cdna Microarray Analysis
|
|
|
- Gillian Franklin
- 5 years ago
- Views:
Transcription
1 cdna Microarray Analysis with BioConductor packages Nolwenn Le Meur Copyright 2007
2 Outline Data acquisition Pre-processing Quality assessment Pre-processing background correction normalization summarization
3 Two-color Microarray Probe (gene reporter) Oligonucleotides or cdna clones Scan Target Test Reference Excitation Laser 2 Laser 1 RT-PCR Label with Fluor dyes PCR amplification and purification Cy5 Cy3 Emission Spotting Hybridize target to microarray Computer analysis (adapted from Duggan et al., Nat. Gen., 1999)
4 Image Analysis 2. Segmentation 3. Quantification Intensity CH2(635) 1. Location CH1(532) Intensity Raw data 60000
5 Terminology Target: DNA hybridized to the array, mobile substrate. Probe: DNA spotted on the array (spot). print-tip-group : collection of spots printed using the same print-tip (or pin), aka. grid. G, Gb: Cy3 signal and background intensities R, Rb: Cy5 signal and background intensities
6 BioC Task View: 129 packages
7 Import data... using limma package limmausersguide() library( limma ) targets <- readtargets( Targets.txt ) RG <- read.maimages(targets$filename, source= genepix )
8 ...into RGList > names(rg) R source G Rb printer Gb targets > RG$R[1,] s1 s2 s3 s4 [1, ] genes
9 Quality Assessment vs Control Quality Assessment: computation and interpretation of metrics that are intended to measure quality Quality Control: possible subsequent action, removing bad array or re-doing part of the experiment
10 Quality Assessment For each array: Diagnostics plots of spot statistics e.g. R and G log-intensities, M, A, spot area. Boxplots; 2D spatial images; Scatter-plots, e.g. MA-plots; Density plots. Stratify plots according to layout parameters, e.g. print-tip-group, plate. summary statistics
11 Quality Filtering Intensity (635) Intensity (635) Intensity (532) Intensity (532) Intensity (532) Background Foreground Intensity (635) Intensity (635) Intensity (532) 6000
12 Spatial Effects Image Plots R Rb R-Rb color scale by rank another array: print-tip color scale ~ log(g) color scale ~ rank(g)
13 Spatial Effects 1 pin 1 block
14 Spotting Pin Quality Decline after delivery of 5x105 spots after delivery of 3x105 spots H. Sueltmann DKFZ/MGA
15 MA plot M = log2(r) - log2(g) A = 1/2(log2(R) + log2(g))
16 Print-tip Effects ECDF plot (a42-u07639vene.txt) by spotting pin ^ F F(q) 0.6 1:1 1:2 1:3 1:4 2:1 2:2 2:3 2:4 3:1 3:2 3:3 3:4 4:1 4:2 4:3 4: q (log-ratio) -0.4 log(fg.green/fg.red)
17 PCR Plates - Boxplots
18 Bioconductor QA packages Several packages arraymagic arrayquality arrayqualitymetrics pdf or html
19 Diagnostic plot with arrayquality
20 Data Exploration with limma (Limma user Guide)
21 Quality Assessment: Summary For each array: Diagnostics plots Stratify BioC packages: arrayquality arraymagic
22 Outline Data acquisition Pre-processing Quality assessment Pre-processing background correction normalization summarization
23 Sources of Variation RNA extraction Systematic similar effect on many reverse transcription measurements labeling efficiencies corrections can be estimated from data Scanner settings PCR DNA concentration Printing or pin cross-hybridization Calibration Stochastic too random to be explicitely accounted for noise Error Model
24 Variance-Bias trade off <- Precision Variance -> <- Bias Accuray ->
25 Background Correction none subtraction, movingmin Minimun,edwards, normexp, More details limma >?backgroundcorrect
26 Background Correction none substraction normexp
27 Why Normalize? Theory Cy5 vs Cy3 Theory Reality Cy5 vs Cy3 Raw Data - Cy5 vs Cy Cy5 Cy5 Raw Cy Cy3 Cy5 Cy Cy Cy3 Raw
28 Normalization Identify and remove the effects of systematic variation Normalization is closely related to quality assessment. In a ideal experiment, no normalization would be necessary, as the technical variations would have been avoided. Normalization is needed to ensure that differences in intensities are indeed due to differential expression, and not some printing, hybridization, or scanning artifact. Normalization is necessary before any analysis which involves within or between slide comparisons of intensities, e.g., clustering, testing.
