Sample Size and Power Calculation in Microarray Studies Using the sizepower package.
|
|
|
- Nathan Wilkins
- 6 years ago
- Views:
Transcription
1 Sample Size and Power Calculation in Microarray Studies Using the sizepower package. Weiliang Qiu Mei-Ling Ting Lee George Alex Whitmore October 30, Introduction The process of obtaining biological samples is often expensive, involved, and time consuming. Thus, the issue of the appropriate sample size is important in planning a study. If the sample size is too large, we waste resources. If the sample size is too small, we cannot draw inferences with the desired precision. Thus, we need to calculate the sample size of a study before proceeding in order to determine the best trade-off between precision and resource use. The calculation of the sample size is closely related to that of power. One goal of microarray studies is to find a subset of genes that are differentially expressed across experimental conditions. The key question is the strength of the claim that a given gene is differentially expressed. In other words, what is the power of the test to determine if a gene is differentially expressed to a specified degree? The R package, sizepower, is used to calculate sample size and power in the planning stage of a microarray study. It helps the user to determine how many samples are needed to achieve a specified power for a test of whether a gene is differentially expressed or, in reverse, to determine the power of a given sample size. This R package provides two functions for sample-size calculations for two types of experimental designs (a completely randomized treatment-control design and a matchedpairs design) and three functions for power calculation for four types of experimental designs (a completely randomized treatment-control design, a matched-pairs design, a multiple-treatment design having an isolated treatment effect, and a randomized block design). 1
2 2 Experiment Designs The following discussion of sample size planning for microarray studies assumes that the investigator has already chosen a particular microarray technology and selected the measure of gene expression. The statistical testing proceeds after the application of appropriate background correction, normalization and mathematical transformations. Thus, expression levels in the following discussion of testing refer to data that have been through these preprocessing steps. 2.1 Completely randomized treatment-control designs In this design, we consider two groups of biological samples: a treatment group and a control group. We are interested in testing if the mean expression level for a given gene is the same for the two groups. The hypotheses of interest for any specific gene are H 0 : θ t = θ c vs H 1 : θ t θ c, (1) where θ t and θ c are the mean expression levels of the treatment and control groups, respectively, for the given gene. When we proceed to consider the required sample size or power level, we will let µ 1 = θ t θ c denote the specific difference in means postulated in the alternative hypothesis H 1. All statistical observations in the design are assumed to be mutually independent. In addition, after appropriate mathematical transformation (such as a log-transformation), all observations in the treatment group are assumed to be drawn from the normal distribution N(θ t, σ 2 ), while samples in the control group are assumed to be drawn from the normal distribution N(θ c, σ 2 ), where the two distributions share a common variance σ 2. To simplify the presentation, we only consider the case where each group has the same number of observations n. For example, in a microarray toxicity study, we randomly assign 2n mice in equal numbers to treatment and control groups. The n mice in the treatment group will be exposed to a toxin and the n mice in the control group will not be exposed to the toxin. We are interested in detecting if any mouse gene on the microarray will be differentially expressed between the treatment and control groups. 2.2 Matched-pairs designs Like the completely randomized treatment-control design, the matched-pairs design consists of two groups of biological samples and we are interested in whether the mean expression levels for genes from the two groups are different. In the matched-pairs design, however, the observations from the two groups are not independent. Instead, each treatment sample is paired with one control sample, creating n pairs of matched (correlated) observations. Within a matched pair, the two observations are dependent. The matched pairs themselves are independent. We still assume that observations in the treatment 2
3 and control groups are normally distributed with means θ t and θ c, respectively, and with common variance σ 2. For example, in a microarray liposarcoma study, we have n patients. From each patient, we obtain one sample of liposarcoma tissue and one sample of normal fat tissue, giving a total of n pairs of tissue samples. Within each pair, the two tissue samples share some features because they are taken from the same patient. We are interested in detecting genes on the microarray that are differentially expressed in the two types of tissue (liposarcoma and normal fat tissues). In other words, for any single gene, we would like to test if the mean difference in expression levels of the two types of tissue is equal to zero. 2.3 Multiple-treatment designs having an isolated treatment effect This design is a generalization of the completely randomized treatment-control designs to multiple treatments. Suppose there are T treatments, designated t = 1,..., T. We first randomly divide nt biological samples into T groups, each with n samples. Then the t-th group is assigned the t-th treatment. The hypotheses are: H 0 : H 1 : θ 1 = = θ T One treatment mean differs from the other T 1 means, (2) where θ t, t = 1,..., T, are the gene expression means of the T treatment groups. The term isolated effect design refers to the fact that, under the alternative hypothesis, all treatments are assumed to have the same mean level of expression except for one treatment that has a different mean level. When we proceed to consider the required sample size or power level, we will let µ 1 denote the specific difference between the mean of the isolated treatment and the common mean of the remaining T 1 treatments under H 1. We assume that all observations for the nt samples are mutually independent and that observations from the t-th treatment group are normally distributed with mean θ t and variance σ 2. Thus, we assume the treatment groups have a common variance. As already seen, we also assume that all treatment groups have the same number of observations. 2.4 Randomized block designs having an isolated treatment effect This design is a generalization of matched-pairs designs to multiple treatments. The benefits of matching are achieved when biological samples within the same block are more homogeneous than samples in different blocks. For example, mice in the same litter tend to be more similar than mice in different litters. A total of n blocks are constructed with each block having T biological samples. The T treatments are randomly applied 3
4 to the samples within each block. For each gene, we are interested in testing if all treatments have the same mean level of expression except for one treatment that has a different mean level. Within a block, the expression levels of a gene are assumed to be dependent. Between blocks, the expression levels of a gene are assumed to be independent. We assume that the expression level of a gene in the t-th treatment group is normally distributed with mean θ t and variance σ 2, i = 1..., T. 3 Sample-size Calculation In this section, we show the method of sample-size calculation for completely randomized treatment-control designs and matched-pairs designs. Denote the total number of genes in a microarray study by G. Next, let G 0 be the true number of genes that are not differentially expressed and R 0 be the number of genes that are not differentially expressed but are falsely declared by the test procedure to be differentially expressed, i.e., R 0 is the number of false positives. The count G 0 is some fixed but unknown number. Prior to applying the test procedure to the data, R 0 is an unknown and random count. The probability α 0 of a type I error for any single gene g among the G 0 genes that are not differentially expressed is given by α 0 = E(R 0) G 0, (3) where E(R 0 ) is the expected number of false positives from applying the test procedure to all of the G 0 genes. Recall that we let µ 1 = θ t θ c denote the difference in means postulated in the alternative hypothesis H 1. For specified values of E(R 0 ) and G 0, we can calculate the requisite sample size to achieve a specified power for completely randomized treatmentcontrol designs and matched-pairs designs as follows: ( ) 2 za + z b n =, (4) µ 1 /σ d where n is the sample size for each group, a = 1 α 0 /2, b is the power of the test, z a and z b are the lower a and b percentiles of the standard normal distribution, and σ d is the standard deviation of the difference in expression between treatment and control samples, as defined below. For a completely randomized treatment-control design, σd 2 is the variance of the difference in expression between randomly chosen treatment and control samples. For a matched-pairs design, σd 2 is the variance of the difference in expression between the treatment and control samples in a matched pair. In both cases, σ 2 d = 2σ 2, (5) 4
