Supplemental Table 1: Strains used in this study
|
|
|
- Regina Tiffany Lewis
- 5 years ago
- Views:
Transcription
1 Supplemental Table : Strains used in this study Name Parent Genotype Reference GA-8 MATa, ade-; can-; his3-,5; trp-; ura3-; leu-3, W33-A GA-3 GA-8 his3-,5::gfp-laci-his3, NUP49::GFP-NUP49-URA3 [] GA-46 GA-3 + ARS67::ARS67 - laco-trp [] GA-95 GA-46 esc::kanmx This study GA-84 GA-46 ptlc-δ48int [as in 3] This study GA-369 GA-3 ARS54::ARS54 - laco-lexa-trp [4] GA-98 MATa, ade-; can-; his3-,5; trp-; ura3-; leu-3, W33-A GA-49 GA-98 nup33::his3 This study GA-438 GA-49 (pun-nup33δn - LEU) This study GA-458 GA-438 pun-nup33δn - leu::kanmx (marker swap) This study GA-45 GA-458 NUP49::CFP-NUP49 - URA3 This study GA-4583 GA-45 +ade-::laci-gfp::ade This study GA-4584 GA-4583 lys::laco.lexa-trp This study GA-46 W33-A +telomere 5R::ADE This study GA-37 GA-46 +tel::his5 This study GA-9 GA-46 +yku7::kanmx This study GA-575 GA-46 MPS3::mps3Δ75-5-KanMx6 [as in 5] This study GA-4954 GA-369 +sir4::hphmx This study GA-3467 GA-5559 W33-A/α GA-46 GA-3467 MATa/MATα, ade-; can-; his3-,5; trp-; ura3-; leu-3, EST/est::HphMX Diploid; see parental phenotypes This study This study Strain references. Heun, P., Laroche, T., Raghuraman, M.K., and Gasser, S.M. (). The positioning and dynamics of origins of replication in the budding yeast nucleus. J Cell Biol 5, Hediger, F., Neumann, F.R., Van Houwe, G., Dubrana, K., and Gasser, S.M. (). Live imaging of telomeres: yku and Sir proteins define redundant telomere-anchoring pathways in yeast. Curr Biol, Stellwagen, A.E., Haimberger, Z.W., Veatch, J.R., and Gottschling, D.E. (3). Ku interacts with telomerase RNA to promote telomere addition at native and broken chromosome ends. Genes Dev 7, Schober, H., Kalck, V., Vega-Palas, M.A., Van Houwe, G., Sage, D., Unser, M., Gartenberg, M.R., and Gasser, S.M. (7). Controlled exchange of chromosomal arms reveals principles driving telomere interactions in yeast. Genome Res.
2 5. Bupp, J.M., Martin, A.E., Stensrud, E.S., and Jaspersen, S.L. (7). Telomere anchoring at the nuclear periphery requires the budding yeast Sad-UNC-84 domain protein Mps3. J Cell Biol. Supplemental Material Simulation and computational modeling The theoretically expected random colocalization of a spot with the pore cluster was calculated geometrically. If the center of the spot is in a certain region of the nucleus the spot is considered as colocalizing with the cluster. If the spot is distributed uniformly inside the nucleus then the is the ratio C/V where C is the volume of the region mentioned above and V is the total volume that is available to the spot. The pore cluster was modeled as a conical layer inside the nucleus (see Supplemental Figure ) and the spot was considered as colocalizing if it at least touches the cluster. The total available volume is the volume of the nucleus minus the volume of the nucleolus (estimated as 3%). Furthermore, the is affected by the position of the spot relative to the nuclear periphery. We calculated this effect by dividing the nucleus into the outermost shell (corresponding to zone ) and the interior (zone and 3). If of N spots εn are in zone then the ς can be calculated as ς = ες + ( - ε)ς 3 where ς = C /V and ς 3 = C 3 /V 3 are the volume fractions for colocalization in zone and zone and 3, respectively. See Supplemental Figure for details on the calculation of ς. Supplemental Figure (A) The pore cluster is modeled as a conical disk at the periphery of the nucleus. The thickness d, the diameter l, and the radius of the spot r where measured in 3D reconstructions of microscopic images. A possible outline of the nucleus is shown for illustration. (B) The for a random spot was calculated analytically as a fraction of volumes (see Supplemental methods). The figure shows a cut through the model. The pore cluster is a conical layer shown in red. The spot is considered as colocalizing if it at least touches the pore cluster which results in the colocalization region shown in green. For the calculation this region is approximated again by a conical
3 layer outlined in dark green (see blow up in the left part of the figure). In the parameter range used here this causes an error in the volume calculation of less than 5%. The nucleolus is shown in grey and is calculated as roughly 3% of the nuclear volume. It is assumed not to overlap with the pore cluster and under this condition its exact position is irrelevant to the calculation. Zone (see text) is shown in light blue. We use the following identifiers, all relative lengths are normalized to R, all relative volumes are normalized to V tot = 4/3πR 3 : R: nuclear radius d: thickness of the colocalization volume ρ d : (R - d) / R ρ = (/3) /3 : relative inner radius of zone θ: angle determining the size of the colocalization volume η: relative height of the nucleolus κ = /4 η (3 - η): relative volume of the nucleolus ε: fraction of spots in zone Note that d is the thickness of the colocalization volume which is not identical to the thickness d of the cluster: d = d + r where r is the radius of the spot. Likewise we have to calculate θ from l and r: θ = θ + sin - (r/r) where θ = sin - (l /R). Without enrichment in zone (i.e. ε = /3) the ς can be calculated as ς = ( ρ d )( cosθ ). κ With enrichment in zone this has to be replaced by ς = ε 3 κ ( ρ )( cosθ ) ( η + ρ ) ( ρ η + ) + ( ε ) 3 ρ ( ρ ρ d )( cosθ ) ( η + ρ ) ( ρ η + ) Two case differentiations are necessary during the derivation of the above formula. In the given form it assumes that both the nucleolus and the colocalization volume extend into zone, i.e. η > ρ and ρ > ρ d, respectively. (C) Expected colocalization versus cluster thickness for three different cluster diameters. The curves do not start at a colocalization value of zero because even for a very thin cluster the colocalization volume is finite. When the cluster thickness approaches the nuclear radius the colocalization volume increases very slowly leading to a very flat.
4 curve. The dashed lines show the measured cluster thickness and the smallest measured colocalization. (D) Expected colocalization versus cluster diameter for three different cluster thicknesses. The vertical dashed line shows the measured cluster diameter. (E) Expected colocalization versus volume fraction occupied by the nucleolus. The increases when the total accessible volume decreases. This can be the case if a certain volume fraction is occupied e.g. by the nucleolus. The vertical dashed line indicates a volume fraction of one third. Cluster thickness and diameter have their default values of 34nm and 8nm, respectively. (F) Expected colocalization versus enrichment in zone. If the spot is more likely to be at the periphery this also increases the expected unspecific colocalization. The vertical dashed lines indicate the uniform distribution (fraction /3) and a typical enriched value of.6. Supplemental Figure : Mlp and Mlp are not required for Yku8 anchoring Targeted anchoring of internally localized ARS67 bearing laco and lexa binding sites was scored in strains GA-765 (W33-A, mlpδ mlpδ, GFP-NUP49, laci-gfp, lacolexa sites). GA lexa alone ((G+S) n=), GA lexa-yku8 ((G+S) n=93), and GA lexa-yku8-4 ((G+S) n=3). * indicates significantly different than random. G and S phase results were not significantly different and were therefore combined. Supplemental Figure 3: Tel6R anchoring is lost in sir4 and est mutants The position of Tel6R tagged with laco sites was scored relative to three zones as described in Figure, in the following strains: GA-459 (Tel6R ) (G n= 87, S phase n=6); GA-867 (Tel6R sir4δ) (G phase n=6, S phase n=57) and GA-867-estΔ (Tel6R estδsir4δ) (G n=5, S phase n=36). * indicates significantly different than random and P values are given. Supplemental Figure 4: tel cells lose sliencing efficiency Telomeric position effect or silencing was monitored in cells bearing the ADE marker at Tel5R in vs telδ cells. GA-46 (W33, ) is restreaked next to isogenic telδ strain GA-37. The loss of pink color indicates loss of repression.
5 A B Average width d = 34nm, n=5 ρ d R pore cluster θ spot Average length l = 8nm, n=63 Spot diameter r = 5nm, n=9 ρ R zone nucleolus ηr C D cluster diameter 6nm cluster diameter 8nm cluster diameter nm cluster thickness nm cluster thickness 34nm cluster thickness 48nm cluster thickness [nm] cluster diameter [nm] E F volume fraction occupied by the nucleolus fraction of spots in zone Schober et al Supplemetal Figure
6 ARS67 lexa in mlp/δ lexa-yku8 in mlp/δ lexa-yku8-4 in mlp/δ random * random % foci Zone G and S phase * * 3 Schober et al. Supplemental Figure Tel6R 8 P =.48 G phase 8 P =. S phase laci-gfp sir4δ sir4δ estδ % foci 6 4 Zone P =.76 * 3 Schober et al. Supplemental Figure 3
7 Sir mediated repression telomere 5R ADE TG (-3) 3-35bp tel telomere 5R ADE TG (-3) -5 bp Schober et al Supplemental Figure 4
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/5/eaar2740/dc1 Supplementary Materials for The fission yeast Stn1-Ten1 complex limits telomerase activity via its SUMO-interacting motif and promotes telomeres
advances.sciencemag.org/cgi/content/full/4/5/eaar2740/dc1 Supplementary Materials for The fission yeast Stn1-Ten1 complex limits telomerase activity via its SUMO-interacting motif and promotes telomeres
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
-CH 2. -S-CH 3 How do I know this?
 What is the major functional difference of Theta vs. Rolling-ircle replication, specifically in regards to the tandem linear repeats found in the R- replication. You won t be responsible for this. Rolling-circle
What is the major functional difference of Theta vs. Rolling-ircle replication, specifically in regards to the tandem linear repeats found in the R- replication. You won t be responsible for this. Rolling-circle
The geneticist s questions. Deleting yeast genes. Functional genomics. From Wikipedia, the free encyclopedia
 From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
ASSAY OF CHROMOSOME MOVEMENT AND PAIRING DURING MEIOSIS IN SACCHAROMYCES CEREVISIAE USING LIVE CELL IMAGING
 ASSAY OF CHROMOSOME MOVEMENT AND PAIRING DURING MEIOSIS IN SACCHAROMYCES CEREVISIAE USING LIVE CELL IMAGING James McGehee Abstract Meiosis is a specialized form of cell division, in which homologous chromosomes
ASSAY OF CHROMOSOME MOVEMENT AND PAIRING DURING MEIOSIS IN SACCHAROMYCES CEREVISIAE USING LIVE CELL IMAGING James McGehee Abstract Meiosis is a specialized form of cell division, in which homologous chromosomes
When one gene is wild type and the other mutant:
 Series 2: Cross Diagrams Linkage Analysis There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Series 2: Cross Diagrams Linkage Analysis There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
The phenotype of this worm is wild type. When both genes are mutant: The phenotype of this worm is double mutant Dpy and Unc phenotype.
