-EM Studies of Drp1 Reveal Cardiolipin Interactions that Activate
 Supplementary Information Cryo-EM Studies of Drp1 Reveal Cardiolipin Interactions that Activate the Helical Oligomer Christopher A. Francy 1, 2, 3, Ryan W. Clinton 1, 2, 3, Chris Fröhlich 4, 5, Colleen
Supplementary Information Cryo-EM Studies of Drp1 Reveal Cardiolipin Interactions that Activate the Helical Oligomer Christopher A. Francy 1, 2, 3, Ryan W. Clinton 1, 2, 3, Chris Fröhlich 4, 5, Colleen
Structure of the α-helix
 Structure of the α-helix Structure of the β Sheet Protein Dynamics Basics of Quenching HDX Hydrogen exchange of amide protons is catalyzed by H 2 O, OH -, and H 3 O +, but it s most dominated by base
Structure of the α-helix Structure of the β Sheet Protein Dynamics Basics of Quenching HDX Hydrogen exchange of amide protons is catalyzed by H 2 O, OH -, and H 3 O +, but it s most dominated by base
Crystallographic Conformers of Actin in a Biologically Active Bundle of Filaments
 doi:10.1016/j.jmb.2007.10.027 J. Mol. Biol. (2008) 375, 331 336 Available online at www.sciencedirect.com COMMUNICATION Crystallographic Conformers of Actin in a Biologically Active Bundle of Filaments
doi:10.1016/j.jmb.2007.10.027 J. Mol. Biol. (2008) 375, 331 336 Available online at www.sciencedirect.com COMMUNICATION Crystallographic Conformers of Actin in a Biologically Active Bundle of Filaments
Scattering Lecture. February 24, 2014
 Scattering Lecture February 24, 2014 Structure Determination by Scattering Waves of radiation scattered by different objects interfere to give rise to an observable pattern! The wavelength needs to close
Scattering Lecture February 24, 2014 Structure Determination by Scattering Waves of radiation scattered by different objects interfere to give rise to an observable pattern! The wavelength needs to close
Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy
 Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy Amit Singer Princeton University, Department of Mathematics and PACM July 23, 2014 Amit Singer (Princeton
Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy Amit Singer Princeton University, Department of Mathematics and PACM July 23, 2014 Amit Singer (Princeton
Cells and the Stuff They re Made of. Indiana University P575 1
 Cells and the Stuff They re Made of Indiana University P575 1 Goal: Establish hierarchy of spatial and temporal scales, and governing processes at each scale of cellular function: o Where does emergent
Cells and the Stuff They re Made of Indiana University P575 1 Goal: Establish hierarchy of spatial and temporal scales, and governing processes at each scale of cellular function: o Where does emergent
NONLINEAR STOCHASTIC TOMOGRAPHY RECONSTRUCTION ALGORITHMS FOR OBJECTS WITH HELICAL SYMMETRY AND APPLICATIONS TO VIRUS STRUCTURES
 NONLINEAR STOCHASTIC TOMOGRAPHY RECONSTRUCTION ALGORITHMS FOR OBJECTS WITH HELICAL SYMMETRY AND APPLICATIONS TO VIRUS STRUCTURES A Thesis Presented to the Faculty of the Graduate School of Cornell University
NONLINEAR STOCHASTIC TOMOGRAPHY RECONSTRUCTION ALGORITHMS FOR OBJECTS WITH HELICAL SYMMETRY AND APPLICATIONS TO VIRUS STRUCTURES A Thesis Presented to the Faculty of the Graduate School of Cornell University
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass
 Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass University of California, Davis, USA http://www.cs.ucdavis.edu/~koehl/ims2017/ Biology = Multiscale. 10 6 m 10 3 m m mm µm nm Å ps
Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass University of California, Davis, USA http://www.cs.ucdavis.edu/~koehl/ims2017/ Biology = Multiscale. 10 6 m 10 3 m m mm µm nm Å ps
The biological motors
 Motor proteins The definition of motor proteins Miklós Nyitrai, November 30, 2016 Molecular machines key to understand biological processes machines in the micro/nano-world (unidirectional steps): nm,
Motor proteins The definition of motor proteins Miklós Nyitrai, November 30, 2016 Molecular machines key to understand biological processes machines in the micro/nano-world (unidirectional steps): nm,
SUPPLEMENTARY FIGURES. Structure of the cholera toxin secretion channel in its. closed state
 SUPPLEMENTARY FIGURES Structure of the cholera toxin secretion channel in its closed state Steve L. Reichow 1,3, Konstantin V. Korotkov 1,3, Wim G. J. Hol 1$ and Tamir Gonen 1,2$ 1, Department of Biochemistry
SUPPLEMENTARY FIGURES Structure of the cholera toxin secretion channel in its closed state Steve L. Reichow 1,3, Konstantin V. Korotkov 1,3, Wim G. J. Hol 1$ and Tamir Gonen 1,2$ 1, Department of Biochemistry
Introduction to" Protein Structure
 Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Protein Crystallography
