Mechanisms of evolution in experiment and theory. Christopher Knight
|
|
|
- Margaret Parks
- 5 years ago
- Views:
Transcription
1 Mechanisms of evolution in experiment and theory Christopher Knight
2 Evolution and molecular biology Nothing in Biology makes sense except in the light of evolution (Dobzhansy 1964 in Biology, molecular and organismic ) Explicit discrete genotypes Only some aligned Long-term evolutionary hypotheses Phylogenetic scale low-level phenotype only e.g. amino acid properties Wright 1932 Arbitrary, continuous genetic axes Scale un-defined Finer scale? high-level phenotype inclusive fitness
3 Evolution and molecular biology Global fitness landscape from genotype? evolution of protein physical properties Focus in on an individual mutational step Adaptive evolution Put mutational steps together In vitro In vivo Focus on feedback between model and experiment
4 Molecular weight (kda) Evolution and molecular biology Global fitness landscape from genotype? evolution of protein physical properties Mass, iso-electric point Predictable from raw sequence Consider across genome Predictable from raw sequence pi pi Knight, C.G. et al. (2004) PNAS
5 Predicted proteomes Global fitness landscape from genotype? evolution of protein physical properties Mass, iso-electric point Predictable from raw sequence Can we predict fitness from predicted proteome distribution? Mycobacterium genetalium Halobacterium sp.nrc1
6 Predicted proteomes Can we predict fitness from predicted proteome distribution? Measure fitness as ability to grow in different environments Gram negative Gram positive Do organisms with similar growth have similar proteomes? Not just because closely related g proteobacteria Pseudomonas 3 4 a proteobacteria 2 1 Bacillus subtilis Sulfolobus solfataricus Sulfolobus tokodaii Knight, C.G. et al. (2004) PNAS
7 Contrasts in Biolog profile Predicted proteomes Can we predict fitness from predicted proteome distribution? Measure fitness as ability to grow in different environments Gram negative Gram positive Do organisms with similar growth have similar proteomes? Yes BUT need better methods Small sample Complexity of omics unclear Hadjipantelis, P.Z. et al. (2013). J R Soc Interface g proteobacteria Pseudomonas 3 4 a proteobacteria Bacillus subtilis Contrasts in proteome distribution Knight, C.G. et al. (2004) PNAS
8 Predicted proteomes Can we predict fitness from predicted proteome distribution? Focus in on the individual, adaptive mutational step Yes BUT need better methods Small sample Complexity of omics unclear Hadjipantelis, P.Z. et al. (2013). J R Soc Interface
9 Adaptive evolution Focus in on the individual, adaptive mutational step
10 Adaptive evolution Can we predict molecular change from adaptive landscape? Smooth SM Wrinkly Spreader (LSWS) Rainey and Travisano (1998) Nature 394:69-72
11 Adaptive evolution Can we predict molecular change from adaptive landscape? + P - P P P SM LSWS Bantinaki, E. et al. (2007) Genetics Yes BUT (Prote) omics complex (pleiotropy) CheV Catechol 1,2, dioxygenase SM LSWS Expectation if predictable Reality for most proteins Andrew Spiers Knight, C.G. et al. (2006). Nat Genet,
12 Adaptive evolution Can we predict molecular change from adaptive landscape? P P P P SM LSWS Bantinaki, E. et al. (2007) Yes BUT (Prote) omics complex (pleiotropy) Time-scale matters Phylogenetic scale Experimental scale Andrew Spiers Farrell and Knight, unpublished
13 Adaptive evolution Can we predict molecular change from adaptive landscape? Yes BUT (Prote) omics complex (pleiotropy) Time-scale matters
14 Evolution and molecular biology Can go both one way or the other in vivo BUT omics complex (e.g. pleiotropy) Evolutionary scale matters Can go both ways in vitro Simplified system Works on average Wright 1932 Generalise over Genotype-Phenotype map Think carefully about rate of movement mutation rate Fomalise and make predictions for in vivo experiments?
15 Can go both one way or the other in vivo BUT omics complex (e.g. pleiotropy) Evolutionary scale matters Can go both ways in vitro Simplified system Works on average Wright 1932 Generalise over Genotype-Phenotype map Think carefully about rate of movement mutation rate Fomalise and make predictions for in vivo experiments?
16 Wright 1932 Different mutation rates ideal at different points in the landscape Generalise over Genotype-Phenotype map Think carefully about rate of movement mutation rate Fomalise and make predictions for in vivo experiments?
