Quantitative Phenetics and Taxonomy of Some Phlebotomine Taxa
|
|
|
- Nora Warner
- 6 years ago
- Views:
Transcription
1 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 94(6): , Nov./Dec Quantitative Phenetics and Taxonomy of Some Phlebotomine Taxa JP Dujardin/ +, F Le Pont*, E Martinez** 735 UMR IRD-CNRS 9926, IRD Montpellier, Av. Agropolis, 911, Montpellier, France *ORSTOM La Paz, Casilla Postal 9214, La Paz, Bolivia **Instituto Boliviano de Biologia de Altura, Calle Claudio Sanjinez, La Paz, Bolivia Elucidating the evolution of Phlebotominae is important not only to revise their taxonomy, but also to help understand the origin of the genus Leishmania and its relationship with humans. Our study is a phenetic portrayal of this history based on the genetic relationships among some New Word and Old Word taxa. We used both multilocus enzyme electrophoresis and morphometry on 24 male specimens of the Old Word genus Phlebotomus (with three of its subgenera: Phlebotomus, Spelaeophlebotomus and Australophlebotomus), and on 67 male specimens of the three New World genera, Warileya, Brumptomyia and Lutzomyia, (with three subgenera of Lutzomyia: Lutzomyia, Oligodontomyia and Psychodopygus). Phenetic trees derived from both techniques were similar, but disclosed relationships that disagree with the present classification of sand flies. The need for a true evolutionary approach is stressed. Key words: Old World New World sand flies - multilocus enzyme electrophoresis - morphometrics The oldest known species of Phlebotominae, Phlebotomites longifilis and P. brevifilis, two fossil records described from Lebanon, lived about 120 millions years ago (MYA) (Lewis 1982). The evolution of Phlebotominae since that time was probably driven by major tectonic events and related climatic changes that affected Pangaea. Phlebotominae are bloodsucking insects of temperate and tropical regions showing significant differences in ecological adaptations, endemism, relictuality and species richness. They include important vectors of Leishmania spp., Bartonella sp. and Phleboviruses. The systematics of this subfamily, especially at the supraspecific level, has always been controversial (Lewis & Dice 1982, Lane 1986). In the last two decades, a conservative approach based on practical criteria (Lewis et al. 1977) led to the present subdivision of the Phlebotominae into six genera: three Old World (OW) genera: Chinius (1 species), Phlebotomus (10 subgenera) and Sergentomyia (7 subgenera), and three New World (NW) genera: Brumptomyia (22 species), Warileya (6 species) and Lutzomyia (26 subgenera) (Lane 1993, Young & Duncan 1994). This stable clas- + Corresponding author. Present address: IRD (ORSTOM) La Paz, Casilla Postal 9214, La Paz, Bolivia. Fax: dujardin@mail.megalink.com. Received 26 October 1998 Accepted 10 June 1999 sification (Lewis et al. 1977) has been generally accepted, though it was considered as premature by some experts of the NW sand fly fauna, and different conceptions have been applied (Forattini 1973, Ready et al. 1980). Recent attempts to bring evolutionary insight into this classification were those of two independent doctoral theses (Rispail 1990, Galati 1992). They used Hennigian methodology (Hennig 1972) based on 100 (Galati 1992) and 28 (Rispail 1990) adult morphological attributes. According to these studies, the NW genus Warileya (Rispail 1990, Galati 1992) and the OW Spelaeophlebotomus and Australophlebotomus (Rispail 1990) (both Phlebotomus subgenera) appeared to have older, more ancient origins than suggested by the present taxonomic classification (Rispail & Leger 1998a). Both studies disagreed, however, on the evolutionary status of the NW genus Brumptomyia that is close either to Lutzomyia (Rispail 1990) or to Phlebotomus (Galati 1992), as well as that of the OW genus Sergentomyia (not included in our material), clustered with Spelaeophlebotomus (Rispail 1990) or with Lutzomyia (Galati 1992, 1995). In the absence of available or relevant outgroup for Phlebotominae, we used a phenetic approach based on dissimilarity indexes, called here quantitative phenetics. By this appellation, we mean a non-phylogenetic approach intended to generate hypotheses about evolutionary history, and based on quantitative characters. It may be regarded as a heuristic device subject to capture some evolutionary trends among not too closely related organisms, where higher probability exists of agreement between phylogenetic and genetic divergences
2 736 A Phenetic Approach to Some Phlebotomine Taxa JP Dujardin et al. (Chavez et al. 1999). We applied it here to the males of some species of sand flies belonging to widely different genera. Quantitative characters are needed because they are more suited to estimate genetic differences than morphological attributes. To increase their likeliness of producing relevant information about evolutionary history, they must be submitted to appropriate analyses. An UPGMA tree was derived from genetic distances for isoenzyme electrophoresis (Nei 1987, Solé-Cava et al. 1994). A canonical variate analysis (multivariate discriminant analysis) was applied to metric data (Pimentel 1992, Sorensen & Foottit 1992). Both statistical techniques have proved informative to outline evolutionary trends in other organisms (Simon 1992, Dujardin et al. 1999). MATERIALS AND METHODS Isoenzyme electrophoresis and morphometry were applied to the same adult males of the following nine species (Table I): Warileya fourgassiensis Le Pont & Desjeux 1984, W. rotundipennis Fairchild & Hertig 1951, Phlebotomus (Spelaeophlebotomus) gigas Parrot & Schwetz 1937, P. (Australophlebotomus) notteghemae Leger & Pesson 1993, P. papatasi Scopoli 1786, Lutzomyia (Lutzomyia) longipalpis (Lutz & Neiva, 1912), L. (Psychodopygus) geniculata (Mangabeira 1941), Brumptomyia pintoi (Lima 1932), and Lutzomyia (Oligodontomyia) toroensis Le Pont, Torres- Espejo & Dujardin 1996 (Table I). The classification adopted here is from Young and Duncan (1994), who consider Psychodopygus as a Lutzomyia subgenus, but we grouped L. toroensis with two related species (L. isopsi and L. oligodonta ) in the recently created Lutzomyia subgenus Oligodontomyia (Galati 1995). Twelve enzyme systems (ALDH, AP, HK, GPD, GPI, IDH, ME, MDH, PEP, PGM, XDH and XO) were run on cellulose acetate according to Richardson et al. (1986) and Dujardin et al. (1996). For genetic relatedness analysis, a phenetic approach was preferred to a cladistic one because our material did not include an outgroup for Phlebotominae (see however comments of Fig. 1). As an immediate measure of genetic divergence, we computed the Jaccard s distances (Jackson et al. 1989) which do not depend on gene frequencies or on locus counting (Table III, below diagonal). From these distances, an UPGMA tree was constructed. Using a camera lucida, we measured eight characters using features of the head (AIII, third antennal segment; EP, epipharynx; P5, fifth palpomere); the wing (WL, wing length, WW, wing width) and the genitalia (GF, genital filaments; GP, genital pump; LL, lateral lobe). Size-free variables could not be derived from our (log-transformed) data because of lack of highly significant correlation between the first pooled within-group principal component and some variables (Dos Reis et al. 1990). Thus, discriminant analysis included size and shape variation. The derived Mahalanobis distances (Mahalanobis 1936) (Table III, above diagonal) were also submitted to the same UPGMA tree-making method as for distances derived from electrophoretic data (Fig. 1, right side). The correlation between Jaccard and Mahalanobis distances was explored by the Mantel test (10,000 runs), and illustrated by the exponential relationship between both distances. TABLE I Origins of male insects and dates of capture Genus Phlebotomus (OW, n = 24) P. (P.) papatasi (n = 10) Laboratory rearing, Montpellier (France), 1996 P. (S.) gigas (n = 4) Meya-N Zouari caves (République du Congo), 1996 P. (A.) notteghemae (n = 10) Touaourou caves (Nouvelle Calédonie), 1997 Genus Warileya (NW, n = 20) W. fourgassiensis (n = 10) Fourgassié caves (Guyane Française), 1985 W. rotundipennis (n = 10) Suapi (Bolivia), 1995 Genus Lutzomyia (NW, n = 40) L. (L.) longipalpis (n = 10) Santa Barbara, North Yungas (Bolivia), 1996 L. (O.) toroensis (n = 10) Toro Toro cave, Potosi (Bolivia), 1995 L. (P.) geniculata (n = 20) Forest, Carrasco, North Yungas (Bolivia), 1997 Genus Brumptomyia (NW, n = 7) B. pintoi (n = 7) Forest, Carrasco, North Yungas (Bolivia), 1997 Fresh material, used for isoenzyme electrophoresis and morphometry, was collected by light traps by one of us (FLP) in the Republic of Congo, New Caledonia and Bolivia between 1995 and P. papatasi originated from the insectary of the Faculté de Médecine, Université de Montpellier (Prof. JP Dedet). W. fourgassiensis were mounted specimens examined by morphometry only. OW: Old World; NW: New World.
