Supplementary Material. Overexpression of a cytochrome P450 and a UDP-glycosyltransferase is associated with
|
|
|
- Raymond Atkinson
- 6 years ago
- Views:
Transcription
1 Supplementary Material Overexpression of a cytochrome P450 and a UDP-glycosyltransferase is associated with imidacloprid resistance in the Colorado potato beetle, Leptinotarsa decemlineata Emine Kaplanoglu 1,2, Patrick Chapman 2, Ian M. Scott 1,2 and Cam Donly 1,2, * 1 Department of Biology, The University of Western Ontario, London, ON, N6A 3K7, Canada 2 London Research and Development Centre, Agriculture and Agri-Food Canada, London, ON, N5V 4T3, Canada *Author for correspondence: Cam Donly cam.donly@agr.gc.ca 1
2 Figure S1. Volcano plot showing differentially expressed contigs between the RS and SS strains of the Colorado potato beetle contigs showed differential expression, and of these 4220 showed increased and 3352 showed decreased transcript levels in the RS beetles. Contigs that were differentially expressed at FDR of and fold change of log2 1 are colored red. 2
3 I U I U I U I U I U I U I U I U Figure S2. Production of dsrna in E. coli HT115 for target genes. I and U indicate lanes loaded with a total RNA sample extracted from bacteria that were induced or not induced with IPTG, respectively. The positions of dsrna species are marked with red arrows. Sizes of nucleic acid markers are as indicated. 3
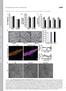
4 A) B) C) D) E) F) Figure S3. Kaplan-Meier survival curves illustrating the percent survival of the RS beetles exposed to LD 20 of imidacloprid after RNAi of target genes. Beetles either ingested E. coli HT115 (control) or E. coli HT115 producing dsrna for A) CYP4Q7, B) EST1, C) GFP, D) UGT1, E) ABC-G, and F) CYP6BQ15 genes. 4
5 Table S1. Summary of mrna-seq data before and after mapping. Sample name Total raw reads Mapped reads % mapped Uniquely mapped % uniquely mapped RS biorep1 58,892,932 51,250, ,586, RS biorep2 62,045,496 53,781, ,718, RS biorep3 55,059,038 48, ,613, SS biorep1 59,130,951 51,889, ,596, SS biorep2 61,911,228 53,098, ,677, SS biorep3 53,963,077 46,221, ,124,
6 Table S2. List of significantly differentially expressed contigs encoding detoxifying enzymes (CYP, EST, UGT, and GST) and ABC transporters in the RS beetles compared to the SS beetles. Contig ID 1 CYPs Sequence Description 2 Read count SS 3 Read count RS 3 Log2 Fold change P-adj 4 Regulation In RS Ld_rep_c34031 CYP6BQ E-32 Up Ld_rep_c51084 CYP6K E-09 Up Ld_c20712 CYP4G E-28 Up Ld_rep_c41850 CYP6BJ E-05 Up Ld_rep_c61559 CYP6EF E-06 Up Ld_rep_c91876 CYP9Z12V E-04 Up Ld_rep_c33314 CYP4Q E-38 Up Ld_rep_c48733 predicted CYP E-12 Up Ld_rep_c25417 CYP4Q E-22 Up Ld_rep_c36308 CYP6BU E-08 Up Ld_rep_c27085 CYP412A E-15 Up Ld_c756 CYP412A E-16 Up Ld_rep_c45335 CYP9Z14V E-05 Up Ld_rep_c63019 CYP413A E-07 Up Ld_c981 CYP12A E-16 Up Ld_c259 CYP6BQ E-06 Up Ld_rep_c30474 CYP301B E-06 Up Ld_c20506 CYP6EH E-04 Up Ld_c72702 CYP12H E-10 Up Ld_rep_c75503 CYP6BQ E-04 Up Ld_rep_c24490 CYP6BQ E-09 Up Ld_c22309 CYP314A E-06 Up Ld_c55986 CYP314A E-05 Up Ld_rep_c68743 CYP4G NA 9.88E-34 Up Ld_rep_c34317 CYP6BQ E-10 Down 6
7 Ld_rep_c75371 CYP412A E-06 Down Ld_c20095 CYP412A E-05 Down Ld_rep_c34168 CYP4Q E-11 Down Ld_rep_c60423 CYP4C E-19 Down Ld_rep_c48659 CYP6BK E-14 Down ESTs Ld_rep_c71421 EST E-32 Up Ld_rep_c36657 Carboxyl EST E-19 Up Ld_rep_c34698 EST E-31 Up Ld_rep_c77075 EST Beta E-05 Up Ld_rep_c35289 Acetlycholin EST E-42 Up Ld_rep_c53802 EST FE E-16 Up Ld_rep_c35399 EST E-22 Up Ld_rep_c46562 EST E-15 Up Ld_c2942 EST E-39 Up Ld_rep_c24217 EST E-10 Up Ld_c2931 Carboxyl EST E-09 Up Ld_rep_c25830 Acetyl Cholinest E-04 Up Ld_c5150 EST E-04 Up Ld_rep_c28597 EST NA 5.25E-46 Up Ld_rep_c36550 EST FE NA 1.27E-21 Down Ld_rep_c34853 Venom Carboxyl NA 7.20E-25 Down EST6 Ld_rep_c68979 Carboxyl EST E-06 Down Ld_rep_c33690 EST E-11 Down Ld_rep_c33908 EST FE E-09 Down Ld_rep_c36417 EST FE E-09 Down Ld_rep_c24505 Acetyl Cholinest E-06 Down Ld_rep_c24395 EST E-14 Down Ld_rep_c26610 Alpha EST E-21 Down 7
8 GSTs Ld_rep_c33018 GST Sigma E- 106 Up Ld_rep_c40253 GST Sigma E-13 Up Ld_rep_c24170 GST E-68 Up Ld_rep_c41971 GST Sigma E-22 Up Ld_rep_c24256 GST Delta E-18 Up Ld_rep_c26032 GST Epsilon E-13 Up Ld_rep_c44006 GST E-09 Up Ld_c19072 GST2C1-Like E-05 Up Ld_rep_c24751 GST Sigma E-04 Up Ld_rep_c38387 GST Epsilon E-06 Up Ld_rep_c33334 GST Omega E-05 Up Ld_rep_c46479 GST Theta E-04 Up Ld_rep_c25066 GST Epsilon E-04 Up Ld_rep_c50771 GST NA 1.38E-04 Up Ld_rep_c48065 GST Delta NA 2.25E-33 Up Ld_rep_c54053 GST Epsilon E-06 Down Ld_rep_c43735 GST E-20 Down Ld_rep_c34301 GST E-13 Down Ld_rep_c38198 GST E-86 Down UGTs Ld_rep_c84840 UGT2C E-52 Up Ld_rep_c41594 UGT E-30 Up Ld_rep_c83152 UGT E-20 Up Ld_rep_c45975 Antennal enriched UGT E-17 Up Ld_rep_c28339 UGT2B E-07 Up Ld_rep_c58571 UGT E-05 Up Ld_c190 UGT E-10 Up Ld_c269 UGT E-04 Up Ld_rep_c39043 UGT2B E-04 Up Ld_rep_c33389 UGT E-04 Up 8
9 Ld_rep_c38005 Antennalenriched UGT NA 1.18E-34 Up Ld_rep_c35232 UGT NA 1.96E-10 Up Ld_rep_c30928 UGT2C NA 1.77E-72 Up Ld_rep_c28388 UGT E-75 Down Ld_rep_c84951 UGT2C1-like E-05 Down ABC transporters Ld_rep_c28427 ABC-G E-05 Up Ld_rep_c27116 MRP E-04 Up Ld_c11003 ABC-B E-07 Up Ld_c571 MRP E-08 Up Ld_rep_c26545 ABC-B6 mitochondrial E-06 Up Ld_c62808 MRP NA 2.90E-27 Up Ld_rep_c34742 MRP NA 4.25E-59 Up Ld_c24118 MRP-4 like NA 5.25E-06 Up Ld_c7947 MRP like E-04 Down Ld_c12043 MRP 4-like E-07 Down Ld_c73069 MRP like E-08 Down Ld_c24114 MRP 4-like E-09 Down Ld_c56678 MRP E-07 Down Ld_c6433 MRP E-41 Down Ld_rep_c91275 MRP E-79 Down 1 Contig ID from the reference transcriptome 33 2 Genes selected for qpcr validation of mrna-seq data are bolded 3 Read counts represent mean normalized counts from three biological replicates 4 Adjusted P-value based on false discovery rate (FDR) 55 -corrected α cut-off of NA = not available 9