 Protein Crystallography Part II Tim Grüne Dept. of Structural Chemistry Prof. G. Sheldrick University of Göttingen http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview The Reciprocal Lattice The
Protein Crystallography Part II Tim Grüne Dept. of Structural Chemistry Prof. G. Sheldrick University of Göttingen http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview The Reciprocal Lattice The
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12045 Supplementary Table 1 Data collection and refinement statistics. Native Pt-SAD X-ray source SSRF BL17U SPring-8 BL41XU Wavelength (Å) 0.97947 1.07171 Space group P2 1 2 1 2 1 P2
doi:10.1038/nature12045 Supplementary Table 1 Data collection and refinement statistics. Native Pt-SAD X-ray source SSRF BL17U SPring-8 BL41XU Wavelength (Å) 0.97947 1.07171 Space group P2 1 2 1 2 1 P2
Part 1 X-ray Crystallography
 Part 1 X-ray Crystallography What happens to electron when it is hit by x-rays? 1. The electron starts vibrating with the same frequency as the x-ray beam 2. As a result, secondary beams will be scattered
Part 1 X-ray Crystallography What happens to electron when it is hit by x-rays? 1. The electron starts vibrating with the same frequency as the x-ray beam 2. As a result, secondary beams will be scattered
Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche
 Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche The molecular structure of a protein can be broken down hierarchically. The primary structure of a protein is simply its
Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche The molecular structure of a protein can be broken down hierarchically. The primary structure of a protein is simply its
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Measuring quaternary structure similarity using global versus local measures.
 Supplementary Figure 1 Measuring quaternary structure similarity using global versus local measures. (a) Structural similarity of two protein complexes can be inferred from a global superposition, which
Supplementary Figure 1 Measuring quaternary structure similarity using global versus local measures. (a) Structural similarity of two protein complexes can be inferred from a global superposition, which
Building a Homology Model of the Transmembrane Domain of the Human Glycine α-1 Receptor
 Building a Homology Model of the Transmembrane Domain of the Human Glycine α-1 Receptor Presented by Stephanie Lee Research Mentor: Dr. Rob Coalson Glycine Alpha 1 Receptor (GlyRa1) Member of the superfamily
Building a Homology Model of the Transmembrane Domain of the Human Glycine α-1 Receptor Presented by Stephanie Lee Research Mentor: Dr. Rob Coalson Glycine Alpha 1 Receptor (GlyRa1) Member of the superfamily
Copyright Mark Brandt, Ph.D A third method, cryogenic electron microscopy has seen increasing use over the past few years.
 Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Lecture CIMST Winter School. 1. What can you see by TEM?
 Lecture CIMST Winter School Cryo-electron microscopy and tomography of biological macromolecules 20.1.2011 9:00-9:45 in Y03G91 Dr. Takashi Ishikawa OFLB/005 Tel: 056 310 4217 e-mail: takashi.ishikawa@psi.ch
Lecture CIMST Winter School Cryo-electron microscopy and tomography of biological macromolecules 20.1.2011 9:00-9:45 in Y03G91 Dr. Takashi Ishikawa OFLB/005 Tel: 056 310 4217 e-mail: takashi.ishikawa@psi.ch
1) NMR is a method of chemical analysis. (Who uses NMR in this way?) 2) NMR is used as a method for medical imaging. (called MRI )
 Uses of NMR: 1) NMR is a method of chemical analysis. (Who uses NMR in this way?) 2) NMR is used as a method for medical imaging. (called MRI ) 3) NMR is used as a method for determining of protein, DNA,
Uses of NMR: 1) NMR is a method of chemical analysis. (Who uses NMR in this way?) 2) NMR is used as a method for medical imaging. (called MRI ) 3) NMR is used as a method for determining of protein, DNA,
SUPPLEMENTARY INFORMATION
 www.nature.com/nature 1 Figure S1 Sequence alignment. a Structure based alignment of the plgic of E. chrysanthemi (ELIC), the acetylcholine binding protein from the snail Lymnea stagnalis (AchBP, PDB code
www.nature.com/nature 1 Figure S1 Sequence alignment. a Structure based alignment of the plgic of E. chrysanthemi (ELIC), the acetylcholine binding protein from the snail Lymnea stagnalis (AchBP, PDB code
Structural Studies of the Flexible Filamentous Plant Viruses
 Structural Studies of the Flexible Filamentous Plant Viruses Caitlin Azzo Dr. Gerald Stubbs BSCI 296 [4] Honors Thesis April 23, 2014 The Plant Viruses of Interest In the Potyviridae family: Wheat streak
