Genome-wide Analysis of Functions Regulated by Sets of Transcription Factors
|
|
|
- Alaina Booth
- 5 years ago
- Views:
Transcription
1 Genome-wide Analysis of Functions Regulated by Sets of Transcription Factors Szymon M. Kiełbasa 1, Nils Blüthgen, Hanspeter Herzel Humboldt University, Institute for Theoretical Biology, Invalidenstraße 43, Berlin, Germany 1 s.kielbasa@itb.biologie.hu-berlin.de both authors contributed equally Abstract: We present a pipeline for inferring biological functions regulated by a combinatorial interaction of transcription factors. Using a robust statistical method the pipeline intersects the presence of transcription factor binding sites in gene upstream sequences with Gene Ontology terms associated with these genes. Positional frequency matrices for the transcription factors constitute the input of the pipeline and significantly enriched biological processes are reported as the output. We demonstrate the usage of the pipeline using two groups of transcription factors: a cell-cycle related family of E2F factors and a NFAT/AP-1 pair involved in immune response. In both cases the reported results match well the experimental knowledge. Furthermore, for the NFAT/AP-1 composite element novel functions are predicted. INTRODUCTION Transcriptional regulation of gene expression is a main mechanism governing the temporal and spatial organization of biological processes in eukaryotic organisms. External and internal stimuli are transduced into activities of transcription factors controlling major biological processes within the cell. Single factors controlling expression in prokaryotic cells evolved into a sophisticated regulatory machinery in eukaryotes, involving the combinatorial action of transcription factors. This phenomenon gives rise to enormously complex responses equipping cells with means to respond adequately to surrounding information [FMZ + 02]. Considering the importance of transcriptional regulation for most biological processes, understanding of the regulatory network is a major challenge in the postgenomic era. Automated inference of the regulatory network could result in predictive models of interactions among genes, that allow a better understanding of gene expression patterns and could hint to targets for drug design. The fundamental building block of the regulatory network is the binding of transcription factors to regulatory cis-sequences located in the promoter regions of genes. Experimentally identified cis-sequences for a single transcription factor can be aligned to find the sequence recognized by the factor. Alignments of such cis-sequences yield positional frequency matrices, which are further used to model the binding affinity of the factor towards
2 DNA sequences [St98]. An in-vitro method, SELEX-SAGE, allows to discover potential binding sites for transcription factors in vitro and to build positional frequency matrices without knowing their target genes. For numerous transcription factors corresponding frequency matrices have been constructed (e.g. [HWR + 98]) for the prediction of transcriptional regulation. Despite advances in this field genome-wide scans of binding sites are difficult to interpret since false, nonfunctional predictions dominate. [WK03] estimate that a simple search using only a single frequency matrix results in only one functional site per 1000 predictions. More recent solutions of this problem take into account further biological properties, like clustering of the functional binding sites [FLW03] and possible conservation of cis-sequences in evolution [DWR + 03]. These approaches can reduce the number of non-functional predictions by about two orders of magnitude [WK03]. There are attempts [WF98, KW01, KMIWK02] to use binding sites predicted in a gene upstream sequence to predict a function of the gene under consideration. This approach originates from the concept that genes sharing the same regulatory elements carry out also the same biological function. Thus it is limited to extend functional families with already well known regulatory properties. New functions of transcription factors (possibly in another cellular context, stage of development, tissue, etc.) are unlikely to be inferred. Utilizing the growing systematic functional annotation provided by the Gene Ontology [ABB + 00] we propose a novel, more function oriented, genome-wide analysis to predict the biological function of transcription factors. Our approach starts from transcription factor frequency matrices and predicts corresponding target genes using established methods. As stated above, many of these predictions are likely to be false. We assume that these false targets are scattered randomly over the biological processes. Contrarily, the true functional predictions are likely to be involved in a few, rather specific biological processes. Consequently we associate significantly enriched biological processes in the predicted target genes as biological functions of the transcription factors under consideration. This approach allows to discover novel regulatory functions without bias towards known functions of a transcription factor. In the following we discuss the details of our prediction pipeline and apply our pipeline to a single transcription factor family (E2F) and to a set containing the transcription factors NFAT and AP-1. MATERIALS AND METHODS Prediction Pipeline We have constructed a pipeline to predict biological functions controlled by a set of transcription factors, as shown in Fig. 1. A set of positional frequency matrices is used in the Cluster-Buster program [FLW03] to predict clusters of binding sites in upstream regions of human genes. We assume, that genes having such clusters in their upstream sequences are potentially regulated by the factors. These genes are annotated with terms describing biological processes from the Gene Ontology [ABB + 00]. Subsequently we identify biological processes which are significantly associated with the potential target genes with
3 GOSSIP [BBC + 04]. Details of these steps of the pipeline are discussed below. binding sites for a set of transcription factors described by weight matrices Gene Ontology DNA sequences upstream of gene-coding regions as annotated in Ensembl predicting regulation with Cluster-Buster biological profiling with GOSSIP predicted biological processes regulated by the set of transcription factors Figure 1: Data processing in our pipeline: 1,000 bp upstream regions of 16,032 human genes predicted by Ensembl were extracted (utilizing HomGL [BKCH04]) giving a collection of 15,362 potential promoter regions. These sequences together with positional frequency matrices of transcription factors binding sites were analyzed by Cluster-Buster to find a subset of genes which are potentially regulated by these factors. Subsequently, GOSSIP [BBC + 04] utilizes the Gene Ontology to find biological processes associated with this subset. Finally, significant processes are mapped to the hierarchy of the Gene Ontology graph. Positional frequency matrices As an example of our method, we have studied two sets of transcription factors. The first set contains only a single positional frequency matrix corresponding to the E2F family of transcription factors. It is taken from the Transfac database [HWR + 98] with the accession number M00427 (V$E2F Q6). As a second set we use the matrices describing sites recognized by AP-1 and NF-AT, as published by Kel et al. [KKMBW99]. Upstream Regions The human part of the UniGene database partitions human transcript sequences into 118,517 UniGene clusters derived from both known genes and ESTs. We could map 16,032 of the clusters to genes reported by the Ensembl database [CEA + 04] by using HomGL [BKCH04]. For these genes we extracted 1,000 bp long DNA sequence upstream of the predicted transcription start sites of the genes. We found, that only 15,362 of these sequences were unique, and we chose them as predicted promoter regions, since the multiple occurrence of the same promoter sequence could bias the further analysis of enriched processes.
