rho Is Not Essential for Viability or Virulence in Staphylococcus aureus
|
|
|
- Domenic Dorsey
- 6 years ago
- Views:
Transcription
1 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 2001, p Vol. 45, No /01/$ DOI: /AAC Copyright 2001, American Society for Microbiology. All Rights Reserved. rho Is Not Essential for Viability or Virulence in Staphylococcus aureus ROBERT S. WASHBURN, ANDREA MARRA, ALEXANDER P. BRYANT, MARTIN ROSENBERG, AND DANIEL R. GENTRY* GlaxoSmithKline, Collegeville, Pennsylvania Received 15 November 2000/Returned for modification 29 December 2000/Accepted 11 January 2001 We have identified the gene for transcription termination factor Rho in Staphylococcus aureus. Deletion of rho in S. aureus reveals that it is not essential for viability or virulence. We also searched the available bacterial genomic sequences for homologs of Rho and found that it is broadly distributed and highly conserved. Exceptions include Streptococcus pneumoniae, Streptococcus pyogenes, Mycoplasma genitalium, Mycoplasma pneumoniae, Ureaplasma urealyticum, and Synechocystis sp. strain PCC6803, all of which appear not to possess a Rho homolog. Complementation studies indicate that S. aureus Rho possesses the same activity as Escherichia coli Rho and that the Rho inhibitor bicyclomycin is active against S. aureus Rho. Our results explain the lack of activity of bicyclomycin against many gram-positive bacteria and raise the possibility that the essentiality of rho may be the exception rather than the rule. The rho gene codes for transcription termination factor Rho (19) and is essential for the viability of Escherichia coli (5). The function of Rho is to catalyze the release of RNA from a transcription complex after a Rho-dependent terminator sequence has been transcribed. Rho has an RNA-dependent NTPase activity, which is required for transcript release (10, 12), and an RNA-DNA helicase activity (1). Several Rhodependent transcription terminators in Escherichia coli and coliphage are known (4, 20). Rho is a hexamer of identical 47-kDa monomers (2, 9, 11, 17). Its ability to bind RNA is thought to be conferred by residues 22 to 116 of the 419-amino-acid protein (14). The ATP binding domain is located between residues 167 and 319 (7). No activity has been assigned to the essential C terminus of Rho, although it has been speculated as being involved in subunit interactions (1, 6, 8). The antibiotic bicyclomycin inhibits Rho (23). With the exception of Micrococcus luteus (15, 16), gram-positive bacteria are resistant to bicyclomycin, and rho is not essential in the gram-positive bacterium Bacillus subtilis (13, 21). This study reports the sequence and characterization of the rho gene in the clinically important gram-positive pathogen Staphylococcus aureus. with E. coli rho. The plasmid pau2442 used in this study is from a genomic library used in the sequencing project. Deletion mutagenesis. A rho deletion was introduced into S. aureus RN4220. This was achieved by electroporating S. aureus RN4220 to tetracycline resistance (Tc r ) with prw101, which has the rho gene with 500 bp of flanking sequence interrupted by a tetracyline resistance marker that replaces the rho gene from codon 1 to codon 313 (Fig. 1). The resulting Tc r colonies were then screened for erythromycin resistance (Em r ), which is carried elsewhere on prw101. Tc r Em s colonies indicated a double-crossover event occurred. The deletion of rho was confirmed by PCR analysis with primers flanking the Tc r marker. To construct a rho strain in a pathogenic background, the rho::tet mutation was moved into the S. aureus clinical isolate WCUH29 by bacteriophage 11 transduction. The resulting strain, RSW101, was used in virulence testing as described below. Complemention of E. coli rho. E. coli rho was replaced with S. aureus rho by introducing the S. aureus rho-containing plasmid pau2442 into the E. coli rho deletion strain BLS107(pPMrho). This strain has a deletion-insertion (rho::kan) mutation in the chromosomal copy of rho. A wild-type copy of rho is carried on the plasmid ppmrho, which has a temperature-sensitive replicon. This strain fails to grow at 42 C due to the loss of rho at this temperature (22). pau2442 was introduced into BLS107(pPMrho) by electroporation and was plated at 30, 37, and 42 C. As a control, the parent strain was also plated at the different temperatures. While the parent strain failed to form isolated colonies at 37 and 42 C, the strain containing pau2442 grew well and formed isolated colonies at all temperatures. The loss of ppmrho was confirmed by scoring for loss of chloramphenicol resistance, which is encoded by this plasmid. To determine if the S. aureus Rho protein is sensitive to bicyclomycin, strain BLS107(pAU2442) was tested for sensitivity by MIC determination. Liquid MICs were determined by a serial dilution method. Strain BLS107 was found to have an MIC of bicyclomycin MATERIALS AND METHODS Bacteria and growth conditions. Staphylococcus aureus was propagated in either tryptic soy broth (DIFCO) or Luria broth (LB). E. coli was grown in LB. For auxotrophy measurements with S. aureus, M9 minimal medium was supplemented with arginine, cysteine, glutamate, glycine, isoleucine, leucine, methionine, proline, and valine, all at 20 g/ml. Nicotinate and thiamine were added at 0.2 g/ml. All cultures were routinely incubated at 37 C. All strains used in this study are described in Table 1. Identification of the S. aureus rho gene. The rho gene from Staphylococcus aureus was identified from an S. aureus genomic sequence database by homology * Corresponding author. Mailing address: GlaxoSmithKline, UP1345, 1250 S. Collegeville Rd., Collegeville, PA Phone: (610) Fax: (610) Dan_R_Gentry@sbphrd.com. Present address: Institute for Cancer Research, Columbia University, New York, NY Present address: Protein Design Laboratories, Inc., Fremont, CA Strain or plasmid TABLE 1. Strains and plasmids used in this study Description Source or reference Strains BLS107 E. coli rho::kan r 22 MG1655 Wild-type E. coli strain Laboratory collection RN4220 S. aureus laboratory strain Laboratory collection RSW100 RN4220 rho::tet r This study RSW101 WCUH29 rho::tet r This study WCUH29 S. aureus clinical isolate Laboratory collection Plasmids pau2442 ppmrho 4.3-kb genomic clone from WCUH29 containing rho Temperature-sensitive replicon, 3.0-kb genomic clone of E. coli containing rho This study