10 Table S3. Estimated fold change differences for seven genes in the RS beetles compared to the SS beetles from qpcr and DESeq analyses. Contig ID 1 Gene 2 Fold change Fold change Trend 4 in qpcr 3 in DESeq Ld_rep_c33314 CYP4Q S Ld_rep_c34031 CYP6BQ S Ld_rep_c34168 CYP4Q O Ld_c2942 EST S Ld_c190 UGT S Ld_rep_c41594 UGT S Ld_rep_c28427 ABC-G S 1 Contig ID from the reference transcriptome 33 2 Genes selected for RNAi studies are bolded 3 qpcr fold changes are means from independent biological samples 4 S = same trend and O = opposite trend in estimated transcript levels of the genes from two methods 10
11 Table S4. Nucleic acid sequences used for production of dsrna in E. coli HT115. Contig ID 1 Sequence Description Sequence 2 Ld_rep_c34031 CYP6BQ15 AACATCCTCACGGACCATTCCATTATTGGCGATGC CATAGACATCAAGGATGTCGTATCCCGATTCACAA CTGACGTGATAGGTTCTGTAGCTTTCGGAATAGAY TGTAATAGTCTCAAGGATCCCGACTCCGAATTCAG GCATTGGGGAAAAAGGATATTCACGTTCGATTTTA TGAGGAGAATCAAGAACAATATAACAATGTTGATT CCGAGGGATATTGTGATCAAGACAGGCATAAAATT GATGTCACGTGACTTGGAAGACTTCTTCATGAACG TGGTCAGAAGCACAGTTCAGTTCAGAGAAACTCAC AACGTCCATAGGAAGGATTTCATGCACTTGCTGTT ACAACTGAAGAATAAAGGACAAATTGCTGAGGAC GATAGCACTGATAAAGAAATCGAAATTAAGGCAC CC Ld_rep_c34168 CYP4Q7 CATCTCCTGACGTCCGAATCCACTAAAATTGTAGT AAATACATGAGATGTATAGCCATGGTTTCTTCAAC CTCTGGARAAATATGTGTCCCATTTTATTAATGGAA GTTATGTATTCTTTATCCTTTTTAGTTTTCTGGTTCA GTTTGGTCCCCATGGATGATTCTGCGATAGTATTCA ATGTAAACTCTGATATCAATGGAAAGACATTGGTC GAGGTTTTTTCAGTTTCATTTCTAAGAACTTCCACT AATTTTTTCGTTTCATTATTGAAAACTCCAACAAAT TGCTGCAAAATACTGRAATGAAAGGCTGGTGTCAM GATTTTTCGACGAGTCTGCCATTTGGAACCTGTACT TGTCAACAAGCCTTCTCCTAGCCAGCGATTCA Ld_rep_c33314 CYP4Q3 TTGGACCAGCAATCGCCTCAAATTCCTCTGCACCT GTAATGTTCACACCAGGGTAGAGAGAAGTCATGTC AATTCTGTAAATTGGTCCGTATGTTCTAGCCCAGTA TCTCACCTTGCGGAATAACGTTATATCGTCGCCCA AAAATTCAAGAACATTTTTCAAGACGGGTATAGGT TTTGGCCCAGGCAGTCCTTTCAAAATTTTCAACGTT CTCAAATGCTCGATGAAACCTCTCACAAGAAAAAA TAATATCACACATCCTGCCACCACAACTGAAAAAT TTACTATAAACATTTCAGTATATTCAATTCAAAAGT TCTTTGTCAAGAACAATCGTCTGAATAAATCTACT ACGCGTTGTGAACGCTGTCAATAATTTTTCTATTCG AACAACTTTTGAAGTGTTTTCGTGCGA 11
12 Ld_c2942 EST1 TCGAATCCAACAAGTGGTGATCCACTTTCCATGAT AGCTCCACGGAAGAGCTCTTCTCCTCCATTTTTCTG TGCTAATACATGCAGACTCACAGATAATGAACCTG AACTTTCACCTACAAGGGTAACTTTTGCAGGATCT CCTCCAAATAAATGAATATTTTCATTCACCCATTTA ATTGCAAACTTTTGATCTTTCAGGCCGATGTTGGCC GGTATAATGTCATCTTCTGTTGTGGTGAATCCAAAT GGCCCCAATCGGTAATTGAAAGTTACTACAATTAT CTCATGATCCATGATGTATTTTGGTCCAGAATATTC ATATGTACTTGATTCTGATGATAAACCTCCACCATG TATCCAAAGTAAAACTGGCAATTTCTTGGATTTACT ACCAGGTTTCAGCGGC Ld_rep_c28427 ABC-G TGGTGACTTTTCCACTGGGAATCGATTCTTTGTCTT GTGGGGTCAACATGTCTCCCCTCGTTACACTGAGT CTCCCCTCATCTTGCTCTTTACTGAGCATCAGAAAA ACGTCTTCTAATGTTTGACTATTGTAAAAAGCCAA AAGCCTTTCCGGGTTCTCTTGAGCCAAAATGATTCC CCCTCTCAACAAACAGATCTTATCAGTCTGCCTAC ATTCCTCAATATAGTGTGTGGTTATTATCACAGAA GTTCCTAGTTTCTTCGTCATATCGACTAGATAGTTC CATATTCTCTCCCTCAAAAGTGGATCAACTCCCACC GTTGGTTCATCCAGTATTAACAACTCTGGTTTGTGG ATAACTGCCGAGGCAAAGGACACTCTTC Ld_rep_c41594 UGT1 TCACTCATGGCGGTTTGTTGAGTACAACAGAAACA ATCTATCATGGTGTACCAATACTCGCTATTCCTGTT TTTGGGGATCAGAAGATGAATGCTGCAAAGGCGGT TGCAGCAGGATATGGTTTATCTTTATCAATAAACG AGTTGAGCGAGGAAAACTTATCTAACAGCATAAAT GAACTTTTGAATAATTCGAAGTATAGGGACAATGC AAAAAGAAGATCCGCAATCATGCACGATCGAAAA GTGAAGCCCATGGATCTTGCAACATATTGGATCGA ATTTGTAGTCCGACATAAAGGAGCACCACATTTGA GGGTAGCTGCCCTTGATTTAACTTGGTACCAATATT TCCTCGTGGATATTGTTCTTCTTGTGGGAGCTGTTG TTGCCAGTTTGATGCTAGCGTC Ld_c190 UGT2 TTTCCATCTCCGCATGAAATATTGCTGAAACTTGGA GAACAACTTATTTATAATGTAGCAGAATAAGAATG AAGTGAAGAATATAAATCCCAAAATATCCAACAA 12
13 GTAAAACTGGTAGAGCGGTATGTCTGAAGCGGGGT TTCGTAGCTCTTTGGCTCCTCTATTCCTTATAACGT ATTCAGTCCACCATACAGCCTTTTCCAAACCACTCA TTGGTTCATCACCAAGAAGACTGGCAAGCTTTCTA ATGGACGATTTATACTTCTCATTATGTGTAACTTCC ATGATGGCATCTCTCAAATCCTTATAGCTCAAAGC CGGTTTATGATAAATTTGTTTCCCAATATTTTTATT CTCTACGATTCGAGCGTTTTTCAGCTGATCTCCGAA GAACGGCATAGCTA - GFP CCATGCCCGAAGGTTATGTACAGGAAAGAACTATA TTTTTCAAAGATGACGGGAACTACAAGACACGTAA GTTTAAACAGTTCGGTACTAACTAACCATACATAT TTAAATTTTCAGGTGCTGAAGTCAAGTTTGAAGGT GATACCCTTGTTAATAGAATCGAGTTAAAAGGTAT TGATTTTAAAGAAGATGGAAACATTCTTGGACACA AATTGGAATACAACTATAACTCACACAATGTATAC ATCATGGCAGACAAACAAAAGAATGGAATCAAAG TTGTAAGTTTAAACATGATTTTACTAACTAACTAAT CTGATTTAAATTTTCAGAACTTCAAAATTAGACAC AACATTGAAGATGGAAGCATTCAACTAGCAGACCA TTATCAACAAAATACTCCAATTGGCGATGGCCCTG TCCTTTTACCAGACAACCATTACCTGTCC 1 Contig ID from reference transcriptome 33 2 Primers used (without restriction enzyme cut sites) are underlined 13
14 Table S5. Topical bioassay results data used to calculate resistance ratio at LD 50 of imidacloprid in adult CPB CPB Strain SS RS Dose (µg) 0.0 (acetone control) (acetone control) 4.8 BR N Time period (days) Total (M+D) M D M D M D M D M D M D M D M= total number of moribund beetles at given day D= total number of dead beetles at a given day BR = biological replicate N= number of adult beetles used per biological replicate % % mortality 1 corrected mortality Approximate LD 50 RR 3 1 Percent mortality was calculated by dividing total number of dead and moribund beetles at the end of 7 th day by total number of beetles used in the bioassay 2 Percent corrected mortality was calculated using Abbott s formula 3 Approximate LD 50 resistance ratio (RR) was calculated by dividing approximate LD 50 of imidacloprid for RS by approximate LD 50 of imidacloprid for SS
