Optimal normalization of DNA-microarray data
|
|
|
- Norman Norton
- 5 years ago
- Views:
Transcription
1 Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La Roche, Pharma Research, Switzerland
2 Contents DNA-microarrays: Basic principle and challenges Optimal transformations Properties & example Application to DNA-microarray data Results Signal variability False positives Improvements Robust estimation Variance stabilization Summary & Outlook
3 DNA-Microarrays Probe 1 Probe 2 mrna-preparation DNA-Array adding Fluorescence Hybridisation
4 DNA-Microarrays: (AG Walz)
5 Challenges From Gene-Expression to numbers biological differences artifacts of fabrication process noise systematic errors correct for systematic errors horizons of comparability combine results from different groups measurements at different times make results comparable What is significant? biology: Log(ratio) > 2 what is the noise level? advanced normalization algorithms
6 Standard techniques: One color design: linear normalization most spots do not change regulation symmetric same mean, median Two color design: Normalization algorithms linear normalization for Log(red/green) the same variability same variance
7 Basic Idea: Real Expression: r g (Gen g) Experiment i measured expression: f i (r g ) Find f i which maximizes correlation Normalization algorithms Assumptions exact replications: none different conditions: most genes do not change robust algorithm biological effects outliers maximize correlation of most genes
8 Optimal Transformations Idea: Rényi-maximalcorrelation: Ψ(x 1, x 2 ) = sup f,g Properties: R(f(x 1 ), g(x 2 )) defined if x 1, x 2 const symmetric, normalized, 0 Ψ 1 Ψ = 0 if and only if x 1, x 2 independent Ψ = 1 if fully dependent p(x 1, x 2 ) = N(µ 1, µ 2, σ 1, σ 2 ) Ψ = R
9 Optimal transformations Advantages: f, g maximize Ψ minimize regression e 2 = e 2 (f, g ) = inf f,g e2 (f, g) e 2 (f, g) = E {[f(x 1 ) g(x 2 )] 2 } E{f 2 (x 1 )}, Ψ 2 = 1 e 2 Correct for every systematic error (Rényi, 1959) No parametric assumptions
10 How to find them? ACE (Breiman & Friedman) iterative algorithm, based on Alternating Conditional Expectation guaranteed convergence Example: Y = exp[sin(2πx) + ɛ/2], X, ɛ N(0, 1)
11 Optimal transformations
12 DNA-microarrays Generalization e 2 = i<j g[φ i (X ig ) Φ j (X jg )] 2 g Φ2 i (X ig) Idea ACE for all pairwise comparisons After every iterative step average all transformations for one measurement Details Rank ordering Expectation value computed by smoothing Transform back using joint distribution The algorithm
13 Results 158 Affymetrix Chips
14 Results
15 Results Variance
16 Results False positive rate
17 Improvements Problems Not robust (L2 norm) Expectation values at the borders of intensity scale Rank ordering Solution Least trimmed squares (LTS) regression to estimate φ i r 2 = N/2 i=1 r2 i:n, r2 i:n ordered residuals In addition: Normalize by variance stabilizing functions Random number Z with mean µ, Variance σ 2 (µ) h(t) = t 1 d u 0 V ar(u)
18 Least Trimmed Squares (LTS) Robust regression: Breakdown points: Least squares: 0 % Least trimmed squares: 50 %
19 Variance stabilization Up to now: smooth functions with µ = 0, σ = 1 Now: smooth functions with σ(µ) = const
20 Variance stabilization Constant variance as a function of intensity by using a common transformation
21 First Results real data (left) real data with artificial systematic errors (right) consistent results
22 The transformations
23 The data
24 Correct for systematic errors No parametric assumptions Significant reduction of signal variability false positive rate Application to data including effects robust because of LTS variance stabilizing functions false negative rate? Interested? Summary & Outlook Please let us know...
25 The algorithm For all pairwise comparisons i, j Φ i (X ig ) = X ig / X ig Repeat For all pairwise comparisons i, j Φ j (X jg ) = E [Φ i (X ig ) X jg ] Φ i (X ig ) = E [Φ j (X jg ) X ig ] / Φ i (X ig ) Compute mean of Φ i same replication while e 2 (Φ i, Φ j ) decreases Back
Lesson 11. Functional Genomics I: Microarray Analysis
 Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Normalization. Example of Replicate Data. Biostatistics Rafael A. Irizarry
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
Microarray Preprocessing
 Microarray Preprocessing Normaliza$on Normaliza$on is needed to ensure that differences in intensi$es are indeed due to differen$al expression, and not some prin$ng, hybridiza$on, or scanning ar$fact.
Microarray Preprocessing Normaliza$on Normaliza$on is needed to ensure that differences in intensi$es are indeed due to differen$al expression, and not some prin$ng, hybridiza$on, or scanning ar$fact.
