Estimation of Transformations for Microarray Data Using Maximum Likelihood and Related Methods
|
|
|
- Chloe Gibson
- 5 years ago
- Views:
Transcription
1 Estimation of Transformations for Microarray Data Using Maximum Likelihood and Related Methods Blythe Durbin, Department of Statistics, UC Davis, Davis, CA David M. Rocke, Department of Applied Science, UC Davis, Davis, CA September 9, 2002 Abstract Motivation and Results Durbin et al (2002), Huber et al (2002) and Munson (2001) independently introduced a family of transformations (the generalized-log family) which stabilizes the variance of microarray data up to the first order. We introduce a method for estimating the transformation parameter in tandem with a linear model based on the procedure outlined in Box and Cox (1964). We also discuss means of finding transformations within the generalized-log family which are optimal under other criteria, such as minimum residual skewness and minimum mean-variance dependency. Contact. bpdurbin@wald.ucdavis.edu Keywords. cdna array, microarray, statistical analysis, transformation, ANOVA normalization. 1 Introduction Many traditional statistical methodologies, such as regression or ANOVA, are based on the assumptions that the data are normally distributed (or at least symmetrically distributed), with constant variance not depending on the mean of the data. If these assumptions are violated, the statistician may choose either to develop some new statistical technique which accounts for the specific ways in which the data fail to comply with the assumptions, or to transform the data. Where possible, data transformation is generally the easier of these two options (see Box and Cox, 1964, and Atkinson, 1985). To whom correspondence should be addressed. 1
2 Data from gene-expression microarrays, which allow measurement of the expression of thousands of genes simultaneously, can yield invaluable information about biology through statistical analysis. However, microarray data fail rather dramatically to conform to the canonical assumptions required for analysis by standard techniques. Rocke and Durbin (2001) demonstrate that the measured expression levels from microarray data can be modeled as y = α + µe η + ε (1) where y is the measured raw expression level for a single color, α is the mean background noise, µ is the true expression level, and η and ε are normallydistributed error terms with mean 0 and variance σ 2 η and σ 2 ε, respectively. This model also works well for Affymetrix GeneChip arrays either applied to the PM-MM data or to individual oligos. The variance of y under this model is Var(y) = µ 2 S 2 η + σ 2 ε, (2) where Sη 2 = e σ2 η (e σ 2 η 1). In Durbin et al. (2002), Huber et al. (2002), and Munson (2001) it was shown that for a random variable z satisfying V (z) = a 2 +b 2 µ 2, with E(y) = µ, there is a transformation that stabilizes the variance to the first order. There are several equivalent ways of writing this transformation, but we will use h λ (z) = ln(z + z 2 + λ), where λ = a 2 /b 2 = σ 2 ε/s 2 η and z = y α or y ˆα. (Use of z rather than y presumes that any requisite background correction and normalization have already been applied to the data so that, to the first order, E(z) = µ. The specific method normalization method used is left to the discretion of the reader.) This transformation converges to ln(z) for large z (up to an additive constant which does not affect the strength of the transformation), and is approximately linear at 0 (Durbin et al. 2002). We shall refer to this transformation as the generalized-log transformation, as in Munson (2001), as the log transformation is a special case of this family for λ = 0. The inverse transformation is f 1 λ (w) = (ez λe z )/2. Both f λ and its inverse are monotonic functions, defined for all values of z and w, with derivatives of all orders. When transforming data from two-color arrays or from complex multi-array experiments, the closed form expression for the transformation parameter shown in (1) is less useful than in the single color, single array case. Even data from different colors on the same two-color array might have different estimated values for the model parameters ση 2 and σε, 2 which makes it unclear exactly how we should obtain the transformation parameter. Pooling of data from different sources in order to estimate parameters could work well for some designs, but is not very flexible. An estimation method which specifically accounts for the structure of the data would be useful in these situations. 2
3 One such approach is to fit a linear model to the data while simultaneously estimating the transformation parameter via maximum likelihood, as was done in Box and Cox (1964). The linear model structure will allow us to account for the different sources of variation in the data, such as variation between arrays, between replicated spots on the same array, and between colors on the same array, in our estimation of the transformation parameter. Furthermore, the linear model, fit to appropriately transformed data, can itself be a useful analysis tool. An example of such an analysis would be the ANOVA normalization method for microarray data developed in Kerr et al (2000). 2 Maximum-Likelihood Estimation The maximum-likelihood estimation of the linear model and transformation parameters can be conducted essentially as in Box and Cox (1964), with the key distinction being that we shall search for an optimal transformation within the family of generalized log transformations, as in (1), rather than among the power transformations. The procedure outlined in Box and Cox (1964) is as follows: Suppose that there exists some λ such that the transformed observations {h i,λ } have independent normal distributions with linear mean structure and constant variance σ 2. That is, suppose there exists lambda such that h λ = (h 1,λ,..., h n,λ ) = Xβ + ε (3) where n is the number of observations in the data set, X is the design matrix from the linear model, β is a fixed vector of unknown linear model parameters, and ε N(0, σ 2 I). The likelihood of the untransformed observations {z i } may therefore be written in terms of the transformed observations {h i,λ } as L(β, σ 2, λ; z) = 1 (2π) n/2 σ n exp ( (h λ Xβ) (h λ Xβ) 2σ 2 ) J(λ), (4) where J(λ) = = n dh i,λ dz i=1 i (5) n 1/ zi 2 + λ. (6) i=1 Fixing λ and maximizing (4) over β and σ 2, we arrive at the usual maximumlikelihood estimates for these parameters (presuming X is of full rank): ˆβ(λ) = (X X) 1 X h λ and ˆσ 2 (λ) = h λ (I H)h λ /n, (7) 3
4 where H = X(X X) 1 X (Box and Cox (1964)). Substituting ˆβ(λ) and ˆσ 2 (λ) into ln(l(β, σ 2, λ; z)), we obtain the partiallymaximized log likelihood lmax(λ) = n 2 ln ˆσ2 (λ) + ln J(λ). (8) For added simplicity, we may scale the transformed observations by J 1/n (λ), defining where w λ = h λ /J 1/n (λ) = ln(z + z 2 + λ)gm( z 2 + λ), gm( ( n ) 1/n z 2 + λ) = zi 2 + λ. The partially-maximized log likelihood of z in terms of w λ can then be closely approximated by i=1 lmax(λ) = n 2 ln ˆσ2 (λ) = n 2 SSE(λ)/n which depends on the data only through SSE(λ), the error sum of squares from the linear model fit to the transformed data. (The approximate nature of the log likelihood arises when we choose to ignore the variability of the Jacobian, which should be quite minimal given the size of most microarray data sets. See Chapter 6 of Ferguson (1996) for further explanation.) This new, partiallymaximized log likelihood is a monotone decreasing function of the error sum of squares from the linear model for fixed λ, SSE(λ), so we may find the MLE of λ, ˆλ, simply by minimizing SSE(λ) (Box and Cox (1964)). This may be accomplished by plotting the error sum of squares as a function of λ, or via numerical optimization methods. Estimates of β and σ 2 on the scale of the transformed data without the Jacobian correction may be obtained by fitting the linear model again using the MLE, ˆλ, as the transformation parameter or by multiplying ˆβ by J 1/n (ˆλ) and multiplying ˆσ 2 by J 2/n (ˆλ). 2.1 Examples We illustrate this estimation method using two example data sets, one from a two-color cdna-array experiment and one from an experiment conducted using Affymetrix oligonucleotide arrays. The first example comes from a toxicology experiment by Bartosiewicz et al (2002) in which male Swiss Webster mice were injected with a toxin. We shall focus on a single slide from this experiment. For this array, the treatment mouse was injected with 0.15 mg/kg of β-napthoflavone dissolved in 10 ml/kg of corn oil, and the control mouse was injected with 10 4