17 % Mutation Distance from peak fitness Can derive various optimal functions 4 All monotonic and decreasing with fitness 2 Belavkin et al.(2015). In review Theory Simulation Different mutation rates ideal at different distances from peak Mutation rate Generalise over Genotype-Phenotype map Think carefully about rate of movement mutation rate Distance from Optimum Fitness
18 % Mutation Distance from peak fitness 4 2 Belavkin et al.(2015). In review Theory Simulation Different mutation rates at different distances from peak? Mutation rate Experiment: Escherichia coli K12 give different fitnesses (numbers of doublings) via different amounts of sugar Assay mutation rate Distance from Optimum Fitness
19 Mutation rate x 10^ Mutation Rate versus Absolute Fitness Consider a subset of ~80 mutations in rpob gene, give antibiotic resistance (Rif R ) 10-9 Mutations Generation Absolute fitness, Wabs w abs (generations/day) Different mutation rates ideal at different points in the landscape Experiment: Escherichia coli K12 give different fitnesses (numbers of doublings) via different amounts of sugar Assay mutation rate
20 % Mutation Mutation rate x 10^ Mutation Rate versus Absolute Fitness Consider a subset of ~80 mutations in rpob gene, give antibiotic resistance (Rif R ) 10-9 Mutations Generation Absolute fitness, Wabs w abs (generations/day) Different mutation rates at different points in the landscape? Yes, Mutation Rate Plasticity (MRP), consistent with theory BUT How is fitness sensed/ transduced? Just one organism Theory Simulation Distance from Optimum Fitness
21 % Mutation Mutation_rate_per10.9 Mutation rate x 10^ Mutations Generation Theory e+08 1e+09 density_per_ml_culture Selective_agent Nal30 Rif50 Population density (10 8 c.f.u. ml -1 ) Simulation Nal R in gyra gene 10-9 Mutations Generation -1 Mutation Rate versus Absolute Fitness Absolute fitness, Wabs w abs (generations/day) Different mutation rates at different points in the landscape? Yes, Mutation Rate Plasticity (MRP), consistent with theory BUT How is fitness sensed/ transduced? Just one organism Statistical association with final population density Distance from Optimum Fitness
22 Mutation rate per cell (10-9 mutations/generation) Mutation rate x 10^ Density dependence in E. coli suggests Quorum Sensing (QS) Does density-dependent mutation rate plasticity (DD-MRP) require luxs QS gene? Test effect of luxs Grows very similarly to WT Effect of LuxS on mutation rate plasticity wild-type quorum-sensing mutant ΔluxS Xavier K B, and Bassler B L J. Bacteriol. 2005;187: e e e e e+08 Population density, cells/ml D (10 8 c.f.u./ml) 4
23 Density dependence in E. coli suggests Quorum Sensing (QS) Does density-dependent mutation rate plasticity (DD-MRP) require luxs QS gene? Yes, needs luxs AND cell-cell interactions But exactly what is the signal? Test effect of luxs Grows very similarly to WT 10-9 Mutations Generation -1 Co-cultures of wild-type and ΔluxS Measure mutation rate in wild-type Population density of signal producing cells Overall population density (10 8 c.f.u./ml) (10 8 c.f.u./ml) Krašovec Rok et al. Nature Communications, (2014).
24 Complement with synthetic AI-2 LuxS Complement with aspartate Yes Activated methyl cycle activated methyl cycle Halliday et al. Anal Biochem., (2010).
25 Mutation rate x 10^ luxs-dependent MRP is mediated by the activated methyl cycle consistent with AI-3 Effect of Aspartate on LuxS mutation rate plasticity 10-9 Mutations Generation -1 ΔluxS - Asp ΔluxS + Asp Krašovec, R. et al. (2014) Nat Commun, Population density cells/ml(10 8 c.f.u./ml) 1.5e e e e e+
26 Mutation_rate_per10.9 Distance from Optimum Fitness Mutation rate x 10^ % Mutation Theory and simulation predict inverse fitness-mutation rate relationship likely to be advantageous Density-Dependent Mutation Rate Plasticity exists in E. coli strains: Higher density, lower mutation Select N R 0.1 Consistent, but not identical, with theory 1e+08 1e+09 density_per_ml_culture Effect of Aspartate on LuxS mutation rate plasticity Depends upon role of luxs in activated methyl cycle 1.5e e e e e cells/ml Signal unknown (AI-3) Downstream mechanism? Truly adaptive? Just E. coli? Collate published mutation rates Estimated by fluctuation test
27 Mutations Generation Population density (10 8 c.f.u. ml -1 ) Richards, Krašovec and Knight, unpublished Just E. coli? Hypothesis: widespread densitydependent MRP? Collate published mutation rates Estimated by fluctuation test
28 10-9 Mutations Generation Baker s yeast Saccharomyces cerevisiae Population density (10 8 c.f.u. ml -1 ) Richards, Krašovec and Knight, unpublished Just E. coli? Hypothesis: widespread densitydependent MRP? Collate published mutation rates Estimated by fluctuation test
29 10-9 Mutations Generation Baker s yeast Saccharomyces cerevisiae Universal DD-MRP? Population density (10 8 c.f.u. ml -1 ) Krašovec and Knight, unpublished Yes even to eukaryotes Just E. coli? No Hypothesis: widespread densitydependent MRP? Collate published mutation rates Estimated by fluctuation test
30 10-9 Mutations Generation Pseudomonas aeruginosa Gamma proteobacterium (like E. coli) No luxs Universal DD-MRP? Population density (10 8 c.f.u. ml -1 ) Richards, Krašovec and Knight, unpublished Just E. coli? Hypothesis: widespread densitydependent MRP? Collate published mutation rates Estimated by fluctuation test