4 Predicting genes regulated by transcription factors Computational prediction of genes regulated by a single transcription factor results in many false positives. Eukaryotic regulation is typically achieved by several regulators, and predictions of clusters of several binding sites for one or more transcription factors may reduce the rate of false predictions by more than an order of magnitude. Therefore we assume here, that a gene is regulated by factors under study if a cluster of several binding sites corresponding to some of the factors is predicted in the gene upstream region. We selected the publicly available Cluster-Buster program [FLW03] to perform the task of cluster searching. The program takes all gene upstream sequences and a set of matrices as input and returns a list of probable locations of functional clusters in these sequences. To keep our pipeline simple we used the Cluster-Buster program with all parameters set to their default values. Functional profiling The Gene Ontology [ABB + 00] specifies a controlled, hierarchical vocabulary for annotating genes. It can be represented by a directed acyclic graph. We utilize the Gene Ontology (GO) to test whether a biological process is significantly associated with a group of genes predicted to be regulated by the set of transcription factors under consideration. This analysis requires four data sources: The group of potentially regulated genes as the test group, all genes with upstream regions as a reference group, GO annotations for these both gene sets, and the Gene Ontology. We annotate the groups of genes with HomGL [BKCH04]. Annotations are usually given as terms close to the leafs of the graph implying a series of more general annotations upward in the GO graph. For each term in the ontology we test whether this particular term is enriched in the test group as compared to the reference group. To do this we categorize each gene in two distinct ways: first, whether it is annotated with the term under consideration or not, and second, whether it belongs to the test group or not. We apply Fisher s exact test that allows to detect and quantify associations between the two distinct categorizations. But since the number of terms in the Gene Ontology is very high, problems arising from multiple testing cannot be left aside. We adjust the p-values to control the number of false positives by calculating the false discovery rate (FDR), which quantifies the ratio of the expected number of false positives among the positives. For this specific problem it can be calculated exactly [BBC + 04]. Assuming, that incorrectly predicted target genes by the prior step of our pipeline do not favor any processes but scatter randomly, the FDR of our pipeline predictions is controlled reliably. In the further analysis, we apply a threshold of F DR 0.05 that keeps the expected ratio of false predictions under five percent. The analysis is been performed by the software package GOSSIP [BBC + 04].
5 Gene_Ontology biological_process cellular process physiological process cell growth and/or maintenance metabolism cell proliferation cell cycle cell organization and biogenesis nuclear organization and biogenesis nucleobase, nucleoside, nucleotide and nucleic acid metabolism DNA metabolism regulation of cell cycle mitotic cell cycle chromosome organization and biogenesis (sensu Eukarya) DNA packaging DNA replication and chromosome cycle S phase of mitotic cell cycle establishment and/or maintenance of chromatin architecture DNA replication chromatin assembly/disassembly DNA dependent DNA replication nucleosome assembly DNA replication initiation Figure 2: A part of the Gene Ontology highlighting the biological processes that are significantly enriched in 273 genes with a cluster of E2F-binding sites predicted in their upstream regions. Black boxes show those processes with F DR 0.01, gray boxes with F DR 0.05 (see Methods for details). RESULTS Processes regulated by E2F E2F s are transcription factors known to be involved in the regulation of the S-phase of the mitotic cell cycle [HS97]. We choose this transcription factor to test whether our pipeline can predict the function regulated by a single family of transcription factor. The Cluster-Buster program predicted 273 genes to contain a cluster of binding sites for E2F. In these 273 target genes, many biological processes related to the cell cycle are significantly enriched (see Figure 2). The more specific processes are related to DNA replication and S-phase of the mitotic cell cycle. Therefore the prediction of our pipeline yields exactly what we know from experiments: E2F is regulating the synthesis phase of the cell cycle.