Structural Studies of the Flexible Filamentous Plant Viruses Caitlin Azzo Dr. Gerald Stubbs BSCI 296 [4] Honors Thesis April 23, 2014 The Plant Viruses of Interest In the Potyviridae family: Wheat streak
Biophysics 490M Project
 Biophysics 490M Project Dan Han Department of Biochemistry Structure Exploration of aa 3 -type Cytochrome c Oxidase from Rhodobacter sphaeroides I. Introduction: All organisms need energy to live. They
Biophysics 490M Project Dan Han Department of Biochemistry Structure Exploration of aa 3 -type Cytochrome c Oxidase from Rhodobacter sphaeroides I. Introduction: All organisms need energy to live. They
Structure Analysis of the Flagellar Cap Filament Complex by Electron Cryomicroscopy and Single-Particle Image Analysis
 Journal of Structural Biology 133, 246 253 (2001) doi:10.1006/jsbi.2000.4345, available online at http://www.idealibrary.com on Structure Analysis of the Flagellar Cap Filament Complex by Electron Cryomicroscopy
Journal of Structural Biology 133, 246 253 (2001) doi:10.1006/jsbi.2000.4345, available online at http://www.idealibrary.com on Structure Analysis of the Flagellar Cap Filament Complex by Electron Cryomicroscopy
Molecular Mechanisms of self-assembly and motion of the bacterial flagellum
 Molecular Mechanisms of self-assembly and motion of the bacterial flagellum Mexico-Japan Workshop on Pharmacobiology and Nanobiology Universidad Nacional Autonoma de México (UNAM) Mexico City, Mexico 2009.02.25-26
Molecular Mechanisms of self-assembly and motion of the bacterial flagellum Mexico-Japan Workshop on Pharmacobiology and Nanobiology Universidad Nacional Autonoma de México (UNAM) Mexico City, Mexico 2009.02.25-26
I690/B680 Structural Bioinformatics Spring Protein Structure Determination by NMR Spectroscopy
 I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
Molecular electron microscopy
 Molecular electron microscopy - Imaging macromolecular assemblies Yifan Cheng Department of Biochemistry & Biophysics office: GH-S427B; email: ycheng@ucsf.edu 2/22/2013 - Introduction of Molecular Microscopy:
Molecular electron microscopy - Imaging macromolecular assemblies Yifan Cheng Department of Biochemistry & Biophysics office: GH-S427B; email: ycheng@ucsf.edu 2/22/2013 - Introduction of Molecular Microscopy:
monomer polymer polymeric network cell
 3.1 Motivation 3.2 Polymerization The structural stability of the cell is provided by the cytoskeleton. Assembling and disassembling dynamically, the cytoskeleton enables cell movement through a highly
3.1 Motivation 3.2 Polymerization The structural stability of the cell is provided by the cytoskeleton. Assembling and disassembling dynamically, the cytoskeleton enables cell movement through a highly
The Potassium Ion Channel: Rahmat Muhammad
 The Potassium Ion Channel: 1952-1998 1998 Rahmat Muhammad Ions: Cell volume regulation Electrical impulse formation (e.g. sodium, potassium) Lipid membrane: the dielectric barrier Pro: compartmentalization
The Potassium Ion Channel: 1952-1998 1998 Rahmat Muhammad Ions: Cell volume regulation Electrical impulse formation (e.g. sodium, potassium) Lipid membrane: the dielectric barrier Pro: compartmentalization
Phase problem: Determining an initial phase angle α hkl for each recorded reflection. 1 ρ(x,y,z) = F hkl cos 2π (hx+ky+ lz - α hkl ) V h k l
 Phase problem: Determining an initial phase angle α hkl for each recorded reflection 1 ρ(x,y,z) = F hkl cos 2π (hx+ky+ lz - α hkl ) V h k l Methods: Heavy atom methods (isomorphous replacement Hg, Pt)
Phase problem: Determining an initial phase angle α hkl for each recorded reflection 1 ρ(x,y,z) = F hkl cos 2π (hx+ky+ lz - α hkl ) V h k l Methods: Heavy atom methods (isomorphous replacement Hg, Pt)
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR
 Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR Residual Dipolar Couplings (RDC s) Relatively new technique ~ 1996 Nico Tjandra, Ad Bax- NIH, Jim Prestegard, UGA Combination of
Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR Residual Dipolar Couplings (RDC s) Relatively new technique ~ 1996 Nico Tjandra, Ad Bax- NIH, Jim Prestegard, UGA Combination of
Atomic structure and handedness of the building block of a biological assembly
 Supporting Information: Atomic structure and handedness of the building block of a biological assembly Antoine Loquet, Birgit Habenstein, Veniamin Chevelkov, Suresh Kumar Vasa, Karin Giller, Stefan Becker,
Supporting Information: Atomic structure and handedness of the building block of a biological assembly Antoine Loquet, Birgit Habenstein, Veniamin Chevelkov, Suresh Kumar Vasa, Karin Giller, Stefan Becker,
Fourier Series. Combination of Waves: Any PERIODIC function f(t) can be written: How to calculate the coefficients?