15 Table S6. List of primers used in qpcr analysis. Contig ID 1 Ld_rep_c33314 Ld_rep_c34031 Ld_rep_c34168 Ld_rep_c28427 Ld_rep_c41594 Ld_c190 Ld_c2942 Gene CYP4Q3 CYP6BQ 15 CYP4Q7 ABC-G UGT1 UGT2 EST1 - L8E - ARF1 Forward and reverse primers (5-3 ) TACCCTGGTGTGAACATTAC AATGAAAGGCTGGTGTCAAG TAGGCTGACCCCAACATTCA AATGGAATGGTCCGTGAGGA CAGCCTAAGACTTCCTTGATG GTTCGAGGGATTTGACACTAC TCACCTCCACTACAGTCAAC GCTCTGGTGGAAAGTCTAAC AGCACCACATTTGAGGGTAG GGTGAGTGAAGATGAGATCC TCTCCGAAGAACGGCATAG GAGTCATCTCTTCCCTTGAATGT ACCCTGCCACTTTTCCACTT ACTGACACAATCGGTGACG GGTAACCATCAACACATTGG TCTTGGCATCCACTTTACC GACTGCAAGTAGGAGAAGTTG TCGGCAGAGTCTACCACAT CAGGGCAAGGTTTGAAAGATAA - EF1Α CCATCAGCACAGTTCCCAT 1 Contig ID from reference transcriptome 33 Primer Amplicon Efficiency (%) 2 size (bp) Primer efficiencies were tested by generating standard curves following the guidelines described 59 3 PCR products were sequenced to confirm amplification of correct sequences 15
16 Table S7. List of primers used in cloning and sequencing of plasmid constructs. Contig ID 1 Gene Forward and reverse primers (5-3 ) 2 Product size (bp) Ld_rep_c34031 Ld_rep_c34168 Ld_rep_c33314 Ld_c2942 Ld_rep_c28427 Ld_rep_c41594 Ld_c190 CYP6BQ15 CYP4Q7 CYP4Q3 EST1 ABC-G UGT1 UGT2 - GFP - L4440 sequencing primers 1 Contig ID from the reference transcriptome 33 2 NotI and SalI restriction enzyme cut sites are underlined TAGCGGCCGCAACATCCTCACGGACCATTC ACAGGTCGACGGGTGCCTTAATTTCGATTTC TAGCGGCCGCCATCTCCTGACGTCCGAATC ACAGGTCGACTGAATCGCTGGCTAGGAGAAG TAGCGGCCGCTTGGACCAGCAATCGCCT ACAGGTCGACTCGCACGAAAACACTTCAAA TAGCGGCCGCTCGAATCCAACAAGTGGTGA ACAGGTCGACGCCGCTGAAACCTGGTAGTA TAGCGGCCGCTGGTGACTTTTCCACTGGG ACAGGTCGACGAAGAGTGTCCTTTGCCTC TAGCGGCCGCTCACTCATGGCGGTTTGTTG ACAGGTCGACGACGCTAGCATCAAACTGGC TAGCGGCCGCTTTCCATCTCCGCATGAAAT ACAGGTCGACTAGCTATGCCGTTCTTCG TAGCGGCCGCCCATGCCCGAAGGTTATGTA ACAGGTCGACGGACAGGTAATGGTTGTCTGG GACCGGCAGATCTGATATCATC CTCACTGGCCGTCGTTTTAC
17 Table S8. Plasmid constructs used in this study. Plasmid 1 Description Source L4440, Amp R RNAi feeding vector, empty backbone Addgene plasmid # 1654 GFP::L4440, Amp R Contains full length GFP sequence Addgene plasmid # GFP-RNAi::L4440, Amp R dsrna production for GFP control This study CYP4Q3::L4440, Amp R dsrna production for CYP4Q3 This study CYP4Q7::L4440, Amp R dsrna production for CYP4Q7 This study CYP6BQ15::L4440, Amp R dsrna production for CYP6BQ15 This study ABC-G::L4440, Amp R dsrna production for ABC-G This study UGT1::L4440, Amp R dsrna production for UGT1 This study UGT2::L4440, Amp R dsrna production for UGT2 This study EST1::L4440, Amp R dsrna production for EST1 This study 1 Amp R is resistance to ampicillin (β-lactamase) 17
the noisy gene Biology of the Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB)
 Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
SUPPLEMENTARY INFORMATION
 VOLUME: 1 ARTICLE NUMBER: 0010 In the format provided by the authors and unedited. Multicopy plasmids potentiate the evolution of antibiotic resistance in bacteria Alvaro San Millan 1,2,*, Jose Antonio
VOLUME: 1 ARTICLE NUMBER: 0010 In the format provided by the authors and unedited. Multicopy plasmids potentiate the evolution of antibiotic resistance in bacteria Alvaro San Millan 1,2,*, Jose Antonio
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Flow of Genetic Information
 presents Flow of Genetic Information A Montagud E Navarro P Fernández de Córdoba JF Urchueguía Elements Nucleic acid DNA RNA building block structure & organization genome building block types Amino acid
presents Flow of Genetic Information A Montagud E Navarro P Fernández de Córdoba JF Urchueguía Elements Nucleic acid DNA RNA building block structure & organization genome building block types Amino acid
2012 Univ Aguilera Lecture. Introduction to Molecular and Cell Biology
 2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
Supplementary Material. An analysis of the role of the ShSUT1 sucrose transporter in sugarcane using RNAi suppression
 10.1071/FP17073_AC CSIRO 2017 Supplementary Material: Functional Plant Biology, 2017, 44(8), 795 808. Supplementary Material An analysis of the role of the ShSUT1 sucrose transporter in sugarcane using
10.1071/FP17073_AC CSIRO 2017 Supplementary Material: Functional Plant Biology, 2017, 44(8), 795 808. Supplementary Material An analysis of the role of the ShSUT1 sucrose transporter in sugarcane using
Genomic expression catalogue of a global collection of BCG vaccine strains. show evidence for highly diverged metabolic and cell-wall adaptations.
 Genomic expression catalogue of a global collection of BCG vaccine strains show evidence for highly diverged metabolic and cell-wall adaptations. Abdallah M. Abdallah 1 *, Grant A. Hill-Cawthorne 1,2,
Genomic expression catalogue of a global collection of BCG vaccine strains show evidence for highly diverged metabolic and cell-wall adaptations. Abdallah M. Abdallah 1 *, Grant A. Hill-Cawthorne 1,2,
Daphnia magna. Genetic and plastic responses in
 Genetic and plastic responses in Daphnia magna Comparison of clonal differences and environmental stress induced changes in alternative splicing and gene expression. Jouni Kvist Institute of Biotechnology,
Genetic and plastic responses in Daphnia magna Comparison of clonal differences and environmental stress induced changes in alternative splicing and gene expression. Jouni Kvist Institute of Biotechnology,
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
The geneticist s questions. Deleting yeast genes. Functional genomics. From Wikipedia, the free encyclopedia
