Photoperiodic effect on protein profiles of potato petiole
|
|
|
- Lester Rodgers
- 5 years ago
- Views:
Transcription
1 Photoperiodic effect on protein profiles of potato petiole Shweta Shah 1, Young Jin-Lee 2, David J. Hannapel 3 and A. Gururaj Rao* 1 1 Department of Biochemistry, Biophysics and Molecular Biology, Iowa State University, Ames, Iowa Department of Chemistry, Iowa State University, Ames, Iowa Plant Biology Major, Iowa State University, Ames, Iowa Abstract In potato (Solanum tuberosum), StBEL5 mrna is a mobile RNA that is involved in the signaling system that activates tuber formation. Short-day (SD) conditions induce tuberization and long-day (LD) inhibits the process. The transcriptional source of this mobile RNA is leaf veins and leaf stalks designated, petioles. Transport of StBEL5 RNA is induced by a SD photoperiod that leads to tuber formation. It is likely that the movement of StBEL5 is facilitated by the formation of RNA-protein complex(s). To further understand this proposed mechanism of downstream signaling we have undertaken a detailed proteomic analysis of proteins isolated from potato petioles (PP) grown under LD and SD photoperiod conditions using 2-dimensional gel electrophoresis (2-DE) followed by MALDI MS/MS and/or LC MS/MS. Proteins that were differentially expressed in response to changes in photoperiods were analyzed by Progenesis SameSpots software. Phosphoproteins and RNA-binding proteins were enriched using immobilized metal affinity chromatography and poly(u) Sepharose columns respectively from SD and LD PP protein extracts and similarly analyzed. In addition to establishing a catalog of PP proteins, we have so far identified nearly sixty-seven proteins that are differentially expressed in response to SD or LD photoperiods. Significantly, the RNA-binding protein, elf-5a, has been detected in multiple isoforms specifically under LD conditions. Numerous other poly(u)-binding proteins which contain RNA recognition motifs have also been isolated and identified. This is the first comprehensive proteomics study that examines and catalogs the proteins that are present in the potato petiole and are potentially differentially regulated by photoperiod.
2 Table 1: Catalogue of differentially expressed proteins of potato petiole extract from plants grown under LD and SD photoperiod. RNA-binding proteins are highlighted in red. Spot # Acession # MW (kda) % coverage # unique peptide Score TAIR accession # Blast E-value 20 gi /8 162 AT4G E- 37 TAIR Annotation Plastocyanin-like domain-containing protein Putative function Electron carrier, copper ion binding a Fold Difference LD: No hit 80 gi /11 99 AT4G Embryo defective RNA binding, translation SD: elongation factor 95 gi /2 90 AT4G Embryo defective 2726 RNA binding, translation elongation factor SD: gi /2 91 AT1G Arabidopsis thaliana Cell division cycle 5 DNA binding/ transcription factor LD: gi /4 229 AT4G Glycine decarboxylase P- protein 1 Glycine dehydrogenase, pyridoxal phosphate binding SD: gi /4 120 AT2G Aconitate hydratase, cytoplasmic, 220 gi /4 120 AT2G Aconitate hydratase, cytoplasmic, Aconitate hydratase, ATP binding Aconitate hydratase, ATP binding 240 No hit SD: 1:6 272 No hit LD: No hit SD: gi /8 139 AT5G UTP--glucose-1- Nucleotidyltransferase LD:2.0 phosphate uridylyltransferase, 484 gi /6 595 AT2G Fumarase 1 Fumarate hydratase LD:1.6 SD:1.6 SD:1.6
3 gi /5 482 AT3G E- 170 Aminopeptidase/ metalloexopeptidase 510 gi /5 301 AT1G Isocitrate dehydrogenase, 520 gi /8 727 AT3G Monodehydroascorba -te reductase, gi / AT5G Sensitive to hot temperatures 5 Metalloexopeptidase Isocitrate dehydrogenase (NADP+), copper ion binding Monodehydroascorbate reductase S-nitrosoglutathione reductase, 564 gi /1 47 AT1G ARG1-LIKE 1 Unfolded protein binding, heat shock protein binding 570 gi /3 119 AT3G Reversibly glycosylated polypeptide 1 Cellulose synthase (UDPforming) 571 gi ,2 13/5 488 AT1G AT-E1 Alpha Oxidoreductase LD1.8 gi /8 520 AT4G Aspartate aminotransferase 602 gi /5 334 AT3G E- 170 Chloroplast stemloop binding protein of 41 kda L-aspartate:2- oxoglutarate aminotransferase mrna binding, poly(u) binding 666 gi /2 105 AT3G Heat shock protein 70 ATP binding 707 gi /3 213 AT2G E- Elongation factor 1- Translation elongation SD: beta, factor gi /5 893 AT1G E gi /5 395 AT3G E gi /5 488 AT1G E- 126 gi /4 323 AT2G E- 123 General regulatory factor 2/ protein PAE2; endopeptidase Ascorbate peroxidase 1 PAG1; endopeptidase Protein phosphorylated amino acid binding/protein binding Threonine-type endopeptidase L-ascorbate peroxidase Threonine-type endopeptidase LD:1.6 SD:3.6 LD:1.6
4 765 gi / E gi /2 293 AT3G E gi /7 249 AT2G E gi /4 562 AT3G E- 116 APX1 (ascorbate peroxidase 1) Triose-phosphate isomerase 29 kda ribonucleoprotein PBF1; peptidase 818 gi /5 149 AT1G e-89 Dehydroascorbate reductase gi /4 189 AT1G Glyceraldehyde-3- phosphate dehydrogenase 853 gi /4 165 AT4G Vacuolar ATP synthase subunit B, 854 gi /1 75 AT4G Vacuolar ATP synthase subunit B, 879 gi /3 379 AT3G E gi /5 604 AT3G E gi /1 93 AT3G E gi /2 94 AT1G E gi /5 263 AT1G E gi /4 229 AT1G E- 144 Nascent polypeptide associated complex alpha chain protein Endoribonuclease L- PSP family protein Ribosomal protein L 12-C subunit P-1 subunit P-1 subunit P-1 L-ascorbate peroxidase Triose-phosphate isomerase RNA binding, poly(u) binding Peptidase/ threonine-type endopeptidase Glutathione binding Glyceraldehyde-3- phosphate dehydrogenase (NADP+) Hydrogen ion transporting ATP synthase Hydrogen ion transporting ATP synthase Involved in response to salt stress Endoribonuclease Structural constituent of ribosome/ translation Poly(U) binding, photosynthesis Poly(U) binding, photosynthesis Poly(U) binding, photosynthesis SD:1.9 SD: 2.1 LD:2.0 LD:2.1 SD: 1.7 LD1.6 SD1.6 SD:2.3
5 940 gi /5 263 AT1G E- 144 subunit P-1 Poly(U) binding, photosynthesis 947 gi /5 753 AT4G39260 GR-RBP8 RNA binding / nucleic acid binding 996 No hit 997 gi AT3G E- 60S acidic ribosomal Structural constituent of 23 protein P2 ribosome 1001 No hit SD: gi AT3G E S acidic ribosomal protein P2 Structural constituent of ribosome 1009 gi AT3G E S acidic ribosomal protein P2 Structural constituent of ribosome SD1.6 SD: No hit SD: gi AT4G E- NDPK1; ATP Nucleoside diphosphate SD: binding / nucleoside diphosphate kinase [ kinase / ATP binding 1071 gi /2 160 AT4G E- Senescence 1 Molecular_function SD: unknown 1078 gi /3 142 AT5G E- Calmodulin 6 Calcium ion binding SD: No hit 1089 gi /2 284 AT1G E gi /2 178 AT3G E gi /2 95 AT1G E b gi AT1G E b gi /1 63 AT3G E- 139 Arabidopsis homolog of anti-oxidant1 Subunit O-2 Metal ion binding Oxygen evolving, poly(u) binding SD:2.0 SD:2.0 Related to Ubiquitin 1 Protein binding SD:1.6 Peroxidase 12 Electron carrier LD:1.9 (S)-2-hydroxy-acid oxidase, peroxisomal, Glycolate oxidase
