Supplementary Figure 1. The proportion S. aureus CFU of the total CFU (S. aureus + E. faecalis CFU) per host in worms
|
|
|
- Coleen Clarke
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1. The proportion S. aureus CFU of the total CFU (S. aureus + E. faecalis CFU) per host in worms alive or dead at 24 hours of exposure. Two sample t-test: t = 1.22, df = 10, P= Sample size for each treatment: 6 biological replicates. Supplementary Figure 2. The relationship between OD630 and siderophore concentration (ug ml ¹) in vitro. Nonlinear (weighted) least-squares estimates of the parameters of a nonlinear model: OD = *exp(- 0.04*concentration). Sample size: 2 biological replicates for each concentration.
2 Supplementary Table 1. Statistical results listed by figure Figure 2a 2b 2c 4a Statistical results Quasibinomial GLM: F=41.96, df=2, P= 5.193e-09 Tukey contrasts: S. aureus vs. E. faecalis: P<1e-04 S. aureus vs. co-colonisation: P<1e-04 E. faecalis vs. co-colonisation: P=0.025 Sample size for each treatment: 10 biological replicates. Welch two sample t test: t=-4.5, df=16.2, P= Sample size for S aureus: 16 biological replicates. Sample size for co-infection: 18 biological replicates. Experiment replicated 3 times. Two sample t test: t=-2.8, df=24, P= Sample size for E faecalis: 11 biological replicates. Sample size for co-infection: 15 biological replicates. Quasibinomial GLM: F=25.7, df=2, P=4.627e-05 Tukey contrasts: Coevolution vs. Ancestor: P<0.001 Coevolution vs. Single evolution: P<0.001 Ancestor vs. Single evolution: P=0.035 Sample size for each treatment: 5 biological replicates (average of 2 technical replicates). 4b 4c Pearson's product-moment correlation: t = 3.38, df = 9, p= R²= Sample size for virulence data: as fig.4a but all ancestral replicates averaged to 1 biological replicate. Sample size for growth rate data: 5 biological replicates (average of 3 technical replicates) for evolved bacteria. 1 biological replicate for ancestral bacteria (average of 8 technical replicates). Anomalous technical replicate points identified and removed using the Dixon test. ANOVA: F=6.55, df=2, P=0.012 Tukey contrasts & 95% family-wise confidence level: diff lwr upr p value Ancestor vs Coevolution: Single evolution vs Coevolution: Single evolution vs. Ancestor: Sample size for each treatment: 5 biological replicates (1 technical replicate).
3 5a 5b ANOVA: F=6.681, df=2, P=0.01 Tukey contrasts & 95% family-wise confidence level: diff lwr upr p value Coevolution vs. Ancestor: Single Evolution vs. Ancestor: Single Evolution vs Coevolution: Sample size for each treatment: 5 biological replicates (average of 2 technical replicates). Pearson's product-moment correlation: t = 2.8, df = 9, P=0.02. R²= Sample size for virulence data: as fig.4a but all ancestral replicates averaged to 1 biological replicate. Sample size for siderophore data: as fig.5a but all ancestral replicates averaged to 1 biological replicate. 7a 7b 7c Pearson's product-moment correlation: t=6.1, df=2, P= R²= Sample size: 1 biological replicate for each concentration (average of 2 technical replicates). ANOVA: F=9.2, df=3, P= Dunnett contrasts: Coevolution S. aureus & ancestral E. faecalis vs. ancestral E. faecalis: P=0.77 Single evolution S. aureus & ancestral E. faecalis vs. ancestral E. faecalis: P=0.023 Ancestor S. aureus & ancestral E. faecalis vs. ancestral E. faecalis: P<0.001 Sample size for each treatment: 5 biological replicates (1 technical replicate). Control ANOVA: F=8.1, df=2, fdr corrected P=0.019 Tukey contrasts & 95% family-wise confidence level: diff lwr upr p value Ancestor vs Coevolution: Single evolution vs Coevolution: Single evolution vs. Ancestor: Fe³+ ANOVA Treatment: F=3.05, df=2, fdr corrected P= Sample size for each treatment of both ANOVAs: 5 biological replicates (1 technical replicate). Anomalous points identified and removed using the Dixon test. Supp 1 Two sample t-test: t = 1.22, df = 10, P= Sample size for each treatment: 6 biological replicates. Supp 2 Nonlinear (weighted) least-squares estimates of the parameters of a nonlinear model: OD = *exp(-0.04*concentration) Parameters: Estimate Std. Error t value Pr(> t ) a e-08 *** b e-15 *** c e-09 *** Signif. codes: 0 *** ** 0.01 * Residual standard error: on 13 degrees of freedom
4 Number of iterations to convergence: 4 Achieved convergence tolerance: 2.732e-07 Sample size: 2 biological replicates for each concentration.