29 Data Transformation measured intensity = offset + Yik = Bik + gain true abundance ik Sk Example: log transformation Intensity measurements adapt a distribution that is closer to the normal distribution Muliplicative noise becomes additive noise: variance more independent of intensity
30 Normalization methods median loess 2D loess print-tip loess variance stabilisation Two-channel Separate-channel Smyth, G. K., and Speed, T. P. (2003). In: METHODS: Selecting Candidate Genes from DNA Array Screens: Application to Neuroscience
31 Two channel normalization Location: centers log-ratios around zero using A and spatial dependent bias
32 Two channel normalization Location: centers log-ratios around zero using A and spatial dependent bias Scale: adjust for different in scale between multiple arrays median centered median centered & MAD scaled Scaling
33 One channel normalization As technology improves the spot-to-spot varation is reduced Development of normalization techniques that work on the absolute intensities Ex: quantile normalization (limma) variance stabilization (vsn)
34 Quantile Normalization Before After Bolstand et al.(2003)
35 Variance Statibilizing Transformation log arsinh intensity Meaningful around 0 Original intensities may be negatives (Huber et al. 2004)
36 Variance stabilization (vsn) linear log arsinh
37 Variance stabilization (vsn) xi log xj log-ratio 'glog' (generalized log-ratio) log xi + xi2 + ci2 xj + xj2 + cj2 - interpretation as "fold change" + interpretation even in cases where genes are off in some conditions (negative values) + visualization + can use standard statistical methods (hypothesis testing, ANOVA, clustering, classification ) without the worries about low-level variability that are often warranted on the log-scale
38 Preprocessing : Summary For each array: Background correction or not Normalization: bias-variance trade-off Diagnostic plots BioC pacakges: marray limma
39
Normalization. Example of Replicate Data. Biostatistics Rafael A. Irizarry
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
Microarray Preprocessing
 Microarray Preprocessing Normaliza$on Normaliza$on is needed to ensure that differences in intensi$es are indeed due to differen$al expression, and not some prin$ng, hybridiza$on, or scanning ar$fact.
Microarray Preprocessing Normaliza$on Normaliza$on is needed to ensure that differences in intensi$es are indeed due to differen$al expression, and not some prin$ng, hybridiza$on, or scanning ar$fact.
Seminar Microarray-Datenanalyse
 Seminar Microarray- Normalization Hans-Ulrich Klein Christian Ruckert Institut für Medizinische Informatik WWU Münster SS 2011 Organisation 1 09.05.11 Normalisierung 2 10.05.11 Bestimmen diff. expr. Gene,
Seminar Microarray- Normalization Hans-Ulrich Klein Christian Ruckert Institut für Medizinische Informatik WWU Münster SS 2011 Organisation 1 09.05.11 Normalisierung 2 10.05.11 Bestimmen diff. expr. Gene,
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Biochip informatics-(i)
 Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
Error models and normalization. Wolfgang Huber DKFZ Heidelberg
 Error models and normalization Wolfgang Huber DKFZ Heidelberg Acknowledgements Anja von Heydebreck, Martin Vingron Andreas Buness, Markus Ruschhaupt, Klaus Steiner, Jörg Schneider, Katharina Finis, Anke
Error models and normalization Wolfgang Huber DKFZ Heidelberg Acknowledgements Anja von Heydebreck, Martin Vingron Andreas Buness, Markus Ruschhaupt, Klaus Steiner, Jörg Schneider, Katharina Finis, Anke
SPOTTED cdna MICROARRAYS
 SPOTTED cdna MICROARRAYS Spot size: 50um - 150um SPOTTED cdna MICROARRAYS Compare the genetic expression in two samples of cells PRINT cdna from one gene on each spot SAMPLES cdna labelled red/green e.g.
SPOTTED cdna MICROARRAYS Spot size: 50um - 150um SPOTTED cdna MICROARRAYS Compare the genetic expression in two samples of cells PRINT cdna from one gene on each spot SAMPLES cdna labelled red/green e.g.
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Low-Level Analysis of High- Density Oligonucleotide Microarray Data
 Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Probe-Level Analysis of Affymetrix GeneChip Microarray Data
 Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley University of Minnesota Mar 30, 2004 Outline
Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley University of Minnesota Mar 30, 2004 Outline
Bioconductor Project Working Papers
 Bioconductor Project Working Papers Bioconductor Project Year 2004 Paper 6 Error models for microarray intensities Wolfgang Huber Anja von Heydebreck Martin Vingron Department of Molecular Genome Analysis,
Bioconductor Project Working Papers Bioconductor Project Year 2004 Paper 6 Error models for microarray intensities Wolfgang Huber Anja von Heydebreck Martin Vingron Department of Molecular Genome Analysis,
Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance
 Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated
Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated
Session 06 (A): Microarray Basic Data Analysis
 1 SJTU-Bioinformatics Summer School 2017 Session 06 (A): Microarray Basic Data Analysis Maoying,Wu ricket.woo@gmail.com Dept. of Bioinformatics & Biostatistics Shanghai Jiao Tong University Summer, 2017
