Fluorescence fluctuation microscopy: FRAP and FCS
|
|
|
- Marybeth McDonald
- 6 years ago
- Views:
Transcription
1 Fluorescence fluctuation microscopy: FRAP and FCS
2 Confocal laser scanning fluorescence microscope for the mobility analysis in living cells
3 Ensemble measurements of protein mobility and interactions pre post: s 1 s 6 s spatial/temporal resolution bleach once FRAP intensity FRAP spatial analysis bleach intensity width 2 time 1 µm 1 ms - hours 1 µm 1 ms - hours continue bleach CP/ FLIP bleach: s 1 s 6 s intensity time 1 µm 1 ms - hours time correlate.3 µm 1 µs - 1 s Wachsmuth, M., Caudron-Herger, M. and Rippe, K. (28). Biochim. Biophys. Acta 1783,
4 Methods comparison FRAP (for slow/immobile particles, ~1 ms time resolution) - diffusion coefficients - rate constants (at immobile substrates) - immobile fractions - photoactivation instead of bleaching (koff measurements) CP/FLIP (for faster particles) - dissociation rate (at immobile substrates) FCS (for fast particles, µs time resolution) - diffusion coefficients - rate constants (at mobile substrates) - anomaly parameters - concentrations
5 The problem: many things affect the observed mobility collision with immobile chromatin heterogenous binding sites active, directed transport complex formation homgenous binding sites caging
6 Protein mobility and interactions in the cell freely mobile trajectory free time time f et acil lo ita ca te tio d n rg or an gan d iza bi ti nd on in g lf- ta co m pl confined/ moving corral se anomalous ex an for d ma bi ti nd on in g immobilízed Mean Squared Displacehment (MSD) MSD = 6 D t α Dependence of diffusion coefficient D and molecular mass M protein: D M 1 3 double mass M =>.8 fold lower D DNA: D M 1 2 double mass M =>.7 fold lower D Wachsmuth, M., Caudron-Herger, M. and Rippe, K. (28). Biochim. Biophys. Acta 1783,
7 Determining diffusion coefficient D, kinetic binding rates kon and koff, and the apparent equilibirium constant Keq * diffusion without binding, α = 1 for free diffusion transient chromatin binding strong chromatin binding D eff k on k off r D, α D free r 2 = 6 D t α D eff = D * K 1+ K * eq = k * on = k S on eq k off [ ] eq k off Erdel, Müller-Ott, Baum, Wachsmuth & Rippe (211) Chromosome Res 19,
8 Pericentric heterochromatin (PCH) in mouse cells telomere telomere euchromatin centromere (minor satellites) pericentric heterochromatin (PCH) (major satellites repeats) Mouse pericentric heterochromatin - chromocenters visualized by DAPI staining - 5meC, H3K9me3, H4K2me3 - HP1, Suv39h, Suv4-2h - silences transcription of repeats (requires Suv39h) - important for chromosome segregation 5 µm Specificity, propagation and epigenetic memory?