31 10-9 Mutations Generation Population density (10 8 c.f.u. ml -1 ) Pseudomonas aeruginosa Gamma proteobacterium (like E. coli) No luxs Universal DD-MRP? No varies with luxs Yes even to eukaryotes Just E. coli? No Hypothesis: widespread densitydependent MRP? Collate published mutation rates Estimated by fluctuation test
32 Mutation_rate_per % Mutation Conclusions Moving between levels of molecules and evolutionary landscapes Can move either way BUT caveats about evolutionary scale and genotypephenotype mapping Going both ways: mutation rate plasticity (MRP) Modify landscape theory for molecules Find processes consistent with theory Find molecules associated with process (luxs) Do molecules perform function for which theory created? Wide-spread density-dependent MRP e+08 1e+09 density_per_ml_culture Selective_agent Krašovec, R. et al. (2014) Nat Commun, Nal30 Rif Distance from Optimum Fitness
33 Rok Krašovec Roman Belavkin John Aston Alastair Channon Liz Aston Bharat Rash Mani Kadirvel Sarah Forbes Acknowledgements Paul Rainey Nicole Zitzmann, Holger Hebestreit Sripardi Prabhakar Chris Brock Andrew Spiers Rees Kassen Rich Lenski Karina Xavier Andrew McBain Dan Smith Casey Bergman Mike Brockhurst Craig Maclean Chris Simms
Evolving Antibiotic Resistance in Bacteria by Controlling Mutation Rate
 Evolving Antibiotic Resistance in Bacteria by Controlling Mutation Rate Roman V. Belavkin 1 1 School of Science and Technology Middlesex University, London NW4 4BT, UK December 11, 2015, Tokyo University
Evolving Antibiotic Resistance in Bacteria by Controlling Mutation Rate Roman V. Belavkin 1 1 School of Science and Technology Middlesex University, London NW4 4BT, UK December 11, 2015, Tokyo University
Rise and Fall of Mutator Bacteria
 Rise and Fall of Mutator Bacteria The Evolution of Mutation Rates in Bacteria Yoav Ram Hadany Evolutionary Theory Lab 29 May 2011 Bacteria Bacteria are unicellular, haploid and asexual Reproduce by binary
Rise and Fall of Mutator Bacteria The Evolution of Mutation Rates in Bacteria Yoav Ram Hadany Evolutionary Theory Lab 29 May 2011 Bacteria Bacteria are unicellular, haploid and asexual Reproduce by binary
Quorum sensing in plantassociated. S. Brook Peterson Parsek Lab UW Microbiology
 Quorum sensing in plantassociated bacteria S. Brook Peterson Parsek Lab UW Microbiology Outline What is quorum sensing? QS in plant associated bacteria What traits are regulated by QS? What benefits does
Quorum sensing in plantassociated bacteria S. Brook Peterson Parsek Lab UW Microbiology Outline What is quorum sensing? QS in plant associated bacteria What traits are regulated by QS? What benefits does
2 Genome evolution: gene fusion versus gene fission
 2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
Using Evolutionary Approaches To Study Biological Pathways. Pathways Have Evolved
 Pathways Have Evolved Using Evolutionary Approaches To Study Biological Pathways Orkun S. Soyer The Microsoft Research - University of Trento Centre for Computational and Systems Biology Protein-protein
Pathways Have Evolved Using Evolutionary Approaches To Study Biological Pathways Orkun S. Soyer The Microsoft Research - University of Trento Centre for Computational and Systems Biology Protein-protein
Introduction to Industrial Biotechnology
 Introduction to Industrial Biotechnology Lecture 2 - Getting things in and out mechanisms of solute transport across membranes and their application to IBBE Learning outcomes Think about the transporter
Introduction to Industrial Biotechnology Lecture 2 - Getting things in and out mechanisms of solute transport across membranes and their application to IBBE Learning outcomes Think about the transporter
Stochastic simulations
 Stochastic simulations Application to molecular networks Literature overview Noise in genetic networks Origins How to measure and distinguish between the two types of noise (intrinsic vs extrinsic)? What
Stochastic simulations Application to molecular networks Literature overview Noise in genetic networks Origins How to measure and distinguish between the two types of noise (intrinsic vs extrinsic)? What
Monotonicity of fitness landscapes and mutation rate control
 J. Math. Biol. (2016) 73:1491 1524 DOI 10.1007/s00285-016-0995-3 Mathematical Biology Monotonicity of fitness landscapes and mutation rate control Roman V. Belavkin 1 Alastair Channon 2 Elizabeth Aston
J. Math. Biol. (2016) 73:1491 1524 DOI 10.1007/s00285-016-0995-3 Mathematical Biology Monotonicity of fitness landscapes and mutation rate control Roman V. Belavkin 1 Alastair Channon 2 Elizabeth Aston
Can evolution paths be explained by chance alone? arxiv: v1 [math.pr] 12 Oct The model
![Can evolution paths be explained by chance alone? arxiv: v1 [math.pr] 12 Oct The model Can evolution paths be explained by chance alone? arxiv: v1 [math.pr] 12 Oct The model](/thumbs/89/98225676.jpg) Can evolution paths be explained by chance alone? Rinaldo B. Schinazi Department of Mathematics University of Colorado at Colorado Springs rinaldo.schinazi@uccs.edu arxiv:1810.05705v1 [math.pr] 12 Oct
Can evolution paths be explained by chance alone? Rinaldo B. Schinazi Department of Mathematics University of Colorado at Colorado Springs rinaldo.schinazi@uccs.edu arxiv:1810.05705v1 [math.pr] 12 Oct
Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling
 Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
Copyright Mark Brandt, Ph.D A third method, cryogenic electron microscopy has seen increasing use over the past few years.
 Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Basic modeling approaches for biological systems. Mahesh Bule
 Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Chapter 19. Microbial Taxonomy
 Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Analysis of Escherichia coli amino acid transporters
 Ph.D thesis Analysis of Escherichia coli amino acid transporters Presented by Attila Szvetnik Supervisor: Dr. Miklós Kálmán Biology Ph.D School University of Szeged Bay Zoltán Foundation for Applied Research
Ph.D thesis Analysis of Escherichia coli amino acid transporters Presented by Attila Szvetnik Supervisor: Dr. Miklós Kálmán Biology Ph.D School University of Szeged Bay Zoltán Foundation for Applied Research
Introduction to Bioinformatics. Shifra Ben-Dor Irit Orr
 Introduction to Bioinformatics Shifra Ben-Dor Irit Orr Lecture Outline: Technical Course Items Introduction to Bioinformatics Introduction to Databases This week and next week What is bioinformatics? A
Introduction to Bioinformatics Shifra Ben-Dor Irit Orr Lecture Outline: Technical Course Items Introduction to Bioinformatics Introduction to Databases This week and next week What is bioinformatics? A
Optimality, Robustness, and Noise in Metabolic Network Control
 Optimality, Robustness, and Noise in Metabolic Network Control Muxing Chen Gal Chechik Daphne Koller Department of Computer Science Stanford University May 18, 2007 Abstract The existence of noise, or
Optimality, Robustness, and Noise in Metabolic Network Control Muxing Chen Gal Chechik Daphne Koller Department of Computer Science Stanford University May 18, 2007 Abstract The existence of noise, or
Fitness constraints on horizontal gene transfer
 Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Optimal Mutation Rate Control under Selection in Hamming Spaces
 DOI: http://dx.doi.org/10.7551/978-0-262-33027-5-ch113 Optimal Mutation Rate Control under Selection in Hamming Spaces Elizabeth Aston 1, Alastair Channon 1, Roman V. Belavkin 2, Rok Krašovec 3 and Christopher
DOI: http://dx.doi.org/10.7551/978-0-262-33027-5-ch113 Optimal Mutation Rate Control under Selection in Hamming Spaces Elizabeth Aston 1, Alastair Channon 1, Roman V. Belavkin 2, Rok Krašovec 3 and Christopher
Stochastic simulations!
 Stochastic simulations! Application to biomolecular networks! Literature overview Noise in genetic networks! Origins! How to measure the noise and distinguish between the two sources of noise (intrinsic
Stochastic simulations! Application to biomolecular networks! Literature overview Noise in genetic networks! Origins! How to measure the noise and distinguish between the two sources of noise (intrinsic
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
ANALYSIS OF MICROBIAL COMPETITION
 ANALYSIS OF MICROBIAL COMPETITION Eric Pomper Microbiology 9 Pittsburgh Central Catholic High School Grade 9 Introduction Escherichia coli (E. coli) and Saccharomyces cerevisiae (Yeast) were grown together
ANALYSIS OF MICROBIAL COMPETITION Eric Pomper Microbiology 9 Pittsburgh Central Catholic High School Grade 9 Introduction Escherichia coli (E. coli) and Saccharomyces cerevisiae (Yeast) were grown together
Регуляция гомеостаза ионов меди у Halobacterium salinarum: компьютерное моделирование и экспериментальный анализ
 Регуляция гомеостаза ионов меди у Halobacterium salinarum: компьютерное моделирование и экспериментальный анализ Regulation of Cu homeostasis in H. salinarum: computational modeling and experimental analysis
Регуляция гомеостаза ионов меди у Halobacterium salinarum: компьютерное моделирование и экспериментальный анализ Regulation of Cu homeostasis in H. salinarum: computational modeling and experimental analysis
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Microbial Taxonomy and the Evolution of Diversity
 19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
Bi 1x Spring 2014: LacI Titration
 Bi 1x Spring 2014: LacI Titration 1 Overview In this experiment, you will measure the effect of various mutated LacI repressor ribosome binding sites in an E. coli cell by measuring the expression of a
Bi 1x Spring 2014: LacI Titration 1 Overview In this experiment, you will measure the effect of various mutated LacI repressor ribosome binding sites in an E. coli cell by measuring the expression of a
The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome
 Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Why EvoSysBio? Combine the rigor from two powerful quantitative modeling traditions: Molecular Systems Biology. Evolutionary Biology
 Why EvoSysBio? Combine the rigor from two powerful quantitative modeling traditions: Molecular Systems Biology rigorous models of molecules... in organisms Modeling Evolutionary Biology rigorous models
Why EvoSysBio? Combine the rigor from two powerful quantitative modeling traditions: Molecular Systems Biology rigorous models of molecules... in organisms Modeling Evolutionary Biology rigorous models
Systems Biology: A Personal View IX. Landscapes. Sitabhra Sinha IMSc Chennai
 Systems Biology: A Personal View IX. Landscapes Sitabhra Sinha IMSc Chennai Fitness Landscapes Sewall Wright pioneered the description of how genotype or phenotypic fitness are related in terms of a fitness
Systems Biology: A Personal View IX. Landscapes Sitabhra Sinha IMSc Chennai Fitness Landscapes Sewall Wright pioneered the description of how genotype or phenotypic fitness are related in terms of a fitness
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Stepping stones towards a new electronic prokaryotic taxonomy. The ultimate goal in taxonomy. Pragmatic towards diagnostics
 Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
Evolution AP Biology
 Darwin s Theory of Evolution How do biologists use evolutionary theory to develop better flu vaccines? Theory: Evolutionary Theory: Why do we need to understand the Theory of Evolution? Charles Darwin:
Darwin s Theory of Evolution How do biologists use evolutionary theory to develop better flu vaccines? Theory: Evolutionary Theory: Why do we need to understand the Theory of Evolution? Charles Darwin:
Molecular Evolution & the Origin of Variation
 Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation
 Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
Molecular Evolution & the Origin of Variation What Is Molecular Evolution? Molecular evolution differs from phenotypic evolution in that mutations and genetic drift are much more important determinants
The Effect of Static Magnetic Field on E. coli, S. aureus and B. subtilis Viability
 The Effect of Static Magnetic Field on E. coli, S. aureus and B. subtilis Viability Khaled A. Al-Khaza'leh 1* Abdullah T. Al-fawwaz 2 1. Department of Physics, Al-albayt University, PO box 130040, Mafraq,
The Effect of Static Magnetic Field on E. coli, S. aureus and B. subtilis Viability Khaled A. Al-Khaza'leh 1* Abdullah T. Al-fawwaz 2 1. Department of Physics, Al-albayt University, PO box 130040, Mafraq,
the noisy gene Biology of the Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB)
 Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Divergent evolution during an experimental adaptive radiation
 Received 8 November 2002 Accepted 21 March 2003 Published online 18 June 2003 Divergent evolution during an experimental adaptive radiation R. Craig MacLean 1* and Graham Bell 1,2 1 Department of Biology,
Received 8 November 2002 Accepted 21 March 2003 Published online 18 June 2003 Divergent evolution during an experimental adaptive radiation R. Craig MacLean 1* and Graham Bell 1,2 1 Department of Biology,
Microbial Diversity and Assessment (II) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Adaptive divergence in experimental populations of Pseudomonas fluorescens. South Parks Road, Oxford OX1 3RB, United Kingdom
 Genetics: Published Articles Ahead of Print, published on March 4, 2007 as 10.1534/genetics.106.069906 Adaptive divergence in experimental populations of Pseudomonas fluorescens. III. Mutational origins
Genetics: Published Articles Ahead of Print, published on March 4, 2007 as 10.1534/genetics.106.069906 Adaptive divergence in experimental populations of Pseudomonas fluorescens. III. Mutational origins
Report. Cooperation Peaks at Intermediate Disturbance. results suggest that they may play a major role in the evolution of social traits in microbes.