 Series 1: Cross Diagrams There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome: When both
Series 1: Cross Diagrams There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome: When both
Functional Characterization of the N Terminus of Sir3p
 MOLECULAR AND CELLULAR BIOLOGY, Oct. 1998, p. 6110 6120 Vol. 18, No. 10 0270-7306/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Functional Characterization of the
MOLECULAR AND CELLULAR BIOLOGY, Oct. 1998, p. 6110 6120 Vol. 18, No. 10 0270-7306/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Functional Characterization of the
Chapter 2 Subnuclear Architecture of Telomeres and Subtelomeres in Yeast
 Chapter 2 Subnuclear Architecture of Telomeres and Subtelomeres in Yeast Emmanuelle Fabre and Maya Spichal Abstract Subtelomeres, upstream telomeres, have a very dynamic spatial positioning along the cell
Chapter 2 Subnuclear Architecture of Telomeres and Subtelomeres in Yeast Emmanuelle Fabre and Maya Spichal Abstract Subtelomeres, upstream telomeres, have a very dynamic spatial positioning along the cell
The phenotype of this worm is wild type. When both genes are mutant: The phenotype of this worm is double mutant Dpy and Unc phenotype.
 Series 2: Cross Diagrams - Complementation There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Series 2: Cross Diagrams - Complementation There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Regulation of Gene Expression in Bacteria and Their Viruses
 11 Regulation of Gene Expression in Bacteria and Their Viruses WORKING WITH THE FIGURES 1. Compare the structure of IPTG shown in Figure 11-7 with the structure of galactose shown in Figure 11-5. Why is
11 Regulation of Gene Expression in Bacteria and Their Viruses WORKING WITH THE FIGURES 1. Compare the structure of IPTG shown in Figure 11-7 with the structure of galactose shown in Figure 11-5. Why is
Table S1. Aspergillus nidulans strains used in this study Strain Genotype Derivation
 Supplemental Material De Souza et al., 211 Table S1. Aspergillus nidulans strains used in this study Strain Genotype Derivation CDS295 pyrg89; pyroa4; pyrg Af ::son promotor::gfp-son nup98/nup96 ; chaa1
Supplemental Material De Souza et al., 211 Table S1. Aspergillus nidulans strains used in this study Strain Genotype Derivation CDS295 pyrg89; pyroa4; pyrg Af ::son promotor::gfp-son nup98/nup96 ; chaa1
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Cartesian Ovals. Kurt Nalty. August 14, 2014
 Cartesian Ovals Kurt Nalty August 14, 2014 Abstract The Cartesian oval is a generalization of the conic sections, where the distance from any two of three fixed points is a generic linear relationship.
Cartesian Ovals Kurt Nalty August 14, 2014 Abstract The Cartesian oval is a generalization of the conic sections, where the distance from any two of three fixed points is a generic linear relationship.
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
EST1 Homology Domain. 100 aa. hest1a / SMG6 PIN TPR TPR. Est1-like DBD? hest1b / SMG5. TPR-like TPR. a helical. hest1c / SMG7.
 hest1a / SMG6 EST1 Homology Domain 100 aa 853 695 761 780 1206 hest1 / SMG5 -like? -like 109 145 214 237 497 165 239 1016 114 207 212 381 583 hest1c / SMG7 a helical 1091 Sc 57 185 267 284 699 Figure S1:
hest1a / SMG6 EST1 Homology Domain 100 aa 853 695 761 780 1206 hest1 / SMG5 -like? -like 109 145 214 237 497 165 239 1016 114 207 212 381 583 hest1c / SMG7 a helical 1091 Sc 57 185 267 284 699 Figure S1:
2. Yeast two-hybrid system
 2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transport between cytosol and nucleus
 of 60 3 Gated trans Lectures 9-15 MBLG 2071 The n GATED TRANSPORT transport between cytoplasm and nucleus (bidirectional) controlled by the nuclear pore complex active transport for macro molecules e.g.
of 60 3 Gated trans Lectures 9-15 MBLG 2071 The n GATED TRANSPORT transport between cytoplasm and nucleus (bidirectional) controlled by the nuclear pore complex active transport for macro molecules e.g.