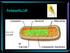
3 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 94(6), Nov./Dec Jaccard 0.1 SPEL WARI AUST BRUM PSYC OLIG LUTZ PHLE 1 2 Mahalanobis 1000 Fig. 1: topology of phenetic trees derived from either isoenzymes (left side) or metric data (right side). Shaded zones refer to Old World taxa. WARI: Warileya; 1: Warileya fourgassiensis (not examined by isoenzymes); 2: W. rotundipennis; AUST: Australophlebotomus; SPEL: Spelaeophlebotomus; BRUM: Brumptomyia; PSYC: Psychodopygus; OLIG: Oligodontomyia; PHLE: Phlebotomus; LUTZ: Lutzomyia. Although the two Warileya species showed striking differences in size, they were clustered together in the metric tree, indicating poor size-effect in the disclosed relationships. Using Warileya as a tentative outgroup, Hennigian methodology on isoenzymes produced a similar tree (results not shown). Presentation and scaling of the trees were obtained by TREE VIEW (Copyright Roderick DM Page, 1998 r.page@bio.gla.ac.uk) on tree files generated by the PHYLIP 3.4 package. RESULTS Isoenzyme analysis - Each enzyme produced bands for each specimen. A total of 95 different electrophoretic band positions was recorded. In the total sample covering both OW and NW species, 37 alleles were found unique to one species, i.e. some species had one or more alleles not shared with other species. Notably, the NW genus Lutzomyia showed no unique allele. The maximum number of shared alleles between two species was found between a NW (L. longipalpis) and an OW (P. papatasi) species, while the minimum number of shared alleles was found between either OW (Spelaeophlebotomus with Australophlebotomus) or NW representatives (see Warileya with either Psychodopygus or Oligodontomyia) (Table II, top). Subdividing the total sample into OW on one hand, and NW on the other hand, unique alleles were found more abundant in the former region (74% versus 56%), although it was represented by one genus only (Phlebotomus). Conversely, the proportion of alleles each region shared among its own taxa was lower in the OW (16/61, 26%) than among the four NW taxa belonging to three different genera (34/77, 44%). Both regions shared a total of 43 alleles (45%, i.e. 43/95) (Table II, bottom). Metric analysis - Although data were log-transformed before multivariate analysis, only three (37%) characters (EP, WW, LL) had a distribution TABLE II Allelic composition of the total sample subdivided by taxa (top) or regions (bottom) Taxa AUST SPEL PHLE LUTZ PSYC OLIG BRUM WARI Total alleles Unique alleles Shared alleles AUST SPEL PHLE LUTZ PSYC OLIG BRUM WARI AUST / SPEL 2 / PHLE 6 4 / LUTZ / PSYC / OLIG / BRUM / WARI / Regions Old World New World Total alleles Unique alleles Shared alleles between species Old World New World Old World 16 (26%) 43 (45%) New World 43 (45%) 34 (44%) Unique alleles are alleles found in one species only ( unshared alleles), either considering the total sample (top) or each region separately (bottom). Region refers to either Old World, with three taxa, or New World, with five taxa (see Materials and Methods). Shared alleles are alleles found in common between two species in the total sample (top), in each region separately (bottom) or between two regions (bottom). The three first taxa are Old World species AUST: Australophlebotomus; SPEL: Spelaeophlebotomus; PHLE: Phlebotomus. New World species are LUTZ: Lutzomyia; PSYC: Psychodopygus; OLIG: Oligodontomyia; BRUM: Brumptomyia; WARI: Warileya (W. rotundipennis).
4 738 A Phenetic Approach to Some Phlebotomine Taxa JP Dujardin et al. TABLE III Jaccard (below diagonal) and Mahalanobis (above diagonal) distances AUST SPEL PHLE LUTZ PSYC OLIG BRUM WARI AUST SPEL PHLE LUTZ PSYC OLIG BRUM WARI Below diagonal are the Jaccard s distances (Dj) derived from electrophoretic data. Above diagonal are the Mahalanobis distances (Dm) computed from discriminant analysis on (log-transformed) measurements of head, wing and genitalia characters. The three first taxa are Old World species AUST: Australophlebotomus; SPEL: Spelaeophlebotomus; PHLE: Phlebotomus. New World species are LUTZ: Lutzomyia; PSYC: Psychodopygus; OLIG: Oligodontomyia; BRUM: Brumptomyia; WARI: Warileya (W. rotundipennis). that conformed with normality, and only two were compatible with homoscedasticity (equal variances among groups) (detailed results not shown). Normality and homoscedasticity are the current assumptions of discriminant analysis (also called canonical variate analysis), however departure from these assumptions are known to have little influence on analysis (Pimentel 1992). Discriminant analysis was highly significant (P < after Wilk s test) (Wilks 1932). Correspondance between analyses - Both trees produced the same topology except for the closer clustering of Warileya and Australophlebotomus after morphometric analysis, or the closer clustering of Warileya and Spelaeophlebotomus after isoenzyme analysis. These three genera were the most external ones in both trees (see Fig. 1). Grouping them into one OTU (organizational taxonomic unit), morphometrics and isoenzyme data produced the same topology. For at least six OTUs to produce by chance an identical topology after different techniques, the estimated probability was (1/945, see Nei 1987 p. 290). A further measure of agreement between both approaches was the significant correlation (Mantel test, P = after 10,000 runs) disclosed between Mahalanobis and Jaccard s distances, which was illustrated here by an exponential relationship (Fig. 2). DISCUSSION In our material composed of male specimens, three taxonomic groups are located on different continents, (i) the OW Phlebotomus and Spelaeophlebotomus, (ii) the Australasian OW Australophlebotomus and (iii) the remaining NW taxa. As vicariant products after fragmentation and resulting drift of Gondwanan plates (Lewis et al. 1977), these groups should have important geological times of divergence between them. For such distant taxa, electrophoresis of isoenzymes is usually not recommended (Thorpe & Solé-Cava 1994), while discriminant analysis on continuous characters, which was used here as an exploratory phylogenetic tool (Sorensen & Footit 1992, Sorensen 1992), remains controversial (Crespi 1992). Neither proteic nor metric trees respected geography (Fig. 1); they were, however, very similar between them, and this congruence was hardly compatible with chance alone. Cladistic trees based on adult morphology showed similar patterns as ours regarding the external positions of Warileya (Rispail 1990, Galati 1992) and Spelaeophlebotomus (Rispail 1990, Rispail & Leger 1998a), but disagreed regarding the respective positions of Brumptomyia Dm Fig. 2: plotting of Mahalanobis (Dm, vertical axis, metric data) against Jaccard (Dj, horizontal axis, isoenzyme data) distances, showing the positive and significant correlation between them (P = after Mantel test, 10,000 runs). The curved line represents the exponential relationship between distances (coefficient of determination was 0.62) Dj 1.0
