Multifractal characterisation of complete genomes
|
|
|
- Meryl Snow
- 6 years ago
- Views:
Transcription
1 arxiv:physics/1854v1 [physics.bio-ph] 28 Aug 21 Multifractal characterisation of complete genomes Vo Anh 1, Ka-Sing Lau 2 and Zu-Guo Yu 1,3 1 Centre in Statistical Science and Industrial Mathematics, Queensland University of Technology, GPO Box 2434, Brisbane, Q41, Australia 2 Department of Mathematics, Chinese University of Hong Kong, Shatin, Hong Kong 3 Department of Mathematics, Xiangtan University, Hunan 41115, P.R. China Abstract This paper develops a theory for characterisation of DNA sequences based on their measure representation. The measures are shown to be random cascades generated by an infinitely divisible distribution. This probability distribution is uniquely determined by the exponent function in the multifractal theory of random cascades. Curve fitting to a large number of complete genomes of bacteria indicates that the Gamma density function provides an excellent fit to the exponent function, and hence to the probability distribution of the complete genomes. 1 Introduction DNA sequences are of fundamental importance in understanding living organisms, since all information on their hereditary evolution is contained in these macromolecules. One of the challenges of DNA sequence analysis is to determine the patterns of these sequences. It is useful to distinguish coding from noncoding sequences. Problems related to the classification and evolution of organisms using DNA sequences are also important. Fractal analysis has proved useful in revealing complex patterns in natural objects. Berthelsen et al. [2] considered the global fractal dimension of human DNA sequences treated as pseudorandom walks. Vieira [3] carried out a low-frequency analysis of the complete DNA of 13 microbial genomes and showed that their fractal behaviour does not always prevail through the entire chain and the autocorrelation functions have a rich variety of behaviours including the presence of anti-persistence. Provata and Almirantis [14] proposed 1 address: v.anh@qut.edu.au, kslau@math.cuhk.edu.hk, yuzg@hotmail.com or z.yu@qut.edu.au 2 The URL of Vo Anh: 1
2 a fractal Cantor pattern of DNA. They mapped coding segments to filled regions and noncoding segments to empty regions of a random Cantor set and then calculated the fractal dimension of this set. They found that the coding/noncoding partition in DNA sequences of lower organisms is homogeneous-like, while in the higher eucariotes the partition is fractal. Yu and Anh [16] proposed a time series model based on the global structure of the complete genome and found that one can get more information from this model than that of fractal Cantor pattern. Some results on the classification and evolution relationship of bacteria were found in [16]. The correlation property of length sequences was discussed in [17]. Although statistical analysis performed directly on DNA sequences has yielded some success, there has been some indication that this method is not powerful enough to amplify thedifference between adnasequence andarandomsequence aswell astodistinguish DNA sequences themselves in more details. One needs more powerful global and visual methods. For this purpose, Hao et al. [7] proposed a visualisation method based on coarse-graining and counting of the frequency of appearance and strings of a given length. They called it the portrait of an organism. They found that there exist some fractal patterns in the portraits which are induced by avoiding and under-represented strings. The fractal dimension of the limit set of portraits was discussed in [18, 8]. There are other graphical methods of sequence patterns, such as chaos game representation (see [1, 5]). In the portrait representation, Hao et al. [7] used squares to represent substrings and discrete colour grades to represent the frequencies of the substrings in the complete genome. It is difficult to know the accurate value of the frequencies of the substrings from the portrait representation. And they did not discuss the classification and evolution problem. In order to improve it, Yu et al [15] used subintervals in one-dimensional space to represent substrings to obtain an accurate histogram of the substrings in the complete genome. The histogram, viewed as a probability measure and was called the measure representation of the complete genome, gives a precise compression of the genome. Multifractal analysis was then proposed in Yu et al [15] to treat the classification and evolution problem based on the measure representation of different organisms. In this paper, we go one step further and provide a complete characterisation of the DNA sequences based on their measure representation. This is given in the form of the probability density function of the measure. We first show that the given measure is in fact a multiplicative cascade generated by an infinitely divisible distribution. This probability distributionisuniquely determined bytheexponent K(q),q,inthemultifractalanalysis of the cascade. This theory will be detailed in the next section. We then apply the theory on a large number of typical genomes. It will be seen that the Gamma density function provides an excellent fit to the K(q) curve of each genome. This characterisation therefore provides a needed tool to study the evolution of organisms. 2
3 2 Measure representation of complete genome We first outline the method of Yu et al [15] in deriving the measure representation of a DNA sequence. Such a sequence is formed by four different nucleotides, namely adenine (a), cytosine (c), guanine (g) and thymine (t). We call any string made of K letters from the set {g,c,a,t} a K-string. For a given K there are in total 4 K different K-strings. In order to count the number of each kind of K-strings in a given DNA sequence, 4 K counters are needed. We divide the interval [,1[ into 4 K disjoint subintervals, and use each subinterval to represent a counter. Letting s = s 1 s K,s i {a,c,g,t},i = 1,,K, be a substring with length K, we define x l (s) = K i=1 x i 4i, (2.1) where x i =, if s i = a, 1, if s i = c, 2, if s i = g, 3, if s i = t, (2.2) and x r (s) = x l (s)+ 1 4K. (2.3) We then use the subinterval [x l (s),x r (s)) to represent substring s. Let N(s) be the times of substring s appearing in the complete genome. If the number of bases in the complete genome is L, we define F(s) = N(s)/(L K +1) (2.4) to be the frequency of substring s. It follows that {s}f(s) = 1. Now we can define a measure µ K on [,1) by where µ K (dx) = Y K (x)dx, Y K (x) = 4 K F K (s), x [x l (s),x r (s)). (2.5) We then have µ K ([,1)) = 1 and µ K ([x l (s),x r (s))) = F K (s). We call µ K (x) the measure representation of an organism. As an example, the measure representation of M. genitalium for K = 3,...,8 is given in Figure 1. Self-similarity is apparent in the measures. More than 33 bacterial complete genomes are now available in public databases. There are six Archaebacteria (Archaeoglobus fulgidus, Pyrococcus abyssi, Methanococcus jannaschii, Pyrococcus horikoshii, Aeropyrum pernix and Methanobacterium thermoautotrophicum); five 3
4 Gram-positive Eubacteria(Mycobacterium tuberculosis, Mycoplasma pneumoniae, Mycoplasma genitalium, Ureaplasma urealyticum, and Bacillus subtilis). The others are Gram-negative Eubacteria, which consist of two Hyperthermophilic bacteria (Aquifex aeolicus and Thermotoga maritima); five Chlamydia (Chlamydia trachomatisserovar, Chlamydia muridarum, Chlamydia pneumoniae and Chlamydia pneumoniae AR39); two Spirochaete(Borrelia burgdorferi and Treponema pallidum); one Cyanobacterium (Synechocystis sp. PCC683); and thirteen Proteobacteria. The thirteen Proteobacteria are divided into four subdivisions, which are alpha subdivision (Rhizobium sp. NGR234 and Rickettsia prowazekii); gamma subdivision (Escherichia coli, Haemophilus influenzae, Xylella fastidiosa, Vibrio cholerae, Pseudomonas aeruginosa and Buchnera sp. APS); beta subdivision (Neisseria meningitidis MC58 and Neisseria meningitidis Z2491); epsilon subdivision (Helicobacter pylori J99, Helicobacter pylori and Campylobacter jejuni). The complete sequences of some chromosomes of higher organisms are also currently available. We selected the sequences of Chromosome 15 of Saccharomyces cerevisiae, Chromosome 3 of Plasmodium falciparum, Chromosome 1 of Caenorhabditis elegans, Chromosome 2 of Arabidopsis thaliana and Chromosome 22 of Homo sapiens. In our previous work (Yu et al [15]), we calculated the numerical dimension spectra D q (defined in next section) for all above organisms and for different K. For small K, there are only a few different K-strings, so there is not enough information for any clear-cut result. We find that the D q curves are very close to one another for K = 6,7,8 for each organism. Hence it would be appropriate to take K = 8 if we want to use the D q curves to discuss the classification and evolution problem. It is still needed to know what is the analytical expression of the dimension spectra. The main aim of this paper is to establish a theoretical model to give such an analytical expression. 3 Multifractal models Let ε(t) be a positive stationary stochastic process on a bounded interval of R, assumed to be the unit interval [,1] for convenience, with Eε(t) = 1. The smoothing of ε(t) at scale r > is defined as ε r (t) = 1 r For < r < u < v, we consider the processes t+r/2 t r/2 X r,v (t) = ε r(t) ε v (t), t [,1]. ε(s)ds. (3.1) Following Novikov [13], we assume the following scale invariance conditions: (i) The random variables X r,u and X u,v are independent; (ii) The probability distribution of each random variable X u,v depends only on the ratio u/v of the corresponding scales. 4