Biochip informatics-(i)
 Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
Seminar Microarray-Datenanalyse
 Seminar Microarray- Normalization Hans-Ulrich Klein Christian Ruckert Institut für Medizinische Informatik WWU Münster SS 2011 Organisation 1 09.05.11 Normalisierung 2 10.05.11 Bestimmen diff. expr. Gene,
Seminar Microarray- Normalization Hans-Ulrich Klein Christian Ruckert Institut für Medizinische Informatik WWU Münster SS 2011 Organisation 1 09.05.11 Normalisierung 2 10.05.11 Bestimmen diff. expr. Gene,
Low-Level Analysis of High- Density Oligonucleotide Microarray Data
 Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Technologie w skali genomowej 2/ Algorytmiczne i statystyczne aspekty sekwencjonowania DNA
 Technologie w skali genomowej 2/ Algorytmiczne i statystyczne aspekty sekwencjonowania DNA Expression analysis for RNA-seq data Ewa Szczurek Instytut Informatyki Uniwersytet Warszawski 1/35 The problem
Technologie w skali genomowej 2/ Algorytmiczne i statystyczne aspekty sekwencjonowania DNA Expression analysis for RNA-seq data Ewa Szczurek Instytut Informatyki Uniwersytet Warszawski 1/35 The problem
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Expression arrays, normalization, and error models
 1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
Chapter 3: Statistical methods for estimation and testing. Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001).
 Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Bayesian Regression of Piecewise Constant Functions
 Marcus Hutter - 1 - Bayesian Regression of Piecewise Constant Functions Bayesian Regression of Piecewise Constant Functions Marcus Hutter Istituto Dalle Molle di Studi sull Intelligenza Artificiale IDSIA,
Marcus Hutter - 1 - Bayesian Regression of Piecewise Constant Functions Bayesian Regression of Piecewise Constant Functions Marcus Hutter Istituto Dalle Molle di Studi sull Intelligenza Artificiale IDSIA,
robustness, efficiency, breakdown point, outliers, rank-based procedures, least absolute regression
 Robust Statistics robustness, efficiency, breakdown point, outliers, rank-based procedures, least absolute regression University of California, San Diego Instructor: Ery Arias-Castro http://math.ucsd.edu/~eariasca/teaching.html
Robust Statistics robustness, efficiency, breakdown point, outliers, rank-based procedures, least absolute regression University of California, San Diego Instructor: Ery Arias-Castro http://math.ucsd.edu/~eariasca/teaching.html
Introduction to Linear regression analysis. Part 2. Model comparisons
 Introduction to Linear regression analysis Part Model comparisons 1 ANOVA for regression Total variation in Y SS Total = Variation explained by regression with X SS Regression + Residual variation SS Residual
Introduction to Linear regression analysis Part Model comparisons 1 ANOVA for regression Total variation in Y SS Total = Variation explained by regression with X SS Regression + Residual variation SS Residual
SUPPLEMENTAL DATA: ROBUST ESTIMATORS FOR EXPRESSION ANALYSIS EARL HUBBELL, WEI-MIN LIU, AND RUI MEI
 SUPPLEMENTAL DATA: ROBUST ESTIMATORS FOR EXPRESSION ANALYSIS EARL HUBBELL, WEI-MIN LIU, AND RUI MEI ABSTRACT. This is supplemental data extracted from the paper Robust Estimators for Expression Analysis
SUPPLEMENTAL DATA: ROBUST ESTIMATORS FOR EXPRESSION ANALYSIS EARL HUBBELL, WEI-MIN LIU, AND RUI MEI ABSTRACT. This is supplemental data extracted from the paper Robust Estimators for Expression Analysis
Discovering Correlation in Data. Vinh Nguyen Research Fellow in Data Science Computing and Information Systems DMD 7.
 Discovering Correlation in Data Vinh Nguyen (vinh.nguyen@unimelb.edu.au) Research Fellow in Data Science Computing and Information Systems DMD 7.14 Discovering Correlation Why is correlation important?
Discovering Correlation in Data Vinh Nguyen (vinh.nguyen@unimelb.edu.au) Research Fellow in Data Science Computing and Information Systems DMD 7.14 Discovering Correlation Why is correlation important?
Bayesian Models for Regularization in Optimization
 Bayesian Models for Regularization in Optimization Aleksandr Aravkin, UBC Bradley Bell, UW Alessandro Chiuso, Padova Michael Friedlander, UBC Gianluigi Pilloneto, Padova Jim Burke, UW MOPTA, Lehigh University,
Bayesian Models for Regularization in Optimization Aleksandr Aravkin, UBC Bradley Bell, UW Alessandro Chiuso, Padova Michael Friedlander, UBC Gianluigi Pilloneto, Padova Jim Burke, UW MOPTA, Lehigh University,
Probe-Level Analysis of Affymetrix GeneChip Microarray Data
 Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley University of Minnesota Mar 30, 2004 Outline
Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley University of Minnesota Mar 30, 2004 Outline
Robust Monte Carlo Methods for Sequential Planning and Decision Making
 Robust Monte Carlo Methods for Sequential Planning and Decision Making Sue Zheng, Jason Pacheco, & John Fisher Sensing, Learning, & Inference Group Computer Science & Artificial Intelligence Laboratory
Robust Monte Carlo Methods for Sequential Planning and Decision Making Sue Zheng, Jason Pacheco, & John Fisher Sensing, Learning, & Inference Group Computer Science & Artificial Intelligence Laboratory
Non-specific filtering and control of false positives
 Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Androgen-independent prostate cancer
 The following tutorial walks through the identification of biological themes in a microarray dataset examining androgen-independent. Visit the GeneSifter Data Center (www.genesifter.net/web/datacenter.html)
The following tutorial walks through the identification of biological themes in a microarray dataset examining androgen-independent. Visit the GeneSifter Data Center (www.genesifter.net/web/datacenter.html)
9. Robust regression
 9. Robust regression Least squares regression........................................................ 2 Problems with LS regression..................................................... 3 Robust regression............................................................