 From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
Topic 1 - The building blocks of. cells! Name:!
 B2 - Revision Topic 1 - The building blocks of Lesson cells Name: Topic B2.1 Plant and Animal Cells B2.2 Inside Bacteria B2.3 DNA B2.4 Extracting DNA: PCA B2.5 DNA Discovery B2.6 Genetic Engineering B2.7
B2 - Revision Topic 1 - The building blocks of Lesson cells Name: Topic B2.1 Plant and Animal Cells B2.2 Inside Bacteria B2.3 DNA B2.4 Extracting DNA: PCA B2.5 DNA Discovery B2.6 Genetic Engineering B2.7
Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis
 Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Statistics for Differential Expression in Sequencing Studies. Naomi Altman
 Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
SUPPLEMENTARY INFORMATION
 reverse 3175 3175 F L C 318 318 3185 3185 319 319 3195 3195 315 8 1 315 3155 315 317 Supplementary Figure 3. Stability of expression of the GFP sensor constructs return to warm conditions. Semi-quantitative
reverse 3175 3175 F L C 318 318 3185 3185 319 319 3195 3195 315 8 1 315 3155 315 317 Supplementary Figure 3. Stability of expression of the GFP sensor constructs return to warm conditions. Semi-quantitative
Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated
 Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated form by recombinant UGT74E2. The naturally occurring auxin
Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated form by recombinant UGT74E2. The naturally occurring auxin
Supplemental Information
 Molecular Cell, Volume 52 Supplemental Information The Translational Landscape of the Mammalian Cell Cycle Craig R. Stumpf, Melissa V. Moreno, Adam B. Olshen, Barry S. Taylor, and Davide Ruggero Supplemental
Molecular Cell, Volume 52 Supplemental Information The Translational Landscape of the Mammalian Cell Cycle Craig R. Stumpf, Melissa V. Moreno, Adam B. Olshen, Barry S. Taylor, and Davide Ruggero Supplemental
Figure S1. UPGMA dendrogram of 24 isolates used in this study based on 17 alleles
 Figure S1. UPGMA dendrogram of 24 isolates used in this study based on 17 alleles from seven genomic microsatellite markers. The scale indicates percent distance. Asterisks indicate isolates with reduced
Figure S1. UPGMA dendrogram of 24 isolates used in this study based on 17 alleles from seven genomic microsatellite markers. The scale indicates percent distance. Asterisks indicate isolates with reduced
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and
 Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1
 Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
What Organelle Makes Proteins According To The Instructions Given By Dna
 What Organelle Makes Proteins According To The Instructions Given By Dna This is because it contains the information needed to make proteins. assemble enzymes and other proteins according to the directions
What Organelle Makes Proteins According To The Instructions Given By Dna This is because it contains the information needed to make proteins. assemble enzymes and other proteins according to the directions
Aquaculture Biology Laboratory
 Aquaculture Biology Laboratory Faculty of Fisheries Nagasaki University Professor: Dr. Atsushi Hagiwara (hagiwara@net.nagasaki-u.ac.jp) Associate Professor: Dr. Yoshitaka Sakakura (sakakura@net.nagasaki-u.ac.jp)
Aquaculture Biology Laboratory Faculty of Fisheries Nagasaki University Professor: Dr. Atsushi Hagiwara (hagiwara@net.nagasaki-u.ac.jp) Associate Professor: Dr. Yoshitaka Sakakura (sakakura@net.nagasaki-u.ac.jp)
Supplementary Information. Drought response transcriptomics are altered in poplar with reduced tonoplast sucrose transporter expression
 Supplementary Information Drought response transcriptomics are altered in poplar with reduced tonoplast sucrose transporter expression Liang Jiao Xue, Christopher J. Frost, Chung Jui Tsai, Scott A. Harding
Supplementary Information Drought response transcriptomics are altered in poplar with reduced tonoplast sucrose transporter expression Liang Jiao Xue, Christopher J. Frost, Chung Jui Tsai, Scott A. Harding
g A n(a, g) n(a, ḡ) = n(a) n(a, g) n(a) B n(b, g) n(a, ḡ) = n(b) n(b, g) n(b) g A,B A, B 2 RNA-seq (D) RNA mrna [3] RNA 2. 2 NGS 2 A, B NGS n(
![g A n(a, g) n(a, ḡ) = n(a) n(a, g) n(a) B n(b, g) n(a, ḡ) = n(b) n(b, g) n(b) g A,B A, B 2 RNA-seq (D) RNA mrna [3] RNA 2. 2 NGS 2 A, B NGS n( g A n(a, g) n(a, ḡ) = n(a) n(a, g) n(a) B n(b, g) n(a, ḡ) = n(b) n(b, g) n(b) g A,B A, B 2 RNA-seq (D) RNA mrna [3] RNA 2. 2 NGS 2 A, B NGS n(](/thumbs/77/74495778.jpg) ,a) RNA-seq RNA-seq Cuffdiff, edger, DESeq Sese Jun,a) Abstract: Frequently used biological experiment technique for observing comprehensive gene expression has been changed from microarray using cdna
,a) RNA-seq RNA-seq Cuffdiff, edger, DESeq Sese Jun,a) Abstract: Frequently used biological experiment technique for observing comprehensive gene expression has been changed from microarray using cdna
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
DNA Technology, Bacteria, Virus and Meiosis Test REVIEW
 Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
AP Bio Module 16: Bacterial Genetics and Operons, Student Learning Guide
 Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Boolean models of gene regulatory networks. Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016
 Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
2. What was the Avery-MacLeod-McCarty experiment and why was it significant? 3. What was the Hershey-Chase experiment and why was it significant?