2 1100 WASHBURN ET AL. ANTIMICROB. AGENTS CHEMOTHER. FIG. 1. Sequence of S. aureus rho and flanking sequences used in this report. Shown is the DNA sequence of the region of the S. aureus chromosome that contains rho. The rho gene product is shown below its coding sequence. DNA sequence with a strikethrough line depicts the sequence deleted in the rho deletion-insertion strain described in this paper. of 125 g/ml. This is twofold lower than that of the wild-type E. coli strain MG1655 (250 g/ml). The MIC for the S. aureus strain RN4220 was found to be 2.5 mg/ml. Virulence testing. The rho deletion strain RSW101 and its wild-type parent, WCUH29, were tested for virulence with two mouse S. aureus infection models. The surgical wound model measures the ability to grow on sutures sewn in the skin of healthy mice, while the hematogenous pyelonephritis model measures the ability of the strains to cause kidney infections following parenteral infection with the bacteria. Surgical wound infections. The mice used for these infections were 4- to 6-week-old male CD-1 mice. Approximately 15-cm sutures were soaked in overnight cultures of WCUH29 and RSW101 for 30 min at room temperature. The backs of the mice were shaved, the animals were anesthetized with isoflurane, and an 2-cm-long incision was made along the animals backs. The infected sutures were secured at one end of the wound and tacked once midway along the wound on the underside of the skin. The suture was then secured at the other end of the wound, and the wound was closed with a single staple. After 5 days, the mice were sacrificed by CO 2 overdose. The skin surrounding the wound was excised and then homogenized in 1 ml of phosphate-buffered saline (PBS) in a stomacher, and bacterial viable counts were enumerated. Both infections yielded over 10 6 bacteria after 5 days, with no significant difference observed between the strains. For the hematogenous pyelonephritis model, bacterial recovery rates (mean standard deviation; n 5) were log 10 CFU/ml for strain WCUH29 and log 10 CFU/ml for strain WCUH29 rho. For the surgical wound model, the recovery rates were log 10 CFU/ml for strain WCUH29 and log 10 CFU/ml for strain WCUH29 rho. One
3 VOL. 45, 2001 S. AUREUS rho DELETION 1101 FIG. 2. Alignment of S. aureus Rho with B. subtilis and E. coli Rho. Sequences were aligned by using the MegAlign program of the DNAstar DNA analysis package. Identical residues are shaded. hundred colonies isolated from the rho mutant infection were checked for tetracycline resistance, indicative of the presence of the mutation, and all were found to be tetracycline resistant. Hematogenous pyelonephritis infections. Six- to eight-week-old male CD-1 mice were used for hematogenous pyelonephritis infections. Cultures of WCUH29 and RSW101 cells were adjusted to an A 600 of 0.6 or 0.3 per ml, and 200 l was injected into each mouse via a tail vein. Mice were sacrificed on day 5 postinfection by CO 2 overdose, and their kidneys were aseptically removed and then homogenized in 1 ml of PBS in a stomacher, and viable counts were enumerated (described above). The inoculum for both strains was 10 7 cells. The number of bacteria recovered from the kidneys infected with RSW101 after 5 days was not decreased. One hundred colonies from the RSW101 infection were checked for the tetracycline resistance marker, and 98% were found to be tetracycline resistant. Nucleotide sequence accession number. The nucleotide sequence of the S. aureus rho gene has been deposited in GenBank under accession no. AF RESULTS The S. aureus Rho protein. S. aureus Rho is predicted to code for a 443-amino-acid protein with a predicted molecular mass of 50 kda. S. aureus Rho is 53% identical and 67% similar to E. coli Rho and 66% identical and 75% similar to B. subtilis Rho (Fig. 2). The predicted ATPase domain (residues 167 to 342 of E. coli Rho) is highly conserved among different bacterial species (18). The predicted ATPase domain of S. aureus Rho (residues 180 to 356) is 80% similar and 70% identical to E. coli Rho and 86% similar and 81% identical to B. subtilis Rho. Finally, the predicted RNA binding domain of S. aureus Rho (residues 40 to 132) has 63% similarity and 41% identity to E. coli Rho and 77% similarity and 60% identity to B. subtilis Rho. Complementation. The S. aureus rho gene can complement the lack of growth of an E. coli strain with rho deleted.. This was determined by curing a rho-deleted strain of a plasmid that carries E. coli rho in the presence of S. aureus rho carried on another plasmid. The sole Rho activity in the resulting strain, BLS107(pAU2442), is from S. aureus Rho. Strain BLS107 (pau2442) is sensitive to bicyclomycin, as indicated by the bicyclomycin MIC of 125 g/ml, indicating that S. aureus Rho is inhibited by bicyclomycin. As further evidence of this, bicyclomycin was found to inhibit the ATPase activity of partially purified S. aureus Rho (data not shown). The S. aureus rho gene is not essential for in vitro growth. A rho strain, RSW100, was constructed by replacing nearly all of the rho gene with a tetracycline resistance cassette (Fig. 1). The resulting strain is viable and grows normally at 30, 37, 42,
4 1102 WASHBURN ET AL. ANTIMICROB. AGENTS CHEMOTHER. Aquificales Aquifex aeolicus TABLE 2. Bacterial phyla with members that possess a Rho homolog a Chlamydiales Chlamydia muridarum Chlamydia pneumoniae Chlamydia trachomatis Cytophaga-Flexibacter-Bacteroides Porphyromonas gingivalis Green sulfur Chlorobium tepidum Gram positive Low G C (not Streptococcus pyogenes, Mycoplasma pneumoniae, Mycoplasma genitalium, orstreptococcus pneumoniae) Bacillus subtilis Clostridium acetobutylicum Listeria monocytogenes High G C (Actinobacteria) Micrococcus luteus Mycobacterium leprae Mycobacterium tuberculosis Streptomyces coelicolor Streptomyces lividans Proteobacteria Rhodobacter sphaeroides Rickettsia prowazekii Neisseria animalis Neisseria cinerea Neisseria elongata Neisseria gonorrhoeae Neisseria lactamica Neisseria meningitidis Neisseria mucosa Neisseria pharyngis Neisseria polysaccharea Neisseria subflava ε Subdivision Campylobacter jejuni Helicobacter pylori Actinobacillus actinomycetemcomitans Actinobacillus pleuropneumoniae Allochromatium vinosum Buchnera aphidicola Escherichia coli Haemophilus influenzae Pseudomonas aeruginosa Pseudomonas fluorescens Salmonella enterica serovar Typhimurium Vibrio cholerae Xylella fastidiosa Yersinia pestis Spirochaetales Borrelia burgdorferi Treponema pallidum Thermotogales Thermotoga maritima Thermus/Deinococcus Deinococcus radiodurans a Groupings taken from and determined by BLAST analysis with S. aureus Rho. and 46 C. RSW100 also grows normally on rich medium (LB or tryptic soy broth) or minimal medium (supplemented M9 medium) and under anaerobic conditions (data not shown). rho is not required for S. aureus virulence. The rho deletion was transduced into the pathogenic clinical isolate WCUH29, yielding strain RSW101. Cultures of WCUH29 and RSW101 were used in the two in vivo models described in Materials and Methods. S. aureus strains mutated in a variety of different genes affecting cell metabolism or virulence have been examined in our laboratories. These strains have displayed reductions of up to 3 logs in the pyelonephritis model and 1 log in the surgical wound infection. In the experiments described here, the in vivo growth of RSW101 was not impaired compared to that of wild-type WCUH29 in either of the model systems. From these data, we conclude that, as with in vitro growth, rho is not essential for the virulence of the strain tested. Review of Rho homologs present in public sequence database. By using the S. aureus Rho sequence as a query, we searched the public databases and some proprietary sequence databases for homologs. Rho was found to be highly conserved; BLAST scores were or lower for all fulllength Rho sequences annotated as such. As indicated previously by Opperman and Richardson (18), Rho is broadly distributed among all the major phyla of the eubacteria (Table 2). A major exception to the universal presence of Rho among the eubacteria are the members of the low-gc, gram-positive bacteria Streptococus pyogenes, Streptococcus pneumoniae, Mycoplasma genitalium, M. pneumoniae, and Ureaplasma urealyticum, all of which do not possess a significant homolog. In the case of S. pneumoniae, we searched several proprietary databases in addition to publicly available databases and failed to identify a homolog. In addition to the mentioned bacteria, Synechocystis sp. strain PCC6803, a cyanobacterium, also apparently lacks a Rho homolog. As