5 These conditions imply the power-law form for the moments of the processes X u,v if they exist. In fact, we may write E(X u,v (t)) q = g q ( u v ), q (3.2) from condition (ii) for some function g which also depends on q. From the identity X r,v (t) = X r,u (t)x u,v (t) and condition (i) we get ( ) ( ) ( ) r r u g q = g q g q. (3.3) v u v Since u is arbitrary, we then have for some function K(q) with K() =. It follows that Writing Y for X r,v we obtain ( ) ( ) r r K(q) g q = (3.4) v v K(q) = lne(x r,v(t)) q. ln(v/r) K (q) = 1 ln(v/r) E(Y q lny), E(Y q ) Since K (q) = 1 (EY q )E ( Y q (lny) 2) (E(Y q lny)) 2 ln(v/r) (EY q ) 2. (E(Y q lny)) 2 = ( E ( Y q/2 Y q/2 lny )) 2 (EY q )E ( Y q (lny) 2) (3.5) by Schwarz s inequality and v/r > 1, we get K (q), that is, K(q) is a convex function. It is noted that equality holds in (3.5) only if K(q) is a linear function of q; other than this, K(q) is a strictly convex function. For < q < 1, we assume that K(q) <, which reflects the fact that, in this range, taking a qth-power necessarily reduces the singularity of X u,v. Also, we assume that the probability density function of X u,v is skewed in the positive direction. This yields that K(q) > for q > 1. These assumptions, in conjunction with the strict convexity of K(q), suggest the assumption that K(1) =. (3.6) 5
6 This implies that EX u,v = 1 for arbitrary < u < v. (3.7) In this paper, we will consider smoothing at discrete scales r j = 2 j+1, j =,1,2,3,... Then the smoothed process at scale r j is X j (t) = ε rj (t) = 1 t+2 j ε(s)ds. (3.8) 2 j+1 t 2 j Under the condition Eε(t) = 1, it is reasonable to assume that Then, at generation J, X (t) = 1, t [,1]. (3.9) X J (t) = X (t) X 1(t) X 2 (t) X (t) X 1 (t)... X J (t) X J 1 (t) = X 1(t) X 2 (t) X (t) X 1 (t)... X J (t) X J 1 (t). (3.1) Under the scale invariance conditions (i) and(ii), the randomvariables X j /X j 1 of (3.1) are independent and have the same probability distribution. Let W denote a generic member of this family. Note that EW = 1 from (3.7). Then (3.1) can be rewritten as X J (t) = X J 1 (t) X J (t) X J 1 (t) = W 1 (t)w 2 (t)...w J (t), t [,1]. (3.11) In other words, X J (t) is a multiplicative cascade process (see Holley and Waymire [9], Gupta and Waymire [6]). Denote by µ J the sequence of random measures defined by the density X J (t), that is, µ J (dt) = X J (t)dt, J = 1,2,3,... It can be checked that µ J a.s. has a weak* limit µ since for each bounded continuous functionf on[,1],thesequence [,1] fdµ J isanl 1 -boundedmartingale(seeholleyandwaymire [9], Mandelbrot [12], Kahane and Peyriere [11]). We denote the density corresponding to µ by X (t). Then it is seen from (3.8) that Summarising, we have established that X (t) = ε(t), t [,1]. (3.12) The positive stationary process ε(t) is the limit of a multiplicative cascade with generator W. 6
7 We next want to characterise this random cascade. We first note that, for j = 1,2,3,..., X j from the positivity of ε(t). Thus, X j 1 = 2 This inequality together with (3.4) imply E t+2 j t 2 j ε(s)ds t+2 (j 1) t 2 ε(s)ds 2 (3.13) (j 1) ( Xj X j 1 ) q 2 q. K(q) q, q. (3.14) We then have ( ) 2q Xj E q= X j 1 1 2q 1 = q=( 2 ) K(2q) 2q =. In other words, the Carleman condition is satisfied (see Feller [4], p. 224). As a result, we get The probability density function f W of the generator W is uniquely determined by the set {K(q), q =,1,2,...}. It is seen that, if the function K(q) has analytic continuation into the complex plane, then the characteristic function of lnw has the form ψ(x) = E ( e ixlnw) ( ) 1 K(ix) =. (3.15) 2 Define ψ n (x) = ( 1/2 1/n) K(ix) for an arbitrary integer n. Then ψn is the characteristic function of the probability distribution corresponding to smoothing with scales ( 2 1/n) j+1. Also, it holds that ψ(x) = (ψ n (x)) n. Thus ψ(x) is infinitely divisible (see Feller [4], p. 532); in other words, ln W has an infinitely divisible distribution. (3.16) It is noted from(3.13) that ln W. The most general formforthe characteristic function 2 ϕ(x) of positive random variables is given by { 1 e ixs } ϕ(x) = exp P (ds)+iax, (3.17) s 7
8 where a and P is a measure on the open interval (, ) such that (1+s) 1 P (ds) < (see Feller [4], p. 539). On the other hand, it follows from (3.2) and (3.4) that the characteristic function of ln W is given by 2 E ( e ixln W 2 ) = 2 ix E(W) ix Using q = ix and equating (3.17) with (3.18) then yields K(q) = = 2 ix ( 1 2) K( ix). (3.18) ( 1 a ) 1 e qs P (ds) q ln2 s ln2. (3.19) As constrained by (3.6), the following condition must be satisfied by the measure P (ds) : 1 e s s P (ds) ln2 = 1 a 1. (3.2) ln2 Equations (3.19) and (3.2) provide the most general form for the K(q) curve of the positive random process {ε(t), t 1}. Inpractice, fittingthisk(q)curvetodatarequiresaproperchoiceofthemeasure P (ds). Novikov [13] suggests the use of the Gamma density function, namely, f (x) = Ax α 1 exp( x/σ), (3.21) where P (dx) = f (x)dx and A,α,σ are positive constants. From (3.19) and (3.21) we get where κ = 1 a/ln2, and from (3.2) we have κ ( ) q (qσ+1)1 α 1 (σ+1) K(q) = 1 α 1, α 1, κ ( ) (3.22) q ln(qσ+1) ln(σ+1), α = 1. A = κln2 σ α 1 Γ(α 1) (1 (σ +1)1 α ) 1. The form (3.22) will be used for data fitting in this paper. It is seen from (3.2) and (3.4) that the data for the K(q) curve is provided by lne(x K(q) = lim J) q J ln2 J+1, (3.23) where it should be noted from (3.12) that X (t) = ε(t), the given positive random process. Since each smoothed process X J may possess long-range dependence (see Anh et al. [1]), the ergodic theorem may not hold for these processes. As a result, the computation of E(X q J) as sample averages may not be sufficiently accurate. There is an alternative form of the ergodic theorem developed by Holley and Waymire [9] for random cascades which we now summarise. 8
9 For random cascades with density ε(t), limit measure µ, branching number b and generator W, define M J (q) = k ( µ ( J k )) q, (3.24) lnm J (q) τ (q) = lim, (3.25) J J lnb D q = τ(q)/(q 1), (3.26) χ b (q) = log b E(W q ) (q 1), (3.27) where the prime in (3.24) indicates a sum over those subintervals J k meet the support of µ. of generation J which Theorem 1 (Holley and Waymire [9]) Assume that W > a for some a > and W < b with probability 1, and that E(W 2q )/(EW q ) 2 < b. Then, with probability 1, τ (q) = χ b (q). (3.28) Inourcaseasdevelopedabove, b = 2,and(3.13)givesW 2. Infactthescaler j = 2 j+1 used in (3.8) is arbitrary; it can be b j+1 and the inequality W b still holds by definition of the smoothing and the positivity of ε(t). In our development, Consequently, K(q) = lim J lne(x q J ) ln2 J+1 = lim J J lne(w q ) (J 1)ln2 1 using (3.11) = lne(wq ). ln2 K(q) = τ (q)+q 1. (3.29) The above formula then provides a way to compute K(q) via (3.25) and (3.29) using sums of q-th powers of the limit measure instead of (3.23) using expectations. In fact, the ergodic theorem now takes the following form lne(x q J lim ) (J 1)ln2 = lim ln ( ( )) q k µ J k J J ln2 J +q 1. 9
10 4 Data fitting and discussion For K = 8, we first calculated K(q) of the measure representation of all the above organisms directly from the definition of K(q) (3.23). Figure 2 shows how to calculate this K(q) curve. We give the K(q) curves of E. coli, S. cerevisiae Chr15, C. elegans Chr1, A. thaliana Chr2, and Homo sapiens Chr22 in Figure 3. From Figure 3, it is seen that the grade of the organism is lower when the K q curve is flatter. Hence the evolution relationship of these organisms is apparent. We denote by K d (q) the value of K(q) computed from the data using its definition (3.23) and define error = J j=1 κ(q j (q jσ +1) 1 α 1 (σ +1) 1 α 1 ) K d(q j ) 2. Then the values of κ, σ and α can be estimated through minimising error. In this minimisation, we assume κ, σ, α 2. After obtaining the value of κ, σ and α, we then get the K(q) curve from (3.22). The data fitting based on the form (3.22) was performed on all the organisms and shown in Table 1 (from top to bottom, in the increasing order of the value of κ). It is found that the form (3.22) gives a perfect fit to the data for all bacteria. As an example, we give the data fitting of E. coli, S. cerevisiae Chr15 and C. elegans Chr1 in Figure 4. But for higher organisms, for example, Homo sapiens Chromosome 22, the fitting is not as good. Note that we only selected one chromosome for each higher organism. If all chromosomes for each higher organism are considered, the data fitting for K q will be better. The fit for Human Chromosome is the worst in Table 1. Since the length of Human Chromosome 22 is not larger than those of the complete genomes of all bacteria, there does not seem to be any relationship between the quality of fit and the length of the complete genome. The parameter κ provides a tool to classify bacteria. From Table 1, one can see Helicobacter pylori and Helicobacter pylori J99 group together, and three Chlamydia almost group together. But this parameter κ alone is not sufficient, it must be combined with other tools to classify bacteria. We also calculated the values of τ(q) using its definition (3.25). We found the values of K d (q) coincide with those obtained from (3.29). Hence we indeed can use (3.29) to calculate K(q). Formula (3.22) gives an analytical expression for the quantity K q. An analytical expressions for τ(q) can therefore be obtained from (3.29) and D q from (3.26). 5 Conclusions The idea of our measure representation is similar to the portrait method proposed by Hao et al.[7]. It provides a simple yet powerful visualisation method to amplify the difference between a DNA sequence and a random sequence as well as to distinguish DNA sequences 1