 Name Date Period AP Exam Review Part 6: Molecular Genetics I. DNA and RNA Basics A. History of finding out what DNA really is 1. What was Griffith s experiment and why was it significant? 1 2. What was
Name Date Period AP Exam Review Part 6: Molecular Genetics I. DNA and RNA Basics A. History of finding out what DNA really is 1. What was Griffith s experiment and why was it significant? 1 2. What was
The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector.
 The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector. Omar S. Akbari*, Igor Antoshechkin*, Henry Amrhein, Brian Williams, Race Diloreto, Jeremy
The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector. Omar S. Akbari*, Igor Antoshechkin*, Henry Amrhein, Brian Williams, Race Diloreto, Jeremy
itraq and RNA-Seq analyses provide new insights of Dendrobium officinale seeds (Orchidaceae)
 Page S-1 Supporting Information (SI) itraq and RNA-Seq analyses provide new insights into regulation mechanism of symbiotic germination of Dendrobium officinale seeds (Orchidaceae) Juan Chen 1, Si Si Liu
Page S-1 Supporting Information (SI) itraq and RNA-Seq analyses provide new insights into regulation mechanism of symbiotic germination of Dendrobium officinale seeds (Orchidaceae) Juan Chen 1, Si Si Liu
Lecture 4: Transcription networks basic concepts
 Lecture 4: Transcription networks basic concepts - Activators and repressors - Input functions; Logic input functions; Multidimensional input functions - Dynamics and response time 2.1 Introduction The
Lecture 4: Transcription networks basic concepts - Activators and repressors - Input functions; Logic input functions; Multidimensional input functions - Dynamics and response time 2.1 Introduction The
*Equal contribution Contact: (TT) 1 Department of Biomedical Engineering, the Engineering Faculty, Tel Aviv
 Supplementary of Complementary Post Transcriptional Regulatory Information is Detected by PUNCH-P and Ribosome Profiling Hadas Zur*,1, Ranen Aviner*,2, Tamir Tuller 1,3 1 Department of Biomedical Engineering,
Supplementary of Complementary Post Transcriptional Regulatory Information is Detected by PUNCH-P and Ribosome Profiling Hadas Zur*,1, Ranen Aviner*,2, Tamir Tuller 1,3 1 Department of Biomedical Engineering,
Bias in RNA sequencing and what to do about it
 Bias in RNA sequencing and what to do about it Walter L. (Larry) Ruzzo Computer Science and Engineering Genome Sciences University of Washington Fred Hutchinson Cancer Research Center Seattle, WA, USA
Bias in RNA sequencing and what to do about it Walter L. (Larry) Ruzzo Computer Science and Engineering Genome Sciences University of Washington Fred Hutchinson Cancer Research Center Seattle, WA, USA
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
A pentose bisphosphate pathway for nucleoside degradation in Archaea. Engineering, Kyoto University, Katsura, Nishikyo-ku, Kyoto , Japan.
 SUPPLEMENTARY INFORMATION A pentose bisphosphate pathway for nucleoside degradation in Archaea Riku Aono 1,, Takaaki Sato 1,, Tadayuki Imanaka, and Haruyuki Atomi 1, * 7 8 9 10 11 1 Department of Synthetic
SUPPLEMENTARY INFORMATION A pentose bisphosphate pathway for nucleoside degradation in Archaea Riku Aono 1,, Takaaki Sato 1,, Tadayuki Imanaka, and Haruyuki Atomi 1, * 7 8 9 10 11 1 Department of Synthetic
Insecticide resistance experiments
 Applied Crop Protection 2015 IX Insecticide resistance experiments Caroline Kaiser, Dorte H. Højland, Karl-Martin V. Jensen and Michael Kristensen Insecticide resistance is less of an issue than that of
Applied Crop Protection 2015 IX Insecticide resistance experiments Caroline Kaiser, Dorte H. Højland, Karl-Martin V. Jensen and Michael Kristensen Insecticide resistance is less of an issue than that of
Lesson 11. Functional Genomics I: Microarray Analysis
 Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
EBSeq: An R package for differential expression analysis using RNA-seq data
 EBSeq: An R package for differential expression analysis using RNA-seq data Ning Leng, John Dawson, and Christina Kendziorski October 14, 2013 Contents 1 Introduction 2 2 Citing this software 2 3 The Model
EBSeq: An R package for differential expression analysis using RNA-seq data Ning Leng, John Dawson, and Christina Kendziorski October 14, 2013 Contents 1 Introduction 2 2 Citing this software 2 3 The Model
Supplemental Data. Wang et al. (2014). Plant Cell /tpc
 Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Genome sequence of Plasmopara viticola and insight into the pathogenic mechanism
 Genome sequence of Plasmopara viticola and insight into the pathogenic mechanism Ling Yin 1,3,, Yunhe An 1,2,, Junjie Qu 3,, Xinlong Li 1, Yali Zhang 1, Ian Dry 5, Huijun Wu 2*, Jiang Lu 1,4** 1 College
Genome sequence of Plasmopara viticola and insight into the pathogenic mechanism Ling Yin 1,3,, Yunhe An 1,2,, Junjie Qu 3,, Xinlong Li 1, Yali Zhang 1, Ian Dry 5, Huijun Wu 2*, Jiang Lu 1,4** 1 College
1. Which of the following has the lowest vapor pressure? A) H 2 O B) NaCl C) NH 3 D) O 2 E) CH 4
 Name: Date: 1. Which of the following has the lowest vapor pressure? A) H O B) NaCl C) NH 3 D) O E) CH 4. Which of the following species exhibit hydrogen bonding? (Check all that apply.) A) HBr B) NO 3
Name: Date: 1. Which of the following has the lowest vapor pressure? A) H O B) NaCl C) NH 3 D) O E) CH 4. Which of the following species exhibit hydrogen bonding? (Check all that apply.) A) HBr B) NO 3
L3.1: Circuits: Introduction to Transcription Networks. Cellular Design Principles Prof. Jenna Rickus
 L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
UNIVERSITY OF YORK. BA, BSc, and MSc Degree Examinations Department : BIOLOGY. Title of Exam: Molecular microbiology
 Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Mini-Tn7 Derivative Construction and Characterization. Mini-Tn7 derivatives for