1 SJTU-Bioinformatics Summer School 2017 Session 06 (A): Microarray Basic Data Analysis Maoying,Wu ricket.woo@gmail.com Dept. of Bioinformatics & Biostatistics Shanghai Jiao Tong University Summer, 2017
Lesson 11. Functional Genomics I: Microarray Analysis
 Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Data Preprocessing. Data Preprocessing
 Data Preprocessing 1 Data Preprocessing Normalization: the process of removing sampleto-sample variations in the measurements not due to differential gene expression. Bringing measurements from the different
Data Preprocessing 1 Data Preprocessing Normalization: the process of removing sampleto-sample variations in the measurements not due to differential gene expression. Bringing measurements from the different
Probe-Level Analysis of Affymetrix GeneChip Microarray Data
 Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley Memorial Sloan-Kettering Cancer Center July
Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley Memorial Sloan-Kettering Cancer Center July
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Expression arrays, normalization, and error models
 1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
(134/ !"#!$%&%' ()*)$+,-.)//01*$2*345/0/ M<39$N1$;<)5$")3/B.)O ;).C0*141=5 " /7$83.9$:
 (134/!"#!$%&%' ()*)$+,-)//01*$2*345/0/ "061335/7$839$: ;$9
(134/!"#!$%&%' ()*)$+,-)//01*$2*345/0/ "061335/7$839$: ;$9
Single gene analysis of differential expression. Giorgio Valentini
 Single gene analysis of differential expression Giorgio Valentini valenti@disi.unige.it Comparing two conditions Each condition may be represented by one or more RNA samples. Using cdna microarrays, samples
Single gene analysis of differential expression Giorgio Valentini valenti@disi.unige.it Comparing two conditions Each condition may be represented by one or more RNA samples. Using cdna microarrays, samples
Chapter 5: Microarray Techniques
 Chapter 5: Microarray Techniques 5.2 Analysis of Microarray Data Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Normalization Clustering Overview 2 1 Processing Microarray Data
Chapter 5: Microarray Techniques 5.2 Analysis of Microarray Data Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Normalization Clustering Overview 2 1 Processing Microarray Data
Gordon Smyth, WEHI Analysis of Replicated Experiments. January 2004 IMS NUS Microarray Tutorial. Some Web Sites. Analysis of Replicated Experiments
 Some Web Sites Statistical Methods in Microarray Analysis Tutorial Institute for Mathematical Sciences National University of Sinapore January, 004 Gordon Smyth Walter and Eliza Hall Institute Technical
Some Web Sites Statistical Methods in Microarray Analysis Tutorial Institute for Mathematical Sciences National University of Sinapore January, 004 Gordon Smyth Walter and Eliza Hall Institute Technical
Improving the identification of differentially expressed genes in cdna microarray experiments
 Improving the identification of differentially expressed genes in cdna microarray experiments Databionics Research Group University of Marburg 33 Marburg, Germany Alfred Ultsch Abstract. The identification
Improving the identification of differentially expressed genes in cdna microarray experiments Databionics Research Group University of Marburg 33 Marburg, Germany Alfred Ultsch Abstract. The identification
ANALYSIS OF DYNAMIC PROTEIN EXPRESSION DATA
 REVSTAT Statistical Journal Volume 3, Number 2, November 2005, 99 111 ANALYSIS OF DYNAMIC PROTEIN EXPRESSION DATA Authors: Klaus Jung Department of Statistics, University of Dortmund, Germany (klaus.jung@uni-dortmund.de)
REVSTAT Statistical Journal Volume 3, Number 2, November 2005, 99 111 ANALYSIS OF DYNAMIC PROTEIN EXPRESSION DATA Authors: Klaus Jung Department of Statistics, University of Dortmund, Germany (klaus.jung@uni-dortmund.de)
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Linear Models and Empirical Bayes Methods for. Assessing Differential Expression in Microarray Experiments
 Linear Models and Empirical Bayes Methods for Assessing Differential Expression in Microarray Experiments by Gordon K. Smyth (as interpreted by Aaron J. Baraff) STAT 572 Intro Talk April 10, 2014 Microarray
Linear Models and Empirical Bayes Methods for Assessing Differential Expression in Microarray Experiments by Gordon K. Smyth (as interpreted by Aaron J. Baraff) STAT 572 Intro Talk April 10, 2014 Microarray
Optimal design of microarray experiments
 University of Groningen e.c.wit@rug.nl http://www.math.rug.nl/ ernst 7 June 2011 What is a cdna Microarray Experiment? GREEN (Cy3) Cancer Tissue mrna Mix tissues in equal amounts G G G R R R R R G R G
University of Groningen e.c.wit@rug.nl http://www.math.rug.nl/ ernst 7 June 2011 What is a cdna Microarray Experiment? GREEN (Cy3) Cancer Tissue mrna Mix tissues in equal amounts G G G R R R R R G R G
Limma = linear models for microarray data. Linear models and Limma. Expression measures. Questions of Interest. y ga = log 2 (R/G) y ga =
 Linear models and Limma Københavns Universitet, 19 August 2009 Mark D. Robinson Bioinformatics, Walter+Eliza Hall Institute Epigenetics Laboratory, Garvan Institute (with many slides taken from Gordon