5 where σ 2 is the variance of the random error component in an ANOVA model. For a completely randomized treatment-control design, the ANOVA model for any single gene is y ij = µ + τ j + ɛ ij, i = 1,..., n, j = 1, 2 (6) where y ij is the expression level of the i-th biological sample in the j-th group, µ is the overall mean, τ j is the j-th treatment effect, and ɛ ij is the random error component and is normally distributed with mean 0 and variance σ 2. Note that θ t = µ + τ 1 and θ c = µ + τ 2 where τ 1 + τ 2 = 0. For a matched-pairs design, the ANOVA model for any single gene is y ij = µ + τ j + β i + ɛ ij, i = 1,..., n, j = 1, 2 (7) where y ij is the expression level of the i-th biological sample in the j-th group, µ is the overall mean, τ j is the j-th treatment effect, β i is the i-th block effect, and ɛ ij is the random error component and is normally distributed with mean 0 and variance σ 2. Note that θ t = µ + τ 1 and θ c = µ + τ 2. To calculate the sample size in formula (4), we have to anticipate the values of G 0 and σ d. The investigator must also specify the desired values of E(R 0 ), µ 1 and the power b. Typically, a high percentage of the genes are not differentially expressed in a microarray study. In this case, the total number of genes G might be substituted for G 0 in the calculation α 0 = E(R 0 )/G 0, giving the approximation α 0 E(R 0 )/G. We let α F denote the probability that one or more false positives will be produced in testing the family of G 0 genes that are not differentially expressed. The probability of the type I error for an individual test α 0, the probability of the type I error for the family of tests α F, and the false positive rate E(R 0 ) are interrelated. Under the Sidák approach to type I error control, we have the following approximation (Lee, 2004, page 202): E(R 0 ) = α 0 G 0 ln (1 α F ). (8) Under the Bonferroni approach to type I error control, we have the following approximation (Lee, 2004, page 203): E(R 0 ) = α 0 G 0 α F. (9) A subject matter expert can estimate G 0. Similarly, a pilot study or closely related study can be used to estimate the value of σ 2 and, hence, σd 2. For example, we can calculate an ANOVA table for each gene from pilot study data and estimate σ 2. (In an ANOVA test, σ 2 is estimated by the mean square error [MSE]). We can then average these estimates to obtain a pooled estimate of σ 2. The user may wish to explore the sensitivity of the sample size to a range of specifications for E(R 0 ), µ 1 and the power b. For the power b, for example, the user may try a series of probabilities, such as 0.7, 0.8, 0.9 and 0.95, to get a series of sample sizes. To remind the user that completely randomized treatment-control designs and matchedpairs designs are different, we provide one function for each design to implement the sample-size calculation. 5
6 4 Power Calculation In this section, we present a unified methodology for computing power for all four designs, namely, completely randomized treatment-control designs, matched-pairs designs, multiple-treatment designs having an isolated treatment effect, and randomized block designs. In this methodology, we obtain the power value by solving the following equation system for (c, b): α 0 = P r(χ 2 > c H 0 true) b = P r(χ 2 (10) > c H 1 true). Here χ 2 is the test statistic. The value c is a cutoff that determines whether to reject H 0 or not. We reject H 0 if χ 2 > c and do not reject it otherwise. The value b is the power of the test and α 0 = E(R 0 )/G 0 is the probability of the type I error for an individual test. For all four designs we discussed (i.e., completely randomized treatment-control designs, matched-pairs designs, multiple-treatment designs having an isolated treatment effect, and randomized block designs), the test statistic χ 2 is chi-square distributed with T 1 degrees of freedom under the null hypothesis H 0, where T is the number of treatments. Under H 1, the test statistic χ 2 has a non-central chi-square distribution with non-centrality parameter ( ) 2 n(t 1) µ1 ψ 1 =, (11) T σ where the notation is that defined earlier. Recall, specifically, that µ 1 denotes the difference θ t θ c for the treatment-control designs and the magnitude of the isolated effect for the multiple treatments designs. The parameter σ 2 in each design is the variance of the ANOVA error term. We obtain estimates of σ and G 0 by the same method discussed in the previous section. Also, the specifications for µ 1, E(R 0 ) and power level b are required from the investigator in the same manner as described previously. We provide one function to calculate the power for completely randomized treatmentcontrol design, one function for matched-pairs designs, and one function for multipletreatment designs having an isolated treatment effect and randomized block designs. 5 Examples To call the functions in the R package sizepower, we first need to load it into R: Consider a completely randomized treatment-control design with an equal number of biological samples in the treatment and control groups. It is anticipated that G 0 = 2000 genes will not be differentially expressed. The mean number of false positives is to be controlled at E(R 0 ) = 1 and power is to be controlled at µ 1 = From similar studies done previously, it is anticipated that σ = 0.40 (i.e., σ d = 2(0.40) = 0.566). If 6
7 we wish the power level to be b = 0.9, we can use the following command to calculate the number of samples needed for each group: > samplesize.randomized(er0=1, G0=2000, power=0.9, absmu1=1, sigmad=0.566) $n [1] 8 $d [1] That is, 8 samples are needed for each group. This sample size is the smallest n that will provide the required power. The returned value d is the statistical distance between treatment and control conditions specified under H 1, i.e., d = µ 1 /σ d. If we want to see what the exact power is if n = 8, we can use the command: > power.randomized(er0=1, G0=2000, absmu1=1, sigmad=0.566, n=8) $power [1] $psi1 [1] That is, the exact power is and the non-centrality parameter of the associated non-central chi-square statistic is Consider a matched-pairs design. Suppose a pilot study shows that σ = 0.35 (i.e., σ d = 2(0.35) = 0.495). We wish E(R 0 ) = 1, µ 1 = 1.00, and the power b = We also expect that G 0 = Then we can use the command: > samplesize.matched(er0=1, G0=2000, power=0.9, absmu1=1, sigmad=0.495) $n [1] 6 $d [1] If we want to see what the exact power is if n = 6, we can use the command: > power.matched(er0=1, G0=2000, absmu1=1, sigmad=0.495, n=6) $power [1] $psi1 [1]
8 That is, the exact power is and the non-centrality parameter is Consider a multiple-treatment design involving T = 5 treatments and G 0 = 2000 undifferentially expressed genes. Suppose we wish to control E(R 0 ) = 1.0 and to detect an isolated effect of µ 1 = 1.00 between one distinguished treatment and all other treatments. We anticipate an experimental error standard deviation of σ = To calculate the power for n = 6, we can use the following command: > power.multi(er0=1, G0=2000, numtrt=5, absmu1=1, sigma=0.4, n=6) $power [1] $psi1 [1] 30 That is, the power is and the non-centrality parameter is 30. Consider a randomized block design with T = 6 treatments, n = 9 blocks, and G 0 = We wish E(R 0 ) = 1 and µ 1 = 0.9. From a pilot study, we anticipate that σ = 0.5. Then the power is > power.multi(er0=1, G0=5000, numtrt=6, absmu1=0.9, sigma=0.5, n=9) $power [1] $psi1 [1] 24.3 References [1] Lee, M.-L. T. Analysis of Microarray Gene Expression Data. Kluwer Academic Publishers, [2] Lee, M.-L. T. and Whitmore, G. A. Power and sample size for DNA microarray studies. Statistics in Medicine, 21: ,
Chapter Seven: Multi-Sample Methods 1/52
 Chapter Seven: Multi-Sample Methods 1/52 7.1 Introduction 2/52 Introduction The independent samples t test and the independent samples Z test for a difference between proportions are designed to analyze
Chapter Seven: Multi-Sample Methods 1/52 7.1 Introduction 2/52 Introduction The independent samples t test and the independent samples Z test for a difference between proportions are designed to analyze
Lec 1: An Introduction to ANOVA
 Ying Li Stockholm University October 31, 2011 Three end-aisle displays Which is the best? Design of the Experiment Identify the stores of the similar size and type. The displays are randomly assigned to
Ying Li Stockholm University October 31, 2011 Three end-aisle displays Which is the best? Design of the Experiment Identify the stores of the similar size and type. The displays are randomly assigned to
The Random Effects Model Introduction
 The Random Effects Model Introduction Sometimes, treatments included in experiment are randomly chosen from set of all possible treatments. Conclusions from such experiment can then be generalized to other
The Random Effects Model Introduction Sometimes, treatments included in experiment are randomly chosen from set of all possible treatments. Conclusions from such experiment can then be generalized to other
Week 14 Comparing k(> 2) Populations
 Week 14 Comparing k(> 2) Populations Week 14 Objectives Methods associated with testing for the equality of k(> 2) means or proportions are presented. Post-testing concepts and analysis are introduced.