 Supplemental Methods Mini-Tn7 Derivative Construction and Characterization. Mini-Tn7 derivatives for constitutive expression of fluorescent proteins in S. oneidensis were constructed as follows. The EcoRI-XbaI
Supplemental Methods Mini-Tn7 Derivative Construction and Characterization. Mini-Tn7 derivatives for constitutive expression of fluorescent proteins in S. oneidensis were constructed as follows. The EcoRI-XbaI
Count ratio model reveals bias affecting NGS fold changes
 Published online 8 July 2015 Nucleic Acids Research, 2015, Vol. 43, No. 20 e136 doi: 10.1093/nar/gkv696 Count ratio model reveals bias affecting NGS fold changes Florian Erhard * and Ralf Zimmer Institut
Published online 8 July 2015 Nucleic Acids Research, 2015, Vol. 43, No. 20 e136 doi: 10.1093/nar/gkv696 Count ratio model reveals bias affecting NGS fold changes Florian Erhard * and Ralf Zimmer Institut
Hiromi Nishida. 1. Introduction. 2. Materials and Methods
 Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
CST and FINAL EXAM REVIEW
 Name Date Period CST and FINAL EXAM REVIEW Directions: Both your final exam and the CST (STAR) test are based on the California Standards. There are five major categories and they include: Investigation
Name Date Period CST and FINAL EXAM REVIEW Directions: Both your final exam and the CST (STAR) test are based on the California Standards. There are five major categories and they include: Investigation
Biology 105/Summer Bacterial Genetics 8/12/ Bacterial Genomes p Gene Transfer Mechanisms in Bacteria p.
 READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
JMJ14-HA. Col. Col. jmj14-1. jmj14-1 JMJ14ΔFYR-HA. Methylene Blue. Methylene Blue
 Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
Fig. S1 JMJ14 JMJ14 JMJ14ΔFYR Methylene Blue Col jmj14-1 JMJ14-HA Methylene Blue Col jmj14-1 JMJ14ΔFYR-HA Fig. S1. The expression level of JMJ14 and truncated JMJ14 with FYR (FYRN + FYRC) domain deletion
Supporting Information
 Supporting Information Theoretical Approach for Determination of True Quantitative Values of Proteins in Pure Membrane Fraction. During the preparation of membrane fractions, some contamination with other
Supporting Information Theoretical Approach for Determination of True Quantitative Values of Proteins in Pure Membrane Fraction. During the preparation of membrane fractions, some contamination with other
Supplementary Information for. Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria
 Supplementary Information for Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria Isabella Santi 1 *, Neeraj Dhar 1, Djenet Bousbaine 1, Yuichi Wakamoto, John D.
Supplementary Information for Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria Isabella Santi 1 *, Neeraj Dhar 1, Djenet Bousbaine 1, Yuichi Wakamoto, John D.
Upstream Elements Regulating mir-241 and mir-48 Abstract Introduction
 Upstream Elements Regulating mir-241 and mir-48 Hanna Vollbrecht, Tamar Resnick, and Ann Rougvie University of Minnesota: Twin Cities Undergraduate Research Scholarship 2012-2013 Abstract Caenorhabditis
Upstream Elements Regulating mir-241 and mir-48 Hanna Vollbrecht, Tamar Resnick, and Ann Rougvie University of Minnesota: Twin Cities Undergraduate Research Scholarship 2012-2013 Abstract Caenorhabditis
Comparative analysis of RNA- Seq data with DESeq2
 Comparative analysis of RNA- Seq data with DESeq2 Simon Anders EMBL Heidelberg Two applications of RNA- Seq Discovery Eind new transcripts Eind transcript boundaries Eind splice junctions Comparison Given
Comparative analysis of RNA- Seq data with DESeq2 Simon Anders EMBL Heidelberg Two applications of RNA- Seq Discovery Eind new transcripts Eind transcript boundaries Eind splice junctions Comparison Given
** * * * Col-0 cau1 CAU1. Actin2 CAS. Actin2. Supplemental Figure 1. CAU1 affects calcium accumulation.
 Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Supplemental Data. Fu et al. Plant Cell. (213). 1.115/tpc.113.113886 A 5 4 3 * Col- cau1 B 4 3 2 Col- cau1 ** * * ** C 2 1 25 2 15 1 5 Shoots Roots *
Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Supplemental Data. Fu et al. Plant Cell. (213). 1.115/tpc.113.113886 A 5 4 3 * Col- cau1 B 4 3 2 Col- cau1 ** * * ** C 2 1 25 2 15 1 5 Shoots Roots *
Gene Expression. Molecular Genetics, March, 2018
 Gene Expression Molecular Genetics, March, 2018 Gene Expression Control of Protein Levels Bacteria Lac Operon Promoter mrna Inducer CAP Control Trp Operon RepressorOperator Control Attenuation Riboswitches
Gene Expression Molecular Genetics, March, 2018 Gene Expression Control of Protein Levels Bacteria Lac Operon Promoter mrna Inducer CAP Control Trp Operon RepressorOperator Control Attenuation Riboswitches
Nature Genetics: doi: /ng Supplementary Figure 1. Plasmid diagrams of primary strains used in this work.
 Supplementary Figure 1 Plasmid diagrams of primary strains used in this work. Refer to Supplementary Table 1 for additional information and full list of strains. Diagrams are not to scale. (a) Strain used
Supplementary Figure 1 Plasmid diagrams of primary strains used in this work. Refer to Supplementary Table 1 for additional information and full list of strains. Diagrams are not to scale. (a) Strain used
Molecular Developmental Physiology and Signal Transduction
 Prof. Dr. J. Vanden Broeck (Animal Physiology and Neurobiology - Dept. of Biology - KU Leuven) Molecular Developmental Physiology and Signal Transduction My Research Team Insect species under study +
Prof. Dr. J. Vanden Broeck (Animal Physiology and Neurobiology - Dept. of Biology - KU Leuven) Molecular Developmental Physiology and Signal Transduction My Research Team Insect species under study +
Supplementary Figure 1 Biochemistry of gene duplication
 Supplementary Figure 1 Biochemistry of gene duplication (a) (b) (c) (d) A B C A (e) Selection (f) reca KO Supplementary Figure 1: Tandem gene duplication: construction, amplification, and stabilization.