5 Coevolution Treatment Replicate type Supplementary Table 2. s found in S. aureus listed by treatment and replicate population. Locus tag is given when mutation occurs in a coding region. Otherwise, the coordinate is given. abundance is the percentage of 40 sampled clones that contain the mutation in the replicate population. Only mutations present in >20% clones are listed to control false positives at the expense of low frequency true positives. Gene names and putative gene function are given where known, along with a reference. A proposed link with siderophore activity is listed where identified. type: S=synonymous. N=non-synonymous. *=stop gain. */= stop loss. Locus tag/ coordinate abundanc e (%) Gene name/annotation Putative function Putative link with siderophores Reference (PMID) 1 SAS_RS09425 N 95 Protein export protein PrsA Foldase. 2 SAS_RS00600 * 98 Hypothetical Between pyrimidine nucleoside transporter NupC & transcriptional regulator CtsR. SAS_RS02825 N 30 Glycosyl transferase family 1. SAS_RS02905 S 100 Uracil-DNA glycosylase. SAS_RS04210 N 23 Cysteine desulfurase. SAS_RS13205 N 98 Holin-like protein CidB Between octanoyl- [GcvH]:protein N- octanoyltransferas e and mevalonate kinase. SAS_RS05930 N 43 Penicillin-binding Between DNA topoisomerase 1 and TrmFO. A facultative facilitator of protein secretion or extracellular folding. Cell wall ABC transporter superfamily. Cell wall DNA baseexcision repair. iron-sulphur cluster Cytolysis. Cellular suicide in response to cellular stress or damage. Cell wall TrmFO: oxidoreductase flavo Excretion or folding of proteins Transport of substrates TRmFO: Siderophoreiron reductase
6 Treatment Replicate type Locus tag/ coordinate abundanc e (%) Gene name/annotation Putative function Putative link with siderophores Reference (PMID) SAS_RS07345 N 98 Sulphite reductase subunit alpha. Sulphite to sulphide reduction. Iron ion binding. Oxidoreductase. ferrisiderophore reduction. Hydrogen sulphide Sulphate assimilation. Siderophoreiron reductase Between MarR family transcriptional regulator and lysophospholipase. SAS_RS11185 N 43 Methicillin resistance protein FmtB. Cell wall SAS_RS13665 N 55 Hypothetical No conserved domains identified. 4 SAS_RS13620 */ 80 Anaerobic ribonucleosidetriphosphate reductase activating Oxidoreductase. Iron-sulfur cluster binding. Siderophoreiron reductase SAS_RS03320 N 93 Membrane Solute carrier families 5 and 6- like. Transports inorganic ions, sugars, amino acids. SAS_RS12635 N 25 Glycerate kinase. Glycine metabolism. Transport of substrates Ferrichrome siderophore is made of glycine residues
7 Single Evolution Treatment Replicate type Locus tag/ coordinate abundanc e (%) Gene name/annotation Putative function Putative link with siderophores Reference (PMID) 1 SAS_RS06355 N 28 RIP metalloprotease RseP. SAS_RS09685 N 50 2-hydroxyacid dehydrogenase. Coordinating gene transcription during extracytoplasmic stress response. Oxidoreductase Between carbohydrate kinase and histidine ammonialyase. SAS_RS01275 N 35 Ribose transporter RbsU. SAS_RS03290 N 28 Ferrichrome ABC transporter permease. Sugar transport across bacterial membrane. Uptake of ribose. Part of the ABC transporter siderophore import. SAS_RS03845 * 30 Helicase. ATP-dependent RNA or DNA unwinding. SAS_RS04065 S 25 Thermonuclease. Catalyses the hydrolysis of both DNA and RNA Between hypothetical protein and hemebinding SAS_RS05650 N 28 Heme uptake system protein IsdE. ABC transport of heme, siderophores and metal ions Between aspartyl/ glutamyltrna(asn/gln) amidotransferase subunit C and sodium:proline symporter. SAS_RS11115 N 38 Membrane No conserved domains identified. Transport of substrates Heme-binding protein; iron Transport of substrates
8 Treatment Replicate type Locus tag/ coordinate abundanc e (%) Gene name/annotation Putative function Putative link with siderophores Reference (PMID) SAS_RS12235 N 33 DNA-binding response regulator. SAS_RS13800 N 30 Surface anchored 4 No mutations >20% 5 SAS_RS12000 N 23 RpiR family transcriptional regulator. Contains a singal receiver domain, an effector domain and a DNA-binding response regulator. No conserved domains identified. Transcription factor.
GO ID GO term Number of members GO: translation 225 GO: nucleosome 50 GO: calcium ion binding 76 GO: structural
 GO ID GO term Number of members GO:0006412 translation 225 GO:0000786 nucleosome 50 GO:0005509 calcium ion binding 76 GO:0003735 structural constituent of ribosome 170 GO:0019861 flagellum 23 GO:0005840
GO ID GO term Number of members GO:0006412 translation 225 GO:0000786 nucleosome 50 GO:0005509 calcium ion binding 76 GO:0003735 structural constituent of ribosome 170 GO:0019861 flagellum 23 GO:0005840
From gene to protein. Premedical biology
 From gene to protein Premedical biology Central dogma of Biology, Molecular Biology, Genetics transcription replication reverse transcription translation DNA RNA Protein RNA chemically similar to DNA,
From gene to protein Premedical biology Central dogma of Biology, Molecular Biology, Genetics transcription replication reverse transcription translation DNA RNA Protein RNA chemically similar to DNA,
(starvation). Description a. Predicted operon members b. Gene no. a. Relative change in expression (n-fold) mutant vs. wild type.