 Fourier Series Combination of Waves: Any PERIODIC function f(t) can be written: How to calculate the coefficients? What is the Fourier Transform It is the representation of an arbitrary NON-periodic signal
Fourier Series Combination of Waves: Any PERIODIC function f(t) can be written: How to calculate the coefficients? What is the Fourier Transform It is the representation of an arbitrary NON-periodic signal
SUPPLEMENTARY INFORMATION
 Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Structure Analysis by Small-Angle X-Ray and Neutron Scattering
 Structure Analysis by Small-Angle X-Ray and Neutron Scattering L. A. Feigin and D. I. Svergun Institute of Crystallography Academy of Sciences of the USSR Moscow, USSR Edited by George W. Taylor Princeton
Structure Analysis by Small-Angle X-Ray and Neutron Scattering L. A. Feigin and D. I. Svergun Institute of Crystallography Academy of Sciences of the USSR Moscow, USSR Edited by George W. Taylor Princeton
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
Polymerization/depolymerization motors
 Polymerization/depolymerization motors Movement formation Kuo Lab, J.H.U. http://www.nature.com/nature/journal/v407/n6807/extref/40 71026a0_S3.mov http://www.bme.jhu.edu/~skuo/movies/macrophchase.mov http://www.bme.jhu.edu/~skuo/movies/gc_filo.mov
Polymerization/depolymerization motors Movement formation Kuo Lab, J.H.U. http://www.nature.com/nature/journal/v407/n6807/extref/40 71026a0_S3.mov http://www.bme.jhu.edu/~skuo/movies/macrophchase.mov http://www.bme.jhu.edu/~skuo/movies/gc_filo.mov
Nature Structural & Molecular Biology: doi: /nsmb.3194
 Supplementary Figure 1 Mass spectrometry and solution NMR data for -syn samples used in this study. (a) Matrix-assisted laser-desorption and ionization time-of-flight (MALDI-TOF) mass spectrum of uniformly-
Supplementary Figure 1 Mass spectrometry and solution NMR data for -syn samples used in this study. (a) Matrix-assisted laser-desorption and ionization time-of-flight (MALDI-TOF) mass spectrum of uniformly-
Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and
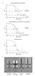 Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and comparisons to other published in situ T3SS structures. a, Resolution estimates after applying Fourier shell correlation
Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and comparisons to other published in situ T3SS structures. a, Resolution estimates after applying Fourier shell correlation
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN
 THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
Protein crystallography. Garry Taylor
 Protein crystallography Garry Taylor X-ray Crystallography - the Basics Grow crystals Collect X-ray data Determine phases Calculate ρ-map Interpret map Refine coordinates Do the biology. Nitrogen at -180
Protein crystallography Garry Taylor X-ray Crystallography - the Basics Grow crystals Collect X-ray data Determine phases Calculate ρ-map Interpret map Refine coordinates Do the biology. Nitrogen at -180
Chapter 6: A Tour of the Cell
 AP Biology Reading Guide Fred and Theresa Holtzclaw Chapter 6: A Tour of the Cell Name Period Chapter 6: A Tour of the Cell Concept 6.1 To study cells, biologists use microscopes and the tools of biochemistry
AP Biology Reading Guide Fred and Theresa Holtzclaw Chapter 6: A Tour of the Cell Name Period Chapter 6: A Tour of the Cell Concept 6.1 To study cells, biologists use microscopes and the tools of biochemistry
X-ray Crystallography. Kalyan Das
 X-ray Crystallography Kalyan Das Electromagnetic Spectrum NMR 10 um - 10 mm 700 to 10 4 nm 400 to 700 nm 10 to 400 nm 10-1 to 10 nm 10-4 to 10-1 nm X-ray radiation was discovered by Roentgen in 1895. X-rays
X-ray Crystallography Kalyan Das Electromagnetic Spectrum NMR 10 um - 10 mm 700 to 10 4 nm 400 to 700 nm 10 to 400 nm 10-1 to 10 nm 10-4 to 10-1 nm X-ray radiation was discovered by Roentgen in 1895. X-rays
Medical Biophysics II. Final exam theoretical questions 2013.