5 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 94(6), Nov./Dec and Psychodopygus, both of which are closer to Lutzomyia than to Phlebotomus. Note, however, that between Lutzomyia (NW) and Phlebotomus (OW), Rispail (1990) did not produce distances significantly higher than zero. Three salient features were shared by electrophoretic and metric data, (i) a high degree of divergence of either Spelaeophlebotomus (OW) or Warileya (NW) with other taxa, (ii) a slightly lower degree of divergence of Australophlebotomus (Australasian region) with South America, relative to Africa, and (iii) the pairing of Phlebotomus (OW) with Lutzomyia (NW), closer together than to any other genera or subgenera with which they were expected to show more affinities. The pairing of the OW subgenus Spelaeophlebotomus with the NW genus Warileya after isoenzyme analysis in a separate, external group (Fig. 1) could be supported by the presence in these two taxa of some plesiomorphic structures recognized in fossil Phlebotomites species (120 MYA), such as rounded wings, a complete inter-ocular suture, legs ratios, as well as the unusual feature of rods near the sperm pump (Lewis 1982). The similarities of these taxa with fossil records of 120 MYA (Lewis 1982), and their external positions on either phenetic (this paper) or cladistic trees (Rispail 1990, Galati 1992), lend support to the idea that they existed in Gondwana before continental fragmentation, and that their populations would have been subjected to division as the continental plates began to separate. According to this hypothesis, they could be assembled within the same separate, sister taxon. The subgenus Australophlebotomus is restricted to the Australasian region. It was classified by Lewis and Dyce (1982) as a subgenus of Phlebotomus, but our study indicated relatively more affinities of Australophlebotomus with South American than with Afrotropical species (Fig. 1). There were fewer electrophoretic differences with the NW genus Lutzomyia (Jaccard 0.81, Table III below diagonal) and related subgenera ( ) than with Phlebotomus (0.85) or Spelaeophlebotomus (0.94). Remarkably, there was a very low metric distance with the genus Warileya (Mahalanobis 293, Table III above diagonal), which parallels the observation of primitive characters, as those described for Warileya, in some Australophlebotomus species (Lewis 1982, Lewis & Dyce 1982). The relatively lower divergence between Australophlebotomus and South American taxa could be attributed to either convergence or common ancestry. For other insects, morphological similarities between Australasian and South American taxa were related to possible exchanges during Eocene and Oligocene (55-25 MYA) (Moore 1978, Parsons 1996). Lewis and Dyce (1982) suggested a Southern origin for Australophlebotomus, and regretted the absence of available data about the Chilean fauna. For this reason we included in our material the Lutzomyia subgenus Oligodontomyia restricted to the south-western parts of South America, including Chile (Galati 1995). However, our data provided no clear evidence of close similarity between Oligodontomyia and Australophlebotomus (Jaccard 0.82/Mahalanobis 944, see Table III). The close proximity of Phlebotomus and Lutzomyia accords with their lack of relevant morphological differences (see Abonnenc & Leger 1976). According to Ashford (1991), these two genera were defined on the unique criterion of geography. Partial ribosomal sequences comparisons showed that some species of Phlebotomus could indeed cluster with some Lutzomyia species (Depaquit et al. 1998). Coincidentally or not, they are the sole genera containing Leishmania vectors. Our measurements of proteic and metric divergence showed that they were closer together than they are to their corresponding subgenera (Spelaeophlebotomus, Australophlebotomus, and Oligodontomyia, Psychodopygus, respectively). It is worth noting that this unexpected proximity was predicted by the hypothesis of a Neotropical origin of the genus Leishmania (Noyes 1998). How far our phenetic approach contains relevant phylogenetic information is however a matter of chance. Provided that the rate of gene substitution does not vary greatly among evolutionary lineages, the UPGMA tree-making method based on electrophoretic data usually contains reliable phylogenetic information (Nei 1987 p. 311, Sole- Cava et al. 1994). On the other hand, since discriminant analysis of metric data focuses upon unshared variation (Pimentel 1992), which may be considered apomorphic variation (Sorensen & Foottit 1992), it also could portray phylogenetic relationships (Sorensen 1992). In this application, discriminant analysis is poorly affected by nonnormality and heteroscedasticity, unless the sample sizes are strongly unequal (Pimentel 1992). Our data regarding OW taxa were in agreement with the cladistic approach proposed by Rispail and Leger (1998a). However, the pairing of Phlebotomus and Lutzomyia should be regarded with caution, since a phenetic approach could be seriously invalidated by homoplasy, convergence, or by strongly unequal evolutionary rates. This prevented us from proposing a new classification to replace the current one. We believe that to help answer these questions, true cladistical methods should be used (Rispail & Leger 1998a,b), based on relevant and practicable outgroups. In this regard, our data, as well as previous ones (Rispail 1990, Galati 1992),
6 740 A Phenetic Approach to Some Phlebotomine Taxa JP Dujardin et al. suggest that Warileya and/or Spelaeophlebotomus could be interesting candidates. There is no doubt that DNA sequencing should provide more accurate estimations of genetic divergences among main groups of sand flies, however such data are presently lacking. Besides, isoenzyme electrophoresis techniques suffer from practical constraints such as accessibility to frozen material, which makes it difficult to assemble specimens from different continents. Also, it is important to gather more information on various representatives of the main sand fly taxa, including the large genus Sergentomyia which we did not study. Thus, morphologic and morphometric studies on adults should be further explored, and could be also improved by the use of larval characters (Vattier-Bernard 1971, Parsons 1996). Other candidate characters, such as the spermatozoa (Dallai et al. 1984) and the chromosomal structures (White & Killick-Kendrick 1976), should be considered, along with molecular data, to provide a more complete analysis (Rangel et al. 1996). ACKNOWLEDGMENTS To Prof. JA Rioux, Dr P Rispail (University of Montpellier I), Mr J Mouchet (ORSTOM Paris), Prof. De Muyzon (MNHN Paris) and Prof. MD Bargues (University of Valencia) for revising this manuscript. To Prof. JP Dedet and Dr E Guilvard for generously providing specimens of P. papatasi. REFERENCES Abonnenc E, Leger N Sur une classification rationnelle des Diptères Phlebotomidae. Cah ORSTOM., sér. Entomol Méd Parasitol 14: Ashford RW A new morphological character to distinguish Sergentomyia and Phlebotomus, p In M Mardi, Proc. 1st Intl Symposium on Phlebotomine Sandflies, Rome, Vol. 33 (Suppl. 