9. Robust regression Least squares regression........................................................ 2 Problems with LS regression..................................................... 3 Robust regression............................................................
Inferring Transcriptional Regulatory Networks from High-throughput Data
 Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Lecture 14 October 13
 STAT 383C: Statistical Modeling I Fall 2015 Lecture 14 October 13 Lecturer: Purnamrita Sarkar Scribe: Some one Disclaimer: These scribe notes have been slightly proofread and may have typos etc. Note:
STAT 383C: Statistical Modeling I Fall 2015 Lecture 14 October 13 Lecturer: Purnamrita Sarkar Scribe: Some one Disclaimer: These scribe notes have been slightly proofread and may have typos etc. Note:
Minimum Hellinger Distance Estimation in a. Semiparametric Mixture Model
 Minimum Hellinger Distance Estimation in a Semiparametric Mixture Model Sijia Xiang 1, Weixin Yao 1, and Jingjing Wu 2 1 Department of Statistics, Kansas State University, Manhattan, Kansas, USA 66506-0802.
Minimum Hellinger Distance Estimation in a Semiparametric Mixture Model Sijia Xiang 1, Weixin Yao 1, and Jingjing Wu 2 1 Department of Statistics, Kansas State University, Manhattan, Kansas, USA 66506-0802.
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data
 GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
Information geometry for bivariate distribution control
 Information geometry for bivariate distribution control C.T.J.Dodson + Hong Wang Mathematics + Control Systems Centre, University of Manchester Institute of Science and Technology Optimal control of stochastic
Information geometry for bivariate distribution control C.T.J.Dodson + Hong Wang Mathematics + Control Systems Centre, University of Manchester Institute of Science and Technology Optimal control of stochastic
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Optimal design of microarray experiments
 University of Groningen e.c.wit@rug.nl http://www.math.rug.nl/ ernst 7 June 2011 What is a cdna Microarray Experiment? GREEN (Cy3) Cancer Tissue mrna Mix tissues in equal amounts G G G R R R R R G R G
University of Groningen e.c.wit@rug.nl http://www.math.rug.nl/ ernst 7 June 2011 What is a cdna Microarray Experiment? GREEN (Cy3) Cancer Tissue mrna Mix tissues in equal amounts G G G R R R R R G R G
Sparsity Models. Tong Zhang. Rutgers University. T. Zhang (Rutgers) Sparsity Models 1 / 28
 Sparsity Models Tong Zhang Rutgers University T. Zhang (Rutgers) Sparsity Models 1 / 28 Topics Standard sparse regression model algorithms: convex relaxation and greedy algorithm sparse recovery analysis:
Sparsity Models Tong Zhang Rutgers University T. Zhang (Rutgers) Sparsity Models 1 / 28 Topics Standard sparse regression model algorithms: convex relaxation and greedy algorithm sparse recovery analysis:
4.1. Introduction: Comparing Means
 4. Analysis of Variance (ANOVA) 4.1. Introduction: Comparing Means Consider the problem of testing H 0 : µ 1 = µ 2 against H 1 : µ 1 µ 2 in two independent samples of two different populations of possibly
4. Analysis of Variance (ANOVA) 4.1. Introduction: Comparing Means Consider the problem of testing H 0 : µ 1 = µ 2 against H 1 : µ 1 µ 2 in two independent samples of two different populations of possibly
Package plw. R topics documented: May 7, Type Package
 Type Package Package plw May 7, 2018 Title Probe level Locally moderated Weighted t-tests. Version 1.40.0 Date 2009-07-22 Author Magnus Astrand Maintainer Magnus Astrand
Type Package Package plw May 7, 2018 Title Probe level Locally moderated Weighted t-tests. Version 1.40.0 Date 2009-07-22 Author Magnus Astrand Maintainer Magnus Astrand
Indian Statistical Institute
 Indian Statistical Institute Introductory Computer programming Robust Regression methods with high breakdown point Author: Roll No: MD1701 February 24, 2018 Contents 1 Introduction 2 2 Criteria for evaluating
Indian Statistical Institute Introductory Computer programming Robust Regression methods with high breakdown point Author: Roll No: MD1701 February 24, 2018 Contents 1 Introduction 2 2 Criteria for evaluating
SPOTTED cdna MICROARRAYS
 SPOTTED cdna MICROARRAYS Spot size: 50um - 150um SPOTTED cdna MICROARRAYS Compare the genetic expression in two samples of cells PRINT cdna from one gene on each spot SAMPLES cdna labelled red/green e.g.
SPOTTED cdna MICROARRAYS Spot size: 50um - 150um SPOTTED cdna MICROARRAYS Compare the genetic expression in two samples of cells PRINT cdna from one gene on each spot SAMPLES cdna labelled red/green e.g.