 1 Table S1. Genes whose expression differ in the phyr mutant 8402 and/or in the ecfg mutant 8404 compared with the wild type when grown to the mid-exponential phase (OD600 0.5-0.7) in rich medium (PSY)
1 Table S1. Genes whose expression differ in the phyr mutant 8402 and/or in the ecfg mutant 8404 compared with the wild type when grown to the mid-exponential phase (OD600 0.5-0.7) in rich medium (PSY)
Strain or plasmid Relevant characteristics Reference
 Table S1. Bacterial strains, and plasmids Strain or plasmid Relevant characteristics Reference M. xanthus strains DK1622 Wild-type M. xanthus 1 AG306 pnbc6::nla6, Kan r 2 AG1151 DK 1622 pkg51::mxan2688
Table S1. Bacterial strains, and plasmids Strain or plasmid Relevant characteristics Reference M. xanthus strains DK1622 Wild-type M. xanthus 1 AG306 pnbc6::nla6, Kan r 2 AG1151 DK 1622 pkg51::mxan2688
Genomic insights into high exopolysaccharide-producing dairy starter. bacterium Streptococcus thermophilus ASCC 1275
 Supplementary information of Genomic insights into high exopolysaccharide-producing dairy starter bacterium Streptococcus thermophilus ASCC 1275 Qinglong Wu, Hein Min Tun, Frederick Chi-Ching Leung, Nagendra
Supplementary information of Genomic insights into high exopolysaccharide-producing dairy starter bacterium Streptococcus thermophilus ASCC 1275 Qinglong Wu, Hein Min Tun, Frederick Chi-Ching Leung, Nagendra
A genome sequence based discriminator for vancomycin intermediate Staphyolococcus aureus Supplementary Methods
 1 Journal of Bacteriology Computational Biology Section 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 Supplementary Information for: A genome sequence based discriminator
1 Journal of Bacteriology Computational Biology Section 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 Supplementary Information for: A genome sequence based discriminator
Table S1. 45 significantly up-regulated P. mirabilis HI4320 genes as determined by SAM during iron-limiting microarray a
 Table S1. 45 significantly up-regulated P. mirabilis HI4320 genes as determined by SAM during iron-limiting microarray a PMI no. Gene Description Log 2 fold-change b PMI0029 exbb biopolymer transport protein
Table S1. 45 significantly up-regulated P. mirabilis HI4320 genes as determined by SAM during iron-limiting microarray a PMI no. Gene Description Log 2 fold-change b PMI0029 exbb biopolymer transport protein
BACTERIAL PHYSIOLOGY SMALL GROUP. Monday, August 25, :00pm. Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D.
 BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
Table 5. Genes unique to G. thermodenitrificans NG80-2 Gene ID Gene name Gene product COG functional category
 Table 5. Genes unique to G. thermodenitrificans NG80-2 GT0030 gt30 Methionine--tRNA ligase/methionyl-trna synthetase COG0143, Translation GT0033 gt33 Unknown GT0106 gt106 Ribosomal protein L3 COG0087,
Table 5. Genes unique to G. thermodenitrificans NG80-2 GT0030 gt30 Methionine--tRNA ligase/methionyl-trna synthetase COG0143, Translation GT0033 gt33 Unknown GT0106 gt106 Ribosomal protein L3 COG0087,
Topic 4 - #14 The Lactose Operon
 Topic 4 - #14 The Lactose Operon The Lactose Operon The lactose operon is an operon which is responsible for the transport and metabolism of the sugar lactose in E. coli. - Lactose is one of many organic
Topic 4 - #14 The Lactose Operon The Lactose Operon The lactose operon is an operon which is responsible for the transport and metabolism of the sugar lactose in E. coli. - Lactose is one of many organic
The body has three primary lines of defense against changes in hydrogen ion concentration in the body fluids.
 ph and Nucleic acids Hydrogen Ion (H+) concentration is precisely regulated. The H+ concentration in the extracellular fluid is maintained at a very low level, averaging 0.00000004Eq/L. normal variations
ph and Nucleic acids Hydrogen Ion (H+) concentration is precisely regulated. The H+ concentration in the extracellular fluid is maintained at a very low level, averaging 0.00000004Eq/L. normal variations
PHOTOSYNTHESIS: A BRIEF STORY!!!!!
 PHOTOSYNTHESIS: A BRIEF STORY!!!!! This is one of the most important biochemical processes in plants and is amongst the most expensive biochemical processes in plant in terms of investment. Photosynthesis
PHOTOSYNTHESIS: A BRIEF STORY!!!!! This is one of the most important biochemical processes in plants and is amongst the most expensive biochemical processes in plant in terms of investment. Photosynthesis
UNIT 5. Protein Synthesis 11/22/16
 UNIT 5 Protein Synthesis IV. Transcription (8.4) A. RNA carries DNA s instruction 1. Francis Crick defined the central dogma of molecular biology a. Replication copies DNA b. Transcription converts DNA
UNIT 5 Protein Synthesis IV. Transcription (8.4) A. RNA carries DNA s instruction 1. Francis Crick defined the central dogma of molecular biology a. Replication copies DNA b. Transcription converts DNA
REGULATION OF GENE EXPRESSION. Bacterial Genetics Lac and Trp Operon
 REGULATION OF GENE EXPRESSION Bacterial Genetics Lac and Trp Operon Levels of Metabolic Control The amount of cellular products can be controlled by regulating: Enzyme activity: alters protein function
REGULATION OF GENE EXPRESSION Bacterial Genetics Lac and Trp Operon Levels of Metabolic Control The amount of cellular products can be controlled by regulating: Enzyme activity: alters protein function
Population transcriptomics uncovers the regulation of gene. expression variation in adaptation to changing environment
 Supplementary information Population transcriptomics uncovers the regulation of gene expression variation in adaptation to changing environment Qin Xu 1, Caiyun Zhu 2,4, Yangyang Fan 1,4, Zhihong Song
Supplementary information Population transcriptomics uncovers the regulation of gene expression variation in adaptation to changing environment Qin Xu 1, Caiyun Zhu 2,4, Yangyang Fan 1,4, Zhihong Song
Transmembrane Domains (TMDs) of ABC transporters
 Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL SUPPLEMENTAL RESULTS Microarray analysis of F. tularensis LVS within hepatocytes: As described in Materials and Methods, the global gene expression of LVS organisms grown in FL83B
SUPPLEMENTAL MATERIAL SUPPLEMENTAL RESULTS Microarray analysis of F. tularensis LVS within hepatocytes: As described in Materials and Methods, the global gene expression of LVS organisms grown in FL83B
Microbiology Helmut Pospiech
 Microbiology 20.03.2018 Helmut Pospiech The control of what gets in Passive transport along a concentration gradient often inefficient Active transport Requires energy consumption and what gets out ABC
Microbiology 20.03.2018 Helmut Pospiech The control of what gets in Passive transport along a concentration gradient often inefficient Active transport Requires energy consumption and what gets out ABC
Chapter 16 Lecture. Concepts Of Genetics. Tenth Edition. Regulation of Gene Expression in Prokaryotes
 Chapter 16 Lecture Concepts Of Genetics Tenth Edition Regulation of Gene Expression in Prokaryotes Chapter Contents 16.1 Prokaryotes Regulate Gene Expression in Response to Environmental Conditions 16.2
Chapter 16 Lecture Concepts Of Genetics Tenth Edition Regulation of Gene Expression in Prokaryotes Chapter Contents 16.1 Prokaryotes Regulate Gene Expression in Response to Environmental Conditions 16.2