 Current Biology 17, 761 765, May 1, 2007 ª2007 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2007.02.057 Cooperation Peaks at Intermediate Disturbance Report Michael A. Brockhurst, 1, * Angus Buckling,
Current Biology 17, 761 765, May 1, 2007 ª2007 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2007.02.057 Cooperation Peaks at Intermediate Disturbance Report Michael A. Brockhurst, 1, * Angus Buckling,
m1 m2 m3 m4 m5 m6 m7 m8 wt m m m m m m m m8 - + wt +
 otherwise, you couldn't grow them!) You perform pairwise infections with each of your mutant bacteriophage strains and get the following results: (+) = pair of phages lysed host cells, (-) = pair of phages
otherwise, you couldn't grow them!) You perform pairwise infections with each of your mutant bacteriophage strains and get the following results: (+) = pair of phages lysed host cells, (-) = pair of phages
Using ADE Mutants to Study Respiration and Fermentation in Yeast Extensions to Basic Lab
 Using ADE Mutants to Study Respiration and Fermentation in Yeast Extensions to Basic Lab Introduction: David Form, Nashoba Regional High School And Jessica Forton, Melrose High School Bioinformatics Component
Using ADE Mutants to Study Respiration and Fermentation in Yeast Extensions to Basic Lab Introduction: David Form, Nashoba Regional High School And Jessica Forton, Melrose High School Bioinformatics Component
Supporting Information
 Supporting Information López et al. 10.1073/pnas.0810940106 1. Ivey DM, et al. (1993) Cloning and characterization of a putative Ca2 /H antiporter gene from Escherichia coli upon functional complementation
Supporting Information López et al. 10.1073/pnas.0810940106 1. Ivey DM, et al. (1993) Cloning and characterization of a putative Ca2 /H antiporter gene from Escherichia coli upon functional complementation
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Microbial Taxonomy. C. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy 1. Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eucaryote, is in a mess we are stuck with it for traditional
Microbial Taxonomy 1. Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eucaryote, is in a mess we are stuck with it for traditional
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Prof. Fahd M. Nasr. Lebanese university Faculty of sciences I Department of Natural Sciences.
 Prof. Fahd M. Nasr Lebanese university Faculty of sciences I Department of Natural Sciences fnasr@ul.edu.lb B3206 Microbial Genetics Eukaryotic M. G. The yeast Saccharomyces cerevisiae as a genetic model
Prof. Fahd M. Nasr Lebanese university Faculty of sciences I Department of Natural Sciences fnasr@ul.edu.lb B3206 Microbial Genetics Eukaryotic M. G. The yeast Saccharomyces cerevisiae as a genetic model
The Evolution of Multicellularity
 The Evolution of Multicellularity agenda Evolution overview Microevolution and macroevolution Evolution of multicellularity--a laboratory investigation possible hypotheses experimental design data collection
The Evolution of Multicellularity agenda Evolution overview Microevolution and macroevolution Evolution of multicellularity--a laboratory investigation possible hypotheses experimental design data collection
Essential knowledge 1.A.2: Natural selection
 Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Microbiology Helmut Pospiech
 Microbiology http://researchmagazine.uga.edu/summer2002/bacteria.htm 05.04.2018 Helmut Pospiech The Species Concept in Microbiology No universally accepted concept of species for prokaryotes Current definition
Microbiology http://researchmagazine.uga.edu/summer2002/bacteria.htm 05.04.2018 Helmut Pospiech The Species Concept in Microbiology No universally accepted concept of species for prokaryotes Current definition
Warm Up. What are some examples of living things? Describe the characteristics of living things
 Warm Up What are some examples of living things? Describe the characteristics of living things Objectives Identify the levels of biological organization and explain their relationships Describe cell structure
Warm Up What are some examples of living things? Describe the characteristics of living things Objectives Identify the levels of biological organization and explain their relationships Describe cell structure
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Verification of the Principles of Genetics by Manipulating Saccharomyces Cerevisiae and Observing Meiotic Products ABSTRACT
 November 27, 2002 Bio251 Dr. Calhoon Verification of the Principles of Genetics by Manipulating Saccharomyces Cerevisiae and Observing Meiotic Products ABSTRACT The experiment was conducted to verify the
November 27, 2002 Bio251 Dr. Calhoon Verification of the Principles of Genetics by Manipulating Saccharomyces Cerevisiae and Observing Meiotic Products ABSTRACT The experiment was conducted to verify the
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Essentiality in B. subtilis
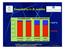 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Text of objective. Investigate and describe the structure and functions of cells including: Cell organelles
 This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
Enduring Understanding: Change in the genetic makeup of a population over time is evolution Pearson Education, Inc.