11 themselves in more details. From our measure representation we can exactly know the frequencies of all the K-string appearing in the complete genome. But the representations alone are not sufficient to discuss the classification and evolution problem. Hence we need further tools. In our previous work (Yu et al [15]), when the measure representations of organisms were viewed as time series, it was found that they are far from being random time series, and in fact exhibit strong long-range dependence. Multifractal analysis of the complete genomes was performed in relation to the problem of classification and evolution of organisms. In this paper, we established a theoretical model of the probability distribution of the complete genomes. This probability distribution, particularly the resulting K(q) curve, provides a precise tool for their characterisation. Numerical results confirm the accuracy of the method of this paper. For a completely random sequence based on the alphabet {a,c,g,t}, we have D q = 1, τ(q) = q 1, K(q) = for all q. From the K(q) curves, it is seen that all complete genomes selected are far from being a completely random sequence. Acknowledgement The Authors would like to express their gratitude to the referees for good comments and suggestions to improve this paper. This research was partially supported by QUT Postdoctoral Research Grant to Zu-Guo Yu, and the RGC Earmarked Grant CUHK 4215/99P. References [1] V. V. Anh, C. C. Heyde, and Q. Tieng. Stochastic models for fractal processes. Journal of Statistical Planning and Inference, 8(1/2): , [2] C. L. Berthelsen, J. A. Glazier, and S. Raghavachari. Phys. Rev. E, 49(3):186, [3] M. de Sousa Vieira. Statistics of DNA sequences: A low-frequency analysis. Phys. Rev. E, 6(5): , [4] W. Feller. An Introduction to Probability Theory and its Applications, volume II. Wiley, New York, [5] N. Goldman. Nucleotide, dinucleotide and trinucleotide frequencies explain patterns observed in chaos game representations of DNA sequences. Nucleic Acids Research, 21(1): , [6] V. K. Gupta and E. C. Waymire. A statistical analysis of mesoscale rainfall as a random cascade. Journal of Applied Meteorology, 32: , [7] B.-L. Hao, H.-C. Lee, and S.-Y. Zhang. Fractals related to long DNA sequences and complete genomes. Chaos, Solitons and Fractals, 11(6): , 2. [8] B.-L. Hao, H.-M. Xie, Z.-G. Yu, and G.-Y. Chen. Avoided strings in bacterial complete genomes and a related combinatorial problem. Ann. of Combinatorics, 4: , 2. [9] R. Holley and E. C. Waymire. Multifractal dimensions and scaling exponents for strongly bounded random cascades. The Annals of Applied Probability, 2(4): ,
12 [1] H. J. Jeffrey. Chaos game representation of gene structure. Nucleic Acids Research, 18(8): , 199. [11] J.-P. Kahane and J. Peyrière. Sur certaines martingales de benoit mandelbrot. Advances in Mathematics, 22: , [12] B. B. Mandelbrot. Intermittent turbulence in self-similar cascades: divergence of high moments and dimension of the carrier. J. Fluid Mech., 62: , [13] E. A. Novikov. Infinitely divisible distributions in turbulence. Physical Review E, 5(5), [14] A. Provata and Y. Almirantis. Fractal cantor patterns in the sequence structure of DNA. Fractals, 8(1):15 27, 2. [15] Z.-G. Yu, V. Anh and K.-S. Lau. Measure representation and multifractal analysis of complete genomes. Phys. Rev. E, Sept. of 21, (To appear). [16] Z.-G. Yu and V. Anh. Time series model based on global structure of complete genome. Chaos, Solitons and Fractals, 12(1): , 21. [17] Z.-G. Yu, V. V. Anh, and B. Wang. Correlation property of length sequences based on global structure of complete genome. Phys. Rev. E, 63: 1193, 21. [18] Z.-G. Yu, B.-L. Hao, H.-M. Xie, and G.-Y. Chen. Dimension of fractals related to language defined by tagged strings in complete genome. Chaos, Solitons and Fractals, 11(14): , 2. 12
13 Table 1: The values of κ, σ, α, error of all the organisms selected. Species κ σ α error Aquifex aeolicus E-3 Haemophilus influenzae E-4 Synechocystis sp. PCC E-4 Mycoplasma pneumoniae E-4 Chlamydia pneumoniae AR E-2 Rhizobium sp. NGR E-5 Chlamydia muridarum E-3 Chlamydia trachomatis E-3 Neisseria meningitidis MC E-4 Helicobacter pylori E-3 Helicobacter pylori J E-3 Methanococcus jannaschii E-5 Rickettsia prowazekii E-4 Neisseria meningitidis Z E-4 Bacillus subtilis E-3 Aeropyrum pernix E-2 Mycoplasma genitalium E-3 Campylobacter jejuni E-3 M. tuberculosis E-4 Borrelia burgdorferi E-3 Thermotoga maritima E-3 Treponema pallidum E-3 Ureaplasma urealyticum E-4 Escherichia coli E-4 M. thermoautotrophicum E-3 Pseudomonas aeruginosa E-5 Caenorhabditis elegans Chr E-2 Chlamydia pneumoniae AR E-3 Archaeoglobus fulgidus E-3 S. cerevisiae Chr E-3 Pyrococcus abyssi E-4 Buchnera sp. APS E-3 Arabidopsis thaliana Chr E-3 Pyrococcus abyssi E-3 Vibrio cholerae E-4 Plasmodium falciparum Chr E-2 Xylella fastidiosa E-2 Homo sapiens Chr E-1 13
14 M. genitalium, K=3.2 M. genitalium, K=4 Probability of substrings Probability of substrings Representation of substrings with length K Representation of substrings with length K.9 M. genitalium, K= M. genitalium, K=6 Probability of substrings Probability of substrings Representation of substrings with length K M. genitalium, K= Representation of substrings with length K.9 M. genitalium, K=8.8.7 Probability of substrings Probability of substrings Representation of substrings with length K Representation of substrings with length K Figure 1: Histograms of substrings with different lengths 14
15 14 12 q=2 q=4 q=6 q=8 1 ln E(Y r q (t)) ln(1/r) Figure 2: An example to show how to obtain the value of K(q) directly using its definition E. coli Yeast Chr15 C. elegans Chr1 A. thaliana Chr2 Human Chr K(q) q Figure 3: The values of K(q) of Chromosome 22 of Homo sapiens, Chromosome 2 of A. thaliana, Chromosome 1 of C. elegans, Chromosome 15 of S. cerevisiae and E. coli. 15
16 3.5 3 E. coli Yeast Chr15 C. elegans Chr K(q) q Figure 4: The data fitting of E. coli,chromosome 15 of S. cerevisiae and chromosome 1 of C. elegans based on the Gamma model. The symbolled curves represent K d (q) computed from data, while the continuous curves represent K(q) computed from formula (3.22). 16
Measure representation and multifractal analysis of complete genomes
 PHYSICAL REVIEW E, VOLUME 64, 031903 Measure representation and multifractal analysis of complete genomes Zu-Guo Yu, 1,2, * Vo Anh, 1 and Ka-Sing Lau 3 1 Centre in Statistical Science and Industrial Mathematics,
PHYSICAL REVIEW E, VOLUME 64, 031903 Measure representation and multifractal analysis of complete genomes Zu-Guo Yu, 1,2, * Vo Anh, 1 and Ka-Sing Lau 3 1 Centre in Statistical Science and Industrial Mathematics,
arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001
![arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001 arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001](/thumbs/77/75258851.jpg) Measure representation and multifractal analysis of complete genomes Zu-Guo Yu,2, Vo Anh and Ka-Sing Lau 3 Centre in Statistical Science and Industrial Mathematics, Queensland University of Technology,
Measure representation and multifractal analysis of complete genomes Zu-Guo Yu,2, Vo Anh and Ka-Sing Lau 3 Centre in Statistical Science and Industrial Mathematics, Queensland University of Technology,
Multifractal characterisation of length arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001
![Multifractal characterisation of length arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001 Multifractal characterisation of length arxiv:physics/ v1 [physics.bio-ph] 28 Aug 2001](/thumbs/96/128544122.jpg) Multifractal characterisation of length arxiv:physics/853v [physics.bio-ph] 28 Aug 2 seuences of coding and noncoding segments in a complete genome Zu-Guo Yu,2, Vo Anh and Ka-Sing Lau 3 Centre in Statistical
Multifractal characterisation of length arxiv:physics/853v [physics.bio-ph] 28 Aug 2 seuences of coding and noncoding segments in a complete genome Zu-Guo Yu,2, Vo Anh and Ka-Sing Lau 3 Centre in Statistical