6 645b gi AT2G Fructosebisphosphate 684b gi /2 101 AT3G E b gi /3 319 AT4G E- 133 aldolase, Glyceraldehyde-3- phosphate dehydrogenase C subunit1 Xyloglucosyl transferase, 817b gi / Hypothetical protein (Solanum tuberosum) Fructose-bisphosphate aldolase Glyceraldehyde-3- phosphate dehydrogenase (phosphorylating) Hydrolase 958b No hit SD: b gi /1 86 AT1G E- LD: PBG1; peptidase/ threonine-type endopeptidase 984b gi /1 109 AT4G Rab GTPASE HomologE1B Peptidase / response to cadmium ion, response to salt stress GTP binding, translation elongation factor 1054b gi /2 64 AT4G e-78 Osmotin 34 Response to salt stress LD: b gi /3 293 AT5G E b gi /3 480 AT5G E- 91 Peptidyl-prolyl cistrans isomerase cyclophilin-type family protein CBS domaincontaining protein Isomerase /protein folding LD:1.7 LD:2.1 SD:2.2 SD: 1.8 Response to salt stress LD: b No hit SD: b gi /4 172 AT2G E- ROC3; peptidylprolyl Isomerase /protein SD: cis-trans folding/signal 1356b gi /2 74 AT3G E- 49 a : higher fold expression under indicated condition. isomerase Pathogenesis related 4 transduction Chitin binding/ response to salt stress LD:1.9
7 Table 2: Catalogue of phosphoproteins enriched from potato petioles extract from plants grown under LD and SD photoperiod. Spot # 5 gi /2 233 AT5G ATGSR1 Glutamate-ammonia ligase, copper ion binding 6 gi /2 101 AT5G E-141 Unknown protein- Molecular function unknown 7 gi /2 177 AT5G E-174 Unknown protein- Molecular function unknown 8 gi /2 177 AT5G E-174 Unknown protein- Molecular function unknown 9 gi /2 177 AT5G E-174 Unknown protein- Molecular function unknown 10 gi /4 452 AT5G Glutmate dehydrogenase 1 ATP binding/ oxidoreductase 11 gi /3 472 AT5G Glutmate dehydrogenase 1 ATP binding/ oxidoreductase 12 gi /2 344 AT5G Glutmate dehydrogenase 1 ATP binding/ oxidoreductase 13 gi /1 153 AT5G Glutmate dehydrogenase 2 ATP binding /oxidoreductase 14 No hit Accession # MW (kda) % coverage/ # unique peptide MOWSE Score TAIR acession # Blast E-value TAIR Annotation Putative function 1 gi /2 161 AT5G e-174 Unknown protein- Biological process unknown 2 gi /3 185 AT1G E-138 Glyoxalase I homolog Lactoylglutathione lyase, metal ion binding 3 gi /3 185 AT1G E-138 Glyoxalase I homolog Lactoylglutathione lyase, metal ion binding 4 gi /2 242 AT5G ATGSR1 Glutamate-ammonia ligase, copper ion binding 15 No hit 16 No hit
8 17 gi /2 115 AT1G E-78 Eukaryotic elongation factor 5A-1 Translation initiation factor 18 gi /3 252 AT1G E-78 Eukaryotic elongation factor 5A-1 Translation initiation factor 19 gi /2 117 AT1G E-75 Eukaryotic elongation factor 5A-1 Translation initiation factor 20 gi /2 99 AT1G E-79 Eukaryotic elongation factor 5A-3 Translation initiation factor 21 gi /2 121 AT5G E-141 Unknown protein Molecular function unknown 22 gi /1 95 AT3G E-99 Arabidopsis thaliana pyrophosphorylase 4 Inorganic diphosphatase 23 No hit 24 No hit 25 no hit 26 gi /4 208 AT4G E-108-1,3-glucanase1 Hydrolase, hydrolyzing O-glycosyl compounds 27 gi /1 68 AT1G Calreticulin 3 Unfolded protein binding, calcium ion binding 28 gi /1 70 AT4G E-16 Aminoacylase, Metallopeptidase, hydrolase, protein dimerization 29 gi /3 264 AT5G E-163 Unknown protein- Molecular function unknown 30 gi /3 185 AT1G11840 Glyoxalase I homolog Lactoylglutathione lyase, metal ion binding 31 gi /3 186 AT1G E-134 Glyoxalase I homolog Lactoylglutathione lyase, metal ion binding 32 gi /3 184 AT1G E-134 Glyoxalase I homolog Lactoylglutathione lyase, metal ion binding 33 gi /2 338 AT5G BIP1; ATP binding Protein folding, ER-associated protein catabolic process, response to heat
9 34 gi /1 56 AT3G Heat shock protein 60 Chaperone-mediated protein complex assembly 35 gi /2 87 AT5G E-177 Endoxyloglucan Hydrolase Transferase A4 36 gi /1 71 AT3G (S)-2-hydroxy-acid oxidase, peroxisomal, Glycolate oxidase 37 gi /2 83 AT5G E-146 Xyloglucan endotransglycosylase 3 38 gi /3 176 AT5G E-174 Unknown protein- 39 gi /3 142 AT5G E-174 Unknown protein- 40 gi /3 143 AT5G E-174 Unknown protein- Hydrolase Molecular function unknown Molecular function unknown Molecular function unknown
GO ID GO term Number of members GO: translation 225 GO: nucleosome 50 GO: calcium ion binding 76 GO: structural
 GO ID GO term Number of members GO:0006412 translation 225 GO:0000786 nucleosome 50 GO:0005509 calcium ion binding 76 GO:0003735 structural constituent of ribosome 170 GO:0019861 flagellum 23 GO:0005840
GO ID GO term Number of members GO:0006412 translation 225 GO:0000786 nucleosome 50 GO:0005509 calcium ion binding 76 GO:0003735 structural constituent of ribosome 170 GO:0019861 flagellum 23 GO:0005840
Biological Process Term Enrichment
 Biological Process Term Enrichment cellular protein localization cellular macromolecule localization intracellular protein transport intracellular transport generation of precursor metabolites and energy
Biological Process Term Enrichment cellular protein localization cellular macromolecule localization intracellular protein transport intracellular transport generation of precursor metabolites and energy
Transcriptome analysis of leaf tissue from Bermudagrass (Cynodon dactylon) using a normalised cdna library
 Ó CSIRO 2008 ISSN 1445-4408 10.1071/FP08133_AC Functional Plant Biology, 2008, 35, 585 594 Transcriptome analysis of leaf tissue from Bermudagrass (Cynodon dactylon) using a normalised cdna library Changsoo
Ó CSIRO 2008 ISSN 1445-4408 10.1071/FP08133_AC Functional Plant Biology, 2008, 35, 585 594 Transcriptome analysis of leaf tissue from Bermudagrass (Cynodon dactylon) using a normalised cdna library Changsoo
Membrane-associated proteomics of chickpea identifies Sad1/UNC-84 protein (CaSUN1), a novel component of dehydration signaling
 Membrane-associated proteomics of chickpea identifies Sad/UNC-84 (CaSUN), a novel component of dehydration signaling Dinesh Kumar Jaiswal, Poonam Mishra, Pratigya Subba, Divya Rathi, Subhra Chakraborty*
Membrane-associated proteomics of chickpea identifies Sad/UNC-84 (CaSUN), a novel component of dehydration signaling Dinesh Kumar Jaiswal, Poonam Mishra, Pratigya Subba, Divya Rathi, Subhra Chakraborty*
Characters related to higher starch accumulation in cassava storage roots
 Characters related to higher starch accumulation in cassava storage roots You-Zhi Li **1, Jian-Yu Zhao *1, San-Min Wu 1, Xian-Wei Fan 1, Xing-Lu Luo 1 & Bao-Shan Chen **1 1 State Key Laboratory for Conservation
Characters related to higher starch accumulation in cassava storage roots You-Zhi Li **1, Jian-Yu Zhao *1, San-Min Wu 1, Xian-Wei Fan 1, Xing-Lu Luo 1 & Bao-Shan Chen **1 1 State Key Laboratory for Conservation
Population transcriptomics uncovers the regulation of gene. expression variation in adaptation to changing environment
 Supplementary information Population transcriptomics uncovers the regulation of gene expression variation in adaptation to changing environment Qin Xu 1, Caiyun Zhu 2,4, Yangyang Fan 1,4, Zhihong Song
Supplementary information Population transcriptomics uncovers the regulation of gene expression variation in adaptation to changing environment Qin Xu 1, Caiyun Zhu 2,4, Yangyang Fan 1,4, Zhihong Song
SUPPLEMENTARY MATERIAL. The table lists the 437 proteins proposed to be myristoylated in A. thaliana. The AGI
 SUPPLEMENTARY MATERIAL Table s1: the N-myristoylome in A. thaliana The table lists the 437 proteins proposed to be myristoylated in A. thaliana. The AGI entries were re-annotated according to the most
SUPPLEMENTARY MATERIAL Table s1: the N-myristoylome in A. thaliana The table lists the 437 proteins proposed to be myristoylated in A. thaliana. The AGI entries were re-annotated according to the most
Chapter
 Chapter 17 17.4-17.6 Molecular Components of Translation A cell interprets a genetic message and builds a polypeptide The message is a series of codons on mrna The interpreter is called transfer (trna)
Chapter 17 17.4-17.6 Molecular Components of Translation A cell interprets a genetic message and builds a polypeptide The message is a series of codons on mrna The interpreter is called transfer (trna)
Enzymes I. Dr. Mamoun Ahram Summer semester,
 Enzymes I Dr. Mamoun Ahram Summer semester, 2017-2018 Resources Mark's Basic Medical Biochemistry Other resources NCBI Bookshelf: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=books The Medical Biochemistry
Enzymes I Dr. Mamoun Ahram Summer semester, 2017-2018 Resources Mark's Basic Medical Biochemistry Other resources NCBI Bookshelf: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=books The Medical Biochemistry
Enzyme Catalysis & Biotechnology
 L28-1 Enzyme Catalysis & Biotechnology Bovine Pancreatic RNase A Biochemistry, Life, and all that L28-2 A brief word about biochemistry traditionally, chemical engineers used organic and inorganic chemistry
L28-1 Enzyme Catalysis & Biotechnology Bovine Pancreatic RNase A Biochemistry, Life, and all that L28-2 A brief word about biochemistry traditionally, chemical engineers used organic and inorganic chemistry
Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport
 Ph.D. thesis Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport Zsigmond Laura Supervisor: Dr. Szabados László Arabidopsis Molecular Genetic Group Institute of Plant
Ph.D. thesis Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport Zsigmond Laura Supervisor: Dr. Szabados László Arabidopsis Molecular Genetic Group Institute of Plant
Table 5. Genes unique to G. thermodenitrificans NG80-2 Gene ID Gene name Gene product COG functional category
 Table 5. Genes unique to G. thermodenitrificans NG80-2 GT0030 gt30 Methionine--tRNA ligase/methionyl-trna synthetase COG0143, Translation GT0033 gt33 Unknown GT0106 gt106 Ribosomal protein L3 COG0087,
Table 5. Genes unique to G. thermodenitrificans NG80-2 GT0030 gt30 Methionine--tRNA ligase/methionyl-trna synthetase COG0143, Translation GT0033 gt33 Unknown GT0106 gt106 Ribosomal protein L3 COG0087,
Center for Academic Services & Advising
 March 2, 2017 Biology I CSI Worksheet 6 1. List the four components of cellular respiration, where it occurs in the cell, and list major products consumed and produced in each step. i. Hint: Think about
March 2, 2017 Biology I CSI Worksheet 6 1. List the four components of cellular respiration, where it occurs in the cell, and list major products consumed and produced in each step. i. Hint: Think about
What is an enzyme? Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics. Margaret A. Daugherty Fall 2004 KEY FEATURES OF ENZYMES
 Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics Margaret A. Daugherty Fall 2004 What is an enzyme? General Properties Mostly proteins, but some are actually RNAs Biological catalysts
Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics Margaret A. Daugherty Fall 2004 What is an enzyme? General Properties Mostly proteins, but some are actually RNAs Biological catalysts
What is an enzyme? Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics. Margaret A. Daugherty Fall General Properties
 Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics Margaret A. Daugherty Fall 2003 ENZYMES: Why, what, when, where, how? All but the who! What: proteins that exert kinetic control over
Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics Margaret A. Daugherty Fall 2003 ENZYMES: Why, what, when, where, how? All but the who! What: proteins that exert kinetic control over
Quiz answers. Allele. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA)
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA) http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Quiz answers Kinase: An enzyme
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA) http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Quiz answers Kinase: An enzyme
Supplementary information
 Supplementary information Comparative proteomic study of Arabidopsis mutants mpk4 and mpk6 Tomáš Takáč 1, Pavol Vadovič 1, Tibor Pechan 2, Ivan Luptovčiak 1, Olga Šamajová 1, Jozef Šamaj 1* 1 Centre of
Supplementary information Comparative proteomic study of Arabidopsis mutants mpk4 and mpk6 Tomáš Takáč 1, Pavol Vadovič 1, Tibor Pechan 2, Ivan Luptovčiak 1, Olga Šamajová 1, Jozef Šamaj 1* 1 Centre of
Module organization and variance in protein-protein interaction networks
 Module organization and variance in protein-protein interaction networks Chun-Yu Lin 1, Tsai-Ling Lee 1, Yi-Yuan Chiu 1, Yi-Wei Lin 1, Yu-Shu Lo 1, Chih-Ta Lin 1, and Jinn-Moon Yang 1,2* 1 Institute of
Module organization and variance in protein-protein interaction networks Chun-Yu Lin 1, Tsai-Ling Lee 1, Yi-Yuan Chiu 1, Yi-Wei Lin 1, Yu-Shu Lo 1, Chih-Ta Lin 1, and Jinn-Moon Yang 1,2* 1 Institute of
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11
 UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
BCH 4054 Spring 2001 Chapter 33 Lecture Notes
 BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Outer Glycolysis mitochondrial membrane Glucose ATP
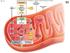 Fig. 7.5 uter Glycolysis mitochondrial membrane Glucose Intermembrane space xidation Mitochondrial matrix Acetyl-oA Krebs FAD e NAD + FAD Inner mitochondrial membrane e Electron e Transport hain hemiosmosis
Fig. 7.5 uter Glycolysis mitochondrial membrane Glucose Intermembrane space xidation Mitochondrial matrix Acetyl-oA Krebs FAD e NAD + FAD Inner mitochondrial membrane e Electron e Transport hain hemiosmosis
Translation and Operons
 Translation and Operons You Should Be Able To 1. Describe the three stages translation. including the movement of trna molecules through the ribosome. 2. Compare and contrast the roles of three different
Translation and Operons You Should Be Able To 1. Describe the three stages translation. including the movement of trna molecules through the ribosome. 2. Compare and contrast the roles of three different
Chapter 17. From Gene to Protein. Biology Kevin Dees
 Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
What is the central dogma of biology?
 Bellringer What is the central dogma of biology? A. RNA DNA Protein B. DNA Protein Gene C. DNA Gene RNA D. DNA RNA Protein Review of DNA processes Replication (7.1) Transcription(7.2) Translation(7.3)
Bellringer What is the central dogma of biology? A. RNA DNA Protein B. DNA Protein Gene C. DNA Gene RNA D. DNA RNA Protein Review of DNA processes Replication (7.1) Transcription(7.2) Translation(7.3)
Cellular Transport. 1. Transport to and across the membrane 1a. Transport of small molecules and ions 1b. Transport of proteins
 Transport Processes Cellular Transport 1. Transport to and across the membrane 1a. Transport of small molecules and ions 1b. Transport of proteins 2. Vesicular transport 3. Transport through the nuclear
Transport Processes Cellular Transport 1. Transport to and across the membrane 1a. Transport of small molecules and ions 1b. Transport of proteins 2. Vesicular transport 3. Transport through the nuclear
Lecture 25: Protein Synthesis Key learning goals: Be able to explain the main stuctural features of ribosomes, and know (roughly) how many DNA and
 Lecture 25: Protein Synthesis Key learning goals: Be able to explain the main stuctural features of ribosomes, and know (roughly) how many DNA and protein subunits they contain. Understand the main functions
Lecture 25: Protein Synthesis Key learning goals: Be able to explain the main stuctural features of ribosomes, and know (roughly) how many DNA and protein subunits they contain. Understand the main functions
Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION
 http://smtom.lecture.ub.ac.id/ Password: https://syukur16tom.wordpress.com/ Password: Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/translation2.gif
http://smtom.lecture.ub.ac.id/ Password: https://syukur16tom.wordpress.com/ Password: Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/translation2.gif
Biologic catalysts 1. Shared properties with chemical catalysts a. Enzymes are neither consumed nor produced during the course of a reaction. b.
 Enzyme definition Enzymes are protein catalysts that increase the velocity of a chemical reaction and are not consumed during the reaction they catalyze. [Note: Some types of RNA can act like enzymes,
Enzyme definition Enzymes are protein catalysts that increase the velocity of a chemical reaction and are not consumed during the reaction they catalyze. [Note: Some types of RNA can act like enzymes,
ENZYMES. by: Dr. Hadi Mozafari
 ENZYMES by: Dr. Hadi Mozafari 1 Specifications Often are Polymers Have a protein structures Enzymes are the biochemical reactions Katalyzers Enzymes are Simple & Complex compounds 2 Enzymatic Reactions
ENZYMES by: Dr. Hadi Mozafari 1 Specifications Often are Polymers Have a protein structures Enzymes are the biochemical reactions Katalyzers Enzymes are Simple & Complex compounds 2 Enzymatic Reactions
Basic Concepts of Enzyme Action. Enzymes. Rate Enhancement 9/17/2015. Stryer Short Course Chapter 6
 Basic Concepts of Enzyme Action Stryer Short Course Chapter 6 Enzymes Biocatalysts Active site Substrate and product Catalyzed rate Uncatalyzed rate Rate Enhancement Which is a better catalyst, carbonic
Basic Concepts of Enzyme Action Stryer Short Course Chapter 6 Enzymes Biocatalysts Active site Substrate and product Catalyzed rate Uncatalyzed rate Rate Enhancement Which is a better catalyst, carbonic
From gene to protein. Premedical biology
 From gene to protein Premedical biology Central dogma of Biology, Molecular Biology, Genetics transcription replication reverse transcription translation DNA RNA Protein RNA chemically similar to DNA,
From gene to protein Premedical biology Central dogma of Biology, Molecular Biology, Genetics transcription replication reverse transcription translation DNA RNA Protein RNA chemically similar to DNA,
Protein synthesis I Biochemistry 302. February 17, 2006
 Protein synthesis I Biochemistry 302 February 17, 2006 Key features and components involved in protein biosynthesis High energy cost (essential metabolic activity of cell Consumes 90% of the chemical energy
Protein synthesis I Biochemistry 302 February 17, 2006 Key features and components involved in protein biosynthesis High energy cost (essential metabolic activity of cell Consumes 90% of the chemical energy
Gene regulation I Biochemistry 302. Bob Kelm February 25, 2005
 Gene regulation I Biochemistry 302 Bob Kelm February 25, 2005 Principles of gene regulation (cellular versus molecular level) Extracellular signals Chemical (e.g. hormones, growth factors) Environmental
Gene regulation I Biochemistry 302 Bob Kelm February 25, 2005 Principles of gene regulation (cellular versus molecular level) Extracellular signals Chemical (e.g. hormones, growth factors) Environmental