 Enduring Understanding: Change in the genetic makeup of a population over time is evolution. Objective: You will be able to identify the key concepts of evolution theory Do Now: Read the enduring understanding
Enduring Understanding: Change in the genetic makeup of a population over time is evolution. Objective: You will be able to identify the key concepts of evolution theory Do Now: Read the enduring understanding
Proteomics. 2 nd semester, Department of Biotechnology and Bioinformatics Laboratory of Nano-Biotechnology and Artificial Bioengineering
 Proteomics 2 nd semester, 2013 1 Text book Principles of Proteomics by R. M. Twyman, BIOS Scientific Publications Other Reference books 1) Proteomics by C. David O Connor and B. David Hames, Scion Publishing
Proteomics 2 nd semester, 2013 1 Text book Principles of Proteomics by R. M. Twyman, BIOS Scientific Publications Other Reference books 1) Proteomics by C. David O Connor and B. David Hames, Scion Publishing
Biology 112 Practice Midterm Questions
 Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc
 OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
Map of AP-Aligned Bio-Rad Kits with Learning Objectives
 Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS. Scientific Background on the Nobel Prize in Medicine 2013
 DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS Scientific Background on the Nobel Prize in Medicine 2013 Daniela Scalet 6/12/2013 The Nobel Prize in Medicine
DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS Scientific Background on the Nobel Prize in Medicine 2013 Daniela Scalet 6/12/2013 The Nobel Prize in Medicine
Introduction to polyphasic taxonomy
 Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
-CH 2. -S-CH 3 How do I know this?
 What is the major functional difference of Theta vs. Rolling-ircle replication, specifically in regards to the tandem linear repeats found in the R- replication. You won t be responsible for this. Rolling-circle
What is the major functional difference of Theta vs. Rolling-ircle replication, specifically in regards to the tandem linear repeats found in the R- replication. You won t be responsible for this. Rolling-circle
-SQA-SCOTTISH QUALIFICATIONS AUTHORITY. Hanover House 24 Douglas Street GLASGOW G2 7NQ NATIONAL CERTIFICATE MODULE DESCRIPTOR
 -SQA-SCOTTISH QUALIFICATIONS AUTHORITY Hanover House 24 Douglas Street GLASGOW G2 7NQ NATIONAL CERTIFICATE MODULE DESCRIPTOR -Module Number- 7312012 -Session-1992-93 -Superclass- RH -Title- EVOLUTION (x
-SQA-SCOTTISH QUALIFICATIONS AUTHORITY Hanover House 24 Douglas Street GLASGOW G2 7NQ NATIONAL CERTIFICATE MODULE DESCRIPTOR -Module Number- 7312012 -Session-1992-93 -Superclass- RH -Title- EVOLUTION (x
Supporting information
 Supporting information Synthesis of the compatible solute proline by Bacillus subtilis: point mutations rendering the osmotically controlled prohj promoter hyperactive Tamara Hoffmann 1*, Monika Bleisteiner
Supporting information Synthesis of the compatible solute proline by Bacillus subtilis: point mutations rendering the osmotically controlled prohj promoter hyperactive Tamara Hoffmann 1*, Monika Bleisteiner
Computational Cell Biology Lecture 4
 Computational Cell Biology Lecture 4 Case Study: Basic Modeling in Gene Expression Yang Cao Department of Computer Science DNA Structure and Base Pair Gene Expression Gene is just a small part of DNA.
Computational Cell Biology Lecture 4 Case Study: Basic Modeling in Gene Expression Yang Cao Department of Computer Science DNA Structure and Base Pair Gene Expression Gene is just a small part of DNA.
The geneticist s questions. Deleting yeast genes. Functional genomics. From Wikipedia, the free encyclopedia
 From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
Bioinformatics. Dept. of Computational Biology & Bioinformatics
 Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Unit # - Title Intro to Biology Unit 1 - Scientific Method Unit 2 - Chemistry
 Intro to Biology Unit 1 - Scientific Method Unit 2 - Chemistry What is Biology? What is Science? What tools, skills, knowledge, and dispositions are needed to conduct scientific inquiry? How do the rules
Intro to Biology Unit 1 - Scientific Method Unit 2 - Chemistry What is Biology? What is Science? What tools, skills, knowledge, and dispositions are needed to conduct scientific inquiry? How do the rules
Evolution in Action: The Power of Mutation in E. coli
 Evolution in Action: The Power of Mutation in E. coli by Merle K. Heidemann, College of Natural Science, Emeritus Peter J. T. White, Lyman Briggs College James J. Smith, Lyman Briggs College Michigan State
Evolution in Action: The Power of Mutation in E. coli by Merle K. Heidemann, College of Natural Science, Emeritus Peter J. T. White, Lyman Briggs College James J. Smith, Lyman Briggs College Michigan State
The phenotype of this worm is wild type. When both genes are mutant: The phenotype of this worm is double mutant Dpy and Unc phenotype.