1). Chavez T, Moreno J, Dujardin JP Isoenzyme electrophoresis of Rhodnius species: a phenetic approach to relationships within the genus. Ann Trop Med Parasitol 93: Crespi BJ Natural selection and morphometrics, p In RG Foottit & JT Sorensen (eds), Ordination in the Study of Morphology, Evolution and Systematics of Insects: Applications and Quantitative Genetic Rationales, Elsevier, New York. Dallai R, Baccetti B, Macceni M, Sabatinelli G The spermatozoan of three species of Phlebotomus (Phlebotominae) and the acrosomal evolution in Nematoceran Dipterans. Intl Insect Morphol Embryol 13: Depaquit J, Perrotey S, Lecointre G, Tillier A, Tillier S, Ferte H, Kaltenbach M, Leger N Molecular systematics of Phlebotominae: a pilot study. Paraphyly of the genus Phlebotomus. CR Acad Sci Paris Life Sciences 321 : Dos Reis SF, Pessoa LM, Strauss RE Application of size-free canonical discriminant analysis to studies of geographic differentiation. Brazil J Genet 13: Dujardin JP, Chavez T, Moreno JM, Machane M, Noireau F, Schofield CJ Rhodniini: the value of traditional morphometrics as a phylogenetic approach. J Med Entomol in press. Dujardin JP, Le Pont F, Cruz M, Leon R, Tarrieu F, Guderian R, Echeverria R, Tibayrenc M Cryptic speciation in Lutzomyia (Nyssomyia) trapidoi (Fairchild & Hertig) (Diptera: Psychodidae) detected by Multilocus Enzyme Electrophoresis. Amer J Trop Med Hyg 54: Forattini OP Entomologia Médica. Psychodidae. Phlebotominae. Leismanioses. Bartonelose, Vol 4, Edgar Blücher Ltda., São Paulo. Galati EAB Sistemática dos Phlebotominae (Diptera, Psychodidae) das Américas. Thesis, São Paulo, 275 pp. Galati EAB Phylogenetic systematics of Phlebotominae (Diptera: Psychodidae) with emphasis on American groups. Bol Dir Malariol Saneam Amb 35: Hennig W Insektenfossilien aus der unteren Kreide. IV. Psychodidae (Phlebotominae). Stutt Beitr Naturk 241: Jackson DA, Somers KM, Harvey HH Similarity coefficients: measures of co-occurrence and association or simply measures of occurrence? The Amer Natur 133: Lane RP Recent advances in the systematics of phlebotomine sandflies. Insect Sci Applic: Lane RP Sandflies (Phlebotominae), p In RP Lane & RW Crosskey (eds), Medical Insects and Arachnids, Chapman & Hall, London, 723 pp. Lewis DJ A taxonomic review of the genus Phlebotomus (Diptera: Psychodidae). Bull Br Mus Nat Hist (Entomol) 45: Lewis DJ, Dyce AL The subgenus Australophlebotomus Theodor of Phlebotomus Rondani and Berté (Diptera: Psychodidae). J Aust Entomol Soc 21: Lewis DJ, Young DG, Fairchild GB, Minter DM Proposals for a stable classification of phlebotomine sandflies. Syst Entomol 2: Mahalanobis PC On the generalized distance in statistics. Proc Natl Inst Sci India 2: Moore BP A new Australian stag beetle (Coleoptera: Lucanidae) with Neotropical affinities. J Aust Entomol Soc 17: Nei M Molecular Evolutionary Genetics, Columbia University Press, New York, 512 pp. Noyes H Implications of Neotropical origin of the Genus Leishmania. Mem Inst Oswaldo Cruz 93: Parsons MJ Gondwana evolution of the troidine swallowtails (Lepidoptera: Papilionidae): cladistic reappraisals using mainly immature stage characters, with focus on the birdwings ornithoptera Boisduval. Bull Kitakyushu Mus Nat Hist 15: Pimentel RA An introduction to ordination, principal components analysis and discriminant analysis, p In RG Foottit & JT Sorensen (eds),
7 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 94(6), Nov./Dec Ordination in the Study of Morphology, Evolution and Systematics of Insects: Applications and Quantitative Genetic Rationales, Elsevier, New York. Rangel EF, Lainson R, Souza AA, Ready P, Azevedo ACR Variation between geographical populations of Lutzomyia (Nyssomyia) whitmani (Antunes & Coutinho, 1939) sensu lato (Diptera: Psychodidae: Phlebotominae) in Brazil. Mem Inst Oswaldo Cruz 91: Ready PD, Fraiha H, Lainson R, Shaw J Psychodopygus as a genus: reasons for a flexible classification of the phlebotomine sand flies (Diptera: Psychodidae). J Med Entomol 17: Richardson BJ, Baverstock PR, Adams SM Allozyme electrophoresis: a Handbook for Animal Systematics and Population Studies, Academic Press, Australia, Orlando, Florida, 409 pp. Rispail P Approche Phénétique et Cladistique du Genre Phlebotomus Rondani & Berté, 1840 (Diptera: Psychodidae). Apport des Caractères Morphologiques Imaginaux, PhD Thesis, Montpellier. Rispail P, Léger N 1998a. Numerical taxonomy of Old World Phlebotominae (Diptera: Psychodidae). 1. Considerations of morphological characters in the genus Phlebotomus Rondani & Berté Mem Inst Oswaldo Cruz 93: Rispail P, Léger N 1998b. Numerical taxonomy of Old World Phlebotominae (Diptera: Psychodidae). 2. Restatement of classification upon subgeneric morphological characters. Mem Inst Oswaldo Cruz 93: Simon C Discriminant analysis of year classes of periodical cicada based on wing morphometric data enhanced by molecular information, p In RG Foottit & JT Sorensen (eds), Ordination in the Study of Morphology, Evolution and Systematics of Insects: Applications and Quantitative Genetic Rationales, Elsevier, New York. Solé-Cava AM, Russo CAM, Araujo ME, Thorpe JP Cladistic and phenetic allozyme data for nine species of sea anemones of the family Actiniidae (Cnidaria: Anthozoa). Biol J Linnean Soc 52: Sorensen JT The use of discriminant function analysis for estimation of phylogeny: partitioning, perspective and problems, p In RG Foottit & JT Sorensen (eds), Ordination in the Study of Morphology, Evolution and Systematics of Insects: Applications and Quantitative Genetic Rationales, Elsevier, New York. Sorensen JT, Foottit RG The evolutionary quantitative genetic rationales for the use of ordination analyses in systematics: phylogenetic implications, p In RG Foottit & JT Sorensen (eds), Ordination in the Study of Morphology, Evolution and Systematics of Insects: Applications and Quantitative Genetic Rationales, Elsevier, New York. Thorpe JP, Solé-Cava AM The use of allozyme electrophoresis in invertebrate systematics. Zoologica scripta 23: Vattier-Bernard G Etude morphologique et biologique des phlébotomes cavernicoles du Congo- Brazzaville. Annal Spél 26: White GB, Killick-Kendrick R Polytene chromosomes of the sandfly Lutzomyia longipalpis and the cytogenetics of Psychodidae in relation to other Diptera. J Entomol A 50: Wilks SS Certain generalizations in the analysis of variance. Biometrika 24: 471. Young DG, Lawyer PG New World vectors of the leishmaniases, p In Kerry F Harris, Current Topics in Vector Research, Vol. 4, Springer-Verlag, New York. Young DG, Duncan MA Guide to the Identification and Geographic Distribution of Lutzomyia Sand Flies in Mexico, the West Indies, Central and South America (Diptera: Psychodidae), Mem Amer Inst Entomol 54, Associate Publishers, Gainesville, 881 pp.
Reconstructing the history of lineages
 Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
PHLEBOTOMINE VECTORS OF HUMAN DISEASE
 C till AD TAXONOMY AND BIOLOGY OF PHLEBOTOMINE VECTORS OF HUMAN DISEASE 11=K REPORT N I O DAMD 17-82-C-2223 D.G. YOUNG DECEMBER 31, 1987 Supported by U. S. ARMY MEDICAL RESEARCH & DEVELOPMENT COMMAND Fort
C till AD TAXONOMY AND BIOLOGY OF PHLEBOTOMINE VECTORS OF HUMAN DISEASE 11=K REPORT N I O DAMD 17-82-C-2223 D.G. YOUNG DECEMBER 31, 1987 Supported by U. S. ARMY MEDICAL RESEARCH & DEVELOPMENT COMMAND Fort
Description of the Fourth Instar Larva of Lutzomyia longipalpis, under Scanning Electron Microscopy
 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 91(5): 571-578, Sep./Oct. 1996 Description of the Fourth Instar Larva of Lutzomyia longipalpis, under Scanning Electron Microscopy Antônio Cesar Rios Leite +,
Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 91(5): 571-578, Sep./Oct. 1996 Description of the Fourth Instar Larva of Lutzomyia longipalpis, under Scanning Electron Microscopy Antônio Cesar Rios Leite +,
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
ESS 345 Ichthyology. Systematic Ichthyology Part II Not in Book
 ESS 345 Ichthyology Systematic Ichthyology Part II Not in Book Thought for today: Now, here, you see, it takes all the running you can do, to keep in the same place. If you want to get somewhere else,
ESS 345 Ichthyology Systematic Ichthyology Part II Not in Book Thought for today: Now, here, you see, it takes all the running you can do, to keep in the same place. If you want to get somewhere else,
Article.