6 Gene_Ontology biological_process development physiological process cellular process regulation of biological process morphogenesis response to stress response to external stimulus cell growth and/or maintenance cell communication regulation of cellular process organogenesis response to wounding response to biotic stimulus cell growth cell adhesion response to pest/pathogen/parasite defense response regulation of cell growth immune response innate immune response inflammatory response Figure 3: A part of the Gene Ontology highlighting the biological processes that are significantly enriched in 1913 genes with a cluster of NFAT and AP-1 binding sites predicted in their upstream regions. Grey boxes display those processes with F DR 0.05 (see Methods for details). Combinatorial regulation by AP-1 and NFAT Kel et al. [KKMBW99] have studied the composite binding of NFAT and AP-1 transcription factors. NFAT plays a role in the regulation of cytokines and other genes during immune response. NFAT co-operates with AP-1 to integrate calcium and PKC-signal transduction. They concluded that the combination of NFAT and AP-1 exhibits a significantly higher specificity than individual factors towards genes induced upon T-cell activation. We applied our pipeline to each of these matrices separately and found no significantly enriched process within the predicted targets of individual factors (NFAT: 2716 predicted target genes no process significant, AP-1: 921 predicted target genes no process significant). However, studying both factors together we found significantly enriched biological processes within the predicted 1913 target genes (see Fig. 3). Many of these overrepresented biological processes are related to immune response: inflammatory response, response to wounding, innate immune response, response to biotic stimulus which is in good agreement with the results of Ref. [KKMBW99]. Moreover we found the processes organogenesis, morphogenesis, development, regulation of cell growth, regulation of cellular process, and cell adhesion as novel processes which might be regulated by the combinatorial action of NFAT and AP-1. DISCUSSION The availability of whole-genome sequences and the growing systematic annotations like the Gene Ontology provide the means for more function oriented data mining that goes beyond the single gene level. In this article we propose an approach, which opens a door
7 to infer a biological function regulated by the combinatorial interaction of transcription factors. Contrary to other widespread techniques our method does not intend to predict which factors control genes of similar expression profile. Instead our search only requires a set of positional frequency matrices representing a set of transcription factors to predict in silico their biological function. First, using Cluster-Buster [FLW03] we predict a list of potential target genes for a set of transcription factors. Afterwards, a rigorous statistical test for over-representation implemented in GOSSIP [BBC + 04] is applied to all biological processes provided by the Gene Ontology. Therefore the search is not biased by any prior knowledge related to the factors and gives a chance to detect novel regulatory associations. Additionally the pipeline provides a list of genes supporting the evidence of the reported enriched processes. This gene list can be understood as the cross section of the list of genes regulated by the studied factors and a list of genes annotated with at least one of the overrepresented terms. Due to the filtering nature of the cross section, the final gene list is expected to have less false predictions than the primary list of potentially regulated genes. Two well studied examples presented in this paper illustrate the pipeline s utility. As expected, the clusters of sites related to the E2F transcription factors family are significantly enriched with terms related to the synthesis phase of the mitotic cell cycle. These terms have been detected despite the fact, that only clusters of binding sites corresponding to a single positional frequency matrix have been searched. Although separate studies of AP-1 and NFAT show no significant biological processes, the combinatorial interaction of NFAT and AP-1 yields several significant terms. Besides terms describing aspects of immune response where NFAT/AP-1 is known to be involved in, several significant terms hint to novel hypothesis that require experimental validation. Notably, no tuning of parameters was necessary within these studies. Our method could be further improved by a better selection of promoters, for example by using databases like dbtss [SYSN04] and EPD [SPD + 04]. Our analysis does not include distal enhancer regions, but might be included when sufficient knowledge about enhancer regions has been accumulated. We show in this paper, that the tools are now in hand to infer the regulatory complexity using genome-wide studies of transcription factor binding sites. In forthcoming studies we plan to investigate systematically combinations of transcription factors to get a better understanding of the combinatorial regulation in eukaryotic organisms. ACKNOWLEDGMENTS The authors thank Dieter Beule, who was involved in the development of GOSSIP, and Martin Frith and Zhiping Weng for providing us the Cluster-Buster program. NB acknowledges support from the German Research Foundation (DFG, SFB 618), and SzMK from German Federal Ministry of Education and Research (BMBF).