RECONSTRUCTING GENE REGULATORY NETWORKS FROM FUNGAL TRANSCRIPTOMIC DATA USING BAYESIAN NETWORK
 RECONSTRUCTING GENE REGULATORY NETWORKS FROM FUNGAL TRANSCRIPTOMIC DATA USING BAYESIAN NETWORK Li Guo Fungal Comparative Genomics Laboratory Department of Biochemistry and Molecular biology University
RECONSTRUCTING GENE REGULATORY NETWORKS FROM FUNGAL TRANSCRIPTOMIC DATA USING BAYESIAN NETWORK Li Guo Fungal Comparative Genomics Laboratory Department of Biochemistry and Molecular biology University
2 4 Chemical Reactions and Enzymes Chemical Reactions
 Chemical Reactions A chemical reaction occurs when chemical bonds are broken and reformed. Rust forms very slowly, while rocket fuel combustion is explosive! The significance of this comparison is that
Chemical Reactions A chemical reaction occurs when chemical bonds are broken and reformed. Rust forms very slowly, while rocket fuel combustion is explosive! The significance of this comparison is that
Midterm Review Guide. Unit 1 : Biochemistry: 1. Give the ph values for an acid and a base. 2. What do buffers do? 3. Define monomer and polymer.
 Midterm Review Guide Name: Unit 1 : Biochemistry: 1. Give the ph values for an acid and a base. 2. What do buffers do? 3. Define monomer and polymer. 4. Fill in the Organic Compounds chart : Elements Monomer
Midterm Review Guide Name: Unit 1 : Biochemistry: 1. Give the ph values for an acid and a base. 2. What do buffers do? 3. Define monomer and polymer. 4. Fill in the Organic Compounds chart : Elements Monomer
Boolean models of gene regulatory networks. Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016
 Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Supplementary Figure 1 The number of differentially expressed genes for uniparental males (green), uniparental females (yellow), biparental males
 Supplementary Figure 1 The number of differentially expressed genes for males (green), females (yellow), males (red), and females (blue) in caring vs. control comparisons in the caring gene set and the
Supplementary Figure 1 The number of differentially expressed genes for males (green), females (yellow), males (red), and females (blue) in caring vs. control comparisons in the caring gene set and the
Chapter 02 The Chemistry of Biology
 Chapter 02 The Chemistry of Biology Multiple Choice Questions 1. Anything that occupies space and has mass is called A. atomic. B. living. C. matter. D. energy. E. space. Learning Outcome: 02.01 Explain
Chapter 02 The Chemistry of Biology Multiple Choice Questions 1. Anything that occupies space and has mass is called A. atomic. B. living. C. matter. D. energy. E. space. Learning Outcome: 02.01 Explain
Lecture 12. Metalloproteins - II
 Lecture 12 Metalloproteins - II Metalloenzymes Metalloproteins with one labile coordination site around the metal centre are known as metalloenzyme. As with all enzymes, the shape of the active site is
Lecture 12 Metalloproteins - II Metalloenzymes Metalloproteins with one labile coordination site around the metal centre are known as metalloenzyme. As with all enzymes, the shape of the active site is
The Prokaryotic World
 The Prokaryotic World A. An overview of prokaryotic life There is no doubt that prokaryotes are everywhere. By everywhere, I mean living in every geographic region, in extremes of environmental conditions,
The Prokaryotic World A. An overview of prokaryotic life There is no doubt that prokaryotes are everywhere. By everywhere, I mean living in every geographic region, in extremes of environmental conditions,
Biology 112 Practice Midterm Questions
 Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Enzyme Catalysis & Biotechnology
 L28-1 Enzyme Catalysis & Biotechnology Bovine Pancreatic RNase A Biochemistry, Life, and all that L28-2 A brief word about biochemistry traditionally, chemical engineers used organic and inorganic chemistry
L28-1 Enzyme Catalysis & Biotechnology Bovine Pancreatic RNase A Biochemistry, Life, and all that L28-2 A brief word about biochemistry traditionally, chemical engineers used organic and inorganic chemistry
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11
 UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
Advanced Higher Biology. Unit 1- Cells and Proteins 2c) Membrane Proteins
 Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Rex-Family Repressor/NADH Complex
 Kasey Royer Michelle Lukosi Rex-Family Repressor/NADH Complex Part A The biological sensing protein that we selected is the Rex-family repressor/nadh complex. We chose this sensor because it is a calcium
Kasey Royer Michelle Lukosi Rex-Family Repressor/NADH Complex Part A The biological sensing protein that we selected is the Rex-family repressor/nadh complex. We chose this sensor because it is a calcium
Reading Assignments. A. Genes and the Synthesis of Polypeptides. Lecture Series 7 From DNA to Protein: Genotype to Phenotype
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
ATP ATP. The energy needs of life. Living economy. Where do we get the energy from? 9/11/2015. Making energy! Organisms are endergonic systems
 Making energy! ATP The energy needs of life rganisms are endergonic systems What do we need energy for? synthesis building biomolecules reproduction movement active transport temperature regulation 2007-2008
Making energy! ATP The energy needs of life rganisms are endergonic systems What do we need energy for? synthesis building biomolecules reproduction movement active transport temperature regulation 2007-2008
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
PROTEIN SYNTHESIS INTRO
 MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
Name Period The Control of Gene Expression in Prokaryotes Notes
 Bacterial DNA contains genes that encode for many different proteins (enzymes) so that many processes have the ability to occur -not all processes are carried out at any one time -what allows expression
Bacterial DNA contains genes that encode for many different proteins (enzymes) so that many processes have the ability to occur -not all processes are carried out at any one time -what allows expression
Cellular Respiration Chapter 5 Notes
 Cellular Respiration Chapter 5 Notes Some Terms to Know Aerobic = WITH oxygen Anaerobic = without oxygen NAD electron carrier = NADH FAD electron carrier = FADH 2 Cellular Respiration a way for cells to
Cellular Respiration Chapter 5 Notes Some Terms to Know Aerobic = WITH oxygen Anaerobic = without oxygen NAD electron carrier = NADH FAD electron carrier = FADH 2 Cellular Respiration a way for cells to
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
Chapter 1. DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d
 Chapter 1 1. Matching Questions DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d 2. Matching Questions : Unbranched polymer that, when folded into its three-dimensional shape,
Chapter 1 1. Matching Questions DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d 2. Matching Questions : Unbranched polymer that, when folded into its three-dimensional shape,
2: CHEMICAL COMPOSITION OF THE BODY
 1 2: CHEMICAL COMPOSITION OF THE BODY CHAPTER OVERVIEW This chapter provides an overview of basic chemical principles that are important to understanding human physiological function and ultimately homeostasis.