5 ml/kg of corn oil. mrna from the livers of these mice was reverse transcribed and fluor labelled, with the treatment sample labelled with Cy5 and the control sample labelled with Cy3. The samples were hybridized to a spotted cdna array on which each of the 138 genes was replicated between 6 and 14 times, resulting in a total of 1008 spots. For the mouse data, we will model the differences of the transformed control and treatment observations rather than the transformed observations themselves. The difference of the transformed observations from replicate j of gene i, h λij, can be modeled as h λij = µ i + ε ij, (9) where µ ij is a gene effect and ε ij is a normally distributed error term. Notice that (9) is a one-way ANOVA model. Figure 1 shows the partially-maximized log likelihood for the mouse data as a function of the transformation parameter, λ. The likelihood is maximized at λ = An asymptotic 95% confidence interval for the MLE, ˆλ, consists of those values of λ for which lmax(ˆλ) lmax(λ) < 1 2 χ2 1,.05, where χ 2 1,.05 is the upper 5% quantile of a χ 2 1 distribution (Box and Cox (1964)). This yields a confidence interval for ˆλ of ( , ), which is bounded well away from 0 and thus definitively excludes the log transformation (corresponding to λ = 0). Figure 2 shows a quantile-quantile plot of the residuals from the linear model (9) fit to the transformed data versus a standard normal distribution. The residuals appear to come from a distribution with heavier tails than a normal distribution. Although the plot appears to exhibit some skewness, this is entirely due to the four observations in the lower left-hand corner. These observations appear to be outliers resulting from phenomena such as dust on the slide, since they all occur in genes which are expressed near background in the control data, and feature a single observation which differs so hugely from the other replicates that it is unlikely to result from actual gene expression. (These observations will be excluded from the analysis of Section 3). Examination of residuals from the linear model appears to facilitate identification of outlying observations, since these outliers were much more obvious in the residuals than they would have been in the raw data. The second example comes from an experiment using 4 Affymetrix HG U95 arrays, which is described in Geller et al (2002). In this experiment, a lymphoblastoid cell line from a single autistic child was grown up in four separate flasks. RNA extraction, cdna synthesis, and in-vitro labelling was conducted separately on each of the 4 samples, and each of the samples was hybridized to a separate array. For the autism data, we model the transformed perfect match minus mismatch observation from gene j on array i, h λij, as h λij = µ i + η j + ε ij, (10) 5
6 where µ i is a fixed array effect, η j is a fixed gene effect, and ε ij is a normally distributed error term. Notice that our model is a two-factor ANOVA model without an interaction term. (We cannot fit the interaction term due to the absence of replicated genes, but we would not expect a gene-array interaction effect anyway.) Figure 3 shows the partially-maximized log likelihood for the autism data as a function of the transformation parameter. The likelihood is maximized at λ = 3873, and a 95% confidence interval for ˆλ is (3751, 4000), which, again, excludes the log transformation. Figure 4 shows a quantile-quantile plot of the residuals from the linear model (10). The residuals, again, appear to come from a symmetric distribution with tails heavier than a normal distribution. 3 Other Methods of Estimating the Transformation Parameter Maximum-likelihood estimation of the transformation parameter in the manner described above in essence simultaneously optimizes constancy of variance, the fit of the transformed residuals to a normal distribution, and the fit to the linear model. In some applications, some of these criteria may be more important than others. For example, for many traditional statistical techniques data that are symmetric are almost as good as data that are normally distributed, and by trying to force the transformed data to fit all of the moments of a normal distribution we may inadvertently compromise those characteristics in which we are most interested. In such cases, we may search within the family of generalized-log transformations for a transformation optimizing the quantity of interest, simply by minimizing the appropriate statistic. For example, to find a transformation minimizing the skewness of residuals from the linear model, we would look for a transformation for which the skewness coefficient of the residuals is equal to 0. To find a transformation for which the fixed effects in an ANOVA model are the most linear, we would look for a transformation minimizing the F-statistic for the interaction term in the model. (Notice that the two estimation procedures just mentioned both incorporate the linear model structure used in the maximum-likelihood estimation.) To find a transformation minimizing the dependency of the replicate mean on the replicate variance, we would regress the replicate standard deviation of the transformed data on the replicate mean and look for the transformation minimizing the t-statistic for the significance of the slope parameter. These other optimal transformations also provide a means of assessing the quality of the maximum-likelihood estimate of the transformation parameter. If the MLE differs too greatly from the optimal transformation parameter under another criterion, this might be cause for concern. We illustrate the skewness-minimizing transformation and the transformation minimizing dependency of the replicate mean and variance using the mouse data. Figure 5 shows a plot of the residual skewness coefficient as a function 6
7 of the transformation parameter for the mouse data, using residuals from the linear model in (9). For these data, the skewness coefficient is non-monotonic in the transformation parameter, so there are two values of λ for which the skewness coefficient is equal to 0, and A asymptotic 95% confidence interval for the skewness-minimizing transformation consists of those values of λ for which the absolute skewness coefficient is not significant at the 5% level. For a sample of size 1008 the absolute skewness is not statistically significant if it is less than , which yields the confidence interval ( , ). The resulting confidence interval is quite large, but notice that it excludes the log transformation (for λ = 0), indicating that the log transformation significantly skews the residuals. It does, however, include the maximum likelihood transformation, (λ = ), implying the the MLE does provide sufficient symmetry. Figure 6 shows the t-statistic for the significance of the slope parameter from the regression of the replicate standard deviation on the replicate mean as a function of the transformation parameter. For the mouse data, the t-statistic is equal to 0 at λ = An asymptotic 95% confidence interval for the t-minimizing transformation consists of those values of λ for which the t-statistic is not significant at the 5% level. For 136 degrees of freedom (since we have 138 genes and lose 2 degrees of freedom from fitting the regression parameters) the cutoff for significance of the t-statistic is ±1.9776, which yields the confidence interval ( , ). This confidence interval excludes both the log transformation and the maximum-likelihood transformation. However, a Wald test to determine if the MLE and the t-minimizing parameters differed significantly (using 1/4th the length of the respective confidence intervals as a crude estimate of the standard deviation of the parameters) yields the test statistic , which is not significant at the 5% level. Figure 7 shows the replicate standard deviation plotted against the replicate mean for the mouse data transformed using the t-minimizing transformation, λ = The plot does not display any obvious mean-variance dependency, indicating that the transformation has effectively stabilized the variance of these data. For purposes of comparison, Figure 8 shows the replicate standard deviation and mean for data transformed using the maximum-likelihood transformation, λ = Again, there does not appear to be any obvious mean-variance dependency, indicating that the MLE also provides sufficient variance stabilization. The MLE provides surprisingly good variance-stabilization and symmetrization, especially in light of the fact that the normal likelihood is almost certainly the wrong likelihood for the transformed data, given the apparent heavy-tailed distribution of the transformed residuals. 4 Conclusions The generalized-log transformation of Durbin et al (2002), Huber et al (2002) and Munson (2001) with parameter λ = a 2 /b 2 stabilizes the variance of data where Var(z) = a 2 + b 2 E 2 (z). Maximum-likelihood estimation in the manner of 7