8 References [ABB + 00] [BBC + 04] Ashburner, M., Ball, C., Blake, J., Botstein, D., Butler, H., Cherry, J., Davis, A., Dolinski, K., Dwight, S., Eppig, J., Harris, M., Hill, D., Issel-Tarver, L., Kasarskis, A., Lewis, S., Matese, J., Richardson, J., Ringwald, M., Rubin, G., and Sherlock, G.: Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 25(1): Blüthgen, N., Brand, K., Cajavec, B., Swat, M., Herzel, H., and Beule, D.: Biological profiling utilizing gene ontology. submitted [BKCH04] Blüthgen, N., Kielbasa, S., Cajavec, B., and Herzel, H.: HOMGL-comparing genelists across species and with different accession numbers. Bioinformatics. 20(1): [CEA + 04] Curwen, V., Eyras, E., Andrews, T., Clarke, L., Mongin, E., Searle, S., and Clamp, M.: The Ensembl automatic gene annotation system. Genome Res. 14(5): [DWR + 03] Dieterich, C., Wang, H., Rateitschak, K., Luz, H., and Vingron, M.: CORG: a database for COmparative Regulatory Genomics. Nucleic Acids Res. 31(1): [FLW03] [FMZ + 02] [HS97] [HWR + 98] Frith, M., Li, M., and Weng, Z.: Cluster-Buster: Finding dense clusters of motifs in DNA sequences. Nucleic Acids Res. 31(13): Fessele, S., Maier, H., Zischek, C., Nelson, P., and Werner, T.: Regulatory context is a crucial part of gene function. Trends Genet. 18(2): Herwig, S. and Strauss, M.: The retinoblastoma protein: a master regulator of cell cycle, differentiation and apoptosis. Eur J Biochem. 246(3): Heinemeyer, T., Wingender, E., Reuter, I., Hermjakob, H., Kel, A., Kel, O., Ignatieva, E., Ananko, E., Podkolodnaya, O., Kolpakov, F., Podkolodny, N., and Kolchanov, N.: Databases on transcriptional regulation: TRANSFAC, TRRD and COMPEL. Nucleic Acids Res. 26(1): [KKMBW99] Kel, A., Kel-Margoulis, O., Babenko, V., and Wingender, E.: Recognition of NFATp/AP-1 composite elements within genes induced upon the activation of immune cells. J Mol Biol. 288(3): [KMIWK02] [KW01] Kel-Margoulis, O., Ivanova, T., Wingender, E., and Kel, A.: Automatic annotation of genomic regulatory sequences by searching for composite clusters. Pac Symp Biocomput. S Krivan, W. and Wasserman, W.: A predictive model for regulatory sequences directing liver-specific transcription. Genome Res. 11(9): [SPD + 04] Schmid, C., Praz, V., Delorenzi, M., Perier, R., and Bucher, P.: The Eukaryotic Promoter Database EPD: the impact of in silico primer extension. Nucleic Acids Res. 32 Database issue:d [St98] Stormo, G.: Information content and free energy in DNA protein interactions. J Theor Biol. 195(1): [SYSN04] Suzuki, Y., Yamashita, R., Sugano, S., and Nakai, K.: DBTSS, DataBase of Transcriptional Start Sites: progress report Nucleic Acids Res. 32 Database issue:d
9 [WF98] Wasserman, W. and Fickett, J.: Identification of regulatory regions which confer muscle-specific gene expression. J Mol Biol. 278(1): [WK03] Wasserman, W. and Krivan, W.: In silico identification of metazoan transcriptional regulatory regions. Naturwissenschaften. 90(4):
Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree
 Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree YOICHI YAMADA, YUKI MIYATA, MASANORI HIGASHIHARA*, KENJI SATOU Graduate School of Natural
Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree YOICHI YAMADA, YUKI MIYATA, MASANORI HIGASHIHARA*, KENJI SATOU Graduate School of Natural
Data visualization and clustering: an application to gene expression data
 Data visualization and clustering: an application to gene expression data Francesco Napolitano Università degli Studi di Salerno Dipartimento di Matematica e Informatica DAA Erice, April 2007 Thanks to
Data visualization and clustering: an application to gene expression data Francesco Napolitano Università degli Studi di Salerno Dipartimento di Matematica e Informatica DAA Erice, April 2007 Thanks to
GOSAP: Gene Ontology Based Semantic Alignment of Biological Pathways
 GOSAP: Gene Ontology Based Semantic Alignment of Biological Pathways Jonas Gamalielsson and Björn Olsson Systems Biology Group, Skövde University, Box 407, Skövde, 54128, Sweden, [jonas.gamalielsson][bjorn.olsson]@his.se,
GOSAP: Gene Ontology Based Semantic Alignment of Biological Pathways Jonas Gamalielsson and Björn Olsson Systems Biology Group, Skövde University, Box 407, Skövde, 54128, Sweden, [jonas.gamalielsson][bjorn.olsson]@his.se,
UDC Comparative Analysis of Methods Recognizing Potential Transcription Factor Binding Sites
 Molecular Biology, Vol. 35, No. 6, 2001, pp. 818 825. Translated from Molekulyarnaya Biologiya, Vol. 35, No. 6, 2001, pp. 961 969. Original Russian Text Copyright 2001 by Pozdnyakov, Vityaev, Ananko, Ignatieva,
Molecular Biology, Vol. 35, No. 6, 2001, pp. 818 825. Translated from Molekulyarnaya Biologiya, Vol. 35, No. 6, 2001, pp. 961 969. Original Russian Text Copyright 2001 by Pozdnyakov, Vityaev, Ananko, Ignatieva,
INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA
 INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA XIUFENG WAN xw6@cs.msstate.edu Department of Computer Science Box 9637 JOHN A. BOYLE jab@ra.msstate.edu Department of Biochemistry and Molecular Biology
INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA XIUFENG WAN xw6@cs.msstate.edu Department of Computer Science Box 9637 JOHN A. BOYLE jab@ra.msstate.edu Department of Biochemistry and Molecular Biology
86 Part 4 SUMMARY INTRODUCTION
