Electron microscopy in molecular cell biology II
|
|
|
- Marsha Jacobs
- 5 years ago
- Views:
Transcription
1 Electron microscopy in molecular cell biology II Cryo-EM and image processing Werner Kühlbrandt Max Planck Institute of Biophysics
2 Sample preparation for cryo-em
3 Preparation laboratory
4 Specimen preparation Carbon coated copper grid
5 Rapid freezing in liquid ethane
6 Rapid freezing in liquid ethane
7 Mounting under liquid nitrogen
8 Grid in EM column
9 Grid in EM column EM grid
10 EM grid in a light microscope 0.2 mm
11 Holey carbon film 3 mm 2 µm 60x 660x 6660x
12 Ice thickness holey C film z = 100 nm
13 Ice thickness holey C film z = 100 nm
14 Ice thickness holey C film z = 2 nm Thin ice squeezes out particles
15 Ice thickness holey C film z = 30 nm
16 Ice thickness holey C film z = 30 nm f = 22.5 nm
17 Ice thickness holey C film z = 30 nm f = 22.5 nm Amphipols may help to achieve optimal ice thickness with membrane proteins
18 A cryo-specimen imaged in the EM electrons camera
19 Cryo-EM film image of Frh complex Janet Vonck, Deryck Mills
20 Fourier transforms for image processing
21 Any physical object can be thought of as the sum of a Fourier series of cosine waves
22 Any physical object can be thought of as the sum of a Fourier series of cosine waves in one dimension spectra, sounds, signals of any kind
23 Any physical object can be thought of as the sum of a Fourier series of cosine waves in one dimension spectra, sounds, signals of any kind in two dimensions electron micrographs, any image
24 Any physical object can be thought of as the sum of a Fourier series of cosine waves in one dimension spectra, sounds, signals of any kind in two dimensions electron micrographs, any image in three dimensions! crystal structures, any 3D object
25 The 3D volume of any object can be reconstructed from its 2D projections
26 The 3D volume of any object can be reconstructed from its 2D projections in Fourier space! 2D crystals, single particles
27 The 3D volume of any object can be reconstructed from its 2D projections in Fourier space! 2D crystals, single particles in real space e.g. electron tomography
28 Why Fourier transforms?
29 Why Fourier transforms? Developed by French mathematician Joseph Fourier ( ) as a method for transforming mathematical functions
30 Why Fourier transforms? Developed by French mathematician Joseph Fourier ( ) as a method for transforming mathematical functions The FT transforms real space into reciprocal space and vice versa
31 Why Fourier transforms? Developed by French mathematician Joseph Fourier ( ) as a method for transforming mathematical functions The FT transforms real space into reciprocal space and vice versa FTs are perfect tools for crystallography and image processing (diffraction, resolution, signal/noise analysis, alignment etc.)
32 Why Fourier transforms? Developed by French mathematician Joseph Fourier ( ) as a method for transforming mathematical functions The FT transforms real space into reciprocal space and vice versa FTs are perfect tools for crystallography and image processing (diffraction, resolution, signal/noise analysis, alignment etc.) The FT of a crystal is its diffraction pattern
33 Why Fourier transforms? Developed by French mathematician Joseph Fourier ( ) as a method for transforming mathematical functions The FT transforms real space into reciprocal space and vice versa FTs are perfect tools for crystallography and image processing (diffraction, resolution, signal/noise analysis, alignment etc.) The FT of a crystal is its diffraction pattern The focal plane of a lens contains the FT of the imaged object
34 Why Fourier transforms? Developed by French mathematician Joseph Fourier ( ) as a method for transforming mathematical functions The FT transforms real space into reciprocal space and vice versa FTs are perfect tools for crystallography and image processing (diffraction, resolution, signal/noise analysis, alignment etc.) The FT of a crystal is its diffraction pattern The focal plane of a lens contains the FT of the imaged object Computers are very good at calculating FTs
35 Why Fourier transforms? Developed by French mathematician Joseph Fourier ( ) as a method for transforming mathematical functions The FT transforms real space into reciprocal space and vice versa FTs are perfect tools for crystallography and image processing (diffraction, resolution, signal/noise analysis, alignment etc.) The FT of a crystal is its diffraction pattern The focal plane of a lens contains the FT of the imaged object Computers are very good at calculating FTs The calculated FT of an object contains all amplitudes and phases of its structure factors
36 Simple one-dimensional Fourier transform periodic structure unit cell x Fourier transform h
37 Simple one-dimensional Fourier transform periodic structure unit cell x Fourier transform h
38 Simple one-dimensional Fourier transform periodic structure unit cell x Fourier transform h
39 Simple one-dimensional Fourier transform periodic structure unit cell x Fourier transform h
40 Simple one-dimensional Fourier transform periodic structure unit cell F 1 α 1 = 0 h = 1 x Fourier transform h
41 Simple one-dimensional Fourier transform periodic structure unit cell F 2 α 2 h = 2 F 1 α 1 = 0 h = 1 x Fourier transform h
42 Simple one-dimensional Fourier transform periodic structure unit cell F 6 α 6 h = 6 F 2 α 2 h = 2 F 1 α 1 = 0 h = 1 x Fourier transform h
43 Simple one-dimensional Fourier transform periodic structure unit cell F 6 α 6 h = 6 F 2 α 2 h = 2 F 1 α 1 = 0 h = 1 x Fourier transform h
44 Simple one-dimensional Fourier transform periodic structure unit cell F 6 α 6 h = 6 F 2 α 2 h = 2 F 1 α 1 = 0 h = 1 x Fourier transform h
45 Simple one-dimensional Fourier transform periodic structure unit cell F 6 α 6 h = 6 F 2 α 2 h = 2 F 1 α 1 = 0 h = 1 x Fourier transform h
46 2D lattice transforms The Fourier transform of a lattice is another lattice with reciprocal dimensions FFT real lattice reciprocal lattice
47 Fourier transforms of lattices and molecules lattice molecule molecular crystal real space convolution reciprocal space FFT lattice transform x FFT molecular transform multiplication FFT molecular transform seen through holes of lattice transform
48 Fourier transforms of lattices and molecules (convolution of two objects is equivalent to multiplication of their Fourier transform) lattice molecule molecular crystal real space convolution reciprocal space FFT lattice transform x FFT molecular transform multiplication FFT molecular transform seen through holes of lattice transform
49 Same molecule on different lattices no lattice real lattice FFT FFT FFT FFT FFT molecular transform reciprocal lattice
50 Same molecule on different lattices no lattice real lattice FFT FFT FFT FFT FFT molecular transform reciprocal lattice
51 Resolution I: crystals real space FFT FFT -1 FFT -1 reciprocal space cutoff cutoff
52 Resolution II: single particles real space FFT FFT -1 FFT -1 reciprocal space cutoff cutoff molecular transform
53 Amplitudes and phases I Helen Saibil, Birkbeck College London
54 real space Amplitudes and phases II
55 Amplitudes and phases II real space FFT
56 Amplitudes and phases II real space FFT reciprocal space (amplitudes and phases)
57 Amplitudes and phases II real space FFT reciprocal space (amplitudes and phases)
58 Amplitudes and phases II real space FFT FFT reciprocal space (amplitudes and phases)
59 cat phases duck amplitudes Amplitudes and phases III
60 Amplitudes and phases III cat phases duck amplitudes duck phases cat amplitudes
61 Amplitudes and phases III cat phases duck amplitudes FFT -1 duck phases cat amplitudes
62 Amplitudes and phases III cat phases duck amplitudes FFT -1 duck phases cat amplitudes FFT -1
63 Amplitudes and phases III cat phases duck amplitudes FFT -1 duck phases cat amplitudes FFT -1 Phases are more important than amplitudes!