This is the author s version of a work that was submitted/accepted for publication in the following source:
 This is the author s version of a work that was submitted/accepted for publication in the following source: Anh, Vo, Lam, K, Leung, Y, & Yu, Zu-Guo (5) Multifractal Characterization of Hong Kong Air Quality
This is the author s version of a work that was submitted/accepted for publication in the following source: Anh, Vo, Lam, K, Leung, Y, & Yu, Zu-Guo (5) Multifractal Characterization of Hong Kong Air Quality
Correlation property of length sequences based on global structure of complete genome
 Correlation property of length sequences based on global structure of complete genome Zu-Guo Yu 1,2,V.V.Anh 1 and Bin Wang 3 1 Centre for Statistical Science and Industrial Mathematics, Queensland University
Correlation property of length sequences based on global structure of complete genome Zu-Guo Yu 1,2,V.V.Anh 1 and Bin Wang 3 1 Centre for Statistical Science and Industrial Mathematics, Queensland University
The genomic tree of living organisms based on a fractal model
 Physics Letters A 317 (2003) 293 302 www.elsevier.com/locate/pla The genomic tree of living organisms based on a fractal model Zu-Guo Yu a,b,,voanh a, Ka-Sing Lau c, Ka-Hou Chu d a Program in Statistics
Physics Letters A 317 (2003) 293 302 www.elsevier.com/locate/pla The genomic tree of living organisms based on a fractal model Zu-Guo Yu a,b,,voanh a, Ka-Sing Lau c, Ka-Hou Chu d a Program in Statistics
Evolutionary Analysis by Whole-Genome Comparisons
 JOURNAL OF BACTERIOLOGY, Apr. 2002, p. 2260 2272 Vol. 184, No. 8 0021-9193/02/$04.00 0 DOI: 184.8.2260 2272.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Evolutionary Analysis
JOURNAL OF BACTERIOLOGY, Apr. 2002, p. 2260 2272 Vol. 184, No. 8 0021-9193/02/$04.00 0 DOI: 184.8.2260 2272.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Evolutionary Analysis
2 Genome evolution: gene fusion versus gene fission
 2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
2 Genome evolution: gene fusion versus gene fission Berend Snel, Peer Bork and Martijn A. Huynen Trends in Genetics 16 (2000) 9-11 13 Chapter 2 Introduction With the advent of complete genome sequencing,
ABSTRACT. As a result of recent successes in genome scale studies, especially genome
 ABSTRACT Title of Dissertation / Thesis: COMPUTATIONAL ANALYSES OF MICROBIAL GENOMES OPERONS, PROTEIN FAMILIES AND LATERAL GENE TRANSFER. Yongpan Yan, Doctor of Philosophy, 2005 Dissertation / Thesis Directed
ABSTRACT Title of Dissertation / Thesis: COMPUTATIONAL ANALYSES OF MICROBIAL GENOMES OPERONS, PROTEIN FAMILIES AND LATERAL GENE TRANSFER. Yongpan Yan, Doctor of Philosophy, 2005 Dissertation / Thesis Directed
The Minimal-Gene-Set -Kapil PHY498BIO, HW 3
 The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
The Minimal-Gene-Set -Kapil Rajaraman(rajaramn@uiuc.edu) PHY498BIO, HW 3 The number of genes in organisms varies from around 480 (for parasitic bacterium Mycoplasma genitalium) to the order of 100,000
Evolutionary Use of Domain Recombination: A Distinction. Between Membrane and Soluble Proteins
 1 Evolutionary Use of Domain Recombination: A Distinction Between Membrane and Soluble Proteins Yang Liu, Mark Gerstein, Donald M. Engelman Department of Molecular Biophysics and Biochemistry, Yale University,
1 Evolutionary Use of Domain Recombination: A Distinction Between Membrane and Soluble Proteins Yang Liu, Mark Gerstein, Donald M. Engelman Department of Molecular Biophysics and Biochemistry, Yale University,
Phylogenetic tree of prokaryotes based on complete genomes using fractal and correlation analyses
 Phylogenetic tree of prokaryotes based on complete genomes using fractal and correlation analyses Zu-Guo Yu, and Vo Anh Program in Statistics and Operations Research, Queensland University of Technology,
Phylogenetic tree of prokaryotes based on complete genomes using fractal and correlation analyses Zu-Guo Yu, and Vo Anh Program in Statistics and Operations Research, Queensland University of Technology,
Correlations between Shine-Dalgarno Sequences and Gene Features Such as Predicted Expression Levels and Operon Structures
 JOURNAL OF BACTERIOLOGY, Oct. 2002, p. 5733 5745 Vol. 184, No. 20 0021-9193/02/$04.00 0 DOI: 10.1128/JB.184.20.5733 5745.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Correlations
JOURNAL OF BACTERIOLOGY, Oct. 2002, p. 5733 5745 Vol. 184, No. 20 0021-9193/02/$04.00 0 DOI: 10.1128/JB.184.20.5733 5745.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Correlations
This is the author s version of a work that was submitted/accepted for publication in the following source:
 This is the author s version of a work that was submitted/accepted for publication in the following source: Yu, Zu-Guo (2004) Fourier Transform and Mean Quadratic Variation of Bernoulli Convolution on
This is the author s version of a work that was submitted/accepted for publication in the following source: Yu, Zu-Guo (2004) Fourier Transform and Mean Quadratic Variation of Bernoulli Convolution on
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
Midterm Exam #1 : In-class questions! MB 451 Microbial Diversity : Spring 2015!
 Midterm Exam #1 : In-class questions MB 451 Microbial Diversity : Spring 2015 Honor pledge: I have neither given nor received unauthorized aid on this test. Signed : Name : Date : TOTAL = 45 points 1.
Midterm Exam #1 : In-class questions MB 451 Microbial Diversity : Spring 2015 Honor pledge: I have neither given nor received unauthorized aid on this test. Signed : Name : Date : TOTAL = 45 points 1.
Base Composition Skews, Replication Orientation, and Gene Orientation in 12 Prokaryote Genomes
 J Mol Evol (1998) 47:691 696 Springer-Verlag New York Inc. 1998 Base Composition Skews, Replication Orientation, and Gene Orientation in 12 Prokaryote Genomes Michael J. McLean, Kenneth H. Wolfe, Kevin
J Mol Evol (1998) 47:691 696 Springer-Verlag New York Inc. 1998 Base Composition Skews, Replication Orientation, and Gene Orientation in 12 Prokaryote Genomes Michael J. McLean, Kenneth H. Wolfe, Kevin
Assessing evolutionary relationships among microbes from whole-genome analysis Jonathan A Eisen
 475 Assessing evolutionary relationships among microbes from whole-genome analysis Jonathan A Eisen The determination and analysis of complete genome sequences have recently enabled many major advances
475 Assessing evolutionary relationships among microbes from whole-genome analysis Jonathan A Eisen The determination and analysis of complete genome sequences have recently enabled many major advances
Genomic analysis of the histidine kinase family in bacteria and archaea
 Microbiology (2001), 147, 1197 1212 Printed in Great Britain Genomic analysis of the histidine kinase family in bacteria and archaea Dong-jin Kim and Steven Forst Author for correspondence: Steven Forst.
Microbiology (2001), 147, 1197 1212 Printed in Great Britain Genomic analysis of the histidine kinase family in bacteria and archaea Dong-jin Kim and Steven Forst Author for correspondence: Steven Forst.
Regulatory Sequence Analysis. Sequence models (Bernoulli and Markov models)
 Regulatory Sequence Analysis Sequence models (Bernoulli and Markov models) 1 Why do we need random models? Any pattern discovery relies on an underlying model to estimate the random expectation. This model
Regulatory Sequence Analysis Sequence models (Bernoulli and Markov models) 1 Why do we need random models? Any pattern discovery relies on an underlying model to estimate the random expectation. This model
# shared OGs (spa, spb) Size of the smallest genome. dist (spa, spb) = 1. Neighbor joining. OG1 OG2 OG3 OG4 sp sp sp
 Bioinformatics and Evolutionary Genomics: Genome Evolution in terms of Gene Content 3/10/2014 1 Gene Content Evolution What about HGT / genome sizes? Genome trees based on gene content: shared genes Haemophilus
Bioinformatics and Evolutionary Genomics: Genome Evolution in terms of Gene Content 3/10/2014 1 Gene Content Evolution What about HGT / genome sizes? Genome trees based on gene content: shared genes Haemophilus
Od vzniku života po evoluci člověka. Václav Pačes Ústav molekulární gene2ky Akademie věd České republiky
 Od vzniku života po evoluci člověka Václav Pačes Ústav molekulární gene2ky Akademie věd České republiky Exon 1 Intron (413 nt) Exon 2 Exon 1 Exon 2 + + 15 nt + 4 nt Genome of RNA-bacteriophage
Od vzniku života po evoluci člověka Václav Pačes Ústav molekulární gene2ky Akademie věd České republiky Exon 1 Intron (413 nt) Exon 2 Exon 1 Exon 2 + + 15 nt + 4 nt Genome of RNA-bacteriophage
Gene Family Content-Based Phylogeny of Prokaryotes: The Effect of Criteria for Inferring Homology
 Syst. Biol. 54(2):268 276, 2005 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150590923335 Gene Family Content-Based Phylogeny of Prokaryotes:
Syst. Biol. 54(2):268 276, 2005 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150590923335 Gene Family Content-Based Phylogeny of Prokaryotes:
Where are the pseudogenes in bacterial genomes?
 Opinion TRENDS in Microbiology Vol.9 No. November Where are the pseudogenes in bacterial genomes? Jeffrey G. Lawrence, Roger W. Hendrix and Sherwood Casjens Most bacterial genomes have very few pseudogenes;
Opinion TRENDS in Microbiology Vol.9 No. November Where are the pseudogenes in bacterial genomes? Jeffrey G. Lawrence, Roger W. Hendrix and Sherwood Casjens Most bacterial genomes have very few pseudogenes;
L Aravind and Eugene V Koonin
 http://genomebiology.com/2000/1/4/research/0007.1 Research The a/b fold uracil DNA glycosylases: a common origin with diverse fates L Aravind and Eugene V Koonin Address: National Center for Biotechnology
http://genomebiology.com/2000/1/4/research/0007.1 Research The a/b fold uracil DNA glycosylases: a common origin with diverse fates L Aravind and Eugene V Koonin Address: National Center for Biotechnology
Deposited research article Short segmental duplication: parsimony in growth of microbial genomes Li-Ching Hsieh*, Liaofu Luo, and Hoong-Chien Lee
 This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article Short segmental duplication: parsimony in growth of microbial genomes
This information has not been peer-reviewed. Responsibility for the findings rests solely with the author(s). Deposited research article Short segmental duplication: parsimony in growth of microbial genomes
rho Is Not Essential for Viability or Virulence in Staphylococcus aureus
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 2001, p. 1099 1103 Vol. 45, No. 4 0066-4804/01/$04.00 0 DOI: 10.1128/AAC.45.4.1099 1103.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 2001, p. 1099 1103 Vol. 45, No. 4 0066-4804/01/$04.00 0 DOI: 10.1128/AAC.45.4.1099 1103.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved.
Mandelbrot s cascade in a Random Environment
 Mandelbrot s cascade in a Random Environment A joint work with Chunmao Huang (Ecole Polytechnique) and Xingang Liang (Beijing Business and Technology Univ) Université de Bretagne-Sud (Univ South Brittany)
Mandelbrot s cascade in a Random Environment A joint work with Chunmao Huang (Ecole Polytechnique) and Xingang Liang (Beijing Business and Technology Univ) Université de Bretagne-Sud (Univ South Brittany)
The use of gene clusters to infer functional coupling
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2896 2901, March 1999 Genetics The use of gene clusters to infer functional coupling ROSS OVERBEEK*, MICHAEL FONSTEIN, MARK D SOUZA*, GORDON D. PUSCH*, AND NATALIA
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2896 2901, March 1999 Genetics The use of gene clusters to infer functional coupling ROSS OVERBEEK*, MICHAEL FONSTEIN, MARK D SOUZA*, GORDON D. PUSCH*, AND NATALIA
MicroGenomics. Universal replication biases in bacteria
 Molecular Microbiology (1999) 32(1), 11±16 MicroGenomics Universal replication biases in bacteria Eduardo P. C. Rocha, 1,2 Antoine Danchin 2 and Alain Viari 1,3 * 1 Atelier de BioInformatique, UniversiteÂ
Molecular Microbiology (1999) 32(1), 11±16 MicroGenomics Universal replication biases in bacteria Eduardo P. C. Rocha, 1,2 Antoine Danchin 2 and Alain Viari 1,3 * 1 Atelier de BioInformatique, UniversiteÂ
Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana
 Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana Table S1 The species marked with belong to the Proteobacteria subset and those marked
Additional file 1 for Structural correlations in bacterial metabolic networks by S. Bernhardsson, P. Gerlee & L. Lizana Table S1 The species marked with belong to the Proteobacteria subset and those marked
Structural Proteomics of Eukaryotic Domain Families ER82 WR66
 Structural Proteomics of Eukaryotic Domain Families ER82 WR66 NIH Protein Structure Initiative Mission Statement To make the three-dimensional atomic level structures of most proteins readily available
Structural Proteomics of Eukaryotic Domain Families ER82 WR66 NIH Protein Structure Initiative Mission Statement To make the three-dimensional atomic level structures of most proteins readily available
Essentiality in B. subtilis
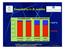 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria)
 Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria) (don t study Concept 27.2) Some phyla Remember: Bacterial cell structure and shapes 1 Usually very small but some are unusually
Ch 27: The Prokaryotes Bacteria & Archaea Older: (Eu)bacteria & Archae(bacteria) (don t study Concept 27.2) Some phyla Remember: Bacterial cell structure and shapes 1 Usually very small but some are unusually
Conserved hypothetical proteins: new hints and new puzzles
 Comparative and Functional Genomics Comp Funct Genom 2001; 2: 14 18. Conference Paper Conserved hypothetical proteins: new hints and new puzzles Michael Y. Galperin* National Center for Biotechnology Information,
Comparative and Functional Genomics Comp Funct Genom 2001; 2: 14 18. Conference Paper Conserved hypothetical proteins: new hints and new puzzles Michael Y. Galperin* National Center for Biotechnology Information,
Chapter 2 Class Notes Words and Probability
 Chapter 2 Class Notes Words and Probability Medical/Genetics Illustration reference Bojesen et al (2003), Integrin 3 Leu33Pro Homozygosity and Risk of Cancer, J. NCI. Women only 2 x 2 table: Stratification
Chapter 2 Class Notes Words and Probability Medical/Genetics Illustration reference Bojesen et al (2003), Integrin 3 Leu33Pro Homozygosity and Risk of Cancer, J. NCI. Women only 2 x 2 table: Stratification
HOBACGEN: Database System for Comparative Genomics in Bacteria
 Resource HOBACGEN: Database System for Comparative Genomics in Bacteria Guy Perrière, 1 Laurent Duret, and Manolo Gouy Laboratoire de Biométrie et Biologie Évolutive, Unité Mixte de Recherche Centre National
Resource HOBACGEN: Database System for Comparative Genomics in Bacteria Guy Perrière, 1 Laurent Duret, and Manolo Gouy Laboratoire de Biométrie et Biologie Évolutive, Unité Mixte de Recherche Centre National
A Specialized Version of the HD Hydrolase Domain Implicated in Signal Transduction
 J. Mol. Microbiol. Biotechnol. (1999) 1(2): 303-305. JMMB Correspondence The HD Hydrolase Domain in Signal Transduction 303 A Specialized Version of the HD Hydrolase Domain Implicated in Signal Transduction
J. Mol. Microbiol. Biotechnol. (1999) 1(2): 303-305. JMMB Correspondence The HD Hydrolase Domain in Signal Transduction 303 A Specialized Version of the HD Hydrolase Domain Implicated in Signal Transduction
Stabilization against Hyperthermal Denaturation through Increased CG Content Can Explain the Discrepancy between Whole Genome and 16S rrna Analyses
 11458 Biochemistry 2005, 44, 11458-11465 Stabilization against Hyperthermal Denaturation through Increased CG Content Can Explain the Discrepancy between Whole Genome and 16S rrna Analyses T. E. Meyer*,
11458 Biochemistry 2005, 44, 11458-11465 Stabilization against Hyperthermal Denaturation through Increased CG Content Can Explain the Discrepancy between Whole Genome and 16S rrna Analyses T. E. Meyer*,
Increasing biological complexity is positively correlated with the relative genome-wide expansion of non-protein-coding DNA sequences
 Increasing biological complexity is positively correlated with the relative genome-wide expansion of non-protein-coding DNA sequences Ryan J. Taft 1, 2 * and John S. Mattick 3 1 Rowe Program in Genetics,
Increasing biological complexity is positively correlated with the relative genome-wide expansion of non-protein-coding DNA sequences Ryan J. Taft 1, 2 * and John S. Mattick 3 1 Rowe Program in Genetics,
Wavelets and Fractals
 Wavelets and Fractals Bikramjit Singh Walia Samir Kagadkar Shreyash Gupta 1 Self-similar sets The best way to study any physical problem with known symmetry is to build a functional basis with symmetry
Wavelets and Fractals Bikramjit Singh Walia Samir Kagadkar Shreyash Gupta 1 Self-similar sets The best way to study any physical problem with known symmetry is to build a functional basis with symmetry
PRED-CLASS: cascading neural networks for generalized protein classification and genome-wide applications
 PRED-CLASS: cascading neural networks for generalized protein classification and genome-wide applications Claude Pasquier, Vasilis Promponas, Stavros Hamodrakas To cite this version: Claude Pasquier, Vasilis
PRED-CLASS: cascading neural networks for generalized protein classification and genome-wide applications Claude Pasquier, Vasilis Promponas, Stavros Hamodrakas To cite this version: Claude Pasquier, Vasilis
Microbial Taxonomy. Classification of living organisms into groups. A group or level of classification
 Lec 2 Oral Microbiology Dr. Chatin Purpose Microbial Taxonomy Classification Systems provide an easy way grouping of diverse and huge numbers of microbes To provide an overview of how physicians think
Lec 2 Oral Microbiology Dr. Chatin Purpose Microbial Taxonomy Classification Systems provide an easy way grouping of diverse and huge numbers of microbes To provide an overview of how physicians think
Applications of algorithmic complexity to network science and synthetic biology
 Applications of algorithmic complexity to network science and synthetic biology Hector Zenil hector.zenil@ki.se Unit of Computational Medicine, KI Conference on Computability, Complexity and Randomness
Applications of algorithmic complexity to network science and synthetic biology Hector Zenil hector.zenil@ki.se Unit of Computational Medicine, KI Conference on Computability, Complexity and Randomness
Origin of Eukaryotic Cell Nuclei by Symbiosis of Archaea in Bacteria supported by the newly clarified origin of functional genes
 Genes Genet. Syst. (2002) 77, p. 369 376 Origin of Eukaryotic Cell Nuclei by Symbiosis of Archaea in Bacteria supported by the newly clarified origin of functional genes Tokumasa Horiike, Kazuo Hamada,
Genes Genet. Syst. (2002) 77, p. 369 376 Origin of Eukaryotic Cell Nuclei by Symbiosis of Archaea in Bacteria supported by the newly clarified origin of functional genes Tokumasa Horiike, Kazuo Hamada,
DNA sequence analysis using Markov chain models
 DNA sequence analysis using Markov chain models Boris Ryabko Siberian University of Telecommunication an Informatics, Institute of Computational Technologies, Siberian Branch of RAS Novosibirsk E-mail:
DNA sequence analysis using Markov chain models Boris Ryabko Siberian University of Telecommunication an Informatics, Institute of Computational Technologies, Siberian Branch of RAS Novosibirsk E-mail:
A thermophilic last universal ancestor inferred from its estimated amino acid composition
 CHAPTER 17 A thermophilic last universal ancestor inferred from its estimated amino acid composition Dawn J. Brooks and Eric A. Gaucher 17.1 Introduction The last universal ancestor (LUA) represents a
CHAPTER 17 A thermophilic last universal ancestor inferred from its estimated amino acid composition Dawn J. Brooks and Eric A. Gaucher 17.1 Introduction The last universal ancestor (LUA) represents a
Fractal Strings and Multifractal Zeta Functions
 Fractal Strings and Multifractal Zeta Functions Michel L. Lapidus, Jacques Lévy-Véhel and John A. Rock August 4, 2006 Abstract. We define a one-parameter family of geometric zeta functions for a Borel
Fractal Strings and Multifractal Zeta Functions Michel L. Lapidus, Jacques Lévy-Véhel and John A. Rock August 4, 2006 Abstract. We define a one-parameter family of geometric zeta functions for a Borel
Biased biological functions of horizontally transferred genes in prokaryotic genomes
 Biased biological functions of horizontally transferred genes in prokaryotic genomes Yoji Nakamura 1,5, Takeshi Itoh 2,3, Hideo Matsuda 4 & Takashi Gojobori 1,2 Horizontal gene transfer is one of the main
Biased biological functions of horizontally transferred genes in prokaryotic genomes Yoji Nakamura 1,5, Takeshi Itoh 2,3, Hideo Matsuda 4 & Takashi Gojobori 1,2 Horizontal gene transfer is one of the main
University of Groningen. Genomics in Bacillus subtilis Noback, Michiel Andries
 University of Groningen Genomics in Bacillus subtilis Noback, Michiel Andries IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check
University of Groningen Genomics in Bacillus subtilis Noback, Michiel Andries IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check
Relationships of exponents in multifractal detrended fluctuation analysis and conventional multifractal analysis
 Relationships of exponents in multifractal detrended fluctuation analysis and conventional multifractal analysis Zhou Yu( ) a), Leung Yee( ) a)b)c), and Yu Zu-Guo( ) d)e) a) Department of Geography and
Relationships of exponents in multifractal detrended fluctuation analysis and conventional multifractal analysis Zhou Yu( ) a), Leung Yee( ) a)b)c), and Yu Zu-Guo( ) d)e) a) Department of Geography and
Combining aperiodic order with structural disorder: branching cellular automata
 Combining aperiodic order with structural disorder: branching cellular automata Michel Dekking Lorentz Center Workshop 30-40 minutes lecture May 31, 2016 Branching cellular automata Branching cellular
Combining aperiodic order with structural disorder: branching cellular automata Michel Dekking Lorentz Center Workshop 30-40 minutes lecture May 31, 2016 Branching cellular automata Branching cellular
THE advances of the last decade on genome sequenc- very different compositional bias, as well as different gene
 Copyright 2003 by the Genetics Society of America Associations Between Inverted Repeats and the Structural Evolution of Bacterial Genomes Guillaume Achaz,*,1 Eric Coissac,*,2 Pierre Netter* and Eduardo
Copyright 2003 by the Genetics Society of America Associations Between Inverted Repeats and the Structural Evolution of Bacterial Genomes Guillaume Achaz,*,1 Eric Coissac,*,2 Pierre Netter* and Eduardo
Chaos game representation of functional protein sequences, and simulation and multifractal analysis of induced measures
 Chaos game representation of functional protein sequences, and simulation and multifractal analysis of induced measures Yu Zu-Guo( 喻祖国 ab, Xiao Qian-Jun( 肖前军 a, Shi Long( 石龙 a, Yu Jun-Wu( 余君武 c, and Vo
Chaos game representation of functional protein sequences, and simulation and multifractal analysis of induced measures Yu Zu-Guo( 喻祖国 ab, Xiao Qian-Jun( 肖前军 a, Shi Long( 石龙 a, Yu Jun-Wu( 余君武 c, and Vo
Intermittency, Fractals, and β-model
 Intermittency, Fractals, and β-model Lecture by Prof. P. H. Diamond, note by Rongjie Hong I. INTRODUCTION An essential assumption of Kolmogorov 1941 theory is that eddies of any generation are space filling
Intermittency, Fractals, and β-model Lecture by Prof. P. H. Diamond, note by Rongjie Hong I. INTRODUCTION An essential assumption of Kolmogorov 1941 theory is that eddies of any generation are space filling
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett. Introduction
 The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
A. Distribution of LCS and their putative functional meaning
 showing the results of two methods at work, namely, the NSEI and a neural network trained by means of LCS lengths to distinguish between coding and non-coding sequences. Finally Section V is devoted to
showing the results of two methods at work, namely, the NSEI and a neural network trained by means of LCS lengths to distinguish between coding and non-coding sequences. Finally Section V is devoted to
Gene Conversion Drives Within Genic Sequences: Concerted Evolution of Ribosomal RNA Genes in Bacteria and Archaea
 J Mol Evol (2000) 51:305 317 DOI: 10.1007/s002390010093 Springer-Verlag New York Inc. 2000 Gene Conversion Drives Within Genic Sequences: Concerted Evolution of Ribosomal RNA Genes in Bacteria and Archaea
J Mol Evol (2000) 51:305 317 DOI: 10.1007/s002390010093 Springer-Verlag New York Inc. 2000 Gene Conversion Drives Within Genic Sequences: Concerted Evolution of Ribosomal RNA Genes in Bacteria and Archaea
Peter Molnar 1 and Paolo Burlando Institute of Environmental Engineering, ETH Zurich, Switzerland