 86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
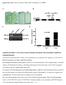 Supplemental Table 1. Primers used for qrt-pcr analysis Name abf2rtf abf2rtr abf3rtf abf3rtf abf4rtf abf4rtr ANAC096RTF ANAC096RTR abf2lp abf2rp abf3lp abf3lp abf4lp abf4rp ANAC096F(XbaI) ANAC096R(BamHI)
Supplemental Table 1. Primers used for qrt-pcr analysis Name abf2rtf abf2rtr abf3rtf abf3rtf abf4rtf abf4rtr ANAC096RTF ANAC096RTR abf2lp abf2rp abf3lp abf3lp abf4lp abf4rp ANAC096F(XbaI) ANAC096R(BamHI)
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Classification of Gene Expression Profiles: Comparison of K-means and Expectation Maximization Algorithms
 Eighth International Conference on Hybrid Intelligent Systems Classification of Gene Expression Profiles: Comparison of K-means and Expectation Maximization Algorithms Cristina Rubio-Escudero Dpto. Lenguajes
Eighth International Conference on Hybrid Intelligent Systems Classification of Gene Expression Profiles: Comparison of K-means and Expectation Maximization Algorithms Cristina Rubio-Escudero Dpto. Lenguajes
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis
 Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Comparison of Human Protein-Protein Interaction Maps
 Comparison of Human Protein-Protein Interaction Maps Matthias E. Futschik 1, Gautam Chaurasia 1,2, Erich Wanker 2 and Hanspeter Herzel 1 1 Institute for Theoretical Biology, Charité, Humboldt-Universität
Comparison of Human Protein-Protein Interaction Maps Matthias E. Futschik 1, Gautam Chaurasia 1,2, Erich Wanker 2 and Hanspeter Herzel 1 1 Institute for Theoretical Biology, Charité, Humboldt-Universität
Rule learning for gene expression data
 Rule learning for gene expression data Stefan Enroth Original slides by Torgeir R. Hvidsten The Linnaeus Centre for Bioinformatics Predicting biological process from gene expression time profiles Papers:
Rule learning for gene expression data Stefan Enroth Original slides by Torgeir R. Hvidsten The Linnaeus Centre for Bioinformatics Predicting biological process from gene expression time profiles Papers:
Gene Ontology. Shifra Ben-Dor. Weizmann Institute of Science
 Gene Ontology Shifra Ben-Dor Weizmann Institute of Science Outline of Session What is GO (Gene Ontology)? What tools do we use to work with it? Combination of GO with other analyses What is Ontology? 1700s
Gene Ontology Shifra Ben-Dor Weizmann Institute of Science Outline of Session What is GO (Gene Ontology)? What tools do we use to work with it? Combination of GO with other analyses What is Ontology? 1700s
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Deciphering regulatory networks by promoter sequence analysis
 Bioinformatics Workshop 2009 Interpreting Gene Lists from -omics Studies Deciphering regulatory networks by promoter sequence analysis Elodie Portales-Casamar University of British Columbia www.cisreg.ca
Bioinformatics Workshop 2009 Interpreting Gene Lists from -omics Studies Deciphering regulatory networks by promoter sequence analysis Elodie Portales-Casamar University of British Columbia www.cisreg.ca
Improving Gene Functional Analysis in Ethylene-induced Leaf Abscission using GO and ProteInOn
 Improving Gene Functional Analysis in Ethylene-induced Leaf Abscission using GO and ProteInOn Sara Domingos 1, Cátia Pesquita 2, Francisco M. Couto 2, Luis F. Goulao 3, Cristina Oliveira 1 1 Instituto
Improving Gene Functional Analysis in Ethylene-induced Leaf Abscission using GO and ProteInOn Sara Domingos 1, Cátia Pesquita 2, Francisco M. Couto 2, Luis F. Goulao 3, Cristina Oliveira 1 1 Instituto
Supplemental table S7.
 Supplemental table S7. GO terms significantly enriched in significantly up-regulated genes of the microarray. K: number of genes from the input cluster in the given category. F: number of total genes in
Supplemental table S7. GO terms significantly enriched in significantly up-regulated genes of the microarray. K: number of genes from the input cluster in the given category. F: number of total genes in
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00.
 Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
How much non-coding DNA do eukaryotes require?
 How much non-coding DNA do eukaryotes require? Andrei Zinovyev UMR U900 Computational Systems Biology of Cancer Institute Curie/INSERM/Ecole de Mine Paritech Dr. Sebastian Ahnert Dr. Thomas Fink Bioinformatics
How much non-coding DNA do eukaryotes require? Andrei Zinovyev UMR U900 Computational Systems Biology of Cancer Institute Curie/INSERM/Ecole de Mine Paritech Dr. Sebastian Ahnert Dr. Thomas Fink Bioinformatics
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
AP Curriculum Framework with Learning Objectives
 Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
simplified Petri nets and analysis uk/gnapn/
 Supplementary information S1 (table) Databases and tools Logical Booleannet GINsim 1 GNaPN 2 MetaReg 3-5 Continuous JigCell 6 Narrator 7 Oscill8 PET Modelling and analysis of RNs. Tools for editing and
Supplementary information S1 (table) Databases and tools Logical Booleannet GINsim 1 GNaPN 2 MetaReg 3-5 Continuous JigCell 6 Narrator 7 Oscill8 PET Modelling and analysis of RNs. Tools for editing and
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Enduring understanding 1.A: Change in the genetic makeup of a population over time is evolution.