Week 14 Comparing k(> 2) Populations Week 14 Objectives Methods associated with testing for the equality of k(> 2) means or proportions are presented. Post-testing concepts and analysis are introduced.
Analysis of Variance
 Statistical Techniques II EXST7015 Analysis of Variance 15a_ANOVA_Introduction 1 Design The simplest model for Analysis of Variance (ANOVA) is the CRD, the Completely Randomized Design This model is also
Statistical Techniques II EXST7015 Analysis of Variance 15a_ANOVA_Introduction 1 Design The simplest model for Analysis of Variance (ANOVA) is the CRD, the Completely Randomized Design This model is also
16.3 One-Way ANOVA: The Procedure
 16.3 One-Way ANOVA: The Procedure Tom Lewis Fall Term 2009 Tom Lewis () 16.3 One-Way ANOVA: The Procedure Fall Term 2009 1 / 10 Outline 1 The background 2 Computing formulas 3 The ANOVA Identity 4 Tom
16.3 One-Way ANOVA: The Procedure Tom Lewis Fall Term 2009 Tom Lewis () 16.3 One-Way ANOVA: The Procedure Fall Term 2009 1 / 10 Outline 1 The background 2 Computing formulas 3 The ANOVA Identity 4 Tom
The legacy of Sir Ronald A. Fisher. Fisher s three fundamental principles: local control, replication, and randomization.
 1 Chapter 1: Research Design Principles The legacy of Sir Ronald A. Fisher. Fisher s three fundamental principles: local control, replication, and randomization. 2 Chapter 2: Completely Randomized Design
1 Chapter 1: Research Design Principles The legacy of Sir Ronald A. Fisher. Fisher s three fundamental principles: local control, replication, and randomization. 2 Chapter 2: Completely Randomized Design
1 Statistical inference for a population mean
 1 Statistical inference for a population mean 1. Inference for a large sample, known variance Suppose X 1,..., X n represents a large random sample of data from a population with unknown mean µ and known
1 Statistical inference for a population mean 1. Inference for a large sample, known variance Suppose X 1,..., X n represents a large random sample of data from a population with unknown mean µ and known
Summary of Chapters 7-9
 Summary of Chapters 7-9 Chapter 7. Interval Estimation 7.2. Confidence Intervals for Difference of Two Means Let X 1,, X n and Y 1, Y 2,, Y m be two independent random samples of sizes n and m from two
Summary of Chapters 7-9 Chapter 7. Interval Estimation 7.2. Confidence Intervals for Difference of Two Means Let X 1,, X n and Y 1, Y 2,, Y m be two independent random samples of sizes n and m from two
IEOR165 Discussion Week 12
 IEOR165 Discussion Week 12 Sheng Liu University of California, Berkeley Apr 15, 2016 Outline 1 Type I errors & Type II errors 2 Multiple Testing 3 ANOVA IEOR165 Discussion Sheng Liu 2 Type I errors & Type
IEOR165 Discussion Week 12 Sheng Liu University of California, Berkeley Apr 15, 2016 Outline 1 Type I errors & Type II errors 2 Multiple Testing 3 ANOVA IEOR165 Discussion Sheng Liu 2 Type I errors & Type
Sample Size Estimation for Studies of High-Dimensional Data
 Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
One-Way Analysis of Variance. With regression, we related two quantitative, typically continuous variables.
 One-Way Analysis of Variance With regression, we related two quantitative, typically continuous variables. Often we wish to relate a quantitative response variable with a qualitative (or simply discrete)
One-Way Analysis of Variance With regression, we related two quantitative, typically continuous variables. Often we wish to relate a quantitative response variable with a qualitative (or simply discrete)
Summary of Chapter 7 (Sections ) and Chapter 8 (Section 8.1)
 Summary of Chapter 7 (Sections 7.2-7.5) and Chapter 8 (Section 8.1) Chapter 7. Tests of Statistical Hypotheses 7.2. Tests about One Mean (1) Test about One Mean Case 1: σ is known. Assume that X N(µ, σ
Summary of Chapter 7 (Sections 7.2-7.5) and Chapter 8 (Section 8.1) Chapter 7. Tests of Statistical Hypotheses 7.2. Tests about One Mean (1) Test about One Mean Case 1: σ is known. Assume that X N(µ, σ
DESAIN EKSPERIMEN Analysis of Variances (ANOVA) Semester Genap 2017/2018 Jurusan Teknik Industri Universitas Brawijaya
 DESAIN EKSPERIMEN Analysis of Variances (ANOVA) Semester Jurusan Teknik Industri Universitas Brawijaya Outline Introduction The Analysis of Variance Models for the Data Post-ANOVA Comparison of Means Sample
DESAIN EKSPERIMEN Analysis of Variances (ANOVA) Semester Jurusan Teknik Industri Universitas Brawijaya Outline Introduction The Analysis of Variance Models for the Data Post-ANOVA Comparison of Means Sample
CH.9 Tests of Hypotheses for a Single Sample
 CH.9 Tests of Hypotheses for a Single Sample Hypotheses testing Tests on the mean of a normal distributionvariance known Tests on the mean of a normal distributionvariance unknown Tests on the variance
CH.9 Tests of Hypotheses for a Single Sample Hypotheses testing Tests on the mean of a normal distributionvariance known Tests on the mean of a normal distributionvariance unknown Tests on the variance
Sociology 6Z03 Review II
 Sociology 6Z03 Review II John Fox McMaster University Fall 2016 John Fox (McMaster University) Sociology 6Z03 Review II Fall 2016 1 / 35 Outline: Review II Probability Part I Sampling Distributions Probability
Sociology 6Z03 Review II John Fox McMaster University Fall 2016 John Fox (McMaster University) Sociology 6Z03 Review II Fall 2016 1 / 35 Outline: Review II Probability Part I Sampling Distributions Probability
Chi-Squared Tests. Semester 1. Chi-Squared Tests
 Semester 1 Goodness of Fit Up to now, we have tested hypotheses concerning the values of population parameters such as the population mean or proportion. We have not considered testing hypotheses about
Semester 1 Goodness of Fit Up to now, we have tested hypotheses concerning the values of population parameters such as the population mean or proportion. We have not considered testing hypotheses about
Power & Sample Size Calculation
 Chapter 7 Power & Sample Size Calculation Yibi Huang Chapter 7 Section 10.3 Power & Sample Size Calculation for CRDs Power & Sample Size for Factorial Designs Chapter 7-1 Power & Sample Size Calculation
Chapter 7 Power & Sample Size Calculation Yibi Huang Chapter 7 Section 10.3 Power & Sample Size Calculation for CRDs Power & Sample Size for Factorial Designs Chapter 7-1 Power & Sample Size Calculation
Analysis of Variance and Design of Experiments-I
 Analysis of Variance and Design of Experiments-I MODULE VIII LECTURE - 35 ANALYSIS OF VARIANCE IN RANDOM-EFFECTS MODEL AND MIXED-EFFECTS MODEL Dr. Shalabh Department of Mathematics and Statistics Indian
Analysis of Variance and Design of Experiments-I MODULE VIII LECTURE - 35 ANALYSIS OF VARIANCE IN RANDOM-EFFECTS MODEL AND MIXED-EFFECTS MODEL Dr. Shalabh Department of Mathematics and Statistics Indian
Chapter 9 Inferences from Two Samples
 Chapter 9 Inferences from Two Samples 9-1 Review and Preview 9-2 Two Proportions 9-3 Two Means: Independent Samples 9-4 Two Dependent Samples (Matched Pairs) 9-5 Two Variances or Standard Deviations Review
Chapter 9 Inferences from Two Samples 9-1 Review and Preview 9-2 Two Proportions 9-3 Two Means: Independent Samples 9-4 Two Dependent Samples (Matched Pairs) 9-5 Two Variances or Standard Deviations Review
Psychology 282 Lecture #4 Outline Inferences in SLR
 Psychology 282 Lecture #4 Outline Inferences in SLR Assumptions To this point we have not had to make any distributional assumptions. Principle of least squares requires no assumptions. Can use correlations
Psychology 282 Lecture #4 Outline Inferences in SLR Assumptions To this point we have not had to make any distributional assumptions. Principle of least squares requires no assumptions. Can use correlations
Questions 3.83, 6.11, 6.12, 6.17, 6.25, 6.29, 6.33, 6.35, 6.50, 6.51, 6.53, 6.55, 6.59, 6.60, 6.65, 6.69, 6.70, 6.77, 6.79, 6.89, 6.