8 Box and Cox (1964) can be used to estimate a transformation parameter for data where observations have different values of a and b. This procedure estimates the transformation parameter while simultaneously fitting a linear model to the data. The maximum-likelihood estimate can be found by finding the transformation minimizing the error sum of squares from the linear model fit to the transformed data. Transformations minimizing residual skewness, mean-variance dependency, and other criteria may be found by minimizing the appropriate statistic. The maximum likelihood estimate appears to perform well compared to transformations specifically minimizing residual skewness and mean-variance dependency, especially in light of the fact that the normal likelihood is a first approximation to the true distribution of the transformed data and was chosen primarily for computational convenience. The residuals from linear model fit to the transformed data appear, in fact, to come from a distribution with heavier tails than a normal distribution. Finally, the log transformation was excluded from the confidence intervals for each of the 3 optimality criteria tested (maximum likelihood, minimum residual skewness, and minimum mean-variance dependency). The generalized-log transformation provides a good alternative to the log transformation for use with gene-expression microarray data. A Numerical Optimization Via Newton s Method Newton s method provides a means of finding a root of a smooth function. We use this technique to perform numerical minimization of the error sum of squares by finding a root of the first derivative of SSE(λ). Plots of the likelihood may be used to confirm that the root found does indeed constitute a global maximum. Denote λ SSE(λ) by SSE (λ) and 2 λ SSE(λ) by SSE (λ). A new estimate of 2 λ, λ (n+1), may be obtained from the previous estimate, λ (n) using λ (n+1) = λ (n) SSE (λ (n) ) SSE (λ (n) ). (11) Convergence is achieved when SSE (λ) is less than the predetermined application tolerance. For the generalized log transformation with parameter λ, n SSE (λ) = 2 (w λi ŵ λi )(w λi ŵ λi), (12) where x i i=1 ŵ λi = x i is the i th row of the design matrix, ˆβ(λ), w λi = λ w λi = [2{z i + z 2 i + λ} z 2 i + λ] 1 J 1/n (λ) + ln(z i + z 2 i + λ) λ J 1/n (λ), 8
9 where and λ J 1/n (λ) = 1 n n J 1/n (λ)/{2(zi 2 + λ)}, i=1 ŵ λi = λŵλi = x i (X X) 1 X w λ, The second derivative of the error sum of squares is SSE (λ) = n (w λi ŵ λi) 2 i=1 n i=1 (w λi ŵ λi )(w λi ŵ λi), where w λi = 2 λ 2 w λi = 1 4 J 1/n (λ)(z i + z 2 i + λ) 1 {z 2 i + λ} J 1/n (λ)(z i + z 2 i + λ) 2 {z 2 i + λ} 1 +(z i + z 2 i + λ) 1 {z 2 i + λ} 1 2 λ J 1/n and and + ln(z i + z 2 i + λ) 2 λ 2 J 1/n 2 λ J 1/n (λ) = 1 n 2 2n i=1 {z2 i + λ} 1 λ J 1/n (λ) 1 2n n i=1 {z2 i + λ} 2 J 1/n (λ) Acknowledgements ŵ λi = 2 λ 2 w λi = x i (X X) 1 X w λ. The research reported in this paper was supported by grants from the National Science Foundation (ACI , and DMS ) and the National Institute of Environmental Health Sciences, National Institutes of Health (P43 ES04699). 9
10 References Atkinson, A.C. (1985) Plots, Transformations, and Regression: An Introduction to Graphical Methods of Diagnostic Regression Analysis Clarendon Press, Oxford. Bartosiewicz, M., Trounstine, M., Barker, D., Johnston, R., and Buckpitt, A. (2000) Development of a toxicological gene array and quantitative assessment of this technology. Archives of Biochemistry and Biophysics., 376, Box, G.E.P., and Cox, D.R. (1964) An analysis of transformations. Journal of the Royal Statistical Society, Series B (Methodological), 26, Durbin, B., Hardin, J., Hawkins, D.M., and Rocke, D.M. (2002) A variancestabilizing transformation for gene-expression microarray data. Bioinformatics, 18, 105S 110S. Ferguson, T.S. (1996) A Course in Large Sample Theory Chapman & Hall, London. Geller, S.C., Gregg, J.P., Hagermann, P., and Rocke, D.M. (2002) Transformation and normalization of oligonucleotide microarray data. (manuscript) Huber, W., von Heydebreck, A., Sültmann, H., Poustka, A., and Vingron, M. (2002) Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics, 18, 96S 104S. Kerr, K., Martin, M., and Churchill, G. (2000) Analysis of variance for gene expression microarray data. Journal of Computational Biology, 7, Munson, P. (2001) A consistency test for determining the significance of gene expression changes on replicate samples and two convenient variance-stabilizing transformations. GeneLogic Workshop of Low Level Analysis of Affymetrix GeneChip Data. Rocke, D.M., and Durbin, B. (2001) A model for measurement error for gene expression arrays. Journal of Computational Biology., 8,
11 0.996 x 104 Figure 1: Log Likelihood by Transformation Parameter, Mouse Data Partially Maximized Log Likelihood Likelihood is maximized at λ = 1.13 X % CI for λ is (8.7 X 10 8, 1.5 X 10 9 ) λ x 10 9
12 Standard Normal Quantiles Figure 2: QQ Plot of Residuals vs. Standard Normal, Maximum Likelihood Transformation, Mouse Data Residual Quantiles
13 1.315 x 105 Figure 3: Log Likelihood by Transformation Parameter, Autism Data 1.32 Likelihood is maximized at λ = 3873 Partially Maximized Log Likelihood % CI for λ is (3751, 4000) λ x 10 4
14 Standard Normal Quantiles 1.5 Figure 4: QQ Plot of Residuals from Linear Model vs. Standard Normal, Mouse Data Residual Quantiles
15 0.025 Figure 5: Residual Skewness by Transformation Parameter, Mouse Data Residual Skewness % CI for λ with 0 skewness is (1.2 X 10 6, 1.7 X 10 9 ) Skewness is 0 at 2.4 X 10 7 and 1.7 X λ x 10 8
16 2 Figure 6: t Statistic for Mean Variance Dependency by Transformation Parameter, Mouse Data 1 0 t statistic 1 T statistic is 0 at 4.0 X % CI for λ with no mean variance dependency is (2.0 X 10 9, 8.1 X 10 9 ) λ x 10 9
17 Replicate Mean 0.35 Figure 7: Replicate Mean and Standard Deviation, t Minimizing Transformation, Mouse Data Replicate Standard Deviation
18 Replicate Mean Figure 8: Replicate Mean and Standard Deviation, Maximum Likelihood Transformation, Mouse Data Replicate Standard Deviation
A variance-stabilizing transformation for gene-expression microarray data
 BIOINFORMATICS Vol. 18 Suppl. 1 00 Pages S105 S110 A variance-stabilizing transformation for gene-expression microarray data B. P. Durbin 1, J. S. Hardin, D. M. Hawins 3 and D. M. Roce 4 1 Department of
BIOINFORMATICS Vol. 18 Suppl. 1 00 Pages S105 S110 A variance-stabilizing transformation for gene-expression microarray data B. P. Durbin 1, J. S. Hardin, D. M. Hawins 3 and D. M. Roce 4 1 Department of
Bioconductor Project Working Papers
 Bioconductor Project Working Papers Bioconductor Project Year 2004 Paper 6 Error models for microarray intensities Wolfgang Huber Anja von Heydebreck Martin Vingron Department of Molecular Genome Analysis,
Bioconductor Project Working Papers Bioconductor Project Year 2004 Paper 6 Error models for microarray intensities Wolfgang Huber Anja von Heydebreck Martin Vingron Department of Molecular Genome Analysis,
Low-Level Analysis of High- Density Oligonucleotide Microarray Data
 Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Chapter 3: Statistical methods for estimation and testing. Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001).
 Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Normalization. Example of Replicate Data. Biostatistics Rafael A. Irizarry
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
Generalized Linear Models
 Generalized Linear Models Lecture 3. Hypothesis testing. Goodness of Fit. Model diagnostics GLM (Spring, 2018) Lecture 3 1 / 34 Models Let M(X r ) be a model with design matrix X r (with r columns) r n
Generalized Linear Models Lecture 3. Hypothesis testing. Goodness of Fit. Model diagnostics GLM (Spring, 2018) Lecture 3 1 / 34 Models Let M(X r ) be a model with design matrix X r (with r columns) r n
Outline of GLMs. Definitions
 Outline of GLMs Definitions This is a short outline of GLM details, adapted from the book Nonparametric Regression and Generalized Linear Models, by Green and Silverman. The responses Y i have density
Outline of GLMs Definitions This is a short outline of GLM details, adapted from the book Nonparametric Regression and Generalized Linear Models, by Green and Silverman. The responses Y i have density
Math 5305 Notes. Diagnostics and Remedial Measures. Jesse Crawford. Department of Mathematics Tarleton State University
 Math 5305 Notes Diagnostics and Remedial Measures Jesse Crawford Department of Mathematics Tarleton State University (Tarleton State University) Diagnostics and Remedial Measures 1 / 44 Model Assumptions
Math 5305 Notes Diagnostics and Remedial Measures Jesse Crawford Department of Mathematics Tarleton State University (Tarleton State University) Diagnostics and Remedial Measures 1 / 44 Model Assumptions
Linear models and their mathematical foundations: Simple linear regression
 Linear models and their mathematical foundations: Simple linear regression Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term 2018/19 1/21 Introduction
Linear models and their mathematical foundations: Simple linear regression Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term 2018/19 1/21 Introduction
Statistics for analyzing and modeling precipitation isotope ratios in IsoMAP
 Statistics for analyzing and modeling precipitation isotope ratios in IsoMAP The IsoMAP uses the multiple linear regression and geostatistical methods to analyze isotope data Suppose the response variable
Statistics for analyzing and modeling precipitation isotope ratios in IsoMAP The IsoMAP uses the multiple linear regression and geostatistical methods to analyze isotope data Suppose the response variable
Regression diagnostics
 Regression diagnostics Kerby Shedden Department of Statistics, University of Michigan November 5, 018 1 / 6 Motivation When working with a linear model with design matrix X, the conventional linear model
Regression diagnostics Kerby Shedden Department of Statistics, University of Michigan November 5, 018 1 / 6 Motivation When working with a linear model with design matrix X, the conventional linear model
Problems. Suppose both models are fitted to the same data. Show that SS Res, A SS Res, B
 Simple Linear Regression 35 Problems 1 Consider a set of data (x i, y i ), i =1, 2,,n, and the following two regression models: y i = β 0 + β 1 x i + ε, (i =1, 2,,n), Model A y i = γ 0 + γ 1 x i + γ 2
Simple Linear Regression 35 Problems 1 Consider a set of data (x i, y i ), i =1, 2,,n, and the following two regression models: y i = β 0 + β 1 x i + ε, (i =1, 2,,n), Model A y i = γ 0 + γ 1 x i + γ 2
Lectures on Simple Linear Regression Stat 431, Summer 2012
 Lectures on Simple Linear Regression Stat 43, Summer 0 Hyunseung Kang July 6-8, 0 Last Updated: July 8, 0 :59PM Introduction Previously, we have been investigating various properties of the population
Lectures on Simple Linear Regression Stat 43, Summer 0 Hyunseung Kang July 6-8, 0 Last Updated: July 8, 0 :59PM Introduction Previously, we have been investigating various properties of the population
Error models and normalization. Wolfgang Huber DKFZ Heidelberg
 Error models and normalization Wolfgang Huber DKFZ Heidelberg Acknowledgements Anja von Heydebreck, Martin Vingron Andreas Buness, Markus Ruschhaupt, Klaus Steiner, Jörg Schneider, Katharina Finis, Anke
Error models and normalization Wolfgang Huber DKFZ Heidelberg Acknowledgements Anja von Heydebreck, Martin Vingron Andreas Buness, Markus Ruschhaupt, Klaus Steiner, Jörg Schneider, Katharina Finis, Anke
Formal Statement of Simple Linear Regression Model
 Formal Statement of Simple Linear Regression Model Y i = β 0 + β 1 X i + ɛ i Y i value of the response variable in the i th trial β 0 and β 1 are parameters X i is a known constant, the value of the predictor
Formal Statement of Simple Linear Regression Model Y i = β 0 + β 1 X i + ɛ i Y i value of the response variable in the i th trial β 0 and β 1 are parameters X i is a known constant, the value of the predictor
Probe-Level Analysis of Affymetrix GeneChip Microarray Data
 Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley University of Minnesota Mar 30, 2004 Outline
Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley University of Minnesota Mar 30, 2004 Outline
Categorical Predictor Variables
 Categorical Predictor Variables We often wish to use categorical (or qualitative) variables as covariates in a regression model. For binary variables (taking on only 2 values, e.g. sex), it is relatively
Categorical Predictor Variables We often wish to use categorical (or qualitative) variables as covariates in a regression model. For binary variables (taking on only 2 values, e.g. sex), it is relatively
Diagnostics and Remedial Measures
 Diagnostics and Remedial Measures Yang Feng http://www.stat.columbia.edu/~yangfeng Yang Feng (Columbia University) Diagnostics and Remedial Measures 1 / 72 Remedial Measures How do we know that the regression
Diagnostics and Remedial Measures Yang Feng http://www.stat.columbia.edu/~yangfeng Yang Feng (Columbia University) Diagnostics and Remedial Measures 1 / 72 Remedial Measures How do we know that the regression
Expression arrays, normalization, and error models
 1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
Simple Linear Regression
 Simple Linear Regression September 24, 2008 Reading HH 8, GIll 4 Simple Linear Regression p.1/20 Problem Data: Observe pairs (Y i,x i ),i = 1,...n Response or dependent variable Y Predictor or independent
Simple Linear Regression September 24, 2008 Reading HH 8, GIll 4 Simple Linear Regression p.1/20 Problem Data: Observe pairs (Y i,x i ),i = 1,...n Response or dependent variable Y Predictor or independent
Approximate Median Regression via the Box-Cox Transformation
 Approximate Median Regression via the Box-Cox Transformation Garrett M. Fitzmaurice,StuartR.Lipsitz, and Michael Parzen Median regression is used increasingly in many different areas of applications. The
Approximate Median Regression via the Box-Cox Transformation Garrett M. Fitzmaurice,StuartR.Lipsitz, and Michael Parzen Median regression is used increasingly in many different areas of applications. The
[y i α βx i ] 2 (2) Q = i=1
![[y i α βx i ] 2 (2) Q = i=1 [y i α βx i ] 2 (2) Q = i=1](/thumbs/75/72005858.jpg) Least squares fits This section has no probability in it. There are no random variables. We are given n points (x i, y i ) and want to find the equation of the line that best fits them. We take the equation
Least squares fits This section has no probability in it. There are no random variables. We are given n points (x i, y i ) and want to find the equation of the line that best fits them. We take the equation
Analysis of 2 n Factorial Experiments with Exponentially Distributed Response Variable
 Applied Mathematical Sciences, Vol. 5, 2011, no. 10, 459-476 Analysis of 2 n Factorial Experiments with Exponentially Distributed Response Variable S. C. Patil (Birajdar) Department of Statistics, Padmashree
Applied Mathematical Sciences, Vol. 5, 2011, no. 10, 459-476 Analysis of 2 n Factorial Experiments with Exponentially Distributed Response Variable S. C. Patil (Birajdar) Department of Statistics, Padmashree
MA 575 Linear Models: Cedric E. Ginestet, Boston University Mixed Effects Estimation, Residuals Diagnostics Week 11, Lecture 1
 MA 575 Linear Models: Cedric E Ginestet, Boston University Mixed Effects Estimation, Residuals Diagnostics Week 11, Lecture 1 1 Within-group Correlation Let us recall the simple two-level hierarchical
MA 575 Linear Models: Cedric E Ginestet, Boston University Mixed Effects Estimation, Residuals Diagnostics Week 11, Lecture 1 1 Within-group Correlation Let us recall the simple two-level hierarchical
Lesson 11. Functional Genomics I: Microarray Analysis
 Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Advanced Quantitative Methods: maximum likelihood
 Advanced Quantitative Methods: Maximum Likelihood University College Dublin 4 March 2014 1 2 3 4 5 6 Outline 1 2 3 4 5 6 of straight lines y = 1 2 x + 2 dy dx = 1 2 of curves y = x 2 4x + 5 of curves y
Advanced Quantitative Methods: Maximum Likelihood University College Dublin 4 March 2014 1 2 3 4 5 6 Outline 1 2 3 4 5 6 of straight lines y = 1 2 x + 2 dy dx = 1 2 of curves y = x 2 4x + 5 of curves y
Correlation and Regression
 Correlation and Regression October 25, 2017 STAT 151 Class 9 Slide 1 Outline of Topics 1 Associations 2 Scatter plot 3 Correlation 4 Regression 5 Testing and estimation 6 Goodness-of-fit STAT 151 Class
Correlation and Regression October 25, 2017 STAT 151 Class 9 Slide 1 Outline of Topics 1 Associations 2 Scatter plot 3 Correlation 4 Regression 5 Testing and estimation 6 Goodness-of-fit STAT 151 Class
Introduction to General and Generalized Linear Models
 Introduction to General and Generalized Linear Models Generalized Linear Models - part II Henrik Madsen Poul Thyregod Informatics and Mathematical Modelling Technical University of Denmark DK-2800 Kgs.