Analysis and modelling of protein interaction networks
 Analysis and modelling of protein interaction networks - A study of the two-hybrid experiment Karin Stibius Jensen Analysis and modelling of protein interaction networks - A study of the two-hybrid experiment
Analysis and modelling of protein interaction networks - A study of the two-hybrid experiment Karin Stibius Jensen Analysis and modelling of protein interaction networks - A study of the two-hybrid experiment
The nature of genomes. Viral genomes. Prokaryotic genome. Nonliving particle. DNA or RNA. Compact genomes with little spacer DNA
 The nature of genomes Genomics: study of structure and function of genomes Genome size variable, by orders of magnitude number of genes roughly proportional to genome size Plasmids symbiotic DNA molecules,
The nature of genomes Genomics: study of structure and function of genomes Genome size variable, by orders of magnitude number of genes roughly proportional to genome size Plasmids symbiotic DNA molecules,
Three Myosins Contribute Uniquely to the Assembly and Constriction of the Fission Yeast Cytokinetic Contractile Ring
 Current Biology Supplemental Information Three Myosins Contribute Uniquely to the Assembly and Constriction of the Fission Yeast Cytokinetic Contractile Ring Caroline Laplante, Julien Berro, Erdem Karatekin,
Current Biology Supplemental Information Three Myosins Contribute Uniquely to the Assembly and Constriction of the Fission Yeast Cytokinetic Contractile Ring Caroline Laplante, Julien Berro, Erdem Karatekin,
The geneticist s questions
 The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
Measuring TF-DNA interactions
 Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Cover Page. The handle holds various files of this Leiden University dissertation
 Cover Page The handle http://hdl.handle.net/1887/41480 holds various files of this Leiden University dissertation Author: Tleis, Mohamed Title: Image analysis for gene expression based phenotype characterization
Cover Page The handle http://hdl.handle.net/1887/41480 holds various files of this Leiden University dissertation Author: Tleis, Mohamed Title: Image analysis for gene expression based phenotype characterization
EMBO. Intracellular trafficking of yeast telomerase components. reports. M. Teresa Teixeira, Klaus Förstemann, Susan M. Gasser 1 & Joachim Lingner +
 EMBO reports Intracellular trafficking of yeast telomerase components M. Teresa Teixeira, Klaus Förstemann, Susan M. Gasser 1 & Joachim Lingner + Swiss Institute for Experimental Cancer Research (ISREC),
EMBO reports Intracellular trafficking of yeast telomerase components M. Teresa Teixeira, Klaus Förstemann, Susan M. Gasser 1 & Joachim Lingner + Swiss Institute for Experimental Cancer Research (ISREC),
AP Physics C. Gauss s Law. Free Response Problems
 AP Physics Gauss s Law Free Response Problems 1. A flat sheet of glass of area 0.4 m 2 is placed in a uniform electric field E = 500 N/. The normal line to the sheet makes an angle θ = 60 ẘith the electric
AP Physics Gauss s Law Free Response Problems 1. A flat sheet of glass of area 0.4 m 2 is placed in a uniform electric field E = 500 N/. The normal line to the sheet makes an angle θ = 60 ẘith the electric
The activities of eukaryotic replication origins in chromatin
 Biochimica et Biophysica Acta 1677 (2004) 142 157 Review The activities of eukaryotic replication origins in chromatin Michael Weinreich a, *, Madeleine A. Palacios DeBeer b, Catherine A. Fox b, * a Laboratory
Biochimica et Biophysica Acta 1677 (2004) 142 157 Review The activities of eukaryotic replication origins in chromatin Michael Weinreich a, *, Madeleine A. Palacios DeBeer b, Catherine A. Fox b, * a Laboratory
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read
 Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read chapter 22 and chapter 10 [section on MATing type gene
Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read chapter 22 and chapter 10 [section on MATing type gene
Ross E Curtis 1,2, Seyoung Kim 2, John L Woolford Jr 3, Wenjie Xu 3 and Eric P Xing 4*
 Curtis et al. BMC Genomics 2013, 14:196 RESEARCH ARTICLE Open Access Structured association analysis leads to insight into Saccharomyces cerevisiae gene regulation by finding multiple contributing eqtl
Curtis et al. BMC Genomics 2013, 14:196 RESEARCH ARTICLE Open Access Structured association analysis leads to insight into Saccharomyces cerevisiae gene regulation by finding multiple contributing eqtl
The nuclear envelope and transcriptional control
 The nuclear envelope and transcriptional control Asifa Akhtar* and Susan M. Gasser Abstract Cells have evolved sophisticated multi-protein complexes that can regulate gene activity at various steps of
The nuclear envelope and transcriptional control Asifa Akhtar* and Susan M. Gasser Abstract Cells have evolved sophisticated multi-protein complexes that can regulate gene activity at various steps of
Explaining Periodic Trends
 Explaining Periodic Trends! Many observable trends in the chemical and physical properties of elements are observable in the periodic table.! On trends you may be familiar with is reactivity, which is
Explaining Periodic Trends! Many observable trends in the chemical and physical properties of elements are observable in the periodic table.! On trends you may be familiar with is reactivity, which is
Mitosis & Meiosis Practice Test
 Name: DO NOT WRITE ON THIS TEST Class: ALL ID: A Mitosis & Meiosis Practice Test Modified True/False Indicate whether the statement is true or false. If false, change the identified word or phrase to make
Name: DO NOT WRITE ON THIS TEST Class: ALL ID: A Mitosis & Meiosis Practice Test Modified True/False Indicate whether the statement is true or false. If false, change the identified word or phrase to make
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Eukaryotic Gene Expression
 Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Bi Lecture 9 Genetic Screens (cont.) Chromosomes
 Bi190-2013 Lecture 9 Genetic Screens (cont.) Chromosomes C. elegans EGF-receptor signaling: a branched signaling pathway LET-23 EGF-R [IP2] PLCγ [IP3] [PIP2] ITR-1 IP3 Receptor SEM-5 Grb2 LET-341 SOS LET-60
Bi190-2013 Lecture 9 Genetic Screens (cont.) Chromosomes C. elegans EGF-receptor signaling: a branched signaling pathway LET-23 EGF-R [IP2] PLCγ [IP3] [PIP2] ITR-1 IP3 Receptor SEM-5 Grb2 LET-341 SOS LET-60
Chapter 2: Chromosomes and cellular reproduction
 Chapter 2: Chromosomes and cellular reproduction I. Contrast between prokaryotes and eukaryotes. See Figure 2.1 Nucleus absent Small diameter 1 to 10 µm Genome usually 1 circular molecule Small genome;
Chapter 2: Chromosomes and cellular reproduction I. Contrast between prokaryotes and eukaryotes. See Figure 2.1 Nucleus absent Small diameter 1 to 10 µm Genome usually 1 circular molecule Small genome;
3.B.1 Gene Regulation. Gene regulation results in differential gene expression, leading to cell specialization.
 3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
Welcome to Class 21!
 Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Exam 1 PBG430/
 1 Exam 1 PBG430/530 2014 1. You read that the genome size of maize is 2,300 Mb and that in this species 2n = 20. This means that there are 2,300 Mb of DNA in a cell that is a. n (e.g. gamete) b. 2n (e.g.
1 Exam 1 PBG430/530 2014 1. You read that the genome size of maize is 2,300 Mb and that in this species 2n = 20. This means that there are 2,300 Mb of DNA in a cell that is a. n (e.g. gamete) b. 2n (e.g.
Unit 3 - Molecular Biology & Genetics - Review Packet
 Name Date Hour Unit 3 - Molecular Biology & Genetics - Review Packet True / False Questions - Indicate True or False for the following statements. 1. Eye color, hair color and the shape of your ears can
Name Date Hour Unit 3 - Molecular Biology & Genetics - Review Packet True / False Questions - Indicate True or False for the following statements. 1. Eye color, hair color and the shape of your ears can
the noisy gene Biology of the Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB)
 Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
UNIVERSITY OF YORK. BA, BSc, and MSc Degree Examinations Department : BIOLOGY. Title of Exam: Molecular microbiology
 Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Supplemental Information: Govindaraghavan, Lad and Osmani
 Supplemental Information: Govindaraghavan, Lad and Osmani Table S1: List of strains used in this study Strain Name Genotype (all strains also carry vea1) Source MG190 nima7; pyrg89; argb2; ndc80-cr::pyroa
Supplemental Information: Govindaraghavan, Lad and Osmani Table S1: List of strains used in this study Strain Name Genotype (all strains also carry vea1) Source MG190 nima7; pyrg89; argb2; ndc80-cr::pyroa
CELL CYCLE, MITOSIS AND MEIOSIS NOTES
 CELL CYCLE, MITOSIS AND MEIOSIS NOTES DNA - Genetic information is stored in the DNA strand in the form of genes. DNA stands for deoxyribose nucleic acid Genes located on the DNA strand 2 Types of DNA
CELL CYCLE, MITOSIS AND MEIOSIS NOTES DNA - Genetic information is stored in the DNA strand in the form of genes. DNA stands for deoxyribose nucleic acid Genes located on the DNA strand 2 Types of DNA
Cell organelles. Cell Wall
 Cell organelles Cell Wall Plant cells have an outermost structure called a cell wall. A cell wall is a rigid structure that gives support to a cell. Plants and algae have cell walls made of a complex sugar.
Cell organelles Cell Wall Plant cells have an outermost structure called a cell wall. A cell wall is a rigid structure that gives support to a cell. Plants and algae have cell walls made of a complex sugar.
What is a sex cell? How are sex cells made? How does meiosis help explain Mendel s results?
 CHAPTER 6 3 Meiosis SECTION Heredity BEFORE YOU READ After you read this section, you should be able to answer these questions: What is a sex cell? How are sex cells made? How does meiosis help explain
CHAPTER 6 3 Meiosis SECTION Heredity BEFORE YOU READ After you read this section, you should be able to answer these questions: What is a sex cell? How are sex cells made? How does meiosis help explain
Unit 6 Test: The Cell Cycle
 Name Date Class Mrs. Knight Biology EHS Unit 6 Test: The Cell Cycle 1. What are the four main stages of the cell cycle (correct order)? A. G 1, S, G 0, M C. G 2, S, G 1, M B. G 1, S, G 2, M D. M, G 2,
Name Date Class Mrs. Knight Biology EHS Unit 6 Test: The Cell Cycle 1. What are the four main stages of the cell cycle (correct order)? A. G 1, S, G 0, M C. G 2, S, G 1, M B. G 1, S, G 2, M D. M, G 2,
Designer Genes C Test
 Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Name Class Date. Term Definition How I m Going to Remember the Meaning
 11.4 Meiosis Lesson Objectives Contrast the number of chromosomes in body cells and in gametes. Summarize the events of meiosis. Contrast meiosis and mitosis. Describe how alleles from different genes
11.4 Meiosis Lesson Objectives Contrast the number of chromosomes in body cells and in gametes. Summarize the events of meiosis. Contrast meiosis and mitosis. Describe how alleles from different genes
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Noble Gas Config. Period Block (s, p, d, f) Group
 LOCATION OF ELEMENTS WORKSHEET Noble Gas Config. Period Block (s, p, d, f) Group 1 [Ne] 3s 2 3p 2 2 [Ar] 4s 2 3d 10 4p 6 3 [Xe] 6s 2 4 [Kr] 5s 2 4d 10 5p 5 5 [Ar] 4s 2 3d 10 4p 1 6 [He] 2s 2 2p 3 7 [Kr]
LOCATION OF ELEMENTS WORKSHEET Noble Gas Config. Period Block (s, p, d, f) Group 1 [Ne] 3s 2 3p 2 2 [Ar] 4s 2 3d 10 4p 6 3 [Xe] 6s 2 4 [Kr] 5s 2 4d 10 5p 5 5 [Ar] 4s 2 3d 10 4p 1 6 [He] 2s 2 2p 3 7 [Kr]
Reading: Chapter 5, pp ; Reference chapter D, pp Problem set F
 Mosaic Analysis Reading: Chapter 5, pp140-141; Reference chapter D, pp820-823 Problem set F Twin spots in Drosophila Although segregation and recombination in mitosis do not occur at the same frequency
Mosaic Analysis Reading: Chapter 5, pp140-141; Reference chapter D, pp820-823 Problem set F Twin spots in Drosophila Although segregation and recombination in mitosis do not occur at the same frequency
Explaining Periodic Trends. Saturday, January 20, 18
 Explaining Periodic Trends Many observable trends in the chemical and physical properties of elements are observable in the periodic table. Let s review a trend that you should already be familiar with,
Explaining Periodic Trends Many observable trends in the chemical and physical properties of elements are observable in the periodic table. Let s review a trend that you should already be familiar with,
Nucleus. The nucleus is a membrane bound organelle that store, protect and express most of the genetic information(dna) found in the cell.
 Nucleus The nucleus is a membrane bound organelle that store, protect and express most of the genetic information(dna) found in the cell. Since regulation of gene expression takes place in the nucleus,
Nucleus The nucleus is a membrane bound organelle that store, protect and express most of the genetic information(dna) found in the cell. Since regulation of gene expression takes place in the nucleus,
Exercise Set 4. D s n ds + + V. s dv = V. After using Stokes theorem, the surface integral becomes
 Exercise Set Exercise - (a) Let s consider a test volum in the pellet. The substract enters the pellet by diffusion and some is created and disappears due to the chemical reaction. The two contribute to
Exercise Set Exercise - (a) Let s consider a test volum in the pellet. The substract enters the pellet by diffusion and some is created and disappears due to the chemical reaction. The two contribute to
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
KEY: Chapter 9 Genetics of Animal Breeding.
 KEY: Chapter 9 Genetics of Animal Breeding. Answer each question using the reading assigned to you. You can access this information by clicking on the following URL: https://drive.google.com/a/meeker.k12.co.us/file/d/0b1yf08xgyhnad08xugxsnfvba28/edit?usp=sh
KEY: Chapter 9 Genetics of Animal Breeding. Answer each question using the reading assigned to you. You can access this information by clicking on the following URL: https://drive.google.com/a/meeker.k12.co.us/file/d/0b1yf08xgyhnad08xugxsnfvba28/edit?usp=sh
Cell Structure, Function & Ultrastructure
 Cell Structure, Function & Ultrastructure Learning Objectives 2.1.2 Components of the cell as seen under the light microscope and their functions. Cell Structure and Function 1. Plant cells: cell wall,
Cell Structure, Function & Ultrastructure Learning Objectives 2.1.2 Components of the cell as seen under the light microscope and their functions. Cell Structure and Function 1. Plant cells: cell wall,
7th 4.2 review. 1. Chromosomes line up single file at the middle of the cell. 2. Two identical nuclei form. 3. Sister chromatids separate.
 Match each activity to the correct phase of mitosis. You may use the same answer more than once. a. prophase b. metaphase c. anaphase d. telophase 1. Chromosomes line up single file at the middle of the
Match each activity to the correct phase of mitosis. You may use the same answer more than once. a. prophase b. metaphase c. anaphase d. telophase 1. Chromosomes line up single file at the middle of the
Classical Selection, Balancing Selection, and Neutral Mutations
 Classical Selection, Balancing Selection, and Neutral Mutations Classical Selection Perspective of the Fate of Mutations All mutations are EITHER beneficial or deleterious o Beneficial mutations are selected
Classical Selection, Balancing Selection, and Neutral Mutations Classical Selection Perspective of the Fate of Mutations All mutations are EITHER beneficial or deleterious o Beneficial mutations are selected
A complementation test would be done by crossing the haploid strains and scoring the phenotype in the diploids.
 Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Review (V1): Phases of Cell Cycle
 Review (V1): Phases of Cell Cycle The cell cycle consists of 4 distinct phases: - G 1 phase, - S phase (synthesis), - G 2 phase - and M phase (mitosis). Interphase: combines G 1, S, and G 2 Activation
Review (V1): Phases of Cell Cycle The cell cycle consists of 4 distinct phases: - G 1 phase, - S phase (synthesis), - G 2 phase - and M phase (mitosis). Interphase: combines G 1, S, and G 2 Activation
Systems Biology of the Cell Cycle
 ICYSB 6th International Course in Yeast Systems Biology Göteborg, Sweden 4 June 213 Systems Biology of the Cell Cycle Assistant Professor University of Amsterdam, E-mail: M.Barberis@uva.nl A Baby of UNICELLSYS
ICYSB 6th International Course in Yeast Systems Biology Göteborg, Sweden 4 June 213 Systems Biology of the Cell Cycle Assistant Professor University of Amsterdam, E-mail: M.Barberis@uva.nl A Baby of UNICELLSYS
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Translation - Prokaryotes
 1 Translation - Prokaryotes Shine-Dalgarno (SD) Sequence rrna 3 -GAUACCAUCCUCCUUA-5 mrna...ggagg..(5-7bp)...aug Influences: Secondary structure!! SD and AUG in unstructured region Start AUG 91% GUG 8 UUG
1 Translation - Prokaryotes Shine-Dalgarno (SD) Sequence rrna 3 -GAUACCAUCCUCCUUA-5 mrna...ggagg..(5-7bp)...aug Influences: Secondary structure!! SD and AUG in unstructured region Start AUG 91% GUG 8 UUG
A redundant function for the N-terminal tail of Ndc80 in kinetochore-microtubule interaction in Saccharomyces cerevisiae.
 Genetics: Published Articles Ahead of Print, published on July 30, 2012 as 10.1534/genetics.112.143818 A redundant function for the N-terminal tail of Ndc80 in kinetochore-microtubule interaction in Saccharomyces
Genetics: Published Articles Ahead of Print, published on July 30, 2012 as 10.1534/genetics.112.143818 A redundant function for the N-terminal tail of Ndc80 in kinetochore-microtubule interaction in Saccharomyces
Chapter 14. Chemical Periodicity
 Chapter 14 Chemical Periodicity Periodic Trends An element s placement in the periodic table determines characteristics like the size of the atom, its ability to attract electrons and the stability of
Chapter 14 Chemical Periodicity Periodic Trends An element s placement in the periodic table determines characteristics like the size of the atom, its ability to attract electrons and the stability of
Engineering Notebook
 Engineering Notebook miscellaneous problems and solutions compiled by C Bond Vol1 no9 Actuator with zero bend angle Rotary Actuator Equations Given two points at which the track radius and gap skew angle
Engineering Notebook miscellaneous problems and solutions compiled by C Bond Vol1 no9 Actuator with zero bend angle Rotary Actuator Equations Given two points at which the track radius and gap skew angle
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
SUPPLEMENTARY INFORMATION
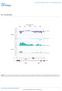 DOI:.8/ncb65 Chromosom XII cdc5- - - RDN7 4585 459 4595 46 465 IGS RDN5- RDN7 Figur S Cdc4 is rquird for rdna silncing. Yast tiling array intnsitis along 5kb of chromosom XII containing IGS rgions ( and
DOI:.8/ncb65 Chromosom XII cdc5- - - RDN7 4585 459 4595 46 465 IGS RDN5- RDN7 Figur S Cdc4 is rquird for rdna silncing. Yast tiling array intnsitis along 5kb of chromosom XII containing IGS rgions ( and
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Texas Biology Standards Review. Houghton Mifflin Harcourt Publishing Company 26 A T
 2.B.6. 1 Which of the following statements best describes the structure of DN? wo strands of proteins are held together by sugar molecules, nitrogen bases, and phosphate groups. B wo strands composed of
2.B.6. 1 Which of the following statements best describes the structure of DN? wo strands of proteins are held together by sugar molecules, nitrogen bases, and phosphate groups. B wo strands composed of
Controlling Gene Expression
 Controlling Gene Expression Control Mechanisms Gene regulation involves turning on or off specific genes as required by the cell Determine when to make more proteins and when to stop making more Housekeeping
Controlling Gene Expression Control Mechanisms Gene regulation involves turning on or off specific genes as required by the cell Determine when to make more proteins and when to stop making more Housekeeping
Cell Division. OpenStax College. 1 Genomic DNA
 OpenStax-CNX module: m44459 1 Cell Division OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will be
OpenStax-CNX module: m44459 1 Cell Division OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you will be
Exam 1 Solutions. Note that there are several variations of some problems, indicated by choices in parentheses. Problem 1
 Exam 1 Solutions Note that there are several variations of some problems, indicated by choices in parentheses. Problem 1 A rod of charge per unit length λ is surrounded by a conducting, concentric cylinder
Exam 1 Solutions Note that there are several variations of some problems, indicated by choices in parentheses. Problem 1 A rod of charge per unit length λ is surrounded by a conducting, concentric cylinder
Biol. 303 EXAM I 9/22/08 Name
 Biol. 303 EXAM I 9/22/08 Name -------------------------------------------------------------------------------------------------------------- This exam consists of 40 multiple choice questions worth 2.5
Biol. 303 EXAM I 9/22/08 Name -------------------------------------------------------------------------------------------------------------- This exam consists of 40 multiple choice questions worth 2.5
Nature Protocols: doi: /nprot Supplementary Figure 1
 Supplementary Figure 1 Photographs of the 3D-MTC device and the confocal fluorescence microscopy. I: The system consists of a Leica SP8-Confocal microscope (with an option of STED), a confocal PC, a 3D-MTC
Supplementary Figure 1 Photographs of the 3D-MTC device and the confocal fluorescence microscopy. I: The system consists of a Leica SP8-Confocal microscope (with an option of STED), a confocal PC, a 3D-MTC
Bacterial Genetics & Operons
 Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Mitosis and Genetics Study Guide Answer Key
 Mitosis and Genetics Study Guide Answer Key 1. Which of the following is true of Interphase? a. It is part of Meiosis b. It occurs before Meiosis c. The cell does normal cell activities during interphase
Mitosis and Genetics Study Guide Answer Key 1. Which of the following is true of Interphase? a. It is part of Meiosis b. It occurs before Meiosis c. The cell does normal cell activities during interphase
Prof. Fahd M. Nasr. Lebanese university Faculty of sciences I Department of Natural Sciences.