 Chapter 7 Reading 7.1, 7.2 Questions 3.83, 6.11, 6.12, 6.17, 6.25, 6.29, 6.33, 6.35, 6.50, 6.51, 6.53, 6.55, 6.59, 6.60, 6.65, 6.69, 6.70, 6.77, 6.79, 6.89, 6.112 Introduction In Chapter 5 and 6, we emphasized
Chapter 7 Reading 7.1, 7.2 Questions 3.83, 6.11, 6.12, 6.17, 6.25, 6.29, 6.33, 6.35, 6.50, 6.51, 6.53, 6.55, 6.59, 6.60, 6.65, 6.69, 6.70, 6.77, 6.79, 6.89, 6.112 Introduction In Chapter 5 and 6, we emphasized
:the actual population proportion are equal to the hypothesized sample proportions 2. H a
 AP Statistics Chapter 14 Chi- Square Distribution Procedures I. Chi- Square Distribution ( χ 2 ) The chi- square test is used when comparing categorical data or multiple proportions. a. Family of only
AP Statistics Chapter 14 Chi- Square Distribution Procedures I. Chi- Square Distribution ( χ 2 ) The chi- square test is used when comparing categorical data or multiple proportions. a. Family of only
Design of Engineering Experiments Part 2 Basic Statistical Concepts Simple comparative experiments
 Design of Engineering Experiments Part 2 Basic Statistical Concepts Simple comparative experiments The hypothesis testing framework The two-sample t-test Checking assumptions, validity Comparing more that
Design of Engineering Experiments Part 2 Basic Statistical Concepts Simple comparative experiments The hypothesis testing framework The two-sample t-test Checking assumptions, validity Comparing more that
Review: General Approach to Hypothesis Testing. 1. Define the research question and formulate the appropriate null and alternative hypotheses.
 1 Review: Let X 1, X,..., X n denote n independent random variables sampled from some distribution might not be normal!) with mean µ) and standard deviation σ). Then X µ σ n In other words, X is approximately
1 Review: Let X 1, X,..., X n denote n independent random variables sampled from some distribution might not be normal!) with mean µ) and standard deviation σ). Then X µ σ n In other words, X is approximately
Statistical Modeling and Analysis of Scientific Inquiry: The Basics of Hypothesis Testing
 Statistical Modeling and Analysis of Scientific Inquiry: The Basics of Hypothesis Testing So, What is Statistics? Theory and techniques for learning from data How to collect How to analyze How to interpret
Statistical Modeling and Analysis of Scientific Inquiry: The Basics of Hypothesis Testing So, What is Statistics? Theory and techniques for learning from data How to collect How to analyze How to interpret
Lecture 21: October 19
 36-705: Intermediate Statistics Fall 2017 Lecturer: Siva Balakrishnan Lecture 21: October 19 21.1 Likelihood Ratio Test (LRT) To test composite versus composite hypotheses the general method is to use
36-705: Intermediate Statistics Fall 2017 Lecturer: Siva Balakrishnan Lecture 21: October 19 21.1 Likelihood Ratio Test (LRT) To test composite versus composite hypotheses the general method is to use
Chapter 15: Nonparametric Statistics Section 15.1: An Overview of Nonparametric Statistics
 Section 15.1: An Overview of Nonparametric Statistics Understand Difference between Parametric and Nonparametric Statistical Procedures Parametric statistical procedures inferential procedures that rely
Section 15.1: An Overview of Nonparametric Statistics Understand Difference between Parametric and Nonparametric Statistical Procedures Parametric statistical procedures inferential procedures that rely
9 One-Way Analysis of Variance
 9 One-Way Analysis of Variance SW Chapter 11 - all sections except 6. The one-way analysis of variance (ANOVA) is a generalization of the two sample t test to k 2 groups. Assume that the populations of
9 One-Way Analysis of Variance SW Chapter 11 - all sections except 6. The one-way analysis of variance (ANOVA) is a generalization of the two sample t test to k 2 groups. Assume that the populations of
Parametric versus Nonparametric Statistics-when to use them and which is more powerful? Dr Mahmoud Alhussami
 Parametric versus Nonparametric Statistics-when to use them and which is more powerful? Dr Mahmoud Alhussami Parametric Assumptions The observations must be independent. Dependent variable should be continuous
Parametric versus Nonparametric Statistics-when to use them and which is more powerful? Dr Mahmoud Alhussami Parametric Assumptions The observations must be independent. Dependent variable should be continuous
The Components of a Statistical Hypothesis Testing Problem
 Statistical Inference: Recall from chapter 5 that statistical inference is the use of a subset of a population (the sample) to draw conclusions about the entire population. In chapter 5 we studied one
Statistical Inference: Recall from chapter 5 that statistical inference is the use of a subset of a population (the sample) to draw conclusions about the entire population. In chapter 5 we studied one
Parametric Empirical Bayes Methods for Microarrays
 Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Lecture 7: Hypothesis Testing and ANOVA
 Lecture 7: Hypothesis Testing and ANOVA Goals Overview of key elements of hypothesis testing Review of common one and two sample tests Introduction to ANOVA Hypothesis Testing The intent of hypothesis
Lecture 7: Hypothesis Testing and ANOVA Goals Overview of key elements of hypothesis testing Review of common one and two sample tests Introduction to ANOVA Hypothesis Testing The intent of hypothesis
22s:152 Applied Linear Regression. Chapter 8: 1-Way Analysis of Variance (ANOVA) 2-Way Analysis of Variance (ANOVA)
 22s:152 Applied Linear Regression Chapter 8: 1-Way Analysis of Variance (ANOVA) 2-Way Analysis of Variance (ANOVA) We now consider an analysis with only categorical predictors (i.e. all predictors are
22s:152 Applied Linear Regression Chapter 8: 1-Way Analysis of Variance (ANOVA) 2-Way Analysis of Variance (ANOVA) We now consider an analysis with only categorical predictors (i.e. all predictors are
Space Telescope Science Institute statistics mini-course. October Inference I: Estimation, Confidence Intervals, and Tests of Hypotheses
 Space Telescope Science Institute statistics mini-course October 2011 Inference I: Estimation, Confidence Intervals, and Tests of Hypotheses James L Rosenberger Acknowledgements: Donald Richards, William
Space Telescope Science Institute statistics mini-course October 2011 Inference I: Estimation, Confidence Intervals, and Tests of Hypotheses James L Rosenberger Acknowledgements: Donald Richards, William
Solutions to Final STAT 421, Fall 2008
 Solutions to Final STAT 421, Fall 2008 Fritz Scholz 1. (8) Two treatments A and B were randomly assigned to 8 subjects (4 subjects to each treatment) with the following responses: 0, 1, 3, 6 and 5, 7,
Solutions to Final STAT 421, Fall 2008 Fritz Scholz 1. (8) Two treatments A and B were randomly assigned to 8 subjects (4 subjects to each treatment) with the following responses: 0, 1, 3, 6 and 5, 7,
Two-Sample Inferential Statistics
 The t Test for Two Independent Samples 1 Two-Sample Inferential Statistics In an experiment there are two or more conditions One condition is often called the control condition in which the treatment is
The t Test for Two Independent Samples 1 Two-Sample Inferential Statistics In an experiment there are two or more conditions One condition is often called the control condition in which the treatment is
HYPOTHESIS TESTING. Hypothesis Testing
 MBA 605 Business Analytics Don Conant, PhD. HYPOTHESIS TESTING Hypothesis testing involves making inferences about the nature of the population on the basis of observations of a sample drawn from the population.
