Rule learning for gene expression data
|
|
|
- Nickolas Nichols
- 6 years ago
- Views:
Transcription
1 Rule learning for gene expression data Stefan Enroth Original slides by Torgeir R. Hvidsten The Linnaeus Centre for Bioinformatics
2 Predicting biological process from gene expression time profiles Papers: I. T. R. Hvidsten, A. Lægreid and J. Komorowski. Learning rule-based models of biological process from gene expression time profiles using gene ontology, Bioinformatics 19(9): , II. A. Lægreid, T. R. Hvidsten, H. Midelfart, J. Komorowski and A. K. Sandvik. Predicting Gene Ontology Biological Process From Temporal Gene Expression Patterns, Genome Research, 13(5): , 2003.
3 Hierarchical clustering 3 Iyer et al., The transcriptional program in the response of human fibroblasts to serum, Science, 283(5398): 83-87, 1999
4 4 Gene Ontology vs. expression clustering
5 Methodology Ontology Process 1. Annotation Transport Defense response Positive control of cell proliferation Cell cycle control g 2... g 2... g 4... g 5 g Extracting features for learning Gene 0HR 15MIN 30MIN 1HR 2HR 4HR 6HR 8HR 12HR 16HR 20HR 24HR Process g Unknown g Transport and defense response g Cell cycle control Positive control of g cell proliferation g Positive control of cell proliferation Inducing minimal decision rules using rough sets (Increasing) AND 6-10(Decreasing) AND 14-18(Constant) => GO(cell proliferation) 5 4. The function of uncharacterized! genes is predicted using the rules
6 Rule Induction IF THEN 0-4 (Constant) AND 0-10(Increasing) GO(prot. met. and mod.) OR GO(mesoderm develop.) OR GO(prot. biosynt.) 6 IF-part (antecedent, premise) the minimal set of discrete changes in expression needed to uphold the discriminatory power of the full data set THEN-part (conclusion) all functions of genes described by the antecedent-side We want rules that describe the expression profiles of several genes with one or a few functions accuracy: the fraction of genes matching the IF-part that are annotated with the process in the THEN-part coverage: the fraction of genes annotated with the process in the THEN-part that matches the IF-part
7 Rule example Rule 0-4(Constant) AND 0-10(Increasing) => GO(protein metabolism and modification) OR GO(mesoderm development) OR GO(protein biosynthesis) Covered genes 4: M35296 J02783 D13748 X : X : D
8 Classification X Function? 8 IF THEN IF THEN IF THEN IF THEN IF THEN IF THEN IF THEN IF THEN IF THEN IF THEN Votes are normalized and processes with vote fractions higher than a selection-threshold are chosen as predictions IF 0-4(Constant) AND 0-10(Increasing) THEN GO(protein metabolism and modification ) OR GO(mesoderm development) OR GO(protein biosynthesis) +1 Process Votes protein metabolism and modification 6 mesoderm development 3 proteolysis and peptidolysis 2 transcription 1 protein biosynthesis 1 vision 1
9 Cross validation Iteration 1 Iteration 2 Iteration 3 Observation 1 Observation 2 9 Fold 1 Fold 2 Fold 3 Training set Training set Test set Training set Test set Training set Test set Training set Training set Observation n
10 Evaluation Evaluation technique divide examples into training set and test set cross validation Evaluation measures: accuracy = (TP+TN)/(TP+FN+TN+FP) sensitivity = TP/(TP+FN) specificity = TN/(TN+FP) Confusion matrix: 10
11 Threshold selection Gene with function protein biosynthesis Gene with a different function sensitivity: TP/(TP+FN) specificity: TN/(TN+FP) 1 Fraction of votes for protein biosynthesis Threshold 1 Threshold 2 g 1 g 2 g 3 g 4 g 5 g 6 g 7 g 8 g 9 g 10 g 11 g 12 g 13 Test set Sensitivity = 2/3, Specificity=1 Sensitivity = 1, Specificity=2/3
12 ROC analysis and classifier evaluation 1 sensitivity Perfect discrimination AUC No discrimination ROC: Receiver Operating Characteristics curve results from plotting sensitivity against specificity for all possible thresholds sensitivity: TP/(TP+FN) specificity: TN/(TN+FP) AUC: Area Under the ROC Curve specificity False alarm 1
13 ROC analysis and classifier evaluation 1 Perfect discrimination A B Which ROC curve is better? A dominants B and C and clearly has a higher AUC sensitivity C No discrimination B and C have approximately the same AUC B is better for some thresholds, C for others specificity 1