1 2: CHEMICAL COMPOSITION OF THE BODY CHAPTER OVERVIEW This chapter provides an overview of basic chemical principles that are important to understanding human physiological function and ultimately homeostasis.
Table S1: Domain based (meta)genome comparison of selected metagenomes of methanotrophs and genomes of selected methanogens*
 ANME-1-s ANME-1-m ANME-2a ANME-2d-h ANME-2d-a AM A M-1 M-2 M-3 H-1 H-2 H-3 H-4 S Table S1: Domain based (meta)genome comparison of selected metagenomes of methanotrophs and genomes of selected methanogens*
ANME-1-s ANME-1-m ANME-2a ANME-2d-h ANME-2d-a AM A M-1 M-2 M-3 H-1 H-2 H-3 H-4 S Table S1: Domain based (meta)genome comparison of selected metagenomes of methanotrophs and genomes of selected methanogens*
Biological Process Term Enrichment
 Biological Process Term Enrichment cellular protein localization cellular macromolecule localization intracellular protein transport intracellular transport generation of precursor metabolites and energy
Biological Process Term Enrichment cellular protein localization cellular macromolecule localization intracellular protein transport intracellular transport generation of precursor metabolites and energy
Curriculum Map. Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1)
 1 Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1) Focus Standards BIO1.LS1.2 Evaluate comparative models of various cell types with a focus on organic molecules
1 Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1) Focus Standards BIO1.LS1.2 Evaluate comparative models of various cell types with a focus on organic molecules
BA, BSc, and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
BCH 400/600 Introductory Biochemistry
 BCH 400/600 Introductory Biochemistry Instructor: David Shintani Office: 311C Fleischmann Ag. Lab: 308 Fleischmann Ag. E-mail: shintani@unr.edu Phone: (775) 784-4631 Before BCH 400 BCH 400 is heavy on
BCH 400/600 Introductory Biochemistry Instructor: David Shintani Office: 311C Fleischmann Ag. Lab: 308 Fleischmann Ag. E-mail: shintani@unr.edu Phone: (775) 784-4631 Before BCH 400 BCH 400 is heavy on
Introduction to Metabolism (Or Energy Management) Chapter 8
 Introduction to Metabolism (Or Energy Management) Chapter 8 Metabolism of the chemical reactions in the organism Building up molecules Breaking down molecules Managing energy and materials Route to end-product
Introduction to Metabolism (Or Energy Management) Chapter 8 Metabolism of the chemical reactions in the organism Building up molecules Breaking down molecules Managing energy and materials Route to end-product
Name: SBI 4U. Gene Expression Quiz. Overall Expectation:
 Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Components of a functional cell. Boundary-membrane Cytoplasm: Cytosol (soluble components) & particulates DNA-information Ribosomes-protein synthesis
 Cell (Outline) - Components of a functional cell - Major Events in the History of Earth: abiotic and biotic phases; anaerobic and aerobic atmosphere - Prokaryotic cells impact on the biosphere - Origin
Cell (Outline) - Components of a functional cell - Major Events in the History of Earth: abiotic and biotic phases; anaerobic and aerobic atmosphere - Prokaryotic cells impact on the biosphere - Origin
Bioinorganic Chemistry
 PRINCIPLES OF Bioinorganic Chemistry Stephen J. Lippard MASSACHUSETTS INSTITUTE OF TECHNOLOGY Jeremy M. Berg JOHNS HOPKINS SCHOOL OF MEDICINE f V University Science Books Mill Valley, California Preface
PRINCIPLES OF Bioinorganic Chemistry Stephen J. Lippard MASSACHUSETTS INSTITUTE OF TECHNOLOGY Jeremy M. Berg JOHNS HOPKINS SCHOOL OF MEDICINE f V University Science Books Mill Valley, California Preface
Advanced Cell Biology. Lecture 6
 Advanced Cell Biology. Lecture 6 Alexey Shipunov Minot State University January 23, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 6 January 23, 2013 1 / 48 Outline Questions and answers Nucleic acids
Advanced Cell Biology. Lecture 6 Alexey Shipunov Minot State University January 23, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 6 January 23, 2013 1 / 48 Outline Questions and answers Nucleic acids
2. Cellular and Molecular Biology
 2. Cellular and Molecular Biology 2.1 Cell Structure 2.2 Transport Across Cell Membranes 2.3 Cellular Metabolism 2.4 DNA Replication 2.5 Cell Division 2.6 Biosynthesis 2.1 Cell Structure What is a cell?