 The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
Gene Ontology and overrepresentation analysis
 Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim Overview How
Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim Overview How
GENOME-WIDE ANALYSIS OF CORE PROMOTER REGIONS IN EMILIANIA HUXLEYI
 1 GENOME-WIDE ANALYSIS OF CORE PROMOTER REGIONS IN EMILIANIA HUXLEYI Justin Dailey and Xiaoyu Zhang Department of Computer Science, California State University San Marcos San Marcos, CA 92096 Email: daile005@csusm.edu,
1 GENOME-WIDE ANALYSIS OF CORE PROMOTER REGIONS IN EMILIANIA HUXLEYI Justin Dailey and Xiaoyu Zhang Department of Computer Science, California State University San Marcos San Marcos, CA 92096 Email: daile005@csusm.edu,
GRAPH-THEORETICAL COMPARISON REVEALS STRUCTURAL DIVERGENCE OF HUMAN PROTEIN INTERACTION NETWORKS
 141 GRAPH-THEORETICAL COMPARISON REVEALS STRUCTURAL DIVERGENCE OF HUMAN PROTEIN INTERACTION NETWORKS MATTHIAS E. FUTSCHIK 1 ANNA TSCHAUT 2 m.futschik@staff.hu-berlin.de tschaut@zedat.fu-berlin.de GAUTAM
141 GRAPH-THEORETICAL COMPARISON REVEALS STRUCTURAL DIVERGENCE OF HUMAN PROTEIN INTERACTION NETWORKS MATTHIAS E. FUTSCHIK 1 ANNA TSCHAUT 2 m.futschik@staff.hu-berlin.de tschaut@zedat.fu-berlin.de GAUTAM
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Big Idea 1: The process of evolution drives the diversity and unity of life.
 Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
Biology Assessment. Eligible Texas Essential Knowledge and Skills
 Biology Assessment Eligible Texas Essential Knowledge and Skills STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules
Biology Assessment Eligible Texas Essential Knowledge and Skills STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
S1 Gene ontology (GO) analysis of the network alignment results
 1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
STAAR Biology Assessment
 STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules as building blocks of cells, and that cells are the basic unit of
STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules as building blocks of cells, and that cells are the basic unit of
Bioinformatics Chapter 1. Introduction
 Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Understanding Science Through the Lens of Computation. Richard M. Karp Nov. 3, 2007
 Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Control of Gene Expression
 Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
Robust Community Detection Methods with Resolution Parameter for Complex Detection in Protein Protein Interaction Networks
 Robust Community Detection Methods with Resolution Parameter for Complex Detection in Protein Protein Interaction Networks Twan van Laarhoven and Elena Marchiori Institute for Computing and Information
Robust Community Detection Methods with Resolution Parameter for Complex Detection in Protein Protein Interaction Networks Twan van Laarhoven and Elena Marchiori Institute for Computing and Information
Valley Central School District 944 State Route 17K Montgomery, NY Telephone Number: (845) ext Fax Number: (845)
 Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
AN EVIDENCE ONTOLOGY FOR USE IN PATHWAY/GENOME DATABASES
 AN EVIDENCE ONTOLOGY FOR USE IN PATHWAY/GENOME DATABASES Appears in Pacific Symposium on Biocomputing 2004 Pages 190-201, World Scientific Publishers, Singapore P.D. KARP, S. PALEY, C.J. KRIEGER SRI International,
AN EVIDENCE ONTOLOGY FOR USE IN PATHWAY/GENOME DATABASES Appears in Pacific Symposium on Biocomputing 2004 Pages 190-201, World Scientific Publishers, Singapore P.D. KARP, S. PALEY, C.J. KRIEGER SRI International,
Supplementary Information 16
 Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
Regulation of gene Expression in Prokaryotes & Eukaryotes
 Regulation of gene Expression in Prokaryotes & Eukaryotes 1 The trp Operon Contains 5 genes coding for proteins (enzymes) required for the synthesis of the amino acid tryptophan. Also contains a promoter
Regulation of gene Expression in Prokaryotes & Eukaryotes 1 The trp Operon Contains 5 genes coding for proteins (enzymes) required for the synthesis of the amino acid tryptophan. Also contains a promoter
Biology. Biology. Slide 1 of 26. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Small RNA in rice genome
 Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Compare and contrast the cellular structures and degrees of complexity of prokaryotic and eukaryotic organisms.