indicated previously (18), the ATP binding motif of Rho shares homology with the and subunits of F1 ATPases. This homology is conserved among the bacteria as well. DISCUSSION In this report, we show that the S. aureus rho gene can complement an E. coli rho deletion strain. The ability of S. aureus rho to replace the essential E. coli rho gene raises the possibility that it performs the same functions in S. aureus and E. coli. rho is not essential in the low-gc, gram-positive bacterium Bacillus subtilis (13, 21), and several gram-positive species, including Staphylococcus aureus, are resistant to the Rho inhibitor bicyclomycin (15). Here we show that rho is not required for viability or virulence in S. aureus. The E. coli strain constructed here, which has had its own rho gene replaced with S. aureus rho, remains sensitive to bicyclomycin. This suggests that the resistance of gram-positive bacteria to bicyclomycin is a result of the nonessentiality of Rho. Supporting this conclusion and extending the lack of essentiality of rho to other genera, we found that the completed genomes of several members of the low-gc, gram-positive bacteria lack a Rho homolog. The cyanobacterium Synechocystis sp. strain PCC6803 was also found to lack a Rho homolog. An open question is why rho should be essential in E. coli
5 VOL. 45, 2001 S. AUREUS rho DELETION 1103 and other gram-negative bacteria, but not be essential in some gram-positive bacteria. It could be that Rho activity is redundant in the bacteria where it is not essential. This possibility cannot be readily discounted, but it is clear that the enzyme responsible for this redundant activity would have to have no sequence homology with Rho for this to be the case. A second possibility, that Rho is essential for organisms with a higher GC content, seems unlikely, given that Synechocystis sp. strain PCC6803 also lacks Rho, and yet its genome is 47% GC, close to that of E. coli. The lack of rho among several members of the low-gc, gram-positive bacteria and a member of the very distantly related cyanobacteria could indicate a trend that the essentiality of rho may be more the exception than the rule. Because of this, a more informative approach to the question is to ask why rho is essential in E. coli and other gram-negative bacteria. There are only a few Rho-dependent terminators in E. coli; the vast majority of E. coli operons contain factorindependent terminators. Furthermore, it is not obvious why inhibition of termination at Rho-dependent terminators would lead to a lethal event. It might be that there is some other function of Rho that better explains its essential nature in some bacteria. REFERENCES 1. Bear, D. G., C. L. Andrews, J. D. Singer, W. D. Morgan, R. A. Grant, P. H. von Hippel, and T. Platt Escherichia coli transcription termination factor Rho has a two-domain structure in its activated form. Proc. Natl. Acad. Sci. USA 82: Bear, D. G., P. S. Hicks, K. W. Escudero, C. L. Andrews, J. A. McSwiggen, and P. H. von Hippel Interactions of Escherichia coli transcription termination factor Rho with RNA. J. Mol. Biol. 199: Brennan, C. A., A. J. Dombroski, and T. Platt Transcription termination factor Rho is an RNA-DNA helicase. Cell 48: Das, A Control of transcription termination by RNA binding proteins. Annu. Rev. Biochem. 62: Das, A., D. Court, and S. Adhya Isolation and characterization of conditional lethal mutants of Escherichia coli defective in transcription termination factor rho. Proc. Natl. Acad. Sci. USA 73: Dolan, J. W., N. F. Marshall, and, J. P. Richardson Transcription termination factor Rho has three distinct structural domains. J. Biol. Chem. 265: Dombroski, A. J., J. R. LaDine, R. L. Cross, and T. Platt The ATP binding site on rho protein. Affinity labelling of Lys 181 by pyridoxal 5 diphospho-5 -adenosine. J. Biol. Chem. 263: Dombroski, A. J., and T. Platt Structure of factor: an RNA-binding domain and a separate region with strong similarity to ATP-binding domains. Proc. Natl. Acad. Sci. USA 85: Finger, L. R., and J. P. Richardson Stabilization of the hexameric form of Escherichia coli protein rho under ATP hydrolysis conditions. J. Mol. Biol. 156: Galluppi, G. R., C. Lowery, and J. P. Richardson Nucleoside triphosphate requirement for termination of RNA synthesis by rho factor, p In R. Losick and M. Chamberlin (ed.), RNA polymerase. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. 11. Gogol, E. P., S. E. Seifred, and P. H. von Hippel Structure and assembly of the Escherichia coli transcription termination factor Rho and its interactions with RNA. I. Cryoelectron microscopic studies. J. Mol. Biol. 221: Howard, B. H., and B. de Crombrugghe ATPase activity required for termination of transcription by the Escherichia coli protein factor. J. Biol. Chem. 251: Ingham, C. J., J. Dennis, and P. A. Furneaux Autogenous regulation of transcription termination factor Rho and the requirement for Nus factors in Bacillus subtilis. Mol. Microbiol. 31: Modrak, D., and J. P. Richardson The RNA-binding domain of transcription termination factor Rho: isolation, characterization and determination of sequence limits. Biochemistry 33: Nishida, M., Y. Mine, and T. Matsubara Bicyclomycin, a new antibiotic. III. In vitro and in vivo antimicrobial activity. J. Antibiot. 25: Nowatzke, W. L., E. Keller, G. Koch, and J. P. Richardson Transcription termination factor Rho is essential for Micrococcus luteus. J. Bacteriol. 179: Oda, T., and M. Takanami Observations on the structure of the termination factor Rho and its attachment to DNA. J. Mol. Biol. 71: Opperman, T., and J. P. Richardson Phylogenetic analysis of sequences from diverse bacteria with homology to the Escherichia coli rho gene. J. Bacteriol. 176: Pinkham, J. L., and T. Platt The nucleotide sequence of the rho gene of E. coli K-12. Nucleic Acids Res. 11: Platt, T., and J. P. Richardson Escherichia coli Rho factor: protein and enzyme of transcription termination, p In S. L. McKnight and K. R. Yamamoto (ed.), Transcriptional regulation. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. 21. Quirk, P. G., E. A. Dunkley, Jr., P. Lee, and T. A. Krulwich Identification of a putative Bacillus subtilis rho gene. J. Bacteriol. 175: Washburn, R. S., and B. L. Stitt In vitro characterization of transcription termination factor Rho from Escherichia coli rho(nusd) mutants. J. Mol. Biol. 260: Zwiefka, A., H. Kohn, and W. R. Widger Transcription termination factor rho: The site of bicyclomycin inhibition in Escherichia coli. Biochemistry 32:
Genetic Basis of Variation in Bacteria
 Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
The Minimal-Gene-Set -Kapil PHY498BIO, HW 3
 The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
Fitness constraints on horizontal gene transfer
 Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Tetracycline Rationale for the EUCAST clinical breakpoints, version th November 2009
 Tetracycline Rationale for the EUCAST clinical breakpoints, version 1.0 20 th November 2009 Introduction The natural tetracyclines, including tetracycline, chlortetracycline, oxytetracycline and demethylchlortetracycline
Tetracycline Rationale for the EUCAST clinical breakpoints, version 1.0 20 th November 2009 Introduction The natural tetracyclines, including tetracycline, chlortetracycline, oxytetracycline and demethylchlortetracycline
2 Genome evolution: gene fusion versus gene fission
 2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana
 Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana Table S1 The species marked with belong to the Proteobacteria subset and those marked
Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana Table S1 The species marked with belong to the Proteobacteria subset and those marked
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Evolutionary Analysis by Whole-Genome Comparisons
 JOURNAL OF BACTERIOLOGY, Apr. 2002, p. 2260 2272 Vol. 184, No. 8 0021-9193/02/$04.00 0 DOI: 184.8.2260 2272.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Evolutionary Analysis
JOURNAL OF BACTERIOLOGY, Apr. 2002, p. 2260 2272 Vol. 184, No. 8 0021-9193/02/$04.00 0 DOI: 184.8.2260 2272.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Evolutionary Analysis
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
Microbial Taxonomy. Classification of living organisms into groups. A group or level of classification
 Lec 2 Oral Microbiology Dr. Chatin Purpose Microbial Taxonomy Classification Systems provide an easy way grouping of diverse and huge numbers of microbes To provide an overview of how physicians think
Lec 2 Oral Microbiology Dr. Chatin Purpose Microbial Taxonomy Classification Systems provide an easy way grouping of diverse and huge numbers of microbes To provide an overview of how physicians think
Figure Page 117 Microbiology: An Introduction, 10e (Tortora/ Funke/ Case)
 Chapter 11 The Prokaryotes: Domains Bacteria and Archaea Objective Questions 1) Which of the following are found primarily in the intestines of humans? A) Gram-negative aerobic rods and cocci B) Aerobic,
Chapter 11 The Prokaryotes: Domains Bacteria and Archaea Objective Questions 1) Which of the following are found primarily in the intestines of humans? A) Gram-negative aerobic rods and cocci B) Aerobic,
MICROBIAL BIOCHEMISTRY BIOT 309. Dr. Leslye Johnson Sept. 30, 2012
 MICROBIAL BIOCHEMISTRY BIOT 309 Dr. Leslye Johnson Sept. 30, 2012 Phylogeny study of evoluhonary relatedness among groups of organisms (e.g. species, populahons), which is discovered through molecular
MICROBIAL BIOCHEMISTRY BIOT 309 Dr. Leslye Johnson Sept. 30, 2012 Phylogeny study of evoluhonary relatedness among groups of organisms (e.g. species, populahons), which is discovered through molecular
ABSTRACT. As a result of recent successes in genome scale studies, especially genome
 ABSTRACT Title of Dissertation / Thesis: COMPUTATIONAL ANALYSES OF MICROBIAL GENOMES OPERONS, PROTEIN FAMILIES AND LATERAL GENE TRANSFER. Yongpan Yan, Doctor of Philosophy, 2005 Dissertation / Thesis Directed
ABSTRACT Title of Dissertation / Thesis: COMPUTATIONAL ANALYSES OF MICROBIAL GENOMES OPERONS, PROTEIN FAMILIES AND LATERAL GENE TRANSFER. Yongpan Yan, Doctor of Philosophy, 2005 Dissertation / Thesis Directed
Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria)
 Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria) (don t study Concept 27.2) Some phyla Remember: Bacterial cell structure and shapes 1 Usually very small but some are unusually
Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria) (don t study Concept 27.2) Some phyla Remember: Bacterial cell structure and shapes 1 Usually very small but some are unusually
INTRODUCTION MATERIALS & METHODS
 Evaluation of Three Bacterial Transport Systems, The New Copan M40 Transystem, Remel Bactiswab And Medical Wire & Equipment Transwab, for Maintenance of Aerobic Fastidious and Non-Fastidious Organisms
Evaluation of Three Bacterial Transport Systems, The New Copan M40 Transystem, Remel Bactiswab And Medical Wire & Equipment Transwab, for Maintenance of Aerobic Fastidious and Non-Fastidious Organisms
ANTIMICROBIAL TESTING. E-Coli K-12 - E-Coli 0157:H7. Salmonella Enterica Servoar Typhimurium LT2 Enterococcus Faecalis
 ANTIMICROBIAL TESTING E-Coli K-12 - E-Coli 0157:H7 Salmonella Enterica Servoar Typhimurium LT2 Enterococcus Faecalis Staphylococcus Aureus (Staph Infection MRSA) Streptococcus Pyrogenes Anti Bacteria effect
ANTIMICROBIAL TESTING E-Coli K-12 - E-Coli 0157:H7 Salmonella Enterica Servoar Typhimurium LT2 Enterococcus Faecalis Staphylococcus Aureus (Staph Infection MRSA) Streptococcus Pyrogenes Anti Bacteria effect
The Prokaryotes: Domains Bacteria and Archaea
 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 11 The Prokaryotes: Domains Bacteria and Archaea Table 11.1 Classification of Selected Prokaryotes*
PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 11 The Prokaryotes: Domains Bacteria and Archaea Table 11.1 Classification of Selected Prokaryotes*
Correlations between Shine-Dalgarno Sequences and Gene Features Such as Predicted Expression Levels and Operon Structures
 JOURNAL OF BACTERIOLOGY, Oct. 2002, p. 5733 5745 Vol. 184, No. 20 0021-9193/02/$04.00 0 DOI: 10.1128/JB.184.20.5733 5745.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Correlations
JOURNAL OF BACTERIOLOGY, Oct. 2002, p. 5733 5745 Vol. 184, No. 20 0021-9193/02/$04.00 0 DOI: 10.1128/JB.184.20.5733 5745.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Correlations
Multifractal characterisation of complete genomes
 arxiv:physics/1854v1 [physics.bio-ph] 28 Aug 21 Multifractal characterisation of complete genomes Vo Anh 1, Ka-Sing Lau 2 and Zu-Guo Yu 1,3 1 Centre in Statistical Science and Industrial Mathematics, Queensland
arxiv:physics/1854v1 [physics.bio-ph] 28 Aug 21 Multifractal characterisation of complete genomes Vo Anh 1, Ka-Sing Lau 2 and Zu-Guo Yu 1,3 1 Centre in Statistical Science and Industrial Mathematics, Queensland
Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: A new family of genes responsible for autoinducer production
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 1639 1644, February 1999 Microbiology Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: A new family of genes responsible for autoinducer
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 1639 1644, February 1999 Microbiology Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: A new family of genes responsible for autoinducer
Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1
 Name I. Multiple Choice (1 point each) Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1 B 1. Which is possessed by eukaryotes but not by prokaryotes? A. Cell wall B. Distinct nucleus
Name I. Multiple Choice (1 point each) Introduction to Microbiology BIOL 220 Summer Session I, 1996 Exam # 1 B 1. Which is possessed by eukaryotes but not by prokaryotes? A. Cell wall B. Distinct nucleus
Overview of the major bacterial pathogens The major bacterial pathogens are presented in this table:
 Practical Microbiology 30/11/2018 University of Sulaimani college of Pharmacy Year2 Lab. 5: Overview of the major bacterial pathogens The major bacterial pathogens are presented in this table: Major Bacterial
Practical Microbiology 30/11/2018 University of Sulaimani college of Pharmacy Year2 Lab. 5: Overview of the major bacterial pathogens The major bacterial pathogens are presented in this table: Major Bacterial
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
The use of gene clusters to infer functional coupling
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2896 2901, March 1999 Genetics The use of gene clusters to infer functional coupling ROSS OVERBEEK*, MICHAEL FONSTEIN, MARK D SOUZA*, GORDON D. PUSCH*, AND NATALIA
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2896 2901, March 1999 Genetics The use of gene clusters to infer functional coupling ROSS OVERBEEK*, MICHAEL FONSTEIN, MARK D SOUZA*, GORDON D. PUSCH*, AND NATALIA
BACTERIAL PHYSIOLOGY SMALL GROUP. Monday, August 25, :00pm. Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D.
 BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
Molecular Systematic Studies of Eubacteria, Using 70 -Type Sigma Factors of Group 1 and Group 2
 JOURNAL OF BACTERIOLOGY, Mar. 1997, p. 1734 1747 Vol. 179, No. 5 0021-9193/97/$04.00 0 Copyright 1997, American Society for Microbiology Molecular Systematic Studies of Eubacteria, Using 70 -Type Sigma
JOURNAL OF BACTERIOLOGY, Mar. 1997, p. 1734 1747 Vol. 179, No. 5 0021-9193/97/$04.00 0 Copyright 1997, American Society for Microbiology Molecular Systematic Studies of Eubacteria, Using 70 -Type Sigma
The Rhodobacter sphaeroides rho Gene: Expression and Genetic Analysis of Structure and Function
 JOURNAL OF BACTERIOLOGY, Apr. 1996, p. 1946 1954 Vol. 178, No. 7 0021-9193/96/$04.00 0 Copyright 1996, American Society for Microbiology The Rhodobacter sphaeroides 2.4.1 rho Gene: Expression and Genetic
JOURNAL OF BACTERIOLOGY, Apr. 1996, p. 1946 1954 Vol. 178, No. 7 0021-9193/96/$04.00 0 Copyright 1996, American Society for Microbiology The Rhodobacter sphaeroides 2.4.1 rho Gene: Expression and Genetic
Supporting online material
 Supporting online material Materials and Methods Target proteins All predicted ORFs in the E. coli genome (1) were downloaded from the Colibri data base (2) (http://genolist.pasteur.fr/colibri/). 737 proteins
Supporting online material Materials and Methods Target proteins All predicted ORFs in the E. coli genome (1) were downloaded from the Colibri data base (2) (http://genolist.pasteur.fr/colibri/). 737 proteins
INTERPRETATION OF THE GRAM STAIN
 INTERPRETATION OF THE GRAM STAIN DISCLOSURE Relevant relationships with commercial entities none Potential for conflicts of interest within this presentation none Steps taken to review and mitigate potential
INTERPRETATION OF THE GRAM STAIN DISCLOSURE Relevant relationships with commercial entities none Potential for conflicts of interest within this presentation none Steps taken to review and mitigate potential
Topology. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 Topology 1 Introduction 2 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 G+C content 7 Codon usage 27 marc.bailly-bechet@univ-lyon1.fr The big picture Eukaryota Bacteria Many linear chromosomes
Topology 1 Introduction 2 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 G+C content 7 Codon usage 27 marc.bailly-bechet@univ-lyon1.fr The big picture Eukaryota Bacteria Many linear chromosomes
Vital Statistics Derived from Complete Genome Sequencing (for E. coli MG1655)
 We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
Bacterial Genetics & Operons
 Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Database and Comparative Identification of Prophages
 Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy
Database and Comparative Identification of Prophages K.V. Srividhya 1, Geeta V Rao 1, Raghavenderan L 1, Preeti Mehta 1, Jaime Prilusky 2, Sankarnarayanan Manicka 1, Joel L. Sussman 3, and S Krishnaswamy
# shared OGs (spa, spb) Size of the smallest genome. dist (spa, spb) = 1. Neighbor joining. OG1 OG2 OG3 OG4 sp sp sp
 Bioinformatics and Evolutionary Genomics: Genome Evolution in terms of Gene Content 3/10/2014 1 Gene Content Evolution What about HGT / genome sizes? Genome trees based on gene content: shared genes Haemophilus
Bioinformatics and Evolutionary Genomics: Genome Evolution in terms of Gene Content 3/10/2014 1 Gene Content Evolution What about HGT / genome sizes? Genome trees based on gene content: shared genes Haemophilus
Taking shape: control of bacterial cell wall biosynthesis
 Blackwell Science, LtdOxford, UKMMIMolecular Microbiology0950-382XBlackwell Publishing Ltd, 2005? 200557511771181CommentaryBacterial morphogenesg. C. Stewart Molecular Microbiology (2005) 57(5), 1177 1181
Blackwell Science, LtdOxford, UKMMIMolecular Microbiology0950-382XBlackwell Publishing Ltd, 2005? 200557511771181CommentaryBacterial morphogenesg. C. Stewart Molecular Microbiology (2005) 57(5), 1177 1181
Measure representation and multifractal analysis of complete genomes
 PHYSICAL REVIEW E, VOLUME 64, 031903 Measure representation and multifractal analysis of complete genomes Zu-Guo Yu, 1,2, * Vo Anh, 1 and Ka-Sing Lau 3 1 Centre in Statistical Science and Industrial Mathematics,
PHYSICAL REVIEW E, VOLUME 64, 031903 Measure representation and multifractal analysis of complete genomes Zu-Guo Yu, 1,2, * Vo Anh, 1 and Ka-Sing Lau 3 1 Centre in Statistical Science and Industrial Mathematics,
1. Prokaryotic Nutritional & Metabolic Adaptations
 Chapter 27B: Bacteria and Archaea 1. Prokaryotic Nutritional & Metabolic Adaptations 2. Survey of Prokaryotic Groups A. Domain Bacteria Gram-negative groups B. Domain Bacteria Gram-positive groups C. Domain
Chapter 27B: Bacteria and Archaea 1. Prokaryotic Nutritional & Metabolic Adaptations 2. Survey of Prokaryotic Groups A. Domain Bacteria Gram-negative groups B. Domain Bacteria Gram-positive groups C. Domain
Chapter 21 PROKARYOTES AND VIRUSES
 Chapter 21 PROKARYOTES AND VIRUSES Bozeman Video classification of life http://www.youtube.com/watch?v=tyl_8gv 7RiE Impacts, Issues: West Nile Virus Takes Off Alexander the Great, 336 B.C., conquered a
Chapter 21 PROKARYOTES AND VIRUSES Bozeman Video classification of life http://www.youtube.com/watch?v=tyl_8gv 7RiE Impacts, Issues: West Nile Virus Takes Off Alexander the Great, 336 B.C., conquered a
EZ-COMP EZ-COMP For Training and Proficiency Testing Product Details
 EZ-COMP For Training and Proficiency Testing Mixed microorganism populations Identified by codes rather than descriptions Refrigerated storage Traceable to reference culture Product warranty Product Details
EZ-COMP For Training and Proficiency Testing Mixed microorganism populations Identified by codes rather than descriptions Refrigerated storage Traceable to reference culture Product warranty Product Details
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
Essentiality in B. subtilis
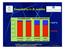 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Heterologous expression and secretion of Clostridium perfringens [beta]-toxoid by Bacillus subtilis Nijland, Reindert
![Heterologous expression and secretion of Clostridium perfringens [beta]-toxoid by Bacillus subtilis Nijland, Reindert Heterologous expression and secretion of Clostridium perfringens [beta]-toxoid by Bacillus subtilis Nijland, Reindert](/thumbs/92/108860012.jpg) University of Groningen Heterologous expression and secretion of Clostridium perfringens [beta]-toxoid by Bacillus subtilis Nijland, Reindert IMPORTANT NOTE: You are advised to consult the publisher's
University of Groningen Heterologous expression and secretion of Clostridium perfringens [beta]-toxoid by Bacillus subtilis Nijland, Reindert IMPORTANT NOTE: You are advised to consult the publisher's
The impact of the neisserial DNA uptake sequence on genome evolution and stability
 The impact of the neisserial DNA uptake sequence on genome evolution and stability Ole Herman Ambur The Tønjum Group Transformation Neisserial transformation requires: DNA uptake sequence (DUS) Transformation
The impact of the neisserial DNA uptake sequence on genome evolution and stability Ole Herman Ambur The Tønjum Group Transformation Neisserial transformation requires: DNA uptake sequence (DUS) Transformation
Stability. Received for publication 1 August to be fl-lactamase-producing strains.