64 Single-particle image processing
65 Noise-free field of single particles Helen Saibil, Birkbeck College London
66 1:1 signal to noise Helen Saibil, Birkbeck College London
67 1:1 signal to noise Helen Saibil, Birkbeck College London
68 3D maps from images Helen Saibil, Birkbeck College London
69 3D maps from images next round Helen Saibil, Birkbeck College London
70 Starting model from conical tilt reconstruction
71 3D reconstruction from 2D projections Helen Saibil, Birkbeck College London
72 Particle selection
73 Particle selection
74 Alignment, classification, class averages raw images aligned images class averages
75 3D reconstruction from class averages
76 3D reconstruction from class averages
77 Reference bias 10,000 images of random noise Janet Vonck
78 Reference bias 10,000 images of random noise noise images aligned to a reference image Janet Vonck
79 Reference bias 10,000 images of random noise Janet Vonck aligned noise images averaged
80 Reference bias 10,000 images of random noise Janet Vonck aligned noise images averaged
81 Reference bias 10,000 images of random noise Janet Vonck aligned noise images averaged
82 Reference bias 10,000 images of random noise Janet Vonck aligned noise images averaged
83 Reference bias 10,000 images of random noise Janet Vonck aligned noise images averaged
84 Gold standard Fourier shell correlation avoids reference bias compare Fourier transforms in resolution shells particle set 1 model 1 3D FT of model 1 Fourier shell correlation (FSC) independent particle set 2 model 2 3D FT of model 2
85 Resolution from FSC plot FSC = 0.5 resolution threshold for gold standard comparison FSC = (FOM = 0.5) (Rosenthal & Henderson JMB 2003) 4.1 Å 3.4 Å
86 Film vs. direct detector Fourier Shell Correlation Film, particles: 5.55 Å Falcon-II, particles: 3.95 Å FSC Resolu'on (Å - 1 ) Allegretti et al, elife 2014
87 Frh at 5.5 Å by cryo-em Mills et al, elife 2013
88 3.4 Å map of Frh complex Allegretti et al, elife 2014
89 3.4 Å map of Frh complex Allegretti et al, elife 2014
90 3.4 Å map of Frh complex Allegretti et al, elife 2014
91 3.4 Å map of Frh complex Allegretti et al, elife 2014
92 Science, 28 March 2014
93 Science, 28 March 2014
94 Three flavours of cryo-em Single-particle cryo-em! complexes > 200 kda up to 3 Å resolution Electron crystallography kda membrane proteins up to 3 Å resolution Electron cryo-tomography whole cells, organelles, membranes up to 8 Å resolution (by subtomogram averaging)
95 macromolecular complexes 2D crystals membranes, organelles, cells Frh ChR2 mito- mitochondrion single-particle processing electron crystallography electron tomography
96 macromolecular complexes 2D crystals membranes, organelles, cells Frh ChR2 mito- mitochondrion single-particle processing electron crystallography electron tomography
97 Electron crystallography
98 Purple membrane and bacteriorhodopsin Freeze-fractured H. halobium purple membranes Blaurock & Stoeckenius, Nature 1971 Henderson & Unwin, Nature 1975
99 2D crystals of LHC-II Kühlbrandt, Nature 1984
100 2D crystals of LHC-II Kühlbrandt et al, JCB 1983
101 Large 2D crystal of LHC-II
102 Electron diffraction patter of LHC-II 3.2 Å
103 Cryo image of LHC-II 2D crystal
104 Fourier transform of 2D crystal 3.28 Å
105 Reciprocal lattice of 2D crystal
106 Amplitudes and phases along lattice lines
107 3.4 Å EM structure of LHC-II Kühlbrandt et al, Nature 1994
108 3.4 Å EM structure of LHC-II Kühlbrandt et al, Nature 1994
109 Electron tomography
110 Two steps in electron tomography Achilleas Frangakis, Frankfurt University
111 Two steps in electron tomography 1. Imaging: convert 3D object into a set of 2D projections Achilleas Frangakis, Frankfurt University
112 Two steps in electron tomography 1. Imaging: convert 3D object into a set of 2D projections Achilleas Frangakis, Frankfurt University
113 Two steps in electron tomography 1. Imaging: convert 3D object into a set of 2D projections 2. reconstruction: generate 3D volume from set of 2D projections Achilleas Frangakis, Frankfurt University
114 Two steps in electron tomography 1. Imaging: convert 3D object into a set of 2D projections 2. reconstruction: generate 3D volume from set of 2D projections Achilleas Frangakis, Frankfurt University
115 Tomographic tilt series of mitochondrion 100 nm
116 Tomographic volume of mitochondrion 100 nm
117 Tomogram of a mouse heart mitochondrion outer membrane intermembrane space inner boundary membrane matrix cristae cristae junctions Tobias Brandt
118 Tomogram of a mouse heart mitochondrion outer membrane intermembrane space inner boundary membrane matrix cristae cristae junctions Tobias Brandt
119 Tomogram of a mouse heart mitochondrion outer membrane intermembrane space inner boundary membrane matrix cristae cristae junctions cristae junctions Tobias Brandt
120 Cryo-ET of Podospora mitochondrion ATP synthase dimers inner membrane outer membrane Davies et al, PNAS 2011
121 Sub-tomogram average of ATP synthase dimer Davies et al, PNAS 2012
122 Fitted X-ray structures F1 head peripheral stalk central stalk rotor ring Davies et al, PNAS 2012
There and back again A short trip to Fourier Space. Janet Vonck 23 April 2014
 There and back again A short trip to Fourier Space Janet Vonck 23 April 2014 Where can I find a Fourier Transform? Fourier Transforms are ubiquitous in structural biology: X-ray diffraction Spectroscopy
There and back again A short trip to Fourier Space Janet Vonck 23 April 2014 Where can I find a Fourier Transform? Fourier Transforms are ubiquitous in structural biology: X-ray diffraction Spectroscopy
Structural biology of mitochondria. Werner Kühlbrandt Max Planck Institute of Biophysics Frankfurt, Germany
 Structural biology of mitochondria Werner Kühlbrandt Max Planck Institute of Biophysics Frankfurt, Germany The mitochondrion Powerhouse of the eukaryotic cell Produces almost all ATP to drive cellular
Structural biology of mitochondria Werner Kühlbrandt Max Planck Institute of Biophysics Frankfurt, Germany The mitochondrion Powerhouse of the eukaryotic cell Produces almost all ATP to drive cellular
sin" =1.22 # D "l =1.22 f# D I: In introduction to molecular electron microscopy - Imaging macromolecular assemblies
 I: In introduction to molecular electron microscopy - Imaging macromolecular assemblies Yifan Cheng Department of Biochemistry & Biophysics office: GH-S472D; email: ycheng@ucsf.edu 2/20/2015 - Introduction
I: In introduction to molecular electron microscopy - Imaging macromolecular assemblies Yifan Cheng Department of Biochemistry & Biophysics office: GH-S472D; email: ycheng@ucsf.edu 2/20/2015 - Introduction
Molecular electron microscopy
 Molecular electron microscopy - Imaging macromolecular assemblies Yifan Cheng Department of Biochemistry & Biophysics office: GH-S427B; email: ycheng@ucsf.edu 2/22/2013 - Introduction of Molecular Microscopy:
Molecular electron microscopy - Imaging macromolecular assemblies Yifan Cheng Department of Biochemistry & Biophysics office: GH-S427B; email: ycheng@ucsf.edu 2/22/2013 - Introduction of Molecular Microscopy:
Lecture CIMST Winter School. 1. What can you see by TEM?