14 Cross validation estimates PROCESS AUC SE P-VALUE Ion homeostasis Protein targeting Blood coagulation DNA metabolism Intracellular signaling cascade Cell cycle Energy pathways Oncogenesis Circulation Cell death Developmental processes Defense (immune) response Transcription Cell adhesion Stress response Protein metabolism and modification Cell motility Cell surface rec linked signal transd Lipid metabolism Cell organization and biogenesis Cell proliferation Transport Amino acid and derivative metabolism AVERAGE Over all classes: Coverage (recall) = TP/(TP+FN) Precision = TP/(TP+FP) Coverage: 84% Precision: 50% Coverage: 71% Precision: 60% Coverage: 39% Precision: 90% *Iyer et al. 14
15 Discovering regulatory binding site modules Paper: III. T. R. Hvidsten, B. Wilczyński, A. Kryshtafovych, J. Tiuryn, J. Komorowski and K. Fidelis. Discovering regulatory binding site modules using rule-based learning, Genome Research 15: , 2005
16 Gene regulation Gene expression is regulated by regulatory proteins (transcription factors) Transcription factors depend on recognizing sequence motifs (binding sites) in order to effect the expression of genes Transcription factors combine to respond to a large number of stress factors (e.g. heat shock) with a large number of expression outcomes Enhancer Silencer Response elements Promoter Regulatory region Binding sites Transcription region Yeast 16 Promoter
17 Background Many studies have used gene expression data to search for overrepresented sequence motifs in co-expressed genes Pilpel et al. (2001) found that genes sharing pairs of binding sites are significantly more likely to be co-expressed than genes with only single binding sites in common Expression coherence score (EC) 17 Pilpel, Y., P. Sudarsanam, and G.M. Church Identifying regulatory networks by combinatorial analysis of promoter elements. Nat Genet 29:
18 Introductory remarks We want to explore the combinatorial nature of gene regulation ASSUMPTION: co-regulated genes (genes regulated by the same transcription factors through the same binding sites) are co-expressed (have similar expression profiles) Genome-wide analysis of combinatorial regulation using sequence and expression data 18 Binding site module: a set of binding sites that is used to co-regulate a set of genes
19 Data and Method Database of 43 known and 313 putative yeast binding site motifs Expression profiles for yeast genes measured under six different conditions: cell cycle and five stress conditions sporulation, diauxic shift, heat shock and cold shock, pheromone and DNA-damaging agents Gene HAP234 Binding sites RAP1 PHO SWI5 ECB MCM1' RPL18A RPS18A RPL16B RPL26A RPS24A RPL RPL14A SST DRS GIT CLN RPO BIT Next gene Similar expression to RPL18A? Rule learning* yes yes yes yes yes yes yes no no no no no no IF RAP1 AND SWI5 AND MCM1' THEN 4 Evaluation: Gene Ontology Binding data Filtering*
20 An example of a binding site module IF RAP1 AND SWI5 AND MCM1' THEN The rule was (re-) discovered in five of the six expression data sets a) Cell cycle: RPL30, RPL18A, RPL26A, RPL14A, RPS18A, (SST2), RPL16B b) Sporulation: RPL30, RPL18A, RPL14A, (RPS18A), RPL16B, RPL26A The central gene in the expression cluster is underlined c) Diauxic shift: RPL30, RPL18A, RPL14A, RPS18A, SST2, RPL16B, RPL26A Genes with differing expression profiles are in parentheses d) Heat and cold shock: RPS24A, RPL26A, RPL14A, RPS18A, RPL16B e) DNA-damaging agents: RPL30, RPL18A, RPL26A, RPL14A, RPS18A, (SST2), RPL16B
21 Evaluation IF RAP1 AND SWI5 AND MCM1' THEN <similar expression> Gene symbol Biological process Molecular function Cellular component Possible transcription factors (P-value < 0.01) RPL16B protein biosynthesis RNA binding, structural constituent of ribosome cytosolic ribosome (sensu Eukarya), large ribosomal subunit FHL1, GAT3, PDR1, RAP1, RGM1, YAP5 RPL26A protein biosynthesis RNA binding, structural constituent of ribosome cytosolic ribosome (sensu Eukarya), large ribosomal subunit FHL1,RAP1 RPS18A protein biosynthesis structural constituent of ribosome cytosolic ribosome (sensu Eukarya), eukaryotic 43S preinitiation complex,eukaryotic 48S initiation complex, mall ribosomal subunit FHL1, GAT3, HIR2, RAP1, RGM1, YAP5 RPL30 protein biosynthesis, rrna processing, mrna splicing, regulation of translation structural constituent of ribosome cytosolic ribosome (sensu Eukarya), cytoplasm, large ribosomal subunit FHL1, GAT3, RAP1, SFP1 RPL18A protein biosynthesis structural constituent of ribosome cytosolic ribosome (sensu Eukarya), large ribosomal subunit FHL1, MAL13, RAP1, YAP5 RPL14A protein biosynthesis RNA binding, structural constituent of ribosome cytosolic ribosome (sensu Eukarya), large ribosomal subunit FHL1, GAT3, GRF10(Pho2), GTS1, RAP1 SST2 signal transduction, adaptation to pheromone during conjugation with cellular fusion GTPase activator activity plasma membrane DIG1, FHL1, RAP1, STE12 RPS24A protein biosynthesis structural constituent of ribosome cytosolic ribosome (sensu Eukarya), eukaryotic 43S pre-initiation complex, eukaryotic 48S initiation complex, small ribosomal subunit FHL1, GAT3, PDR1, RAP1, RGM1, SMP1, YAP5 21 P-VALUE 2.35E-04 (protein biosynthesis) 2.36E-06 (structural constituent of ribosome) 5.66E-07 (cytosolic ribosome) 2.38E-10 (FHL1), 3.38E-08 (RAP1), 2.96E-05 (GAT3)