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 1978, p. 584-588 0066-4804/78/0013-0584$02.00/0 Copyright X) 1978 American Society for Microbiology Vol. 13, No. 4 Printed in U.S.A. Cefaclor: In Vitro Spectrum
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 1978, p. 584-588 0066-4804/78/0013-0584$02.00/0 Copyright X) 1978 American Society for Microbiology Vol. 13, No. 4 Printed in U.S.A. Cefaclor: In Vitro Spectrum
The Effect of Static Magnetic Field on E. coli, S. aureus and B. subtilis Viability
 The Effect of Static Magnetic Field on E. coli, S. aureus and B. subtilis Viability Khaled A. Al-Khaza'leh 1* Abdullah T. Al-fawwaz 2 1. Department of Physics, Al-albayt University, PO box 130040, Mafraq,
The Effect of Static Magnetic Field on E. coli, S. aureus and B. subtilis Viability Khaled A. Al-Khaza'leh 1* Abdullah T. Al-fawwaz 2 1. Department of Physics, Al-albayt University, PO box 130040, Mafraq,
chapter 5 the mammalian cell entry 1 (mce1) operon of Mycobacterium Ieprae and Mycobacterium tuberculosis
 chapter 5 the mammalian cell entry 1 (mce1) operon of Mycobacterium Ieprae and Mycobacterium tuberculosis chapter 5 Harald G. Wiker, Eric Spierings, Marc A. B. Kolkman, Tom H. M. Ottenhoff, and Morten
chapter 5 the mammalian cell entry 1 (mce1) operon of Mycobacterium Ieprae and Mycobacterium tuberculosis chapter 5 Harald G. Wiker, Eric Spierings, Marc A. B. Kolkman, Tom H. M. Ottenhoff, and Morten
Domain Bacteria. BIO 220 Microbiology Jackson Community College
 Domain Bacteria BIO 220 Microbiology Jackson Community College John Ireland, Ph.D. 2006 Scientific Nomenclature Domain - Bacteria Phylum Important for gross characteristics Class Intermediate characteristics
Domain Bacteria BIO 220 Microbiology Jackson Community College John Ireland, Ph.D. 2006 Scientific Nomenclature Domain - Bacteria Phylum Important for gross characteristics Class Intermediate characteristics
CLASSIFICATION OF BACTERIA
 CLASSIFICATION OF BACTERIA DISCLOSURE Relevant relationships with commercial entities none Potential for conflicts of interest within this presentation none Steps taken to review and mitigate potential
CLASSIFICATION OF BACTERIA DISCLOSURE Relevant relationships with commercial entities none Potential for conflicts of interest within this presentation none Steps taken to review and mitigate potential
Bacteria Outline. 1. Overview. 2. Structural & Functional Features. 3. Taxonomy. 4. Communities
 Bacteria Outline 1. Overview 2. Structural & Functional Features 3. Taxonomy 4. Communities Bacteria - Taxonomy PHYLUM CLASS ORDER FAMILY GENUS SPECIES SUB-SPECIES & STRAINS Bacteria - Phyla Firmicutes
Bacteria Outline 1. Overview 2. Structural & Functional Features 3. Taxonomy 4. Communities Bacteria - Taxonomy PHYLUM CLASS ORDER FAMILY GENUS SPECIES SUB-SPECIES & STRAINS Bacteria - Phyla Firmicutes
Microbial Genetics, Mutation and Repair. 2. State the function of Rec A proteins in homologous genetic recombination.
 Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Answer the following questions 1. Define genetic recombination. Microbial Genetics, Mutation and Repair 2. State the function of Rec A proteins in homologous genetic recombination. 3. List 3 types of bacterial
Nitroxoline Rationale for the NAK clinical breakpoints, version th October 2013
 Nitroxoline Rationale for the NAK clinical breakpoints, version 1.0 4 th October 2013 Foreword NAK The German Antimicrobial Susceptibility Testing Committee (NAK - Nationales Antibiotika-Sensitivitätstest
Nitroxoline Rationale for the NAK clinical breakpoints, version 1.0 4 th October 2013 Foreword NAK The German Antimicrobial Susceptibility Testing Committee (NAK - Nationales Antibiotika-Sensitivitätstest
Objects of the Medical Microbiology revision a) Pathogenic microbes (causing diseases of human beings or animals) b) Normal microflora (microbes commo
 Institute for Microbiology, Medical Faculty of Masaryk University and St. Anna Faculty Hospital in Brno Miroslav Votava MORPHOLOGY AND STRUCTURE OF BACTERIAL CELL The 2nd lecture for 2nd-year students
Institute for Microbiology, Medical Faculty of Masaryk University and St. Anna Faculty Hospital in Brno Miroslav Votava MORPHOLOGY AND STRUCTURE OF BACTERIAL CELL The 2nd lecture for 2nd-year students
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
Mouth animalcules (bacteria)
 Mouth animalcules (bacteria) 1684 http://en.citizendium.org/images/thumb/9/94/leeuwenhoek.jpg/300px-leeuwenhoek.jpg Prokaryotic Cell Shapes Coccus - cocci Bacillus - bacillus Spirillum - spirilli Vibrio
Mouth animalcules (bacteria) 1684 http://en.citizendium.org/images/thumb/9/94/leeuwenhoek.jpg/300px-leeuwenhoek.jpg Prokaryotic Cell Shapes Coccus - cocci Bacillus - bacillus Spirillum - spirilli Vibrio
By signing below, you acknowledge that you have ensured that you are complying with the above statement.
 Instructions: This exam consists of 31 multiple choice questions on 8 pages, including this one. Please submit your answers on the scantron sheet provided and on this copy of the exam. This exam is closed
Instructions: This exam consists of 31 multiple choice questions on 8 pages, including this one. Please submit your answers on the scantron sheet provided and on this copy of the exam. This exam is closed
UNIVERSITY OF YORK. BA, BSc, and MSc Degree Examinations Department : BIOLOGY. Title of Exam: Molecular microbiology
 Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
UKPMC Funders Group Author Manuscript J Microbiol. Author manuscript; available in PMC 2007 March 28.
 UKPMC Funders Group Author Manuscript Published in final edited form as: J Microbiol. 2006 February ; 44(1): 11 22. Rho-dependent Transcription Termination: More Questions than Answers Sharmistha Banerjee,
UKPMC Funders Group Author Manuscript Published in final edited form as: J Microbiol. 2006 February ; 44(1): 11 22. Rho-dependent Transcription Termination: More Questions than Answers Sharmistha Banerjee,
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Tiffany Samaroo MB&B 452a December 8, Take Home Final. Topic 1
 Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Prokaryotic phylogenies inferred from protein structural domains
 Letter Prokaryotic phylogenies inferred from protein structural domains Eric J. Deeds, 1 Hooman Hennessey, 2 and Eugene I. Shakhnovich 3,4 1 Department of Molecular and Cellular Biology, Harvard University,
Letter Prokaryotic phylogenies inferred from protein structural domains Eric J. Deeds, 1 Hooman Hennessey, 2 and Eugene I. Shakhnovich 3,4 1 Department of Molecular and Cellular Biology, Harvard University,
Bio Microbiology - Spring 2014 Learning Guide 04.
 Bio 230 - Microbiology - Spring 2014 Learning Guide 04 http://pessimistcomic.blogspot.com/ Cell division is a part of a replication cycle that takes place throughout the life of the bacterium A septum
Bio 230 - Microbiology - Spring 2014 Learning Guide 04 http://pessimistcomic.blogspot.com/ Cell division is a part of a replication cycle that takes place throughout the life of the bacterium A septum
(Lys), resulting in translation of a polypeptide without the Lys amino acid. resulting in translation of a polypeptide without the Lys amino acid.
 1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
Gene Family Content-Based Phylogeny of Prokaryotes: The Effect of Criteria for Inferring Homology
 Syst. Biol. 54(2):268 276, 2005 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150590923335 Gene Family Content-Based Phylogeny of Prokaryotes:
Syst. Biol. 54(2):268 276, 2005 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150590923335 Gene Family Content-Based Phylogeny of Prokaryotes:
Obligate anaerobes - cannot grow in the presence of oxygen Facultative anaerobes - can grow with or without oxygen Aerobic - require oxygen
 PROKARYOTES *include bacteria and archaea *singular: bacterium / plural: bacteria PROPERTIES 1. Bacteria are classified into two kingdoms: Eubacteria (true bacteria) and Archaebacteria (Ancient Bacteria).
PROKARYOTES *include bacteria and archaea *singular: bacterium / plural: bacteria PROPERTIES 1. Bacteria are classified into two kingdoms: Eubacteria (true bacteria) and Archaebacteria (Ancient Bacteria).