 Lecture CIMST Winter School Cryo-electron microscopy and tomography of biological macromolecules 20.1.2011 9:00-9:45 in Y03G91 Dr. Takashi Ishikawa OFLB/005 Tel: 056 310 4217 e-mail: takashi.ishikawa@psi.ch
Lecture CIMST Winter School Cryo-electron microscopy and tomography of biological macromolecules 20.1.2011 9:00-9:45 in Y03G91 Dr. Takashi Ishikawa OFLB/005 Tel: 056 310 4217 e-mail: takashi.ishikawa@psi.ch
History of 3D Electron Microscopy and Helical Reconstruction
 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N History of 3D Electron Microscopy and Helical Reconstruction For students of HI
T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N History of 3D Electron Microscopy and Helical Reconstruction For students of HI
Electron microscopy in molecular cell biology I
 Electron microscopy in molecular cell biology I Electron optics and image formation Werner Kühlbrandt Max Planck Institute of Biophysics chemistry biology Objects of interest Galaxy 10 6 light years 10
Electron microscopy in molecular cell biology I Electron optics and image formation Werner Kühlbrandt Max Planck Institute of Biophysics chemistry biology Objects of interest Galaxy 10 6 light years 10
Part 1 X-ray Crystallography
 Part 1 X-ray Crystallography What happens to electron when it is hit by x-rays? 1. The electron starts vibrating with the same frequency as the x-ray beam 2. As a result, secondary beams will be scattered
Part 1 X-ray Crystallography What happens to electron when it is hit by x-rays? 1. The electron starts vibrating with the same frequency as the x-ray beam 2. As a result, secondary beams will be scattered
From electron crystallography to single particle cryoem
 From electron crystallography to single particle cryoem Nobel Lectures in Chemistry 8 th December 2017 Richard Henderson From Baker & Henderson (2001) Int.Tab.Cryst.Vol.F, on-line (2006), revised (2011)
From electron crystallography to single particle cryoem Nobel Lectures in Chemistry 8 th December 2017 Richard Henderson From Baker & Henderson (2001) Int.Tab.Cryst.Vol.F, on-line (2006), revised (2011)
HIGH RESOLUTION ELECTRON MICROSCOPY
 HIGH RESOLUTION ELECTRON MICROSCOPY BC530 Fall Quarter 2014 Wim G. J. Hol http://www.bmsc.washington.edu/wimhol/ Structure Determination by ELECTRON MICROSCOPY STRENGTHS: - Small amounts of material usually
HIGH RESOLUTION ELECTRON MICROSCOPY BC530 Fall Quarter 2014 Wim G. J. Hol http://www.bmsc.washington.edu/wimhol/ Structure Determination by ELECTRON MICROSCOPY STRENGTHS: - Small amounts of material usually
Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy
 Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy Amit Singer Princeton University, Department of Mathematics and PACM July 23, 2014 Amit Singer (Princeton
Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy Amit Singer Princeton University, Department of Mathematics and PACM July 23, 2014 Amit Singer (Princeton
Three-Dimensional Electron Microscopy of Macromolecular Assemblies
 Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil
 Review R45 What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil Chaperonins form large, cage-like structures that act as protein folding machines. Recent developments in
Review R45 What can electron microscopy tell us about chaperoned protein folding? Helen R Saibil Chaperonins form large, cage-like structures that act as protein folding machines. Recent developments in
GBS765 Electron microscopy
 GBS765 Electron microscopy Lecture 1 Waves and Fourier transforms 10/14/14 9:05 AM Some fundamental concepts: Periodicity! If there is some a, for a function f(x), such that f(x) = f(x + na) then function
GBS765 Electron microscopy Lecture 1 Waves and Fourier transforms 10/14/14 9:05 AM Some fundamental concepts: Periodicity! If there is some a, for a function f(x), such that f(x) = f(x + na) then function
ParM filament images were extracted and from the electron micrographs and
 Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Laboratory-Based Cryogenic Soft X-ray Tomography and Correlative Microscopy: 3D Visualization Inside the Cell
 Laboratory-Based Cryogenic Soft X-ray Tomography and Correlative Microscopy: 3D Visualization Inside the Cell David Carlson, Jeff Gelb, Vadim Palshin and James Evans Pacific Northwest National Laboratory
Laboratory-Based Cryogenic Soft X-ray Tomography and Correlative Microscopy: 3D Visualization Inside the Cell David Carlson, Jeff Gelb, Vadim Palshin and James Evans Pacific Northwest National Laboratory
Protein Structure Determination 9/25/2007
 One-dimensional NMR spectra Ethanol Cellulase (36 a.a.) Branden & Tooze, Fig. 18.16 1D and 2D NMR spectra of inhibitor K (57 a.a.) K. Wuthrich, NMR of Proteins and Nucleic Acids. (Wiley, 1986.) p. 54-55.
One-dimensional NMR spectra Ethanol Cellulase (36 a.a.) Branden & Tooze, Fig. 18.16 1D and 2D NMR spectra of inhibitor K (57 a.a.) K. Wuthrich, NMR of Proteins and Nucleic Acids. (Wiley, 1986.) p. 54-55.
General theory of diffraction
 General theory of diffraction X-rays scatter off the charge density (r), neutrons scatter off the spin density. Coherent scattering (diffraction) creates the Fourier transform of (r) from real to reciprocal
General theory of diffraction X-rays scatter off the charge density (r), neutrons scatter off the spin density. Coherent scattering (diffraction) creates the Fourier transform of (r) from real to reciprocal
CIMST Summer School Cryo-electron microscopy for structural biology 11 September, 2014 Dr. Takashi Ishikawa
 CIMST Summer School Cryo-electron microscopy for structural biology 11 September, 2014 Dr. Takashi Ishikawa Tel: 056 310 4217 e-mail: takashi.ishikawa@psi.ch Lab webpage: http://www.psi.ch/lbr/takashi-ishikawa
CIMST Summer School Cryo-electron microscopy for structural biology 11 September, 2014 Dr. Takashi Ishikawa Tel: 056 310 4217 e-mail: takashi.ishikawa@psi.ch Lab webpage: http://www.psi.ch/lbr/takashi-ishikawa
Transmission Electron Microscopy
 L. Reimer H. Kohl Transmission Electron Microscopy Physics of Image Formation Fifth Edition el Springer Contents 1 Introduction... 1 1.1 Transmission Electron Microscopy... 1 1.1.1 Conventional Transmission
L. Reimer H. Kohl Transmission Electron Microscopy Physics of Image Formation Fifth Edition el Springer Contents 1 Introduction... 1 1.1 Transmission Electron Microscopy... 1 1.1.1 Conventional Transmission
Electron cryo- microscopy at Yale
 Electron cryo- microscopy at Yale A discussion with the Basic Science Strategic Planning Commi
Electron cryo- microscopy at Yale A discussion with the Basic Science Strategic Planning Commi
Fourier Series. Combination of Waves: Any PERIODIC function f(t) can be written: How to calculate the coefficients?