9 Writing, reading and transmitting epigenetic signals pioneering factor writer reader spreading of modification? histone modification from Molecular Biology of the Cell
10 Distinct chromatin states can be established and maintained via interlinked feedback loops silenced H3K9me3 heterochromatin H3K9me3 histone H3 lysine 9 methylation DNA methylase DNMT histone methylase histone acetylase HP1 SUV39h HAT SUV39h, DNMT Jmjd2 HAT HDAC histone H3 lysine 9 acetylation 5meC MeCP1 HDAC histone deacetylase histone demethylase H3K9ac active H3K9ac euchromatin
11 Colocalization of heterochromatin protein 1 (HP1), Suv39H1 histone methylase and H3K9 methylation DAPI GFP-HP1A H3K9me3-Alexa568 Merge GFP-HP1A TagRFP-Suv39H1 Merge pericentric heterochromatin DAPI 1 µm
12 Fluorescence Recovery After Photobleaching (FRAP)
13 Fluorescence Recovery After Photobleaching (FRAP) Protocol: 1. Bleach particles (with a laser) 2. Wait and watch during they diffuse away 3. Fit the fluorescence-over-time curve or profile Source: Wikipedia
14 example: ER marker protein Reits et al. 21 (From fixed to FRAP: measuring protein mobility and activity in living cells)
15 mobile protein immobile protein
16 Typical FRAP curve F i : f.i. before bleaching F : f.i. just after bleaching F : f.i. in recovery region mobile fraction Equation for the recovery curve in the absence of binding: characteristic diffusion time bleach radius w diffusion coefficient D τ D = w 2 4D F i (t) = e 2τ D 2τ t I D 2τ + I D t 1 t I, I 1 : Modified Bessel functions
17 FRAP with fast binding Fast binding: Reduction of the diffusion coefficient, shape of the curve remains unchanged Fast means: Many binding events occur during translocations on the length scale of the bleach spot Only one fit parameter: Effective diffusion coefficient D eff = D 1 + k * on k off If D is known, the ratio of the rate constants is obtained
18 FRAP with slow binding Long-lived binding events lead to different shape of recovery curve Data slow binding Effective diffusion (no or fast binding):.8.6 free/effective diffusion F i (t) = e 2τ D 2τ t I D t Slow binding: 2τ + I D 1 t.4 F i (t) =1 e k off t time (s) dissociation rate Moderately fast binding: no analytical solution
19 Intensity analysis FRAP resolves HP1 diffusion and interactions on the 1 µm and second time scale immobilized fraction (~ 1%) only in heterochromatin residuals relative intensity diffusion model reaction model diffusion-reaction model time (s) time (s) time (s) diffusion-reaction analysis (Sprague & McNally 24, Biophys. J. 86, 3473) yields k off =.15 ±.7 s -1 in heterochromatin
20 FRAP profile analysis yields an effective nuclear diffusion coefficient of Deff = 1.4 µm² s -1 of HP1 (1 µm and second scale) fit to a confined diffusion model
21 Diffusion versus binding in FRAP profile analysis diffusion dominant case y position 5 [mm] -5 1 relative.5 intensity -5 postbleach time s 13.3 s s 5 5 x position [mm] s binding dominant case y position 5 [mm] -5 1 relative.5 intensity -5 postbleach time s 13.3 s x position [mm] 43.3 s s Calculations Malte Wachsmuth
22 Mobility and interaction analysis in living cells Fluorescence recovery after photobleaching (FRAP) intensity 5 µm prebleach postbleach.3 s postbleach 4 s time Fluorescence correlation spectroscopy (FCS) intensity time (µs) correlation fct. log (time) Müller et al. (29). Biophys. J. 97, ; Erdel et al. (211) Chromosome Res 19,
23 Fluorescence correlation spectroscopy
24 The concept: measuring number fluctuations of fluorescent particles in the focus Diffusion induces fluctuations of the number of molecules N = 3 N = 2 N = 4 <N> = 3 F(t) <F> F(t) F(t) F This results in fluctuations of the fluorescence signal t
25 Fluctuations of the particle number of a 1 nm rhodamine solution in dependence of the observation volume Size [mm] Volume [l] particles N N/N [%]
26 FCS data analysis F(t) <F> We want to know: the average number of molecules in the focus concentration the dwell time in the focus diffusion coefficient
27 The concept: autocorrelation analysis δf(t) t τ G(τ) = 2 log τ