Supplementary Figure 1 Biochemistry of gene duplication (a) (b) (c) (d) A B C A (e) Selection (f) reca KO Supplementary Figure 1: Tandem gene duplication: construction, amplification, and stabilization.
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data
 DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
RNA-seq to study rice defense responses upon parasitic nematode infections. Tina Kyndt
 RNA-seq to study rice defense responses upon parasitic nematode infections Tina Kyndt 1 Rice (Oryza sativa L.) Most important staple food for at least half of the human population Monocot model plant Paddy
RNA-seq to study rice defense responses upon parasitic nematode infections Tina Kyndt 1 Rice (Oryza sativa L.) Most important staple food for at least half of the human population Monocot model plant Paddy
Supplementary Information
 Supplementary Information Supplementary figures % Occupancy 90 80 70 60 50 40 30 20 Wt tol-1(nr2033) Figure S1. Avoidance behavior to S. enterica was not observed in wild-type or tol-1(nr2033) mutant nematodes.
Supplementary Information Supplementary figures % Occupancy 90 80 70 60 50 40 30 20 Wt tol-1(nr2033) Figure S1. Avoidance behavior to S. enterica was not observed in wild-type or tol-1(nr2033) mutant nematodes.
BTRY 7210: Topics in Quantitative Genomics and Genetics
 BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu February 12, 2015 Lecture 3:
BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu February 12, 2015 Lecture 3:
DEXSeq paper discussion
 DEXSeq paper discussion L Collado-Torres December 10th, 2012 1 / 23 1 Background 2 DEXSeq paper 3 Results 2 / 23 Gene Expression 1 Background 1 Source: http://www.ncbi.nlm.nih.gov/projects/genome/probe/doc/applexpression.shtml
DEXSeq paper discussion L Collado-Torres December 10th, 2012 1 / 23 1 Background 2 DEXSeq paper 3 Results 2 / 23 Gene Expression 1 Background 1 Source: http://www.ncbi.nlm.nih.gov/projects/genome/probe/doc/applexpression.shtml
Fitness constraints on horizontal gene transfer
 Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Title: A novel mechanism of protein thermostability: a unique N-terminal domain confers
 1 2 Title: A novel mechanism of protein thermostability: a unique N-terminal domain confers heat resistance to Fe/Mn-SODs 3 4 Running Title: Thermostability-improving peptide for SODs 5 6 7 8 Authors Wei
1 2 Title: A novel mechanism of protein thermostability: a unique N-terminal domain confers heat resistance to Fe/Mn-SODs 3 4 Running Title: Thermostability-improving peptide for SODs 5 6 7 8 Authors Wei
The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas
 The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas Is there a Role for the CCM in Increasing Biological CO 2 Capture? Generalized
The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas Is there a Role for the CCM in Increasing Biological CO 2 Capture? Generalized
Lesson Overview. Gene Regulation and Expression. Lesson Overview Gene Regulation and Expression
 13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
Name: SBI 4U. Gene Expression Quiz. Overall Expectation:
 Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Supplementary Table 1. Primers used in this study.
 Supplementary Tale 1. Primers used in this study. Name Primer sequence (5'-3') Primers of PCR-ased molecular markers developed in this study M1 F M1 R M2 F M2 R M3 F M3 R M4 F M4 R M5 F M5 R M6 F M6 R
Supplementary Tale 1. Primers used in this study. Name Primer sequence (5'-3') Primers of PCR-ased molecular markers developed in this study M1 F M1 R M2 F M2 R M3 F M3 R M4 F M4 R M5 F M5 R M6 F M6 R
Characters related to higher starch accumulation in cassava storage roots
 Characters related to higher starch accumulation in cassava storage roots You-Zhi Li **1, Jian-Yu Zhao *1, San-Min Wu 1, Xian-Wei Fan 1, Xing-Lu Luo 1 & Bao-Shan Chen **1 1 State Key Laboratory for Conservation
Characters related to higher starch accumulation in cassava storage roots You-Zhi Li **1, Jian-Yu Zhao *1, San-Min Wu 1, Xian-Wei Fan 1, Xing-Lu Luo 1 & Bao-Shan Chen **1 1 State Key Laboratory for Conservation
By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food Supervised by Henrik Hasman, PhD
 By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food elibi@food.dtu.dk Supervised by Henrik Hasman, PhD 1. Introduction to plasmid biology 2. Plasmid encoded resistance to β- lactams (basic theories)
By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food elibi@food.dtu.dk Supervised by Henrik Hasman, PhD 1. Introduction to plasmid biology 2. Plasmid encoded resistance to β- lactams (basic theories)
Insect/Bacterial Symbioses Aphid/Buchnera association
 Insect/Bacterial Symbioses Aphid/Buchnera association I. Introduction A. Intracellular symbioses are common in the order Homoptera, which includes aphids, mealy bugs, whiteflies, and cicadas, Blattaria,
Insect/Bacterial Symbioses Aphid/Buchnera association I. Introduction A. Intracellular symbioses are common in the order Homoptera, which includes aphids, mealy bugs, whiteflies, and cicadas, Blattaria,
Chapter 12. Genes: Expression and Regulation
 Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
Supporting Information
 Supporting Information Cao et al. 10.1073/pnas.1306220110 Gram - bacteria Hemolymph Cytoplasm PGRP-LC TAK1 signalosome Imd dfadd Dredd Dnr1 Ikk signalosome P Relish Nucleus AMP and effector genes Fig.
Supporting Information Cao et al. 10.1073/pnas.1306220110 Gram - bacteria Hemolymph Cytoplasm PGRP-LC TAK1 signalosome Imd dfadd Dredd Dnr1 Ikk signalosome P Relish Nucleus AMP and effector genes Fig.
Normalization and differential analysis of RNA-seq data
 Normalization and differential analysis of RNA-seq data Nathalie Villa-Vialaneix INRA, Toulouse, MIAT (Mathématiques et Informatique Appliquées de Toulouse) nathalie.villa@toulouse.inra.fr http://www.nathalievilla.org
Normalization and differential analysis of RNA-seq data Nathalie Villa-Vialaneix INRA, Toulouse, MIAT (Mathématiques et Informatique Appliquées de Toulouse) nathalie.villa@toulouse.inra.fr http://www.nathalievilla.org
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Expression of nuclearencoded. photosynthesis in sea slug (Elysia chlorotica)
 Expression of nuclearencoded proteins for photosynthesis in sea slug (Elysia chlorotica) Timothy Youngblood Writer s Comment: This paper was written as an assignment for UWP 102B instructed by Jared Haynes.
Expression of nuclearencoded proteins for photosynthesis in sea slug (Elysia chlorotica) Timothy Youngblood Writer s Comment: This paper was written as an assignment for UWP 102B instructed by Jared Haynes.