 Zootaxa 3838 (5): 501 517 www.mapress.com/zootaxa/ Copyright 2014 Magnolia Press Article http://dx.doi.org/10.11646/zootaxa.3838.5.1 http://zoobank.org/urn:lsid:zoobank.org:pub:76550bdf-c89d-44cd-9847-170a34d18cde
Zootaxa 3838 (5): 501 517 www.mapress.com/zootaxa/ Copyright 2014 Magnolia Press Article http://dx.doi.org/10.11646/zootaxa.3838.5.1 http://zoobank.org/urn:lsid:zoobank.org:pub:76550bdf-c89d-44cd-9847-170a34d18cde
8/23/2014. Phylogeny and the Tree of Life
 Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
UoN, CAS, DBSC BIOL102 lecture notes by: Dr. Mustafa A. Mansi. The Phylogenetic Systematics (Phylogeny and Systematics)
 - Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
- Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
Relationships of New World Phlebotomine Sand Flies (Diptera: Psychodidae) Based on Fossil Evidence
 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 98(Suppl. I): 145-149, 2003 145 Relationships of New World Phlebotomine Sand Flies (Diptera: Psychodidae) Based on Fossil Evidence José Dilermando Andrade Filho,
Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 98(Suppl. I): 145-149, 2003 145 Relationships of New World Phlebotomine Sand Flies (Diptera: Psychodidae) Based on Fossil Evidence José Dilermando Andrade Filho,
Gustavo Mayr de Lima Carvalho 1 / +, Reginaldo Peçanha Brazil 2, Cristiani de Castilho Sanguinette 1, José Dilermando Andrade Filho 1
 336 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 105(3): 336-340, May 2010 Description of a new phlebotomine species, Martinsmyia reginae sp. nov. (Diptera: Psychodidae: Phlebotominae) from a cave in the
336 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 105(3): 336-340, May 2010 Description of a new phlebotomine species, Martinsmyia reginae sp. nov. (Diptera: Psychodidae: Phlebotominae) from a cave in the
Outline. Classification of Living Things
 Outline Classification of Living Things Chapter 20 Mader: Biology 8th Ed. Taxonomy Binomial System Species Identification Classification Categories Phylogenetic Trees Tracing Phylogeny Cladistic Systematics
Outline Classification of Living Things Chapter 20 Mader: Biology 8th Ed. Taxonomy Binomial System Species Identification Classification Categories Phylogenetic Trees Tracing Phylogeny Cladistic Systematics
PHYLOGENY AND SYSTEMATICS
 AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
Chapter 27: Evolutionary Genetics
 Chapter 27: Evolutionary Genetics Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand what the term species means to biology. 2. Recognize the various patterns
Chapter 27: Evolutionary Genetics Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand what the term species means to biology. 2. Recognize the various patterns
Classification, Phylogeny yand Evolutionary History
 Classification, Phylogeny yand Evolutionary History The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize
Classification, Phylogeny yand Evolutionary History The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize
Morphometric Analysis of Longipalpis (Diptera: Psychodidae) Complex Populations in Mato Grosso do Sul, Brazil
 Journal of Medical Entomology Advance Access published March 29, 2015 MORPHOLOGY, SYSTEMATICS, EVOLUTION Morphometric Analysis of Longipalpis (Diptera: Psychodidae) Complex Populations in Mato Grosso do
Journal of Medical Entomology Advance Access published March 29, 2015 MORPHOLOGY, SYSTEMATICS, EVOLUTION Morphometric Analysis of Longipalpis (Diptera: Psychodidae) Complex Populations in Mato Grosso do
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Chapter 26: Phylogeny and the Tree of Life Phylogenies Show Evolutionary Relationships
 Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Geography of Evolution
 Geography of Evolution Biogeography - the study of the geographic distribution of organisms. The current distribution of organisms can be explained by historical events and current climatic patterns. Darwin
Geography of Evolution Biogeography - the study of the geographic distribution of organisms. The current distribution of organisms can be explained by historical events and current climatic patterns. Darwin
Lecture 11 Friday, October 21, 2011
 Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
CHAPTER 10 Taxonomy and Phylogeny of Animals
 CHAPTER 10 Taxonomy and Phylogeny of Animals 10-1 10-2 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Linnaeus and Taxonomy More than 1.5 million species of
CHAPTER 10 Taxonomy and Phylogeny of Animals 10-1 10-2 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Linnaeus and Taxonomy More than 1.5 million species of
Modern Evolutionary Classification. Section 18-2 pgs
 Modern Evolutionary Classification Section 18-2 pgs 451-455 Modern Evolutionary Classification In a sense, organisms determine who belongs to their species by choosing with whom they will mate. Taxonomic
Modern Evolutionary Classification Section 18-2 pgs 451-455 Modern Evolutionary Classification In a sense, organisms determine who belongs to their species by choosing with whom they will mate. Taxonomic
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
The Life System and Environmental & Evolutionary Biology II
 The Life System and Environmental & Evolutionary Biology II EESC V2300y / ENVB W2002y Laboratory 1 (01/28/03) Systematics and Taxonomy 1 SYNOPSIS In this lab we will give an overview of the methodology
The Life System and Environmental & Evolutionary Biology II EESC V2300y / ENVB W2002y Laboratory 1 (01/28/03) Systematics and Taxonomy 1 SYNOPSIS In this lab we will give an overview of the methodology
Lecture V Phylogeny and Systematics Dr. Kopeny
 Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
Wing Geometry as a Tool for Studying the Lutzomyia longipalpis (Diptera: Psychodidae) Complex
 Mem Inst Oswaldo Cruz, io de Janeiro, Vol. 96(8): 1089-1094, ovember 2001 Wing Geometry as a ool for Studying the Lutzomyia longipalpis (Diptera: Psychodidae) Complex J De la iva/ +, F Le Pont*, V Ali**,
Mem Inst Oswaldo Cruz, io de Janeiro, Vol. 96(8): 1089-1094, ovember 2001 Wing Geometry as a ool for Studying the Lutzomyia longipalpis (Diptera: Psychodidae) Complex J De la iva/ +, F Le Pont*, V Ali**,
What do we mean by a species? Morphological species concept. Morphological species concept BIOL2007 SPECIES AND BIODIVERSITY. Kanchon Dasmahapatra
 BIOL2007 SPECIES AND BIODIVERSITY Kanchon Dasmahapatra What are species? How do species differ from each other? Biodiversity: How many species are there? What do we mean by a species? Darwin proved species
BIOL2007 SPECIES AND BIODIVERSITY Kanchon Dasmahapatra What are species? How do species differ from each other? Biodiversity: How many species are there? What do we mean by a species? Darwin proved species
Name. Ecology & Evolutionary Biology 2245/2245W Exam 2 1 March 2014
 Name 1 Ecology & Evolutionary Biology 2245/2245W Exam 2 1 March 2014 1. Use the following matrix of nucleotide sequence data and the corresponding tree to answer questions a. through h. below. (16 points)
Name 1 Ecology & Evolutionary Biology 2245/2245W Exam 2 1 March 2014 1. Use the following matrix of nucleotide sequence data and the corresponding tree to answer questions a. through h. below. (16 points)
DNA Sequencing as a Method for Larval Identification in Odonates
 DNA Sequencing as a Method for Larval Identification in Odonates Adeline Harris 121 North St Apt 3 Farmington, ME 04938 Adeline.harris@maine.edu Christopher Stevens 147 Main St Apt 1 Farmington, ME 04938
DNA Sequencing as a Method for Larval Identification in Odonates Adeline Harris 121 North St Apt 3 Farmington, ME 04938 Adeline.harris@maine.edu Christopher Stevens 147 Main St Apt 1 Farmington, ME 04938
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
Historical Biogeography. Historical Biogeography. Systematics
 Historical Biogeography I. Definitions II. Fossils: problems with fossil record why fossils are important III. Phylogeny IV. Phenetics VI. Phylogenetic Classification Disjunctions debunked: Examples VII.
Historical Biogeography I. Definitions II. Fossils: problems with fossil record why fossils are important III. Phylogeny IV. Phenetics VI. Phylogenetic Classification Disjunctions debunked: Examples VII.