2. Cellular and Molecular Biology 2.1 Cell Structure 2.2 Transport Across Cell Membranes 2.3 Cellular Metabolism 2.4 DNA Replication 2.5 Cell Division 2.6 Biosynthesis 2.1 Cell Structure What is a cell?
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Table S1. 10μM_rfp_B1 2,074, ohr 10μM_rfp_C2 2,074, ohr
 Table S1 Selection without RpoD Insert Start Position Insert End Position Potential Tolerance Gene Candidates 0μM_rfp_A1 2,076,184 -- ohr 0μM_rfp_B1 2,082,864 -- ohr 0μM_rfp_C1 2,076,798 -- ohr 1μM_rfp_A1
Table S1 Selection without RpoD Insert Start Position Insert End Position Potential Tolerance Gene Candidates 0μM_rfp_A1 2,076,184 -- ohr 0μM_rfp_B1 2,082,864 -- ohr 0μM_rfp_C1 2,076,798 -- ohr 1μM_rfp_A1
The geneticist s questions. Deleting yeast genes. Functional genomics. From Wikipedia, the free encyclopedia
 From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
Agrobacterium tumefasciens, the Ti Plasmid, and Crown Gall Tumorigenesis
 Agrobacterium tumefasciens, the Ti Plasmid, and Crown Gall Tumorigenesis BOM-11: 10.9 Plasmids: General Principles (review) p. 274 10.11 Conjugation: Essential Features (review) p. 278 19.21 Agrobacterium
Agrobacterium tumefasciens, the Ti Plasmid, and Crown Gall Tumorigenesis BOM-11: 10.9 Plasmids: General Principles (review) p. 274 10.11 Conjugation: Essential Features (review) p. 278 19.21 Agrobacterium
the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Chemical structure Covalent bond Ionic bond
 Chemical structure the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Covalent bond bond formed by the sharing of valence electrons between atoms Ionic bond
Chemical structure the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Covalent bond bond formed by the sharing of valence electrons between atoms Ionic bond
Metabolism and Enzymes
 Energy Basics Metabolism and Enzymes Chapter 5 Pgs. 77 86 Chapter 8 Pgs. 142 162 Energy is the capacity to cause change, and is required to do work. Very difficult to define quantity. Two types of energy:
Energy Basics Metabolism and Enzymes Chapter 5 Pgs. 77 86 Chapter 8 Pgs. 142 162 Energy is the capacity to cause change, and is required to do work. Very difficult to define quantity. Two types of energy:
AP BIOLOGY BIOCHEMISTRY MULTIPLE CHOICE EXAM (RAVEN CHAPTERS 2, 3)
 Period Date AP BIOLOGY BIOCHEMISTRY MULTIPLE CHOICE EXAM (RAVEN CHAPTERS 2, 3) 1. Which of the following is an example of a hydrogen bond? (90:09) A. The peptide bond between amino acids in a protein B.
Period Date AP BIOLOGY BIOCHEMISTRY MULTIPLE CHOICE EXAM (RAVEN CHAPTERS 2, 3) 1. Which of the following is an example of a hydrogen bond? (90:09) A. The peptide bond between amino acids in a protein B.
Biology I Fall Semester Exam Review 2014
 Biology I Fall Semester Exam Review 2014 Biomolecules and Enzymes (Chapter 2) 8 questions Macromolecules, Biomolecules, Organic Compunds Elements *From the Periodic Table of Elements Subunits Monomers,
Biology I Fall Semester Exam Review 2014 Biomolecules and Enzymes (Chapter 2) 8 questions Macromolecules, Biomolecules, Organic Compunds Elements *From the Periodic Table of Elements Subunits Monomers,
Berg Tymoczko Stryer Biochemistry Sixth Edition Chapter 1:
 Berg Tymoczko Stryer Biochemistry Sixth Edition Chapter 1: Biochemistry: An Evolving Science Tips on note taking... Remember copies of my lectures are available on my webpage If you forget to print them
Berg Tymoczko Stryer Biochemistry Sixth Edition Chapter 1: Biochemistry: An Evolving Science Tips on note taking... Remember copies of my lectures are available on my webpage If you forget to print them
Molecular Biology (9)
 Molecular Biology (9) Translation Mamoun Ahram, PhD Second semester, 2017-2018 1 Resources This lecture Cooper, Ch. 8 (297-319) 2 General information Protein synthesis involves interactions between three
Molecular Biology (9) Translation Mamoun Ahram, PhD Second semester, 2017-2018 1 Resources This lecture Cooper, Ch. 8 (297-319) 2 General information Protein synthesis involves interactions between three
Courtesy of Elsevier. Used with permission.