 Prof. Fahd M. Nasr Lebanese university Faculty of sciences I Department of Natural Sciences fnasr@ul.edu.lb B3206 Microbial Genetics Eukaryotic M. G. The yeast Saccharomyces cerevisiae as a genetic model
Prof. Fahd M. Nasr Lebanese university Faculty of sciences I Department of Natural Sciences fnasr@ul.edu.lb B3206 Microbial Genetics Eukaryotic M. G. The yeast Saccharomyces cerevisiae as a genetic model
Homeotic genes in flies. Sem 9.3.B.6 Animal Science
 Homeotic genes in flies Sem 9.3.B.6 Animal Science So far We have seen that identities of each segment is determined by various regulators of segment polarity genes In arthopods, and in flies, each segment
Homeotic genes in flies Sem 9.3.B.6 Animal Science So far We have seen that identities of each segment is determined by various regulators of segment polarity genes In arthopods, and in flies, each segment
A thermodynamic origin of a universal mass-angular momentum relationship
 A thermodynamic origin of a universal mass-angular momentum relationship Astrophysics and Space Science, v133, pp 403-410 (1987) M. N. Macrossan Department of Mechanical Engineering University of Queensland,
A thermodynamic origin of a universal mass-angular momentum relationship Astrophysics and Space Science, v133, pp 403-410 (1987) M. N. Macrossan Department of Mechanical Engineering University of Queensland,
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
Meiosis and Fertilization Understanding How Genes Are Inherited 1
 Meiosis and Fertilization Understanding How Genes Are Inherited 1 How does a child inherit one copy of each gene from each parent? Compare what you already know with this flowchart. 1. Fill in each blank
Meiosis and Fertilization Understanding How Genes Are Inherited 1 How does a child inherit one copy of each gene from each parent? Compare what you already know with this flowchart. 1. Fill in each blank
Principles of chromosomal organization: lessons from yeast
 JCB: Review Principles of chromosomal organization: lessons from yeast Christophe Zimmer 1,4 and Emmanuelle Fabre 2,3,5 1 Groupe Imagerie et Modélisation, Département Biologie Cellulaire et Infection;
JCB: Review Principles of chromosomal organization: lessons from yeast Christophe Zimmer 1,4 and Emmanuelle Fabre 2,3,5 1 Groupe Imagerie et Modélisation, Département Biologie Cellulaire et Infection;
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
Chapter 22. Dr. Armen Kocharian. Gauss s Law Lecture 4
 Chapter 22 Dr. Armen Kocharian Gauss s Law Lecture 4 Field Due to a Plane of Charge E must be perpendicular to the plane and must have the same magnitude at all points equidistant from the plane Choose
Chapter 22 Dr. Armen Kocharian Gauss s Law Lecture 4 Field Due to a Plane of Charge E must be perpendicular to the plane and must have the same magnitude at all points equidistant from the plane Choose
4. Why not make all enzymes all the time (even if not needed)? Enzyme synthesis uses a lot of energy.
 1 C2005/F2401 '10-- Lecture 15 -- Last Edited: 11/02/10 01:58 PM Copyright 2010 Deborah Mowshowitz and Lawrence Chasin Department of Biological Sciences Columbia University New York, NY. Handouts: 15A
1 C2005/F2401 '10-- Lecture 15 -- Last Edited: 11/02/10 01:58 PM Copyright 2010 Deborah Mowshowitz and Lawrence Chasin Department of Biological Sciences Columbia University New York, NY. Handouts: 15A
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/5/e1602426/dc1 Supplementary Materials for In vivo mapping of tissue- and subcellular-specific proteomes in Caenorhabditis elegans The PDF file includes: Aaron
advances.sciencemag.org/cgi/content/full/3/5/e1602426/dc1 Supplementary Materials for In vivo mapping of tissue- and subcellular-specific proteomes in Caenorhabditis elegans The PDF file includes: Aaron
Control of Gene Expression in Prokaryotes
 Why? Control of Expression in Prokaryotes How do prokaryotes use operons to control gene expression? Houses usually have a light source in every room, but it would be a waste of energy to leave every light
Why? Control of Expression in Prokaryotes How do prokaryotes use operons to control gene expression? Houses usually have a light source in every room, but it would be a waste of energy to leave every light
Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes.
 Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes. Enduring understanding 3.A: Heritable information provides for continuity of life. Essential
Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes. Enduring understanding 3.A: Heritable information provides for continuity of life. Essential
Development Team. Regulation of gene expression in Prokaryotes: Lac Operon. Molecular Cell Biology. Department of Zoology, University of Delhi
 Paper Module : 15 : 23 Development Team Principal Investigator : Prof. Neeta Sehgal Department of Zoology, University of Delhi Co-Principal Investigator : Prof. D.K. Singh Department of Zoology, University
Paper Module : 15 : 23 Development Team Principal Investigator : Prof. Neeta Sehgal Department of Zoology, University of Delhi Co-Principal Investigator : Prof. D.K. Singh Department of Zoology, University
