Design of microarray experiments
|
|
|
- Egbert Long
- 5 years ago
- Views:
Transcription
1 Design of microarray experiments Ulrich Mansmann Practical microarray analysis March 23 Heidelberg Heidelberg, March 23
2 Experiments Scientists deal mostly with experiments of the following form: A number of alternative conditions / treatments one of which is applied to each experimental unit an observation (or several observations) then being made on each unit. The objective is: Separate out differences between the conditions / treatments from the uncontrolled variation that is assumed to be present. Take steps towards understanding the phenomena under investigation. Heidelberg, March 23 2
3 Statistical thinking Uncertain knowledge Knowledge of the extent of uncertainty in it + = Useable knowledge Measurement model m = µ + e m measurement with error, µ - true but unknown value What is the mean of e? What is the variance of e? Is there dependence between e and µ? What is the distribution of e (and µ)? Typically but not always: e ~ N(,σ²) Gaussian / Normal measurement model Decisions on the experimental design influence the measurement model. Heidelberg, March 23 3
4 Main requirements for experiments Once the conditions / treatments, experimental units, and the nature of the observations have been fixed, the main requirements are: Experimental units receiving different treatments should differ in no systematic way from one another Assumptions that certain sources of variation are absent or negligible should, as far as practical, be avoided; Random errors of estimation should be suitably small, and this should be achieved with as few experimental units as possible; The conclusions of the experiment should have a wide range of validity; The experiment should be simple in design and analysis; A proper statistical analysis of the results should be possible without making artificial assumptions. Taken from Cox DR (958) Planning of experiments, Wiley & Sons, New York (page 3) Heidelberg, March 23 4
5 The most simple measurement model in microarray experiments Situation: m arrays (Affimetrix) from control population n arrays (Affimetrix) from population with special condition /treatment Observation of interest: Mean difference of log-transformed gene expression ( logfc) logfc obs = logfc true + e e ~ N(, σ² [/n+/m]) In an experiment with 5 arrays per population and the same variance for the expression of a gene of interest, the above formula implies that the variance of the logfc is only 4% (/5+/5 = 2/5 =.4) of the variability of a single measurement taming of uncertainty. Heidelberg, March 23 5
6 Separate out differences between the conditions / treatments from the uncontrolled variation that is assumed to be present. Is logfc true? How to decide? Special Decision rules: Statistical Tests When the probability model for the mechanism generating the observed data is known, hypotheses about the model can be tested. This involves the question: Could the presented data reasonable have come from the model if the hypothesis is correct? Usually a decision must be made on the basis of the available data, and some degree of uncertainty is tolerated about the correctness of that decision. These four components: data, model, hypothesis, and decision are basic to the statistical problem of hypothesis testing. Heidelberg, March 23 6
7 Quality of decision True state of gene Decision Gene is diff. expr. Gene is not diff. expr. Gene is diff. expr. OK false positive decision happens with probability α Gene is not diff. expr. OK false negative decision happens with probability β Two sources of error: Power of a test: False positive rate α False negative rate β Ability to detect a difference if there is a true difference Power true positive rate or Power = - β Heidelberg, March 23 7
8 Question of interest (Alterative): The Statistical test Is the gene G differentially expressed between two cell populations? Answer the question via a proof by contradiction: Show that there is no evidence to support the logical contrary of the alternative. The logical contrary of the alternative is called null hypothesis. Null hypothesis: The gene G is not differentially expressed between two cell populations of interest. A test statistic T is introduced which measures the fit of the observed data to the null hypothesis. The test statistics T implies a prob. distribution P to quantify its variability when the null hypothesis is true. It will be checked if the test statistic evaluated at the observed data t obs behaves typically (not extreme) with respect to the test distribution. The p-value is the probability under the null hypothesis of an observation which is more extreme as the observation given by the data: P( T t obs ) = p. A criteria is needed to asses extreme behaviour of the test statistic via the p value which is called the level of the test: a. The observed data does not fit to the null hypothesis if p < a or t obs > t* where t* is the -α or -α/2 quantile of the prob. distribution P. t* is also called the critical value. The conditions p < a and t obs > t* are equivalent. If p < a or t obs > t* the null hypothesis will be rejected. If p a or t obs t* the null hypothesis can not be rejected this does not mean that it is true Absence of evidence for a difference is no evidence for an absence of difference. Heidelberg, March 23 8
9 Controlling the power sample size calculations The test should produce a significant result (level α) with a power of -β if logfc true = δ null hypothesis: logfc true = alternative: logfc true = δ δ z -α/2 σ n,m z -β σ n,m σ 2 n,m = σ 2 (/n+/m) The above requirement is fulfilled if: δ = (z -α/2 + z -β ) σ n,m or 2 n m (z α / 2 + z β) = n + m δ 2 σ 2 Heidelberg, March 23 9
10 Controlling the power sample size calculations 2 n m (z α / 2 + z β) = n + m δ 2 σ 2 n = N γ and m = N (-γ) with M total size of experiment and γ ],[ 2 (z α / 2 + z β ) N = γ ( γ) 2 δ 2 σ The size of the experiment is minimal if g = ½ Gamma Heidelberg, March 23
11 Sample size calculation for a microarray experiment I Truth Test result diff. expr. (H ) not diff. expr. (H ) diff. expr. D D D not diff. expr. U U U Number of genes on array G G G α = E[D ]/G β = E[U ]/G FDR=E[D /D] E: expectation / mean number family type I error probability: α F = P[D >] family type II error probability: β F = P[U >] Heidelberg, March 23
12 Sample size calculation for a microarray experiment II Independent genes Dependent Genes P[D =] = (-α ) Go = -α F D ~ inomial(g, α ) E[D ] = G α Poissonapprox.: E[D ] ~ -ln(-α F ) P[U =] = (-β ) G = -β F E[U ] = G (-β ) onferroni: α = α F / G No direct link between the probability for D and α F. -β F max{,- G β } No direct link between the probability for U and β F. Heidelberg, March 23 2
13 Sample size calculation for a microarray experiment III for an array with 33 independent genes What are useful α and β? α F =.8 E[D ] = - ln(-.8) =.6 = λ P(exactly k false pos.) = exp(-λ) λ k / (k!) false pos Prob P(at least six false positives) = unexpressed genes: α =.6/325 = expressed genes, set E[D ] = 45 -β = 45/5 =.9 β =. -β F = (-β ) G < -23 E[FDR] =.35 95% quantile of FDR:.89 (calculated by simulation) Heidelberg, March 23 3
14 Sample size calculation for a microarray experiment IV In order to complete the sample size calculation for a microarray experiment, information on σ 2 is needed. The size of the experiment, N, needed to detect a logfc true of δ on a significance level α and with power -β is: 2 2 (z α / 2 + z β ) σ N = 4 2 δ In a similar set of experiments σ 2 for a set of 2 VSN transformed arrays was between.55 and.85. One may choose the value σ 2 = 2. δ log(.5) log(2) log(3) log(5) log() N (σ 2 = 2) N (σ 2 = ) Sample size with α =.495, β =. Heidelberg, March 23 4
15 Sample size formula for a one group test The test should produce a significant result (level α) with a power of -β if T = δ null hypothesis: T = alternative: T = δ δ z -α/2 σ n z -β σ n σ 2 n = σ 2 /n The above requirement is fulfilled if: δ = (z -α/2 + z -β ) σ n or 2 2 (z α / 2 + z β ) σ n = 2 δ Heidelberg, March 23 5
16 Measurement model for cdna arrays Gene expression under condition A intensity of red colour, Gene expression under condition intensity of green colour Measurement: m A/ = I red,a 2 I green, Log = γ A/ + δ + e γ A/ log-transformed true fold change of gene of condition A with respect to condition δ - dye effect, e measurement error with E[e] = and Var(e) = σ 2 Measurement m A/ is used to estimate unknown γ A/ Vertices Edges mrna samples hybridization Direction Dye assignment Green Red A Heidelberg, March 23 6
17 Estimation of log fold change g A/ Reference Design Dye swap design A A R R A/ Estimate of γ A/ DS g = m A/R m /R g A/ = (m A/ m /A )/2 Variability of estimate g ) = 2 σ 2 Var( g DS A/ ) =.5 σ 2 Sample Size increases proportional to the variance of the measurement! Var( R A/ Heidelberg, March 23 7
18 2x2 factorial experiments I treatment / condition Wild type Mutation before treatment β β+µ after treatment β+τ β+τ+µ+ψ β - baseline effect; τ - effect of treatment; µ - effect of mutation ψ - differential effect on treatment between WT and MUT treatment effect on gene expr. in WT cells: WT = (β + τ) - β = τ treatment effect on gene expr. in MUT cells: MUT = (β + τ + µ + ψ) (β + µ)= τ + ψ differential treatment effect: MUT WT or ψ How many cdna arrays are needed to show ψ with significance α and power -β if ψ > ln(5)? Heidelberg, March 23 8
19 2x2 factorial experiments II Study the joint effect of two conditions / treatment, A and, on the gene expression of a cell population of interest. There are four possible condition / treatment combinations: A: treatment applied to MUT cells A: treatment applied to WT cells : no treatment applied to MUT cells : no treatment applied to WT cells A A Design with 2 slides Heidelberg, March 23 9
20 2x2 factorial experiments III Array m A/ m /A m / m / m A/ m /A m A/A m A/A m A/ m /A m A/ m /A Measurement γ A/ + δ + e = τ + δ + e -γ A/ + δ + e = -τ + δ + e γ / + δ + e = µ + δ + e -γ / + δ + e = -µ + δ + e γ A/ + δ + e = µ + τ + ψ + δ + e -γ A/ + δ + e = - (µ + τ + ψ) + δ + e γ A/A + δ + e = µ + ψ + δ + e -γ A/A + δ + e = - (µ + ψ) + δ + e γ A/ + δ + e = µ + ψ + δ + e -γ A/ + δ + e = - (µ + ψ) + δ + e γ A/ + δ + e = τ - µ + δ + e -γ A/ + δ + e = - (τ - µ) + δ + e Each measurement has variance σ 2 Parameter β is confounded with the dye effect Heidelberg, March 23 2
21 Heidelberg, March 23 2 Regression analysis ψ µ τ δ = M M M M M M M M M M M M E A / A / A / A / A A / A A / A / A / / / A / A / A A For parameter θ = (δ, τ, µ, ψ) define the design matrix X such that E(M) = Xθ. For each gene, compute least square estimate θ* = (X X) - X M (LUE) Obtain measures of precision of estimated effects. Use all possibilities of the theory of linear models. Design problem: Each measurement M is made with variability σ 2. How precise can we estimate the components or contrasts of θ? Answer: Look at (X X) -
22 A 2 x 2 factorial designs IV A total.2.by.2.design.mat delta alpha beta psi > precision.2.by.2.rfc(x.mat) A/ $inv.mat /A - / tau mu psi / - tau A/ mu /A psi A/A A/A - - $effects A/ tau mu psi tau-mu /A /A - A/ - Var(A-) = Var(A) + Var() 2 Cov(A,) Heidelberg, March 23 22
23 Sample size for differential treatment effect (DTE) in a 2 x 2 factorial designs I Array has 2. genes: 95 without DTE, 5 with DTE α F =.9, using onferroni adjustment: α =.9/2. =.462 Mean number of correct positives is set to 45: -β =.9 σ 2 =.7, taken from similar experiments A total dye swap design (2 arrays) estimates ψ with precision σ 2 /2 =.35 N = [ ] 2.35 / ln(5) 2 = The experiment would need in total 4 x 2 = 48 arrays Is there a chance to get the same result cheaper? Heidelberg, March 23 23
24 2 x 2 factorial designs V Design I Common ref. Design II Common ref. Design III Connected Design IV Connected A A A A A A A A A Design V All-pairs A Scaled variances of estimated effects D.I D.II D.III D.IV D.V D.tot tau mu psi # chips Heidelberg, March 23 24
25 Sample size for differential treatment effect (DTE) in a 2 x 2 factorial designs II Is there a chance to get the same result cheaper? Using total dye swap design, the experiment would need in total 4 x 2 = 48 arrays Using Design III, the effect of interest is estimated with doubled variance (4 8) but by using a design which need only 4 arrays (2 4). This reduces the number of arrays needed from 48 to 32. Heidelberg, March 23 25
26 Designs for time course experiments Experimental Design - Conclusions In addition to experimental constraints, design decisions should be guided by knowledge of which effects are of greater interest to the investigator. The unrealistic planning based on independent genes may be put into a more realistic framework by using simulation studies speak to your bio statistician/informatician How to collect and present experience from performed microarray experiments on which to base assumptions for planing (σ 2 )? Further reading: Kerr MK, Churchill GA (2) Experimental design for gene expression microarrays, iostatistics, 2:83-2 Lee MLT, Whitmore GA (22), Power and sample size for DNA microarray studies, Stat. in Med., 2: Heidelberg, March 23 26
Design of microarray experiments
 Design of microarray experiments Ulrich ansmann mansmann@imbi.uni-heidelberg.de Practical microarray analysis September Heidelberg Heidelberg, September otivation The lab biologist and theoretician need
Design of microarray experiments Ulrich ansmann mansmann@imbi.uni-heidelberg.de Practical microarray analysis September Heidelberg Heidelberg, September otivation The lab biologist and theoretician need
Design of Microarray Experiments. Xiangqin Cui
 Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Chapter 3: Statistical methods for estimation and testing. Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001).
 Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
LECTURE 5. Introduction to Econometrics. Hypothesis testing
 LECTURE 5 Introduction to Econometrics Hypothesis testing October 18, 2016 1 / 26 ON TODAY S LECTURE We are going to discuss how hypotheses about coefficients can be tested in regression models We will
LECTURE 5 Introduction to Econometrics Hypothesis testing October 18, 2016 1 / 26 ON TODAY S LECTURE We are going to discuss how hypotheses about coefficients can be tested in regression models We will
Two-Color Microarray Experimental Design Notation. Simple Examples of Analysis for a Single Gene. Microarray Experimental Design Notation
 Simple Examples of Analysis for a Single Gene wo-olor Microarray Experimental Design Notation /3/0 opyright 0 Dan Nettleton Microarray Experimental Design Notation Microarray Experimental Design Notation
Simple Examples of Analysis for a Single Gene wo-olor Microarray Experimental Design Notation /3/0 opyright 0 Dan Nettleton Microarray Experimental Design Notation Microarray Experimental Design Notation
Probability Methods in Civil Engineering Prof. Dr. Rajib Maity Department of Civil Engineering Indian Institution of Technology, Kharagpur
 Probability Methods in Civil Engineering Prof. Dr. Rajib Maity Department of Civil Engineering Indian Institution of Technology, Kharagpur Lecture No. # 36 Sampling Distribution and Parameter Estimation
Probability Methods in Civil Engineering Prof. Dr. Rajib Maity Department of Civil Engineering Indian Institution of Technology, Kharagpur Lecture No. # 36 Sampling Distribution and Parameter Estimation
Hypothesis testing (cont d)
 Hypothesis testing (cont d) Ulrich Heintz Brown University 4/12/2016 Ulrich Heintz - PHYS 1560 Lecture 11 1 Hypothesis testing Is our hypothesis about the fundamental physics correct? We will not be able
Hypothesis testing (cont d) Ulrich Heintz Brown University 4/12/2016 Ulrich Heintz - PHYS 1560 Lecture 11 1 Hypothesis testing Is our hypothesis about the fundamental physics correct? We will not be able
Tests about a population mean
 October 2 nd, 2017 Overview Week 1 Week 2 Week 4 Week 7 Week 10 Week 12 Chapter 1: Descriptive statistics Chapter 6: Statistics and Sampling Distributions Chapter 7: Point Estimation Chapter 8: Confidence
October 2 nd, 2017 Overview Week 1 Week 2 Week 4 Week 7 Week 10 Week 12 Chapter 1: Descriptive statistics Chapter 6: Statistics and Sampling Distributions Chapter 7: Point Estimation Chapter 8: Confidence
Biochip informatics-(i)
 Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
STAT 515 fa 2016 Lec Statistical inference - hypothesis testing
 STAT 515 fa 2016 Lec 20-21 Statistical inference - hypothesis testing Karl B. Gregory Wednesday, Oct 12th Contents 1 Statistical inference 1 1.1 Forms of the null and alternate hypothesis for µ and p....................
STAT 515 fa 2016 Lec 20-21 Statistical inference - hypothesis testing Karl B. Gregory Wednesday, Oct 12th Contents 1 Statistical inference 1 1.1 Forms of the null and alternate hypothesis for µ and p....................
Introductory Econometrics. Review of statistics (Part II: Inference)
 Introductory Econometrics Review of statistics (Part II: Inference) Jun Ma School of Economics Renmin University of China October 1, 2018 1/16 Null and alternative hypotheses Usually, we have two competing
Introductory Econometrics Review of statistics (Part II: Inference) Jun Ma School of Economics Renmin University of China October 1, 2018 1/16 Null and alternative hypotheses Usually, we have two competing
Hypothesis testing. Data to decisions
 Hypothesis testing Data to decisions The idea Null hypothesis: H 0 : the DGP/population has property P Under the null, a sample statistic has a known distribution If, under that that distribution, the
Hypothesis testing Data to decisions The idea Null hypothesis: H 0 : the DGP/population has property P Under the null, a sample statistic has a known distribution If, under that that distribution, the
Mathematical statistics
 November 1 st, 2018 Lecture 18: Tests about a population mean Overview 9.1 Hypotheses and test procedures test procedures errors in hypothesis testing significance level 9.2 Tests about a population mean
November 1 st, 2018 Lecture 18: Tests about a population mean Overview 9.1 Hypotheses and test procedures test procedures errors in hypothesis testing significance level 9.2 Tests about a population mean
Expression arrays, normalization, and error models
 1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
A Sequential Bayesian Approach with Applications to Circadian Rhythm Microarray Gene Expression Data
 A Sequential Bayesian Approach with Applications to Circadian Rhythm Microarray Gene Expression Data Faming Liang, Chuanhai Liu, and Naisyin Wang Texas A&M University Multiple Hypothesis Testing Introduction
A Sequential Bayesian Approach with Applications to Circadian Rhythm Microarray Gene Expression Data Faming Liang, Chuanhai Liu, and Naisyin Wang Texas A&M University Multiple Hypothesis Testing Introduction
Section 10.1 (Part 2 of 2) Significance Tests: Power of a Test
 1 Section 10.1 (Part 2 of 2) Significance Tests: Power of a Test Learning Objectives After this section, you should be able to DESCRIBE the relationship between the significance level of a test, P(Type
1 Section 10.1 (Part 2 of 2) Significance Tests: Power of a Test Learning Objectives After this section, you should be able to DESCRIBE the relationship between the significance level of a test, P(Type
Introduction to Statistical Hypothesis Testing
 Introduction to Statistical Hypothesis Testing Arun K. Tangirala Power of Hypothesis Tests Arun K. Tangirala, IIT Madras Intro to Statistical Hypothesis Testing 1 Learning objectives I Computing Pr(Type
Introduction to Statistical Hypothesis Testing Arun K. Tangirala Power of Hypothesis Tests Arun K. Tangirala, IIT Madras Intro to Statistical Hypothesis Testing 1 Learning objectives I Computing Pr(Type
Linear Models and Empirical Bayes Methods for. Assessing Differential Expression in Microarray Experiments
 Linear Models and Empirical Bayes Methods for Assessing Differential Expression in Microarray Experiments by Gordon K. Smyth (as interpreted by Aaron J. Baraff) STAT 572 Intro Talk April 10, 2014 Microarray
Linear Models and Empirical Bayes Methods for Assessing Differential Expression in Microarray Experiments by Gordon K. Smyth (as interpreted by Aaron J. Baraff) STAT 572 Intro Talk April 10, 2014 Microarray
Bio 183 Statistics in Research. B. Cleaning up your data: getting rid of problems
 Bio 183 Statistics in Research A. Research designs B. Cleaning up your data: getting rid of problems C. Basic descriptive statistics D. What test should you use? What is science?: Science is a way of knowing.(anon.?)
Bio 183 Statistics in Research A. Research designs B. Cleaning up your data: getting rid of problems C. Basic descriptive statistics D. What test should you use? What is science?: Science is a way of knowing.(anon.?)
The regression model with one fixed regressor cont d
 The regression model with one fixed regressor cont d 3150/4150 Lecture 4 Ragnar Nymoen 27 January 2012 The model with transformed variables Regression with transformed variables I References HGL Ch 2.8
The regression model with one fixed regressor cont d 3150/4150 Lecture 4 Ragnar Nymoen 27 January 2012 The model with transformed variables Regression with transformed variables I References HGL Ch 2.8
Parametric Empirical Bayes Methods for Microarrays
 Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Resampling and the Bootstrap
 Resampling and the Bootstrap Axel Benner Biostatistics, German Cancer Research Center INF 280, D-69120 Heidelberg benner@dkfz.de Resampling and the Bootstrap 2 Topics Estimation and Statistical Testing
Resampling and the Bootstrap Axel Benner Biostatistics, German Cancer Research Center INF 280, D-69120 Heidelberg benner@dkfz.de Resampling and the Bootstrap 2 Topics Estimation and Statistical Testing
6.4 Type I and Type II Errors
 6.4 Type I and Type II Errors Ulrich Hoensch Friday, March 22, 2013 Null and Alternative Hypothesis Neyman-Pearson Approach to Statistical Inference: A statistical test (also known as a hypothesis test)
6.4 Type I and Type II Errors Ulrich Hoensch Friday, March 22, 2013 Null and Alternative Hypothesis Neyman-Pearson Approach to Statistical Inference: A statistical test (also known as a hypothesis test)
The University of Hong Kong Department of Statistics and Actuarial Science STAT2802 Statistical Models Tutorial Solutions Solutions to Problems 71-80
 The University of Hong Kong Department of Statistics and Actuarial Science STAT2802 Statistical Models Tutorial Solutions Solutions to Problems 71-80 71. Decide in each case whether the hypothesis is simple
The University of Hong Kong Department of Statistics and Actuarial Science STAT2802 Statistical Models Tutorial Solutions Solutions to Problems 71-80 71. Decide in each case whether the hypothesis is simple
Experimental Design. Experimental design. Outline. Choice of platform Array design. Target samples
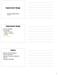 Experimental Design Credit for some of today s materials: Jean Yang, Terry Speed, and Christina Kendziorski Experimental design Choice of platform rray design Creation of probes Location on the array Controls
Experimental Design Credit for some of today s materials: Jean Yang, Terry Speed, and Christina Kendziorski Experimental design Choice of platform rray design Creation of probes Location on the array Controls
Statistical Tests. Matthieu de Lapparent
 Statistical Tests Matthieu de Lapparent matthieu.delapparent@epfl.ch Transport and Mobility Laboratory, School of Architecture, Civil and Environmental Engineering, Ecole Polytechnique Fédérale de Lausanne
Statistical Tests Matthieu de Lapparent matthieu.delapparent@epfl.ch Transport and Mobility Laboratory, School of Architecture, Civil and Environmental Engineering, Ecole Polytechnique Fédérale de Lausanne
Sample Size and Power Calculation in Microarray Studies Using the sizepower package.
 Sample Size and Power Calculation in Microarray Studies Using the sizepower package. Weiliang Qiu email: weiliang.qiu@gmail.com Mei-Ling Ting Lee email: meilinglee@sph.osu.edu George Alex Whitmore email:
Sample Size and Power Calculation in Microarray Studies Using the sizepower package. Weiliang Qiu email: weiliang.qiu@gmail.com Mei-Ling Ting Lee email: meilinglee@sph.osu.edu George Alex Whitmore email:
Bayesian methods in economics and finance
 1/26 Bayesian methods in economics and finance Linear regression: Bayesian model selection and sparsity priors Linear Regression 2/26 Linear regression Model for relationship between (several) independent
1/26 Bayesian methods in economics and finance Linear regression: Bayesian model selection and sparsity priors Linear Regression 2/26 Linear regression Model for relationship between (several) independent
Statistical Methods for Particle Physics Lecture 1: parameter estimation, statistical tests
 Statistical Methods for Particle Physics Lecture 1: parameter estimation, statistical tests http://benasque.org/2018tae/cgi-bin/talks/allprint.pl TAE 2018 Benasque, Spain 3-15 Sept 2018 Glen Cowan Physics
Statistical Methods for Particle Physics Lecture 1: parameter estimation, statistical tests http://benasque.org/2018tae/cgi-bin/talks/allprint.pl TAE 2018 Benasque, Spain 3-15 Sept 2018 Glen Cowan Physics
Interpreting Regression Results
 Interpreting Regression Results Carlo Favero Favero () Interpreting Regression Results 1 / 42 Interpreting Regression Results Interpreting regression results is not a simple exercise. We propose to split
Interpreting Regression Results Carlo Favero Favero () Interpreting Regression Results 1 / 42 Interpreting Regression Results Interpreting regression results is not a simple exercise. We propose to split
Economics 520. Lecture Note 19: Hypothesis Testing via the Neyman-Pearson Lemma CB 8.1,
 Economics 520 Lecture Note 9: Hypothesis Testing via the Neyman-Pearson Lemma CB 8., 8.3.-8.3.3 Uniformly Most Powerful Tests and the Neyman-Pearson Lemma Let s return to the hypothesis testing problem
Economics 520 Lecture Note 9: Hypothesis Testing via the Neyman-Pearson Lemma CB 8., 8.3.-8.3.3 Uniformly Most Powerful Tests and the Neyman-Pearson Lemma Let s return to the hypothesis testing problem
UCLA STAT 251. Statistical Methods for the Life and Health Sciences. Hypothesis Testing. Instructor: Ivo Dinov,
 UCLA STAT 251 Statistical Methods for the Life and Health Sciences Instructor: Ivo Dinov, Asst. Prof. In Statistics and Neurology University of California, Los Angeles, Winter 22 http://www.stat.ucla.edu/~dinov/
UCLA STAT 251 Statistical Methods for the Life and Health Sciences Instructor: Ivo Dinov, Asst. Prof. In Statistics and Neurology University of California, Los Angeles, Winter 22 http://www.stat.ucla.edu/~dinov/
Computer exercises on Experimental Design, ANOVA, and the BOOTSTRAP
 Computer exercises on Experimental Design, ANOVA, and the BOOTSTRAP Exercice 1: Copy the workspace ExpDes_1.Rdata to your local D drive and load it into a R-session. Perform a sample size calculation for
Computer exercises on Experimental Design, ANOVA, and the BOOTSTRAP Exercice 1: Copy the workspace ExpDes_1.Rdata to your local D drive and load it into a R-session. Perform a sample size calculation for
Optimal design of microarray experiments
 University of Groningen e.c.wit@rug.nl http://www.math.rug.nl/ ernst 7 June 2011 What is a cdna Microarray Experiment? GREEN (Cy3) Cancer Tissue mrna Mix tissues in equal amounts G G G R R R R R G R G
University of Groningen e.c.wit@rug.nl http://www.math.rug.nl/ ernst 7 June 2011 What is a cdna Microarray Experiment? GREEN (Cy3) Cancer Tissue mrna Mix tissues in equal amounts G G G R R R R R G R G
Alpha-Investing. Sequential Control of Expected False Discoveries
 Alpha-Investing Sequential Control of Expected False Discoveries Dean Foster Bob Stine Department of Statistics Wharton School of the University of Pennsylvania www-stat.wharton.upenn.edu/ stine Joint
Alpha-Investing Sequential Control of Expected False Discoveries Dean Foster Bob Stine Department of Statistics Wharton School of the University of Pennsylvania www-stat.wharton.upenn.edu/ stine Joint
Partitioning the Parameter Space. Topic 18 Composite Hypotheses
 Topic 18 Composite Hypotheses Partitioning the Parameter Space 1 / 10 Outline Partitioning the Parameter Space 2 / 10 Partitioning the Parameter Space Simple hypotheses limit us to a decision between one
Topic 18 Composite Hypotheses Partitioning the Parameter Space 1 / 10 Outline Partitioning the Parameter Space 2 / 10 Partitioning the Parameter Space Simple hypotheses limit us to a decision between one
Mathematical Statistics
 Mathematical Statistics MAS 713 Chapter 8 Previous lecture: 1 Bayesian Inference 2 Decision theory 3 Bayesian Vs. Frequentist 4 Loss functions 5 Conjugate priors Any questions? Mathematical Statistics
Mathematical Statistics MAS 713 Chapter 8 Previous lecture: 1 Bayesian Inference 2 Decision theory 3 Bayesian Vs. Frequentist 4 Loss functions 5 Conjugate priors Any questions? Mathematical Statistics
Introductory Econometrics
 Session 4 - Testing hypotheses Roland Sciences Po July 2011 Motivation After estimation, delivering information involves testing hypotheses Did this drug had any effect on the survival rate? Is this drug
Session 4 - Testing hypotheses Roland Sciences Po July 2011 Motivation After estimation, delivering information involves testing hypotheses Did this drug had any effect on the survival rate? Is this drug
Lecture 2: Basic Concepts and Simple Comparative Experiments Montgomery: Chapter 2
 Lecture 2: Basic Concepts and Simple Comparative Experiments Montgomery: Chapter 2 Fall, 2013 Page 1 Random Variable and Probability Distribution Discrete random variable Y : Finite possible values {y
Lecture 2: Basic Concepts and Simple Comparative Experiments Montgomery: Chapter 2 Fall, 2013 Page 1 Random Variable and Probability Distribution Discrete random variable Y : Finite possible values {y
Parameter Estimation and Fitting to Data
 Parameter Estimation and Fitting to Data Parameter estimation Maximum likelihood Least squares Goodness-of-fit Examples Elton S. Smith, Jefferson Lab 1 Parameter estimation Properties of estimators 3 An
Parameter Estimation and Fitting to Data Parameter estimation Maximum likelihood Least squares Goodness-of-fit Examples Elton S. Smith, Jefferson Lab 1 Parameter estimation Properties of estimators 3 An
Statistical Modeling and Analysis of Scientific Inquiry: The Basics of Hypothesis Testing
 Statistical Modeling and Analysis of Scientific Inquiry: The Basics of Hypothesis Testing So, What is Statistics? Theory and techniques for learning from data How to collect How to analyze How to interpret
Statistical Modeling and Analysis of Scientific Inquiry: The Basics of Hypothesis Testing So, What is Statistics? Theory and techniques for learning from data How to collect How to analyze How to interpret
Lesson 11. Functional Genomics I: Microarray Analysis
 Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Example 1: Two-Treatment CRD
 Introduction to Mixed Linear Models in Microarray Experiments //0 Copyright 0 Dan Nettleton Statistical Models A statistical model describes a formal mathematical data generation mechanism from which an
Introduction to Mixed Linear Models in Microarray Experiments //0 Copyright 0 Dan Nettleton Statistical Models A statistical model describes a formal mathematical data generation mechanism from which an
Low-Level Analysis of High- Density Oligonucleotide Microarray Data
 Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
f (1 0.5)/n Z =
 Math 466/566 - Homework 4. We want to test a hypothesis involving a population proportion. The unknown population proportion is p. The null hypothesis is p = / and the alternative hypothesis is p > /.
Math 466/566 - Homework 4. We want to test a hypothesis involving a population proportion. The unknown population proportion is p. The null hypothesis is p = / and the alternative hypothesis is p > /.
An inferential procedure to use sample data to understand a population Procedures
 Hypothesis Test An inferential procedure to use sample data to understand a population Procedures Hypotheses, the alpha value, the critical region (z-scores), statistics, conclusion Two types of errors
Hypothesis Test An inferential procedure to use sample data to understand a population Procedures Hypotheses, the alpha value, the critical region (z-scores), statistics, conclusion Two types of errors
How is the Statistical Power of Hypothesis Tests affected by Dose Uncertainty?
 How is the Statistical Power of Hypothesis Tests affected by Dose Uncertainty? by Eduard Hofer Workshop on Uncertainties in Radiation Dosimetry and their Impact on Risk Analysis, Washington, DC, May 2009
How is the Statistical Power of Hypothesis Tests affected by Dose Uncertainty? by Eduard Hofer Workshop on Uncertainties in Radiation Dosimetry and their Impact on Risk Analysis, Washington, DC, May 2009
Statistical Inference
 Statistical Inference Classical and Bayesian Methods Class 6 AMS-UCSC Thu 26, 2012 Winter 2012. Session 1 (Class 6) AMS-132/206 Thu 26, 2012 1 / 15 Topics Topics We will talk about... 1 Hypothesis testing
Statistical Inference Classical and Bayesian Methods Class 6 AMS-UCSC Thu 26, 2012 Winter 2012. Session 1 (Class 6) AMS-132/206 Thu 26, 2012 1 / 15 Topics Topics We will talk about... 1 Hypothesis testing
Analysis of Variance
 Statistical Techniques II EXST7015 Analysis of Variance 15a_ANOVA_Introduction 1 Design The simplest model for Analysis of Variance (ANOVA) is the CRD, the Completely Randomized Design This model is also
Statistical Techniques II EXST7015 Analysis of Variance 15a_ANOVA_Introduction 1 Design The simplest model for Analysis of Variance (ANOVA) is the CRD, the Completely Randomized Design This model is also
Sample Size Estimation for Studies of High-Dimensional Data
 Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
Class 19. Daniel B. Rowe, Ph.D. Department of Mathematics, Statistics, and Computer Science. Marquette University MATH 1700
 Class 19 Daniel B. Rowe, Ph.D. Department of Mathematics, Statistics, and Computer Science Copyright 2017 by D.B. Rowe 1 Agenda: Recap Chapter 8.3-8.4 Lecture Chapter 8.5 Go over Exam. Problem Solving
Class 19 Daniel B. Rowe, Ph.D. Department of Mathematics, Statistics, and Computer Science Copyright 2017 by D.B. Rowe 1 Agenda: Recap Chapter 8.3-8.4 Lecture Chapter 8.5 Go over Exam. Problem Solving
Ch. 5 Hypothesis Testing
 Ch. 5 Hypothesis Testing The current framework of hypothesis testing is largely due to the work of Neyman and Pearson in the late 1920s, early 30s, complementing Fisher s work on estimation. As in estimation,
Ch. 5 Hypothesis Testing The current framework of hypothesis testing is largely due to the work of Neyman and Pearson in the late 1920s, early 30s, complementing Fisher s work on estimation. As in estimation,
Statistical Data Analysis Stat 3: p-values, parameter estimation
 Statistical Data Analysis Stat 3: p-values, parameter estimation London Postgraduate Lectures on Particle Physics; University of London MSci course PH4515 Glen Cowan Physics Department Royal Holloway,
Statistical Data Analysis Stat 3: p-values, parameter estimation London Postgraduate Lectures on Particle Physics; University of London MSci course PH4515 Glen Cowan Physics Department Royal Holloway,
STAT 450: Final Examination Version 1. Richard Lockhart 16 December 2002
 Name: Last Name 1, First Name 1 Stdnt # StudentNumber1 STAT 450: Final Examination Version 1 Richard Lockhart 16 December 2002 Instructions: This is an open book exam. You may use notes, books and a calculator.
Name: Last Name 1, First Name 1 Stdnt # StudentNumber1 STAT 450: Final Examination Version 1 Richard Lockhart 16 December 2002 Instructions: This is an open book exam. You may use notes, books and a calculator.
Sequential Procedure for Testing Hypothesis about Mean of Latent Gaussian Process
 Applied Mathematical Sciences, Vol. 4, 2010, no. 62, 3083-3093 Sequential Procedure for Testing Hypothesis about Mean of Latent Gaussian Process Julia Bondarenko Helmut-Schmidt University Hamburg University
Applied Mathematical Sciences, Vol. 4, 2010, no. 62, 3083-3093 Sequential Procedure for Testing Hypothesis about Mean of Latent Gaussian Process Julia Bondarenko Helmut-Schmidt University Hamburg University
8.1-4 Test of Hypotheses Based on a Single Sample
 8.1-4 Test of Hypotheses Based on a Single Sample Example 1 (Example 8.6, p. 312) A manufacturer of sprinkler systems used for fire protection in office buildings claims that the true average system-activation
8.1-4 Test of Hypotheses Based on a Single Sample Example 1 (Example 8.6, p. 312) A manufacturer of sprinkler systems used for fire protection in office buildings claims that the true average system-activation
Statistical Methods in Particle Physics Lecture 1: Bayesian methods
 Statistical Methods in Particle Physics Lecture 1: Bayesian methods SUSSP65 St Andrews 16 29 August 2009 Glen Cowan Physics Department Royal Holloway, University of London g.cowan@rhul.ac.uk www.pp.rhul.ac.uk/~cowan
Statistical Methods in Particle Physics Lecture 1: Bayesian methods SUSSP65 St Andrews 16 29 August 2009 Glen Cowan Physics Department Royal Holloway, University of London g.cowan@rhul.ac.uk www.pp.rhul.ac.uk/~cowan
Statistical inference
 Statistical inference Contents 1. Main definitions 2. Estimation 3. Testing L. Trapani MSc Induction - Statistical inference 1 1 Introduction: definition and preliminary theory In this chapter, we shall
Statistical inference Contents 1. Main definitions 2. Estimation 3. Testing L. Trapani MSc Induction - Statistical inference 1 1 Introduction: definition and preliminary theory In this chapter, we shall
Mathematical statistics
 October 20 th, 2018 Lecture 17: Tests of Hypotheses Overview Week 1 Week 2 Week 4 Week 7 Week 10 Week 14 Probability reviews Chapter 6: Statistics and Sampling Distributions Chapter 7: Point Estimation
October 20 th, 2018 Lecture 17: Tests of Hypotheses Overview Week 1 Week 2 Week 4 Week 7 Week 10 Week 14 Probability reviews Chapter 6: Statistics and Sampling Distributions Chapter 7: Point Estimation
Section 5.4: Hypothesis testing for μ
 Section 5.4: Hypothesis testing for μ Possible claims or hypotheses: Ball bearings have μ = 1 cm Medicine decreases blood pressure For testing hypotheses, we set up a null (H 0 ) and alternative (H a )
Section 5.4: Hypothesis testing for μ Possible claims or hypotheses: Ball bearings have μ = 1 cm Medicine decreases blood pressure For testing hypotheses, we set up a null (H 0 ) and alternative (H a )
Bayesian Regression (1/31/13)
 STA613/CBB540: Statistical methods in computational biology Bayesian Regression (1/31/13) Lecturer: Barbara Engelhardt Scribe: Amanda Lea 1 Bayesian Paradigm Bayesian methods ask: given that I have observed
STA613/CBB540: Statistical methods in computational biology Bayesian Regression (1/31/13) Lecturer: Barbara Engelhardt Scribe: Amanda Lea 1 Bayesian Paradigm Bayesian methods ask: given that I have observed
ORF 245 Fundamentals of Statistics Chapter 9 Hypothesis Testing
 ORF 245 Fundamentals of Statistics Chapter 9 Hypothesis Testing Robert Vanderbei Fall 2014 Slides last edited on November 24, 2014 http://www.princeton.edu/ rvdb Coin Tossing Example Consider two coins.
ORF 245 Fundamentals of Statistics Chapter 9 Hypothesis Testing Robert Vanderbei Fall 2014 Slides last edited on November 24, 2014 http://www.princeton.edu/ rvdb Coin Tossing Example Consider two coins.
Mixtures and Hidden Markov Models for analyzing genomic data
 Mixtures and Hidden Markov Models for analyzing genomic data Marie-Laure Martin-Magniette UMR AgroParisTech/INRA Mathématique et Informatique Appliquées, Paris UMR INRA/UEVE ERL CNRS Unité de Recherche
Mixtures and Hidden Markov Models for analyzing genomic data Marie-Laure Martin-Magniette UMR AgroParisTech/INRA Mathématique et Informatique Appliquées, Paris UMR INRA/UEVE ERL CNRS Unité de Recherche
Topics on statistical design and analysis. of cdna microarray experiment
 Topics on statistical design and analysis of cdna microarray experiment Ximin Zhu A Dissertation Submitted to the University of Glasgow for the degree of Doctor of Philosophy Department of Statistics May
Topics on statistical design and analysis of cdna microarray experiment Ximin Zhu A Dissertation Submitted to the University of Glasgow for the degree of Doctor of Philosophy Department of Statistics May
CH.9 Tests of Hypotheses for a Single Sample
 CH.9 Tests of Hypotheses for a Single Sample Hypotheses testing Tests on the mean of a normal distributionvariance known Tests on the mean of a normal distributionvariance unknown Tests on the variance
CH.9 Tests of Hypotheses for a Single Sample Hypotheses testing Tests on the mean of a normal distributionvariance known Tests on the mean of a normal distributionvariance unknown Tests on the variance
PASS Sample Size Software. Poisson Regression
 Chapter 870 Introduction Poisson regression is used when the dependent variable is a count. Following the results of Signorini (99), this procedure calculates power and sample size for testing the hypothesis
Chapter 870 Introduction Poisson regression is used when the dependent variable is a count. Following the results of Signorini (99), this procedure calculates power and sample size for testing the hypothesis
High-Throughput Sequencing Course. Introduction. Introduction. Multiple Testing. Biostatistics and Bioinformatics. Summer 2018
 High-Throughput Sequencing Course Multiple Testing Biostatistics and Bioinformatics Summer 2018 Introduction You have previously considered the significance of a single gene Introduction You have previously
High-Throughput Sequencing Course Multiple Testing Biostatistics and Bioinformatics Summer 2018 Introduction You have previously considered the significance of a single gene Introduction You have previously
Lecture on Null Hypothesis Testing & Temporal Correlation
 Lecture on Null Hypothesis Testing & Temporal Correlation CS 590.21 Analysis and Modeling of Brain Networks Department of Computer Science University of Crete Acknowledgement Resources used in the slides
Lecture on Null Hypothesis Testing & Temporal Correlation CS 590.21 Analysis and Modeling of Brain Networks Department of Computer Science University of Crete Acknowledgement Resources used in the slides
Political Science 236 Hypothesis Testing: Review and Bootstrapping
 Political Science 236 Hypothesis Testing: Review and Bootstrapping Rocío Titiunik Fall 2007 1 Hypothesis Testing Definition 1.1 Hypothesis. A hypothesis is a statement about a population parameter The
Political Science 236 Hypothesis Testing: Review and Bootstrapping Rocío Titiunik Fall 2007 1 Hypothesis Testing Definition 1.1 Hypothesis. A hypothesis is a statement about a population parameter The
Power and Sample Size + Principles of Simulation. Benjamin Neale March 4 th, 2010 International Twin Workshop, Boulder, CO
 Power and Sample Size + Principles of Simulation Benjamin Neale March 4 th, 2010 International Twin Workshop, Boulder, CO What is power? What affects power? How do we calculate power? What is simulation?
Power and Sample Size + Principles of Simulation Benjamin Neale March 4 th, 2010 International Twin Workshop, Boulder, CO What is power? What affects power? How do we calculate power? What is simulation?
Probabilistic Inference for Multiple Testing
 This is the title page! This is the title page! Probabilistic Inference for Multiple Testing Chuanhai Liu and Jun Xie Department of Statistics, Purdue University, West Lafayette, IN 47907. E-mail: chuanhai,
This is the title page! This is the title page! Probabilistic Inference for Multiple Testing Chuanhai Liu and Jun Xie Department of Statistics, Purdue University, West Lafayette, IN 47907. E-mail: chuanhai,
Statistical analysis of microarray data: a Bayesian approach
 Biostatistics (003), 4, 4,pp. 597 60 Printed in Great Britain Statistical analysis of microarray data: a Bayesian approach RAPHAEL GTTARD University of Washington, Department of Statistics, Box 3543, Seattle,
Biostatistics (003), 4, 4,pp. 597 60 Printed in Great Britain Statistical analysis of microarray data: a Bayesian approach RAPHAEL GTTARD University of Washington, Department of Statistics, Box 3543, Seattle,
Exam: high-dimensional data analysis February 28, 2014
 Exam: high-dimensional data analysis February 28, 2014 Instructions: - Write clearly. Scribbles will not be deciphered. - Answer each main question (not the subquestions) on a separate piece of paper.
Exam: high-dimensional data analysis February 28, 2014 Instructions: - Write clearly. Scribbles will not be deciphered. - Answer each main question (not the subquestions) on a separate piece of paper.
Sizing Mixture (RSM) Designs for Adequate Precision via Fraction of Design Space (FDS)
 for Adequate Precision via Fraction of Design Space (FDS) Pat Whitcomb Stat-Ease, Inc. 61.746.036 036 fax 61.746.056 pat@statease.com Gary W. Oehlert School of Statistics University it of Minnesota 61.65.1557
for Adequate Precision via Fraction of Design Space (FDS) Pat Whitcomb Stat-Ease, Inc. 61.746.036 036 fax 61.746.056 pat@statease.com Gary W. Oehlert School of Statistics University it of Minnesota 61.65.1557
Design and Analysis of Gene Expression Experiments
 Design and Analysis of Gene Expression Experiments Guilherme J. M. Rosa Department of Animal Sciences Department of Biostatistics & Medical Informatics University of Wisconsin - Madison OUTLINE Æ Linear
Design and Analysis of Gene Expression Experiments Guilherme J. M. Rosa Department of Animal Sciences Department of Biostatistics & Medical Informatics University of Wisconsin - Madison OUTLINE Æ Linear
SMAM 314 Exam 3 Name. F A. A null hypothesis that is rejected at α =.05 will always be rejected at α =.01.
 SMAM 314 Exam 3 Name 1. Indicate whether the following statements are true (T) or false (F) (6 points) F A. A null hypothesis that is rejected at α =.05 will always be rejected at α =.01. T B. A course
SMAM 314 Exam 3 Name 1. Indicate whether the following statements are true (T) or false (F) (6 points) F A. A null hypothesis that is rejected at α =.05 will always be rejected at α =.01. T B. A course
Optimal normalization of DNA-microarray data
 Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La
Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La
Single gene analysis of differential expression. Giorgio Valentini
 Single gene analysis of differential expression Giorgio Valentini valenti@disi.unige.it Comparing two conditions Each condition may be represented by one or more RNA samples. Using cdna microarrays, samples
Single gene analysis of differential expression Giorgio Valentini valenti@disi.unige.it Comparing two conditions Each condition may be represented by one or more RNA samples. Using cdna microarrays, samples
Stat 535 C - Statistical Computing & Monte Carlo Methods. Arnaud Doucet.
 Stat 535 C - Statistical Computing & Monte Carlo Methods Arnaud Doucet Email: arnaud@cs.ubc.ca 1 CS students: don t forget to re-register in CS-535D. Even if you just audit this course, please do register.
Stat 535 C - Statistical Computing & Monte Carlo Methods Arnaud Doucet Email: arnaud@cs.ubc.ca 1 CS students: don t forget to re-register in CS-535D. Even if you just audit this course, please do register.
Data analysis and Geostatistics - lecture VI
 Data analysis and Geostatistics - lecture VI Statistical testing with population distributions Statistical testing - the steps 1. Define a hypothesis to test in statistics only a hypothesis rejection is
Data analysis and Geostatistics - lecture VI Statistical testing with population distributions Statistical testing - the steps 1. Define a hypothesis to test in statistics only a hypothesis rejection is
Hypothesis Testing. Part I. James J. Heckman University of Chicago. Econ 312 This draft, April 20, 2006
 Hypothesis Testing Part I James J. Heckman University of Chicago Econ 312 This draft, April 20, 2006 1 1 A Brief Review of Hypothesis Testing and Its Uses values and pure significance tests (R.A. Fisher)
Hypothesis Testing Part I James J. Heckman University of Chicago Econ 312 This draft, April 20, 2006 1 1 A Brief Review of Hypothesis Testing and Its Uses values and pure significance tests (R.A. Fisher)
Inferential Statistical Analysis of Microarray Experiments 2007 Arizona Microarray Workshop
 Inferential Statistical Analysis of Microarray Experiments 007 Arizona Microarray Workshop μ!! Robert J Tempelman Department of Animal Science tempelma@msuedu HYPOTHESIS TESTING (as if there was only one
Inferential Statistical Analysis of Microarray Experiments 007 Arizona Microarray Workshop μ!! Robert J Tempelman Department of Animal Science tempelma@msuedu HYPOTHESIS TESTING (as if there was only one
Non-specific filtering and control of false positives
 Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
BIOS 312: Precision of Statistical Inference
 and Power/Sample Size and Standard Errors BIOS 312: of Statistical Inference Chris Slaughter Department of Biostatistics, Vanderbilt University School of Medicine January 3, 2013 Outline Overview and Power/Sample
and Power/Sample Size and Standard Errors BIOS 312: of Statistical Inference Chris Slaughter Department of Biostatistics, Vanderbilt University School of Medicine January 3, 2013 Outline Overview and Power/Sample
" M A #M B. Standard deviation of the population (Greek lowercase letter sigma) σ 2
 Notation and Equations for Final Exam Symbol Definition X The variable we measure in a scientific study n The size of the sample N The size of the population M The mean of the sample µ The mean of the
Notation and Equations for Final Exam Symbol Definition X The variable we measure in a scientific study n The size of the sample N The size of the population M The mean of the sample µ The mean of the
Power and sample size calculations
 Patrick Breheny October 20 Patrick Breheny University of Iowa Biostatistical Methods I (BIOS 5710) 1 / 26 Planning a study Introduction What is power? Why is it important? Setup One of the most important
Patrick Breheny October 20 Patrick Breheny University of Iowa Biostatistical Methods I (BIOS 5710) 1 / 26 Planning a study Introduction What is power? Why is it important? Setup One of the most important
1 Descriptive statistics. 2 Scores and probability distributions. 3 Hypothesis testing and one-sample t-test. 4 More on t-tests
 Overall Overview INFOWO Statistics lecture S3: Hypothesis testing Peter de Waal Department of Information and Computing Sciences Faculty of Science, Universiteit Utrecht 1 Descriptive statistics 2 Scores
Overall Overview INFOWO Statistics lecture S3: Hypothesis testing Peter de Waal Department of Information and Computing Sciences Faculty of Science, Universiteit Utrecht 1 Descriptive statistics 2 Scores
Preface Introduction to Statistics and Data Analysis Overview: Statistical Inference, Samples, Populations, and Experimental Design The Role of
 Preface Introduction to Statistics and Data Analysis Overview: Statistical Inference, Samples, Populations, and Experimental Design The Role of Probability Sampling Procedures Collection of Data Measures
Preface Introduction to Statistics and Data Analysis Overview: Statistical Inference, Samples, Populations, and Experimental Design The Role of Probability Sampling Procedures Collection of Data Measures
Lecture 7: Interaction Analysis. Summer Institute in Statistical Genetics 2017
 Lecture 7: Interaction Analysis Timothy Thornton and Michael Wu Summer Institute in Statistical Genetics 2017 1 / 39 Lecture Outline Beyond main SNP effects Introduction to Concept of Statistical Interaction
Lecture 7: Interaction Analysis Timothy Thornton and Michael Wu Summer Institute in Statistical Genetics 2017 1 / 39 Lecture Outline Beyond main SNP effects Introduction to Concept of Statistical Interaction
Seminar Microarray-Datenanalyse
 Seminar Microarray- Normalization Hans-Ulrich Klein Christian Ruckert Institut für Medizinische Informatik WWU Münster SS 2011 Organisation 1 09.05.11 Normalisierung 2 10.05.11 Bestimmen diff. expr. Gene,
Seminar Microarray- Normalization Hans-Ulrich Klein Christian Ruckert Institut für Medizinische Informatik WWU Münster SS 2011 Organisation 1 09.05.11 Normalisierung 2 10.05.11 Bestimmen diff. expr. Gene,
Theory. Pattern and Process
 Theory Pattern and Process Definition of Science: Science is based on evidence and always changing. Scientists test explanations and predictions of natural phenomena. Some questions are outside the realm
Theory Pattern and Process Definition of Science: Science is based on evidence and always changing. Scientists test explanations and predictions of natural phenomena. Some questions are outside the realm
CHAPTER 8. Test Procedures is a rule, based on sample data, for deciding whether to reject H 0 and contains:
 CHAPTER 8 Test of Hypotheses Based on a Single Sample Hypothesis testing is the method that decide which of two contradictory claims about the parameter is correct. Here the parameters of interest are
CHAPTER 8 Test of Hypotheses Based on a Single Sample Hypothesis testing is the method that decide which of two contradictory claims about the parameter is correct. Here the parameters of interest are
Setting Precision Requirements for Estimating Proportions
 Setting Precision Requirements for Estimating Proportions James R. Chromy August 1, 2007 Joint Statistical Meetings, Salt Lake City, Utah 3040 Cornwallis Road P.O. Box 12194 Research Triangle Park, North
Setting Precision Requirements for Estimating Proportions James R. Chromy August 1, 2007 Joint Statistical Meetings, Salt Lake City, Utah 3040 Cornwallis Road P.O. Box 12194 Research Triangle Park, North
Bios 6649: Clinical Trials - Statistical Design and Monitoring
 Bios 6649: Clinical Trials - Statistical Design and Monitoring Spring Semester 2015 John M. Kittelson Department of Biostatistics & nformatics Colorado School of Public Health University of Colorado Denver
Bios 6649: Clinical Trials - Statistical Design and Monitoring Spring Semester 2015 John M. Kittelson Department of Biostatistics & nformatics Colorado School of Public Health University of Colorado Denver
DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS
 Sankhyā : The Indian Journal of Statistics Special issue in memory of D. Basu 2002, Volume 64, Series A, Pt. 3, pp 706-720 DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS By TERENCE P. SPEED
Sankhyā : The Indian Journal of Statistics Special issue in memory of D. Basu 2002, Volume 64, Series A, Pt. 3, pp 706-720 DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS By TERENCE P. SPEED
Simple Linear Regression for the Climate Data
 Prediction Prediction Interval Temperature 0.2 0.0 0.2 0.4 0.6 0.8 320 340 360 380 CO 2 Simple Linear Regression for the Climate Data What do we do with the data? y i = Temperature of i th Year x i =CO
Prediction Prediction Interval Temperature 0.2 0.0 0.2 0.4 0.6 0.8 320 340 360 380 CO 2 Simple Linear Regression for the Climate Data What do we do with the data? y i = Temperature of i th Year x i =CO
Introduction 1. STA442/2101 Fall See last slide for copyright information. 1 / 33
 Introduction 1 STA442/2101 Fall 2016 1 See last slide for copyright information. 1 / 33 Background Reading Optional Chapter 1 of Linear models with R Chapter 1 of Davison s Statistical models: Data, and
Introduction 1 STA442/2101 Fall 2016 1 See last slide for copyright information. 1 / 33 Background Reading Optional Chapter 1 of Linear models with R Chapter 1 of Davison s Statistical models: Data, and
Practice Problems Section Problems
 Practice Problems Section 4-4-3 4-4 4-5 4-6 4-7 4-8 4-10 Supplemental Problems 4-1 to 4-9 4-13, 14, 15, 17, 19, 0 4-3, 34, 36, 38 4-47, 49, 5, 54, 55 4-59, 60, 63 4-66, 68, 69, 70, 74 4-79, 81, 84 4-85,
Practice Problems Section 4-4-3 4-4 4-5 4-6 4-7 4-8 4-10 Supplemental Problems 4-1 to 4-9 4-13, 14, 15, 17, 19, 0 4-3, 34, 36, 38 4-47, 49, 5, 54, 55 4-59, 60, 63 4-66, 68, 69, 70, 74 4-79, 81, 84 4-85,
A variance-stabilizing transformation for gene-expression microarray data
 BIOINFORMATICS Vol. 18 Suppl. 1 00 Pages S105 S110 A variance-stabilizing transformation for gene-expression microarray data B. P. Durbin 1, J. S. Hardin, D. M. Hawins 3 and D. M. Roce 4 1 Department of
BIOINFORMATICS Vol. 18 Suppl. 1 00 Pages S105 S110 A variance-stabilizing transformation for gene-expression microarray data B. P. Durbin 1, J. S. Hardin, D. M. Hawins 3 and D. M. Roce 4 1 Department of