Introduction to General and Generalized Linear Models Generalized Linear Models - part II Henrik Madsen Poul Thyregod Informatics and Mathematical Modelling Technical University of Denmark DK-2800 Kgs.
Least Absolute Value vs. Least Squares Estimation and Inference Procedures in Regression Models with Asymmetric Error Distributions
 Journal of Modern Applied Statistical Methods Volume 8 Issue 1 Article 13 5-1-2009 Least Absolute Value vs. Least Squares Estimation and Inference Procedures in Regression Models with Asymmetric Error
Journal of Modern Applied Statistical Methods Volume 8 Issue 1 Article 13 5-1-2009 Least Absolute Value vs. Least Squares Estimation and Inference Procedures in Regression Models with Asymmetric Error
Generalized Linear Models. Kurt Hornik
 Generalized Linear Models Kurt Hornik Motivation Assuming normality, the linear model y = Xβ + e has y = β + ε, ε N(0, σ 2 ) such that y N(μ, σ 2 ), E(y ) = μ = β. Various generalizations, including general
Generalized Linear Models Kurt Hornik Motivation Assuming normality, the linear model y = Xβ + e has y = β + ε, ε N(0, σ 2 ) such that y N(μ, σ 2 ), E(y ) = μ = β. Various generalizations, including general
Regression. Machine Learning and Pattern Recognition. Chris Williams. School of Informatics, University of Edinburgh.
 Regression Machine Learning and Pattern Recognition Chris Williams School of Informatics, University of Edinburgh September 24 (All of the slides in this course have been adapted from previous versions
Regression Machine Learning and Pattern Recognition Chris Williams School of Informatics, University of Edinburgh September 24 (All of the slides in this course have been adapted from previous versions
LOGISTIC REGRESSION Joseph M. Hilbe
 LOGISTIC REGRESSION Joseph M. Hilbe Arizona State University Logistic regression is the most common method used to model binary response data. When the response is binary, it typically takes the form of
LOGISTIC REGRESSION Joseph M. Hilbe Arizona State University Logistic regression is the most common method used to model binary response data. When the response is binary, it typically takes the form of
Multivariate Linear Models
 Multivariate Linear Models Stanley Sawyer Washington University November 7, 2001 1. Introduction. Suppose that we have n observations, each of which has d components. For example, we may have d measurements
Multivariate Linear Models Stanley Sawyer Washington University November 7, 2001 1. Introduction. Suppose that we have n observations, each of which has d components. For example, we may have d measurements
SCHOOL OF MATHEMATICS AND STATISTICS. Linear and Generalised Linear Models
 SCHOOL OF MATHEMATICS AND STATISTICS Linear and Generalised Linear Models Autumn Semester 2017 18 2 hours Attempt all the questions. The allocation of marks is shown in brackets. RESTRICTED OPEN BOOK EXAMINATION
SCHOOL OF MATHEMATICS AND STATISTICS Linear and Generalised Linear Models Autumn Semester 2017 18 2 hours Attempt all the questions. The allocation of marks is shown in brackets. RESTRICTED OPEN BOOK EXAMINATION
Microarray Preprocessing
 Microarray Preprocessing Normaliza$on Normaliza$on is needed to ensure that differences in intensi$es are indeed due to differen$al expression, and not some prin$ng, hybridiza$on, or scanning ar$fact.
Microarray Preprocessing Normaliza$on Normaliza$on is needed to ensure that differences in intensi$es are indeed due to differen$al expression, and not some prin$ng, hybridiza$on, or scanning ar$fact.
Hypothesis Testing hypothesis testing approach
 Hypothesis Testing In this case, we d be trying to form an inference about that neighborhood: Do people there shop more often those people who are members of the larger population To ascertain this, we
Hypothesis Testing In this case, we d be trying to form an inference about that neighborhood: Do people there shop more often those people who are members of the larger population To ascertain this, we
Module 6: Model Diagnostics
 St@tmaster 02429/MIXED LINEAR MODELS PREPARED BY THE STATISTICS GROUPS AT IMM, DTU AND KU-LIFE Module 6: Model Diagnostics 6.1 Introduction............................... 1 6.2 Linear model diagnostics........................
St@tmaster 02429/MIXED LINEAR MODELS PREPARED BY THE STATISTICS GROUPS AT IMM, DTU AND KU-LIFE Module 6: Model Diagnostics 6.1 Introduction............................... 1 6.2 Linear model diagnostics........................
STATISTICS 141 Final Review
 STATISTICS 141 Final Review Bin Zou bzou@ualberta.ca Department of Mathematical & Statistical Sciences University of Alberta Winter 2015 Bin Zou (bzou@ualberta.ca) STAT 141 Final Review Winter 2015 1 /
STATISTICS 141 Final Review Bin Zou bzou@ualberta.ca Department of Mathematical & Statistical Sciences University of Alberta Winter 2015 Bin Zou (bzou@ualberta.ca) STAT 141 Final Review Winter 2015 1 /
Semester 2, 2015/2016
 ECN 3202 APPLIED ECONOMETRICS 2. Simple linear regression B Mr. Sydney Armstrong Lecturer 1 The University of Guyana 1 Semester 2, 2015/2016 PREDICTION The true value of y when x takes some particular
ECN 3202 APPLIED ECONOMETRICS 2. Simple linear regression B Mr. Sydney Armstrong Lecturer 1 The University of Guyana 1 Semester 2, 2015/2016 PREDICTION The true value of y when x takes some particular
Spring 2017 Econ 574 Roger Koenker. Lecture 14 GEE-GMM
 University of Illinois Department of Economics Spring 2017 Econ 574 Roger Koenker Lecture 14 GEE-GMM Throughout the course we have emphasized methods of estimation and inference based on the principle
University of Illinois Department of Economics Spring 2017 Econ 574 Roger Koenker Lecture 14 GEE-GMM Throughout the course we have emphasized methods of estimation and inference based on the principle
Statistics 203: Introduction to Regression and Analysis of Variance Penalized models
 Statistics 203: Introduction to Regression and Analysis of Variance Penalized models Jonathan Taylor - p. 1/15 Today s class Bias-Variance tradeoff. Penalized regression. Cross-validation. - p. 2/15 Bias-variance
Statistics 203: Introduction to Regression and Analysis of Variance Penalized models Jonathan Taylor - p. 1/15 Today s class Bias-Variance tradeoff. Penalized regression. Cross-validation. - p. 2/15 Bias-variance
Seminar Microarray-Datenanalyse
 Seminar Microarray- Normalization Hans-Ulrich Klein Christian Ruckert Institut für Medizinische Informatik WWU Münster SS 2011 Organisation 1 09.05.11 Normalisierung 2 10.05.11 Bestimmen diff. expr. Gene,
Seminar Microarray- Normalization Hans-Ulrich Klein Christian Ruckert Institut für Medizinische Informatik WWU Münster SS 2011 Organisation 1 09.05.11 Normalisierung 2 10.05.11 Bestimmen diff. expr. Gene,
Logistic regression: Miscellaneous topics
 Logistic regression: Miscellaneous topics April 11 Introduction We have covered two approaches to inference for GLMs: the Wald approach and the likelihood ratio approach I claimed that the likelihood ratio
Logistic regression: Miscellaneous topics April 11 Introduction We have covered two approaches to inference for GLMs: the Wald approach and the likelihood ratio approach I claimed that the likelihood ratio
VARIANCE COMPONENT ANALYSIS
 VARIANCE COMPONENT ANALYSIS T. KRISHNAN Cranes Software International Limited Mahatma Gandhi Road, Bangalore - 560 001 krishnan.t@systat.com 1. Introduction In an experiment to compare the yields of two
VARIANCE COMPONENT ANALYSIS T. KRISHNAN Cranes Software International Limited Mahatma Gandhi Road, Bangalore - 560 001 krishnan.t@systat.com 1. Introduction In an experiment to compare the yields of two
Proportional hazards regression
 Proportional hazards regression Patrick Breheny October 8 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/28 Introduction The model Solving for the MLE Inference Today we will begin discussing regression
Proportional hazards regression Patrick Breheny October 8 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/28 Introduction The model Solving for the MLE Inference Today we will begin discussing regression
Generalized linear models
 Generalized linear models Søren Højsgaard Department of Mathematical Sciences Aalborg University, Denmark October 29, 202 Contents Densities for generalized linear models. Mean and variance...............................