Prohorov s theorem. Bengt Ringnér. October 26, 2008
 Prohorov s theorem Bengt Ringnér October 26, 2008 1 The theorem Definition 1 A set Π of probability measures defined on the Borel sets of a topological space is called tight if, for each ε > 0, there is
Prohorov s theorem Bengt Ringnér October 26, 2008 1 The theorem Definition 1 A set Π of probability measures defined on the Borel sets of a topological space is called tight if, for each ε > 0, there is
Bayesian ANalysis of Variance for Microarray Analysis
 Bayesian ANalysis of Variance for Microarray Analysis c These notes are copyrighted by the authors. Unauthorized use is not permitted. Bayesian ANalysis of Variance p.1/19 Normalization Nuisance effects,
Bayesian ANalysis of Variance for Microarray Analysis c These notes are copyrighted by the authors. Unauthorized use is not permitted. Bayesian ANalysis of Variance p.1/19 Normalization Nuisance effects,
STA205 Probability: Week 8 R. Wolpert
 INFINITE COIN-TOSS AND THE LAWS OF LARGE NUMBERS The traditional interpretation of the probability of an event E is its asymptotic frequency: the limit as n of the fraction of n repeated, similar, and
INFINITE COIN-TOSS AND THE LAWS OF LARGE NUMBERS The traditional interpretation of the probability of an event E is its asymptotic frequency: the limit as n of the fraction of n repeated, similar, and
Multiple comparisons The problem with the one-pair-at-a-time approach is its error rate.
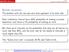 Multiple comparisons The problem with the one-pair-at-a-time approach is its error rate. Each confidence interval has a 95% probability of making a correct statement, and hence a 5% probability of making
Multiple comparisons The problem with the one-pair-at-a-time approach is its error rate. Each confidence interval has a 95% probability of making a correct statement, and hence a 5% probability of making
Asymptotic Statistics-III. Changliang Zou
 Asymptotic Statistics-III Changliang Zou The multivariate central limit theorem Theorem (Multivariate CLT for iid case) Let X i be iid random p-vectors with mean µ and and covariance matrix Σ. Then n (
Asymptotic Statistics-III Changliang Zou The multivariate central limit theorem Theorem (Multivariate CLT for iid case) Let X i be iid random p-vectors with mean µ and and covariance matrix Σ. Then n (
Introduction to Functional Data Analysis A CSCU Workshop. Giles Hooker Biological Statistics and Computational Biology
 Introduction to Functional Data Analysis A CSCU Workshop Giles Hooker Biological Statistics and Computational Biology gjh27@cornell.edu www.bscb.cornell.edu/ hooker/fdaworkshop 1 / 26 Agenda What is Functional
Introduction to Functional Data Analysis A CSCU Workshop Giles Hooker Biological Statistics and Computational Biology gjh27@cornell.edu www.bscb.cornell.edu/ hooker/fdaworkshop 1 / 26 Agenda What is Functional
Chapter 5: Microarray Techniques
 Chapter 5: Microarray Techniques 5.2 Analysis of Microarray Data Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Normalization Clustering Overview 2 1 Processing Microarray Data
Chapter 5: Microarray Techniques 5.2 Analysis of Microarray Data Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Normalization Clustering Overview 2 1 Processing Microarray Data
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Design of microarray experiments
 Design of microarray experiments Ulrich ansmann mansmann@imbi.uni-heidelberg.de Practical microarray analysis September Heidelberg Heidelberg, September otivation The lab biologist and theoretician need
Design of microarray experiments Ulrich ansmann mansmann@imbi.uni-heidelberg.de Practical microarray analysis September Heidelberg Heidelberg, September otivation The lab biologist and theoretician need
Lecture 12 Robust Estimation
 Lecture 12 Robust Estimation Prof. Dr. Svetlozar Rachev Institute for Statistics and Mathematical Economics University of Karlsruhe Financial Econometrics, Summer Semester 2007 Copyright These lecture-notes
Lecture 12 Robust Estimation Prof. Dr. Svetlozar Rachev Institute for Statistics and Mathematical Economics University of Karlsruhe Financial Econometrics, Summer Semester 2007 Copyright These lecture-notes
Bioconductor Project Working Papers
 Bioconductor Project Working Papers Bioconductor Project Year 2004 Paper 6 Error models for microarray intensities Wolfgang Huber Anja von Heydebreck Martin Vingron Department of Molecular Genome Analysis,
Bioconductor Project Working Papers Bioconductor Project Year 2004 Paper 6 Error models for microarray intensities Wolfgang Huber Anja von Heydebreck Martin Vingron Department of Molecular Genome Analysis,
Joint Probability Distributions
 Joint Probability Distributions ST 370 In many random experiments, more than one quantity is measured, meaning that there is more than one random variable. Example: Cell phone flash unit A flash unit is
Joint Probability Distributions ST 370 In many random experiments, more than one quantity is measured, meaning that there is more than one random variable. Example: Cell phone flash unit A flash unit is
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Nonlinear Programming Models
 Nonlinear Programming Models Fabio Schoen 2008 http://gol.dsi.unifi.it/users/schoen Nonlinear Programming Models p. Introduction Nonlinear Programming Models p. NLP problems minf(x) x S R n Standard form:
Nonlinear Programming Models Fabio Schoen 2008 http://gol.dsi.unifi.it/users/schoen Nonlinear Programming Models p. Introduction Nonlinear Programming Models p. NLP problems minf(x) x S R n Standard form:
2. Mathematical descriptions. (i) the master equation (ii) Langevin theory. 3. Single cell measurements
 1. Why stochastic?. Mathematical descriptions (i) the master equation (ii) Langevin theory 3. Single cell measurements 4. Consequences Any chemical reaction is stochastic. k P d φ dp dt = k d P deterministic
1. Why stochastic?. Mathematical descriptions (i) the master equation (ii) Langevin theory 3. Single cell measurements 4. Consequences Any chemical reaction is stochastic. k P d φ dp dt = k d P deterministic
LECTURE NOTE #NEW 6 PROF. ALAN YUILLE
 LECTURE NOTE #NEW 6 PROF. ALAN YUILLE 1. Introduction to Regression Now consider learning the conditional distribution p(y x). This is often easier than learning the likelihood function p(x y) and the
LECTURE NOTE #NEW 6 PROF. ALAN YUILLE 1. Introduction to Regression Now consider learning the conditional distribution p(y x). This is often easier than learning the likelihood function p(x y) and the
Practical Applications and Properties of the Exponentially. Modified Gaussian (EMG) Distribution. A Thesis. Submitted to the Faculty
 Practical Applications and Properties of the Exponentially Modified Gaussian (EMG) Distribution A Thesis Submitted to the Faculty of Drexel University by Scott Haney in partial fulfillment of the requirements
Practical Applications and Properties of the Exponentially Modified Gaussian (EMG) Distribution A Thesis Submitted to the Faculty of Drexel University by Scott Haney in partial fulfillment of the requirements
Sub-Gaussian Estimators of the Mean of a Random Matrix with Entries Possessing Only Two Moments
 Sub-Gaussian Estimators of the Mean of a Random Matrix with Entries Possessing Only Two Moments Stas Minsker University of Southern California July 21, 2016 ICERM Workshop Simple question: how to estimate
Sub-Gaussian Estimators of the Mean of a Random Matrix with Entries Possessing Only Two Moments Stas Minsker University of Southern California July 21, 2016 ICERM Workshop Simple question: how to estimate
Chapter 12. Analysis of variance
 Serik Sagitov, Chalmers and GU, January 9, 016 Chapter 1. Analysis of variance Chapter 11: I = samples independent samples paired samples Chapter 1: I 3 samples of equal size J one-way layout two-way layout
Serik Sagitov, Chalmers and GU, January 9, 016 Chapter 1. Analysis of variance Chapter 11: I = samples independent samples paired samples Chapter 1: I 3 samples of equal size J one-way layout two-way layout
Fuzzy Clustering of Gene Expression Data
 Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
High Breakdown Point Estimation in Regression
 WDS'08 Proceedings of Contributed Papers, Part I, 94 99, 2008. ISBN 978-80-7378-065-4 MATFYZPRESS High Breakdown Point Estimation in Regression T. Jurczyk Charles University, Faculty of Mathematics and
WDS'08 Proceedings of Contributed Papers, Part I, 94 99, 2008. ISBN 978-80-7378-065-4 MATFYZPRESS High Breakdown Point Estimation in Regression T. Jurczyk Charles University, Faculty of Mathematics and
Statistical Estimation
 Statistical Estimation Use data and a model. The plug-in estimators are based on the simple principle of applying the defining functional to the ECDF. Other methods of estimation: minimize residuals from
Statistical Estimation Use data and a model. The plug-in estimators are based on the simple principle of applying the defining functional to the ECDF. Other methods of estimation: minimize residuals from
Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance
 Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated
Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated
Regression Clustering
 Regression Clustering In regression clustering, we assume a model of the form y = f g (x, θ g ) + ɛ g for observations y and x in the g th group. Usually, of course, we assume linear models of the form
Regression Clustering In regression clustering, we assume a model of the form y = f g (x, θ g ) + ɛ g for observations y and x in the g th group. Usually, of course, we assume linear models of the form
More on Unsupervised Learning
 More on Unsupervised Learning Two types of problems are to find association rules for occurrences in common in observations (market basket analysis), and finding the groups of values of observational data
More on Unsupervised Learning Two types of problems are to find association rules for occurrences in common in observations (market basket analysis), and finding the groups of values of observational data
Variations on Nonparametric Additive Models: Computational and Statistical Aspects
 Variations on Nonparametric Additive Models: Computational and Statistical Aspects John Lafferty Department of Statistics & Department of Computer Science University of Chicago Collaborators Sivaraman
Variations on Nonparametric Additive Models: Computational and Statistical Aspects John Lafferty Department of Statistics & Department of Computer Science University of Chicago Collaborators Sivaraman
Mixture distributions with application to microarray data analysis
 University of South Florida Scholar Commons Graduate Theses and Dissertations Graduate School 2009 Mixture distributions with application to microarray data analysis O'Neil Lynch University of South Florida