MBA 605 Business Analytics Don Conant, PhD. HYPOTHESIS TESTING Hypothesis testing involves making inferences about the nature of the population on the basis of observations of a sample drawn from the population.
Table of Outcomes. Table of Outcomes. Table of Outcomes. Table of Outcomes. Table of Outcomes. Table of Outcomes. T=number of type 2 errors
 The Multiple Testing Problem Multiple Testing Methods for the Analysis of Microarray Data 3/9/2009 Copyright 2009 Dan Nettleton Suppose one test of interest has been conducted for each of m genes in a
The Multiple Testing Problem Multiple Testing Methods for the Analysis of Microarray Data 3/9/2009 Copyright 2009 Dan Nettleton Suppose one test of interest has been conducted for each of m genes in a
Statistical methods for comparing multiple groups. Lecture 7: ANOVA. ANOVA: Definition. ANOVA: Concepts
 Statistical methods for comparing multiple groups Lecture 7: ANOVA Sandy Eckel seckel@jhsph.edu 30 April 2008 Continuous data: comparing multiple means Analysis of variance Binary data: comparing multiple
Statistical methods for comparing multiple groups Lecture 7: ANOVA Sandy Eckel seckel@jhsph.edu 30 April 2008 Continuous data: comparing multiple means Analysis of variance Binary data: comparing multiple
I i=1 1 I(J 1) j=1 (Y ij Ȳi ) 2. j=1 (Y j Ȳ )2 ] = 2n( is the two-sample t-test statistic.
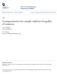
![I i=1 1 I(J 1) j=1 (Y ij Ȳi ) 2. j=1 (Y j Ȳ )2 ] = 2n( is the two-sample t-test statistic. I i=1 1 I(J 1) j=1 (Y ij Ȳi ) 2. j=1 (Y j Ȳ )2 ] = 2n( is the two-sample t-test statistic.](/thumbs/85/92729432.jpg) Serik Sagitov, Chalmers and GU, February, 08 Solutions chapter Matlab commands: x = data matrix boxplot(x) anova(x) anova(x) Problem.3 Consider one-way ANOVA test statistic For I = and = n, put F = MS
Serik Sagitov, Chalmers and GU, February, 08 Solutions chapter Matlab commands: x = data matrix boxplot(x) anova(x) anova(x) Problem.3 Consider one-way ANOVA test statistic For I = and = n, put F = MS
High-Throughput Sequencing Course. Introduction. Introduction. Multiple Testing. Biostatistics and Bioinformatics. Summer 2018
 High-Throughput Sequencing Course Multiple Testing Biostatistics and Bioinformatics Summer 2018 Introduction You have previously considered the significance of a single gene Introduction You have previously
High-Throughput Sequencing Course Multiple Testing Biostatistics and Bioinformatics Summer 2018 Introduction You have previously considered the significance of a single gene Introduction You have previously
STAT 111 Recitation 9
 STAT 111 Recitation 9 Linjun Zhang November 10, 2017 Hypothesis Testing Basic concepts: H 0 (null hypothesis), H 1 (alternative hypothesis) 1 Hypothesis Testing Basic concepts: H 0 (null hypothesis), H
STAT 111 Recitation 9 Linjun Zhang November 10, 2017 Hypothesis Testing Basic concepts: H 0 (null hypothesis), H 1 (alternative hypothesis) 1 Hypothesis Testing Basic concepts: H 0 (null hypothesis), H
One-Way Analysis of Variance (ANOVA)
 1 One-Way Analysis of Variance (ANOVA) One-Way Analysis of Variance (ANOVA) is a method for comparing the means of a populations. This kind of problem arises in two different settings 1. When a independent
1 One-Way Analysis of Variance (ANOVA) One-Way Analysis of Variance (ANOVA) is a method for comparing the means of a populations. This kind of problem arises in two different settings 1. When a independent
Sleep data, two drugs Ch13.xls
 Model Based Statistics in Biology. Part IV. The General Linear Mixed Model.. Chapter 13.3 Fixed*Random Effects (Paired t-test) ReCap. Part I (Chapters 1,2,3,4), Part II (Ch 5, 6, 7) ReCap Part III (Ch
Model Based Statistics in Biology. Part IV. The General Linear Mixed Model.. Chapter 13.3 Fixed*Random Effects (Paired t-test) ReCap. Part I (Chapters 1,2,3,4), Part II (Ch 5, 6, 7) ReCap Part III (Ch
Statistical Inference: Estimation and Confidence Intervals Hypothesis Testing
 Statistical Inference: Estimation and Confidence Intervals Hypothesis Testing 1 In most statistics problems, we assume that the data have been generated from some unknown probability distribution. We desire
Statistical Inference: Estimation and Confidence Intervals Hypothesis Testing 1 In most statistics problems, we assume that the data have been generated from some unknown probability distribution. We desire
ME3620. Theory of Engineering Experimentation. Spring Chapter IV. Decision Making for a Single Sample. Chapter IV
 Theory of Engineering Experimentation Chapter IV. Decision Making for a Single Sample Chapter IV 1 4 1 Statistical Inference The field of statistical inference consists of those methods used to make decisions
Theory of Engineering Experimentation Chapter IV. Decision Making for a Single Sample Chapter IV 1 4 1 Statistical Inference The field of statistical inference consists of those methods used to make decisions
10.2: The Chi Square Test for Goodness of Fit
 10.2: The Chi Square Test for Goodness of Fit We can perform a hypothesis test to determine whether the distribution of a single categorical variable is following a proposed distribution. We call this
10.2: The Chi Square Test for Goodness of Fit We can perform a hypothesis test to determine whether the distribution of a single categorical variable is following a proposed distribution. We call this
STAT 135 Lab 5 Bootstrapping and Hypothesis Testing
 STAT 135 Lab 5 Bootstrapping and Hypothesis Testing Rebecca Barter March 2, 2015 The Bootstrap Bootstrap Suppose that we are interested in estimating a parameter θ from some population with members x 1,...,
STAT 135 Lab 5 Bootstrapping and Hypothesis Testing Rebecca Barter March 2, 2015 The Bootstrap Bootstrap Suppose that we are interested in estimating a parameter θ from some population with members x 1,...,
Lecture 2: Basic Concepts and Simple Comparative Experiments Montgomery: Chapter 2
 Lecture 2: Basic Concepts and Simple Comparative Experiments Montgomery: Chapter 2 Fall, 2013 Page 1 Random Variable and Probability Distribution Discrete random variable Y : Finite possible values {y
Lecture 2: Basic Concepts and Simple Comparative Experiments Montgomery: Chapter 2 Fall, 2013 Page 1 Random Variable and Probability Distribution Discrete random variable Y : Finite possible values {y
Chapter 12. Analysis of variance
 Serik Sagitov, Chalmers and GU, January 9, 016 Chapter 1. Analysis of variance Chapter 11: I = samples independent samples paired samples Chapter 1: I 3 samples of equal size J one-way layout two-way layout