 Subject Area - 3: Science and Technology and Engineering Education Standard Area - 3.1: Biological Sciences Organizing Category - 3.1.A: Organisms and Cells Course - 3.1.B.A: BIOLOGY Standard - 3.1.B.A1:
Subject Area - 3: Science and Technology and Engineering Education Standard Area - 3.1: Biological Sciences Organizing Category - 3.1.A: Organisms and Cells Course - 3.1.B.A: BIOLOGY Standard - 3.1.B.A1:
COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry.
 North Carolina Draft Standard Course of Study and Grade Level Competencies, Biology BIOLOGY COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry. 1.01
North Carolina Draft Standard Course of Study and Grade Level Competencies, Biology BIOLOGY COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry. 1.01
Supplemental Material
 Supplemental Material Protocol S1 The PromPredict algorithm considers the average free energy over a 100nt window and compares it with the the average free energy over a downstream 100nt window separated
Supplemental Material Protocol S1 The PromPredict algorithm considers the average free energy over a 100nt window and compares it with the the average free energy over a downstream 100nt window separated
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón
 GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation
 Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation Authors: Fan Zhang, Runsheng Liu and Jie Zheng Presented by: Fan Wu School of Computer Science and
Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation Authors: Fan Zhang, Runsheng Liu and Jie Zheng Presented by: Fan Wu School of Computer Science and
Sequence analysis and comparison
 The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs
 Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Map of AP-Aligned Bio-Rad Kits with Learning Objectives
 Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
networks in molecular biology Wolfgang Huber
 networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
CSCE555 Bioinformatics. Protein Function Annotation
 CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
Genome-wide co-occurrence of promoter elements reveals a cisregulatory. cassette of rrna transcription motifs in S. cerevisiae
 Genome-wide co-occurrence of promoter elements reveals a cisregulatory cassette of rrna transcription motifs in S. cerevisiae Priya Sudarsanam #, Yitzhak Pilpel #, and George M. Church * Department of
Genome-wide co-occurrence of promoter elements reveals a cisregulatory cassette of rrna transcription motifs in S. cerevisiae Priya Sudarsanam #, Yitzhak Pilpel #, and George M. Church * Department of
Essential knowledge 1.A.2: Natural selection
 Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
An introduction to SYSTEMS BIOLOGY
 An introduction to SYSTEMS BIOLOGY Paolo Tieri CNR Consiglio Nazionale delle Ricerche, Rome, Italy 10 February 2015 Universidade Federal de Minas Gerais, Belo Horizonte, Brasil Course outline Day 1: intro
An introduction to SYSTEMS BIOLOGY Paolo Tieri CNR Consiglio Nazionale delle Ricerche, Rome, Italy 10 February 2015 Universidade Federal de Minas Gerais, Belo Horizonte, Brasil Course outline Day 1: intro
Fuzzy Clustering of Gene Expression Data
 Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes.
 I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Dynamical Modeling in Biology: a semiotic perspective. Junior Barrera BIOINFO-USP
 Dynamical Modeling in Biology: a semiotic perspective Junior Barrera BIOINFO-USP Layout Introduction Dynamical Systems System Families System Identification Genetic networks design Cell Cycle Modeling
Dynamical Modeling in Biology: a semiotic perspective Junior Barrera BIOINFO-USP Layout Introduction Dynamical Systems System Families System Identification Genetic networks design Cell Cycle Modeling
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Total
 Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
Science Online Instructional Materials Correlation to the 2010 Biology Standards of Learning and Curriculum Framework
 and Curriculum Framework Provider York County School Divison Course Title Biology Last Updated 2010-11 Course Syllabus URL http://yorkcountyschools.org/virtuallearning/coursecatalog.aspx BIO.1 The student
and Curriculum Framework Provider York County School Divison Course Title Biology Last Updated 2010-11 Course Syllabus URL http://yorkcountyschools.org/virtuallearning/coursecatalog.aspx BIO.1 The student
 1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
Campbell Biology AP Edition 11 th Edition, 2018
 A Correlation and Narrative Summary of Campbell Biology AP Edition 11 th Edition, 2018 To the AP Biology Curriculum Framework AP is a trademark registered and/or owned by the College Board, which was not
A Correlation and Narrative Summary of Campbell Biology AP Edition 11 th Edition, 2018 To the AP Biology Curriculum Framework AP is a trademark registered and/or owned by the College Board, which was not
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Whole Genome Human/Mouse Phylogenetic Footprinting of Potential Transcription Regulatory Signals. E. Cheremushkin, A. Kel
 Whole Genome Human/Mouse Phylogenetic Footprinting of Potential Transcription Regulatory Signals E. Cheremushkin, A. Kel Pacific Symposium on Biocomputing 8:9-30(003) WHOLE GEN OM E HU M A N / M OU S E
Whole Genome Human/Mouse Phylogenetic Footprinting of Potential Transcription Regulatory Signals E. Cheremushkin, A. Kel Pacific Symposium on Biocomputing 8:9-30(003) WHOLE GEN OM E HU M A N / M OU S E