28 The concept: autocorrelation analysis δf(t) t τ G(τ) = 2 log τ
29 The concept: autocorrelation analysis δf(t) t τ G(τ) = 2 log τ
30 The concept: autocorrelation analysis δf(t) t τ G(τ) = 2 log τ
31 Results from FCS experiments G(τ) G( ) F( t) F( t ) F( t) 2 1/N 1/c τ corr 1/D log τ variance of fluctuations concentration length of fluctuations diffusion coefficient
32 Theoretical approach - formulas Properties of the optical system Properties of the diffusion process = I (r) =... c (r,t) =... G(τ) Analytical autocorrelation function concentration, brightness, diffusion properties of particles, interactions, cellular environment... log τ
33 FCS correlation function for free diffusion in 3D The correlation function G(τ) gives the probability to detect a particle at time t and at time t + τ That s how a point in the confocal microscope looks like detection efficiency t=1 That s how a diffusing molecule spreads out over time (in 1 dimension) t=2 Green s function for free 3D diffusion
34 FCS correlation function for free diffusion in 3D Probability to detect a particle before and after it has diffused for time τ definition of the correlation function correlation function diffusion time, concentration, focal volume, structure parameter, focus radius G(τ) τ diff resulting function log τ
35 Determining diffusion coefficients small molecules generate short fluctuations... F(t) larger complexes generate longer fluctuations... F(t) t t G(τ)... and rapidly decaying correlation functions log τ... and slowly decaying correlation functions
36 Measuring ligand binding affinity + kon k off I(t) I(t) G(τ) t t log τ Properties of ligand-receptor interactions: dissociation constant, reaction rates, concentrations
37 Fluorescence correlation spectroscopy (FCS) of the H3K9me3 histone methylase Suv39h1 FCS measurements at different locations in the cell G(τ) normalized cytoplasm D = 17 µm² s -1 euchromatin heterochromatin τ (µs)
38 Pixel-wise Photobleaching Profile Evolution Analysis - 3PEA bleach image intensity distribution during scanning acquisition low mobility low mobility 1. bleach line t = 3 τ l 2. bleach line t = 4 τ l 3. bleach line t = 5 τ l high mobility high mobility Erdel & Rippe (212). PNAS 19, E
39 3PEA analysis of Snf2H chromatin remodeler: Deff = 6.5 µm2 s-1 (FRAP:.7 µm2 s-1) RFP Merge experiment Snf2H-GFP D = 44 µm2s-1 fit result D = 6.5 µm2s Snf2H-GFP RFP SSR D (µm2s-1) D (µm2s-1) D (µm2s-1) 8 1
40 camera (area) Wide-field verus confocal microscopy setup fluorescent probe objective reconstruction of the spatial fluorescence distribution laser objective probe synchronisation of scan mirror and signal acquisition light source detector (point) filters pinhole dichroic mirror rotating scan mirror
41 Dynamic processes in the cell nucleus take place on the microsecond to hour time scale molecular diffusion molecular interactions processes particle/structural dynamics fluctuation spectroscopy photobleaching methods methods imaging/tracking 1 µs 1 ms 1 s 1 s Wachsmuth, M., Caudron-Herger, M. and Rippe, K. (28). Biochim. Biophys. Acta 1783,
42 typical resolution acquisition rate (frames/sec) light exposure wide field 25 nm (x,y) > 2 µm (z) 2 low confocal 25 nm (x,y) 6 nm (z) 1-1 high line scanner confocal 25 nm (x) 38 nm (y) 7 nm (z) 3 low
43 Typical values for diffusion coefficients in the nucleus Dmin (µm 2 s -1 ) accesible corral radius methods chromatin/ telomeres µm.2 µm.3-.8 µm CLSM, single particle tracking transcription factor 1-15 (free) -.1 up to 1 µm (nucleus) FRAP (bound) FCS (free, transiently bound) membrane proteins 2-2 (2-D) 1 µm (nuclear membrane) FRAP FCS
44 Accessible range of diffusion coefficients for CLSM, FRAP and FCS measurments tmin / tmax typical analysis volume Dmin / Dmax confocal (single particle tracking).4-2 sec / infinite 1 x 1 µm (x,y) 6 nm (z) µm 2 s -1 / 1 µm 2 s -1 FRAP.4-2 sec / infinite 2 x 2 µm (x,y).6-5 µm (z) µm 2 s -1 / 1 µm 2 s -1 FCS 1 µsec / 1 sec 25 nm (x,y) 6 nm (z).5 µm 2 s -1 / 2 µm 2 s -1
FROM LOCALIZATION TO INTERACTION