 Chemistry 5.07 2013 Problem Set 5 Answers Problem 1 Succinate dehydrogenase (SDH) is a heterotetramer enzyme complex that catalyzes the oxidation of succinate to fumarate with concomitant reduction of
Chemistry 5.07 2013 Problem Set 5 Answers Problem 1 Succinate dehydrogenase (SDH) is a heterotetramer enzyme complex that catalyzes the oxidation of succinate to fumarate with concomitant reduction of
Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1
 Name I. Multiple Choice (1 point each) Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1 B 1. Which is possessed by eukaryotes but not by prokaryotes? A. Cell wall B. Distinct nucleus
Name I. Multiple Choice (1 point each) Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1 B 1. Which is possessed by eukaryotes but not by prokaryotes? A. Cell wall B. Distinct nucleus
Number of questions TEK (Learning Target) Biomolecules & Enzymes
 Unit Biomolecules & Enzymes Number of questions TEK (Learning Target) on Exam 8 questions 9A I can compare and contrast the structure and function of biomolecules. 9C I know the role of enzymes and how
Unit Biomolecules & Enzymes Number of questions TEK (Learning Target) on Exam 8 questions 9A I can compare and contrast the structure and function of biomolecules. 9C I know the role of enzymes and how
Membranes 2: Transportation
 Membranes 2: Transportation Steven E. Massey, Ph.D. Associate Professor Bioinformatics Department of Biology University of Puerto Rico Río Piedras Office & Lab: NCN#343B Tel: 787-764-0000 ext. 7798 E-mail:
Membranes 2: Transportation Steven E. Massey, Ph.D. Associate Professor Bioinformatics Department of Biology University of Puerto Rico Río Piedras Office & Lab: NCN#343B Tel: 787-764-0000 ext. 7798 E-mail:
2: CHEMICAL COMPOSITION OF THE BODY
 1 2: CHEMICAL COMPOSITION OF THE BODY Although most students of human physiology have had at least some chemistry, this chapter serves very well as a review and as a glossary of chemical terms. In particular,
1 2: CHEMICAL COMPOSITION OF THE BODY Although most students of human physiology have had at least some chemistry, this chapter serves very well as a review and as a glossary of chemical terms. In particular,
Basic Structure of a Cell
 Basic Structure of a Cell 1 Review Facts About Living Things copyright cmassengale 2 What is life? Alive? Alive? What is life? Thus life defies a one sentence answer/definition Instead, life is recognized
Basic Structure of a Cell 1 Review Facts About Living Things copyright cmassengale 2 What is life? Alive? Alive? What is life? Thus life defies a one sentence answer/definition Instead, life is recognized
Cell (Learning Objectives)
 Cell (Learning Objectives) 1. Understand & describe the basic components necessary for a functional cell. 2. Review the order of appearance of cells on earth and explain the endosymbiotic theory. 3. Compare
Cell (Learning Objectives) 1. Understand & describe the basic components necessary for a functional cell. 2. Review the order of appearance of cells on earth and explain the endosymbiotic theory. 3. Compare
GENERAL BIOLOGY LABORATORY EXERCISE Amino Acid Sequence Analysis of Cytochrome C in Bacteria and Eukarya Using Bioinformatics
 GENERAL BIOLOGY LABORATORY EXERCISE Amino Acid Sequence Analysis of Cytochrome C in Bacteria and Eukarya Using Bioinformatics INTRODUCTION: All life forms undergo metabolic processes to obtain energy.
GENERAL BIOLOGY LABORATORY EXERCISE Amino Acid Sequence Analysis of Cytochrome C in Bacteria and Eukarya Using Bioinformatics INTRODUCTION: All life forms undergo metabolic processes to obtain energy.
Chapter 02 Testbank. 1. Anything that occupies space and has mass is called. A. an electron. B. living. C. matter. D. energy. E. space.
 Chapter 02 Testbank Student: 1. Anything that occupies space and has mass is called A. an electron. B. living. C. matter. D. energy. E. space. 2. The electrons of an atom are A. always equal to the number
Chapter 02 Testbank Student: 1. Anything that occupies space and has mass is called A. an electron. B. living. C. matter. D. energy. E. space. 2. The electrons of an atom are A. always equal to the number
REVIEW 1: BIOCHEMISTRY UNIT. A. Top 10 If you learned anything from this unit, you should have learned:
 Period Date REVIEW 1: BIOCHEMISTRY UNIT A. Top 10 If you learned anything from this unit, you should have learned: 1. All living matter made up of CHONPS 2. Bonds a. covalent bonds are strong b. hydrogen
Period Date REVIEW 1: BIOCHEMISTRY UNIT A. Top 10 If you learned anything from this unit, you should have learned: 1. All living matter made up of CHONPS 2. Bonds a. covalent bonds are strong b. hydrogen
S1 Gene ontology (GO) analysis of the network alignment results
 1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus
 Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus Primary Source for figures and content: Tortora, G.J. Microbiology
Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus Primary Source for figures and content: Tortora, G.J. Microbiology
Chapter 002 The Chemistry of Biology
 Chapter 002 The Chemistry of Biology Multiple Choice Questions 1. Anything that occupies space and has mass is called A. Atomic B. Living C. Matter D. Energy E. Space 2. The electrons of an atom are A.
Chapter 002 The Chemistry of Biology Multiple Choice Questions 1. Anything that occupies space and has mass is called A. Atomic B. Living C. Matter D. Energy E. Space 2. The electrons of an atom are A.
Chapter 02 Testbank. 1. Anything that occupies space and has mass is called. A. an electron. B. living. C. matter. D. energy. E. space.
 Chapter 02 Testbank Student: 1. Anything that occupies space and has mass is called A. an electron. B. living. C. matter. D. energy. E. space. 2. The electrons of an atom are A. always equal to the number
Chapter 02 Testbank Student: 1. Anything that occupies space and has mass is called A. an electron. B. living. C. matter. D. energy. E. space. 2. The electrons of an atom are A. always equal to the number
The biomolecules of terrestrial life
 Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Patrick: An Introduction to Medicinal Chemistry 5e Chapter 04
 01) Which of the following statements is not true about receptors? a. Most receptors are proteins situated inside the cell. b. Receptors contain a hollow or cleft on their surface which is known as a binding
01) Which of the following statements is not true about receptors? a. Most receptors are proteins situated inside the cell. b. Receptors contain a hollow or cleft on their surface which is known as a binding
SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question.
 ch 2 chemical basis of life Name SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question. Fill in the blank or provide a short answer: 1) When a change in matter
ch 2 chemical basis of life Name SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question. Fill in the blank or provide a short answer: 1) When a change in matter
Nature Genetics: doi: /ng Supplementary Figure 1. Icm/Dot secretion system region I in 41 Legionella species.