22 Model-based detection of periodic expression Paper: IV. C.R. Andersson, T.R. Hvidsten, A. Isaksson, M.G. Gustafsson and J. Komorowski. Revealing cell cycle control by combining model-based detection of periodic expression with novel cisregulatory descriptors, BMC Systems Biology, 1: 45, 2007.
23 Experiment Induction Prior knowledge Hypothesis Experiment Induction A technique that infers generalizations from the information in the data. Inference Inference the reasoning involved in drawing a conclusion or making a logical judgment on the basis of circumstantial evidence and prior conclusions rather than on the basis of direct observation 23
24 Synchronization To recover mrna-levels, cultures must be synchronized. Synchronization halts cells at a particular point in the cell cycle. Typically the mating system or temperature sensitive mutants are used (cdc28, cdc15). Spellman (1998). S cerevisae under various synchronizations (alpha-factor, ts cdc15, cdc28 and elutriation) Synchronization reveals periodic expression Periodic expression is related to the cell cycle 24
25 Detecting periodically expressed genes Signal Expression Time Which model is most similar to the signal? => Probability (Periodical expression) Model 0 Model 1 Expression Expression 25 Time Time T
26 The models H 0 : y t = μ + ε H = μ + ω + φ 1 : y t A cos( t ) + ε Prior knowledge: The period T 26
27 27 Conditional periodicity
28 28 Method
29 Comparison to clustering Known cell cycle related: sequence motifs CCA, ECB, MCB, SWI5, SCB, SFF, MCM1, SFF' and MCM1. transcription factors ACE2, FKH1, FKH2, GTS1, HIR1, HIR2, MBP1, MCM1, NDD1, STB1, SWI4,SWI5, SWI6, XBP1, YBR267W, YHP1 and YOX1. 29
30 Examples of interactions The point (0.034, 0.73) on the curve is associated with the 145 rules with p-value lower than These rules include 19 of the 26 known phase specific regulators (73%) and 18 other regulators (3.4%). Furthermore, they describe 24% of the genes in the periodic classes. 30
31 Examples of interactions Ellipses/rectangles: transcription factors/sequence motifs Green/Blue: Cell cycle related transcription factors/sequence motifs Red: interactions between transcription factors 31
32 Summary Gene expression data can be used in many types of applications in which rule-based methods can be used to synthesis hypothesis. Discovered rules/classes must always be validated against real world knowledge! If appropriate, prior real world knowledge can be incorporated into the models. 32
Inferring Transcriptional Regulatory Networks from Gene Expression Data II
 Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Discovering modules in expression profiles using a network
 Discovering modules in expression profiles using a network Igor Ulitsky 1 2 Protein-protein interactions (PPIs) Low throughput measurements: accurate, scarce High throughput: more abundant, noisy Large,
Discovering modules in expression profiles using a network Igor Ulitsky 1 2 Protein-protein interactions (PPIs) Low throughput measurements: accurate, scarce High throughput: more abundant, noisy Large,
Bioinformatics. Transcriptome
 Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
CLUSTER, FUNCTION AND PROMOTER: ANALYSIS OF YEAST EXPRESSION ARRAY
 CLUSTER, FUNCTION AND PROMOTER: ANALYSIS OF YEAST EXPRESSION ARRAY J. ZHU, M. Q. ZHANG Cold Spring Harbor Lab, P. O. Box 100 Cold Spring Harbor, NY 11724 Gene clusters could be derived based on expression
CLUSTER, FUNCTION AND PROMOTER: ANALYSIS OF YEAST EXPRESSION ARRAY J. ZHU, M. Q. ZHANG Cold Spring Harbor Lab, P. O. Box 100 Cold Spring Harbor, NY 11724 Gene clusters could be derived based on expression
Understanding Science Through the Lens of Computation. Richard M. Karp Nov. 3, 2007
 Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Central postgenomic. Transcription regulation: a genomic network. Transcriptome: the set of all mrnas expressed in a cell at a specific time.
 Transcription regulation: a genomic network Nicholas Luscombe Laboratory of Mark Gerstein Department of Molecular Biophysics and Biochemistry Yale University Understand Proteins, through analyzing populations
Transcription regulation: a genomic network Nicholas Luscombe Laboratory of Mark Gerstein Department of Molecular Biophysics and Biochemistry Yale University Understand Proteins, through analyzing populations
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
Name: SBI 4U. Gene Expression Quiz. Overall Expectation:
 Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
S1 Gene ontology (GO) analysis of the network alignment results
 1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
GO ID GO term Number of members GO: translation 225 GO: nucleosome 50 GO: calcium ion binding 76 GO: structural
 GO ID GO term Number of members GO:0006412 translation 225 GO:0000786 nucleosome 50 GO:0005509 calcium ion binding 76 GO:0003735 structural constituent of ribosome 170 GO:0019861 flagellum 23 GO:0005840
GO ID GO term Number of members GO:0006412 translation 225 GO:0000786 nucleosome 50 GO:0005509 calcium ion binding 76 GO:0003735 structural constituent of ribosome 170 GO:0019861 flagellum 23 GO:0005840
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Supplementary Information 16
 Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p
 Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis
 Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Biological Process Term Enrichment
 Biological Process Term Enrichment cellular protein localization cellular macromolecule localization intracellular protein transport intracellular transport generation of precursor metabolites and energy
Biological Process Term Enrichment cellular protein localization cellular macromolecule localization intracellular protein transport intracellular transport generation of precursor metabolites and energy
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
Supplemental table S7.
 Supplemental table S7. GO terms significantly enriched in significantly up-regulated genes of the microarray. K: number of genes from the input cluster in the given category. F: number of total genes in
Supplemental table S7. GO terms significantly enriched in significantly up-regulated genes of the microarray. K: number of genes from the input cluster in the given category. F: number of total genes in
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs
 Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón
 GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
Reading Assignments. A. Genes and the Synthesis of Polypeptides. Lecture Series 7 From DNA to Protein: Genotype to Phenotype
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Eukaryotic Gene Expression
 Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Computational Cell Biology Lecture 4
 Computational Cell Biology Lecture 4 Case Study: Basic Modeling in Gene Expression Yang Cao Department of Computer Science DNA Structure and Base Pair Gene Expression Gene is just a small part of DNA.
Computational Cell Biology Lecture 4 Case Study: Basic Modeling in Gene Expression Yang Cao Department of Computer Science DNA Structure and Base Pair Gene Expression Gene is just a small part of DNA.
Total
 Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
STAAR Biology Assessment
 STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules as building blocks of cells, and that cells are the basic unit of
STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules as building blocks of cells, and that cells are the basic unit of
Signal Transduction. Dr. Chaidir, Apt
 Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
The Research Plan. Functional Genomics Research Stream. Transcription Factors. Tuning In Is A Good Idea
 Functional Genomics Research Stream The Research Plan Tuning In Is A Good Idea Research Meeting: March 23, 2010 The Road to Publication Transcription Factors Protein that binds specific DNA sequences controlling
Functional Genomics Research Stream The Research Plan Tuning In Is A Good Idea Research Meeting: March 23, 2010 The Road to Publication Transcription Factors Protein that binds specific DNA sequences controlling
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11
 UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
Welcome to Class 21!
 Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
Gene Control Mechanisms at Transcription and Translation Levels
 Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Biology Assessment. Eligible Texas Essential Knowledge and Skills
 Biology Assessment Eligible Texas Essential Knowledge and Skills STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules
Biology Assessment Eligible Texas Essential Knowledge and Skills STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules
Boolean models of gene regulatory networks. Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016
 Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Computational Genomics. Reconstructing dynamic regulatory networks in multiple species
 02-710 Computational Genomics Reconstructing dynamic regulatory networks in multiple species Methods for reconstructing networks in cells CRH1 SLT2 SLR3 YPS3 YPS1 Amit et al Science 2009 Pe er et al Recomb
02-710 Computational Genomics Reconstructing dynamic regulatory networks in multiple species Methods for reconstructing networks in cells CRH1 SLT2 SLR3 YPS3 YPS1 Amit et al Science 2009 Pe er et al Recomb
identifiers matched to homologous genes. Probeset annotation files for each array platform were used to
 SUPPLEMENTARY METHODS Data combination and normalization Prior to data analysis we first had to appropriately combine all 1617 arrays such that probeset identifiers matched to homologous genes. Probeset
SUPPLEMENTARY METHODS Data combination and normalization Prior to data analysis we first had to appropriately combine all 1617 arrays such that probeset identifiers matched to homologous genes. Probeset
56:198:582 Biological Networks Lecture 10
 56:198:582 Biological Networks Lecture 10 Temporal Programs and the Global Structure The single-input module (SIM) network motif The network motifs we have studied so far all had a defined number of nodes.
56:198:582 Biological Networks Lecture 10 Temporal Programs and the Global Structure The single-input module (SIM) network motif The network motifs we have studied so far all had a defined number of nodes.
From Gene to Protein
 From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
1. In most cases, genes code for and it is that
 Name Chapter 10 Reading Guide From DNA to Protein: Gene Expression Concept 10.1 Genetics Shows That Genes Code for Proteins 1. In most cases, genes code for and it is that determine. 2. Describe what Garrod
Name Chapter 10 Reading Guide From DNA to Protein: Gene Expression Concept 10.1 Genetics Shows That Genes Code for Proteins 1. In most cases, genes code for and it is that determine. 2. Describe what Garrod
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Whole-genome analysis of GCN4 binding in S.cerevisiae
 Whole-genome analysis of GCN4 binding in S.cerevisiae Lillian Dai Alex Mallet Gcn4/DNA diagram (CREB symmetric site and AP-1 asymmetric site: Song Tan, 1999) removed for copyright reasons. What is GCN4?
Whole-genome analysis of GCN4 binding in S.cerevisiae Lillian Dai Alex Mallet Gcn4/DNA diagram (CREB symmetric site and AP-1 asymmetric site: Song Tan, 1999) removed for copyright reasons. What is GCN4?
Prokaryotic Regulation
 Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
Regulation of gene expression. Premedical - Biology
 Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Bioinformatics Chapter 1. Introduction
 Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Lecture 3: A basic statistical concept
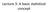 Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Model Accuracy Measures
 Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
A general co-expression network-based approach to gene expression analysis: comparison and applications
 BMC Systems Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. A general co-expression
BMC Systems Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. A general co-expression
Evidence for dynamically organized modularity in the yeast protein-protein interaction network
 Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Lecture 8: Temporal programs and the global structure of transcription networks. Chap 5 of Alon. 5.1 Introduction
 Lecture 8: Temporal programs and the global structure of transcription networks Chap 5 of Alon 5. Introduction We will see in this chapter that sensory transcription networks are largely made of just four
Lecture 8: Temporal programs and the global structure of transcription networks Chap 5 of Alon 5. Introduction We will see in this chapter that sensory transcription networks are largely made of just four
Topographic Independent Component Analysis of Gene Expression Time Series Data
 Topographic Independent Component Analysis of Gene Expression Time Series Data Sookjeong Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong,
Topographic Independent Component Analysis of Gene Expression Time Series Data Sookjeong Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong,
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
What Organelle Makes Proteins According To The Instructions Given By Dna
 What Organelle Makes Proteins According To The Instructions Given By Dna This is because it contains the information needed to make proteins. assemble enzymes and other proteins according to the directions
What Organelle Makes Proteins According To The Instructions Given By Dna This is because it contains the information needed to make proteins. assemble enzymes and other proteins according to the directions
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION
 http://smtom.lecture.ub.ac.id/ Password: https://syukur16tom.wordpress.com/ Password: Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/translation2.gif
http://smtom.lecture.ub.ac.id/ Password: https://syukur16tom.wordpress.com/ Password: Lecture 13: PROTEIN SYNTHESIS II- TRANSLATION http://hyperphysics.phy-astr.gsu.edu/hbase/organic/imgorg/translation2.gif
Eukaryotic vs. Prokaryotic genes
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
BioControl - Week 6, Lecture 1
 BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
9/11/18. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
CISC 636 Computational Biology & Bioinformatics (Fall 2016)
 CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
Computational Systems Biology
 Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
Gene regulation II Biochemistry 302. Bob Kelm February 28, 2005
 Gene regulation II Biochemistry 302 Bob Kelm February 28, 2005 Catabolic operons: Regulation by multiple signals targeting different TFs Catabolite repression: Activity of lac operon is restricted when
Gene regulation II Biochemistry 302 Bob Kelm February 28, 2005 Catabolic operons: Regulation by multiple signals targeting different TFs Catabolite repression: Activity of lac operon is restricted when
Quantitative Molecular Biology
 Quantitative Molecular Biology PHYS 176/276 Instructor: Terry Hwa Winter 2018 What is quantitative biology? èquantitative biology biology + numbers/equations biology-inspired physics application of existing
Quantitative Molecular Biology PHYS 176/276 Instructor: Terry Hwa Winter 2018 What is quantitative biology? èquantitative biology biology + numbers/equations biology-inspired physics application of existing
Inferring Transcriptional Regulatory Networks from High-throughput Data
 Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Functional Genomics Research Stream. Lecture: May 5, 2009 The Road to Publication
 Functional Genomics Research Stream Lecture: May 5, 2009 The Road to Publication Final Lab Issues Lab Hours this Week: Wednesday 10 AM - 6 PM Thursday 10 AM - 6 PM One Hour Required (sign in, sign out)
Functional Genomics Research Stream Lecture: May 5, 2009 The Road to Publication Final Lab Issues Lab Hours this Week: Wednesday 10 AM - 6 PM Thursday 10 AM - 6 PM One Hour Required (sign in, sign out)
Chapter 12. Genes: Expression and Regulation
 Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
9/2/17. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter
 Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter 9/10/2008 1 Learning Objectives Explain why a cell cycle was selected for during evolution
Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter 9/10/2008 1 Learning Objectives Explain why a cell cycle was selected for during evolution
02/02/ Living things are organized. Analyze the functional inter-relationship of cell structures. Learning Outcome B1
 Analyze the functional inter-relationship of cell structures Learning Outcome B1 Describe the following cell structures and their functions: Cell membrane Cell wall Chloroplast Cytoskeleton Cytoplasm Golgi
Analyze the functional inter-relationship of cell structures Learning Outcome B1 Describe the following cell structures and their functions: Cell membrane Cell wall Chloroplast Cytoskeleton Cytoplasm Golgi
Outline. Terminologies and Ontologies. Communication and Computation. Communication. Outline. Terminologies and Vocabularies.
 Page 1 Outline 1. Why do we need terminologies and ontologies? Terminologies and Ontologies Iwei Yeh yeh@smi.stanford.edu 04/16/2002 2. Controlled Terminologies Enzyme Classification Gene Ontology 3. Ontologies
Page 1 Outline 1. Why do we need terminologies and ontologies? Terminologies and Ontologies Iwei Yeh yeh@smi.stanford.edu 04/16/2002 2. Controlled Terminologies Enzyme Classification Gene Ontology 3. Ontologies
Compare and contrast the cellular structures and degrees of complexity of prokaryotic and eukaryotic organisms.
 Subject Area - 3: Science and Technology and Engineering Education Standard Area - 3.1: Biological Sciences Organizing Category - 3.1.A: Organisms and Cells Course - 3.1.B.A: BIOLOGY Standard - 3.1.B.A1:
Subject Area - 3: Science and Technology and Engineering Education Standard Area - 3.1: Biological Sciences Organizing Category - 3.1.A: Organisms and Cells Course - 3.1.B.A: BIOLOGY Standard - 3.1.B.A1:
GENE ACTIVITY Gene structure Transcription Transcript processing mrna transport mrna stability Translation Posttranslational modifications
 1 GENE ACTIVITY Gene structure Transcription Transcript processing mrna transport mrna stability Translation Posttranslational modifications 2 DNA Promoter Gene A Gene B Termination Signal Transcription
1 GENE ACTIVITY Gene structure Transcription Transcript processing mrna transport mrna stability Translation Posttranslational modifications 2 DNA Promoter Gene A Gene B Termination Signal Transcription
REVIEW SESSION. Wednesday, September 15 5:30 PM SHANTZ 242 E
 REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
Regulation of Transcription in Eukaryotes. Nelson Saibo
 Regulation of Transcription in Eukaryotes Nelson Saibo saibo@itqb.unl.pt In eukaryotes gene expression is regulated at different levels 1 - Transcription 2 Post-transcriptional modifications 3 RNA transport
Regulation of Transcription in Eukaryotes Nelson Saibo saibo@itqb.unl.pt In eukaryotes gene expression is regulated at different levels 1 - Transcription 2 Post-transcriptional modifications 3 RNA transport
L3.1: Circuits: Introduction to Transcription Networks. Cellular Design Principles Prof. Jenna Rickus
 L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
Activation of a receptor. Assembly of the complex
 Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Simulation of Gene Regulatory Networks
 Simulation of Gene Regulatory Networks Overview I have been assisting Professor Jacques Cohen at Brandeis University to explore and compare the the many available representations and interpretations of
Simulation of Gene Regulatory Networks Overview I have been assisting Professor Jacques Cohen at Brandeis University to explore and compare the the many available representations and interpretations of
Initiation of translation in eukaryotic cells:connecting the head and tail
 Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Prokaryotic Gene Expression (Learning Objectives)
 Prokaryotic Gene Expression (Learning Objectives) 1. Learn how bacteria respond to changes of metabolites in their environment: short-term and longer-term. 2. Compare and contrast transcriptional control
Prokaryotic Gene Expression (Learning Objectives) 1. Learn how bacteria respond to changes of metabolites in their environment: short-term and longer-term. 2. Compare and contrast transcriptional control
Discovering MultipleLevels of Regulatory Networks
 Discovering MultipleLevels of Regulatory Networks IAS EXTENDED WORKSHOP ON GENOMES, CELLS, AND MATHEMATICS Hong Kong, July 25, 2018 Gary D. Stormo Department of Genetics Outline of the talk 1. Transcriptional
Discovering MultipleLevels of Regulatory Networks IAS EXTENDED WORKSHOP ON GENOMES, CELLS, AND MATHEMATICS Hong Kong, July 25, 2018 Gary D. Stormo Department of Genetics Outline of the talk 1. Transcriptional
Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read
 Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read chapter 22 and chapter 10 [section on MATing type gene
Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read chapter 22 and chapter 10 [section on MATing type gene
Biology 112 Practice Midterm Questions
 Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Connectivity in the Yeast Cell Cycle Transcription Network: Inferences from Neural Networks
 1 Connectivity in the Yeast Cell Cycle Transcription Network: Inferences from Neural Networks Christopher E. Hart 1,2, Eric Mjolsness 3,5, Barbara J. Wold 2,4,5 1 Current Address: Department of Molecular,
1 Connectivity in the Yeast Cell Cycle Transcription Network: Inferences from Neural Networks Christopher E. Hart 1,2, Eric Mjolsness 3,5, Barbara J. Wold 2,4,5 1 Current Address: Department of Molecular,
CSCE555 Bioinformatics. Protein Function Annotation
 CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
Quiz answers. Allele. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA)
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA) http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Quiz answers Kinase: An enzyme
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA) http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Quiz answers Kinase: An enzyme
Protein-protein interaction networks Prof. Peter Csermely
 Protein-Protein Interaction Networks 1 Department of Medical Chemistry Semmelweis University, Budapest, Hungary www.linkgroup.hu csermely@eok.sote.hu Advantages of multi-disciplinarity Networks have general
Protein-Protein Interaction Networks 1 Department of Medical Chemistry Semmelweis University, Budapest, Hungary www.linkgroup.hu csermely@eok.sote.hu Advantages of multi-disciplinarity Networks have general