 EPFL SV PTBIOP FROM LOCALIZATION TO INTERACTION BIOP COURSE 2015 COLOCALIZATION TYPICAL EXAMPLE EPFL SV PTBIOP Vinculin Alexa568 Actin Alexa488 http://www.olympusconfocal.com/applications/colocalization.html
EPFL SV PTBIOP FROM LOCALIZATION TO INTERACTION BIOP COURSE 2015 COLOCALIZATION TYPICAL EXAMPLE EPFL SV PTBIOP Vinculin Alexa568 Actin Alexa488 http://www.olympusconfocal.com/applications/colocalization.html
In order to confirm the contribution of diffusion to the FRAP recovery curves of
 Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
Solution structure and dynamics of biopolymers
 Solution structure and dynamics of biopolymers Atomic-detail vs. low resolution structure Information available at different scales Mobility of macromolecules in solution Brownian motion, random walk,
Solution structure and dynamics of biopolymers Atomic-detail vs. low resolution structure Information available at different scales Mobility of macromolecules in solution Brownian motion, random walk,
A short story about FRAP (fluourescence recovery after photobleaching)
 1 A short story about FRAP (fluourescence recovery after photobleaching) Bob Pego (Carnegie Mellon) work involving team leader: James McNally (NCI Lab of Receptor Biology & Gene Expression) Biophysicists:
1 A short story about FRAP (fluourescence recovery after photobleaching) Bob Pego (Carnegie Mellon) work involving team leader: James McNally (NCI Lab of Receptor Biology & Gene Expression) Biophysicists:
The Photon Counting Histogram: Statistical Analysis of Single Molecule Populations
 The Photon Counting Histogram: Statistical Analysis of Single Molecule Populations E. Gratton Laboratory for Fluorescence Dynamics University of California, Irvine Transition from FCS The Autocorrelation
The Photon Counting Histogram: Statistical Analysis of Single Molecule Populations E. Gratton Laboratory for Fluorescence Dynamics University of California, Irvine Transition from FCS The Autocorrelation
FLUORESCENCE MICROSCOPY TECHNIQUES PRACTICAL MANUAL FOR
 FLUORESCENCE PRACTICAL MANUAL FOR MICROSCOPY TECHNIQUES Sohail Ahmed Sudhaharan Thankiah Radek Machán Martin Hof Andrew H. A. Clayton Graham Wright Jean-Baptiste Sibarita Thomas Korte Andreas Herrmann
FLUORESCENCE PRACTICAL MANUAL FOR MICROSCOPY TECHNIQUES Sohail Ahmed Sudhaharan Thankiah Radek Machán Martin Hof Andrew H. A. Clayton Graham Wright Jean-Baptiste Sibarita Thomas Korte Andreas Herrmann
Rice/TCU REU on Computational Neuroscience. Fundamentals of Molecular Imaging
 Rice/TCU REU on Computational Neuroscience Fundamentals of Molecular Imaging June 3, 2008 Neal Waxham 713-500-5621 m.n.waxham@uth.tmc.edu Objectives Brief discussion of optical resolution and lasers as
Rice/TCU REU on Computational Neuroscience Fundamentals of Molecular Imaging June 3, 2008 Neal Waxham 713-500-5621 m.n.waxham@uth.tmc.edu Objectives Brief discussion of optical resolution and lasers as
Retrieving the intracellular topology from multi-scale protein mobility mapping in living cells
 ARTICLE Received 24 Oct 2013 Accepted 24 Jun 2014 Published 24 Jul 2014 Retrieving the intracellular topology from multi-scale protein mobility mapping in living cells Michael Baum 1, Fabian Erdel 1, Malte
ARTICLE Received 24 Oct 2013 Accepted 24 Jun 2014 Published 24 Jul 2014 Retrieving the intracellular topology from multi-scale protein mobility mapping in living cells Michael Baum 1, Fabian Erdel 1, Malte
Interactions between proteins, DNA and RNA. The energy, length and time coordinate system to find your way in the cell
 Interactions between proteins, DNA and RNA The energy, length and time coordinate system to find your way in the cell Scanning/atomic force microscope (SFM/AFM) mirror laser beam photo diode fluid cell
Interactions between proteins, DNA and RNA The energy, length and time coordinate system to find your way in the cell Scanning/atomic force microscope (SFM/AFM) mirror laser beam photo diode fluid cell
BMB Class 17, November 30, Single Molecule Biophysics (II)