Linear models and Limma Københavns Universitet, 19 August 2009 Mark D. Robinson Bioinformatics, Walter+Eliza Hall Institute Epigenetics Laboratory, Garvan Institute (with many slides taken from Gordon
Lecture 5 Processing microarray data
 Lecture 5 Processin microarray data (1)Transform the data into a scale suitable for analysis ()Remove the effects of systematic and obfuscatin sources of variation (3)Identify discrepant observations Preprocessin
Lecture 5 Processin microarray data (1)Transform the data into a scale suitable for analysis ()Remove the effects of systematic and obfuscatin sources of variation (3)Identify discrepant observations Preprocessin
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data
 DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
ncounter PlexSet Data Analysis Guidelines
 ncounter PlexSet Data Analysis Guidelines NanoString Technologies, Inc. 530 airview Ave North Seattle, Washington 98109 USA Telephone: 206.378.6266 888.358.6266 E-mail: info@nanostring.com Molecules That
ncounter PlexSet Data Analysis Guidelines NanoString Technologies, Inc. 530 airview Ave North Seattle, Washington 98109 USA Telephone: 206.378.6266 888.358.6266 E-mail: info@nanostring.com Molecules That
Tools and topics for microarray analysis
 Tools and topics for microarray analysis USSES Conference, Blowing Rock, North Carolina, June, 2005 Jason A. Osborne, osborne@stat.ncsu.edu Department of Statistics, North Carolina State University 1 Outline
Tools and topics for microarray analysis USSES Conference, Blowing Rock, North Carolina, June, 2005 Jason A. Osborne, osborne@stat.ncsu.edu Department of Statistics, North Carolina State University 1 Outline
Optimal normalization of DNA-microarray data
 Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La
Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La
Clustering & microarray technology
 Clustering & microarray technology A large scale way to measure gene expression levels. Thanks to Kevin Wayne, Matt Hibbs, & SMD for a few of the slides 1 Why is expression important? Proteins Gene Expression
Clustering & microarray technology A large scale way to measure gene expression levels. Thanks to Kevin Wayne, Matt Hibbs, & SMD for a few of the slides 1 Why is expression important? Proteins Gene Expression
Statistics for Differential Expression in Sequencing Studies. Naomi Altman
 Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
Linear Models and Empirical Bayes Methods for Microarrays
 Methods for Microarrays by Gordon Smyth Alex Sánchez and Carme Ruíz de Villa Department d Estadística Universitat de Barcelona 16-12-2004 Outline 1 An introductory example Paper overview 2 3 Lönnsted and
Methods for Microarrays by Gordon Smyth Alex Sánchez and Carme Ruíz de Villa Department d Estadística Universitat de Barcelona 16-12-2004 Outline 1 An introductory example Paper overview 2 3 Lönnsted and
Lecture 1: The Structure of Microarray Data
 INTRODUCTION TO MICROARRAYS 1 Lecture 1: The Structure of Microarray Data So, Why are we here? What are we trying to measure, and how? Mechanics the data files produced and used Getting numbers: normalization
INTRODUCTION TO MICROARRAYS 1 Lecture 1: The Structure of Microarray Data So, Why are we here? What are we trying to measure, and how? Mechanics the data files produced and used Getting numbers: normalization
Systematic Variation in Genetic Microarray Data
 Biostatistics (2004), 1, 1, pp 1 47 Printed in Great Britain Systematic Variation in Genetic Microarray Data By KIMBERLY F SELLERS, JEFFREY MIECZNIKOWSKI, andwilliam F EDDY Department of Statistics, Carnegie
Biostatistics (2004), 1, 1, pp 1 47 Printed in Great Britain Systematic Variation in Genetic Microarray Data By KIMBERLY F SELLERS, JEFFREY MIECZNIKOWSKI, andwilliam F EDDY Department of Statistics, Carnegie
Technologie w skali genomowej 2/ Algorytmiczne i statystyczne aspekty sekwencjonowania DNA
 Technologie w skali genomowej 2/ Algorytmiczne i statystyczne aspekty sekwencjonowania DNA Expression analysis for RNA-seq data Ewa Szczurek Instytut Informatyki Uniwersytet Warszawski 1/35 The problem
Technologie w skali genomowej 2/ Algorytmiczne i statystyczne aspekty sekwencjonowania DNA Expression analysis for RNA-seq data Ewa Szczurek Instytut Informatyki Uniwersytet Warszawski 1/35 The problem
Processing microarray data with Bioconductor
 Processing microarray data with Bioconductor Statistical analysis of gene expression data with R and Bioconductor University of Copenhagen Copenhagen Biocenter Laurent Gautier 1, 2 August 17-21 2009 Contents
Processing microarray data with Bioconductor Statistical analysis of gene expression data with R and Bioconductor University of Copenhagen Copenhagen Biocenter Laurent Gautier 1, 2 August 17-21 2009 Contents
Design and Analysis of Gene Expression Experiments
 Design and Analysis of Gene Expression Experiments Guilherme J. M. Rosa Department of Animal Sciences Department of Biostatistics & Medical Informatics University of Wisconsin - Madison OUTLINE Æ Linear
Design and Analysis of Gene Expression Experiments Guilherme J. M. Rosa Department of Animal Sciences Department of Biostatistics & Medical Informatics University of Wisconsin - Madison OUTLINE Æ Linear