Biology 112 Practice Midterm Questions
 Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Microbial Typing by Machine Learned DNA Melt Signatures
 Microbial Typing by Machine Learned DNA Melt Signatures Nadya Andini 1, Bo Wang 2, Pornpat Athamanolap 3, Justin Hardick 4, Billie J. Masek 5, Simone Thair 1, Annie Hu 1, Gideon Avornu 5, Stephen Peterson
Microbial Typing by Machine Learned DNA Melt Signatures Nadya Andini 1, Bo Wang 2, Pornpat Athamanolap 3, Justin Hardick 4, Billie J. Masek 5, Simone Thair 1, Annie Hu 1, Gideon Avornu 5, Stephen Peterson
Genetics 304 Lecture 6
 Genetics 304 Lecture 6 00/01/27 Assigned Readings Busby, S. and R.H. Ebright (1994). Promoter structure, promoter recognition, and transcription activation in prokaryotes. Cell 79:743-746. Reed, W.L. and
Genetics 304 Lecture 6 00/01/27 Assigned Readings Busby, S. and R.H. Ebright (1994). Promoter structure, promoter recognition, and transcription activation in prokaryotes. Cell 79:743-746. Reed, W.L. and
1. Which of the following species have strains that are capable of undergoing the process of conjugation?
 Biology 3340 Summer 2005 Second Examination Version A Name Be sure to put your name on the mark-sense sheet as well Directions: Write your name in the correct space on the mark-sense sheet and the exam
Biology 3340 Summer 2005 Second Examination Version A Name Be sure to put your name on the mark-sense sheet as well Directions: Write your name in the correct space on the mark-sense sheet and the exam
Gentamicin Rationale for the EUCAST clinical breakpoints, version th February, 2009
 Gentamicin Rationale for the EUCAST clinical breakpoints, version 1.2 16 th February, 2009 Introduction The aminoglycosides are a group of naturally occurring or semi-synthetic compounds with bactericidal
Gentamicin Rationale for the EUCAST clinical breakpoints, version 1.2 16 th February, 2009 Introduction The aminoglycosides are a group of naturally occurring or semi-synthetic compounds with bactericidal
Studies with RP1 deletion plasmids: Incompatibility properties, plasmid curing and the spontaneous loss of resistance
 J. Biosci., Vol. 1, Number 3, September 1979, pp. 345 354. Printed in India. Studies with RP1 deletion plasmids: Incompatibility properties, plasmid curing and the spontaneous loss of resistance HABIBUL
J. Biosci., Vol. 1, Number 3, September 1979, pp. 345 354. Printed in India. Studies with RP1 deletion plasmids: Incompatibility properties, plasmid curing and the spontaneous loss of resistance HABIBUL
Mini-Tn7 Derivative Construction and Characterization. Mini-Tn7 derivatives for
 Supplemental Methods Mini-Tn7 Derivative Construction and Characterization. Mini-Tn7 derivatives for constitutive expression of fluorescent proteins in S. oneidensis were constructed as follows. The EcoRI-XbaI
Supplemental Methods Mini-Tn7 Derivative Construction and Characterization. Mini-Tn7 derivatives for constitutive expression of fluorescent proteins in S. oneidensis were constructed as follows. The EcoRI-XbaI
Chapter 20. Initiation of transcription. Eukaryotic transcription initiation
 Chapter 20. Initiation of transcription Eukaryotic transcription initiation 2003. 5.22 Prokaryotic vs eukaryotic Bacteria = one RNA polymerase Eukaryotes have three RNA polymerases (I, II, and III) in
Chapter 20. Initiation of transcription Eukaryotic transcription initiation 2003. 5.22 Prokaryotic vs eukaryotic Bacteria = one RNA polymerase Eukaryotes have three RNA polymerases (I, II, and III) in
By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food Supervised by Henrik Hasman, PhD
 By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food elibi@food.dtu.dk Supervised by Henrik Hasman, PhD 1. Introduction to plasmid biology 2. Plasmid encoded resistance to β- lactams (basic theories)
By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food elibi@food.dtu.dk Supervised by Henrik Hasman, PhD 1. Introduction to plasmid biology 2. Plasmid encoded resistance to β- lactams (basic theories)
THE THIRD GENERAL TRANSPORT SYSTEM BRANCHED-CHAIN AMINO ACIDS IN SALMONELLA T YPHIMURI UM KEIKO MATSUBARA, KUNIHARU OHNISHI, AND KAZUYOSHI KIRITANI
 J. Gen. Appl. Microbiol., 34, 183-189 (1988) THE THIRD GENERAL TRANSPORT SYSTEM BRANCHED-CHAIN AMINO ACIDS IN SALMONELLA T YPHIMURI UM FOR KEIKO MATSUBARA, KUNIHARU OHNISHI, AND KAZUYOSHI KIRITANI Department
J. Gen. Appl. Microbiol., 34, 183-189 (1988) THE THIRD GENERAL TRANSPORT SYSTEM BRANCHED-CHAIN AMINO ACIDS IN SALMONELLA T YPHIMURI UM FOR KEIKO MATSUBARA, KUNIHARU OHNISHI, AND KAZUYOSHI KIRITANI Department
TER 26. Preview for 2/6/02 Dr. Kopeny. Bacteria and Archaea: The Prokaryotic Domains. Nitrogen cycle
 Preview for 2/6/02 Dr. Kopeny Bacteria and Archaea: The Prokaryotic Domains TER 26 Nitrogen cycle Mycobacterium tuberculosis Color-enhanced images shows rod-shaped bacterium responsible for tuberculosis
Preview for 2/6/02 Dr. Kopeny Bacteria and Archaea: The Prokaryotic Domains TER 26 Nitrogen cycle Mycobacterium tuberculosis Color-enhanced images shows rod-shaped bacterium responsible for tuberculosis
Three types of RNA polymerase in eukaryotic nuclei
 Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
Microbiology Helmut Pospiech
 Microbiology http://researchmagazine.uga.edu/summer2002/bacteria.htm 05.04.2018 Helmut Pospiech The Species Concept in Microbiology No universally accepted concept of species for prokaryotes Current definition
Microbiology http://researchmagazine.uga.edu/summer2002/bacteria.htm 05.04.2018 Helmut Pospiech The Species Concept in Microbiology No universally accepted concept of species for prokaryotes Current definition
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett. Introduction
 The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
Pseudogenes are considered to be dysfunctional genes
 Pseudogenes and bacterial genome decay Jean O Micks Contrary to the evolutionary idea of junk DNA, many pseudogenes still have function in the genomes of archaea, bacteria, and also eukaryotes, such as
Pseudogenes and bacterial genome decay Jean O Micks Contrary to the evolutionary idea of junk DNA, many pseudogenes still have function in the genomes of archaea, bacteria, and also eukaryotes, such as
DNA Technology, Bacteria, Virus and Meiosis Test REVIEW
 Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
Be prepared to turn in a completed test review before your test. In addition to the questions below you should be able to make and analyze a plasmid map. Prokaryotic Gene Regulation 1. What is meant by
BACTERIA. CLS 212: Medical Microbiology Miss Zeina Alkudmani
 BACTERIA CLS 212: Medical Microbiology Miss Zeina Alkudmani Prokaryotes Prokaryotic cells possess simpler structures than eukaryotic cells, since they do not have a nucleus or a lot of cytoplasmic organelles.
BACTERIA CLS 212: Medical Microbiology Miss Zeina Alkudmani Prokaryotes Prokaryotic cells possess simpler structures than eukaryotic cells, since they do not have a nucleus or a lot of cytoplasmic organelles.
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools. Evaluation. Course Homepage.
 CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
A Gene (sleb) Encoding a Spore Cortex-Lytic Enzyme from Bacillus subtilis and Response of the Enzyme to
 JOURNAL OF BACTERIOLOGY, Oct. 1996, p. 6059 6063 Vol. 178, No. 20 0021-9193/96/$04.00 0 Copyright 1996, American Society for Microbiology A Gene (sleb) Encoding a Spore Cortex-Lytic Enzyme from Bacillus
JOURNAL OF BACTERIOLOGY, Oct. 1996, p. 6059 6063 Vol. 178, No. 20 0021-9193/96/$04.00 0 Copyright 1996, American Society for Microbiology A Gene (sleb) Encoding a Spore Cortex-Lytic Enzyme from Bacillus
Considerations with Antibiotic Therapy PART
 Considerations with Antibiotic Therapy PART 1 The Wonderful World of Microbiology 1 Despite the promises of the household-products industry, almost every surface is covered in microorganisms almost all
Considerations with Antibiotic Therapy PART 1 The Wonderful World of Microbiology 1 Despite the promises of the household-products industry, almost every surface is covered in microorganisms almost all
Most common dose (mg) 1g x 1 1g x 1 1g x 1 1g x 1 1g x 1 1g x 1. Maximum dose schedule (mg) 1g x 1 1g x 1 1g x 1 1g x 1 1g x 1 1g x 1
 Ertapenem Rationale for the EUCAST clinical breakpoints, version 1.3 1 st June 2009 Introduction Ertapenem is a carbapenem, available only for parenteral use. Ertapenem is relevant for therapy of septicaemia,
Ertapenem Rationale for the EUCAST clinical breakpoints, version 1.3 1 st June 2009 Introduction Ertapenem is a carbapenem, available only for parenteral use. Ertapenem is relevant for therapy of septicaemia,
Telithromycin in vitro
 in vitro Telithromycin in vitro MBCMIC Staphylococcus aureus Enterococcus faecalis0 Streptococcus pneumoniae Haemophilus influenzae telithromycin MIC MBC erythromycin Aclarithromycin azithromycin josamycin
in vitro Telithromycin in vitro MBCMIC Staphylococcus aureus Enterococcus faecalis0 Streptococcus pneumoniae Haemophilus influenzae telithromycin MIC MBC erythromycin Aclarithromycin azithromycin josamycin
Biased biological functions of horizontally transferred genes in prokaryotic genomes
 Biased biological functions of horizontally transferred genes in prokaryotic genomes Yoji Nakamura 1,5, Takeshi Itoh 2,3, Hideo Matsuda 4 & Takashi Gojobori 1,2 Horizontal gene transfer is one of the main
Biased biological functions of horizontally transferred genes in prokaryotic genomes Yoji Nakamura 1,5, Takeshi Itoh 2,3, Hideo Matsuda 4 & Takashi Gojobori 1,2 Horizontal gene transfer is one of the main
N o hal June 2007
 LABORATOIRE D INFORMATIQUE DE NANTES-ATLANTIQUE A large-scale analysis for significance assessment of frequencies relative to potentially strong sigma 70 promoters: comparison of 32 prokaryotic Christine
LABORATOIRE D INFORMATIQUE DE NANTES-ATLANTIQUE A large-scale analysis for significance assessment of frequencies relative to potentially strong sigma 70 promoters: comparison of 32 prokaryotic Christine
Kharkov National Medical University. Head of Microbiology, Virology and Immunology Department Minukhin Valeriy Vladimirivich
 Kharkov National Medical University Head of Microbiology, Virology and Immunology Department Minukhin Valeriy Vladimirivich Tkachenko Victoria 1, 5, 11, 14, 19, 21, 30 Kovalenko Natalia 2, 12, 25, 29 Siritsa
Kharkov National Medical University Head of Microbiology, Virology and Immunology Department Minukhin Valeriy Vladimirivich Tkachenko Victoria 1, 5, 11, 14, 19, 21, 30 Kovalenko Natalia 2, 12, 25, 29 Siritsa
arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001
![arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001 arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001](/thumbs/77/75258851.jpg) Measure representation and multifractal analysis of complete genomes Zu-Guo Yu,2, Vo Anh and Ka-Sing Lau 3 Centre in Statistical Science and Industrial Mathematics, Queensland University of Technology,
Measure representation and multifractal analysis of complete genomes Zu-Guo Yu,2, Vo Anh and Ka-Sing Lau 3 Centre in Statistical Science and Industrial Mathematics, Queensland University of Technology,
Microbiology. Definition of a Microorganism. Microorganisms in the Lab. The Study of Microorganisms
 Microbiology The Study of Microorganisms Definition of a Microorganism Derived from the Greek: Mikros, «small» and Organismos, organism Microscopic organism which is single celled (unicellular) or a mass
Microbiology The Study of Microorganisms Definition of a Microorganism Derived from the Greek: Mikros, «small» and Organismos, organism Microscopic organism which is single celled (unicellular) or a mass
Product Catalogue 2015 Clinical and Industrial Microbiology
 Acinetobacter baumannii ATCC BAA-747 * 89141 Actinomyces odontolyticus ATCC 17929 * 89114 Aeromonas hydrophila ATCC 7966 * 89119 Aggregatibacter aphrophilus ATCC 7901 * 89091 Aspergillus brasiliensis ATCC
Acinetobacter baumannii ATCC BAA-747 * 89141 Actinomyces odontolyticus ATCC 17929 * 89114 Aeromonas hydrophila ATCC 7966 * 89119 Aggregatibacter aphrophilus ATCC 7901 * 89091 Aspergillus brasiliensis ATCC
Antibiotic Resistance in Escherichia coli Iron Transport Mutants
 Bowling Green State University ScholarWorks@BGSU Honors Projects Honors College Fall 12-11-2017 Antibiotic Resistance in Escherichia coli Iron Transport Mutants Madeline Brandt mbrandt@bgsu.edu Follow
Bowling Green State University ScholarWorks@BGSU Honors Projects Honors College Fall 12-11-2017 Antibiotic Resistance in Escherichia coli Iron Transport Mutants Madeline Brandt mbrandt@bgsu.edu Follow
LECTURE 13. THE BACTERIA (cont.) Photosynthetic Bacteria, phylogenetically widespread. And many Proteobacteria. Photosynthetic Bacteria
 Photosynthetic Bacteria, phylogenetically widespread LECTURE 13 THE BACTERIA (cont.) And many Proteobacteria Photosynthetic Bacteria > Green Sulfur > Green Nonsulfur > Purple Sulfur > Purple Nonsulfur
Photosynthetic Bacteria, phylogenetically widespread LECTURE 13 THE BACTERIA (cont.) And many Proteobacteria Photosynthetic Bacteria > Green Sulfur > Green Nonsulfur > Purple Sulfur > Purple Nonsulfur
phenomenon called cross resistance. As a consequence of cross resistance the entire class of aminoglycosides looses its therapeutic potential.
 Experiment 25 Laboratory to Biology III Diversity of Microorganisms / Wintersemester / page 1 Mechanisms of aminoglycoside resistance in mycobacteria Advisor P.D. Dr. Peter Sander, psander@immv.unizh.ch,
Experiment 25 Laboratory to Biology III Diversity of Microorganisms / Wintersemester / page 1 Mechanisms of aminoglycoside resistance in mycobacteria Advisor P.D. Dr. Peter Sander, psander@immv.unizh.ch,
ydci GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC
 Table S1. DNA primers used in this study. Name ydci P1ydcIkd3 Sequence GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC Kd3ydcIp2 lacz fusion YdcIendP1 YdcItrgP2 GAC AGC
Table S1. DNA primers used in this study. Name ydci P1ydcIkd3 Sequence GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC Kd3ydcIp2 lacz fusion YdcIendP1 YdcItrgP2 GAC AGC
PROTEIN SYNTHESIS INTRO
 MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
Bio Microbiology - Spring 2012 Learning Guide 04.
 Bio 230 - Microbiology - Spring 2012 Learning Guide 04 http://pessimistcomic.blogspot.com/ A septum assembles at the center of the cell. This molecular "purse string" is linked to the inner surface of
Bio 230 - Microbiology - Spring 2012 Learning Guide 04 http://pessimistcomic.blogspot.com/ A septum assembles at the center of the cell. This molecular "purse string" is linked to the inner surface of