 BMB 178 2018 Class 17, November 30, 2018 15. Single Molecule Biophysics (II) New Advances in Single Molecule Techniques Atomic Force Microscopy Single Molecule Manipulation - optical traps and tweezers
BMB 178 2018 Class 17, November 30, 2018 15. Single Molecule Biophysics (II) New Advances in Single Molecule Techniques Atomic Force Microscopy Single Molecule Manipulation - optical traps and tweezers
Rotational Brownian motion; Fluorescence correlation spectroscpy; Photobleaching and FRET. David A. Case Rutgers, Spring 2009
 Rotational Brownian motion; Fluorescence correlation spectroscpy; Photobleaching and FRET David A. Case Rutgers, Spring 2009 Techniques based on rotational motion What we studied last time probed translational
Rotational Brownian motion; Fluorescence correlation spectroscpy; Photobleaching and FRET David A. Case Rutgers, Spring 2009 Techniques based on rotational motion What we studied last time probed translational
Modular scanning FCS quantifies receptor-ligand interactions in living multicellular organisms
 nature methods Modular scanning FCS quantifies receptor-ligand interactions in living multicellular organisms Jonas Ries, Shuizi Rachel Yu, Markus Burkhardt, Michael Brand & Petra Schwille Supplementary
nature methods Modular scanning FCS quantifies receptor-ligand interactions in living multicellular organisms Jonas Ries, Shuizi Rachel Yu, Markus Burkhardt, Michael Brand & Petra Schwille Supplementary
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2011 Monday, 27 June, 9.30 am 12.30 pm Answer
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2011 Monday, 27 June, 9.30 am 12.30 pm Answer
Analysis of Active Transport by Fluorescence Recovery after Photobleaching. MV Ciocanel, JA Kreiling, JA Gagnon, KL Mowry, B Sandstede
 Analysis of Active Transport by Fluorescence Recovery after Photobleaching MV Ciocanel, JA Kreiling, JA Gagnon, KL Mowry, B Sandstede Analysis of Active Transport by Fluorescence Recovery after Photobleaching
Analysis of Active Transport by Fluorescence Recovery after Photobleaching MV Ciocanel, JA Kreiling, JA Gagnon, KL Mowry, B Sandstede Analysis of Active Transport by Fluorescence Recovery after Photobleaching
The Photon Counting Histogram (PCH)
 The Photon Counting Histogram (PCH) While Fluorescence Correlation Spectroscopy (FCS) can measure the diffusion coefficient and concentration of fluorophores, a complementary method, a photon counting
The Photon Counting Histogram (PCH) While Fluorescence Correlation Spectroscopy (FCS) can measure the diffusion coefficient and concentration of fluorophores, a complementary method, a photon counting
Quantitative fluorescence loss in photobleaching for analysis of protein transport and aggregation
 Wüstner et al. BMC Bioinformatics 2012, 13:296 METHODOLOGY ARTICLE Open Access Quantitative fluorescence loss in photobleaching for analysis of protein transport and aggregation Daniel Wüstner 1*, Lukasz
Wüstner et al. BMC Bioinformatics 2012, 13:296 METHODOLOGY ARTICLE Open Access Quantitative fluorescence loss in photobleaching for analysis of protein transport and aggregation Daniel Wüstner 1*, Lukasz
Fluorescence Resonance Energy Transfer (FRET) Microscopy
 Fluorescence Resonance Energy Transfer () Microscopy Mike Lorenz Optical Technology Development mlorenz@mpi-cbg.de -FLM course, May 2009 What is fluorescence? Stoke s shift Fluorescence light is always
Fluorescence Resonance Energy Transfer () Microscopy Mike Lorenz Optical Technology Development mlorenz@mpi-cbg.de -FLM course, May 2009 What is fluorescence? Stoke s shift Fluorescence light is always
Regulation of gene Expression in Prokaryotes & Eukaryotes
 Regulation of gene Expression in Prokaryotes & Eukaryotes 1 The trp Operon Contains 5 genes coding for proteins (enzymes) required for the synthesis of the amino acid tryptophan. Also contains a promoter
Regulation of gene Expression in Prokaryotes & Eukaryotes 1 The trp Operon Contains 5 genes coding for proteins (enzymes) required for the synthesis of the amino acid tryptophan. Also contains a promoter
BMB November 17, Single Molecule Biophysics (I)