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
ENCODE DCC Antibody Validation Document
 ENCODE DCC Antibody Validation Document Date of Submission Name: Email: Lab Antibody Name: Target: Company/ Source: Catalog Number, database ID, laboratory Lot Number Antibody Description: Target Description:
ENCODE DCC Antibody Validation Document Date of Submission Name: Email: Lab Antibody Name: Target: Company/ Source: Catalog Number, database ID, laboratory Lot Number Antibody Description: Target Description:
Oxidative Phosphorylation versus. Photophosphorylation
 Photosynthesis Oxidative Phosphorylation versus Photophosphorylation Oxidative Phosphorylation Electrons from the reduced cofactors NADH and FADH 2 are passed to proteins in the respiratory chain. In eukaryotes,
Photosynthesis Oxidative Phosphorylation versus Photophosphorylation Oxidative Phosphorylation Electrons from the reduced cofactors NADH and FADH 2 are passed to proteins in the respiratory chain. In eukaryotes,
Chapter 19 Overview. Protein Synthesis. for amino acid. n Protein Synthesis genetic info encoded in nucleic acids translated into standard amino acids
 Chapter 19 Overview Protein Synthesis n Protein Synthesis genetic info encoded in nucleic acids translated into standard amino acids n Genetic code dictionary defining meaning for base sequence n Codon
Chapter 19 Overview Protein Synthesis n Protein Synthesis genetic info encoded in nucleic acids translated into standard amino acids n Genetic code dictionary defining meaning for base sequence n Codon
Genomic insights into high exopolysaccharide-producing dairy starter. bacterium Streptococcus thermophilus ASCC 1275
 Supplementary information of Genomic insights into high exopolysaccharide-producing dairy starter bacterium Streptococcus thermophilus ASCC 1275 Qinglong Wu, Hein Min Tun, Frederick Chi-Ching Leung, Nagendra
Supplementary information of Genomic insights into high exopolysaccharide-producing dairy starter bacterium Streptococcus thermophilus ASCC 1275 Qinglong Wu, Hein Min Tun, Frederick Chi-Ching Leung, Nagendra
Chapter 12. Genes: Expression and Regulation
 Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
REVIEW 1: BIOCHEMISTRY UNIT. A. Top 10 If you learned anything from this unit, you should have learned:
 Period Date REVIEW 1: BIOCHEMISTRY UNIT A. Top 10 If you learned anything from this unit, you should have learned: 1. All living matter made up of CHONPS 2. Bonds a. covalent bonds are strong b. hydrogen
Period Date REVIEW 1: BIOCHEMISTRY UNIT A. Top 10 If you learned anything from this unit, you should have learned: 1. All living matter made up of CHONPS 2. Bonds a. covalent bonds are strong b. hydrogen
Chapter 6: Outline-2. Chapter 6: Outline Properties of Enzymes. Introduction. Activation Energy, E act. Activation Energy-2
 Chapter 6: Outline- Properties of Enzymes Classification of Enzymes Enzyme inetics Michaelis-Menten inetics Lineweaver-Burke Plots Enzyme Inhibition Catalysis Catalytic Mechanisms Cofactors Chapter 6:
Chapter 6: Outline- Properties of Enzymes Classification of Enzymes Enzyme inetics Michaelis-Menten inetics Lineweaver-Burke Plots Enzyme Inhibition Catalysis Catalytic Mechanisms Cofactors Chapter 6:
Protein synthesis I Biochemistry 302. Bob Kelm February 23, 2004
 Protein synthesis I Biochemistry 302 Bob Kelm February 23, 2004 Key features of protein synthesis Energy glutton Essential metabolic activity of the cell. Consumes 90% of the chemical energy (ATP,GTP).
Protein synthesis I Biochemistry 302 Bob Kelm February 23, 2004 Key features of protein synthesis Energy glutton Essential metabolic activity of the cell. Consumes 90% of the chemical energy (ATP,GTP).
AP Bio-Ms.Bell Unit#3 Cellular Energies Name
 AP Bio-Ms.Bell Unit#3 Cellular Energies Name 1. Base your answer to the following question on the image below. 7. Base your answer to the following question on Which of the following choices correctly
AP Bio-Ms.Bell Unit#3 Cellular Energies Name 1. Base your answer to the following question on the image below. 7. Base your answer to the following question on Which of the following choices correctly
S1 Gene ontology (GO) analysis of the network alignment results
 1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
Activation of a receptor. Assembly of the complex
 Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Midterm Review Guide. Unit 1 : Biochemistry: 1. Give the ph values for an acid and a base. 2. What do buffers do? 3. Define monomer and polymer.
 Midterm Review Guide Name: Unit 1 : Biochemistry: 1. Give the ph values for an acid and a base. 2. What do buffers do? 3. Define monomer and polymer. 4. Fill in the Organic Compounds chart : Elements Monomer
Midterm Review Guide Name: Unit 1 : Biochemistry: 1. Give the ph values for an acid and a base. 2. What do buffers do? 3. Define monomer and polymer. 4. Fill in the Organic Compounds chart : Elements Monomer
CHAPTER4 Translation
 CHAPTER4 Translation 4.1 Outline of Translation 4.2 Genetic Code 4.3 trna and Anticodon 4.4 Ribosome 4.5 Protein Synthesis 4.6 Posttranslational Events 4.1 Outline of Translation From mrna to protein
CHAPTER4 Translation 4.1 Outline of Translation 4.2 Genetic Code 4.3 trna and Anticodon 4.4 Ribosome 4.5 Protein Synthesis 4.6 Posttranslational Events 4.1 Outline of Translation From mrna to protein
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Scale in the biological world
 Scale in the biological world 2 A cell seen by TEM 3 4 From living cells to atoms 5 Compartmentalisation in the cell: internal membranes and the cytosol 6 The Origin of mitochondria: The endosymbion hypothesis
Scale in the biological world 2 A cell seen by TEM 3 4 From living cells to atoms 5 Compartmentalisation in the cell: internal membranes and the cytosol 6 The Origin of mitochondria: The endosymbion hypothesis
Chapter 8 Metabolism: Energy, Enzymes, and Regulation
 Chapter 8 Metabolism: Energy, Enzymes, and Regulation Energy: Capacity to do work or cause a particular change. Thus, all physical and chemical processes are the result of the application or movement of
Chapter 8 Metabolism: Energy, Enzymes, and Regulation Energy: Capacity to do work or cause a particular change. Thus, all physical and chemical processes are the result of the application or movement of