Serik Sagitov, Chalmers and GU, January 9, 016 Chapter 1. Analysis of variance Chapter 11: I = samples independent samples paired samples Chapter 1: I 3 samples of equal size J one-way layout two-way layout
Extending the Robust Means Modeling Framework. Alyssa Counsell, Phil Chalmers, Matt Sigal, Rob Cribbie
 Extending the Robust Means Modeling Framework Alyssa Counsell, Phil Chalmers, Matt Sigal, Rob Cribbie One-way Independent Subjects Design Model: Y ij = µ + τ j + ε ij, j = 1,, J Y ij = score of the ith
Extending the Robust Means Modeling Framework Alyssa Counsell, Phil Chalmers, Matt Sigal, Rob Cribbie One-way Independent Subjects Design Model: Y ij = µ + τ j + ε ij, j = 1,, J Y ij = score of the ith
Section 9.4. Notation. Requirements. Definition. Inferences About Two Means (Matched Pairs) Examples
 Objective Section 9.4 Inferences About Two Means (Matched Pairs) Compare of two matched-paired means using two samples from each population. Hypothesis Tests and Confidence Intervals of two dependent means
Objective Section 9.4 Inferences About Two Means (Matched Pairs) Compare of two matched-paired means using two samples from each population. Hypothesis Tests and Confidence Intervals of two dependent means
CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity
 CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity Prof. Kevin E. Thorpe Dept. of Public Health Sciences University of Toronto Objectives 1. Be able to distinguish among the various
CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity Prof. Kevin E. Thorpe Dept. of Public Health Sciences University of Toronto Objectives 1. Be able to distinguish among the various
Design of Microarray Experiments. Xiangqin Cui
 Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
CIVL /8904 T R A F F I C F L O W T H E O R Y L E C T U R E - 8
 CIVL - 7904/8904 T R A F F I C F L O W T H E O R Y L E C T U R E - 8 Chi-square Test How to determine the interval from a continuous distribution I = Range 1 + 3.322(logN) I-> Range of the class interval
CIVL - 7904/8904 T R A F F I C F L O W T H E O R Y L E C T U R E - 8 Chi-square Test How to determine the interval from a continuous distribution I = Range 1 + 3.322(logN) I-> Range of the class interval
Research Article A Nonparametric Two-Sample Wald Test of Equality of Variances
 Advances in Decision Sciences Volume 211, Article ID 74858, 8 pages doi:1.1155/211/74858 Research Article A Nonparametric Two-Sample Wald Test of Equality of Variances David Allingham 1 andj.c.w.rayner
Advances in Decision Sciences Volume 211, Article ID 74858, 8 pages doi:1.1155/211/74858 Research Article A Nonparametric Two-Sample Wald Test of Equality of Variances David Allingham 1 andj.c.w.rayner
CHAPTER 17 CHI-SQUARE AND OTHER NONPARAMETRIC TESTS FROM: PAGANO, R. R. (2007)
 FROM: PAGANO, R. R. (007) I. INTRODUCTION: DISTINCTION BETWEEN PARAMETRIC AND NON-PARAMETRIC TESTS Statistical inference tests are often classified as to whether they are parametric or nonparametric Parameter
FROM: PAGANO, R. R. (007) I. INTRODUCTION: DISTINCTION BETWEEN PARAMETRIC AND NON-PARAMETRIC TESTS Statistical inference tests are often classified as to whether they are parametric or nonparametric Parameter
ANOVA Analysis of Variance
 ANOVA Analysis of Variance ANOVA Analysis of Variance Extends independent samples t test ANOVA Analysis of Variance Extends independent samples t test Compares the means of groups of independent observations
ANOVA Analysis of Variance ANOVA Analysis of Variance Extends independent samples t test ANOVA Analysis of Variance Extends independent samples t test Compares the means of groups of independent observations
Chapter 10: Analysis of variance (ANOVA)
 Chapter 10: Analysis of variance (ANOVA) ANOVA (Analysis of variance) is a collection of techniques for dealing with more general experiments than the previous one-sample or two-sample tests. We first
Chapter 10: Analysis of variance (ANOVA) ANOVA (Analysis of variance) is a collection of techniques for dealing with more general experiments than the previous one-sample or two-sample tests. We first
PSY 307 Statistics for the Behavioral Sciences. Chapter 20 Tests for Ranked Data, Choosing Statistical Tests
 PSY 307 Statistics for the Behavioral Sciences Chapter 20 Tests for Ranked Data, Choosing Statistical Tests What To Do with Non-normal Distributions Tranformations (pg 382): The shape of the distribution
PSY 307 Statistics for the Behavioral Sciences Chapter 20 Tests for Ranked Data, Choosing Statistical Tests What To Do with Non-normal Distributions Tranformations (pg 382): The shape of the distribution
Chapter 10. Chapter 10. Multinomial Experiments and. Multinomial Experiments and Contingency Tables. Contingency Tables.
 Chapter 10 Multinomial Experiments and Contingency Tables 1 Chapter 10 Multinomial Experiments and Contingency Tables 10-1 1 Overview 10-2 2 Multinomial Experiments: of-fitfit 10-3 3 Contingency Tables:
Chapter 10 Multinomial Experiments and Contingency Tables 1 Chapter 10 Multinomial Experiments and Contingency Tables 10-1 1 Overview 10-2 2 Multinomial Experiments: of-fitfit 10-3 3 Contingency Tables:
STAT 461/561- Assignments, Year 2015
 STAT 461/561- Assignments, Year 2015 This is the second set of assignment problems. When you hand in any problem, include the problem itself and its number. pdf are welcome. If so, use large fonts and
STAT 461/561- Assignments, Year 2015 This is the second set of assignment problems. When you hand in any problem, include the problem itself and its number. pdf are welcome. If so, use large fonts and
Chapter 1. Scientific Process and Themes of Biology
 Chapter 1 Scientific Process and Themes of Biology What is Science? u Scientific knowledge is acquired using a rigorous process u Science is an organized way of gathering and analyzing evidence about the
Chapter 1 Scientific Process and Themes of Biology What is Science? u Scientific knowledge is acquired using a rigorous process u Science is an organized way of gathering and analyzing evidence about the
An Omnibus Consistent Adaptive Percentile Modified. Wilcoxon Rank Sum Test with Applicaitions in Gene. Expression Studies
 An Omnibus Consistent Adaptive Percentile Modified Wilcoxon Rank Sum Test with Applicaitions in Gene Expression Studies O. Thas, L. Clement, J.C.W. Rayner, B. Carvalho, and W. Van Criekinge Supplementary
An Omnibus Consistent Adaptive Percentile Modified Wilcoxon Rank Sum Test with Applicaitions in Gene Expression Studies O. Thas, L. Clement, J.C.W. Rayner, B. Carvalho, and W. Van Criekinge Supplementary
Adaptive Designs: Why, How and When?