Generalized linear models Søren Højsgaard Department of Mathematical Sciences Aalborg University, Denmark October 29, 202 Contents Densities for generalized linear models. Mean and variance...............................
Simple linear regression
 Simple linear regression Biometry 755 Spring 2008 Simple linear regression p. 1/40 Overview of regression analysis Evaluate relationship between one or more independent variables (X 1,...,X k ) and a single
Simple linear regression Biometry 755 Spring 2008 Simple linear regression p. 1/40 Overview of regression analysis Evaluate relationship between one or more independent variables (X 1,...,X k ) and a single
Non-specific filtering and control of false positives
 Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Systematic Variation in Genetic Microarray Data
 Biostatistics (2004), 1, 1, pp 1 47 Printed in Great Britain Systematic Variation in Genetic Microarray Data By KIMBERLY F SELLERS, JEFFREY MIECZNIKOWSKI, andwilliam F EDDY Department of Statistics, Carnegie
Biostatistics (2004), 1, 1, pp 1 47 Printed in Great Britain Systematic Variation in Genetic Microarray Data By KIMBERLY F SELLERS, JEFFREY MIECZNIKOWSKI, andwilliam F EDDY Department of Statistics, Carnegie
Probe-Level Analysis of Affymetrix GeneChip Microarray Data
 Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley Memorial Sloan-Kettering Cancer Center July
Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley Memorial Sloan-Kettering Cancer Center July
Institute of Actuaries of India
 Institute of Actuaries of India Subject CT3 Probability & Mathematical Statistics May 2011 Examinations INDICATIVE SOLUTION Introduction The indicative solution has been written by the Examiners with the
Institute of Actuaries of India Subject CT3 Probability & Mathematical Statistics May 2011 Examinations INDICATIVE SOLUTION Introduction The indicative solution has been written by the Examiners with the
Example 1: Two-Treatment CRD
 Introduction to Mixed Linear Models in Microarray Experiments //0 Copyright 0 Dan Nettleton Statistical Models A statistical model describes a formal mathematical data generation mechanism from which an
Introduction to Mixed Linear Models in Microarray Experiments //0 Copyright 0 Dan Nettleton Statistical Models A statistical model describes a formal mathematical data generation mechanism from which an
STAT 525 Fall Final exam. Tuesday December 14, 2010
 STAT 525 Fall 2010 Final exam Tuesday December 14, 2010 Time: 2 hours Name (please print): Show all your work and calculations. Partial credit will be given for work that is partially correct. Points will
STAT 525 Fall 2010 Final exam Tuesday December 14, 2010 Time: 2 hours Name (please print): Show all your work and calculations. Partial credit will be given for work that is partially correct. Points will
Multiple Linear Regression
 Multiple Linear Regression University of California, San Diego Instructor: Ery Arias-Castro http://math.ucsd.edu/~eariasca/teaching.html 1 / 42 Passenger car mileage Consider the carmpg dataset taken from
Multiple Linear Regression University of California, San Diego Instructor: Ery Arias-Castro http://math.ucsd.edu/~eariasca/teaching.html 1 / 42 Passenger car mileage Consider the carmpg dataset taken from
Regression Diagnostics for Survey Data
 Regression Diagnostics for Survey Data Richard Valliant Joint Program in Survey Methodology, University of Maryland and University of Michigan USA Jianzhu Li (Westat), Dan Liao (JPSM) 1 Introduction Topics
Regression Diagnostics for Survey Data Richard Valliant Joint Program in Survey Methodology, University of Maryland and University of Michigan USA Jianzhu Li (Westat), Dan Liao (JPSM) 1 Introduction Topics
You can compute the maximum likelihood estimate for the correlation
 Stat 50 Solutions Comments on Assignment Spring 005. (a) _ 37.6 X = 6.5 5.8 97.84 Σ = 9.70 4.9 9.70 75.05 7.80 4.9 7.80 4.96 (b) 08.7 0 S = Σ = 03 9 6.58 03 305.6 30.89 6.58 30.89 5.5 (c) You can compute
Stat 50 Solutions Comments on Assignment Spring 005. (a) _ 37.6 X = 6.5 5.8 97.84 Σ = 9.70 4.9 9.70 75.05 7.80 4.9 7.80 4.96 (b) 08.7 0 S = Σ = 03 9 6.58 03 305.6 30.89 6.58 30.89 5.5 (c) You can compute
DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS
 Sankhyā : The Indian Journal of Statistics Special issue in memory of D. Basu 2002, Volume 64, Series A, Pt. 3, pp 706-720 DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS By TERENCE P. SPEED
Sankhyā : The Indian Journal of Statistics Special issue in memory of D. Basu 2002, Volume 64, Series A, Pt. 3, pp 706-720 DIRECT VERSUS INDIRECT DESIGNS FOR edna MICROARRAY EXPERIMENTS By TERENCE P. SPEED
Estimating σ 2. We can do simple prediction of Y and estimation of the mean of Y at any value of X.
 Estimating σ 2 We can do simple prediction of Y and estimation of the mean of Y at any value of X. To perform inferences about our regression line, we must estimate σ 2, the variance of the error term.
Estimating σ 2 We can do simple prediction of Y and estimation of the mean of Y at any value of X. To perform inferences about our regression line, we must estimate σ 2, the variance of the error term.