Biology I Fall Semester Exam Review 2014
 Biology I Fall Semester Exam Review 2014 Biomolecules and Enzymes (Chapter 2) 8 questions Macromolecules, Biomolecules, Organic Compunds Elements *From the Periodic Table of Elements Subunits Monomers,
Biology I Fall Semester Exam Review 2014 Biomolecules and Enzymes (Chapter 2) 8 questions Macromolecules, Biomolecules, Organic Compunds Elements *From the Periodic Table of Elements Subunits Monomers,
Chapter 5: Photosynthesis: The Energy of Life pg : Pathways of Photosynthesis pg
 UNIT 2: Metabolic Processes Chapter 5: Photosynthesis: The Energy of Life pg. 210-240 5.2: Pathways of Photosynthesis pg. 220-228 Light Dependent Reactions Photosystem II and I are the two light capturing
UNIT 2: Metabolic Processes Chapter 5: Photosynthesis: The Energy of Life pg. 210-240 5.2: Pathways of Photosynthesis pg. 220-228 Light Dependent Reactions Photosystem II and I are the two light capturing
Proteome mapping of mature pollen of Arabidopsis thaliana
 4864 DOI 10.1002/pmic.200402011 Proteomics 2005, 5, 4864 4884 REGULAR ARTICLE Proteome mapping of mature pollen of Arabidopsis thaliana Rachel Holmes-Davis 1, Charlene K. Tanaka 2, William H. Vensel 2,
4864 DOI 10.1002/pmic.200402011 Proteomics 2005, 5, 4864 4884 REGULAR ARTICLE Proteome mapping of mature pollen of Arabidopsis thaliana Rachel Holmes-Davis 1, Charlene K. Tanaka 2, William H. Vensel 2,
Number of questions TEK (Learning Target) Biomolecules & Enzymes
 Unit Biomolecules & Enzymes Number of questions TEK (Learning Target) on Exam 8 questions 9A I can compare and contrast the structure and function of biomolecules. 9C I know the role of enzymes and how
Unit Biomolecules & Enzymes Number of questions TEK (Learning Target) on Exam 8 questions 9A I can compare and contrast the structure and function of biomolecules. 9C I know the role of enzymes and how
Pathways that Harvest and Store Chemical Energy
 6 Pathways that Harvest and Store Chemical Energy Energy is stored in chemical bonds and can be released and transformed by metabolic pathways. Chemical energy available to do work is termed free energy
6 Pathways that Harvest and Store Chemical Energy Energy is stored in chemical bonds and can be released and transformed by metabolic pathways. Chemical energy available to do work is termed free energy
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Gene regulation II Biochemistry 302. February 27, 2006
 Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
Protein synthesis II Biochemistry 302. Bob Kelm February 25, 2004
 Protein synthesis II Biochemistry 302 Bob Kelm February 25, 2004 Two idealized views of the 70S ribosomal complex during translation 70S cavity Fig. 27.25 50S tunnel View with 30S subunit in front, 50S
Protein synthesis II Biochemistry 302 Bob Kelm February 25, 2004 Two idealized views of the 70S ribosomal complex during translation 70S cavity Fig. 27.25 50S tunnel View with 30S subunit in front, 50S
GENETICS - CLUTCH CH.11 TRANSLATION.
 !! www.clutchprep.com CONCEPT: GENETIC CODE Nucleotides and amino acids are translated in a 1 to 1 method The triplet code states that three nucleotides codes for one amino acid - A codon is a term for
!! www.clutchprep.com CONCEPT: GENETIC CODE Nucleotides and amino acids are translated in a 1 to 1 method The triplet code states that three nucleotides codes for one amino acid - A codon is a term for
Assimilation of Carbon in Plants
 Module 0210101: Molecular Biology and Biochemistry of the Cell Lecture 18 Assimilation of Carbon in Plants Dale Sanders 17 March 2009 Objectives By the end of the lecture you should understand: 1. How
Module 0210101: Molecular Biology and Biochemistry of the Cell Lecture 18 Assimilation of Carbon in Plants Dale Sanders 17 March 2009 Objectives By the end of the lecture you should understand: 1. How
Molecular Biology (9)
 Molecular Biology (9) Translation Mamoun Ahram, PhD Second semester, 2017-2018 1 Resources This lecture Cooper, Ch. 8 (297-319) 2 General information Protein synthesis involves interactions between three
Molecular Biology (9) Translation Mamoun Ahram, PhD Second semester, 2017-2018 1 Resources This lecture Cooper, Ch. 8 (297-319) 2 General information Protein synthesis involves interactions between three
Biochemistry Prokaryotic translation
 1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 2. Understand the concept of genetic code 3. Understand the concept of wobble hypothesis
1 Description of Module Subject Name Paper Name Module Name/Title Dr. Vijaya Khader Dr. MC Varadaraj 2 1. Objectives 2. Understand the concept of genetic code 3. Understand the concept of wobble hypothesis
THYLAKOID GRANUM / GRANA
 THYLAKOID GRANUM / GRANA THYLAKOID GRANUM + STACKED THYLAKOID VESICLES THYLAKOID GRANUM THYLAKOID GRANUM + STACKED THYLAKOID VESICLES --- SITE: LIGHT RXT THYLAKOID GRANUM THYLAKOID GRANUM G STACKED THYLAKOID
THYLAKOID GRANUM / GRANA THYLAKOID GRANUM + STACKED THYLAKOID VESICLES THYLAKOID GRANUM THYLAKOID GRANUM + STACKED THYLAKOID VESICLES --- SITE: LIGHT RXT THYLAKOID GRANUM THYLAKOID GRANUM G STACKED THYLAKOID
Laith AL-Mustafa. Protein synthesis. Nabil Bashir 10\28\ First
 Laith AL-Mustafa Protein synthesis Nabil Bashir 10\28\2015 http://1drv.ms/1gigdnv 01 First 0 Protein synthesis In previous lectures we started talking about DNA Replication (DNA synthesis) and we covered
Laith AL-Mustafa Protein synthesis Nabil Bashir 10\28\2015 http://1drv.ms/1gigdnv 01 First 0 Protein synthesis In previous lectures we started talking about DNA Replication (DNA synthesis) and we covered
Biochemical and Biophysical Research Communications 337 (2005)
 Biochemical and Biophysical Research Communications 337 (2005) 1257 1266 BBRC www.elsevier.com/locate/ybbrc A reference map of the Arabidopsis thaliana mature pollen proteome q Sandra Noir, Anne Bräutigam,
Biochemical and Biophysical Research Communications 337 (2005) 1257 1266 BBRC www.elsevier.com/locate/ybbrc A reference map of the Arabidopsis thaliana mature pollen proteome q Sandra Noir, Anne Bräutigam,
Reading Assignments. A. Genes and the Synthesis of Polypeptides. Lecture Series 7 From DNA to Protein: Genotype to Phenotype
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
f) Adding an enzyme does not change the Gibbs free energy. It only increases the rate of the reaction by lowering the activation energy.