Networks & pathways. Hedi Peterson MTAT Bioinformatics
 Networks & pathways Hedi Peterson (peterson@quretec.com) MTAT.03.239 Bioinformatics 03.11.2010 Networks are graphs Nodes Edges Edges Directed, undirected, weighted Nodes Genes Proteins Metabolites Enzymes
Networks & pathways Hedi Peterson (peterson@quretec.com) MTAT.03.239 Bioinformatics 03.11.2010 Networks are graphs Nodes Edges Edges Directed, undirected, weighted Nodes Genes Proteins Metabolites Enzymes
Measuring TF-DNA interactions
 Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Keystone Exams: Biology Assessment Anchors and Eligible Content. Pennsylvania Department of Education
 Assessment Anchors and Pennsylvania Department of Education www.education.state.pa.us 2010 PENNSYLVANIA DEPARTMENT OF EDUCATION General Introduction to the Keystone Exam Assessment Anchors Introduction
Assessment Anchors and Pennsylvania Department of Education www.education.state.pa.us 2010 PENNSYLVANIA DEPARTMENT OF EDUCATION General Introduction to the Keystone Exam Assessment Anchors Introduction
Large-scale GO analysis reveals gene expression level and variation are functional and location related
 Large-scale GO analysis reveals gene expression level and variation are functional and location related Simon M. Lin 1, Pan Du 1, and Warren A. Kibbe 1 1 Robert H. Lurie Comprehensive Cancer Center, Northwestern
Large-scale GO analysis reveals gene expression level and variation are functional and location related Simon M. Lin 1, Pan Du 1, and Warren A. Kibbe 1 1 Robert H. Lurie Comprehensive Cancer Center, Northwestern
Science Textbook and Instructional Materials Correlation to the 2010 Biology Standards of Learning and Curriculum Framework. Publisher Information
 Publisher Information Copyright date 2013 Contact Carol Kornfeind Phone# 847-486-2065 E-mail carol.kornfeind@pearson.com Biology 1 of 12 Virginia Department of Education Text Miller Levine Biology, Virginia
Publisher Information Copyright date 2013 Contact Carol Kornfeind Phone# 847-486-2065 E-mail carol.kornfeind@pearson.com Biology 1 of 12 Virginia Department of Education Text Miller Levine Biology, Virginia
Biological Networks: Comparison, Conservation, and Evolution via Relative Description Length By: Tamir Tuller & Benny Chor
 Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
identifiers matched to homologous genes. Probeset annotation files for each array platform were used to
 SUPPLEMENTARY METHODS Data combination and normalization Prior to data analysis we first had to appropriately combine all 1617 arrays such that probeset identifiers matched to homologous genes. Probeset
SUPPLEMENTARY METHODS Data combination and normalization Prior to data analysis we first had to appropriately combine all 1617 arrays such that probeset identifiers matched to homologous genes. Probeset
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 SUMOylation of proteins changes drastically upon heat shock, MG-132 treatment and PR-619 treatment. (a) Schematic overview of all SUMOylation proteins identified to be differentially
Supplementary Figure 1 SUMOylation of proteins changes drastically upon heat shock, MG-132 treatment and PR-619 treatment. (a) Schematic overview of all SUMOylation proteins identified to be differentially
Francisco M. Couto Mário J. Silva Pedro Coutinho
 Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Drosophila melanogaster and D. simulans, two fruit fly species that are nearly
 Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Chapter 20. Initiation of transcription. Eukaryotic transcription initiation
 Chapter 20. Initiation of transcription Eukaryotic transcription initiation 2003. 5.22 Prokaryotic vs eukaryotic Bacteria = one RNA polymerase Eukaryotes have three RNA polymerases (I, II, and III) in
Chapter 20. Initiation of transcription Eukaryotic transcription initiation 2003. 5.22 Prokaryotic vs eukaryotic Bacteria = one RNA polymerase Eukaryotes have three RNA polymerases (I, II, and III) in
Package neat. February 23, 2018
 Type Package Title Efficient Network Enrichment Analysis Test Version 1.1.3 Date 2018-02-23 Depends R (>= 3.3.0) Package neat February 23, 2018 Author Mirko Signorelli, Veronica Vinciotti and Ernst C.
Type Package Title Efficient Network Enrichment Analysis Test Version 1.1.3 Date 2018-02-23 Depends R (>= 3.3.0) Package neat February 23, 2018 Author Mirko Signorelli, Veronica Vinciotti and Ernst C.
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Matrix-based pattern discovery algorithms
 Regulatory Sequence Analysis Matrix-based pattern discovery algorithms Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe)
Regulatory Sequence Analysis Matrix-based pattern discovery algorithms Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe)
Initiation of translation in eukaryotic cells:connecting the head and tail
 Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Computation-Based Discovery of Cis-Regulatory. Modules by Hidden Markov Model
 Computation-Based Discovery of Cis-Regulatory Modules by Hidden Markov Model Jing Wu and Jun Xie Department of Statistics Purdue University 150 N. University Street West Lafayette, IN 47907 Tel: 765-494-6032
Computation-Based Discovery of Cis-Regulatory Modules by Hidden Markov Model Jing Wu and Jun Xie Department of Statistics Purdue University 150 N. University Street West Lafayette, IN 47907 Tel: 765-494-6032
Evidence for dynamically organized modularity in the yeast protein-protein interaction network
 Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