University of South Florida Scholar Commons Graduate Theses and Dissertations Graduate School 2009 Mixture distributions with application to microarray data analysis O'Neil Lynch University of South Florida
Exhaustive search. CS 466 Saurabh Sinha
 Exhaustive search CS 466 Saurabh Sinha Agenda Two different problems Restriction mapping Motif finding Common theme: exhaustive search of solution space Reading: Chapter 4. Restriction Mapping Restriction
Exhaustive search CS 466 Saurabh Sinha Agenda Two different problems Restriction mapping Motif finding Common theme: exhaustive search of solution space Reading: Chapter 4. Restriction Mapping Restriction
cdna Microarray Analysis
 cdna Microarray Analysis with BioConductor packages Nolwenn Le Meur Copyright 2007 Outline Data acquisition Pre-processing Quality assessment Pre-processing background correction normalization summarization
cdna Microarray Analysis with BioConductor packages Nolwenn Le Meur Copyright 2007 Outline Data acquisition Pre-processing Quality assessment Pre-processing background correction normalization summarization
4 Invariant Statistical Decision Problems
 4 Invariant Statistical Decision Problems 4.1 Invariant decision problems Let G be a group of measurable transformations from the sample space X into itself. The group operation is composition. Note that
4 Invariant Statistical Decision Problems 4.1 Invariant decision problems Let G be a group of measurable transformations from the sample space X into itself. The group operation is composition. Note that
Probe-Level Analysis of Affymetrix GeneChip Microarray Data
 Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley Memorial Sloan-Kettering Cancer Center July
Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley Memorial Sloan-Kettering Cancer Center July
Experimental Design and Data Analysis for Biologists
 Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Data Preprocessing. Data Preprocessing
 Data Preprocessing 1 Data Preprocessing Normalization: the process of removing sampleto-sample variations in the measurements not due to differential gene expression. Bringing measurements from the different
Data Preprocessing 1 Data Preprocessing Normalization: the process of removing sampleto-sample variations in the measurements not due to differential gene expression. Bringing measurements from the different
Statistical Applications in Genetics and Molecular Biology
 Statistical Applications in Genetics and Molecular Biology Volume 6, Issue 1 2007 Article 28 A Comparison of Methods to Control Type I Errors in Microarray Studies Jinsong Chen Mark J. van der Laan Martyn
Statistical Applications in Genetics and Molecular Biology Volume 6, Issue 1 2007 Article 28 A Comparison of Methods to Control Type I Errors in Microarray Studies Jinsong Chen Mark J. van der Laan Martyn
S The Over-Reliance on the Central Limit Theorem
 S04-2008 The Over-Reliance on the Central Limit Theorem Abstract The objective is to demonstrate the theoretical and practical implication of the central limit theorem. The theorem states that as n approaches
S04-2008 The Over-Reliance on the Central Limit Theorem Abstract The objective is to demonstrate the theoretical and practical implication of the central limit theorem. The theorem states that as n approaches
On Modifications to Linking Variance Estimators in the Fay-Herriot Model that Induce Robustness
 Statistics and Applications {ISSN 2452-7395 (online)} Volume 16 No. 1, 2018 (New Series), pp 289-303 On Modifications to Linking Variance Estimators in the Fay-Herriot Model that Induce Robustness Snigdhansu
Statistics and Applications {ISSN 2452-7395 (online)} Volume 16 No. 1, 2018 (New Series), pp 289-303 On Modifications to Linking Variance Estimators in the Fay-Herriot Model that Induce Robustness Snigdhansu
Introduction to Empirical Processes and Semiparametric Inference Lecture 13: Entropy Calculations
 Introduction to Empirical Processes and Semiparametric Inference Lecture 13: Entropy Calculations Michael R. Kosorok, Ph.D. Professor and Chair of Biostatistics Professor of Statistics and Operations Research
Introduction to Empirical Processes and Semiparametric Inference Lecture 13: Entropy Calculations Michael R. Kosorok, Ph.D. Professor and Chair of Biostatistics Professor of Statistics and Operations Research
1 General problem. 2 Terminalogy. Estimation. Estimate θ. (Pick a plausible distribution from family. ) Or estimate τ = τ(θ).
 Estimation February 3, 206 Debdeep Pati General problem Model: {P θ : θ Θ}. Observe X P θ, θ Θ unknown. Estimate θ. (Pick a plausible distribution from family. ) Or estimate τ = τ(θ). Examples: θ = (µ,
Estimation February 3, 206 Debdeep Pati General problem Model: {P θ : θ Θ}. Observe X P θ, θ Θ unknown. Estimate θ. (Pick a plausible distribution from family. ) Or estimate τ = τ(θ). Examples: θ = (µ,
On the interior of the simplex, we have the Hessian of d(x), Hd(x) is diagonal with ith. µd(w) + w T c. minimize. subject to w T 1 = 1,
 Math 30 Winter 05 Solution to Homework 3. Recognizing the convexity of g(x) := x log x, from Jensen s inequality we get d(x) n x + + x n n log x + + x n n where the equality is attained only at x = (/n,...,
Math 30 Winter 05 Solution to Homework 3. Recognizing the convexity of g(x) := x log x, from Jensen s inequality we get d(x) n x + + x n n log x + + x n n where the equality is attained only at x = (/n,...,
Robustness and Distribution Assumptions
 Chapter 1 Robustness and Distribution Assumptions 1.1 Introduction In statistics, one often works with model assumptions, i.e., one assumes that data follow a certain model. Then one makes use of methodology
Chapter 1 Robustness and Distribution Assumptions 1.1 Introduction In statistics, one often works with model assumptions, i.e., one assumes that data follow a certain model. Then one makes use of methodology
Gaussian processes for inference in stochastic differential equations
 Gaussian processes for inference in stochastic differential equations Manfred Opper, AI group, TU Berlin November 6, 2017 Manfred Opper, AI group, TU Berlin (TU Berlin) inference in SDE November 6, 2017
Gaussian processes for inference in stochastic differential equations Manfred Opper, AI group, TU Berlin November 6, 2017 Manfred Opper, AI group, TU Berlin (TU Berlin) inference in SDE November 6, 2017
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
Inferential Statistical Analysis of Microarray Experiments 2007 Arizona Microarray Workshop
 Inferential Statistical Analysis of Microarray Experiments 007 Arizona Microarray Workshop μ!! Robert J Tempelman Department of Animal Science tempelma@msuedu HYPOTHESIS TESTING (as if there was only one
Inferential Statistical Analysis of Microarray Experiments 007 Arizona Microarray Workshop μ!! Robert J Tempelman Department of Animal Science tempelma@msuedu HYPOTHESIS TESTING (as if there was only one
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Robust statistics. Michael Love 7/10/2016
 Robust statistics Michael Love 7/10/2016 Robust topics Median MAD Spearman Wilcoxon rank test Weighted least squares Cook's distance M-estimators Robust topics Median => middle MAD => spread Spearman =>
Robust statistics Michael Love 7/10/2016 Robust topics Median MAD Spearman Wilcoxon rank test Weighted least squares Cook's distance M-estimators Robust topics Median => middle MAD => spread Spearman =>
Fractal functional regression for classification of gene expression data by wavelets
 Fractal functional regression for classification of gene expression data by wavelets Margarita María Rincón 1 and María Dolores Ruiz-Medina 2 1 University of Granada Campus Fuente Nueva 18071 Granada,
Fractal functional regression for classification of gene expression data by wavelets Margarita María Rincón 1 and María Dolores Ruiz-Medina 2 1 University of Granada Campus Fuente Nueva 18071 Granada,
Efficient and Robust Scale Estimation
 Efficient and Robust Scale Estimation Garth Tarr, Samuel Müller and Neville Weber School of Mathematics and Statistics THE UNIVERSITY OF SYDNEY Outline Introduction and motivation The robust scale estimator
Efficient and Robust Scale Estimation Garth Tarr, Samuel Müller and Neville Weber School of Mathematics and Statistics THE UNIVERSITY OF SYDNEY Outline Introduction and motivation The robust scale estimator
Correlation analysis 2: Measures of correlation
 Correlation analsis 2: Measures of correlation Ran Tibshirani Data Mining: 36-462/36-662 Februar 19 2013 1 Review: correlation Pearson s correlation is a measure of linear association In the population:
Correlation analsis 2: Measures of correlation Ran Tibshirani Data Mining: 36-462/36-662 Februar 19 2013 1 Review: correlation Pearson s correlation is a measure of linear association In the population:
Error models and normalization. Wolfgang Huber DKFZ Heidelberg
 Error models and normalization Wolfgang Huber DKFZ Heidelberg Acknowledgements Anja von Heydebreck, Martin Vingron Andreas Buness, Markus Ruschhaupt, Klaus Steiner, Jörg Schneider, Katharina Finis, Anke
Error models and normalization Wolfgang Huber DKFZ Heidelberg Acknowledgements Anja von Heydebreck, Martin Vingron Andreas Buness, Markus Ruschhaupt, Klaus Steiner, Jörg Schneider, Katharina Finis, Anke
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Design of microarray experiments
 Design of microarray experiments Ulrich Mansmann mansmann@imbi.uni-heidelberg.de Practical microarray analysis March 23 Heidelberg Heidelberg, March 23 Experiments Scientists deal mostly with experiments
Design of microarray experiments Ulrich Mansmann mansmann@imbi.uni-heidelberg.de Practical microarray analysis March 23 Heidelberg Heidelberg, March 23 Experiments Scientists deal mostly with experiments
networks in molecular biology Wolfgang Huber
 networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
Linear Algebra Massoud Malek
 CSUEB Linear Algebra Massoud Malek Inner Product and Normed Space In all that follows, the n n identity matrix is denoted by I n, the n n zero matrix by Z n, and the zero vector by θ n An inner product
CSUEB Linear Algebra Massoud Malek Inner Product and Normed Space In all that follows, the n n identity matrix is denoted by I n, the n n zero matrix by Z n, and the zero vector by θ n An inner product
Accelerated Block-Coordinate Relaxation for Regularized Optimization
 Accelerated Block-Coordinate Relaxation for Regularized Optimization Stephen J. Wright Computer Sciences University of Wisconsin, Madison October 09, 2012 Problem descriptions Consider where f is smooth
Accelerated Block-Coordinate Relaxation for Regularized Optimization Stephen J. Wright Computer Sciences University of Wisconsin, Madison October 09, 2012 Problem descriptions Consider where f is smooth