 Problem Set 2-Answer Key BILD1 SP16 1) How does an enzyme catalyze a chemical reaction? Define the terms and substrate and active site. An enzyme lowers the energy of activation so the reaction proceeds
Problem Set 2-Answer Key BILD1 SP16 1) How does an enzyme catalyze a chemical reaction? Define the terms and substrate and active site. An enzyme lowers the energy of activation so the reaction proceeds
Turns sunlight, water & carbon dioxide (CO 2 ) into sugar & oxygen through photosynthesis
 CELL PART/ ORGANELLE FUNCTION (what it does) PICTURE Plant, Animal, or Both Cell Membrane controls what goes in & out of the cell protects the cell Nucleus directs all the cell s activities contains cell
CELL PART/ ORGANELLE FUNCTION (what it does) PICTURE Plant, Animal, or Both Cell Membrane controls what goes in & out of the cell protects the cell Nucleus directs all the cell s activities contains cell
Translation. Genetic code
 Translation Genetic code If genes are segments of DNA and if DNA is just a string of nucleotide pairs, then how does the sequence of nucleotide pairs dictate the sequence of amino acids in proteins? Simple
Translation Genetic code If genes are segments of DNA and if DNA is just a string of nucleotide pairs, then how does the sequence of nucleotide pairs dictate the sequence of amino acids in proteins? Simple
Harvesting energy: photosynthesis & cellular respiration part 1
 Harvesting energy: photosynthesis & cellular respiration part 1 Agenda I. Overview (Big Pictures) of Photosynthesis & Cellular Respiration II. Making Glucose - Photosynthesis III. Making ATP - Cellular
Harvesting energy: photosynthesis & cellular respiration part 1 Agenda I. Overview (Big Pictures) of Photosynthesis & Cellular Respiration II. Making Glucose - Photosynthesis III. Making ATP - Cellular
9 The Process of Translation
 9 The Process of Translation 9.1 Stages of Translation Process We are familiar with the genetic code, we can begin to study the mechanism by which amino acids are assembled into proteins. Because more
9 The Process of Translation 9.1 Stages of Translation Process We are familiar with the genetic code, we can begin to study the mechanism by which amino acids are assembled into proteins. Because more
DNA Technology, Bacteria, Virus and Meiosis Test REVIEW
 Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
Supporting Information
 Supporting Information Proteomic Analyses of Cysteine Redox in High-fat-fed and Fasted Mouse Livers: Implications for Liver Metabolic Homeostasis Yixing Li 1#, Zupeng Luo 1#, Xilong Wu 2, Jun Zhu 2, Kai
Supporting Information Proteomic Analyses of Cysteine Redox in High-fat-fed and Fasted Mouse Livers: Implications for Liver Metabolic Homeostasis Yixing Li 1#, Zupeng Luo 1#, Xilong Wu 2, Jun Zhu 2, Kai
Translation. A ribosome, mrna, and trna.
 Translation The basic processes of translation are conserved among prokaryotes and eukaryotes. Prokaryotic Translation A ribosome, mrna, and trna. In the initiation of translation in prokaryotes, the Shine-Dalgarno
Translation The basic processes of translation are conserved among prokaryotes and eukaryotes. Prokaryotic Translation A ribosome, mrna, and trna. In the initiation of translation in prokaryotes, the Shine-Dalgarno
Transport between cytosol and nucleus
 of 60 3 Gated trans Lectures 9-15 MBLG 2071 The n GATED TRANSPORT transport between cytoplasm and nucleus (bidirectional) controlled by the nuclear pore complex active transport for macro molecules e.g.
of 60 3 Gated trans Lectures 9-15 MBLG 2071 The n GATED TRANSPORT transport between cytoplasm and nucleus (bidirectional) controlled by the nuclear pore complex active transport for macro molecules e.g.
PETER PAZMANY CATHOLIC UNIVERSITY Consortium members SEMMELWEIS UNIVERSITY, DIALOG CAMPUS PUBLISHER
 PETER PAZMANY SEMMELWEIS CATHOLIC UNIVERSITY UNIVERSITY Development of Complex Curricula for Molecular Bionics and Infobionics Programs within a consortial* framework** Consortium leader PETER PAZMANY
PETER PAZMANY SEMMELWEIS CATHOLIC UNIVERSITY UNIVERSITY Development of Complex Curricula for Molecular Bionics and Infobionics Programs within a consortial* framework** Consortium leader PETER PAZMANY
Table S1. 10μM_rfp_B1 2,074, ohr 10μM_rfp_C2 2,074, ohr
 Table S1 Selection without RpoD Insert Start Position Insert End Position Potential Tolerance Gene Candidates 0μM_rfp_A1 2,076,184 -- ohr 0μM_rfp_B1 2,082,864 -- ohr 0μM_rfp_C1 2,076,798 -- ohr 1μM_rfp_A1
Table S1 Selection without RpoD Insert Start Position Insert End Position Potential Tolerance Gene Candidates 0μM_rfp_A1 2,076,184 -- ohr 0μM_rfp_B1 2,082,864 -- ohr 0μM_rfp_C1 2,076,798 -- ohr 1μM_rfp_A1
AQA Biology A-level Topic 5: Energy transfers in and between organisms
 AQA Biology A-level Topic 5: Energy transfers in and between organisms Notes Photosynthesis Photosynthesis is a reaction in which light energy is used to produce glucose in plants. The process requires
AQA Biology A-level Topic 5: Energy transfers in and between organisms Notes Photosynthesis Photosynthesis is a reaction in which light energy is used to produce glucose in plants. The process requires
PHOTOSYNTHESIS. The Details
 PHOTOSYNTHESIS The Details Photosynthesis is divided into 2 sequential processes: 1. The Light Dependent Reactions (stages 1 & 2) 2. The Light Independent Reactions (stage 3) a.k.a. the Calvin Cycle THE
PHOTOSYNTHESIS The Details Photosynthesis is divided into 2 sequential processes: 1. The Light Dependent Reactions (stages 1 & 2) 2. The Light Independent Reactions (stage 3) a.k.a. the Calvin Cycle THE
2/25/2013. Electronic Configurations
 1 2 3 4 5 Chapter 2 Chemical Principles The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions
1 2 3 4 5 Chapter 2 Chemical Principles The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions
Mean Hes score. Threshold
 Mean Hes score 0.1 0.09 Common 0.08 Family 0.07 0.06 0.05 0.04 0.03 0.02 0.01 0 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 Threshold SUPPLEMENTARY FIG. 1. Trend of mean Hes scores calculated based on the
Mean Hes score 0.1 0.09 Common 0.08 Family 0.07 0.06 0.05 0.04 0.03 0.02 0.01 0 0 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 Threshold SUPPLEMENTARY FIG. 1. Trend of mean Hes scores calculated based on the
Gene regulation II Biochemistry 302. Bob Kelm February 28, 2005
 Gene regulation II Biochemistry 302 Bob Kelm February 28, 2005 Catabolic operons: Regulation by multiple signals targeting different TFs Catabolite repression: Activity of lac operon is restricted when
Gene regulation II Biochemistry 302 Bob Kelm February 28, 2005 Catabolic operons: Regulation by multiple signals targeting different TFs Catabolite repression: Activity of lac operon is restricted when
Biophysics Lectures Three and Four
 Biophysics Lectures Three and Four Kevin Cahill cahill@unm.edu http://dna.phys.unm.edu/ 1 The Atoms and Molecules of Life Cells are mostly made from the most abundant chemical elements, H, C, O, N, Ca,
Biophysics Lectures Three and Four Kevin Cahill cahill@unm.edu http://dna.phys.unm.edu/ 1 The Atoms and Molecules of Life Cells are mostly made from the most abundant chemical elements, H, C, O, N, Ca,
1 GO: regulation of cell size E-04 2 GO: negative regulation of cell growth GO:
 Table S2: The biological modulated by mir-5701 Sr. No Term Id 1 Term Name 2 Hit Gene Number 3 P-Value 4 1 GO:0008361 regulation of cell size 9 4.37E-04 2 GO:0030308 negative regulation of cell growth 8
Table S2: The biological modulated by mir-5701 Sr. No Term Id 1 Term Name 2 Hit Gene Number 3 P-Value 4 1 GO:0008361 regulation of cell size 9 4.37E-04 2 GO:0030308 negative regulation of cell growth 8
Ch 7: Cell Structure and Functions. AP Biology
 Ch 7: Cell Structure and Functions AP Biology The Cell Theory 1. All living things are made of cells. 2. New cells come from existing cells. 3. Cells are the basic units of structure and function of living
Ch 7: Cell Structure and Functions AP Biology The Cell Theory 1. All living things are made of cells. 2. New cells come from existing cells. 3. Cells are the basic units of structure and function of living
PHOTOSYNTHESIS: A BRIEF STORY!!!!!