 Supplementary Figure 1 Icm/Dot secretion system region I in 41 Legionella species. Homologs of the effector-coding gene lega15 (orange) were found within Icm/Dot region I in 13 Legionella species. In four
Supplementary Figure 1 Icm/Dot secretion system region I in 41 Legionella species. Homologs of the effector-coding gene lega15 (orange) were found within Icm/Dot region I in 13 Legionella species. In four
DNA Technology, Bacteria, Virus and Meiosis Test REVIEW
 Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
What is an enzyme? Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics. Margaret A. Daugherty Fall General Properties
 Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics Margaret A. Daugherty Fall 2003 ENZYMES: Why, what, when, where, how? All but the who! What: proteins that exert kinetic control over
Lecture 12: Enzymes & Kinetics I Introduction to Enzymes and Kinetics Margaret A. Daugherty Fall 2003 ENZYMES: Why, what, when, where, how? All but the who! What: proteins that exert kinetic control over
Lecture 3: A basic statistical concept
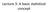 Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
BACTERIA AND ARCHAEA 10/15/2012
 BACTERIA AND ARCHAEA Chapter 27 KEY CONCEPTS: Structural and functional adaptations contribute to prokaryotic success Rapid reproduction, mutation, and genetic recombination promote genetic diversity in
BACTERIA AND ARCHAEA Chapter 27 KEY CONCEPTS: Structural and functional adaptations contribute to prokaryotic success Rapid reproduction, mutation, and genetic recombination promote genetic diversity in
Human Biology, 7e (Johnson) Chapter 2 The Chemistry of Living Things. 2.1 Multiple Choice Questions
 Human Biology, 7e (Johnson) Chapter 2 The Chemistry of Living Things 2.1 Multiple Choice Questions 1) Which one of the following characteristics applies to both living organisms and nonliving things? A)
Human Biology, 7e (Johnson) Chapter 2 The Chemistry of Living Things 2.1 Multiple Choice Questions 1) Which one of the following characteristics applies to both living organisms and nonliving things? A)
Ch 7: Cell Structure and Functions. AP Biology
 Ch 7: Cell Structure and Functions AP Biology The Cell Theory 1. All living things are made of cells. 2. New cells come from existing cells. 3. Cells are the basic units of structure and function of living
Ch 7: Cell Structure and Functions AP Biology The Cell Theory 1. All living things are made of cells. 2. New cells come from existing cells. 3. Cells are the basic units of structure and function of living
Old FINAL EXAM BIO409/509 NAME. Please number your answers and write them on the attached, lined paper.
 Old FINAL EXAM BIO409/509 NAME Please number your answers and write them on the attached, lined paper. Gene expression can be regulated at several steps. Describe one example for each of the following:
Old FINAL EXAM BIO409/509 NAME Please number your answers and write them on the attached, lined paper. Gene expression can be regulated at several steps. Describe one example for each of the following:
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Chapter 12. Genes: Expression and Regulation
 Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
Translational Initiation
 Translational Initiation Lecture Outline 1. Process of Initiation. Alternative mechanisms of Initiation 3. Key Experiments on Initiation 4. Regulation of Initiation Translation is a process with three
Translational Initiation Lecture Outline 1. Process of Initiation. Alternative mechanisms of Initiation 3. Key Experiments on Initiation 4. Regulation of Initiation Translation is a process with three
Electrochemical Potential and the Thermodynamic Basis of Solute Transport Mechanisms
 Electrochemical Potential and the Thermodynamic Basis of Solute Transport Mechanisms A. Electrochemical Potential The electrochemical potential arising from the distribution of a solute A across a membrane
Electrochemical Potential and the Thermodynamic Basis of Solute Transport Mechanisms A. Electrochemical Potential The electrochemical potential arising from the distribution of a solute A across a membrane
3.B.1 Gene Regulation. Gene regulation results in differential gene expression, leading to cell specialization.
 3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Metabolism and enzymes
 Metabolism and enzymes 4-11-16 What is a chemical reaction? A chemical reaction is a process that forms or breaks the chemical bonds that hold atoms together Chemical reactions convert one set of chemical
Metabolism and enzymes 4-11-16 What is a chemical reaction? A chemical reaction is a process that forms or breaks the chemical bonds that hold atoms together Chemical reactions convert one set of chemical
What is the central dogma of biology?
 Bellringer What is the central dogma of biology? A. RNA DNA Protein B. DNA Protein Gene C. DNA Gene RNA D. DNA RNA Protein Review of DNA processes Replication (7.1) Transcription(7.2) Translation(7.3)
Bellringer What is the central dogma of biology? A. RNA DNA Protein B. DNA Protein Gene C. DNA Gene RNA D. DNA RNA Protein Review of DNA processes Replication (7.1) Transcription(7.2) Translation(7.3)
(Lys), resulting in translation of a polypeptide without the Lys amino acid. resulting in translation of a polypeptide without the Lys amino acid.
 1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
Chemistry Basics. Matter anything that occupies space and has mass Energy the ability to do work. Chemical Electrical Mechanical Radiant. Slide 2.
 Chemistry Basics Matter anything that occupies space and has mass Energy the ability to do work Chemical Electrical Mechanical Radiant Slide 2.1 Composition of Matter Elements Fundamental units of matter
Chemistry Basics Matter anything that occupies space and has mass Energy the ability to do work Chemical Electrical Mechanical Radiant Slide 2.1 Composition of Matter Elements Fundamental units of matter
number Done by Corrected by Doctor Nafeth Abu Tarboush
 number 8 Done by Ali Yaghi Corrected by Mamoon Mohamad Alqtamin Doctor Nafeth Abu Tarboush 0 P a g e Oxidative phosphorylation Oxidative phosphorylation has 3 major aspects: 1. It involves flow of electrons
number 8 Done by Ali Yaghi Corrected by Mamoon Mohamad Alqtamin Doctor Nafeth Abu Tarboush 0 P a g e Oxidative phosphorylation Oxidative phosphorylation has 3 major aspects: 1. It involves flow of electrons