Image Cartoon-Texture Decomposition and Feature Selection using the Total Variation Regularized L 1 Functional
 Image Cartoon-Texture Decomposition and Feature Selection using the Total Variation Regularized L 1 Functional Wotao Yin 1, Donald Goldfarb 1, and Stanley Osher 2 1 Department of Industrial Engineering
Image Cartoon-Texture Decomposition and Feature Selection using the Total Variation Regularized L 1 Functional Wotao Yin 1, Donald Goldfarb 1, and Stanley Osher 2 1 Department of Industrial Engineering
APMA 2811Q. Homework #1. Due: 9/25/13. 1 exp ( f (x) 2) dx, I[f] =
![APMA 2811Q. Homework #1. Due: 9/25/13. 1 exp ( f (x) 2) dx, I[f] = APMA 2811Q. Homework #1. Due: 9/25/13. 1 exp ( f (x) 2) dx, I[f] =](/thumbs/74/71070037.jpg) APMA 8Q Homework # Due: 9/5/3. Ill-posed problems a) Consider I : W,, ) R defined by exp f x) ) dx, where W,, ) = f W,, ) : f) = f) = }. Show that I has no minimizer in A. This problem is not coercive
APMA 8Q Homework # Due: 9/5/3. Ill-posed problems a) Consider I : W,, ) R defined by exp f x) ) dx, where W,, ) = f W,, ) : f) = f) = }. Show that I has no minimizer in A. This problem is not coercive
Statistical analysis of microarray data: a Bayesian approach
 Biostatistics (003), 4, 4,pp. 597 60 Printed in Great Britain Statistical analysis of microarray data: a Bayesian approach RAPHAEL GTTARD University of Washington, Department of Statistics, Box 3543, Seattle,
Biostatistics (003), 4, 4,pp. 597 60 Printed in Great Britain Statistical analysis of microarray data: a Bayesian approach RAPHAEL GTTARD University of Washington, Department of Statistics, Box 3543, Seattle,
Robust Statistics, Revisited
 Robust Statistics, Revisited Ankur Moitra (MIT) joint work with Ilias Diakonikolas, Jerry Li, Gautam Kamath, Daniel Kane and Alistair Stewart CLASSIC PARAMETER ESTIMATION Given samples from an unknown
Robust Statistics, Revisited Ankur Moitra (MIT) joint work with Ilias Diakonikolas, Jerry Li, Gautam Kamath, Daniel Kane and Alistair Stewart CLASSIC PARAMETER ESTIMATION Given samples from an unknown
Support Vector Machines
 Support Vector Machines Here we approach the two-class classification problem in a direct way: We try and find a plane that separates the classes in feature space. If we cannot, we get creative in two
Support Vector Machines Here we approach the two-class classification problem in a direct way: We try and find a plane that separates the classes in feature space. If we cannot, we get creative in two
If we want to analyze experimental or simulated data we might encounter the following tasks:
 Chapter 1 Introduction If we want to analyze experimental or simulated data we might encounter the following tasks: Characterization of the source of the signal and diagnosis Studying dependencies Prediction
Chapter 1 Introduction If we want to analyze experimental or simulated data we might encounter the following tasks: Characterization of the source of the signal and diagnosis Studying dependencies Prediction
A Resampling Method on Pivotal Estimating Functions
 A Resampling Method on Pivotal Estimating Functions Kun Nie Biostat 277,Winter 2004 March 17, 2004 Outline Introduction A General Resampling Method Examples - Quantile Regression -Rank Regression -Simulation
A Resampling Method on Pivotal Estimating Functions Kun Nie Biostat 277,Winter 2004 March 17, 2004 Outline Introduction A General Resampling Method Examples - Quantile Regression -Rank Regression -Simulation
EECS E6690: Statistical Learning for Biological and Information Systems Lecture1: Introduction
 EECS E6690: Statistical Learning for Biological and Information Systems Lecture1: Introduction Prof. Predrag R. Jelenković Time: Tuesday 4:10-6:40pm 1127 Seeley W. Mudd Building Dept. of Electrical Engineering
EECS E6690: Statistical Learning for Biological and Information Systems Lecture1: Introduction Prof. Predrag R. Jelenković Time: Tuesday 4:10-6:40pm 1127 Seeley W. Mudd Building Dept. of Electrical Engineering
Regression Shrinkage and Selection via the Lasso
 Regression Shrinkage and Selection via the Lasso ROBERT TIBSHIRANI, 1996 Presenter: Guiyun Feng April 27 () 1 / 20 Motivation Estimation in Linear Models: y = β T x + ɛ. data (x i, y i ), i = 1, 2,...,
Regression Shrinkage and Selection via the Lasso ROBERT TIBSHIRANI, 1996 Presenter: Guiyun Feng April 27 () 1 / 20 Motivation Estimation in Linear Models: y = β T x + ɛ. data (x i, y i ), i = 1, 2,...,
Statistical Applications in Genetics and Molecular Biology
 Statistical Applications in Genetics and Molecular Biology Volume 4, Issue 1 2005 Article 30 Weighted Analysis of Paired Microarray Experiments Erik Kristiansson Anders Sjögren Mats Rudemo Olle Nerman
Statistical Applications in Genetics and Molecular Biology Volume 4, Issue 1 2005 Article 30 Weighted Analysis of Paired Microarray Experiments Erik Kristiansson Anders Sjögren Mats Rudemo Olle Nerman
Statistical Inference on Large Contingency Tables: Convergence, Testability, Stability. COMPSTAT 2010 Paris, August 23, 2010
 Statistical Inference on Large Contingency Tables: Convergence, Testability, Stability Marianna Bolla Institute of Mathematics Budapest University of Technology and Economics marib@math.bme.hu COMPSTAT
Statistical Inference on Large Contingency Tables: Convergence, Testability, Stability Marianna Bolla Institute of Mathematics Budapest University of Technology and Economics marib@math.bme.hu COMPSTAT