 Fourier Series Combination of Waves: Any PERIODIC function f(t) can be written: How to calculate the coefficients? What is the Fourier Transform It is the representation of an arbitrary NON-periodic signal
Fourier Series Combination of Waves: Any PERIODIC function f(t) can be written: How to calculate the coefficients? What is the Fourier Transform It is the representation of an arbitrary NON-periodic signal
New algoritms for electron diffraction of 3D protein crystals. JP Abrahams, D Georgieva, L Jiang, I Sikhuralidze, NS Pannu
 New algoritms for electron diffraction of 3D protein crystals JP Abrahams, D Georgieva, L Jiang, I Sikhuralidze, NS Pannu Why new algorithms? New research questions New experimental techniques Better insight
New algoritms for electron diffraction of 3D protein crystals JP Abrahams, D Georgieva, L Jiang, I Sikhuralidze, NS Pannu Why new algorithms? New research questions New experimental techniques Better insight
Opportunity and Challenge: the new era of cryo-em. Ning Gao School of Life Sciences Tsinghua University
 Opportunity and Challenge: the new era of cryo-em Ning Gao School of Life Sciences Tsinghua University 2015-10-20 Education and Research Experience Principal Investigator 2008.11-present Postdoc, Department
Opportunity and Challenge: the new era of cryo-em Ning Gao School of Life Sciences Tsinghua University 2015-10-20 Education and Research Experience Principal Investigator 2008.11-present Postdoc, Department
Image Assessment San Diego, November 2005
 Image Assessment San Diego, November 005 Pawel A. Penczek The University of Texas Houston Medical School Department of Biochemistry and Molecular Biology 6431 Fannin, MSB6.18, Houston, TX 77030, USA phone:
Image Assessment San Diego, November 005 Pawel A. Penczek The University of Texas Houston Medical School Department of Biochemistry and Molecular Biology 6431 Fannin, MSB6.18, Houston, TX 77030, USA phone:
CHEM-E5225 :Electron Microscopy Imaging
 CHEM-E5225 :Electron Microscopy Imaging 2016.10 Yanling Ge Outline Planar Defects Image strain field WBDF microscopy HRTEM information theory Discuss of question homework? Planar Defects - Internal Interface
CHEM-E5225 :Electron Microscopy Imaging 2016.10 Yanling Ge Outline Planar Defects Image strain field WBDF microscopy HRTEM information theory Discuss of question homework? Planar Defects - Internal Interface
PSD '17 -- Xray Lecture 5, 6. Patterson Space, Molecular Replacement and Heavy Atom Isomorphous Replacement
 PSD '17 -- Xray Lecture 5, 6 Patterson Space, Molecular Replacement and Heavy Atom Isomorphous Replacement The Phase Problem We can t measure the phases! X-ray detectors (film, photomultiplier tubes, CCDs,
PSD '17 -- Xray Lecture 5, 6 Patterson Space, Molecular Replacement and Heavy Atom Isomorphous Replacement The Phase Problem We can t measure the phases! X-ray detectors (film, photomultiplier tubes, CCDs,
How To Evaluate Electron Crystallographic Data
 How To Evaluate Electron Crystallographic Data Vinzenz Unger, Dept of Molecular Biosciences, Northwestern University Chemistry of Life Sciences Institute, Northwestern University Unstained = Resolution
How To Evaluate Electron Crystallographic Data Vinzenz Unger, Dept of Molecular Biosciences, Northwestern University Chemistry of Life Sciences Institute, Northwestern University Unstained = Resolution
Fourier Syntheses, Analyses, and Transforms
 Fourier Syntheses, Analyses, and Transforms http://homepages.utoledo.edu/clind/ The electron density The electron density in a crystal can be described as a periodic function - same contents in each unit
Fourier Syntheses, Analyses, and Transforms http://homepages.utoledo.edu/clind/ The electron density The electron density in a crystal can be described as a periodic function - same contents in each unit
What can we see with a transmission electron microscope?
 What can we see with a transmission electron microscope? β-galactosidase Subramaniam Lab o.lambert@cbmn.u-bordeaux.fr Renafobis 2017 1 Film or camera Renafobis 2017 2 1897 J.J. Thomson Electron discovery
What can we see with a transmission electron microscope? β-galactosidase Subramaniam Lab o.lambert@cbmn.u-bordeaux.fr Renafobis 2017 1 Film or camera Renafobis 2017 2 1897 J.J. Thomson Electron discovery
Application examples of single particle 3D reconstruction. Ning Gao Tsinghua University
 Application examples of single particle 3D reconstruction Ning Gao Tsinghua University ninggao@tsinghua.edu.cn Electron Microscopes First electron microscope constructed by Ernst Ruska in 1930 s (1986
Application examples of single particle 3D reconstruction Ning Gao Tsinghua University ninggao@tsinghua.edu.cn Electron Microscopes First electron microscope constructed by Ernst Ruska in 1930 s (1986
Macromolecular X-ray Crystallography
 Protein Structural Models for CHEM 641 Fall 07 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware Macromolecular X-ray Crystallography Purified Protein X-ray Diffraction Data collection
Protein Structural Models for CHEM 641 Fall 07 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware Macromolecular X-ray Crystallography Purified Protein X-ray Diffraction Data collection
Structure, Dynamics and Function of Membrane Proteins using 4D Electron Microscopy
 Structure, Dynamics and Function of Membrane Proteins using 4D Electron Microscopy Anthony W. P. Fitzpatrick Department of Chemistry, Lensfield Road, Cambridge, CB2 1EW My research aims to understand the
Structure, Dynamics and Function of Membrane Proteins using 4D Electron Microscopy Anthony W. P. Fitzpatrick Department of Chemistry, Lensfield Road, Cambridge, CB2 1EW My research aims to understand the
Chapter 9. Electron mean free path Microscopy principles of SEM, TEM, LEEM
 Chapter 9 Electron mean free path Microscopy principles of SEM, TEM, LEEM 9.1 Electron Mean Free Path 9. Scanning Electron Microscopy (SEM) -SEM design; Secondary electron imaging; Backscattered electron
Chapter 9 Electron mean free path Microscopy principles of SEM, TEM, LEEM 9.1 Electron Mean Free Path 9. Scanning Electron Microscopy (SEM) -SEM design; Secondary electron imaging; Backscattered electron
Protein Crystallography
 Protein Crystallography Part II Tim Grüne Dept. of Structural Chemistry Prof. G. Sheldrick University of Göttingen http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview The Reciprocal Lattice The
Protein Crystallography Part II Tim Grüne Dept. of Structural Chemistry Prof. G. Sheldrick University of Göttingen http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview The Reciprocal Lattice The
ABC s of Electrochemistry series Materials Characterization techniques: SEM and EDS Ana María Valenzuela-Muñiz November 3, 2011
 ABC s of Electrochemistry series Materials Characterization techniques: SEM and EDS Ana María Valenzuela-Muñiz November 3, 2011 CEER, Department of Chemical and Biomolecular Engineering Outline Introduction
ABC s of Electrochemistry series Materials Characterization techniques: SEM and EDS Ana María Valenzuela-Muñiz November 3, 2011 CEER, Department of Chemical and Biomolecular Engineering Outline Introduction
Scattering Lecture. February 24, 2014
 Scattering Lecture February 24, 2014 Structure Determination by Scattering Waves of radiation scattered by different objects interfere to give rise to an observable pattern! The wavelength needs to close
Scattering Lecture February 24, 2014 Structure Determination by Scattering Waves of radiation scattered by different objects interfere to give rise to an observable pattern! The wavelength needs to close
Resolution: maximum limit of diffraction (asymmetric)