changes in gene expression, we developed and tested several models. Each model was
 Additional Files Additional File 1 File format: PDF Title: Experimental design and linear models Description: This additional file describes in detail the experimental design and linear models used to
Additional Files Additional File 1 File format: PDF Title: Experimental design and linear models Description: This additional file describes in detail the experimental design and linear models used to
Laurent Gautier 1, 2. August 18 th, 2009
 s Processing of Laurent Gautier 1, 2 1 DTU Multi-Assay Core(DMAC) 2 Center for Biological Sequence (CBS) August 18 th, 2009 s 1 s 2 3 4 s 1 s 2 3 4 Photolithography s Courtesy of The Science Creative Quarterly,
s Processing of Laurent Gautier 1, 2 1 DTU Multi-Assay Core(DMAC) 2 Center for Biological Sequence (CBS) August 18 th, 2009 s 1 s 2 3 4 s 1 s 2 3 4 Photolithography s Courtesy of The Science Creative Quarterly,
Experimental Design. Experimental design. Outline. Choice of platform Array design. Target samples
 Experimental Design Credit for some of today s materials: Jean Yang, Terry Speed, and Christina Kendziorski Experimental design Choice of platform rray design Creation of probes Location on the array Controls
Experimental Design Credit for some of today s materials: Jean Yang, Terry Speed, and Christina Kendziorski Experimental design Choice of platform rray design Creation of probes Location on the array Controls
ABSSeq: a new RNA-Seq analysis method based on modelling absolute expression differences
 ABSSeq: a new RNA-Seq analysis method based on modelling absolute expression differences Wentao Yang October 30, 2018 1 Introduction This vignette is intended to give a brief introduction of the ABSSeq
ABSSeq: a new RNA-Seq analysis method based on modelling absolute expression differences Wentao Yang October 30, 2018 1 Introduction This vignette is intended to give a brief introduction of the ABSSeq
Practical Statistics for the Analytical Scientist Table of Contents
 Practical Statistics for the Analytical Scientist Table of Contents Chapter 1 Introduction - Choosing the Correct Statistics 1.1 Introduction 1.2 Choosing the Right Statistical Procedures 1.2.1 Planning
Practical Statistics for the Analytical Scientist Table of Contents Chapter 1 Introduction - Choosing the Correct Statistics 1.1 Introduction 1.2 Choosing the Right Statistical Procedures 1.2.1 Planning
networks in molecular biology Wolfgang Huber
 networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
Design of Microarray Experiments. Xiangqin Cui
 Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Color Compensation Set 1
 Color Compensation Set 1 Version 2, March 2012 Solutions for the generation of color compensation objects for the LightCycler System. Order No. A 500 08 For 5 calibration runs. Store in dark at -15 to
Color Compensation Set 1 Version 2, March 2012 Solutions for the generation of color compensation objects for the LightCycler System. Order No. A 500 08 For 5 calibration runs. Store in dark at -15 to
Electronic Supplementary Information Increasing Hybridization Rate and Sensitivity of DNA Microarrays Using Isotachophoresis
 Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Increasing Hybridization Rate and Sensitivity of DNA
Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2014 Electronic Supplementary Information Increasing Hybridization Rate and Sensitivity of DNA
Estimation of Transformations for Microarray Data Using Maximum Likelihood and Related Methods
 Estimation of Transformations for Microarray Data Using Maximum Likelihood and Related Methods Blythe Durbin, Department of Statistics, UC Davis, Davis, CA 95616 David M. Rocke, Department of Applied Science,
Estimation of Transformations for Microarray Data Using Maximum Likelihood and Related Methods Blythe Durbin, Department of Statistics, UC Davis, Davis, CA 95616 David M. Rocke, Department of Applied Science,
Bioinformatics. Transcriptome
 Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
DATA ANALYSIS FOR MICROARRAY EXPERIMENT AND DNA BARCODE OF LIFE
 DATA ANALYSIS FOR MICROARRAY EXPERIMENT AND DNA BARCODE OF LIFE BY CHING-RAY YU A dissertation submitted to the Graduate School New Brunswick Rutgers, The State University of New Jersey in partial fulfillment
DATA ANALYSIS FOR MICROARRAY EXPERIMENT AND DNA BARCODE OF LIFE BY CHING-RAY YU A dissertation submitted to the Graduate School New Brunswick Rutgers, The State University of New Jersey in partial fulfillment
Practical Applications and Properties of the Exponentially. Modified Gaussian (EMG) Distribution. A Thesis. Submitted to the Faculty
 Practical Applications and Properties of the Exponentially Modified Gaussian (EMG) Distribution A Thesis Submitted to the Faculty of Drexel University by Scott Haney in partial fulfillment of the requirements
Practical Applications and Properties of the Exponentially Modified Gaussian (EMG) Distribution A Thesis Submitted to the Faculty of Drexel University by Scott Haney in partial fulfillment of the requirements
A calibration method for estimating absolute expression levels from microarray data