Supplementary Figure 1 The number of differentially expressed genes for uniparental males (green), uniparental females (yellow), biparental males
 Supplementary Figure 1 The number of differentially expressed genes for males (green), females (yellow), males (red), and females (blue) in caring vs. control comparisons in the caring gene set and the
Supplementary Figure 1 The number of differentially expressed genes for males (green), females (yellow), males (red), and females (blue) in caring vs. control comparisons in the caring gene set and the
The geneticist s questions
 The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
Other host processes. Key. Orthogonal translation
 Core host processes s e s e s e p T s i s i s i p E Metabolism Other host processes p H Host gene expression X {E, T, H} e e e Key Host mrnas Circuit mrnas Proteins ω X (e) e Core ribosomal mrnas h-rrna
Core host processes s e s e s e p T s i s i s i p E Metabolism Other host processes p H Host gene expression X {E, T, H} e e e Key Host mrnas Circuit mrnas Proteins ω X (e) e Core ribosomal mrnas h-rrna
Supplementary Materials for
 www.advances.sciencemag.org/cgi/content/full/1/5/e1500358/dc1 Supplementary Materials for Transplantability of a circadian clock to a noncircadian organism Anna H. Chen, David Lubkowicz, Vivian Yeong,
www.advances.sciencemag.org/cgi/content/full/1/5/e1500358/dc1 Supplementary Materials for Transplantability of a circadian clock to a noncircadian organism Anna H. Chen, David Lubkowicz, Vivian Yeong,
Variation in the genetic response to high temperature in Montastraea faveolata from the Florida Keys & Mexico
 Variation in the genetic response to high temperature in Montastraea faveolata from the Florida Keys & Mexico Nicholas R. Polato 1, Christian R. Voolstra 2, Julia Schnetzer 3, Michael K. DeSalvo 4, Carly
Variation in the genetic response to high temperature in Montastraea faveolata from the Florida Keys & Mexico Nicholas R. Polato 1, Christian R. Voolstra 2, Julia Schnetzer 3, Michael K. DeSalvo 4, Carly
4. Identify one bird that would most likely compete for food with the large tree finch. Support your answer. [1]
![4. Identify one bird that would most likely compete for food with the large tree finch. Support your answer. [1] 4. Identify one bird that would most likely compete for food with the large tree finch. Support your answer. [1]](/thumbs/84/91138454.jpg) Name: Topic 5B 1. A hawk has a genetic trait that gives it much better eyesight than other hawks of the same species in the same area. Explain how this could lead to evolutionary change within this species
Name: Topic 5B 1. A hawk has a genetic trait that gives it much better eyesight than other hawks of the same species in the same area. Explain how this could lead to evolutionary change within this species
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene Regulation and Expression
 THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium that contains more than 4000 genes.
THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium that contains more than 4000 genes.
56:198:582 Biological Networks Lecture 8
 56:198:582 Biological Networks Lecture 8 Course organization Two complementary approaches to modeling and understanding biological networks Constraint-based modeling (Palsson) System-wide Metabolism Steady-state
56:198:582 Biological Networks Lecture 8 Course organization Two complementary approaches to modeling and understanding biological networks Constraint-based modeling (Palsson) System-wide Metabolism Steady-state
Supplementary Figure 1. The proportion S. aureus CFU of the total CFU (S. aureus + E. faecalis CFU) per host in worms
 Supplementary Figure 1. The proportion S. aureus CFU of the total CFU (S. aureus + E. faecalis CFU) per host in worms alive or dead at 24 hours of exposure. Two sample t-test: t = 1.22, df = 10, P= 0.25.
Supplementary Figure 1. The proportion S. aureus CFU of the total CFU (S. aureus + E. faecalis CFU) per host in worms alive or dead at 24 hours of exposure. Two sample t-test: t = 1.22, df = 10, P= 0.25.
Introduction to molecular biology. Mitesh Shrestha
 Introduction to molecular biology Mitesh Shrestha Molecular biology: definition Molecular biology is the study of molecular underpinnings of the process of replication, transcription and translation of
Introduction to molecular biology Mitesh Shrestha Molecular biology: definition Molecular biology is the study of molecular underpinnings of the process of replication, transcription and translation of
2. Yeast two-hybrid system
 2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids. Prepared by Woojoo Choi
 Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids Prepared by Woojoo Choi Dead or alive? 1) In this chapter we will explore the twilight zone of biology and the gene creature who live there.
Chapter 19. Gene creatures, Part 1: viruses, viroids and plasmids Prepared by Woojoo Choi Dead or alive? 1) In this chapter we will explore the twilight zone of biology and the gene creature who live there.
Microbiome: 16S rrna Sequencing 3/30/2018
 Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
1.9 Practice Problems
 1.9 Practice Problems 1. Solution: B It s not only chlorophyll a but a combination of pigments. 2. Solution: D See at what wavelength rate of photosynthesis is the highest. 3. Solution: D It s a fact.
1.9 Practice Problems 1. Solution: B It s not only chlorophyll a but a combination of pigments. 2. Solution: D See at what wavelength rate of photosynthesis is the highest. 3. Solution: D It s a fact.
AP Biology. Free-Response Questions
 2018 AP Biology Free-Response Questions College Board, Advanced Placement Program, AP, AP Central, and the acorn logo are registered trademarks of the College Board. AP Central is the official online home
2018 AP Biology Free-Response Questions College Board, Advanced Placement Program, AP, AP Central, and the acorn logo are registered trademarks of the College Board. AP Central is the official online home
Microbial DNA qpcr Multi-Assay Kit Clostridium perfringens Pathogenicity
 Microbial DNA qpcr Multi-Assay Kit Clostridium perfringens Pathogenicity Cat. no. 330043 BBID-1507ZR-3 For real-time PCR-based, application-specific microbial identification or profiling The Clostridium
Microbial DNA qpcr Multi-Assay Kit Clostridium perfringens Pathogenicity Cat. no. 330043 BBID-1507ZR-3 For real-time PCR-based, application-specific microbial identification or profiling The Clostridium
MicroRNA mir-34 provides robustness to environmental stress response via
 MicroRNA mir-34 provides robustness to environmental stress respoe via the DAF-16 network in C. elega Meltem Isik 1,2, T. Keith Blackwell 2, and Eugene Berezikov 1,3 1 Hubrecht Ititute-KNAW and University
MicroRNA mir-34 provides robustness to environmental stress respoe via the DAF-16 network in C. elega Meltem Isik 1,2, T. Keith Blackwell 2, and Eugene Berezikov 1,3 1 Hubrecht Ititute-KNAW and University
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Bacterial Genetics & Operons
 Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
pglo/amp R Bacterial Transformation Lab
 pglo/amp R Bacterial Transformation Lab Name: Date: Purpose: To gain an understanding of the techniques of culturing E. coli bacteria and transforming E. coli bacteria using genetic engineering. Introduction:
pglo/amp R Bacterial Transformation Lab Name: Date: Purpose: To gain an understanding of the techniques of culturing E. coli bacteria and transforming E. coli bacteria using genetic engineering. Introduction:
