Improving Gene Functional Analysis in Ethylene-induced Leaf Abscission using GO and ProteInOn
|
|
|
- Mariah Bradford
- 6 years ago
- Views:
Transcription
1 Improving Gene Functional Analysis in Ethylene-induced Leaf Abscission using GO and ProteInOn Sara Domingos 1, Cátia Pesquita 2, Francisco M. Couto 2, Luis F. Goulao 3, Cristina Oliveira 1 1 Instituto Superior de Agronomia, Universidade Técnica de Lisboa, Tapada da Ajuda, Lisboa, Portugal saradomingos@gmail.com, crismoniz@isa.utl.pt 2 Faculdade de Ciências, Universidade de Lisboa, Campo Grande, Lisboa, Portugal cpesquita@xldb.di.fc.ul.pt, fcouto@di.fc.ul.pt 3 Eco-Bio, Instituto de Investigação Científica Tropical IP, Av. da República, Quinta do Marquês, Oeiras, Portugal goulao@iict.pt Abstract. In this study we apply a new strategy to select candidate genes involved in the abscission of citrus leaves based on the identification of the most meaningful functional aspects of differentially expressed genes using ProteInOn, a Gene Ontology (GO) based web tool. We submitted a previously analyzed dataset obtained from a microarray experiment to the tool, to obtain the most meaningful GO terms annotated to the given gene products, highlighting the relevant functional processes involved with a greater detail than the original analysis and supporting identification of the genes more likely to be involved in the studied phenomenon. This strategy proved to be effective by complementing the previous analysis with a wider knowledge about the biological processes and gene products more relevant to ethylene-induced leaf abscission. Keywords: ethylene-induced leaf abscission, Citrus clementina, functional analysis, meaningful terms, semantic similarity. 1 Introduction Abscission is a cell separation process by which plant organs are detached, by reduction of cell adhesion and cell wall disassembly in specialized cells, named the abscission zone, and represents an important agricultural phenomenon which affects final yield [1]. The accumulation of data generated by high-throughput techniques of gene expression analysis requires bioinformatics tools and defined vocabularies to describe genes functions that allow its integration and computation. This study aims to improve functional analysis of a list of differentially expressed genes, using the Gene Ontology (GO) and the ProteInOn web tool. To perform it, we
2 used microarray experiment results obtained in [1] where a functional categorization from MIPS (Munich Information Center for Protein Sequences) was applied. MIPS functional catalogue database (FunCat) provides hierarchical information derived from a few species [2] assigning a p-value for each category. This requires translation into Arabidopsis thaliana orthologues. Nowadays, functional categorization of transcripts is mostly based on GO which provides a controlled vocabulary to describe protein function for all eukaryotes, based on 3 orthogonal ontologies: biological processes, cellular components and molecular functions [3]. To analyze the gene set, we chose ProteInOn, a GO-based web tool focused on identifying the most meaningful functional aspects of related gene products [4]. It uses the hierarchical structure of GO to find representative terms, while typical enrichment strategies use direct annotations and usually fail to find general but meaningful annotations. ProteInOn also measures the specificity of each GO term based on information content (IC) and calculates semantic similarity (SSM) which is a function that, given two sets of terms annotating two entities, returns a value reflecting the closeness in meaning between them. There are more than other 68 enrichment analysis tools available (e.g. Onto-Express, FatiGO), based on the premise that co-functioning genes should have a higher potential to be selected, if a biological process is irregular in a study [6]. Our main goal was to apply the functional analysis strategy detailed in [7] and to compare these results with the original ones, while comparing different gene products functional classifications, GO and FunCat. By using updated databases and bioinformatics tools to analyze previously results, we aim to provide insight into the transcriptome abscission-related to support further experiments. 2 Methods The data set used for functional analysis was obtained from [1]. The authors aimed to identify and classify genes involved in the ethylene-induced leaf abscission process in Citrus clementina, and used a cdna microarray to analyze transcriptome of laminar abscission zone (LAZ) and petiole of leaf explants (Pet). The functional analysis strategy applied here was as follows: i) A list of genes was organized according to moment and organ where genes were over-represented [1] and translated into UniProtKB accession numbers. ii) GO annotations were analyzed through ProteInOn, using UniProtKB accession numbers as input. For genes absent in UniProtKB database, the code of a putative A.th orthologue was used. iii) The ten most meaningful GO terms were selected. Meaningfulness was given by the term representativity measure (e-value) and calculated using the global
3 frequency of each GO term annotation in the database as an estimator of its probability of occurrence. For each GO term, a protein taken randomly from the dataset is a random event with two outcomes: success if annotated to t, or failure. The probability of observing at least k successes in a random set of n gene products is given by the probability of success P(t) (1). IC values were calculated using (2), where f(t) is the frequency of annotation of the term t [7]. n P (x1 k) = Σ ( i n ) P(t) I (1-p(t)) n-i. i=k (1) IC(t) = log 2 f(t). iv. The SSM between selected gene products was calculated with simgic measure [5], which has been shown to outperform other measures. SimGIC is given by the sum of the IC of each shared term by two gene products, A and B, divided by the sum of the IC of all their terms (3): (2) simgic (A,B) = Σ te{go(a) GO(B)} IC(t) Σ te{go(a)ugo(b)}ic(t). (3) v. Gene products with SSM > 80% were clustered and the remaining ones were analyzed. vi. GO annotations for both the clusters and relevant individual gene products were analyzed and compared with their previous MIPS classification. 3 Results The list of differentially expressed genes in LAZ and Pet was divided by the two main moments after ethylene treatment (Supplementary Data 1). In the first moment (6h and 12h after treatment) ten differently expressed genes were found in the LAZ. In the second moment (24h and 36 h after treatment) 23 over-represented genes were found in the LAZ and 31 in the Pet, three of them in common with the first phase. The initial set was composed by 61 gene products of which 56 had UniProtKB accession number, 52 had annotations on GO, of which 84% were inferred from electronic annotations. The authors of the experimental assay associated each over-represented transcript to the matching gene, first identified in a single species, which we used as input in UniProtKB, instead of finding the A.th orthologue to use MIPS [1]. From the term representativity analysis we obtained a list of relevant GO terms ranked by e-value. Figure 1 and Supplementary Data 2 show the ten most meaningful biological process terms of each set, since it is the most interesting GO type for the identification of the relevant genes in a biological phenomenon. For the LAZ, in the
4 first moment, the most meaningful GO terms had a low IC, but in the second moment we found annotations with a higher specificity, such as lipid transport and cell wall organization (Supplementary Data 2). Figure 1 shows the ten most meaningful terms of over-represented gene products in the Pet, which have a high IC and relevant biologic meaning for the abscission process, as chitin and other cell wall molecular catabolic processes, cell wall biosynthesis, polysaccharide metabolism, response to stress and oxidative stress and response to stimulus and chemical stimulus. We calculated the SSM for the subset of gene products that are annotated with these meaningful GO terms and built clusters of gene products with SSM > 80% (Table 1). Relevant GO terms associated to remaining gene products were also selected (Supplementary Data 3). Then the annotations of the selected gene products were analyzed (Table 2). In the LAZ, in the first phase, we found gene products involved in photosynthetic processes, response to biotic and abiotic stimulus, protein metabolism, lipid metabolism and floral organ abscission, while in the second phase gene products were related to carbohydrate and protein metabolism, molecular transport and cell wall organization (Tables 1, 2 and Supplementary Data 3). In the PET, there was more processes involved (Supplementary Data 4): DNA transcription, cell wall organization and biosynthesis, response to abiotic stress, oxidation-reduction, hormonal regulation, protein metabolism, molecular catabolism and transport Frequency of the 10 most meaningful GO terms Term Id Term Name IC GO: chitin catabolic process 0.45 GO: response to oxidative stress 0.37 GO: cell wall macromolecule catabolic p GO: response to chemical stimulus 0.25 GO: cell wall organization or biogenesis 0.27 GO: polysaccharide metabolic process 0.24 GO: response to stress 0.19 GO: response to stimulus 0.16 GO: catabolic process 0.20 GO: cellular response to chemical stimulus 0.39 Fig. 1. Ten most meaningful GO terms and corresponding IC, of biological process type, annotated to gene products over-represented in Pet, in the second moment (Source: ProteInOn). Table 1. Clusters of gene products over-represented in LAZ e Pet. LAZ/ second moment Pet/ second moment Clusters of gene products (Uniprot ac. number) Term name Q6EV47, Q8L5S8 lipid transport Q81746, Q94C86 cellular cell wall organization A0MKC8, Q8L4F2, Q9SF40 translation O82547, Q43752, Q8H985, Q8H986 A9PHA0, A9UFX7, Q0ZA67 chitin catabolic process cell wall macromolecule catabolic proc. response to oxidative stress oxidation-reduction process
5 Table 2. Functional characterization of isolated gene products over-represented in LAZ, in the first moment after ethylene treatment, of biological process GO terms type. LAZ/ first moment Term Name O regulation of stomatal complex patterning regulation of stomatal complex development floral organ abscission jasmonic acid metabolic process plant-type hypersensitive response defense response, incompatible interaction oxylipin biosynthetic process protein phosphorylation response to wounding response to stress Q 8 L 6 Y 9 Q 9 A R Discussion and Conclusion Using the current versions of UniProtKB and GO annotations, we obtained an improvement on the coverage of the initial gene list with only four terms without GO annotations associated comparing with MIPS categorization [1]. Over-represented gene products with manual annotations were 15.38% of the transcripts, which probably means that the studied phenomenon has been gaining interest. By selecting the clusters of gene products with high SSM and the relevant isolated gene products, we found gene products with an interesting set of annotations for ethylene-induced leaf abscission. Analyzing this restricted list, for example in the Pet, in the second moment after ethylene treatment (Supplementary Data 4) we found that they are associated with other important processes such as hormonal regulation, molecular transport, DNA transcription, protein metabolism, aromatic compounds metabolism and organ formation. In the last phase of abscission, a protective layer is generated in the organ that remains attached to the plant [1], which can be related with the gene product annotated with organ formation term. MIPS categorization indicated that transcriptional activation during abscission was involved in defense, transport mechanisms, signal transduction and protein biosynthesis, comparing it with our strategy we obtained a more comprehensive characterization, including cell wall biosynthesis, hormonal regulation and biosynthesis, protein metabolism, catabolic process, molecular transport and response to biotic/abiotic stresses, and other detailed information. The adopted strategy using ProteInOn, with the empirical choice of the ten most meaningful terms and clustering based on SSM > 80%, proved to be effective in providing a wider understanding of the process under study. A gene product that seems to have a key role is ATMKK4 (O80397) which is annotated with very relevant terms: floral organ abscission, stomatal complex patterning and
6 development regulation, plant-type hypersensitive response, defense, response to stress and protein phosphorylation. For this specific gene product, and to account for the bias in basing our analysis in a GO 2011 version and comparing it to results obtained with a MIPS 2008 version, we investigated its functional description in MIPS 2011 and GO In MIPS 2011 it is related to defense, response to biotic stimulus, protein phosporylation and protein kinase function, while in GO 2008 it is annotated with sensitive response, protein phosphorylation and defense. Hence, GO did gain relevant information about this gene since 2008, while MIPS did not. By reanalyzing previously obtained results with more up to date bioinformatics tools and databases, we were able to improve the functional analysis of differentially expressed genes and the identification of gene products relevant to the studied phenomenon. This strategy can be an important tool to guide future experiments, saving time and materials by avoiding experiment replication. Supplementary Data. Supplementary data are available at Acknowldegements. This work was supported by the Portuguese Fundação para a Ciência e Tecnologia through the Multiannual Funding Programme for LaSIGE, and the PhD grants SFRH/BD/42481/2007 and SFRH/BD/69076/ References 1. Agusti, J., Merelo, P., Cercós, M., Tadeo, F.R., Talón, M.: Ethylene-induced Differential Gene Expression during Abscission of Citrus Leaves. J. Exp. Bot. 59, (2008) 2. Ruepp, A., Zollner, A., Maier, D., Albermann, K., Hani, J., Mokrejs, M., Tetko, I., Guldener, U., Mannhaupt, G., Munsterkotter, M., Mewes, H.W.: The FunCat, a Functional Annotation Scheme for Systematic Classification of Proteins from Whole Genomes. Nucleic Acids Res. 32, (2004) 3. The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nat. Genet. 25(1):25-29 (2000) 4. Faria, D., Pesquita, C., Couto, F., Falcao, A.,: ProteInOn: A Web Tool for Protein Semantic Similarity. DI/FCUL TR 07-6, Department of Informatics, University of Lisbon (2007) 5. Pesquita C, Faria D, Bastos H, Falcão AO, Couto F.: Evaluating GO-based Semantic Similarity Measures. ISMB/ECCB 2007 SIG Meeting Program Materials, International Society for Computational Biology (2007) 6. Huang, daw., Sherman, B.T., Lempicki, R.A.: Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37,1-13 (2009) 7. Bastos, H., Tavares, B., Pesquita, C., Faria, D., Couto F.: Application of Gene Ontology to Gene Identification. In: Yu, B., Hinchcliffe, M., (eds) In Silico Tools for Gene Discovery. Springer Science (in press)
Francisco M. Couto Mário J. Silva Pedro Coutinho
 Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
Mixture models for analysing transcriptome and ChIP-chip data
 Mixture models for analysing transcriptome and ChIP-chip data Marie-Laure Martin-Magniette French National Institute for agricultural research (INRA) Unit of Applied Mathematics and Informatics at AgroParisTech,
Mixture models for analysing transcriptome and ChIP-chip data Marie-Laure Martin-Magniette French National Institute for agricultural research (INRA) Unit of Applied Mathematics and Informatics at AgroParisTech,
Network by Weighted Graph Mining
 2012 4th International Conference on Bioinformatics and Biomedical Technology IPCBEE vol.29 (2012) (2012) IACSIT Press, Singapore + Prediction of Protein Function from Protein-Protein Interaction Network
2012 4th International Conference on Bioinformatics and Biomedical Technology IPCBEE vol.29 (2012) (2012) IACSIT Press, Singapore + Prediction of Protein Function from Protein-Protein Interaction Network
4CitySemantics. GIS-Semantic Tool for Urban Intervention Areas
 4CitySemantics GIS-Semantic Tool for Urban Intervention Areas Nuno MONTENEGRO 1 ; Jorge GOMES 2 ; Paulo URBANO 2 José P. DUARTE 1 1 Faculdade de Arquitectura da Universidade Técnica de Lisboa, Rua Sá Nogueira,
4CitySemantics GIS-Semantic Tool for Urban Intervention Areas Nuno MONTENEGRO 1 ; Jorge GOMES 2 ; Paulo URBANO 2 José P. DUARTE 1 1 Faculdade de Arquitectura da Universidade Técnica de Lisboa, Rua Sá Nogueira,
Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis
 Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Integration of functional genomics data
 Integration of functional genomics data Laboratoire Bordelais de Recherche en Informatique (UMR) Centre de Bioinformatique de Bordeaux (Plateforme) Rennes Oct. 2006 1 Observations and motivations Genomics
Integration of functional genomics data Laboratoire Bordelais de Recherche en Informatique (UMR) Centre de Bioinformatique de Bordeaux (Plateforme) Rennes Oct. 2006 1 Observations and motivations Genomics
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes.
 I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs
 Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón
 GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
Virginia Western Community College BIO 101 General Biology I
 BIO 101 General Biology I Prerequisites Successful completion of MTE 1, 2, 3, 4, and 5; and a placement recommendation for ENG 111, co-enrollment in ENF 3/ENG 111, or successful completion of all developmental
BIO 101 General Biology I Prerequisites Successful completion of MTE 1, 2, 3, 4, and 5; and a placement recommendation for ENG 111, co-enrollment in ENF 3/ENG 111, or successful completion of all developmental
I. Molecules & Cells. A. Unit One: The Nature of Science. B. Unit Two: The Chemistry of Life. C. Unit Three: The Biology of the Cell.
 I. Molecules & Cells A. Unit One: The Nature of Science a. How is the scientific method used to solve problems? b. What is the importance of controls? c. How does Darwin s theory of evolution illustrate
I. Molecules & Cells A. Unit One: The Nature of Science a. How is the scientific method used to solve problems? b. What is the importance of controls? c. How does Darwin s theory of evolution illustrate
Compare and contrast the cellular structures and degrees of complexity of prokaryotic and eukaryotic organisms.
 Subject Area - 3: Science and Technology and Engineering Education Standard Area - 3.1: Biological Sciences Organizing Category - 3.1.A: Organisms and Cells Course - 3.1.B.A: BIOLOGY Standard - 3.1.B.A1:
Subject Area - 3: Science and Technology and Engineering Education Standard Area - 3.1: Biological Sciences Organizing Category - 3.1.A: Organisms and Cells Course - 3.1.B.A: BIOLOGY Standard - 3.1.B.A1:
Toponym Disambiguation using Ontology-based Semantic Similarity
 Toponym Disambiguation using Ontology-based Semantic Similarity David S Batista 1, João D Ferreira 2, Francisco M Couto 2, and Mário J Silva 1 1 IST/INESC-ID Lisbon, Portugal {dsbatista,msilva}@inesc-id.pt
Toponym Disambiguation using Ontology-based Semantic Similarity David S Batista 1, João D Ferreira 2, Francisco M Couto 2, and Mário J Silva 1 1 IST/INESC-ID Lisbon, Portugal {dsbatista,msilva}@inesc-id.pt
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
AP BIOLOGY SUMMER ASSIGNMENT
 AP BIOLOGY SUMMER ASSIGNMENT Welcome to EDHS Advanced Placement Biology! The attached summer assignment is required for all AP Biology students for the 2011-2012 school year. The assignment consists of
AP BIOLOGY SUMMER ASSIGNMENT Welcome to EDHS Advanced Placement Biology! The attached summer assignment is required for all AP Biology students for the 2011-2012 school year. The assignment consists of
Biology Assessment. Eligible Texas Essential Knowledge and Skills
 Biology Assessment Eligible Texas Essential Knowledge and Skills STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules
Biology Assessment Eligible Texas Essential Knowledge and Skills STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules
STAAR Biology Assessment
 STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules as building blocks of cells, and that cells are the basic unit of
STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules as building blocks of cells, and that cells are the basic unit of
2 GENE FUNCTIONAL SIMILARITY. 2.1 Semantic values of GO terms
 Bioinformatics Advance Access published March 7, 2007 The Author (2007). Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Bioinformatics Advance Access published March 7, 2007 The Author (2007). Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Inductive logic programming for gene regulation prediction
 Mach Learn (2008) 70: 225 240 DOI 10.1007/s10994-007-5037-3 Inductive logic programming for gene regulation prediction Sebastian Fröhler Stefan Kramer Received: 10 October 2007 / Accepted: 11 October 2007
Mach Learn (2008) 70: 225 240 DOI 10.1007/s10994-007-5037-3 Inductive logic programming for gene regulation prediction Sebastian Fröhler Stefan Kramer Received: 10 October 2007 / Accepted: 11 October 2007
Data visualization and clustering: an application to gene expression data
 Data visualization and clustering: an application to gene expression data Francesco Napolitano Università degli Studi di Salerno Dipartimento di Matematica e Informatica DAA Erice, April 2007 Thanks to
Data visualization and clustering: an application to gene expression data Francesco Napolitano Università degli Studi di Salerno Dipartimento di Matematica e Informatica DAA Erice, April 2007 Thanks to
Bio 101 General Biology 1
 Revised: Fall 2016 Bio 101 General Biology 1 COURSE OUTLINE Prerequisites: Prerequisite: Successful completion of MTE 1, 2, 3, 4, and 5, and a placement recommendation for ENG 111, co-enrollment in ENF
Revised: Fall 2016 Bio 101 General Biology 1 COURSE OUTLINE Prerequisites: Prerequisite: Successful completion of MTE 1, 2, 3, 4, and 5, and a placement recommendation for ENG 111, co-enrollment in ENF
2. Cellular and Molecular Biology
 2. Cellular and Molecular Biology 2.1 Cell Structure 2.2 Transport Across Cell Membranes 2.3 Cellular Metabolism 2.4 DNA Replication 2.5 Cell Division 2.6 Biosynthesis 2.1 Cell Structure What is a cell?
2. Cellular and Molecular Biology 2.1 Cell Structure 2.2 Transport Across Cell Membranes 2.3 Cellular Metabolism 2.4 DNA Replication 2.5 Cell Division 2.6 Biosynthesis 2.1 Cell Structure What is a cell?
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
A transcriptome meta-analysis identifies the response of plant to stresses. Etienne Delannoy, Rim Zaag, Guillem Rigaill, Marie-Laure Martin-Magniette
 A transcriptome meta-analysis identifies the response of plant to es Etienne Delannoy, Rim Zaag, Guillem Rigaill, Marie-Laure Martin-Magniette Biological context Multiple biotic and abiotic es impacting
A transcriptome meta-analysis identifies the response of plant to es Etienne Delannoy, Rim Zaag, Guillem Rigaill, Marie-Laure Martin-Magniette Biological context Multiple biotic and abiotic es impacting
Course Descriptions Biology
 Course Descriptions Biology BIOL 1010 (F/S) Human Anatomy and Physiology I. An introductory study of the structure and function of the human organ systems including the nervous, sensory, muscular, skeletal,
Course Descriptions Biology BIOL 1010 (F/S) Human Anatomy and Physiology I. An introductory study of the structure and function of the human organ systems including the nervous, sensory, muscular, skeletal,
Miller & Levine Biology 2010
 A Correlation of 2010 to the Pennsylvania Assessment Anchors Grades 9-12 INTRODUCTION This document demonstrates how 2010 meets the Pennsylvania Assessment Anchors, grades 9-12. Correlation page references
A Correlation of 2010 to the Pennsylvania Assessment Anchors Grades 9-12 INTRODUCTION This document demonstrates how 2010 meets the Pennsylvania Assessment Anchors, grades 9-12. Correlation page references
Genome-Scale Gene Function Prediction Using Multiple Sources of High-Throughput Data in Yeast Saccharomyces cerevisiae ABSTRACT
 OMICS A Journal of Integrative Biology Volume 8, Number 4, 2004 Mary Ann Liebert, Inc. Genome-Scale Gene Function Prediction Using Multiple Sources of High-Throughput Data in Yeast Saccharomyces cerevisiae
OMICS A Journal of Integrative Biology Volume 8, Number 4, 2004 Mary Ann Liebert, Inc. Genome-Scale Gene Function Prediction Using Multiple Sources of High-Throughput Data in Yeast Saccharomyces cerevisiae
Slide 1 / Describe the setup of Stanley Miller s experiment and the results. What was the significance of his results?
 Slide 1 / 57 1 Describe the setup of Stanley Miller s experiment and the results. What was the significance of his results? Slide 2 / 57 2 Explain how dehydration synthesis and hydrolysis are related.
Slide 1 / 57 1 Describe the setup of Stanley Miller s experiment and the results. What was the significance of his results? Slide 2 / 57 2 Explain how dehydration synthesis and hydrolysis are related.
Marine Resources Development Foundation/MarineLab Grades: 9, 10, 11, 12 States: AP Biology Course Description Subjects: Science
 Marine Resources Development Foundation/MarineLab Grades: 9, 10, 11, 12 States: AP Biology Course Description Subjects: Science Highlighted components are included in Tallahassee Museum s 2016 program
Marine Resources Development Foundation/MarineLab Grades: 9, 10, 11, 12 States: AP Biology Course Description Subjects: Science Highlighted components are included in Tallahassee Museum s 2016 program
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Supplementary Information
 Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Outline. Terminologies and Ontologies. Communication and Computation. Communication. Outline. Terminologies and Vocabularies.
 Page 1 Outline 1. Why do we need terminologies and ontologies? Terminologies and Ontologies Iwei Yeh yeh@smi.stanford.edu 04/16/2002 2. Controlled Terminologies Enzyme Classification Gene Ontology 3. Ontologies
Page 1 Outline 1. Why do we need terminologies and ontologies? Terminologies and Ontologies Iwei Yeh yeh@smi.stanford.edu 04/16/2002 2. Controlled Terminologies Enzyme Classification Gene Ontology 3. Ontologies
Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport
 Ph.D. thesis Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport Zsigmond Laura Supervisor: Dr. Szabados László Arabidopsis Molecular Genetic Group Institute of Plant
Ph.D. thesis Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport Zsigmond Laura Supervisor: Dr. Szabados László Arabidopsis Molecular Genetic Group Institute of Plant
Computational Genomics. Reconstructing dynamic regulatory networks in multiple species
 02-710 Computational Genomics Reconstructing dynamic regulatory networks in multiple species Methods for reconstructing networks in cells CRH1 SLT2 SLR3 YPS3 YPS1 Amit et al Science 2009 Pe er et al Recomb
02-710 Computational Genomics Reconstructing dynamic regulatory networks in multiple species Methods for reconstructing networks in cells CRH1 SLT2 SLR3 YPS3 YPS1 Amit et al Science 2009 Pe er et al Recomb
COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry.
 North Carolina Draft Standard Course of Study and Grade Level Competencies, Biology BIOLOGY COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry. 1.01
North Carolina Draft Standard Course of Study and Grade Level Competencies, Biology BIOLOGY COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry. 1.01
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
October 08-11, Co-Organizer Dr. S D Samantaray Professor & Head,
 A Journey towards Systems system Biology biology :: Biocomputing of of Hi-throughput Omics omics Data data October 08-11, 2018 Coordinator Dr. Anil Kumar Gaur Professor & Head Department of Molecular Biology
A Journey towards Systems system Biology biology :: Biocomputing of of Hi-throughput Omics omics Data data October 08-11, 2018 Coordinator Dr. Anil Kumar Gaur Professor & Head Department of Molecular Biology
An introduction to SYSTEMS BIOLOGY
 An introduction to SYSTEMS BIOLOGY Paolo Tieri CNR Consiglio Nazionale delle Ricerche, Rome, Italy 10 February 2015 Universidade Federal de Minas Gerais, Belo Horizonte, Brasil Course outline Day 1: intro
An introduction to SYSTEMS BIOLOGY Paolo Tieri CNR Consiglio Nazionale delle Ricerche, Rome, Italy 10 February 2015 Universidade Federal de Minas Gerais, Belo Horizonte, Brasil Course outline Day 1: intro
EASTERN ARIZONA COLLEGE General Biology I
 EASTERN ARIZONA COLLEGE General Biology I Course Design 2013-2014 Course Information Division Science Course Number BIO 181 (SUN# BIO 1181) Title General Biology I Credits 4 Developed by David J. Henson
EASTERN ARIZONA COLLEGE General Biology I Course Design 2013-2014 Course Information Division Science Course Number BIO 181 (SUN# BIO 1181) Title General Biology I Credits 4 Developed by David J. Henson
Field 045: Science Life Science Assessment Blueprint
 Field 045: Science Life Science Assessment Blueprint Domain I Foundations of Science 0001 The Nature and Processes of Science (Standard 1) 0002 Central Concepts and Connections in Science (Standard 2)
Field 045: Science Life Science Assessment Blueprint Domain I Foundations of Science 0001 The Nature and Processes of Science (Standard 1) 0002 Central Concepts and Connections in Science (Standard 2)
Vance County Early College High School Pacing Guide Course: Introduction to Biology (Semester I)
 Vance County Early College High School Pacing Guide Course: Introduction to Biology (Semester I) Week(s ) Dates Unit Unit Title Essential Questions / Topic Questions 1 1 Introduction to Biology 1. How
Vance County Early College High School Pacing Guide Course: Introduction to Biology (Semester I) Week(s ) Dates Unit Unit Title Essential Questions / Topic Questions 1 1 Introduction to Biology 1. How
Total
 Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
SPRINGFIELD TECHNICAL COMMUNITY COLLEGE ACADEMIC AFFAIRS
 SPRINGFIELD TECHNICAL COMMUNITY COLLEGE ACADEMIC AFFAIRS Course Number: BIOL 102 Department: Biological Sciences Course Title: Principles of Biology 1 Semester: Spring Year: 1997 Objectives/ 1. Summarize
SPRINGFIELD TECHNICAL COMMUNITY COLLEGE ACADEMIC AFFAIRS Course Number: BIOL 102 Department: Biological Sciences Course Title: Principles of Biology 1 Semester: Spring Year: 1997 Objectives/ 1. Summarize
What is Systems Biology
 What is Systems Biology 2 CBS, Department of Systems Biology 3 CBS, Department of Systems Biology Data integration In the Big Data era Combine different types of data, describing different things or the
What is Systems Biology 2 CBS, Department of Systems Biology 3 CBS, Department of Systems Biology Data integration In the Big Data era Combine different types of data, describing different things or the
Unit 1: Chemistry of Life Guided Reading Questions (80 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 1 Exploring Life Unit 1: Chemistry of Life Guided Reading Questions
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 1 Exploring Life Unit 1: Chemistry of Life Guided Reading Questions
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 SUMOylation of proteins changes drastically upon heat shock, MG-132 treatment and PR-619 treatment. (a) Schematic overview of all SUMOylation proteins identified to be differentially
Supplementary Figure 1 SUMOylation of proteins changes drastically upon heat shock, MG-132 treatment and PR-619 treatment. (a) Schematic overview of all SUMOylation proteins identified to be differentially
Contra Costa College Course Outline
 Contra Costa College Course Outline Department & Number: BIOSC 110 Course Title: Introduction to Biological Science Pre-requisite: None Corequisite: None Advisory: None Entry Skill: None Lecture Hours:
Contra Costa College Course Outline Department & Number: BIOSC 110 Course Title: Introduction to Biological Science Pre-requisite: None Corequisite: None Advisory: None Entry Skill: None Lecture Hours:
Goal 1: Develop knowledge and understanding of core content in biology
 Indiana State University» College of Arts & Sciences» Biology BA/BS in Biology Standing Requirements s Library Goal 1: Develop knowledge and understanding of core content in biology 1: Illustrate and examine
Indiana State University» College of Arts & Sciences» Biology BA/BS in Biology Standing Requirements s Library Goal 1: Develop knowledge and understanding of core content in biology 1: Illustrate and examine
ADVANCED PLACEMENT BIOLOGY
 ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
Peddie Summer Day School
 Peddie Summer Day School Course Syllabus: BIOLOGY Teacher: Mr. Jeff Tuliszewski Text: Biology by Miller and Levine, Prentice Hall, 2010 edition ISBN 9780133669510 Guided Reading Workbook for Biology ISBN
Peddie Summer Day School Course Syllabus: BIOLOGY Teacher: Mr. Jeff Tuliszewski Text: Biology by Miller and Levine, Prentice Hall, 2010 edition ISBN 9780133669510 Guided Reading Workbook for Biology ISBN
BIOLOGY IB DIPLOMA PROGRAM PARENTS ORIENTATION DAY
 BIOLOGY IB DIPLOMA PROGRAM PARENTS ORIENTATION DAY 2018-2019 ADITYA NUGRAHA WARDHANA EDUCATION BACKGROUND: Bachelor of Science, Biology major University of Indonesia, Depok WORK EXPERIENCE: Laboratory
BIOLOGY IB DIPLOMA PROGRAM PARENTS ORIENTATION DAY 2018-2019 ADITYA NUGRAHA WARDHANA EDUCATION BACKGROUND: Bachelor of Science, Biology major University of Indonesia, Depok WORK EXPERIENCE: Laboratory
Supplemental Material
 Supplemental Material Protocol S1 The PromPredict algorithm considers the average free energy over a 100nt window and compares it with the the average free energy over a downstream 100nt window separated
Supplemental Material Protocol S1 The PromPredict algorithm considers the average free energy over a 100nt window and compares it with the the average free energy over a downstream 100nt window separated
Abiotic Stress in Crop Plants
 1 Abiotic Stress in Crop Plants Mirza Hasanuzzaman, PhD Professor Department of Agronomy Sher-e-Bangla Agricultural University E-mail: mhzsauag@yahoo.com Stress Stress is usually defined as an external
1 Abiotic Stress in Crop Plants Mirza Hasanuzzaman, PhD Professor Department of Agronomy Sher-e-Bangla Agricultural University E-mail: mhzsauag@yahoo.com Stress Stress is usually defined as an external
Text of objective. Investigate and describe the structure and functions of cells including: Cell organelles
 This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
86 Part 4 SUMMARY INTRODUCTION
 86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
Curriculum Map. Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1)
 1 Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1) Focus Standards BIO1.LS1.2 Evaluate comparative models of various cell types with a focus on organic molecules
1 Biology, Quarter 1 Big Ideas: From Molecules to Organisms: Structures and Processes (BIO1.LS1) Focus Standards BIO1.LS1.2 Evaluate comparative models of various cell types with a focus on organic molecules
MINISYMPOSIUM LOGICAL MODELLING OF (MULTI)CELLULAR NETWORKS
 Friday, July 27th, 11:00 MINISYMPOSIUM LOGICAL MODELLING OF (MULTI)CELLULAR NETWORKS Organizer Pedro T. Monteiro INESC-ID/IST - Univ. de Lisboa 1000-029 Lisboa, PT Pedro.Tiago.Monteiro@tecnico.ulisboa.pt
Friday, July 27th, 11:00 MINISYMPOSIUM LOGICAL MODELLING OF (MULTI)CELLULAR NETWORKS Organizer Pedro T. Monteiro INESC-ID/IST - Univ. de Lisboa 1000-029 Lisboa, PT Pedro.Tiago.Monteiro@tecnico.ulisboa.pt
Miller & Levine Biology
 A Correlation of To the Science Biology A Correlation of, 2014 to the, Table of Contents From Molecules to Organisms: Structures and Processes... 3 Ecosystems: Interactions, Energy, and Dynamics... 4 Heredity:
A Correlation of To the Science Biology A Correlation of, 2014 to the, Table of Contents From Molecules to Organisms: Structures and Processes... 3 Ecosystems: Interactions, Energy, and Dynamics... 4 Heredity:
VCE BIOLOGY Relationship between the key knowledge and key skills of the Study Design and the Study Design
 VCE BIOLOGY 2006 2014 Relationship between the key knowledge and key skills of the 2000 2005 Study Design and the 2006 2014 Study Design The following table provides a comparison of the key knowledge (and
VCE BIOLOGY 2006 2014 Relationship between the key knowledge and key skills of the 2000 2005 Study Design and the 2006 2014 Study Design The following table provides a comparison of the key knowledge (and
Modesto Junior College Course Outline of Record BIO 101
 Modesto Junior College Course Outline of Record BIO 101 I. OVERVIEW The following information will appear in the 2010-2011 catalog BIO 101 Biological Principles 5 Units Prerequisite: Satisfactory completion
Modesto Junior College Course Outline of Record BIO 101 I. OVERVIEW The following information will appear in the 2010-2011 catalog BIO 101 Biological Principles 5 Units Prerequisite: Satisfactory completion
GACE Biology Assessment Test I (026) Curriculum Crosswalk
 Subarea I. Cell Biology: Cell Structure and Function (50%) Objective 1: Understands the basic biochemistry and metabolism of living organisms A. Understands the chemical structures and properties of biologically
Subarea I. Cell Biology: Cell Structure and Function (50%) Objective 1: Understands the basic biochemistry and metabolism of living organisms A. Understands the chemical structures and properties of biologically
Area of Focus: Biology. Learning Objective 1: Describe the structure and function of organs. Pre-Learning Evaluation: Teaching Methods and Process:
 Area of Focus: Biology Learning Objective 1: Describe the structure and function of organs. Pre- Diagram and label the structure of the primary components of representative organs in plants and animals
Area of Focus: Biology Learning Objective 1: Describe the structure and function of organs. Pre- Diagram and label the structure of the primary components of representative organs in plants and animals
Introduction to Biology
 Introduction to Biology Course Description Introduction to Biology is an introductory course in the biological sciences. Topics included are biological macromolecules, cell biology and metabolism, DNA
Introduction to Biology Course Description Introduction to Biology is an introductory course in the biological sciences. Topics included are biological macromolecules, cell biology and metabolism, DNA
Name Date Period Unit 1 Basic Biological Principles 1. What are the 7 characteristics of life?
 Unit 1 Basic Biological Principles 1. What are the 7 characteristics of life? Eukaryotic cell parts you should be able a. to identify and label: Nucleus b. Nucleolus c. Rough/smooth ER Ribosomes d. Golgi
Unit 1 Basic Biological Principles 1. What are the 7 characteristics of life? Eukaryotic cell parts you should be able a. to identify and label: Nucleus b. Nucleolus c. Rough/smooth ER Ribosomes d. Golgi
CATEGORY a TERM COUNT b P VALUE GENE FAMILIES: REPRESENTATIVE GENE SYMBOLS c. Annotation Cluster 1 Enrichment Score: 1.
 Table S7 GO term and functional classification enrichment analysis using DAVID for gene families that are expanded in the Drosophila suzukii genome as compared to 14 Drosophila species analyzed in this
Table S7 GO term and functional classification enrichment analysis using DAVID for gene families that are expanded in the Drosophila suzukii genome as compared to 14 Drosophila species analyzed in this
Local topological signatures for network-based prediction of biological function
 Local topological signatures for network-based prediction of biological function Wynand Winterbach 1,2, Piet Van Mieghem 1, Marcel Reinders 2,3,4, Huijuan Wang 1, and Dick de Ridder 2,3,4 1 Network Architectures
Local topological signatures for network-based prediction of biological function Wynand Winterbach 1,2, Piet Van Mieghem 1, Marcel Reinders 2,3,4, Huijuan Wang 1, and Dick de Ridder 2,3,4 1 Network Architectures
Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function
 Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function Brian D. Gregory Department of Biology Penn Genome Frontiers Institute University of Pennsylvania
Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function Brian D. Gregory Department of Biology Penn Genome Frontiers Institute University of Pennsylvania
Ph.D. thesis. Study of proline accumulation and transcriptional regulation of genes involved in this process in Arabidopsis thaliana
 Ph.D. thesis Study of proline accumulation and transcriptional regulation of genes involved in this process in Arabidopsis thaliana Written by: Edit Ábrahám Temesváriné Supervisors: Dr. László Szabados
Ph.D. thesis Study of proline accumulation and transcriptional regulation of genes involved in this process in Arabidopsis thaliana Written by: Edit Ábrahám Temesváriné Supervisors: Dr. László Szabados
CHAPTER 1 Life: Biological Principles and the Science of Zoology
 CHAPTER 1 Life: Biological Principles and the Science of Zoology 1-1 Zoology: The Uses of Principles The scientific study of animal life Does Life Have Defining Properties? No simple definition The history
CHAPTER 1 Life: Biological Principles and the Science of Zoology 1-1 Zoology: The Uses of Principles The scientific study of animal life Does Life Have Defining Properties? No simple definition The history
2. What properties or characteristics distinguish living organisms? Substance Description Example(s)
 PREIB BIOLOGY FIRST SEMESTER REVIEW (I) 2015-16 Life on Earth 1. Describe the hierarchy of life on Earth from broadest to narrowest category. 2. What properties or characteristics distinguish living organisms?
PREIB BIOLOGY FIRST SEMESTER REVIEW (I) 2015-16 Life on Earth 1. Describe the hierarchy of life on Earth from broadest to narrowest category. 2. What properties or characteristics distinguish living organisms?
Gene Ontology and overrepresentation analysis
 Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim Overview How
Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim Overview How
Computational Structural Bioinformatics
 Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Supplementary Materials for R3P-Loc Web-server
 Supplementary Materials for R3P-Loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to R3P-Loc Server Contents 1 Introduction to R3P-Loc
Supplementary Materials for R3P-Loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to R3P-Loc Server Contents 1 Introduction to R3P-Loc
2012 Biology 1 End-of-Course (EOC) Assessment Form 1
 read How should use of Reports be limited? provided on page 5 of 01 Biology 1 End-of-Course (EOC) Assessment Form 1 SC.91.L.1.1 Cell theory and advances in science 1 SC.91.L.1. Cell membrane; Comparing
read How should use of Reports be limited? provided on page 5 of 01 Biology 1 End-of-Course (EOC) Assessment Form 1 SC.91.L.1.1 Cell theory and advances in science 1 SC.91.L.1. Cell membrane; Comparing
Identify stages of plant life cycle Botany Oral/written pres, exams
 DPI Standards Biology Education (for students) 1. Characteristics of organisms Know Properties of living organisms, including: Acquire and use energy and materials Sense and respond to stimuli Reproduce
DPI Standards Biology Education (for students) 1. Characteristics of organisms Know Properties of living organisms, including: Acquire and use energy and materials Sense and respond to stimuli Reproduce
Measuring Semantic Similarity between Gene Ontology Terms
 Measuring Semantic Similarity between Gene Ontology Terms Francisco M. Couto a Mário J. Silva a Pedro M. Coutinho b a Departamento de Informática, Faculdade de Ciências da Universidade de Lisboa, Portugal
Measuring Semantic Similarity between Gene Ontology Terms Francisco M. Couto a Mário J. Silva a Pedro M. Coutinho b a Departamento de Informática, Faculdade de Ciências da Universidade de Lisboa, Portugal
Function Prediction Using Neighborhood Patterns
 Function Prediction Using Neighborhood Patterns Petko Bogdanov Department of Computer Science, University of California, Santa Barbara, CA 93106 petko@cs.ucsb.edu Ambuj Singh Department of Computer Science,
Function Prediction Using Neighborhood Patterns Petko Bogdanov Department of Computer Science, University of California, Santa Barbara, CA 93106 petko@cs.ucsb.edu Ambuj Singh Department of Computer Science,
Dynamical Modeling in Biology: a semiotic perspective. Junior Barrera BIOINFO-USP
 Dynamical Modeling in Biology: a semiotic perspective Junior Barrera BIOINFO-USP Layout Introduction Dynamical Systems System Families System Identification Genetic networks design Cell Cycle Modeling
Dynamical Modeling in Biology: a semiotic perspective Junior Barrera BIOINFO-USP Layout Introduction Dynamical Systems System Families System Identification Genetic networks design Cell Cycle Modeling
Mixtures and Hidden Markov Models for analyzing genomic data