Bio 2 Plant and Animal Biology
 Bio 2 Plant and Animal Biology Evolution Evolution as the explanation for life s unity and diversity Darwinian Revolution Two main Points Descent with Modification Natural Selection Biological Species
Bio 2 Plant and Animal Biology Evolution Evolution as the explanation for life s unity and diversity Darwinian Revolution Two main Points Descent with Modification Natural Selection Biological Species
Molecular Markers, Natural History, and Evolution
 Molecular Markers, Natural History, and Evolution Second Edition JOHN C. AVISE University of Georgia Sinauer Associates, Inc. Publishers Sunderland, Massachusetts Contents PART I Background CHAPTER 1:
Molecular Markers, Natural History, and Evolution Second Edition JOHN C. AVISE University of Georgia Sinauer Associates, Inc. Publishers Sunderland, Massachusetts Contents PART I Background CHAPTER 1:
SPECIATION. REPRODUCTIVE BARRIERS PREZYGOTIC: Barriers that prevent fertilization. Habitat isolation Populations can t get together
 SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
A Phylogenetic Network Construction due to Constrained Recombination
 A Phylogenetic Network Construction due to Constrained Recombination Mohd. Abdul Hai Zahid Research Scholar Research Supervisors: Dr. R.C. Joshi Dr. Ankush Mittal Department of Electronics and Computer
A Phylogenetic Network Construction due to Constrained Recombination Mohd. Abdul Hai Zahid Research Scholar Research Supervisors: Dr. R.C. Joshi Dr. Ankush Mittal Department of Electronics and Computer
CLASSIFICATION AND EVOLUTION OF CAMINALCULES:
 CLASSIFICATION AND EVOLUTION OF CAMINALCULES: One of the main goals of the lab is to illustrate the intimate connection between the classification of living species and their evolutionary relationships.
CLASSIFICATION AND EVOLUTION OF CAMINALCULES: One of the main goals of the lab is to illustrate the intimate connection between the classification of living species and their evolutionary relationships.
Vertebrate Biogeography and Evolution
 Vertebrate Biogeography and Evolution Phylogeny, Plate Tectonics, and Climate Less Digitigrady More Location 1 Location 2 Location 3 Location 4 Biogeography The study of the distribution of species, organisms,
Vertebrate Biogeography and Evolution Phylogeny, Plate Tectonics, and Climate Less Digitigrady More Location 1 Location 2 Location 3 Location 4 Biogeography The study of the distribution of species, organisms,
Early theories: Joseph Hooker (1853) vs. Charles Darwin (1859)
 Gondwanan Plants of the Sydney Region Presentation Dr Peter Weston 25/11/2017 Honorary Research Associate, Science and Conservation Branch, Royal Botanic Gardens and Domain Trust Summary: Dr Marilyn Cross,
Gondwanan Plants of the Sydney Region Presentation Dr Peter Weston 25/11/2017 Honorary Research Associate, Science and Conservation Branch, Royal Botanic Gardens and Domain Trust Summary: Dr Marilyn Cross,
Classification and Phylogeny
 Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
The practice of naming and classifying organisms is called taxonomy.
 Chapter 18 Key Idea: Biologists use taxonomic systems to organize their knowledge of organisms. These systems attempt to provide consistent ways to name and categorize organisms. The practice of naming
Chapter 18 Key Idea: Biologists use taxonomic systems to organize their knowledge of organisms. These systems attempt to provide consistent ways to name and categorize organisms. The practice of naming
Biology 2. Lecture Material. For. Macroevolution. Systematics
 Biology 2 Macroevolution & Systematics 1 Biology 2 Lecture Material For Macroevolution & Systematics Biology 2 Macroevolution & Systematics 2 Microevolution: Biological Species: Two Patterns of Evolutionary
Biology 2 Macroevolution & Systematics 1 Biology 2 Lecture Material For Macroevolution & Systematics Biology 2 Macroevolution & Systematics 2 Microevolution: Biological Species: Two Patterns of Evolutionary
Research Article. Article History : Received 05 June 2013, Accepted 27 July 2013
 International Journal for Life Sciences and Educational Research Vol.1(2), pp. 91-95, July- 2013 Available online at http://www.ijlser.com E-ISSN : 2321-1229; P ISSN : 2321-1180 Research Article Geometric
International Journal for Life Sciences and Educational Research Vol.1(2), pp. 91-95, July- 2013 Available online at http://www.ijlser.com E-ISSN : 2321-1229; P ISSN : 2321-1180 Research Article Geometric
How should we organize the diversity of animal life?
 How should we organize the diversity of animal life? The difference between Taxonomy Linneaus, and Cladistics Darwin What are phylogenies? How do we read them? How do we estimate them? Classification (Taxonomy)
How should we organize the diversity of animal life? The difference between Taxonomy Linneaus, and Cladistics Darwin What are phylogenies? How do we read them? How do we estimate them? Classification (Taxonomy)
Applications of Genetics to Conservation Biology
 Applications of Genetics to Conservation Biology Molecular Taxonomy Populations, Gene Flow, Phylogeography Relatedness - Kinship, Paternity, Individual ID Conservation Biology Population biology Physiology
Applications of Genetics to Conservation Biology Molecular Taxonomy Populations, Gene Flow, Phylogeography Relatedness - Kinship, Paternity, Individual ID Conservation Biology Population biology Physiology
Classification and Phylogeny
 Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS
 ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS JAMES S. FARRIS Museum of Zoology, The University of Michigan, Ann Arbor Accepted March 30, 1966 The concept of conservatism
ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS JAMES S. FARRIS Museum of Zoology, The University of Michigan, Ann Arbor Accepted March 30, 1966 The concept of conservatism
Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them?
 Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them? Carolus Linneaus:Systema Naturae (1735) Swedish botanist &
Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them? Carolus Linneaus:Systema Naturae (1735) Swedish botanist &
Phylogeny 9/8/2014. Evolutionary Relationships. Data Supporting Phylogeny. Chapter 26
 Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Philippe Rispail/ +, Nicole Léger*
 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 93(6): 773-785, Nov./Dec. 1998 Numerical Taxonomy of Old World Phlebotominae (Diptera: Psychodidae). 1. Considerations of Morphological Characters in the Genus
Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 93(6): 773-785, Nov./Dec. 1998 Numerical Taxonomy of Old World Phlebotominae (Diptera: Psychodidae). 1. Considerations of Morphological Characters in the Genus
Integrating Fossils into Phylogenies. Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
 IB 200B Principals of Phylogenetic Systematics Spring 2011 Integrating Fossils into Phylogenies Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
IB 200B Principals of Phylogenetic Systematics Spring 2011 Integrating Fossils into Phylogenies Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
1/17/2012. Class Aves. Avian Systematics. Avian Systematics. Subclass Sauriurae
 Systematics deals with evolutionary relationships among organisms. Allied with classification (or taxonomy). All birds are classified within the single Class Aves 2 Subclasses 4 Infraclasses Class Aves
Systematics deals with evolutionary relationships among organisms. Allied with classification (or taxonomy). All birds are classified within the single Class Aves 2 Subclasses 4 Infraclasses Class Aves
Species diversification in space: biogeographic patterns
 Species diversification in space: biogeographic patterns Outline Endemism and cosmopolitanism Disjunctions Biogeographic regions Barriers and interchanges Divergence and convergence Biogeographic patterns
Species diversification in space: biogeographic patterns Outline Endemism and cosmopolitanism Disjunctions Biogeographic regions Barriers and interchanges Divergence and convergence Biogeographic patterns
Systematics - BIO 615
 A Knyght ther was, and that a worthy man, A Knyght ther was, and he was a worthy man, A Knyght ther was, and that a worthy man, A Knyght ther was, and he was a worthy man, A Knyght ther was, and he wasn
A Knyght ther was, and that a worthy man, A Knyght ther was, and he was a worthy man, A Knyght ther was, and that a worthy man, A Knyght ther was, and he was a worthy man, A Knyght ther was, and he wasn
Chapter 16: Reconstructing and Using Phylogenies
 Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/65602 holds various files of this Leiden University dissertation. Author: Ruchisansakun, S. Title: Balsaminaceae in Southeast Asia: systematics, evolution,
Cover Page The handle http://hdl.handle.net/1887/65602 holds various files of this Leiden University dissertation. Author: Ruchisansakun, S. Title: Balsaminaceae in Southeast Asia: systematics, evolution,
Chapter 26. Phylogeny and the Tree of Life. Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Pearson Education, Inc.