Simple and Multiple Linear Regression
 Sta. 113 Chapter 12 and 13 of Devore March 12, 2010 Table of contents 1 Simple Linear Regression 2 Model Simple Linear Regression A simple linear regression model is given by Y = β 0 + β 1 x + ɛ where
Sta. 113 Chapter 12 and 13 of Devore March 12, 2010 Table of contents 1 Simple Linear Regression 2 Model Simple Linear Regression A simple linear regression model is given by Y = β 0 + β 1 x + ɛ where
2.2 Classical Regression in the Time Series Context
 48 2 Time Series Regression and Exploratory Data Analysis context, and therefore we include some material on transformations and other techniques useful in exploratory data analysis. 2.2 Classical Regression
48 2 Time Series Regression and Exploratory Data Analysis context, and therefore we include some material on transformations and other techniques useful in exploratory data analysis. 2.2 Classical Regression
9 Correlation and Regression
 9 Correlation and Regression SW, Chapter 12. Suppose we select n = 10 persons from the population of college seniors who plan to take the MCAT exam. Each takes the test, is coached, and then retakes the
9 Correlation and Regression SW, Chapter 12. Suppose we select n = 10 persons from the population of college seniors who plan to take the MCAT exam. Each takes the test, is coached, and then retakes the
Chapter 1 Statistical Inference
 Chapter 1 Statistical Inference causal inference To infer causality, you need a randomized experiment (or a huge observational study and lots of outside information). inference to populations Generalizations
Chapter 1 Statistical Inference causal inference To infer causality, you need a randomized experiment (or a huge observational study and lots of outside information). inference to populations Generalizations
Design of Microarray Experiments. Xiangqin Cui
 Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Math 423/533: The Main Theoretical Topics
 Math 423/533: The Main Theoretical Topics Notation sample size n, data index i number of predictors, p (p = 2 for simple linear regression) y i : response for individual i x i = (x i1,..., x ip ) (1 p)
Math 423/533: The Main Theoretical Topics Notation sample size n, data index i number of predictors, p (p = 2 for simple linear regression) y i : response for individual i x i = (x i1,..., x ip ) (1 p)
Linear Models in Machine Learning
 CS540 Intro to AI Linear Models in Machine Learning Lecturer: Xiaojin Zhu jerryzhu@cs.wisc.edu We briefly go over two linear models frequently used in machine learning: linear regression for, well, regression,
CS540 Intro to AI Linear Models in Machine Learning Lecturer: Xiaojin Zhu jerryzhu@cs.wisc.edu We briefly go over two linear models frequently used in machine learning: linear regression for, well, regression,
Section 4.6 Simple Linear Regression
 Section 4.6 Simple Linear Regression Objectives ˆ Basic philosophy of SLR and the regression assumptions ˆ Point & interval estimation of the model parameters, and how to make predictions ˆ Point and interval
Section 4.6 Simple Linear Regression Objectives ˆ Basic philosophy of SLR and the regression assumptions ˆ Point & interval estimation of the model parameters, and how to make predictions ˆ Point and interval
Tied survival times; estimation of survival probabilities
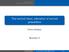 Tied survival times; estimation of survival probabilities Patrick Breheny November 5 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/22 Introduction Tied survival times Introduction Breslow approximation
Tied survival times; estimation of survival probabilities Patrick Breheny November 5 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/22 Introduction Tied survival times Introduction Breslow approximation
Experimental Design. Experimental design. Outline. Choice of platform Array design. Target samples
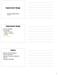 Experimental Design Credit for some of today s materials: Jean Yang, Terry Speed, and Christina Kendziorski Experimental design Choice of platform rray design Creation of probes Location on the array Controls
Experimental Design Credit for some of today s materials: Jean Yang, Terry Speed, and Christina Kendziorski Experimental design Choice of platform rray design Creation of probes Location on the array Controls
5.2 Expounding on the Admissibility of Shrinkage Estimators
 STAT 383C: Statistical Modeling I Fall 2015 Lecture 5 September 15 Lecturer: Purnamrita Sarkar Scribe: Ryan O Donnell Disclaimer: These scribe notes have been slightly proofread and may have typos etc
STAT 383C: Statistical Modeling I Fall 2015 Lecture 5 September 15 Lecturer: Purnamrita Sarkar Scribe: Ryan O Donnell Disclaimer: These scribe notes have been slightly proofread and may have typos etc
Lecture 5 Processing microarray data
 Lecture 5 Processin microarray data (1)Transform the data into a scale suitable for analysis ()Remove the effects of systematic and obfuscatin sources of variation (3)Identify discrepant observations Preprocessin
Lecture 5 Processin microarray data (1)Transform the data into a scale suitable for analysis ()Remove the effects of systematic and obfuscatin sources of variation (3)Identify discrepant observations Preprocessin
Diagnostics and Remedial Measures: An Overview
 Diagnostics and Remedial Measures: An Overview Residuals Model diagnostics Graphical techniques Hypothesis testing Remedial measures Transformation Later: more about all this for multiple regression W.
Diagnostics and Remedial Measures: An Overview Residuals Model diagnostics Graphical techniques Hypothesis testing Remedial measures Transformation Later: more about all this for multiple regression W.
7. Response Surface Methodology (Ch.10. Regression Modeling Ch. 11. Response Surface Methodology)
 7. Response Surface Methodology (Ch.10. Regression Modeling Ch. 11. Response Surface Methodology) Hae-Jin Choi School of Mechanical Engineering, Chung-Ang University 1 Introduction Response surface methodology,
7. Response Surface Methodology (Ch.10. Regression Modeling Ch. 11. Response Surface Methodology) Hae-Jin Choi School of Mechanical Engineering, Chung-Ang University 1 Introduction Response surface methodology,
STAT 4385 Topic 03: Simple Linear Regression
 STAT 4385 Topic 03: Simple Linear Regression Xiaogang Su, Ph.D. Department of Mathematical Science University of Texas at El Paso xsu@utep.edu Spring, 2017 Outline The Set-Up Exploratory Data Analysis
STAT 4385 Topic 03: Simple Linear Regression Xiaogang Su, Ph.D. Department of Mathematical Science University of Texas at El Paso xsu@utep.edu Spring, 2017 Outline The Set-Up Exploratory Data Analysis
LAB 5 INSTRUCTIONS LINEAR REGRESSION AND CORRELATION
 LAB 5 INSTRUCTIONS LINEAR REGRESSION AND CORRELATION In this lab you will learn how to use Excel to display the relationship between two quantitative variables, measure the strength and direction of the
LAB 5 INSTRUCTIONS LINEAR REGRESSION AND CORRELATION In this lab you will learn how to use Excel to display the relationship between two quantitative variables, measure the strength and direction of the
TESTING FOR NORMALITY IN THE LINEAR REGRESSION MODEL: AN EMPIRICAL LIKELIHOOD RATIO TEST
 Econometrics Working Paper EWP0402 ISSN 1485-6441 Department of Economics TESTING FOR NORMALITY IN THE LINEAR REGRESSION MODEL: AN EMPIRICAL LIKELIHOOD RATIO TEST Lauren Bin Dong & David E. A. Giles Department
Econometrics Working Paper EWP0402 ISSN 1485-6441 Department of Economics TESTING FOR NORMALITY IN THE LINEAR REGRESSION MODEL: AN EMPIRICAL LIKELIHOOD RATIO TEST Lauren Bin Dong & David E. A. Giles Department
Semiparametric Regression
 Semiparametric Regression Patrick Breheny October 22 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/23 Introduction Over the past few weeks, we ve introduced a variety of regression models under
Semiparametric Regression Patrick Breheny October 22 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/23 Introduction Over the past few weeks, we ve introduced a variety of regression models under
Ma 3/103: Lecture 24 Linear Regression I: Estimation
 Ma 3/103: Lecture 24 Linear Regression I: Estimation March 3, 2017 KC Border Linear Regression I March 3, 2017 1 / 32 Regression analysis Regression analysis Estimate and test E(Y X) = f (X). f is the
Ma 3/103: Lecture 24 Linear Regression I: Estimation March 3, 2017 KC Border Linear Regression I March 3, 2017 1 / 32 Regression analysis Regression analysis Estimate and test E(Y X) = f (X). f is the
Statistical Methods for Handling Incomplete Data Chapter 2: Likelihood-based approach
 Statistical Methods for Handling Incomplete Data Chapter 2: Likelihood-based approach Jae-Kwang Kim Department of Statistics, Iowa State University Outline 1 Introduction 2 Observed likelihood 3 Mean Score
Statistical Methods for Handling Incomplete Data Chapter 2: Likelihood-based approach Jae-Kwang Kim Department of Statistics, Iowa State University Outline 1 Introduction 2 Observed likelihood 3 Mean Score
Now consider the case where E(Y) = µ = Xβ and V (Y) = σ 2 G, where G is diagonal, but unknown.