 Hydrology Days 6 Seasonal and regional variability in scaling properties and correlation structure of high resolution precipitation data in a highly heterogeneous mountain environment (Switzerland) Peter
Hydrology Days 6 Seasonal and regional variability in scaling properties and correlation structure of high resolution precipitation data in a highly heterogeneous mountain environment (Switzerland) Peter
Prokaryotic phylogenies inferred from protein structural domains
 Letter Prokaryotic phylogenies inferred from protein structural domains Eric J. Deeds, 1 Hooman Hennessey, 2 and Eugene I. Shakhnovich 3,4 1 Department of Molecular and Cellular Biology, Harvard University,
Letter Prokaryotic phylogenies inferred from protein structural domains Eric J. Deeds, 1 Hooman Hennessey, 2 and Eugene I. Shakhnovich 3,4 1 Department of Molecular and Cellular Biology, Harvard University,
6.096 Algorithms for Computational Biology. Prof. Manolis Kellis
 6.096 Algorithms for Computational Biology Prof. Manolis Kellis Today s Goals Introduction Class introduction Challenges in Computational Biology Gene Regulation: Regulatory Motif Discovery Exhaustive
6.096 Algorithms for Computational Biology Prof. Manolis Kellis Today s Goals Introduction Class introduction Challenges in Computational Biology Gene Regulation: Regulatory Motif Discovery Exhaustive
Advanced Machine Learning & Perception
 Advanced Machine Learning & Perception Instructor: Tony Jebara Topic 6 Standard Kernels Unusual Input Spaces for Kernels String Kernels Probabilistic Kernels Fisher Kernels Probability Product Kernels
Advanced Machine Learning & Perception Instructor: Tony Jebara Topic 6 Standard Kernels Unusual Input Spaces for Kernels String Kernels Probabilistic Kernels Fisher Kernels Probability Product Kernels
Computational Structural Bioinformatics
 Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
MICROBIAL BIOCHEMISTRY BIOT 309. Dr. Leslye Johnson Sept. 30, 2012
 MICROBIAL BIOCHEMISTRY BIOT 309 Dr. Leslye Johnson Sept. 30, 2012 Phylogeny study of evoluhonary relatedness among groups of organisms (e.g. species, populahons), which is discovered through molecular
MICROBIAL BIOCHEMISTRY BIOT 309 Dr. Leslye Johnson Sept. 30, 2012 Phylogeny study of evoluhonary relatedness among groups of organisms (e.g. species, populahons), which is discovered through molecular
Power Law Distributions of Gene Family Sizes. Elizabeth Rach
 CORNELL UNIVERSITY MATHEMATICS DEPARTMENT SENIOR THESIS Power Law Distributions of Gene Family Sizes A THESIS PRESENTED IN PARTIAL FULFILLMENT OF CRITERIA FOR HONORS IN MATHEMATICS Elizabeth Rach May 2004
CORNELL UNIVERSITY MATHEMATICS DEPARTMENT SENIOR THESIS Power Law Distributions of Gene Family Sizes A THESIS PRESENTED IN PARTIAL FULFILLMENT OF CRITERIA FOR HONORS IN MATHEMATICS Elizabeth Rach May 2004
Multifractal analysis of Bernoulli convolutions associated with Salem numbers
 Multifractal analysis of Bernoulli convolutions associated with Salem numbers De-Jun Feng The Chinese University of Hong Kong Fractals and Related Fields II, Porquerolles - France, June 13th-17th 2011
Multifractal analysis of Bernoulli convolutions associated with Salem numbers De-Jun Feng The Chinese University of Hong Kong Fractals and Related Fields II, Porquerolles - France, June 13th-17th 2011
BATMAS30: Amino Acid Substitution Matrix for Alignment of Bacterial Transporters
 PROTEINS: Structure, Function, and Genetics 51:85 95 (2003) BATMAS30: Amino Acid Substitution Matrix for Alignment of Bacterial Transporters Roman A. Sutormin, 1 * Aleksandra B. Rakhmaninova, 2 and Mikhail
PROTEINS: Structure, Function, and Genetics 51:85 95 (2003) BATMAS30: Amino Acid Substitution Matrix for Alignment of Bacterial Transporters Roman A. Sutormin, 1 * Aleksandra B. Rakhmaninova, 2 and Mikhail
BIOINFORMATICS. Sequence complexity profiles of prokaryotic genomic sequences: A fast algorithm for calculating linguistic complexity
 BIOINFORMATICS Vol. 18 no. 5 2002 Pages 679 688 Sequence complexity profiles of prokaryotic genomic sequences: A fast algorithm for calculating linguistic complexity Olga G. Troyanskaya 1,4, Ora Arbell
BIOINFORMATICS Vol. 18 no. 5 2002 Pages 679 688 Sequence complexity profiles of prokaryotic genomic sequences: A fast algorithm for calculating linguistic complexity Olga G. Troyanskaya 1,4, Ora Arbell
Advanced Algorithms and Models for Computational Biology
 Advanced Algorithms and Models for Computational Biology Introduction to cell biology genomics development and probability Eric ing Lecture January 3 006 Reading: Chap. DTM book Introduction to cell biology
Advanced Algorithms and Models for Computational Biology Introduction to cell biology genomics development and probability Eric ing Lecture January 3 006 Reading: Chap. DTM book Introduction to cell biology
arxiv:cond-mat/ v1 [cond-mat.dis-nn] 3 Apr 2002
![arxiv:cond-mat/ v1 [cond-mat.dis-nn] 3 Apr 2002 arxiv:cond-mat/ v1 [cond-mat.dis-nn] 3 Apr 2002](/thumbs/95/123949582.jpg) arxiv:cond-mat/0204076v1 [cond-mat.dis-nn] 3 Apr 2002 In: M. Suzuki and N. Kawashima (eds.) Coherent Approaches to Fluctuations (Proc. Hayashibara Forum 95), pp. 59-64, World Scientific (singapore, 1995).
arxiv:cond-mat/0204076v1 [cond-mat.dis-nn] 3 Apr 2002 In: M. Suzuki and N. Kawashima (eds.) Coherent Approaches to Fluctuations (Proc. Hayashibara Forum 95), pp. 59-64, World Scientific (singapore, 1995).
Membrane structure prediction 1
 title: Computational Biology 1 - Protein structure: Membrane structure prediction 1 short title: cb1_tmh1 lecture: Protein Prediction 1 - Protein structure Computational Biology 1 - TUM Summer 2015 1/00
title: Computational Biology 1 - Protein structure: Membrane structure prediction 1 short title: cb1_tmh1 lecture: Protein Prediction 1 - Protein structure Computational Biology 1 - TUM Summer 2015 1/00
INTERPRETATION OF THE GRAM STAIN
 INTERPRETATION OF THE GRAM STAIN DISCLOSURE Relevant relationships with commercial entities none Potential for conflicts of interest within this presentation none Steps taken to review and mitigate potential
INTERPRETATION OF THE GRAM STAIN DISCLOSURE Relevant relationships with commercial entities none Potential for conflicts of interest within this presentation none Steps taken to review and mitigate potential
Exponential Distribution and Poisson Process
 Exponential Distribution and Poisson Process Stochastic Processes - Lecture Notes Fatih Cavdur to accompany Introduction to Probability Models by Sheldon M. Ross Fall 215 Outline Introduction Exponential
Exponential Distribution and Poisson Process Stochastic Processes - Lecture Notes Fatih Cavdur to accompany Introduction to Probability Models by Sheldon M. Ross Fall 215 Outline Introduction Exponential
1. Prokaryotic Nutritional & Metabolic Adaptations
 Chapter 27B: Bacteria and Archaea 1. Prokaryotic Nutritional & Metabolic Adaptations 2. Survey of Prokaryotic Groups A. Domain Bacteria Gram-negative groups B. Domain Bacteria Gram-positive groups C. Domain
Chapter 27B: Bacteria and Archaea 1. Prokaryotic Nutritional & Metabolic Adaptations 2. Survey of Prokaryotic Groups A. Domain Bacteria Gram-negative groups B. Domain Bacteria Gram-positive groups C. Domain
Two Families of Mechanosensitive Channel Proteins
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Mar. 2003, p. 66 85 Vol. 67, No. 1 1092-2172/03/$08.00 0 DOI: 10.1128/MMBR.67.1.66 85.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Mar. 2003, p. 66 85 Vol. 67, No. 1 1092-2172/03/$08.00 0 DOI: 10.1128/MMBR.67.1.66 85.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
Comparing function and structure between entire proteomes
 Comparing function and structure between entire proteomes JINFENG LIU 1,2 AND BURKHARD ROST 1 1 CUBIC, Department of Biochemistry and Molecular Biophysics, Columbia University, New York, New York 10032,
Comparing function and structure between entire proteomes JINFENG LIU 1,2 AND BURKHARD ROST 1 1 CUBIC, Department of Biochemistry and Molecular Biophysics, Columbia University, New York, New York 10032,
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools. Evaluation. Course Homepage.
 CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
AUTOREGRESSIVE MODELING OF CODING SEQUENCE LENGTHS IN BACTERIAL GENOME
 Fluctuation and oise Letters Vol. 0, o. 0 (0000) 000 000 World Scientific Publishing Company AUTOREGRESSIVE MODELIG OF CODIG SEQUECE LEGTHS I BACTERIAL GEOME VASILE V. MORARIU 1,, LUIZA BUIMAGA-IARICA
Fluctuation and oise Letters Vol. 0, o. 0 (0000) 000 000 World Scientific Publishing Company AUTOREGRESSIVE MODELIG OF CODIG SEQUECE LEGTHS I BACTERIAL GEOME VASILE V. MORARIU 1,, LUIZA BUIMAGA-IARICA
Modeling Noise in Genetic Sequences
 Modeling Noise in Genetic Sequences M. Radavičius 1 and T. Rekašius 2 1 Institute of Mathematics and Informatics, Vilnius, Lithuania 2 Vilnius Gediminas Technical University, Vilnius, Lithuania 1. Introduction:
Modeling Noise in Genetic Sequences M. Radavičius 1 and T. Rekašius 2 1 Institute of Mathematics and Informatics, Vilnius, Lithuania 2 Vilnius Gediminas Technical University, Vilnius, Lithuania 1. Introduction:
New families of non-travelling wave solutions to a new (3+1)-dimensional potential-ytsf equation
 MM Research Preprints, 376 381 MMRC, AMSS, Academia Sinica, Beijing No., December 3 New families of non-travelling wave solutions to a new (3+1-dimensional potential-ytsf equation Zhenya Yan Key Laboratory
MM Research Preprints, 376 381 MMRC, AMSS, Academia Sinica, Beijing No., December 3 New families of non-travelling wave solutions to a new (3+1-dimensional potential-ytsf equation Zhenya Yan Key Laboratory
1. Let X and Y be independent exponential random variables with rate α. Find the densities of the random variables X 3, X Y, min(x, Y 3 )
 1 Introduction These problems are meant to be practice problems for you to see if you have understood the material reasonably well. They are neither exhaustive (e.g. Diffusions, continuous time branching
1 Introduction These problems are meant to be practice problems for you to see if you have understood the material reasonably well. They are neither exhaustive (e.g. Diffusions, continuous time branching
Introns. Illusions as neuro-signs Nicholas Wade
 R364 Current Biology, Vol 8 No 11 Introns Illusions as neuro-signs Nicholas Wade Almost the whole of visual science is concerned with illusions if they are defined as images in which the physical description
R364 Current Biology, Vol 8 No 11 Introns Illusions as neuro-signs Nicholas Wade Almost the whole of visual science is concerned with illusions if they are defined as images in which the physical description
A multifractal random walk
 A multifractal random walk E. Bacry Centre de Mathématiques Appliquées, Ecole Polytechnique, 91128 Palaiseau Cedex, France J. Delour and J.F. Muzy Centre de Recherche Paul Pascal, Avenue Schweitzer 33600
A multifractal random walk E. Bacry Centre de Mathématiques Appliquées, Ecole Polytechnique, 91128 Palaiseau Cedex, France J. Delour and J.F. Muzy Centre de Recherche Paul Pascal, Avenue Schweitzer 33600
A Tapestry of Complex Dimensions
 A Tapestry of Complex Dimensions John A. Rock May 7th, 2009 1 f (α) dim M(supp( β)) (a) (b) α Figure 1: (a) Construction of the binomial measure β. spectrum f(α) of the measure β. (b) The multifractal
A Tapestry of Complex Dimensions John A. Rock May 7th, 2009 1 f (α) dim M(supp( β)) (a) (b) α Figure 1: (a) Construction of the binomial measure β. spectrum f(α) of the measure β. (b) The multifractal
Keywords: gene order, operon, phylogenetic analysis, functional signal, local alignment, dot-plot, conserved gene pair
 Igor B. Rogozin is a staff scientist at the National Center for Biotechnology Information NLM/NIH (Bethesda, MD, USA) and a research scientist at the Institute of Cytology and Genetics RAS (Novosibirsk,
Igor B. Rogozin is a staff scientist at the National Center for Biotechnology Information NLM/NIH (Bethesda, MD, USA) and a research scientist at the Institute of Cytology and Genetics RAS (Novosibirsk,
Global dinucleotide signatures and analysis of genomic heterogeneity Samuel Karlin
 598 Global dinucleotide signatures and analysis of genomic heterogeneity Samuel Karlin Early biochemical experiments measuring nearest neighbor frequencies established that the set of dinucleotide relative
598 Global dinucleotide signatures and analysis of genomic heterogeneity Samuel Karlin Early biochemical experiments measuring nearest neighbor frequencies established that the set of dinucleotide relative
A Structural Equation Model Study of Shannon Entropy Effect on CG content of Thermophilic 16S rrna and Bacterial Radiation Repair Rec-A Gene Sequences
 A Structural Equation Model Study of Shannon Entropy Effect on CG content of Thermophilic 16S rrna and Bacterial Radiation Repair Rec-A Gene Sequences T. Holden, P. Schneider, E. Cheung, J. Prayor, R.
A Structural Equation Model Study of Shannon Entropy Effect on CG content of Thermophilic 16S rrna and Bacterial Radiation Repair Rec-A Gene Sequences T. Holden, P. Schneider, E. Cheung, J. Prayor, R.
Chaos game representation (CGR)-walk model for DNA sequences
 Vol 18 No 1, January 2009 c 2009 Chin. Phys. Soc. 1674-1056/2009/18(01)/0370-07 Chinese Physics B and IOP Publishing Ltd Chaos game representation (CGR)-walk model for DNA sequences Gao Jie( ) a)b) and
Vol 18 No 1, January 2009 c 2009 Chin. Phys. Soc. 1674-1056/2009/18(01)/0370-07 Chinese Physics B and IOP Publishing Ltd Chaos game representation (CGR)-walk model for DNA sequences Gao Jie( ) a)b) and
N o hal June 2007
 LABORATOIRE D INFORMATIQUE DE NANTES-ATLANTIQUE A large-scale analysis for significance assessment of frequencies relative to potentially strong sigma 70 promoters: comparison of 32 prokaryotic Christine
LABORATOIRE D INFORMATIQUE DE NANTES-ATLANTIQUE A large-scale analysis for significance assessment of frequencies relative to potentially strong sigma 70 promoters: comparison of 32 prokaryotic Christine
The impact of the neisserial DNA uptake sequence on genome evolution and stability
 The impact of the neisserial DNA uptake sequence on genome evolution and stability Ole Herman Ambur The Tønjum Group Transformation Neisserial transformation requires: DNA uptake sequence (DUS) Transformation
The impact of the neisserial DNA uptake sequence on genome evolution and stability Ole Herman Ambur The Tønjum Group Transformation Neisserial transformation requires: DNA uptake sequence (DUS) Transformation
Amino acid composition of genomes, lifestyles of organisms, and evolutionary trends: a global picture with correspondence analysis
 Gene 297 (2002) 51 60 www.elsevier.com/locate/gene Amino acid composition of genomes, lifestyles of organisms, and evolutionary trends: a global picture with correspondence analysis Fredj Tekaia a, *,
Gene 297 (2002) 51 60 www.elsevier.com/locate/gene Amino acid composition of genomes, lifestyles of organisms, and evolutionary trends: a global picture with correspondence analysis Fredj Tekaia a, *,
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Tree of Life: An Introduction to Microbial Phylogeny Beverly Brown, Sam Fan, LeLeng To Isaacs, and Min-Ken Liao
 Microbes Count! 191 Tree of Life: An Introduction to Microbial Phylogeny Beverly Brown, Sam Fan, LeLeng To Isaacs, and Min-Ken Liao Video VI: Microbial Evolution Introduction Bioinformatics tools allow
Microbes Count! 191 Tree of Life: An Introduction to Microbial Phylogeny Beverly Brown, Sam Fan, LeLeng To Isaacs, and Min-Ken Liao Video VI: Microbial Evolution Introduction Bioinformatics tools allow
A Martingale Central Limit Theorem
 A Martingale Central Limit Theorem Sunder Sethuraman We present a proof of a martingale central limit theorem (Theorem 2) due to McLeish (974). Then, an application to Markov chains is given. Lemma. For
A Martingale Central Limit Theorem Sunder Sethuraman We present a proof of a martingale central limit theorem (Theorem 2) due to McLeish (974). Then, an application to Markov chains is given. Lemma. For
Remarkable sequence signatures in archaeal genomes
 Archaea 1, 185 190 2003 Heron Publishing Victoria, Canada Remarkable sequence signatures in archaeal genomes AHMED FADIEL, 1,2 STUART LITHWICK, 3 GOPI GANJI 3 and STEPHEN W. SCHERER 1 1 The Center for
Archaea 1, 185 190 2003 Heron Publishing Victoria, Canada Remarkable sequence signatures in archaeal genomes AHMED FADIEL, 1,2 STUART LITHWICK, 3 GOPI GANJI 3 and STEPHEN W. SCHERER 1 1 The Center for
Fractals and Fractal Dimensions
 Fractals and Fractal Dimensions John A. Rock July 13th, 2009 begin with [0,1] remove 1 of length 1/3 remove 2 of length 1/9 remove 4 of length 1/27 remove 8 of length 1/81...................................
Fractals and Fractal Dimensions John A. Rock July 13th, 2009 begin with [0,1] remove 1 of length 1/3 remove 2 of length 1/9 remove 4 of length 1/27 remove 8 of length 1/81...................................
FE610 Stochastic Calculus for Financial Engineers. Stevens Institute of Technology
 FE610 Stochastic Calculus for Financial Engineers Lecture 3. Calculaus in Deterministic and Stochastic Environments Steve Yang Stevens Institute of Technology 01/31/2012 Outline 1 Modeling Random Behavior
FE610 Stochastic Calculus for Financial Engineers Lecture 3. Calculaus in Deterministic and Stochastic Environments Steve Yang Stevens Institute of Technology 01/31/2012 Outline 1 Modeling Random Behavior
Bayesian Inference and the Symbolic Dynamics of Deterministic Chaos. Christopher C. Strelioff 1,2 Dr. James P. Crutchfield 2
 How Random Bayesian Inference and the Symbolic Dynamics of Deterministic Chaos Christopher C. Strelioff 1,2 Dr. James P. Crutchfield 2 1 Center for Complex Systems Research and Department of Physics University
How Random Bayesian Inference and the Symbolic Dynamics of Deterministic Chaos Christopher C. Strelioff 1,2 Dr. James P. Crutchfield 2 1 Center for Complex Systems Research and Department of Physics University
Simultaneous Accumulation Points to Sets of d-tuples
 ISSN 1749-3889 print, 1749-3897 online International Journal of Nonlinear Science Vol.92010 No.2,pp.224-228 Simultaneous Accumulation Points to Sets of d-tuples Zhaoxin Yin, Meifeng Dai Nonlinear Scientific
ISSN 1749-3889 print, 1749-3897 online International Journal of Nonlinear Science Vol.92010 No.2,pp.224-228 Simultaneous Accumulation Points to Sets of d-tuples Zhaoxin Yin, Meifeng Dai Nonlinear Scientific
Membrane structure prediction 1
 title: Membrane structure prediction 1 short title: cb1_tmh1 lecture: Computational Biology 1 - Protein structure (for Informatics) - TUM summer semester 1/77 Announcements Videos: THANKS : Michael Leunig
title: Membrane structure prediction 1 short title: cb1_tmh1 lecture: Computational Biology 1 - Protein structure (for Informatics) - TUM summer semester 1/77 Announcements Videos: THANKS : Michael Leunig