 Adaptive Designs: Why, How and When? Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj ISBS Conference Shanghai, July 2008 1 Adaptive designs:
Adaptive Designs: Why, How and When? Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj ISBS Conference Shanghai, July 2008 1 Adaptive designs:
Central Limit Theorem ( 5.3)
 Central Limit Theorem ( 5.3) Let X 1, X 2,... be a sequence of independent random variables, each having n mean µ and variance σ 2. Then the distribution of the partial sum S n = X i i=1 becomes approximately
Central Limit Theorem ( 5.3) Let X 1, X 2,... be a sequence of independent random variables, each having n mean µ and variance σ 2. Then the distribution of the partial sum S n = X i i=1 becomes approximately
3. (a) (8 points) There is more than one way to correctly express the null hypothesis in matrix form. One way to state the null hypothesis is
 Stat 501 Solutions and Comments on Exam 1 Spring 005-4 0-4 1. (a) (5 points) Y ~ N, -1-4 34 (b) (5 points) X (X,X ) = (5,8) ~ N ( 11.5, 0.9375 ) 3 1 (c) (10 points, for each part) (i), (ii), and (v) are
Stat 501 Solutions and Comments on Exam 1 Spring 005-4 0-4 1. (a) (5 points) Y ~ N, -1-4 34 (b) (5 points) X (X,X ) = (5,8) ~ N ( 11.5, 0.9375 ) 3 1 (c) (10 points, for each part) (i), (ii), and (v) are
Answer keys for Assignment 10: Measurement of study variables (The correct answer is underlined in bold text)
 Answer keys for Assignment 10: Measurement of study variables (The correct answer is underlined in bold text) 1. A quick and easy indicator of dispersion is a. Arithmetic mean b. Variance c. Standard deviation
Answer keys for Assignment 10: Measurement of study variables (The correct answer is underlined in bold text) 1. A quick and easy indicator of dispersion is a. Arithmetic mean b. Variance c. Standard deviation
Statistical Inference. Hypothesis Testing
 Statistical Inference Hypothesis Testing Previously, we introduced the point and interval estimation of an unknown parameter(s), say µ and σ 2. However, in practice, the problem confronting the scientist
Statistical Inference Hypothesis Testing Previously, we introduced the point and interval estimation of an unknown parameter(s), say µ and σ 2. However, in practice, the problem confronting the scientist
Contrasts and Multiple Comparisons Supplement for Pages
 Contrasts and Multiple Comparisons Supplement for Pages 302-323 Brian Habing University of South Carolina Last Updated: July 20, 2001 The F-test from the ANOVA table allows us to test the null hypothesis
Contrasts and Multiple Comparisons Supplement for Pages 302-323 Brian Habing University of South Carolina Last Updated: July 20, 2001 The F-test from the ANOVA table allows us to test the null hypothesis
Tests about a population mean
 October 2 nd, 2017 Overview Week 1 Week 2 Week 4 Week 7 Week 10 Week 12 Chapter 1: Descriptive statistics Chapter 6: Statistics and Sampling Distributions Chapter 7: Point Estimation Chapter 8: Confidence
October 2 nd, 2017 Overview Week 1 Week 2 Week 4 Week 7 Week 10 Week 12 Chapter 1: Descriptive statistics Chapter 6: Statistics and Sampling Distributions Chapter 7: Point Estimation Chapter 8: Confidence
SEVERAL μs AND MEDIANS: MORE ISSUES. Business Statistics
 SEVERAL μs AND MEDIANS: MORE ISSUES Business Statistics CONTENTS Post-hoc analysis ANOVA for 2 groups The equal variances assumption The Kruskal-Wallis test Old exam question Further study POST-HOC ANALYSIS
SEVERAL μs AND MEDIANS: MORE ISSUES Business Statistics CONTENTS Post-hoc analysis ANOVA for 2 groups The equal variances assumption The Kruskal-Wallis test Old exam question Further study POST-HOC ANALYSIS
T.I.H.E. IT 233 Statistics and Probability: Sem. 1: 2013 ESTIMATION AND HYPOTHESIS TESTING OF TWO POPULATIONS
 ESTIMATION AND HYPOTHESIS TESTING OF TWO POPULATIONS In our work on hypothesis testing, we used the value of a sample statistic to challenge an accepted value of a population parameter. We focused only
ESTIMATION AND HYPOTHESIS TESTING OF TWO POPULATIONS In our work on hypothesis testing, we used the value of a sample statistic to challenge an accepted value of a population parameter. We focused only
1 One-way Analysis of Variance
 1 One-way Analysis of Variance Suppose that a random sample of q individuals receives treatment T i, i = 1,,... p. Let Y ij be the response from the jth individual to be treated with the ith treatment
1 One-way Analysis of Variance Suppose that a random sample of q individuals receives treatment T i, i = 1,,... p. Let Y ij be the response from the jth individual to be treated with the ith treatment
Single Sample Means. SOCY601 Alan Neustadtl
 Single Sample Means SOCY601 Alan Neustadtl The Central Limit Theorem If we have a population measured by a variable with a mean µ and a standard deviation σ, and if all possible random samples of size
Single Sample Means SOCY601 Alan Neustadtl The Central Limit Theorem If we have a population measured by a variable with a mean µ and a standard deviation σ, and if all possible random samples of size
STAT 135 Lab 9 Multiple Testing, One-Way ANOVA and Kruskal-Wallis
 STAT 135 Lab 9 Multiple Testing, One-Way ANOVA and Kruskal-Wallis Rebecca Barter April 6, 2015 Multiple Testing Multiple Testing Recall that when we were doing two sample t-tests, we were testing the equality
STAT 135 Lab 9 Multiple Testing, One-Way ANOVA and Kruskal-Wallis Rebecca Barter April 6, 2015 Multiple Testing Multiple Testing Recall that when we were doing two sample t-tests, we were testing the equality
Probabilities & Statistics Revision
 Probabilities & Statistics Revision Christopher Ting Christopher Ting http://www.mysmu.edu/faculty/christophert/ : christopherting@smu.edu.sg : 6828 0364 : LKCSB 5036 January 6, 2017 Christopher Ting QF
Probabilities & Statistics Revision Christopher Ting Christopher Ting http://www.mysmu.edu/faculty/christophert/ : christopherting@smu.edu.sg : 6828 0364 : LKCSB 5036 January 6, 2017 Christopher Ting QF
4.1. Introduction: Comparing Means
 4. Analysis of Variance (ANOVA) 4.1. Introduction: Comparing Means Consider the problem of testing H 0 : µ 1 = µ 2 against H 1 : µ 1 µ 2 in two independent samples of two different populations of possibly
4. Analysis of Variance (ANOVA) 4.1. Introduction: Comparing Means Consider the problem of testing H 0 : µ 1 = µ 2 against H 1 : µ 1 µ 2 in two independent samples of two different populations of possibly
Statistical inference (estimation, hypothesis tests, confidence intervals) Oct 2018
 Statistical inference (estimation, hypothesis tests, confidence intervals) Oct 2018 Sampling A trait is measured on each member of a population. f(y) = propn of individuals in the popn with measurement
Statistical inference (estimation, hypothesis tests, confidence intervals) Oct 2018 Sampling A trait is measured on each member of a population. f(y) = propn of individuals in the popn with measurement
LECTURE 5 HYPOTHESIS TESTING
 October 25, 2016 LECTURE 5 HYPOTHESIS TESTING Basic concepts In this lecture we continue to discuss the normal classical linear regression defined by Assumptions A1-A5. Let θ Θ R d be a parameter of interest.
October 25, 2016 LECTURE 5 HYPOTHESIS TESTING Basic concepts In this lecture we continue to discuss the normal classical linear regression defined by Assumptions A1-A5. Let θ Θ R d be a parameter of interest.
Purposes of Data Analysis. Variables and Samples. Parameters and Statistics. Part 1: Probability Distributions
 Part 1: Probability Distributions Purposes of Data Analysis True Distributions or Relationships in the Earths System Probability Distribution Normal Distribution Student-t Distribution Chi Square Distribution
Part 1: Probability Distributions Purposes of Data Analysis True Distributions or Relationships in the Earths System Probability Distribution Normal Distribution Student-t Distribution Chi Square Distribution
Application of Variance Homogeneity Tests Under Violation of Normality Assumption
 Application of Variance Homogeneity Tests Under Violation of Normality Assumption Alisa A. Gorbunova, Boris Yu. Lemeshko Novosibirsk State Technical University Novosibirsk, Russia e-mail: gorbunova.alisa@gmail.com
Application of Variance Homogeneity Tests Under Violation of Normality Assumption Alisa A. Gorbunova, Boris Yu. Lemeshko Novosibirsk State Technical University Novosibirsk, Russia e-mail: gorbunova.alisa@gmail.com
Much of the material we will be covering for a while has to do with designing an experimental study that concerns some phenomenon of interest.