 Medical Biophysics II. Final exam theoretical questions 2013. 1. Early atomic models. Rutherford-experiment. Franck-Hertz experiment. Bohr model of atom. 2. Quantum mechanical atomic model. Quantum numbers.
Medical Biophysics II. Final exam theoretical questions 2013. 1. Early atomic models. Rutherford-experiment. Franck-Hertz experiment. Bohr model of atom. 2. Quantum mechanical atomic model. Quantum numbers.
Scattering of Neutrons: Basics. Jill Trewhella University of Sydney
 Scattering of Neutrons: Basics Jill Trewhella University of Sydney Small-angle scattering of x-rays (or neutrons) tells us about the size and shape of macromolecules r r Sample -randomly oriented particles
Scattering of Neutrons: Basics Jill Trewhella University of Sydney Small-angle scattering of x-rays (or neutrons) tells us about the size and shape of macromolecules r r Sample -randomly oriented particles
Protein Folding Prof. Eugene Shakhnovich
 Protein Folding Eugene Shakhnovich Department of Chemistry and Chemical Biology Harvard University 1 Proteins are folded on various scales As of now we know hundreds of thousands of sequences (Swissprot)
Protein Folding Eugene Shakhnovich Department of Chemistry and Chemical Biology Harvard University 1 Proteins are folded on various scales As of now we know hundreds of thousands of sequences (Swissprot)
Overview - Macromolecular Crystallography
 Overview - Macromolecular Crystallography 1. Overexpression and crystallization 2. Crystal characterization and data collection 3. The diffraction experiment 4. Phase problem 1. MIR (Multiple Isomorphous
Overview - Macromolecular Crystallography 1. Overexpression and crystallization 2. Crystal characterization and data collection 3. The diffraction experiment 4. Phase problem 1. MIR (Multiple Isomorphous
SUPPLEMENTARY INFORMATION
 SUPPLMTARY IFORMATIO a doi:10.108/nature10402 b 100 nm 100 nm c SAXS Model d ulers assigned to reference- Back-projected free class averages class averages Refinement against single particles Reconstructed
SUPPLMTARY IFORMATIO a doi:10.108/nature10402 b 100 nm 100 nm c SAXS Model d ulers assigned to reference- Back-projected free class averages class averages Refinement against single particles Reconstructed
research papers 1. Introduction Thomas C. Terwilliger a * and Joel Berendzen b
 Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 Discrimination of solvent from protein regions in native Fouriers as a means of evaluating heavy-atom solutions in the MIR and
Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 Discrimination of solvent from protein regions in native Fouriers as a means of evaluating heavy-atom solutions in the MIR and
Full wwpdb X-ray Structure Validation Report i
 Full wwpdb X-ray Structure Validation Report i Mar 14, 2018 02:00 pm GMT PDB ID : 3RRQ Title : Crystal structure of the extracellular domain of human PD-1 Authors : Lazar-Molnar, E.; Ramagopal, U.A.; Nathenson,
Full wwpdb X-ray Structure Validation Report i Mar 14, 2018 02:00 pm GMT PDB ID : 3RRQ Title : Crystal structure of the extracellular domain of human PD-1 Authors : Lazar-Molnar, E.; Ramagopal, U.A.; Nathenson,
Table of contents. Saturday 24 June Sunday 25 June Monday 26 June
 Table of contents Saturday 24 June 2017... 1 Sunday 25 June 2017... 3 Monday 26 June 2017... 5 i Saturday 24 June 2017 Quantitative Biology 2017: Computational and Single-Molecule Biophysics Saturday 24
Table of contents Saturday 24 June 2017... 1 Sunday 25 June 2017... 3 Monday 26 June 2017... 5 i Saturday 24 June 2017 Quantitative Biology 2017: Computational and Single-Molecule Biophysics Saturday 24
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11744 Supplementary Table 1. Crystallographic data collection and refinement statistics. Wild-type Se-Met-BcsA-B SmCl 3 -soaked EMTS-soaked Data collection Space
SUPPLEMENTARY INFORMATION doi:10.1038/nature11744 Supplementary Table 1. Crystallographic data collection and refinement statistics. Wild-type Se-Met-BcsA-B SmCl 3 -soaked EMTS-soaked Data collection Space
Guided Reading Activities