 BIOINFORMATICS ORIGINAL PAPER Vol. 22 no. 10 2006, pages 1251 1258 doi:10.1093/bioinformatics/btl068 Gene expression A calibration method for estimating absolute expression levels from microarray data
BIOINFORMATICS ORIGINAL PAPER Vol. 22 no. 10 2006, pages 1251 1258 doi:10.1093/bioinformatics/btl068 Gene expression A calibration method for estimating absolute expression levels from microarray data
Microarray Data Analysis - II. FIOCRUZ Bioinformatics Workshop 6 June, 2001
 Microarray Data Analysis - II FIOCRUZ Bioinformatics Workshop 6 June, 2001 Challenges in Microarray Data Analysis Spot Identification and Quantitation. Normalization of data from each experiment. Identification
Microarray Data Analysis - II FIOCRUZ Bioinformatics Workshop 6 June, 2001 Challenges in Microarray Data Analysis Spot Identification and Quantitation. Normalization of data from each experiment. Identification
Non-specific filtering and control of false positives
 Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Topics on statistical design and analysis. of cdna microarray experiment
 Topics on statistical design and analysis of cdna microarray experiment Ximin Zhu A Dissertation Submitted to the University of Glasgow for the degree of Doctor of Philosophy Department of Statistics May
Topics on statistical design and analysis of cdna microarray experiment Ximin Zhu A Dissertation Submitted to the University of Glasgow for the degree of Doctor of Philosophy Department of Statistics May
Proteomics. Areas of Interest
 Introduction to BioMEMS & Medical Microdevices Proteomics and Protein Microarrays Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices Proteomics and Protein Microarrays Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Statistics for Managers using Microsoft Excel 6 th Edition
 Statistics for Managers using Microsoft Excel 6 th Edition Chapter 3 Numerical Descriptive Measures 3-1 Learning Objectives In this chapter, you learn: To describe the properties of central tendency, variation,
Statistics for Managers using Microsoft Excel 6 th Edition Chapter 3 Numerical Descriptive Measures 3-1 Learning Objectives In this chapter, you learn: To describe the properties of central tendency, variation,
Multiplicative background correction for spotted. microarrays to improve reproducibility
 Multiplicative background correction for spotted microarrays to improve reproducibility DABAO ZHANG,, MIN ZHANG, MARTIN T. WELLS, March 12, 2006 Department of Statistics, Purdue University, West Lafayette,
Multiplicative background correction for spotted microarrays to improve reproducibility DABAO ZHANG,, MIN ZHANG, MARTIN T. WELLS, March 12, 2006 Department of Statistics, Purdue University, West Lafayette,
FEATURE SELECTION COMBINED WITH RANDOM SUBSPACE ENSEMBLE FOR GENE EXPRESSION BASED DIAGNOSIS OF MALIGNANCIES
 FEATURE SELECTION COMBINED WITH RANDOM SUBSPACE ENSEMBLE FOR GENE EXPRESSION BASED DIAGNOSIS OF MALIGNANCIES Alberto Bertoni, 1 Raffaella Folgieri, 1 Giorgio Valentini, 1 1 DSI, Dipartimento di Scienze
FEATURE SELECTION COMBINED WITH RANDOM SUBSPACE ENSEMBLE FOR GENE EXPRESSION BASED DIAGNOSIS OF MALIGNANCIES Alberto Bertoni, 1 Raffaella Folgieri, 1 Giorgio Valentini, 1 1 DSI, Dipartimento di Scienze
BMC Bioinformatics. Open Access. Abstract. Background Microarray technology has become a useful tool for quantitatively.
 BMC Bioinformatics BioMed Central Methodology article A robust two-way semi-linear model for normalization of cdna microarray data Deli Wang, Jian Huang, Hehuang Xie 3, Liliana Manzella 3 and Marcelo Bento
BMC Bioinformatics BioMed Central Methodology article A robust two-way semi-linear model for normalization of cdna microarray data Deli Wang, Jian Huang, Hehuang Xie 3, Liliana Manzella 3 and Marcelo Bento
Analyses biostatistiques de données RNA-seq
 Analyses biostatistiques de données RNA-seq Ignacio Gonzàlez, Annick Moisan & Nathalie Villa-Vialaneix prenom.nom@toulouse.inra.fr Toulouse, 18/19 mai 2017 IG, AM, NV 2 (INRA) Biostatistique RNA-seq Toulouse,
Analyses biostatistiques de données RNA-seq Ignacio Gonzàlez, Annick Moisan & Nathalie Villa-Vialaneix prenom.nom@toulouse.inra.fr Toulouse, 18/19 mai 2017 IG, AM, NV 2 (INRA) Biostatistique RNA-seq Toulouse,
MAIZE GENE EXPRESSION UV RESPONSE PATTERNS REVEAL COORDINATE REGULATION OF MANY GENES. Carletha R. Blanding
 MAIZE GENE EXPRESSION UV RESPONSE PATTERNS REVEAL COORDINATE REGULATION OF MANY GENES Carletha R. Blanding A Thesis Submitted to the University of North Carolina Wilmington in Partial Fulfillment Of the
MAIZE GENE EXPRESSION UV RESPONSE PATTERNS REVEAL COORDINATE REGULATION OF MANY GENES Carletha R. Blanding A Thesis Submitted to the University of North Carolina Wilmington in Partial Fulfillment Of the
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Three-Dimensional Electron Microscopy of Macromolecular Assemblies
 Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
EE/CpE 345. Modeling and Simulation. Fall Class 9
 EE/CpE 345 Modeling and Simulation Class 9 208 Input Modeling Inputs(t) Actual System Outputs(t) Parameters? Simulated System Outputs(t) The input data is the driving force for the simulation - the behavior