 BMB 178 2017 November 17, 2017 14. Single Molecule Biophysics (I) Goals 1. Understand the information SM experiments can provide 2. Be acquainted with different SM approaches 3. Learn to interpret SM results
BMB 178 2017 November 17, 2017 14. Single Molecule Biophysics (I) Goals 1. Understand the information SM experiments can provide 2. Be acquainted with different SM approaches 3. Learn to interpret SM results
Measuring Colocalization within Fluorescence Microscopy Images
 from photonics.com: 03/01/2007 http://www.photonics.com/article.aspx?aid=39341 Measuring Colocalization within Fluorescence Microscopy Images Two-color fluorescence-based methods are uncovering molecular
from photonics.com: 03/01/2007 http://www.photonics.com/article.aspx?aid=39341 Measuring Colocalization within Fluorescence Microscopy Images Two-color fluorescence-based methods are uncovering molecular
Gene Expression. Molecular Genetics, March, 2018
 Gene Expression Molecular Genetics, March, 2018 Gene Expression Control of Protein Levels Bacteria Lac Operon Promoter mrna Inducer CAP Control Trp Operon RepressorOperator Control Attenuation Riboswitches
Gene Expression Molecular Genetics, March, 2018 Gene Expression Control of Protein Levels Bacteria Lac Operon Promoter mrna Inducer CAP Control Trp Operon RepressorOperator Control Attenuation Riboswitches
Supporting Information
 Simultaneous Electro-Optical Tracking for Nanoparticle Recognition and Counting Elena Angeli*,1, Andrea Volpe 1, Paola Fanzio 1, Luca Repetto 1, Giuseppe Firpo 1, Patrizia Guida 1, Roberto Lo Savio 1,
Simultaneous Electro-Optical Tracking for Nanoparticle Recognition and Counting Elena Angeli*,1, Andrea Volpe 1, Paola Fanzio 1, Luca Repetto 1, Giuseppe Firpo 1, Patrizia Guida 1, Roberto Lo Savio 1,
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Single-Molecule Methods I - in vitro
 Single-Molecule Methods I - in vitro Bo Huang Macromolecules 2014.03.10 F 1 -ATPase: a case study Membrane ADP ATP Rotation of the axle when hydrolyzing ATP Kinosita group, 1997-2005 Single Molecule Methods
Single-Molecule Methods I - in vitro Bo Huang Macromolecules 2014.03.10 F 1 -ATPase: a case study Membrane ADP ATP Rotation of the axle when hydrolyzing ATP Kinosita group, 1997-2005 Single Molecule Methods
FLUORESCENCE CORRELATION SPECTROSCOPY
 FLUORESCENCE CORRELATION SPECTROSCOPY ABSTRACT Fluorescence correlation spectroscopy (FCS) is a technique that detects diffusion of fluorescent species through a small focal volume. From FCS measurements,
FLUORESCENCE CORRELATION SPECTROSCOPY ABSTRACT Fluorescence correlation spectroscopy (FCS) is a technique that detects diffusion of fluorescent species through a small focal volume. From FCS measurements,
The nature of genomes. Viral genomes. Prokaryotic genome. Nonliving particle. DNA or RNA. Compact genomes with little spacer DNA
 The nature of genomes Genomics: study of structure and function of genomes Genome size variable, by orders of magnitude number of genes roughly proportional to genome size Plasmids symbiotic DNA molecules,
The nature of genomes Genomics: study of structure and function of genomes Genome size variable, by orders of magnitude number of genes roughly proportional to genome size Plasmids symbiotic DNA molecules,
Detection of Single Photon Emission by Hanbury-Brown Twiss Interferometry
 Detection of Single Photon Emission by Hanbury-Brown Twiss Interferometry Greg Howland and Steven Bloch May 11, 009 Abstract We prepare a solution of nano-diamond particles on a glass microscope slide
Detection of Single Photon Emission by Hanbury-Brown Twiss Interferometry Greg Howland and Steven Bloch May 11, 009 Abstract We prepare a solution of nano-diamond particles on a glass microscope slide
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Investigating the motility of Ecoli using Fluorescence Correlation spectroscopy. 30 may 2005
 Investigating the motility of Ecoli using Fluorescence Correlation spectroscopy Anders Rosengren Peter Bjødstrup jensen 30 may 2005 Contents 1 Introduction 2 1.1 E.coli... 2 2 Theory 4 2.1 Diffusionandrandomwalk...