 Name Period Chapter 4: A Tour of the Cell Guided Reading Activities Big Idea: Introduction to the Cell Answer the following questions as you read Modules 4.1 4.4: 1. A(n) uses a beam of light to illuminate
Name Period Chapter 4: A Tour of the Cell Guided Reading Activities Big Idea: Introduction to the Cell Answer the following questions as you read Modules 4.1 4.4: 1. A(n) uses a beam of light to illuminate
Some Optimization and Statistical Learning Problems in Structural Biology
 Some Optimization and Statistical Learning Problems in Structural Biology Amit Singer Princeton University, Department of Mathematics and PACM January 8, 2013 Amit Singer (Princeton University) January
Some Optimization and Statistical Learning Problems in Structural Biology Amit Singer Princeton University, Department of Mathematics and PACM January 8, 2013 Amit Singer (Princeton University) January
X RAY DIFFRACTION STUDIES OF MUSCLE AND THE CROSSBRIDGE CYCLE
 X RAY DIFFRACTION STUDIES OF MUSCLE AND THE CROSSBRIDGE CYCLE By JOHN M. SQUIRE* AND CARLO KNUPP { *Biological Structure & Function Section, Biomedical Sciences Division, Imperial College Faculty of Medicine,
X RAY DIFFRACTION STUDIES OF MUSCLE AND THE CROSSBRIDGE CYCLE By JOHN M. SQUIRE* AND CARLO KNUPP { *Biological Structure & Function Section, Biomedical Sciences Division, Imperial College Faculty of Medicine,
Diphthamide biosynthesis requires a radical iron-sulfur enzyme. Pennsylvania State University, University Park, Pennsylvania 16802, USA
 Diphthamide biosynthesis requires a radical iron-sulfur enzyme Yang Zhang, 1,4 Xuling Zhu, 1,4 Andrew T. Torelli, 1 Michael Lee, 2 Boris Dzikovski, 1 Rachel Koralewski, 1 Eileen Wang, 1 Jack Freed, 1 Carsten
Diphthamide biosynthesis requires a radical iron-sulfur enzyme Yang Zhang, 1,4 Xuling Zhu, 1,4 Andrew T. Torelli, 1 Michael Lee, 2 Boris Dzikovski, 1 Rachel Koralewski, 1 Eileen Wang, 1 Jack Freed, 1 Carsten
Supplemental Information for: Mechanical properties of a complete microtubule revealed through molecular dynamics simulation
 Supplemental Information for: Mechanical properties of a complete microtubule revealed through molecular dynamics simulation avid Wells and Aleksei Aksimentiev Water placement and histidine protonation
Supplemental Information for: Mechanical properties of a complete microtubule revealed through molecular dynamics simulation avid Wells and Aleksei Aksimentiev Water placement and histidine protonation
What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil
 Review R45 What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil Chaperonins form large, cage-like structures that act as protein folding machines. Recent developments in
Review R45 What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil Chaperonins form large, cage-like structures that act as protein folding machines. Recent developments in
From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available
 From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available Scripps EM course, November 14, 2007 What aspects of contemporary
From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available Scripps EM course, November 14, 2007 What aspects of contemporary
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
SUPPLEMENTARY INFORMATION
 Dph2 SeMet (iron-free) # Dph2 (iron-free) Dph2-[4Fe-4S] Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 58.26, 82.08, 160.42 58.74, 81.87, 160.01 55.70, 80.53,
Dph2 SeMet (iron-free) # Dph2 (iron-free) Dph2-[4Fe-4S] Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 58.26, 82.08, 160.42 58.74, 81.87, 160.01 55.70, 80.53,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Tools for Cryo-EM Map Fitting. Paul Emsley MRC Laboratory of Molecular Biology
 Tools for Cryo-EM Map Fitting Paul Emsley MRC Laboratory of Molecular Biology April 2017 Cryo-EM model-building typically need to move more atoms that one does for crystallography the maps are lower resolution
Tools for Cryo-EM Map Fitting Paul Emsley MRC Laboratory of Molecular Biology April 2017 Cryo-EM model-building typically need to move more atoms that one does for crystallography the maps are lower resolution