 Series 2: Cross Diagrams - Complementation There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Series 2: Cross Diagrams - Complementation There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Supplemental Data. Perea-Resa et al. Plant Cell. (2012) /tpc
 Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
The Evolution of Stress-Induced Mutagenesis in Finite and Infinite Populations. Yoav Ram & Lilach Hadany Tel-Aviv University
 The Evolution of Stress-Induced Mutagenesis in Finite and Infinite Populations Yoav Ram & Lilach Hadany Tel-Aviv University PopGroup45, Nottingham UK, January 2012 Evolution of the Mutation Rate Deleterious
The Evolution of Stress-Induced Mutagenesis in Finite and Infinite Populations Yoav Ram & Lilach Hadany Tel-Aviv University PopGroup45, Nottingham UK, January 2012 Evolution of the Mutation Rate Deleterious
Critical Mutation Rate Has an Exponential Dependence on Population Size
 Critical Mutation Rate Has an Exponential Dependence on Population Size Alastair Channon 1, Elizabeth Aston 1, Charles Day 1, Roman V. Belavkin 2 and Christopher G. Knight 3 1 Research Institute for the
Critical Mutation Rate Has an Exponential Dependence on Population Size Alastair Channon 1, Elizabeth Aston 1, Charles Day 1, Roman V. Belavkin 2 and Christopher G. Knight 3 1 Research Institute for the
Chapter 15: Darwin and Evolution
 Chapter 15: Darwin and Evolution AP Curriculum Alignment Big Idea 1 is about evolution. Charles Darwin is called the father of evolution because his theory of natural selection explains how evolution occurs.
Chapter 15: Darwin and Evolution AP Curriculum Alignment Big Idea 1 is about evolution. Charles Darwin is called the father of evolution because his theory of natural selection explains how evolution occurs.
BIOLOGY STANDARDS BASED RUBRIC
 BIOLOGY STANDARDS BASED RUBRIC STUDENTS WILL UNDERSTAND THAT THE FUNDAMENTAL PROCESSES OF ALL LIVING THINGS DEPEND ON A VARIETY OF SPECIALIZED CELL STRUCTURES AND CHEMICAL PROCESSES. First Semester Benchmarks:
BIOLOGY STANDARDS BASED RUBRIC STUDENTS WILL UNDERSTAND THAT THE FUNDAMENTAL PROCESSES OF ALL LIVING THINGS DEPEND ON A VARIETY OF SPECIALIZED CELL STRUCTURES AND CHEMICAL PROCESSES. First Semester Benchmarks:
Enduring understanding 1.A: Change in the genetic makeup of a population over time is evolution.
 The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
Vital Statistics Derived from Complete Genome Sequencing (for E. coli MG1655)
 We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
ECOL/MCB 320 and 320H Genetics
 ECOL/MCB 320 and 320H Genetics Instructors Dr. C. William Birky, Jr. Dept. of Ecology and Evolutionary Biology Lecturing on Molecular genetics Transmission genetics Population and evolutionary genetics
ECOL/MCB 320 and 320H Genetics Instructors Dr. C. William Birky, Jr. Dept. of Ecology and Evolutionary Biology Lecturing on Molecular genetics Transmission genetics Population and evolutionary genetics
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
μ gyra parc Escherichia coli Klebsiella pneumoniae Pseudomonas aeruginosa E. coli gyra E. coli parc gyra parc gyra Escherichia coli E. coli E.
 gyra parc Escherichia coli Klebsiella pneumoniae Pseudomonas aeruginosa E. coli μ E. coli gyra parc gyra parc gyra parc μ μ gyra parc Key words Escherichia coli gyra parc Escherichia coli E. coli gyra
gyra parc Escherichia coli Klebsiella pneumoniae Pseudomonas aeruginosa E. coli μ E. coli gyra parc gyra parc gyra parc μ μ gyra parc Key words Escherichia coli gyra parc Escherichia coli E. coli gyra
1. CHEMISTRY OF LIFE. Tutorial Outline
 Tutorial Outline North Carolina Tutorials are designed specifically for the Common Core State Standards for English language arts, the North Carolina Standard Course of Study for Math, and the North Carolina
Tutorial Outline North Carolina Tutorials are designed specifically for the Common Core State Standards for English language arts, the North Carolina Standard Course of Study for Math, and the North Carolina
A redundant function for the N-terminal tail of Ndc80 in kinetochore-microtubule interaction in Saccharomyces cerevisiae.
 Genetics: Published Articles Ahead of Print, published on July 30, 2012 as 10.1534/genetics.112.143818 A redundant function for the N-terminal tail of Ndc80 in kinetochore-microtubule interaction in Saccharomyces
Genetics: Published Articles Ahead of Print, published on July 30, 2012 as 10.1534/genetics.112.143818 A redundant function for the N-terminal tail of Ndc80 in kinetochore-microtubule interaction in Saccharomyces
7.32/7.81J/8.591J. Rm Rm (under construction) Alexander van Oudenaarden Jialing Li. Bernardo Pando. Rm.