Investigating the motility of Ecoli using Fluorescence Correlation spectroscopy Anders Rosengren Peter Bjødstrup jensen 30 may 2005 Contents 1 Introduction 2 1.1 E.coli... 2 2 Theory 4 2.1 Diffusionandrandomwalk...
Co-localization, FRET
 Co-localization, FRET Last class FRAP Diffusion This class Co-localization Correlation FRET Co-localization Can you infer function of protein from it s intracellular location How do you measure if two
Co-localization, FRET Last class FRAP Diffusion This class Co-localization Correlation FRET Co-localization Can you infer function of protein from it s intracellular location How do you measure if two
Supplementary Figures Supplementary Figure 1: Estimation of the error of the number and brightness of molecules in a single cluster; Simulation
 Supplementary Figures Supplementary Figure 1: Estimation of the error of the number and brightness of molecules in a single cluster; Simulation (a,c) Relative estimated numbers of molecules ; (b,d) relative
Supplementary Figures Supplementary Figure 1: Estimation of the error of the number and brightness of molecules in a single cluster; Simulation (a,c) Relative estimated numbers of molecules ; (b,d) relative
Supplementary Figures
 Supplementary Figures Supplementary Figure. X-ray diffraction pattern of CH 3 NH 3 PbI 3 film. Strong reflections of the () family of planes is characteristics of the preferred orientation of the perovskite
Supplementary Figures Supplementary Figure. X-ray diffraction pattern of CH 3 NH 3 PbI 3 film. Strong reflections of the () family of planes is characteristics of the preferred orientation of the perovskite
Anti-Bunching from a Quantum Dot
 Anti-Bunching from a Quantum Dot Gerardo I. Viza 1, 1 Department of Physics and Astronomy, University of Rochester, Rochester, NY 14627 We study the nature of non-classical single emitter light experimentally
Anti-Bunching from a Quantum Dot Gerardo I. Viza 1, 1 Department of Physics and Astronomy, University of Rochester, Rochester, NY 14627 We study the nature of non-classical single emitter light experimentally
Ultrafast Dynamics and Single Particle Spectroscopy of Au-CdSe Nanorods
 Supporting Information Ultrafast Dynamics and Single Particle Spectroscopy of Au-CdSe Nanorods G. Sagarzazu a, K. Inoue b, M. Saruyama b, M. Sakamoto b, T. Teranishi b, S. Masuo a and N. Tamai a a Department
Supporting Information Ultrafast Dynamics and Single Particle Spectroscopy of Au-CdSe Nanorods G. Sagarzazu a, K. Inoue b, M. Saruyama b, M. Sakamoto b, T. Teranishi b, S. Masuo a and N. Tamai a a Department
Confocal Microscopy Imaging of Single Emitter Fluorescence and Hanbury Brown and Twiss Photon Antibunching Setup
 1 Confocal Microscopy Imaging of Single Emitter Fluorescence and Hanbury Brown and Twiss Photon Antibunching Setup Abstract Jacob Begis The purpose of this lab was to prove that a source of light can be
1 Confocal Microscopy Imaging of Single Emitter Fluorescence and Hanbury Brown and Twiss Photon Antibunching Setup Abstract Jacob Begis The purpose of this lab was to prove that a source of light can be
Simultaneous intracellular chloride and ph measurements using a GFPbased
 nature methods Simultaneous intracellular chloride and ph measurements using a GFPbased sensor Daniele Arosio, Fernanda Ricci, Laura Marchetti, Roberta Gualdani, Lorenzo Albertazzi & Fabio Beltram Supplementary
nature methods Simultaneous intracellular chloride and ph measurements using a GFPbased sensor Daniele Arosio, Fernanda Ricci, Laura Marchetti, Roberta Gualdani, Lorenzo Albertazzi & Fabio Beltram Supplementary
Nonlinear Optics. Single-Molecule Microscopy Group. Physical Optics Maria Dienerowitz.