Table 1. Crystallographic data collection, phasing and refinement statistics. Native Hg soaked Mn soaked 1 Mn soaked 2
 Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Table 1. Crystallographic data collection, phasing and refinement statistics Native Hg soaked Mn soaked 1 Mn soaked 2 Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell
Scandium and Yttrium Metallocene Borohydride Complexes: Comparisons of (BH 4 ) 1 vs (BPh 4 ) 1 Coordination and Reactivity
 Scandium and Yttrium Metallocene Borohydride Complexes: Comparisons of (BH 4 ) 1 vs (BPh 4 ) 1 Coordination and Reactivity Selvan Demir, Nathan A. Siladke, Joseph W. Ziller, and William J. Evans * Department
Scandium and Yttrium Metallocene Borohydride Complexes: Comparisons of (BH 4 ) 1 vs (BPh 4 ) 1 Coordination and Reactivity Selvan Demir, Nathan A. Siladke, Joseph W. Ziller, and William J. Evans * Department
A Primer in X-ray Crystallography for Redox Biologists. Mark Wilson Karolinska Institute June 3 rd, 2014
 A Primer in X-ray Crystallography for Redox Biologists Mark Wilson Karolinska Institute June 3 rd, 2014 X-ray Crystallography Basics Optimistic workflow for crystallography Experiment Schematic Fourier
A Primer in X-ray Crystallography for Redox Biologists Mark Wilson Karolinska Institute June 3 rd, 2014 X-ray Crystallography Basics Optimistic workflow for crystallography Experiment Schematic Fourier
Crystal lattice Real Space. Reflections Reciprocal Space. I. Solving Phases II. Model Building for CHEM 645. Purified Protein. Build model.
 I. Solving Phases II. Model Building for CHEM 645 Purified Protein Solve Phase Build model and refine Crystal lattice Real Space Reflections Reciprocal Space ρ (x, y, z) pronounced rho F hkl 2 I F (h,
I. Solving Phases II. Model Building for CHEM 645 Purified Protein Solve Phase Build model and refine Crystal lattice Real Space Reflections Reciprocal Space ρ (x, y, z) pronounced rho F hkl 2 I F (h,
Biological Opportunities with Solution Scattering. Brian R. Crane
 Biological Opportunities with Solution Scattering XDL 2011 Brian R. Crane Cornell University, Ithaca NY bc69@cornell.edu Bacterial Transmembrane Receptors Histidine kinases, adenylyl cyclases, methyl accepting
Biological Opportunities with Solution Scattering XDL 2011 Brian R. Crane Cornell University, Ithaca NY bc69@cornell.edu Bacterial Transmembrane Receptors Histidine kinases, adenylyl cyclases, methyl accepting
Chapter 4: Cells: The Working Units of Life
 Name Period Chapter 4: Cells: The Working Units of Life 1. What are the three critical components of the cell theory? 2. What are the two important conceptual implications of the cell theory? 3. Which
Name Period Chapter 4: Cells: The Working Units of Life 1. What are the three critical components of the cell theory? 2. What are the two important conceptual implications of the cell theory? 3. Which
Full wwpdb X-ray Structure Validation Report i
 Full wwpdb X-ray Structure Validation Report i Jan 17, 2019 09:42 AM EST PDB ID : 6D3Z Title : Protease SFTI complex Authors : Law, R.H.P.; Wu, G. Deposited on : 2018-04-17 Resolution : 2.00 Å(reported)
Full wwpdb X-ray Structure Validation Report i Jan 17, 2019 09:42 AM EST PDB ID : 6D3Z Title : Protease SFTI complex Authors : Law, R.H.P.; Wu, G. Deposited on : 2018-04-17 Resolution : 2.00 Å(reported)
Esser et al. Crystal Structures of R. sphaeroides bc 1
 Esser et al. Crystal Structures of R. sphaeroides bc Supplementary Information Trivariate Gaussian Probability Analysis The superposition of six structures results in sextets of 3D coordinates for every
Esser et al. Crystal Structures of R. sphaeroides bc Supplementary Information Trivariate Gaussian Probability Analysis The superposition of six structures results in sextets of 3D coordinates for every
Electron microscopy in molecular cell biology II
 Electron microscopy in molecular cell biology II Cryo-EM and image processing Werner Kühlbrandt Max Planck Institute of Biophysics Sample preparation for cryo-em Preparation laboratory Specimen preparation