 Weighting We have seen that if E(Y) = Xβ and V (Y) = σ 2 G, where G is known, the model can be rewritten as a linear model. This is known as generalized least squares or, if G is diagonal, with trace(g)
Weighting We have seen that if E(Y) = Xβ and V (Y) = σ 2 G, where G is known, the model can be rewritten as a linear model. This is known as generalized least squares or, if G is diagonal, with trace(g)
The Model Building Process Part I: Checking Model Assumptions Best Practice
 The Model Building Process Part I: Checking Model Assumptions Best Practice Authored by: Sarah Burke, PhD 31 July 2017 The goal of the STAT T&E COE is to assist in developing rigorous, defensible test
The Model Building Process Part I: Checking Model Assumptions Best Practice Authored by: Sarah Burke, PhD 31 July 2017 The goal of the STAT T&E COE is to assist in developing rigorous, defensible test
Let us first identify some classes of hypotheses. simple versus simple. H 0 : θ = θ 0 versus H 1 : θ = θ 1. (1) one-sided
 Let us first identify some classes of hypotheses. simple versus simple H 0 : θ = θ 0 versus H 1 : θ = θ 1. (1) one-sided H 0 : θ θ 0 versus H 1 : θ > θ 0. (2) two-sided; null on extremes H 0 : θ θ 1 or
Let us first identify some classes of hypotheses. simple versus simple H 0 : θ = θ 0 versus H 1 : θ = θ 1. (1) one-sided H 0 : θ θ 0 versus H 1 : θ > θ 0. (2) two-sided; null on extremes H 0 : θ θ 1 or
cdna Microarray Analysis
 cdna Microarray Analysis with BioConductor packages Nolwenn Le Meur Copyright 2007 Outline Data acquisition Pre-processing Quality assessment Pre-processing background correction normalization summarization
cdna Microarray Analysis with BioConductor packages Nolwenn Le Meur Copyright 2007 Outline Data acquisition Pre-processing Quality assessment Pre-processing background correction normalization summarization
Y it = µ + τ i + ε it ; ε it ~ N(0, σ 2 ) (1)
 DOE (Linear Model) Strategy for checking experimental model assumptions Part 1 When we discuss experiments whose data are described or analyzed by the one-way analysis of variance ANOVA model, we note
DOE (Linear Model) Strategy for checking experimental model assumptions Part 1 When we discuss experiments whose data are described or analyzed by the one-way analysis of variance ANOVA model, we note
Spatial inference. Spatial inference. Accounting for spatial correlation. Multivariate normal distributions
 Spatial inference I will start with a simple model, using species diversity data Strong spatial dependence, Î = 0.79 what is the mean diversity? How precise is our estimate? Sampling discussion: The 64
Spatial inference I will start with a simple model, using species diversity data Strong spatial dependence, Î = 0.79 what is the mean diversity? How precise is our estimate? Sampling discussion: The 64
a = 4 levels of treatment A = Poison b = 3 levels of treatment B = Pretreatment n = 4 replicates for each treatment combination
 In Box, Hunter, and Hunter Statistics for Experimenters is a two factor example of dying times for animals, let's say cockroaches, using 4 poisons and pretreatments with n=4 values for each combination
In Box, Hunter, and Hunter Statistics for Experimenters is a two factor example of dying times for animals, let's say cockroaches, using 4 poisons and pretreatments with n=4 values for each combination
Optimal normalization of DNA-microarray data
 Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La
Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La
Ch 2: Simple Linear Regression
 Ch 2: Simple Linear Regression 1. Simple Linear Regression Model A simple regression model with a single regressor x is y = β 0 + β 1 x + ɛ, where we assume that the error ɛ is independent random component
Ch 2: Simple Linear Regression 1. Simple Linear Regression Model A simple regression model with a single regressor x is y = β 0 + β 1 x + ɛ, where we assume that the error ɛ is independent random component
Some General Types of Tests
 Some General Types of Tests We may not be able to find a UMP or UMPU test in a given situation. In that case, we may use test of some general class of tests that often have good asymptotic properties.
Some General Types of Tests We may not be able to find a UMP or UMPU test in a given situation. In that case, we may use test of some general class of tests that often have good asymptotic properties.
Review of Statistics 101
 Review of Statistics 101 We review some important themes from the course 1. Introduction Statistics- Set of methods for collecting/analyzing data (the art and science of learning from data). Provides methods
Review of Statistics 101 We review some important themes from the course 1. Introduction Statistics- Set of methods for collecting/analyzing data (the art and science of learning from data). Provides methods
Robust model selection criteria for robust S and LT S estimators
 Hacettepe Journal of Mathematics and Statistics Volume 45 (1) (2016), 153 164 Robust model selection criteria for robust S and LT S estimators Meral Çetin Abstract Outliers and multi-collinearity often
Hacettepe Journal of Mathematics and Statistics Volume 45 (1) (2016), 153 164 Robust model selection criteria for robust S and LT S estimators Meral Çetin Abstract Outliers and multi-collinearity often
The Model Building Process Part I: Checking Model Assumptions Best Practice (Version 1.1)
 The Model Building Process Part I: Checking Model Assumptions Best Practice (Version 1.1) Authored by: Sarah Burke, PhD Version 1: 31 July 2017 Version 1.1: 24 October 2017 The goal of the STAT T&E COE
The Model Building Process Part I: Checking Model Assumptions Best Practice (Version 1.1) Authored by: Sarah Burke, PhD Version 1: 31 July 2017 Version 1.1: 24 October 2017 The goal of the STAT T&E COE
ACOVA and Interactions
 Chapter 15 ACOVA and Interactions Analysis of covariance (ACOVA) incorporates one or more regression variables into an analysis of variance. As such, we can think of it as analogous to the two-way ANOVA
Chapter 15 ACOVA and Interactions Analysis of covariance (ACOVA) incorporates one or more regression variables into an analysis of variance. As such, we can think of it as analogous to the two-way ANOVA
1 Outline. 1. Motivation. 2. SUR model. 3. Simultaneous equations. 4. Estimation
 1 Outline. 1. Motivation 2. SUR model 3. Simultaneous equations 4. Estimation 2 Motivation. In this chapter, we will study simultaneous systems of econometric equations. Systems of simultaneous equations
1 Outline. 1. Motivation 2. SUR model 3. Simultaneous equations 4. Estimation 2 Motivation. In this chapter, we will study simultaneous systems of econometric equations. Systems of simultaneous equations
Regression Review. Statistics 149. Spring Copyright c 2006 by Mark E. Irwin
 Regression Review Statistics 149 Spring 2006 Copyright c 2006 by Mark E. Irwin Matrix Approach to Regression Linear Model: Y i = β 0 + β 1 X i1 +... + β p X ip + ɛ i ; ɛ i iid N(0, σ 2 ), i = 1,..., n
Regression Review Statistics 149 Spring 2006 Copyright c 2006 by Mark E. Irwin Matrix Approach to Regression Linear Model: Y i = β 0 + β 1 X i1 +... + β p X ip + ɛ i ; ɛ i iid N(0, σ 2 ), i = 1,..., n
Introduction to Linear regression analysis. Part 2. Model comparisons
 Introduction to Linear regression analysis Part Model comparisons 1 ANOVA for regression Total variation in Y SS Total = Variation explained by regression with X SS Regression + Residual variation SS Residual
Introduction to Linear regression analysis Part Model comparisons 1 ANOVA for regression Total variation in Y SS Total = Variation explained by regression with X SS Regression + Residual variation SS Residual
Improving the identification of differentially expressed genes in cdna microarray experiments
 Improving the identification of differentially expressed genes in cdna microarray experiments Databionics Research Group University of Marburg 33 Marburg, Germany Alfred Ultsch Abstract. The identification
Improving the identification of differentially expressed genes in cdna microarray experiments Databionics Research Group University of Marburg 33 Marburg, Germany Alfred Ultsch Abstract. The identification
BIOS 2083 Linear Models c Abdus S. Wahed
 Chapter 5 206 Chapter 6 General Linear Model: Statistical Inference 6.1 Introduction So far we have discussed formulation of linear models (Chapter 1), estimability of parameters in a linear model (Chapter
Chapter 5 206 Chapter 6 General Linear Model: Statistical Inference 6.1 Introduction So far we have discussed formulation of linear models (Chapter 1), estimability of parameters in a linear model (Chapter
Confidence Intervals, Testing and ANOVA Summary
 Confidence Intervals, Testing and ANOVA Summary 1 One Sample Tests 1.1 One Sample z test: Mean (σ known) Let X 1,, X n a r.s. from N(µ, σ) or n > 30. Let The test statistic is H 0 : µ = µ 0. z = x µ 0
Confidence Intervals, Testing and ANOVA Summary 1 One Sample Tests 1.1 One Sample z test: Mean (σ known) Let X 1,, X n a r.s. from N(µ, σ) or n > 30. Let The test statistic is H 0 : µ = µ 0. z = x µ 0
Advanced Regression Topics: Violation of Assumptions
 Advanced Regression Topics: Violation of Assumptions Lecture 7 February 15, 2005 Applied Regression Analysis Lecture #7-2/15/2005 Slide 1 of 36 Today s Lecture Today s Lecture rapping Up Revisiting residuals.
Advanced Regression Topics: Violation of Assumptions Lecture 7 February 15, 2005 Applied Regression Analysis Lecture #7-2/15/2005 Slide 1 of 36 Today s Lecture Today s Lecture rapping Up Revisiting residuals.