 PHOTOSYNTHESIS: A BRIEF STORY!!!!! This is one of the most important biochemical processes in plants and is amongst the most expensive biochemical processes in plant in terms of investment. Photosynthesis
PHOTOSYNTHESIS: A BRIEF STORY!!!!! This is one of the most important biochemical processes in plants and is amongst the most expensive biochemical processes in plant in terms of investment. Photosynthesis
2011 The Simple Homeschool Simple Days Unit Studies Cells
 1 We have a full line of high school biology units and courses at CurrClick and as online courses! Subscribe to our interactive unit study classroom and make science fun and exciting! 2 A cell is a small
1 We have a full line of high school biology units and courses at CurrClick and as online courses! Subscribe to our interactive unit study classroom and make science fun and exciting! 2 A cell is a small
From Gene to Protein
 From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
SUMMARY Chlorophyll biosynthesis:
 Summary SUMMARY Plants are severely affected by various abiotic stresses due to their static nature. Abiotic stresses may be in the form of drought, salt, flooding, temperature, light or metal stress.
Summary SUMMARY Plants are severely affected by various abiotic stresses due to their static nature. Abiotic stresses may be in the form of drought, salt, flooding, temperature, light or metal stress.
Study Guide: Fall Final Exam H O N O R S B I O L O G Y : U N I T S 1-5
 Study Guide: Fall Final Exam H O N O R S B I O L O G Y : U N I T S 1-5 Directions: The list below identifies topics, terms, and concepts that will be addressed on your Fall Final Exam. This list should
Study Guide: Fall Final Exam H O N O R S B I O L O G Y : U N I T S 1-5 Directions: The list below identifies topics, terms, and concepts that will be addressed on your Fall Final Exam. This list should
Lecture 10 (10/4/17) Lecture 10 (10/4/17)
 Lecture 10 (10/4/17) Reading: Ch4; 125, 138-141, 141-142 Problems: Ch4 (text); 7, 9, 11 Ch4 (study guide); 1, 2 NEXT Reading: Ch4; 125, 132-136 (structure determination) Ch4; 12-130 (Collagen) Problems:
Lecture 10 (10/4/17) Reading: Ch4; 125, 138-141, 141-142 Problems: Ch4 (text); 7, 9, 11 Ch4 (study guide); 1, 2 NEXT Reading: Ch4; 125, 132-136 (structure determination) Ch4; 12-130 (Collagen) Problems:
Organelles & Cells Student Edition. A. chromosome B. gene C. mitochondrion D. vacuole
 Name: Date: 1. Which structure is outside the nucleus of a cell and contains DNA? A. chromosome B. gene C. mitochondrion D. vacuole 2. A potato core was placed in a beaker of water as shown in the figure
Name: Date: 1. Which structure is outside the nucleus of a cell and contains DNA? A. chromosome B. gene C. mitochondrion D. vacuole 2. A potato core was placed in a beaker of water as shown in the figure
Types of RNA. 1. Messenger RNA(mRNA): 1. Represents only 5% of the total RNA in the cell.
 RNAs L.Os. Know the different types of RNA & their relative concentration Know the structure of each RNA Understand their functions Know their locations in the cell Understand the differences between prokaryotic
RNAs L.Os. Know the different types of RNA & their relative concentration Know the structure of each RNA Understand their functions Know their locations in the cell Understand the differences between prokaryotic
Biochemistry 3300 Problems (and Solutions) Metabolism I
 (1) Provide a reasonable systematic name for an enzyme that catalyzes the following reaction: fructose + ATP > fructose-1 phosphate + ADP (2) The IUBMB has a developed a set of rules for classifying enzymes
(1) Provide a reasonable systematic name for an enzyme that catalyzes the following reaction: fructose + ATP > fructose-1 phosphate + ADP (2) The IUBMB has a developed a set of rules for classifying enzymes
GACE Biology Assessment Test I (026) Curriculum Crosswalk
 Subarea I. Cell Biology: Cell Structure and Function (50%) Objective 1: Understands the basic biochemistry and metabolism of living organisms A. Understands the chemical structures and properties of biologically
Subarea I. Cell Biology: Cell Structure and Function (50%) Objective 1: Understands the basic biochemistry and metabolism of living organisms A. Understands the chemical structures and properties of biologically
Photosynthesis: Carbon Reactions. Dr. Obaidur Rahman
 Photosynthesis: Carbon Reactions Dr. Obaidur Rahman Topics: Part I THE CALVIN CYCLE The Calvin Cycle Has Three Stages: Carboxylation, Reduction, and Regeneration The Carboxylation of Ribulose Bisphosphate
Photosynthesis: Carbon Reactions Dr. Obaidur Rahman Topics: Part I THE CALVIN CYCLE The Calvin Cycle Has Three Stages: Carboxylation, Reduction, and Regeneration The Carboxylation of Ribulose Bisphosphate
1 (a) Fig. 1.1 is a diagram representing a three-dimensional view of a chloroplast. space B. Fig (i) Name parts A to C in Fig A... B...
 1 (a) Fig. 1.1 is a diagram representing a three-dimensional view of a chloroplast. A space B C Fig. 1.1 (i) Name parts A to C in Fig. 1.1. A... B... C... [3] (ii) Describe two ways in which the structure
1 (a) Fig. 1.1 is a diagram representing a three-dimensional view of a chloroplast. A space B C Fig. 1.1 (i) Name parts A to C in Fig. 1.1. A... B... C... [3] (ii) Describe two ways in which the structure
Chapter 2 The Chemistry of Biology. Dr. Ramos BIO 370
 Chapter 2 The Chemistry of Biology Dr. Ramos BIO 370 2 Atoms, Bonds, and Molecules Matter - all materials that occupy space and have mass Matter is composed of atoms. Atom simplest form of matter not divisible
Chapter 2 The Chemistry of Biology Dr. Ramos BIO 370 2 Atoms, Bonds, and Molecules Matter - all materials that occupy space and have mass Matter is composed of atoms. Atom simplest form of matter not divisible