 Mixtures and Hidden Markov Models for analyzing genomic data Marie-Laure Martin-Magniette UMR AgroParisTech/INRA Mathématique et Informatique Appliquées, Paris UMR INRA/UEVE ERL CNRS Unité de Recherche
Mixtures and Hidden Markov Models for analyzing genomic data Marie-Laure Martin-Magniette UMR AgroParisTech/INRA Mathématique et Informatique Appliquées, Paris UMR INRA/UEVE ERL CNRS Unité de Recherche
Valley Central School District 944 State Route 17K Montgomery, NY Telephone Number: (845) ext Fax Number: (845)
 Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
CSCE555 Bioinformatics. Protein Function Annotation
 CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
Information Extraction from Biomedical Text. BMI/CS 776 Mark Craven
 Information Extraction from Biomedical Text BMI/CS 776 www.biostat.wisc.edu/bmi776/ Mark Craven craven@biostat.wisc.edu Spring 2012 Goals for Lecture the key concepts to understand are the following! named-entity
Information Extraction from Biomedical Text BMI/CS 776 www.biostat.wisc.edu/bmi776/ Mark Craven craven@biostat.wisc.edu Spring 2012 Goals for Lecture the key concepts to understand are the following! named-entity
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA
 INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA XIUFENG WAN xw6@cs.msstate.edu Department of Computer Science Box 9637 JOHN A. BOYLE jab@ra.msstate.edu Department of Biochemistry and Molecular Biology
INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA XIUFENG WAN xw6@cs.msstate.edu Department of Computer Science Box 9637 JOHN A. BOYLE jab@ra.msstate.edu Department of Biochemistry and Molecular Biology
Supplementary Information 16
 Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
Cluster Analysis of Gene Expression Microarray Data. BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002
 Cluster Analysis of Gene Expression Microarray Data BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002 1 Data representations Data are relative measurements log 2 ( red
Cluster Analysis of Gene Expression Microarray Data BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002 1 Data representations Data are relative measurements log 2 ( red
Gabriella Rustici EMBL-EBI
 Towards a cellular phenotype ontology Gabriella Rustici EMBL-EBI gabry@ebi.ac.uk Systems Microscopy NoE at a glance The Systems Microscopy NoE (2011 2015) is a FP7 funded project, involving 15 research
Towards a cellular phenotype ontology Gabriella Rustici EMBL-EBI gabry@ebi.ac.uk Systems Microscopy NoE at a glance The Systems Microscopy NoE (2011 2015) is a FP7 funded project, involving 15 research
Context dependent visualization of protein function
 Article III Context dependent visualization of protein function In: Juho Rousu, Samuel Kaski and Esko Ukkonen (eds.). Probabilistic Modeling and Machine Learning in Structural and Systems Biology. 2006,
Article III Context dependent visualization of protein function In: Juho Rousu, Samuel Kaski and Esko Ukkonen (eds.). Probabilistic Modeling and Machine Learning in Structural and Systems Biology. 2006,
Lowndes County Biology II Pacing Guide Approximate
 Lowndes County Biology II Pacing Guide 2009-2010 MS Frameworks Pacing Guide Worksheet Grade Level: Biology II Grading Period: 1 st 9 weeks Chapter/Unit Lesson Topic Objective Number 1 The Process of 1.
Lowndes County Biology II Pacing Guide 2009-2010 MS Frameworks Pacing Guide Worksheet Grade Level: Biology II Grading Period: 1 st 9 weeks Chapter/Unit Lesson Topic Objective Number 1 The Process of 1.
Miller & Levine Biology 2014
 A Correlation of Miller & Levine Biology To the Essential Standards for Biology High School Introduction This document demonstrates how meets the North Carolina Essential Standards for Biology, grades
A Correlation of Miller & Levine Biology To the Essential Standards for Biology High School Introduction This document demonstrates how meets the North Carolina Essential Standards for Biology, grades
Lecture 3: A basic statistical concept
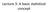 Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Model Worksheet Student Handout
 Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Chetek-Weyerhaeuser High School
 Chetek-Weyerhaeuser High School Unit 1 The Science of Biology (5 days) Biology I Units and s Biology I A s 1. I can design a scientific experiment that includes a control group, experimental group, constants,
Chetek-Weyerhaeuser High School Unit 1 The Science of Biology (5 days) Biology I Units and s Biology I A s 1. I can design a scientific experiment that includes a control group, experimental group, constants,
1. CHEMISTRY OF LIFE. Tutorial Outline
 Tutorial Outline North Carolina Tutorials are designed specifically for the Common Core State Standards for English language arts, the North Carolina Standard Course of Study for Math, and the North Carolina
Tutorial Outline North Carolina Tutorials are designed specifically for the Common Core State Standards for English language arts, the North Carolina Standard Course of Study for Math, and the North Carolina
Introduction to the Study of Life
 1 Introduction to the Study of Life Bio 103 Lecture GMU Dr. Largen 2 Outline Biology is the science of life The process of science Evolution, unity and diversity Core principles of biology 3 The Science
1 Introduction to the Study of Life Bio 103 Lecture GMU Dr. Largen 2 Outline Biology is the science of life The process of science Evolution, unity and diversity Core principles of biology 3 The Science
Administrative - Master Syllabus COVER SHEET
 Administrative - Master Syllabus COVER SHEET Purpose: It is the intention of this to provide a general description of the course, outline the required elements of the course and to lay the foundation for
Administrative - Master Syllabus COVER SHEET Purpose: It is the intention of this to provide a general description of the course, outline the required elements of the course and to lay the foundation for
Unit # - Title Intro to Biology Unit 1 - Scientific Method Unit 2 - Chemistry
 Intro to Biology Unit 1 - Scientific Method Unit 2 - Chemistry What is Biology? What is Science? What tools, skills, knowledge, and dispositions are needed to conduct scientific inquiry? How do the rules
Intro to Biology Unit 1 - Scientific Method Unit 2 - Chemistry What is Biology? What is Science? What tools, skills, knowledge, and dispositions are needed to conduct scientific inquiry? How do the rules