 Experimental Design: Much of the material we will be covering for a while has to do with designing an experimental study that concerns some phenomenon of interest We wish to use our subjects in the best
Experimental Design: Much of the material we will be covering for a while has to do with designing an experimental study that concerns some phenomenon of interest We wish to use our subjects in the best
OHSU OGI Class ECE-580-DOE :Design of Experiments Steve Brainerd
 Why We Use Analysis of Variance to Compare Group Means and How it Works The question of how to compare the population means of more than two groups is an important one to researchers. Let us suppose that
Why We Use Analysis of Variance to Compare Group Means and How it Works The question of how to compare the population means of more than two groups is an important one to researchers. Let us suppose that
Tutorial 1: Power and Sample Size for the One-sample t-test. Acknowledgements:
 Tutorial 1: Power and Sample Size for the One-sample t-test Anna E. Barón, Keith E. Muller, Sarah M. Kreidler, and Deborah H. Glueck Acknowledgements: The project was supported in large part by the National
Tutorial 1: Power and Sample Size for the One-sample t-test Anna E. Barón, Keith E. Muller, Sarah M. Kreidler, and Deborah H. Glueck Acknowledgements: The project was supported in large part by the National
A nonparametric two-sample wald test of equality of variances
 University of Wollongong Research Online Faculty of Informatics - Papers (Archive) Faculty of Engineering and Information Sciences 211 A nonparametric two-sample wald test of equality of variances David
University of Wollongong Research Online Faculty of Informatics - Papers (Archive) Faculty of Engineering and Information Sciences 211 A nonparametric two-sample wald test of equality of variances David
Lecture 28. Ingo Ruczinski. December 3, Department of Biostatistics Johns Hopkins Bloomberg School of Public Health Johns Hopkins University
 Lecture 28 Department of Biostatistics Johns Hopkins Bloomberg School of Public Health Johns Hopkins University December 3, 2015 1 2 3 4 5 1 Familywise error rates 2 procedure 3 Performance of with multiple
Lecture 28 Department of Biostatistics Johns Hopkins Bloomberg School of Public Health Johns Hopkins University December 3, 2015 1 2 3 4 5 1 Familywise error rates 2 procedure 3 Performance of with multiple
Econ 3790: Business and Economic Statistics. Instructor: Yogesh Uppal
 Econ 3790: Business and Economic Statistics Instructor: Yogesh Uppal Email: yuppal@ysu.edu Chapter 13, Part A: Analysis of Variance and Experimental Design Introduction to Analysis of Variance Analysis
Econ 3790: Business and Economic Statistics Instructor: Yogesh Uppal Email: yuppal@ysu.edu Chapter 13, Part A: Analysis of Variance and Experimental Design Introduction to Analysis of Variance Analysis
In a one-way ANOVA, the total sums of squares among observations is partitioned into two components: Sums of squares represent:
 Activity #10: AxS ANOVA (Repeated subjects design) Resources: optimism.sav So far in MATH 300 and 301, we have studied the following hypothesis testing procedures: 1) Binomial test, sign-test, Fisher s
Activity #10: AxS ANOVA (Repeated subjects design) Resources: optimism.sav So far in MATH 300 and 301, we have studied the following hypothesis testing procedures: 1) Binomial test, sign-test, Fisher s
Classroom Activity 7 Math 113 Name : 10 pts Intro to Applied Stats
 Classroom Activity 7 Math 113 Name : 10 pts Intro to Applied Stats Materials Needed: Bags of popcorn, watch with second hand or microwave with digital timer. Instructions: Follow the instructions on the
Classroom Activity 7 Math 113 Name : 10 pts Intro to Applied Stats Materials Needed: Bags of popcorn, watch with second hand or microwave with digital timer. Instructions: Follow the instructions on the
22s:152 Applied Linear Regression. 1-way ANOVA visual:
 22s:152 Applied Linear Regression 1-way ANOVA visual: Chapter 8: 1-Way Analysis of Variance (ANOVA) 2-Way Analysis of Variance (ANOVA) 0.00 0.05 0.10 0.15 0.20 0.25 0.30 0.35 Y We now consider an analysis
22s:152 Applied Linear Regression 1-way ANOVA visual: Chapter 8: 1-Way Analysis of Variance (ANOVA) 2-Way Analysis of Variance (ANOVA) 0.00 0.05 0.10 0.15 0.20 0.25 0.30 0.35 Y We now consider an analysis
Introduction 1. STA442/2101 Fall See last slide for copyright information. 1 / 33
 Introduction 1 STA442/2101 Fall 2016 1 See last slide for copyright information. 1 / 33 Background Reading Optional Chapter 1 of Linear models with R Chapter 1 of Davison s Statistical models: Data, and
Introduction 1 STA442/2101 Fall 2016 1 See last slide for copyright information. 1 / 33 Background Reading Optional Chapter 1 of Linear models with R Chapter 1 of Davison s Statistical models: Data, and
What If There Are More Than. Two Factor Levels?
 What If There Are More Than Chapter 3 Two Factor Levels? Comparing more that two factor levels the analysis of variance ANOVA decomposition of total variability Statistical testing & analysis Checking
What If There Are More Than Chapter 3 Two Factor Levels? Comparing more that two factor levels the analysis of variance ANOVA decomposition of total variability Statistical testing & analysis Checking
ANOVA: Comparing More Than Two Means
 1 ANOVA: Comparing More Than Two Means 10.1 ANOVA: The Completely Randomized Design Elements of a Designed Experiment Before we begin any calculations, we need to discuss some terminology. To make this
1 ANOVA: Comparing More Than Two Means 10.1 ANOVA: The Completely Randomized Design Elements of a Designed Experiment Before we begin any calculations, we need to discuss some terminology. To make this
CHAPTER 8. Test Procedures is a rule, based on sample data, for deciding whether to reject H 0 and contains:
 CHAPTER 8 Test of Hypotheses Based on a Single Sample Hypothesis testing is the method that decide which of two contradictory claims about the parameter is correct. Here the parameters of interest are
CHAPTER 8 Test of Hypotheses Based on a Single Sample Hypothesis testing is the method that decide which of two contradictory claims about the parameter is correct. Here the parameters of interest are
We need to define some concepts that are used in experiments.
 Chapter 0 Analysis of Variance (a.k.a. Designing and Analysing Experiments) Section 0. Introduction In Chapter we mentioned some different ways in which we could get data: Surveys, Observational Studies,
Chapter 0 Analysis of Variance (a.k.a. Designing and Analysing Experiments) Section 0. Introduction In Chapter we mentioned some different ways in which we could get data: Surveys, Observational Studies,
BIO5312 Biostatistics Lecture 6: Statistical hypothesis testings
 BIO5312 Biostatistics Lecture 6: Statistical hypothesis testings Yujin Chung October 4th, 2016 Fall 2016 Yujin Chung Lec6: Statistical hypothesis testings Fall 2016 1/30 Previous Two types of statistical
BIO5312 Biostatistics Lecture 6: Statistical hypothesis testings Yujin Chung October 4th, 2016 Fall 2016 Yujin Chung Lec6: Statistical hypothesis testings Fall 2016 1/30 Previous Two types of statistical
Design & Analysis of Experiments 7E 2009 Montgomery
 1 What If There Are More Than Two Factor Levels? The t-test does not directly apply ppy There are lots of practical situations where there are either more than two levels of interest, or there are several
1 What If There Are More Than Two Factor Levels? The t-test does not directly apply ppy There are lots of practical situations where there are either more than two levels of interest, or there are several
Statistical Inference
 Statistical Inference Classical and Bayesian Methods Revision Class for Midterm Exam AMS-UCSC Th Feb 9, 2012 Winter 2012. Session 1 (Revision Class) AMS-132/206 Th Feb 9, 2012 1 / 23 Topics Topics We will
Statistical Inference Classical and Bayesian Methods Revision Class for Midterm Exam AMS-UCSC Th Feb 9, 2012 Winter 2012. Session 1 (Revision Class) AMS-132/206 Th Feb 9, 2012 1 / 23 Topics Topics We will