Electron microscopy in molecular cell biology II Cryo-EM and image processing Werner Kühlbrandt Max Planck Institute of Biophysics Sample preparation for cryo-em Preparation laboratory Specimen preparation
CAP 5510 Lecture 3 Protein Structures
 CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/5/e1603193/dc1 Supplementary Materials for Molecular surgery on a 23-gold-atom nanoparticle Qi Li, Tian-Yi Luo, Michael G. Taylor, Shuxin Wang, Xiaofan Zhu, Yongbo
advances.sciencemag.org/cgi/content/full/3/5/e1603193/dc1 Supplementary Materials for Molecular surgery on a 23-gold-atom nanoparticle Qi Li, Tian-Yi Luo, Michael G. Taylor, Shuxin Wang, Xiaofan Zhu, Yongbo
Biology Tutorial. Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan
 Biology Tutorial Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan Viruses A T4 bacteriophage injecting DNA into a cell. Influenza A virus Electron micrograph of HIV. Cone-shaped cores are
Biology Tutorial Aarti Balasubramani Anusha Bharadwaj Massa Shoura Stefan Giovan Viruses A T4 bacteriophage injecting DNA into a cell. Influenza A virus Electron micrograph of HIV. Cone-shaped cores are
SUPPLEMENTARY INFORMATION. doi: /nature07461
 Figure S1 Electrophysiology. a ph-activation of. Two-electrode voltage clamp recordings of Xenopus oocytes expressing in comparison to waterinjected oocytes. Currents were recorded at 40 mv. The ph of
Figure S1 Electrophysiology. a ph-activation of. Two-electrode voltage clamp recordings of Xenopus oocytes expressing in comparison to waterinjected oocytes. Currents were recorded at 40 mv. The ph of
Molecular Motors. Structural and Mechanistic Overview! Kimberly Nguyen - December 6, 2013! MOLECULAR MOTORS - KIMBERLY NGUYEN
 Molecular Motors Structural and Mechanistic Overview!! Kimberly Nguyen - December 6, 2013!! 1 Molecular Motors: A Structure and Mechanism Overview! Introduction! Molecular motors are fundamental agents
Molecular Motors Structural and Mechanistic Overview!! Kimberly Nguyen - December 6, 2013!! 1 Molecular Motors: A Structure and Mechanism Overview! Introduction! Molecular motors are fundamental agents
Bioinformatics and Biology Insights
 Bioinformatics and Biology Insights Original Research Open Access Full open access to this and thousands of other papers at http://www.la-press.com. Integrative Structural Modelling of the Cardiac Thin
Bioinformatics and Biology Insights Original Research Open Access Full open access to this and thousands of other papers at http://www.la-press.com. Integrative Structural Modelling of the Cardiac Thin
O.k., Now Starts the Good Stuff (Part II) Eukaryotic Cell Structure and Function
 O.k., Now Starts the Good Stuff (Part II) Eukaryotic Cell Structure and Function Eukaryotic Cells These cells have membrane-bound structures called organelles. Cell processes occur in these organelles.
O.k., Now Starts the Good Stuff (Part II) Eukaryotic Cell Structure and Function Eukaryotic Cells These cells have membrane-bound structures called organelles. Cell processes occur in these organelles.
Structure of the SPRY domain of human DDX1 helicase, a putative interaction platform within a DEAD-box protein
 Supporting information Volume 71 (2015) Supporting information for article: Structure of the SPRY domain of human DDX1 helicase, a putative interaction platform within a DEAD-box protein Julian Kellner
Supporting information Volume 71 (2015) Supporting information for article: Structure of the SPRY domain of human DDX1 helicase, a putative interaction platform within a DEAD-box protein Julian Kellner
TER 26. Preview for 2/6/02 Dr. Kopeny. Bacteria and Archaea: The Prokaryotic Domains. Nitrogen cycle
 Preview for 2/6/02 Dr. Kopeny Bacteria and Archaea: The Prokaryotic Domains TER 26 Nitrogen cycle Mycobacterium tuberculosis Color-enhanced images shows rod-shaped bacterium responsible for tuberculosis
Preview for 2/6/02 Dr. Kopeny Bacteria and Archaea: The Prokaryotic Domains TER 26 Nitrogen cycle Mycobacterium tuberculosis Color-enhanced images shows rod-shaped bacterium responsible for tuberculosis