EE/CpE 345 Modeling and Simulation Class 9 208 Input Modeling Inputs(t) Actual System Outputs(t) Parameters? Simulated System Outputs(t) The input data is the driving force for the simulation - the behavior
RNASeq Differential Expression
 12/06/2014 RNASeq Differential Expression Le Corguillé v1.01 1 Introduction RNASeq No previous genomic sequence information is needed In RNA-seq the expression signal of a transcript is limited by the
12/06/2014 RNASeq Differential Expression Le Corguillé v1.01 1 Introduction RNASeq No previous genomic sequence information is needed In RNA-seq the expression signal of a transcript is limited by the
Answer keys for Assignment 10: Measurement of study variables (The correct answer is underlined in bold text)
 Answer keys for Assignment 10: Measurement of study variables (The correct answer is underlined in bold text) 1. A quick and easy indicator of dispersion is a. Arithmetic mean b. Variance c. Standard deviation
Answer keys for Assignment 10: Measurement of study variables (The correct answer is underlined in bold text) 1. A quick and easy indicator of dispersion is a. Arithmetic mean b. Variance c. Standard deviation
CONJOINT 541. Translating a Transcriptome at Specific Times and Places. David Morris. Department of Biochemistry
 CONJOINT 541 Translating a Transcriptome at Specific Times and Places David Morris Department of Biochemistry http://faculty.washington.edu/dmorris/ Lecture 1 The Biology and Experimental Analysis of mrna
CONJOINT 541 Translating a Transcriptome at Specific Times and Places David Morris Department of Biochemistry http://faculty.washington.edu/dmorris/ Lecture 1 The Biology and Experimental Analysis of mrna
EE/CpE 345. Modeling and Simulation. Fall Class 10 November 18, 2002
 EE/CpE 345 Modeling and Simulation Class 0 November 8, 2002 Input Modeling Inputs(t) Actual System Outputs(t) Parameters? Simulated System Outputs(t) The input data is the driving force for the simulation
EE/CpE 345 Modeling and Simulation Class 0 November 8, 2002 Input Modeling Inputs(t) Actual System Outputs(t) Parameters? Simulated System Outputs(t) The input data is the driving force for the simulation
Descriptive Data Summarization
 Descriptive Data Summarization Descriptive data summarization gives the general characteristics of the data and identify the presence of noise or outliers, which is useful for successful data cleaning
Descriptive Data Summarization Descriptive data summarization gives the general characteristics of the data and identify the presence of noise or outliers, which is useful for successful data cleaning
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Design of microarray experiments
 Design of microarray experiments Ulrich ansmann mansmann@imbi.uni-heidelberg.de Practical microarray analysis September Heidelberg Heidelberg, September otivation The lab biologist and theoretician need
Design of microarray experiments Ulrich ansmann mansmann@imbi.uni-heidelberg.de Practical microarray analysis September Heidelberg Heidelberg, September otivation The lab biologist and theoretician need
Statistics Toolbox 6. Apply statistical algorithms and probability models
 Statistics Toolbox 6 Apply statistical algorithms and probability models Statistics Toolbox provides engineers, scientists, researchers, financial analysts, and statisticians with a comprehensive set of
Statistics Toolbox 6 Apply statistical algorithms and probability models Statistics Toolbox provides engineers, scientists, researchers, financial analysts, and statisticians with a comprehensive set of
Evaluation of the performance of Amersham HyPer5 dye in comparative genomic hybridization microarray applications
 GE Healthcare Application note 28-9304-76 AA Nucleic acid labeling and detection Evaluation of the performance of Amersham HyPer5 dye in comparative genomic hybridization microarray applications Key words:
GE Healthcare Application note 28-9304-76 AA Nucleic acid labeling and detection Evaluation of the performance of Amersham HyPer5 dye in comparative genomic hybridization microarray applications Key words:
Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking
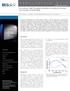 A p p l i c a t i o n N o t e Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking Peter J. Brescia, Jr., Chris Wilson, and Peter Banks, BioTek Instruments, Inc., Winooski,
A p p l i c a t i o n N o t e Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking Peter J. Brescia, Jr., Chris Wilson, and Peter Banks, BioTek Instruments, Inc., Winooski,
Wavelet-Based Preprocessing Methods for Mass Spectrometry Data
 Wavelet-Based Preprocessing Methods for Mass Spectrometry Data Jeffrey S. Morris Department of Biostatistics and Applied Mathematics UT M.D. Anderson Cancer Center Overview Background and Motivation Preprocessing
Wavelet-Based Preprocessing Methods for Mass Spectrometry Data Jeffrey S. Morris Department of Biostatistics and Applied Mathematics UT M.D. Anderson Cancer Center Overview Background and Motivation Preprocessing
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
Chapter 3: Statistical methods for estimation and testing. Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001).
 Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Fuzzy Clustering of Gene Expression Data
 Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Application Note # AN112 Failure analysis of packaging materials
 Application Note # AN112 Failure analysis of packaging materials Introduction Packaging materials are often composed of various layers that fulfill different functions. The actual packaging foil has a
Application Note # AN112 Failure analysis of packaging materials Introduction Packaging materials are often composed of various layers that fulfill different functions. The actual packaging foil has a
The Union and Intersection for Different Configurations of Two Events Mutually Exclusive vs Independency of Events
 Section 1: Introductory Probability Basic Probability Facts Probabilities of Simple Events Overview of Set Language Venn Diagrams Probabilities of Compound Events Choices of Events The Addition Rule Combinations
Section 1: Introductory Probability Basic Probability Facts Probabilities of Simple Events Overview of Set Language Venn Diagrams Probabilities of Compound Events Choices of Events The Addition Rule Combinations
g A n(a, g) n(a, ḡ) = n(a) n(a, g) n(a) B n(b, g) n(a, ḡ) = n(b) n(b, g) n(b) g A,B A, B 2 RNA-seq (D) RNA mrna [3] RNA 2. 2 NGS 2 A, B NGS n(
![g A n(a, g) n(a, ḡ) = n(a) n(a, g) n(a) B n(b, g) n(a, ḡ) = n(b) n(b, g) n(b) g A,B A, B 2 RNA-seq (D) RNA mrna [3] RNA 2. 2 NGS 2 A, B NGS n( g A n(a, g) n(a, ḡ) = n(a) n(a, g) n(a) B n(b, g) n(a, ḡ) = n(b) n(b, g) n(b) g A,B A, B 2 RNA-seq (D) RNA mrna [3] RNA 2. 2 NGS 2 A, B NGS n(](/thumbs/77/74495778.jpg) ,a) RNA-seq RNA-seq Cuffdiff, edger, DESeq Sese Jun,a) Abstract: Frequently used biological experiment technique for observing comprehensive gene expression has been changed from microarray using cdna
,a) RNA-seq RNA-seq Cuffdiff, edger, DESeq Sese Jun,a) Abstract: Frequently used biological experiment technique for observing comprehensive gene expression has been changed from microarray using cdna
Two-Color Microarray Experimental Design Notation. Simple Examples of Analysis for a Single Gene. Microarray Experimental Design Notation
 Simple Examples of Analysis for a Single Gene wo-olor Microarray Experimental Design Notation /3/0 opyright 0 Dan Nettleton Microarray Experimental Design Notation Microarray Experimental Design Notation
Simple Examples of Analysis for a Single Gene wo-olor Microarray Experimental Design Notation /3/0 opyright 0 Dan Nettleton Microarray Experimental Design Notation Microarray Experimental Design Notation
Supplementary Information
 Supplementary Information A versatile genome-scale PCR-based pipeline for high-definition DNA FISH Magda Bienko,, Nicola Crosetto,, Leonid Teytelman, Sandy Klemm, Shalev Itzkovitz & Alexander van Oudenaarden,,
Supplementary Information A versatile genome-scale PCR-based pipeline for high-definition DNA FISH Magda Bienko,, Nicola Crosetto,, Leonid Teytelman, Sandy Klemm, Shalev Itzkovitz & Alexander van Oudenaarden,,
Differential expression analysis for sequencing count data. Simon Anders
 Differential expression analysis for sequencing count data Simon Anders RNA-Seq Count data in HTS RNA-Seq Tag-Seq Gene 13CDNA73 A2BP1 A2M A4GALT AAAS AACS AADACL1 [...] ChIP-Seq Bar-Seq... GliNS1 4 19
Differential expression analysis for sequencing count data Simon Anders RNA-Seq Count data in HTS RNA-Seq Tag-Seq Gene 13CDNA73 A2BP1 A2M A4GALT AAAS AACS AADACL1 [...] ChIP-Seq Bar-Seq... GliNS1 4 19
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides
 Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
Microarray data analysis
 Microarray data analysis September 20, 2006 Jonathan Pevsner, Ph.D. Introduction to Bioinformatics pevsner@kennedykrieger.org Johns Hopkins School of Public Health (260.602.01) Copyright notice Many of
Microarray data analysis September 20, 2006 Jonathan Pevsner, Ph.D. Introduction to Bioinformatics pevsner@kennedykrieger.org Johns Hopkins School of Public Health (260.602.01) Copyright notice Many of
Linker Dependent Bond Rupture Force Measurements in Single-Molecule Junctions
 Supplemental Information Linker Dependent Bond Rupture Force Measurements in Single-Molecule Junctions M. Frei 1, S Aradhya 1, M. S. Hybertsen 2, L. Venkataraman 1 1 Department of Applied Physics and Applied
Supplemental Information Linker Dependent Bond Rupture Force Measurements in Single-Molecule Junctions M. Frei 1, S Aradhya 1, M. S. Hybertsen 2, L. Venkataraman 1 1 Department of Applied Physics and Applied
DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS
 Sankhyā : The Indian Journal of Statistics Special issue in memory of D. Basu 2002, Volume 64, Series A, Pt. 3, pp 706-720 DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS By TERENCE P. SPEED
Sankhyā : The Indian Journal of Statistics Special issue in memory of D. Basu 2002, Volume 64, Series A, Pt. 3, pp 706-720 DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS By TERENCE P. SPEED
BTRY 7210: Topics in Quantitative Genomics and Genetics
 BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu February 12, 2015 Lecture 3:
BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu February 12, 2015 Lecture 3:
Data Sheet. Azide Cy5 RNA T7 Transcription Kit
 Cat. No. Size 1. Description PP-501-Cy5 10 reactions à 40 µl For in vitro use only Quality guaranteed for 12 months Store all components at -20 C. Avoid freeze and thaw cycles. DBCO-Sulfo-Cy5 must be stored
Cat. No. Size 1. Description PP-501-Cy5 10 reactions à 40 µl For in vitro use only Quality guaranteed for 12 months Store all components at -20 C. Avoid freeze and thaw cycles. DBCO-Sulfo-Cy5 must be stored