 Introducing... 7.32/7.81J/8.591J Systems Biology modeling biological networks Lectures: Recitations: ti TR 1:00-2:30 PM W 4:00-5:00 PM Rm. 6-120 Rm. 26-204 (under construction) Alexander van Oudenaarden
Introducing... 7.32/7.81J/8.591J Systems Biology modeling biological networks Lectures: Recitations: ti TR 1:00-2:30 PM W 4:00-5:00 PM Rm. 6-120 Rm. 26-204 (under construction) Alexander van Oudenaarden
Statistical Models in Evolutionary Biology An Introductory Discussion
 Statistical Models in Evolutionary Biology An Introductory Discussion Christopher R. Genovese Department of Statistics Carnegie Mellon University http://www.stat.cmu.edu/ ~ genovese/ JSM 8 August 2006
Statistical Models in Evolutionary Biology An Introductory Discussion Christopher R. Genovese Department of Statistics Carnegie Mellon University http://www.stat.cmu.edu/ ~ genovese/ JSM 8 August 2006
The geneticist s questions
 The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
PROTEIN SYNTHESIS INTRO
 MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
Computational Biology From The Perspective Of A Physical Scientist
 Computational Biology From The Perspective Of A Physical Scientist Dr. Arthur Dong PP1@TUM 26 November 2013 Bioinformatics Education Curriculum Math, Physics, Computer Science (Statistics and Programming)
Computational Biology From The Perspective Of A Physical Scientist Dr. Arthur Dong PP1@TUM 26 November 2013 Bioinformatics Education Curriculum Math, Physics, Computer Science (Statistics and Programming)
When one gene is wild type and the other mutant:
 Series 2: Cross Diagrams Linkage Analysis There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Series 2: Cross Diagrams Linkage Analysis There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
WHERE DOES THE VARIATION COME FROM IN THE FIRST PLACE?
 What factors contribute to phenotypic variation? The world s tallest man, Sultan Kosen (8 feet 1 inch) towers over the world s smallest, He Ping (2 feet 5 inches). WHERE DOES THE VARIATION COME FROM IN
What factors contribute to phenotypic variation? The world s tallest man, Sultan Kosen (8 feet 1 inch) towers over the world s smallest, He Ping (2 feet 5 inches). WHERE DOES THE VARIATION COME FROM IN
The type IV pilus - WWU Münster Kalvi Institute,
 The type IV pilus - a multifunctional bacterial motor Berenike Maier WWU Münster Kalvi Institute, 1.2.2011 1 Type IV pili are ubiquitous multifunctional polymers 2 Type IV pili are ubiquitous multifunctional
The type IV pilus - a multifunctional bacterial motor Berenike Maier WWU Münster Kalvi Institute, 1.2.2011 1 Type IV pili are ubiquitous multifunctional polymers 2 Type IV pili are ubiquitous multifunctional
The Double Helix. CSE 417: Algorithms and Computational Complexity! The Central Dogma of Molecular Biology! DNA! RNA! Protein! Protein!
 The Double Helix SE 417: lgorithms and omputational omplexity! Winter 29! W. L. Ruzzo! Dynamic Programming, II" RN Folding! http://www.rcsb.org/pdb/explore.do?structureid=1t! Los lamos Science The entral
The Double Helix SE 417: lgorithms and omputational omplexity! Winter 29! W. L. Ruzzo! Dynamic Programming, II" RN Folding! http://www.rcsb.org/pdb/explore.do?structureid=1t! Los lamos Science The entral
Microbial Genetics, Mutation and Repair. 2. State the function of Rec A proteins in homologous genetic recombination.
 Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
AP Biology Curriculum Framework
 AP Biology Curriculum Framework This chart correlates the College Board s Advanced Placement Biology Curriculum Framework to the corresponding chapters and Key Concept numbers in Campbell BIOLOGY IN FOCUS,
AP Biology Curriculum Framework This chart correlates the College Board s Advanced Placement Biology Curriculum Framework to the corresponding chapters and Key Concept numbers in Campbell BIOLOGY IN FOCUS,
Stoichiometric analysis of metabolic networks
 1 SGN-6156 Computational systems biology II Antti Larjo antti.larjo@tut.fi Computational Systems Biology group 30.4.2008 30.4.2008 2 Contents Networks in cells Metabolism Metabolic networks and pathways
1 SGN-6156 Computational systems biology II Antti Larjo antti.larjo@tut.fi Computational Systems Biology group 30.4.2008 30.4.2008 2 Contents Networks in cells Metabolism Metabolic networks and pathways
Miller & Levine Biology 2014
 A Correlation of Miller & Levine Biology To the Essential Standards for Biology High School Introduction This document demonstrates how meets the North Carolina Essential Standards for Biology, grades
A Correlation of Miller & Levine Biology To the Essential Standards for Biology High School Introduction This document demonstrates how meets the North Carolina Essential Standards for Biology, grades
phenomenon called cross resistance. As a consequence of cross resistance the entire class of aminoglycosides looses its therapeutic potential.
 Experiment 25 Laboratory to Biology III Diversity of Microorganisms / Wintersemester / page 1 Mechanisms of aminoglycoside resistance in mycobacteria Advisor P.D. Dr. Peter Sander, psander@immv.unizh.ch,
Experiment 25 Laboratory to Biology III Diversity of Microorganisms / Wintersemester / page 1 Mechanisms of aminoglycoside resistance in mycobacteria Advisor P.D. Dr. Peter Sander, psander@immv.unizh.ch,
Phylogenetics in the Age of Genomics: Prospects and Challenges
 Phylogenetics in the Age of Genomics: Prospects and Challenges Antonis Rokas Department of Biological Sciences, Vanderbilt University http://as.vanderbilt.edu/rokaslab http://pubmed2wordle.appspot.com/
Phylogenetics in the Age of Genomics: Prospects and Challenges Antonis Rokas Department of Biological Sciences, Vanderbilt University http://as.vanderbilt.edu/rokaslab http://pubmed2wordle.appspot.com/