 Resolution: maximum limit of diffraction (asymmetric) crystal Y X-ray source 2θ X direct beam tan 2θ = Y X d = resolution 2d sinθ = λ detector 1 Unit Cell: two vectors in plane of image c* Observe: b*
Resolution: maximum limit of diffraction (asymmetric) crystal Y X-ray source 2θ X direct beam tan 2θ = Y X d = resolution 2d sinθ = λ detector 1 Unit Cell: two vectors in plane of image c* Observe: b*
(i.e. what you should be able to answer at end of lecture)
 Today s Announcements 1. Test given back next Wednesday 2. HW assigned next Wednesday. 3. Next Monday 1 st discussion about Individual Projects. Today s take-home lessons (i.e. what you should be able
Today s Announcements 1. Test given back next Wednesday 2. HW assigned next Wednesday. 3. Next Monday 1 st discussion about Individual Projects. Today s take-home lessons (i.e. what you should be able
Protein crystallography. Garry Taylor
 Protein crystallography Garry Taylor X-ray Crystallography - the Basics Grow crystals Collect X-ray data Determine phases Calculate ρ-map Interpret map Refine coordinates Do the biology. Nitrogen at -180
Protein crystallography Garry Taylor X-ray Crystallography - the Basics Grow crystals Collect X-ray data Determine phases Calculate ρ-map Interpret map Refine coordinates Do the biology. Nitrogen at -180
X-Ray Diffraction as a key to the Structure of Materials Interpretation of scattering patterns in real and reciprocal space
 X-Ray Diffraction as a key to the Structure of Materials Interpretation of scattering patterns in real and reciprocal space Tobias U. Schülli, X-ray nanoprobe group ESRF OUTLINE 1 Internal structure of
X-Ray Diffraction as a key to the Structure of Materials Interpretation of scattering patterns in real and reciprocal space Tobias U. Schülli, X-ray nanoprobe group ESRF OUTLINE 1 Internal structure of
Scattering by two Electrons
 Scattering by two Electrons p = -r k in k in p r e 2 q k in /λ θ θ k out /λ S q = r k out p + q = r (k out - k in ) e 1 Phase difference of wave 2 with respect to wave 1: 2π λ (k out - k in ) r= 2π S r
Scattering by two Electrons p = -r k in k in p r e 2 q k in /λ θ θ k out /λ S q = r k out p + q = r (k out - k in ) e 1 Phase difference of wave 2 with respect to wave 1: 2π λ (k out - k in ) r= 2π S r
Part 3 - Image Formation
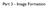 Part 3 - Image Formation Three classes of scattering outcomes Types of electron microscopes Example SEM image: fly nose Example TEM image: muscle Skeletal muscle. Cell and Tissue Ultrastructure Mercer
Part 3 - Image Formation Three classes of scattering outcomes Types of electron microscopes Example SEM image: fly nose Example TEM image: muscle Skeletal muscle. Cell and Tissue Ultrastructure Mercer
Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and
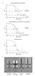 Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and comparisons to other published in situ T3SS structures. a, Resolution estimates after applying Fourier shell correlation
Supplementary Figure 1. Fourier shell correlation curves for sub-tomogram averages and comparisons to other published in situ T3SS structures. a, Resolution estimates after applying Fourier shell correlation
From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available
 From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available Scripps EM course, November 14, 2007 What aspects of contemporary
From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available Scripps EM course, November 14, 2007 What aspects of contemporary
Copyright Mark Brandt, Ph.D A third method, cryogenic electron microscopy has seen increasing use over the past few years.
 Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
2.1 CELL STRUCTURE. The cell is the smallest unit of living organisms that shows the characteristics of life.
 2.1.1 Microscopy The cell is the smallest unit of living organisms that shows the characteristics of life. A general introduction to the microscope. The light microscope All cells are microscopic which
2.1.1 Microscopy The cell is the smallest unit of living organisms that shows the characteristics of life. A general introduction to the microscope. The light microscope All cells are microscopic which
CRYSTALLOGRAPHY AND STORYTELLING WITH DATA. President, Association of Women in Science, Bethesda Chapter STEM Consultant
 CRYSTALLOGRAPHY AND STORYTELLING WITH DATA President, Association of Women in Science, Bethesda Chapter STEM Consultant MY STORY Passion for Science BS Biology Major MS Biotechnology & Project in Bioinformatics
CRYSTALLOGRAPHY AND STORYTELLING WITH DATA President, Association of Women in Science, Bethesda Chapter STEM Consultant MY STORY Passion for Science BS Biology Major MS Biotechnology & Project in Bioinformatics
MODULE 2 : FOUNDATIONS IN BIOLOGY
 OCR A LEVEL BIOLOGY MODULE 2 : FOUNDATIONS IN BIOLOGY REVISION NOTES For 2015 onwards specification Miss T Banda All living things are primarily made from 4 key elements: Carbon (C) Hydrogen (H) Oxygen
OCR A LEVEL BIOLOGY MODULE 2 : FOUNDATIONS IN BIOLOGY REVISION NOTES For 2015 onwards specification Miss T Banda All living things are primarily made from 4 key elements: Carbon (C) Hydrogen (H) Oxygen
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/310/5753/1513/dc1 Supporting Online Material for Structural Roles for Human Translation Factor eif3 in Initiation of Protein Synthesis Bunpote Siridechadilok, Christopher
www.sciencemag.org/cgi/content/full/310/5753/1513/dc1 Supporting Online Material for Structural Roles for Human Translation Factor eif3 in Initiation of Protein Synthesis Bunpote Siridechadilok, Christopher
Schematic representation of relation between disorder and scattering
 Crystal lattice Reciprocal lattice FT Schematic representation of relation between disorder and scattering ρ = Δρ + Occupational disorder Diffuse scattering Bragg scattering ρ = Δρ + Positional
Crystal lattice Reciprocal lattice FT Schematic representation of relation between disorder and scattering ρ = Δρ + Occupational disorder Diffuse scattering Bragg scattering ρ = Δρ + Positional
Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples.
 Supplementary Figure 1 Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples. (a-b) SDS-PAGE analysis of the hexamer and heptamer samples. The eluted hexamer
Supplementary Figure 1 Purification, SDS-PAGE and cryo-em characterization of the MCM hexamer and Cdt1 MCM heptamer samples. (a-b) SDS-PAGE analysis of the hexamer and heptamer samples. The eluted hexamer
MSE 321 Structural Characterization
 Auger Spectroscopy Auger Electron Spectroscopy (AES) Scanning Auger Microscopy (SAM) Incident Electron Ejected Electron Auger Electron Initial State Intermediate State Final State Physical Electronics
Auger Spectroscopy Auger Electron Spectroscopy (AES) Scanning Auger Microscopy (SAM) Incident Electron Ejected Electron Auger Electron Initial State Intermediate State Final State Physical Electronics
Electron Microscopy. SEM = Scanning Electron Microscopy TEM = Transmission Electron Microscopy. E. coli, William E. Bentley, Maryland, USA
 Electron Microscopy Transmission Electron Microscopy SEM = Scanning Electron Microscopy TEM = Transmission Electron Microscopy Sara Henriksson, UCEM 2017-02-16 E. coli, William E. Bentley, Maryland, USA
Electron Microscopy Transmission Electron Microscopy SEM = Scanning Electron Microscopy TEM = Transmission Electron Microscopy Sara Henriksson, UCEM 2017-02-16 E. coli, William E. Bentley, Maryland, USA
Lecture 7 Cell Biolog y ٢٢٢ ١
 Lecture 7 ١ Mitochondria ٢ Mitochondria Mitochondria are the energy factories of the cells. The energy currency for the work that animals must do is the energy-rich molecule adenosine triphosphate (ATP).