 Chapter 26 Phylogeny and the Tree of Life Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Investigating the Tree of Life Phylogeny is the evolutionary history of a species or group of
Chapter 26 Phylogeny and the Tree of Life Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Investigating the Tree of Life Phylogeny is the evolutionary history of a species or group of
Need for systematics. Applications of systematics. Linnaeus plus Darwin. Approaches in systematics. Principles of cladistics
 Topics Need for systematics Applications of systematics Linnaeus plus Darwin Approaches in systematics Principles of cladistics Systematics pp. 474-475. Systematics - Study of diversity and evolutionary
Topics Need for systematics Applications of systematics Linnaeus plus Darwin Approaches in systematics Principles of cladistics Systematics pp. 474-475. Systematics - Study of diversity and evolutionary
2 Earth s Changing Continents
 CHAPTER 9 SECTION The History of Life on Earth 2 Earth s Changing Continents California Science Standards 7.4.a, 7.4.e, 7.4.f BEFORE YOU READ After you read this section, you should be able to answer these
CHAPTER 9 SECTION The History of Life on Earth 2 Earth s Changing Continents California Science Standards 7.4.a, 7.4.e, 7.4.f BEFORE YOU READ After you read this section, you should be able to answer these
Simone M da Costa/ +, Michela Cechinel *, Valdenir Bandeira *, José C Zannuncio **, Ralph Lainson ***, Elizabeth F Rangel
 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 102(2): 149-153, March 2007 149 Lutzomyia (Nyssomyia) whitmani s.l. (Antunes & Coutinho, 1939) (Diptera: Psychodidae: Phlebotominae): geographical distribution
Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 102(2): 149-153, March 2007 149 Lutzomyia (Nyssomyia) whitmani s.l. (Antunes & Coutinho, 1939) (Diptera: Psychodidae: Phlebotominae): geographical distribution
Evolution Problem Drill 09: The Tree of Life
 Evolution Problem Drill 09: The Tree of Life Question No. 1 of 10 Question 1. The age of the Earth is estimated to be about 4.0 to 4.5 billion years old. All of the following methods may be used to estimate
Evolution Problem Drill 09: The Tree of Life Question No. 1 of 10 Question 1. The age of the Earth is estimated to be about 4.0 to 4.5 billion years old. All of the following methods may be used to estimate
Plant of the Day Isoetes andicola
 Plant of the Day Isoetes andicola Endemic to central and southern Peru Found in scattered populations above 4000 m Restricted to the edges of bogs and lakes Leaves lack stomata and so CO 2 is obtained,
Plant of the Day Isoetes andicola Endemic to central and southern Peru Found in scattered populations above 4000 m Restricted to the edges of bogs and lakes Leaves lack stomata and so CO 2 is obtained,
PHYLOGENY & THE TREE OF LIFE
 PHYLOGENY & THE TREE OF LIFE PREFACE In this powerpoint we learn how biologists distinguish and categorize the millions of species on earth. Early we looked at the process of evolution here we look at
PHYLOGENY & THE TREE OF LIFE PREFACE In this powerpoint we learn how biologists distinguish and categorize the millions of species on earth. Early we looked at the process of evolution here we look at
Chapter 19: Taxonomy, Systematics, and Phylogeny
 Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Supplementary Information
 Supplementary Information For the article"comparable system-level organization of Archaea and ukaryotes" by J. Podani, Z. N. Oltvai, H. Jeong, B. Tombor, A.-L. Barabási, and. Szathmáry (reference numbers
Supplementary Information For the article"comparable system-level organization of Archaea and ukaryotes" by J. Podani, Z. N. Oltvai, H. Jeong, B. Tombor, A.-L. Barabási, and. Szathmáry (reference numbers
Ph ylogeography. A guide to the study of the spatial distribution of Seahorses. By Leila Mougoui Bakhtiari
 Ph ylogeography A guide to the study of the spatial distribution of Seahorses By Leila Mougoui Bakhtiari Contents An Introduction to Phylogeography JT Bohem s Resarch Map of erectu s migration Conservation
Ph ylogeography A guide to the study of the spatial distribution of Seahorses By Leila Mougoui Bakhtiari Contents An Introduction to Phylogeography JT Bohem s Resarch Map of erectu s migration Conservation
3 Hours 18 / 06 / 2012 EXAMS OFFICE USE ONLY University of the Witwatersrand, Johannesburg Course or topic No(s) ANAT 4000
 3 Hours 18 / 06 / 2012 EXAMS OFFICE USE ONLY University of the Witwatersrand, Johannesburg Course or topic No(s) ANAT 4000 Course or topic name(s) Paper Number & title HUMAN BIOLOGY HONOURS: PAPER 1 Examination
3 Hours 18 / 06 / 2012 EXAMS OFFICE USE ONLY University of the Witwatersrand, Johannesburg Course or topic No(s) ANAT 4000 Course or topic name(s) Paper Number & title HUMAN BIOLOGY HONOURS: PAPER 1 Examination
Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food.
 Prokaryotic Cell Eukaryotic Cell Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food. Heterotrophs must get energy by eating autotrophs or other heterotrophs.
Prokaryotic Cell Eukaryotic Cell Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food. Heterotrophs must get energy by eating autotrophs or other heterotrophs.
SCIENTIFIC EVIDENCE TO SUPPORT THE THEORY OF EVOLUTION. Using Anatomy, Embryology, Biochemistry, and Paleontology
 SCIENTIFIC EVIDENCE TO SUPPORT THE THEORY OF EVOLUTION Using Anatomy, Embryology, Biochemistry, and Paleontology Scientific Fields Different fields of science have contributed evidence for the theory of
SCIENTIFIC EVIDENCE TO SUPPORT THE THEORY OF EVOLUTION Using Anatomy, Embryology, Biochemistry, and Paleontology Scientific Fields Different fields of science have contributed evidence for the theory of
B. Phylogeny and Systematics:
 Tracing Phylogeny A. Fossils: Some fossils form as is weathered and eroded from the land and carried by rivers to seas and where the particles settle to the bottom. Deposits pile up and the older sediments
Tracing Phylogeny A. Fossils: Some fossils form as is weathered and eroded from the land and carried by rivers to seas and where the particles settle to the bottom. Deposits pile up and the older sediments
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Phylogeography and genetic differentiation between Loxigilla noctis and L. barbadensis in the Lesser Antilles
 Phylogeography and genetic differentiation between Loxigilla noctis and L. barbadensis in the Lesser Antilles Sophie Arnaud-Haond 1, Carla Daniel 2, Sébastien Motreuil 3, Julia Horrocks 2 & Frank Cézilly
Phylogeography and genetic differentiation between Loxigilla noctis and L. barbadensis in the Lesser Antilles Sophie Arnaud-Haond 1, Carla Daniel 2, Sébastien Motreuil 3, Julia Horrocks 2 & Frank Cézilly
AP Biology. Cladistics
 Cladistics Kingdom Summary Review slide Review slide Classification Old 5 Kingdom system Eukaryote Monera, Protists, Plants, Fungi, Animals New 3 Domain system reflects a greater understanding of evolution
Cladistics Kingdom Summary Review slide Review slide Classification Old 5 Kingdom system Eukaryote Monera, Protists, Plants, Fungi, Animals New 3 Domain system reflects a greater understanding of evolution
How Biological Diversity Evolves
 CHAPTER 14 How Biological Diversity Evolves PowerPoint Lectures for Essential Biology, Third Edition Neil Campbell, Jane Reece, and Eric Simon Essential Biology with Physiology, Second Edition Neil Campbell,
CHAPTER 14 How Biological Diversity Evolves PowerPoint Lectures for Essential Biology, Third Edition Neil Campbell, Jane Reece, and Eric Simon Essential Biology with Physiology, Second Edition Neil Campbell,
Mechanisms of Evolution Darwinian Evolution
 Mechanisms of Evolution Darwinian Evolution Descent with modification by means of natural selection All life has descended from a common ancestor The mechanism of modification is natural selection Concept
Mechanisms of Evolution Darwinian Evolution Descent with modification by means of natural selection All life has descended from a common ancestor The mechanism of modification is natural selection Concept
A Multivariate Study of Variation in Two Species of Rock Crab of the Genus Leptograpsus
 Awt. J. Zool. 1974, 22,417-25 A Multivariate Study of Variation in Two Species of Rock Crab of the Genus Leptograpsus N. A. ~arn~bell* and R. J. MahonB A Division of Mathematical Statistics, CSIRO, Floreat
Awt. J. Zool. 1974, 22,417-25 A Multivariate Study of Variation in Two Species of Rock Crab of the Genus Leptograpsus N. A. ~arn~bell* and R. J. MahonB A Division of Mathematical Statistics, CSIRO, Floreat
Evidence of Evolution
 Evidence of Evolution There is a gigantic body of evidence supporting evolution. Six major areas of study contribute to that body of evidence: 1. The Fossil Record 2. Comparative Anatomy 3. Comparative
Evidence of Evolution There is a gigantic body of evidence supporting evolution. Six major areas of study contribute to that body of evidence: 1. The Fossil Record 2. Comparative Anatomy 3. Comparative
Name: Class: Date: ID: A
 Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
Speciation. Today s OUTLINE: Mechanisms of Speciation. Mechanisms of Speciation. Geographic Models of speciation. (1) Mechanisms of Speciation
 Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny
 CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny To trace phylogeny or the evolutionary history of life, biologists use evidence from paleontology, molecular data, comparative
CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny To trace phylogeny or the evolutionary history of life, biologists use evidence from paleontology, molecular data, comparative
Phylogeny and systematics. Why are these disciplines important in evolutionary biology and how are they related to each other?
 Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Biology 211 (2) Week 1 KEY!
 Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Speciation. Today s OUTLINE: Mechanisms of Speciation. Mechanisms of Speciation. Geographic Models of speciation. (1) Mechanisms of Speciation
 Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Speciation. Today s OUTLINE: Mechanisms of Speciation. Mechanisms of Speciation. Geographic Models of speciation. (1) Mechanisms of Speciation
 Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Evidence of Evolution *
 OpenStax-CNX module: m45491 1 Evidence of Evolution * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
OpenStax-CNX module: m45491 1 Evidence of Evolution * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will
Macroevolution Part I: Phylogenies
 Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
What is the purpose of the Classifying System? To allow the accurate identification of a particular organism
 What is the purpose of the Classifying System? To allow the accurate identification of a particular organism Taxonomy The practice of classifying organisms -Taxonomy was founded nearly 300 years ago by
What is the purpose of the Classifying System? To allow the accurate identification of a particular organism Taxonomy The practice of classifying organisms -Taxonomy was founded nearly 300 years ago by
Chapter 19 Organizing Information About Species: Taxonomy and Cladistics
 Chapter 19 Organizing Information About Species: Taxonomy and Cladistics An unexpected family tree. What are the evolutionary relationships among a human, a mushroom, and a tulip? Molecular systematics
Chapter 19 Organizing Information About Species: Taxonomy and Cladistics An unexpected family tree. What are the evolutionary relationships among a human, a mushroom, and a tulip? Molecular systematics
BIOL 428: Introduction to Systematics Midterm Exam
 Midterm exam page 1 BIOL 428: Introduction to Systematics Midterm Exam Please, write your name on each page! The exam is worth 150 points. Verify that you have all 8 pages. Read the questions carefully,
Midterm exam page 1 BIOL 428: Introduction to Systematics Midterm Exam Please, write your name on each page! The exam is worth 150 points. Verify that you have all 8 pages. Read the questions carefully,
A. Incorrect! In the binomial naming convention the Kingdom is not part of the name.
 Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Concepts and Methods in Molecular Divergence Time Estimation
 Concepts and Methods in Molecular Divergence Time Estimation 26 November 2012 Prashant P. Sharma American Museum of Natural History Overview 1. Why do we date trees? 2. The molecular clock 3. Local clocks
Concepts and Methods in Molecular Divergence Time Estimation 26 November 2012 Prashant P. Sharma American Museum of Natural History Overview 1. Why do we date trees? 2. The molecular clock 3. Local clocks
MOLECULAR PHYLOGENY AND GENETIC DIVERSITY ANALYSIS. Masatoshi Nei"
 MOLECULAR PHYLOGENY AND GENETIC DIVERSITY ANALYSIS Masatoshi Nei" Abstract: Phylogenetic trees: Recent advances in statistical methods for phylogenetic reconstruction and genetic diversity analysis were
MOLECULAR PHYLOGENY AND GENETIC DIVERSITY ANALYSIS Masatoshi Nei" Abstract: Phylogenetic trees: Recent advances in statistical methods for phylogenetic reconstruction and genetic diversity analysis were
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
--Therefore, congruence among all postulated homologies provides a test of any single character in question [the central epistemological advance].
![--Therefore, congruence among all postulated homologies provides a test of any single character in question [the central epistemological advance]. --Therefore, congruence among all postulated homologies provides a test of any single character in question [the central epistemological advance].](/thumbs/86/93197257.jpg) Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2008 University of California, Berkeley B.D. Mishler Jan. 29, 2008. The Hennig Principle: Homology, Synapomorphy, Rooting issues The fundamental
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2008 University of California, Berkeley B.D. Mishler Jan. 29, 2008. The Hennig Principle: Homology, Synapomorphy, Rooting issues The fundamental
Homework Assignment, Evolutionary Systems Biology, Spring Homework Part I: Phylogenetics:
 Homework Assignment, Evolutionary Systems Biology, Spring 2009. Homework Part I: Phylogenetics: Introduction. The objective of this assignment is to understand the basics of phylogenetic relationships
Homework Assignment, Evolutionary Systems Biology, Spring 2009. Homework Part I: Phylogenetics: Introduction. The objective of this assignment is to understand the basics of phylogenetic relationships
Chapters 25 and 26. Searching for Homology. Phylogeny
 Chapters 25 and 26 The Origin of Life as we know it. Phylogeny traces evolutionary history of taxa Systematics- analyzes relationships (modern and past) of organisms Figure 25.1 A gallery of fossils The
Chapters 25 and 26 The Origin of Life as we know it. Phylogeny traces evolutionary history of taxa Systematics- analyzes relationships (modern and past) of organisms Figure 25.1 A gallery of fossils The
Lab 06 Phylogenetics, part 1
 Lab 06 Phylogenetics, part 1 phylogeny is a visual representation of a hypothesis about the relationships among a set of organisms. Phylogenetics is the study of phylogenies and and their development.
Lab 06 Phylogenetics, part 1 phylogeny is a visual representation of a hypothesis about the relationships among a set of organisms. Phylogenetics is the study of phylogenies and and their development.
PRINCIPLES OF MENDELIAN GENETICS APPLICABLE IN FORESTRY. by Erich Steiner 1/
 PRINCIPLES OF MENDELIAN GENETICS APPLICABLE IN FORESTRY by Erich Steiner 1/ It is well known that the variation exhibited by living things has two components, one hereditary, the other environmental. One
PRINCIPLES OF MENDELIAN GENETICS APPLICABLE IN FORESTRY by Erich Steiner 1/ It is well known that the variation exhibited by living things has two components, one hereditary, the other environmental. One
ELE4120 Bioinformatics Tutorial 8
 ELE4120 ioinformatics Tutorial 8 ontent lassifying Organisms Systematics and Speciation Taxonomy and phylogenetics Phenetics versus cladistics Phylogenetic trees iological classification Goal: To develop
ELE4120 ioinformatics Tutorial 8 ontent lassifying Organisms Systematics and Speciation Taxonomy and phylogenetics Phenetics versus cladistics Phylogenetic trees iological classification Goal: To develop
Phylogenetic Analysis
 Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis
 Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