Lecture 7 ١ Mitochondria ٢ Mitochondria Mitochondria are the energy factories of the cells. The energy currency for the work that animals must do is the energy-rich molecule adenosine triphosphate (ATP).
Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass
 Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass University of California, Davis, USA http://www.cs.ucdavis.edu/~koehl/ims2017/ Biology = Multiscale. 10 6 m 10 3 m m mm µm nm Å ps
Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass University of California, Davis, USA http://www.cs.ucdavis.edu/~koehl/ims2017/ Biology = Multiscale. 10 6 m 10 3 m m mm µm nm Å ps
Fourier Transform. sin(n# x)), where! = 2" / L and
 Fourier Transform Henning Stahlberg Introduction The tools provided by the Fourier transform are helpful for the analysis of 1D signals (time and frequency (or Fourier) domains), as well as 2D/3D signals
Fourier Transform Henning Stahlberg Introduction The tools provided by the Fourier transform are helpful for the analysis of 1D signals (time and frequency (or Fourier) domains), as well as 2D/3D signals
Life Depends on Photosynthesis
 Photosynthesis Life Depends on Photosynthesis Most energy Comes from the Sun Life Depends on Photosynthesis Most energy Comes from the Sun Life Depends on Photosynthesis Most energy Comes from the Sun
Photosynthesis Life Depends on Photosynthesis Most energy Comes from the Sun Life Depends on Photosynthesis Most energy Comes from the Sun Life Depends on Photosynthesis Most energy Comes from the Sun
Characterisation of Nanoparticle Structure by High Resolution Electron Microscopy
 Journal of Physics: Conference Series OPEN ACCESS Characterisation of Nanoparticle Structure by High Resolution Electron Microscopy To cite this article: Robert D Boyd et al 2014 J. Phys.: Conf. Ser. 522
Journal of Physics: Conference Series OPEN ACCESS Characterisation of Nanoparticle Structure by High Resolution Electron Microscopy To cite this article: Robert D Boyd et al 2014 J. Phys.: Conf. Ser. 522
MSE 321 Structural Characterization
 Auger Spectroscopy Auger Electron Spectroscopy (AES) Scanning Auger Microscopy (SAM) Incident Electron Ejected Electron Auger Electron Initial State Intermediate State Final State Physical Electronics
Auger Spectroscopy Auger Electron Spectroscopy (AES) Scanning Auger Microscopy (SAM) Incident Electron Ejected Electron Auger Electron Initial State Intermediate State Final State Physical Electronics
Guided Reading Activities
 Name Period Chapter 4: A Tour of the Cell Guided Reading Activities Big Idea: Introduction to the Cell Answer the following questions as you read Modules 4.1 4.4: 1. A(n) uses a beam of light to illuminate
Name Period Chapter 4: A Tour of the Cell Guided Reading Activities Big Idea: Introduction to the Cell Answer the following questions as you read Modules 4.1 4.4: 1. A(n) uses a beam of light to illuminate
Mitochondria Mitochondria were first seen by kollicker in 1850 in muscles and called them sarcosomes. Flemming (1882) described these organelles as
 Mitochondria Mitochondria were first seen by kollicker in 1850 in muscles and called them sarcosomes. Flemming (1882) described these organelles as filia Altmann (1890) observed these structures and named
Mitochondria Mitochondria were first seen by kollicker in 1850 in muscles and called them sarcosomes. Flemming (1882) described these organelles as filia Altmann (1890) observed these structures and named
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.2259 An infinite chainmail of M 6 L 6 metallacycles featuring multiple Borromean links Flora L. Thorp-Greenwood, Alexander N. Kulak and Michaele J. Hardie * School of Chemistry, University
DOI: 10.1038/NCHEM.2259 An infinite chainmail of M 6 L 6 metallacycles featuring multiple Borromean links Flora L. Thorp-Greenwood, Alexander N. Kulak and Michaele J. Hardie * School of Chemistry, University
Supporting Information
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supporting Information to True ferroelectric switching in thin films of trialkylbenzene-1,3,5-tricarboxamide
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supporting Information to True ferroelectric switching in thin films of trialkylbenzene-1,3,5-tricarboxamide
Reduced radiation damage in transmission electron microscopy of. proteins in graphene liquid cells
 Supplementary Information Reduced radiation damage in transmission electron microscopy of proteins in graphene liquid cells Sercan Keskin, and Niels de Jonge, INM Leibniz Institute for New Materials, D-66123
Supplementary Information Reduced radiation damage in transmission electron microscopy of proteins in graphene liquid cells Sercan Keskin, and Niels de Jonge, INM Leibniz Institute for New Materials, D-66123
Fourier transforms and convolution
 Fourier transforms and convolution (without the agonizing pain) CS/CME/BioE/Biophys/BMI 279 Oct. 26, 2017 Ron Dror 1 Outline Why do we care? Fourier transforms Writing functions as sums of sinusoids The
Fourier transforms and convolution (without the agonizing pain) CS/CME/BioE/Biophys/BMI 279 Oct. 26, 2017 Ron Dror 1 Outline Why do we care? Fourier transforms Writing functions as sums of sinusoids The
Improving cryo-em maps: focused 2D & 3D classifications and focused refinements
 Improving cryo-em maps: focused 2D & 3D classifications and focused refinements 3rd International Symposium on Cryo-3D Image Analysis 23.3.2018, Tahoe Bruno Klaholz Centre for Integrative Biology, IGBMC,
Improving cryo-em maps: focused 2D & 3D classifications and focused refinements 3rd International Symposium on Cryo-3D Image Analysis 23.3.2018, Tahoe Bruno Klaholz Centre for Integrative Biology, IGBMC,
 Supplemental Figure and Movie Legends Figure S1. Time course experiments on the thermal stability of apo AT cpn-α by native PAGE (related to Figure 2B). The samples were heated, respectively, to (A) 45
Supplemental Figure and Movie Legends Figure S1. Time course experiments on the thermal stability of apo AT cpn-α by native PAGE (related to Figure 2B). The samples were heated, respectively, to (A) 45
Comparison of single-particle analysis and electron tomography approaches: an overview
 Journal of Microscopy, Vol. 232, Pt 3 2008, pp. 562 579 Received 4 July 2007; accepted 2 March 2008 Comparison of single-particle analysis and electron tomography approaches: an overview S. JONIĆ, C.O.S.
Journal of Microscopy, Vol. 232, Pt 3 2008, pp. 562 579 Received 4 July 2007; accepted 2 March 2008 Comparison of single-particle analysis and electron tomography approaches: an overview S. JONIĆ, C.O.S.
Integrating Globus into a Science Gateway for Cryo-EM
 Integrating Globus into a Science Gateway for Cryo-EM Michael Cianfrocco Life Sciences Institute Department of Biological Chemistry University of Michigan Globus World 2018 Impact & growth of Executive
Integrating Globus into a Science Gateway for Cryo-EM Michael Cianfrocco Life Sciences Institute Department of Biological Chemistry University of Michigan Globus World 2018 Impact & growth of Executive
A short overview of electron tomography
 A short overview of electron tomography Ozan Öktem ozan.oktem@sidec.com Sidec Technologies January 17, 2006 IMA Workshop: New Mathematics and Algorithms for 3-D Image Analysis, January 9-12, 2006 Outline
A short overview of electron tomography Ozan Öktem ozan.oktem@sidec.com Sidec Technologies January 17, 2006 IMA Workshop: New Mathematics and Algorithms for 3-D Image Analysis, January 9-12, 2006 Outline
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary References
 Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary References Supplementary Figure 1. SEM images of perovskite single-crystal patterned thin film with
Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary References Supplementary Figure 1. SEM images of perovskite single-crystal patterned thin film with
Some Optimization and Statistical Learning Problems in Structural Biology
 Some Optimization and Statistical Learning Problems in Structural Biology Amit Singer Princeton University, Department of Mathematics and PACM January 8, 2013 Amit Singer (Princeton University) January
Some Optimization and Statistical Learning Problems in Structural Biology Amit Singer Princeton University, Department of Mathematics and PACM January 8, 2013 Amit Singer (Princeton University) January
Focused-ion-beam BRIEF COMMUNICATIONS. Michael Marko 1, Chyongere Hsieh 1, Richard Schalek 2, Joachim Frank 1,3 & Carmen Mannella 1
 BRIEF COMMUNICATIONS 2007 Nature Publishing Group http://www.nature.com/naturemethods Focused-ion-beam thinning of frozenhydrated biological specimens for cryoelectron microscopy Michael Marko 1, Chyongere
BRIEF COMMUNICATIONS 2007 Nature Publishing Group http://www.nature.com/naturemethods Focused-ion-beam thinning of frozenhydrated biological specimens for cryoelectron microscopy Michael Marko 1, Chyongere
Supplementary Information
 SSZ-52, a zeolite with an 18-layer aluminosilicate framework structure related to that of the DeNOx catalyst Cu-SSZ-13 Dan Xie 1,2, Lynne B. McCusker 2 *, Christian Baerlocher 2, Stacey I. Zones 1 *, Wei
SSZ-52, a zeolite with an 18-layer aluminosilicate framework structure related to that of the DeNOx catalyst Cu-SSZ-13 Dan Xie 1,2, Lynne B. McCusker 2 *, Christian Baerlocher 2, Stacey I. Zones 1 *, Wei
Scanning Electron Microscopy
 Scanning Electron Microscopy Amanpreet Kaur 1 www.reading.ac.uk/emlab Scanning Electron Microscopy What is scanning electron microscopy? Basic features of conventional SEM Limitations of conventional SEM
Scanning Electron Microscopy Amanpreet Kaur 1 www.reading.ac.uk/emlab Scanning Electron Microscopy What is scanning electron microscopy? Basic features of conventional SEM Limitations of conventional SEM
3. Bonding Ionic Bonding
 3. Bonding Ionic Bonding Metal atoms lose electrons to form +ve ions. on-metal atoms gain electrons to form -ve ions. Mg goes from 1s 2 2s 2 2p 6 3s 2 to Mg 2+ 1s 2 2s 2 2p 6 goes from 1s 2 2s 2 2p 4 to
3. Bonding Ionic Bonding Metal atoms lose electrons to form +ve ions. on-metal atoms gain electrons to form -ve ions. Mg goes from 1s 2 2s 2 2p 6 3s 2 to Mg 2+ 1s 2 2s 2 2p 6 goes from 1s 2 2s 2 2p 4 to
High-Resolution. Transmission. Electron Microscopy
 Part 4 High-Resolution Transmission Electron Microscopy 186 Significance high-resolution transmission electron microscopy (HRTEM): resolve object details smaller than 1nm (10 9 m) image the interior of
Part 4 High-Resolution Transmission Electron Microscopy 186 Significance high-resolution transmission electron microscopy (HRTEM): resolve object details smaller than 1nm (10 9 m) image the interior of
The Tecnai Arctica (TEM-9) Cryo-EM workflow. Contact: Svetla Stoilova-McPhie, PhD Advanced bioimaging scientist
 The Tecnai Arctica (TEM-9) Cryo-EM workflow Contact: Svetla Stoilova-McPhie, PhD Advanced bioimaging scientist stoilovamcphie@fas.harvard.edu The Arctica Cryo-EM training chart Cryo-TEM training involves
The Tecnai Arctica (TEM-9) Cryo-EM workflow Contact: Svetla Stoilova-McPhie, PhD Advanced bioimaging scientist stoilovamcphie@fas.harvard.edu The Arctica Cryo-EM training chart Cryo-TEM training involves
University of Groningen
 University of Groningen Megacomplex organization of the oxidative phosphorylation system by structural analysis of respiratory supercomplexes from potato Bultema, Jelle B.; Braun, Hans-Peter; Boekema,
University of Groningen Megacomplex organization of the oxidative phosphorylation system by structural analysis of respiratory supercomplexes from potato Bultema, Jelle B.; Braun, Hans-Peter; Boekema,
Introduction to Cryo Microscopy. Nikolaus Grigorieff
 Introduction to Cryo Microscopy Nikolaus Grigorieff Early TEM and Biology 1906-1988 Bacterial culture in negative stain. Friedrich Krause, 1937 Nobel Prize in Physics 1986 Early design, 1931 Decades of
Introduction to Cryo Microscopy Nikolaus Grigorieff Early TEM and Biology 1906-1988 Bacterial culture in negative stain. Friedrich Krause, 1937 Nobel Prize in Physics 1986 Early design, 1931 Decades of
channel/ryanodine receptor from sarcoplasmic reticulum
 Cryo-EM of the native structure of the calcium release channel/ryanodine receptor from sarcoplasmic reticulum M. Radermacher, T. Wagenknecht, R. Grassucci, J. Frank, M. Inui,* C. Chadwick,* and S. Fleischer*
Cryo-EM of the native structure of the calcium release channel/ryanodine receptor from sarcoplasmic reticulum M. Radermacher, T. Wagenknecht, R. Grassucci, J. Frank, M. Inui,* C. Chadwick,* and S. Fleischer*
F. Piazza Center for Molecular Biophysics and University of Orléans, France. Selected topic in Physical Biology. Lecture 1
 Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Methoden moderner Röntgenphysik I. Coherence based techniques II. Christian Gutt DESY, Hamburg
 Methoden moderner Röntgenphysik I Coherence based techniques II Christian Gutt DESY Hamburg christian.gutt@desy.de 8. January 009 Outline 18.1. 008 Introduction to Coherence 8.01. 009 Structure determination
Methoden moderner Röntgenphysik I Coherence based techniques II Christian Gutt DESY Hamburg christian.gutt@desy.de 8. January 009 Outline 18.1. 008 Introduction to Coherence 8.01. 009 Structure determination
Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Information: Long range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
HOW TO APPROACH SCANNING ELECTRON MICROSCOPY AND ENERGY DISPERSIVE SPECTROSCOPY ANALYSIS. SCSAM Short Course Amir Avishai
 HOW TO APPROACH SCANNING ELECTRON MICROSCOPY AND ENERGY DISPERSIVE SPECTROSCOPY ANALYSIS SCSAM Short Course Amir Avishai RESEARCH QUESTIONS Sea Shell Cast Iron EDS+SE Fe Cr C Objective Ability to ask the
HOW TO APPROACH SCANNING ELECTRON MICROSCOPY AND ENERGY DISPERSIVE SPECTROSCOPY ANALYSIS SCSAM Short Course Amir Avishai RESEARCH QUESTIONS Sea Shell Cast Iron EDS+SE Fe Cr C Objective Ability to ask the
Crystals, X-rays and Proteins
 Crystals, X-rays and Proteins Comprehensive Protein Crystallography Dennis Sherwood MA (Hons), MPhil, PhD Jon Cooper BA (Hons), PhD OXFORD UNIVERSITY PRESS Contents List of symbols xiv PART I FUNDAMENTALS
Crystals, X-rays and Proteins Comprehensive Protein Crystallography Dennis Sherwood MA (Hons), MPhil, PhD Jon Cooper BA (Hons), PhD OXFORD UNIVERSITY PRESS Contents List of symbols xiv PART I FUNDAMENTALS
Halorhodopsin (hr) is a light-driven chloride pump in the
 The three-dimensional structure of halorhodopsin to 5 Å by electron crystallography: A new unbending procedure for two-dimensional crystals by using a global reference structure Edmund R. S. Kunji*, Susanne
The three-dimensional structure of halorhodopsin to 5 Å by electron crystallography: A new unbending procedure for two-dimensional crystals by using a global reference structure Edmund R. S. Kunji*, Susanne
Determination of the Substructure
 Monday, June 15 th, 2009 Determination of the Substructure EMBO / MAX-INF2 Practical Course http://shelx.uni-ac.gwdg.de Overview Substructure Definition and Motivation Extracting Substructure Data from
Monday, June 15 th, 2009 Determination of the Substructure EMBO / MAX-INF2 Practical Course http://shelx.uni-ac.gwdg.de Overview Substructure Definition and Motivation Extracting Substructure Data from
Summary: Crystallography in a nutshell. Lecture no. 4. (Crystallography without tears, part 2)
 Structure Determination Summary: Crystallography in a nutshell. Lecture no. 4. (Crystallography without tears, part 2) Cele Abad-Zapatero University of Illinois at Chicago Center for Pharmaceutical Biotechnology.
Structure Determination Summary: Crystallography in a nutshell. Lecture no. 4. (Crystallography without tears, part 2) Cele Abad-Zapatero University of Illinois at Chicago Center for Pharmaceutical Biotechnology.
Development of a Radiation Hard CMOS Monolithic Pixel Sensor
 Development of a Radiation Hard CMOS Monolithic Pixel Sensor M. Battaglia 1,2, D. Bisello 3, D. Contarato 2, P. Denes 2, D. Doering 2, P. Giubilato 2,3, T.S. Kim 2, Z. Lee 2, S. Mattiazzo 3, V. Radmilovic
Development of a Radiation Hard CMOS Monolithic Pixel Sensor M. Battaglia 1,2, D. Bisello 3, D. Contarato 2, P. Denes 2, D. Doering 2, P. Giubilato 2,3, T.S. Kim 2, Z. Lee 2, S. Mattiazzo 3, V. Radmilovic
Fan, Hai-fu Institute of Physics, Chinese Academy of Sciences, Beijing , China
 Direct Methods in Crystallography Fan, Hai-fu Institute of Physics, Chinese Academy of Sciences, Beijing 100080, China An important branch of crystallography is the X-ray diffraction analysis of crystal
Direct Methods in Crystallography Fan, Hai-fu Institute of Physics, Chinese Academy of Sciences, Beijing 100080, China An important branch of crystallography is the X-ray diffraction analysis of crystal
Phase problem: Determining an initial phase angle α hkl for each recorded reflection. 1 ρ(x,y,z) = F hkl cos 2π (hx+ky+ lz - α hkl ) V h k l
 Phase problem: Determining an initial phase angle α hkl for each recorded reflection 1 ρ(x,y,z) = F hkl cos 2π (hx+ky+ lz - α hkl ) V h k l Methods: Heavy atom methods (isomorphous replacement Hg, Pt)
Phase problem: Determining an initial phase angle α hkl for each recorded reflection 1 ρ(x,y,z) = F hkl cos 2π (hx+ky+ lz - α hkl ) V h k l Methods: Heavy atom methods (isomorphous replacement Hg, Pt)
Computer Vision. Filtering in the Frequency Domain
 Computer Vision Filtering in the Frequency Domain Filippo Bergamasco (filippo.bergamasco@unive.it) http://www.dais.unive.it/~bergamasco DAIS, Ca Foscari University of Venice Academic year 2016/2017 Introduction
Computer Vision Filtering in the Frequency Domain Filippo Bergamasco (filippo.bergamasco@unive.it) http://www.dais.unive.it/~bergamasco DAIS, Ca Foscari University of Venice Academic year 2016/2017 Introduction
4. Other diffraction techniques
 4. Other diffraction techniques 4.1 Reflection High Energy Electron Diffraction (RHEED) Setup: - Grazing-incidence high energy electron beam (3-5 kev: MEED,
4. Other diffraction techniques 4.1 Reflection High Energy Electron Diffraction (RHEED) Setup: - Grazing-incidence high energy electron beam (3-5 kev: MEED,
Supplementary Figure 1
 Supplementary Figure 1 Electron-dense inter-mitochondrial junctions (IMJs) and associated cristae organization are evolutionary conserved features of mitochondria-mitochondria contacts. Shown are electron
Supplementary Figure 1 Electron-dense inter-mitochondrial junctions (IMJs) and associated cristae organization are evolutionary conserved features of mitochondria-mitochondria contacts. Shown are electron
Christopher Pavlik Bioanalytical Chemistry March 2, 2011
 Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. DOI: 10.1038/NCHEM.2675 Real-time molecular scale observation of crystal formation Roy E. Schreiber, 1 Lothar Houben, 2 Sharon G. Wolf, 2 Gregory Leitus,
In the format provided by the authors and unedited. DOI: 10.1038/NCHEM.2675 Real-time molecular scale observation of crystal formation Roy E. Schreiber, 1 Lothar Houben, 2 Sharon G. Wolf, 2 Gregory Leitus,
Supplementary Figure 1: Example non-overlapping, binary probe functions P1 (~q) and P2 (~q), that add to form a top hat function A(~q).
 Supplementary Figures P(q) A(q) + Function Value P(q) qmax = Supplementary Figure : Example non-overlapping, binary probe functions P (~q) and P (~q), that add to form a top hat function A(~q). qprobe
Supplementary Figures P(q) A(q) + Function Value P(q) qmax = Supplementary Figure : Example non-overlapping, binary probe functions P (~q) and P (~q), that add to form a top hat function A(~q). qprobe
Chapter 10: PHOTOSYNTHESIS
 Chapter 10: PHOTOSYNTHESIS 1. Overview of Photosynthesis 2. Light Absorption 3. The Light Reactions 4. The Calvin Cycle 1. Overview of Photosynthesis Chapter Reading pp. 185-190, 206-207 What is Photosynthesis?
Chapter 10: PHOTOSYNTHESIS 1. Overview of Photosynthesis 2. Light Absorption 3. The Light Reactions 4. The Calvin Cycle 1. Overview of Photosynthesis Chapter Reading pp. 185-190, 206-207 What is Photosynthesis?