 Single-Molecule Microscopy Group Nonlinear Optics Physical Optics 21-06-2017 Maria Dienerowitz maria.dienerowitz@med.uni-jena.de www.single-molecule-microscopy.uniklinikum-jena.de Contents Introduction
Single-Molecule Microscopy Group Nonlinear Optics Physical Optics 21-06-2017 Maria Dienerowitz maria.dienerowitz@med.uni-jena.de www.single-molecule-microscopy.uniklinikum-jena.de Contents Introduction
Supporting Information. Even Hard Sphere Colloidal Suspensions Display. Fickian yet Non-Gaussian Diffusion
 Supporting Information Even Hard Sphere Colloidal Suspensions Display Fickian yet Non-Gaussian Diffusion Juan Guan, a Bo Wang, a and Steve Granick a,b,c,* Departments of Materials Science, a Chemistry,
Supporting Information Even Hard Sphere Colloidal Suspensions Display Fickian yet Non-Gaussian Diffusion Juan Guan, a Bo Wang, a and Steve Granick a,b,c,* Departments of Materials Science, a Chemistry,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer
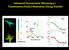 Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
nucleus: DNA & chromosomes
 nucleus: DNA & chromosomes chapter 5 nuclear organization nuclear structure nuclear envelope nucleoplasm nuclear matrix nucleolus nuclear envelope nucleolus nuclear matrix nucleoplasm nuclear pore nuclear
nucleus: DNA & chromosomes chapter 5 nuclear organization nuclear structure nuclear envelope nucleoplasm nuclear matrix nucleolus nuclear envelope nucleolus nuclear matrix nucleoplasm nuclear pore nuclear
Laboratory 3&4: Confocal Microscopy Imaging of Single-Emitter Fluorescence and Hanbury Brown and Twiss setup for Photon Antibunching
 Laboratory 3&4: Confocal Microscopy Imaging of Single-Emitter Fluorescence and Hanbury Brown and Twiss setup for Photon Antibunching Jose Alejandro Graniel Institute of Optics University of Rochester,
Laboratory 3&4: Confocal Microscopy Imaging of Single-Emitter Fluorescence and Hanbury Brown and Twiss setup for Photon Antibunching Jose Alejandro Graniel Institute of Optics University of Rochester,
Regulation of Transcription in Eukaryotes
 Regulation of Transcription in Eukaryotes Leucine zipper and helix-loop-helix proteins contain DNA-binding domains formed by dimerization of two polypeptide chains. Different members of each family can
Regulation of Transcription in Eukaryotes Leucine zipper and helix-loop-helix proteins contain DNA-binding domains formed by dimerization of two polypeptide chains. Different members of each family can
When do diffusion-limited trajectories become memoryless?
 When do diffusion-limited trajectories become memoryless? Maciej Dobrzyński CWI (Center for Mathematics and Computer Science) Kruislaan 413, 1098 SJ Amsterdam, The Netherlands Abstract Stochastic description
When do diffusion-limited trajectories become memoryless? Maciej Dobrzyński CWI (Center for Mathematics and Computer Science) Kruislaan 413, 1098 SJ Amsterdam, The Netherlands Abstract Stochastic description
This document contains the following supporting information: 1. Wide field scanning electron microscope image
 Supporting information for Self-assembled nanoparticle dimer antennas for plasmonic-enhanced single-molecule fluorescence detection at micromolar concentrations Deep Punj, Raju Regmi, Alexis Devilez, Robin
Supporting information for Self-assembled nanoparticle dimer antennas for plasmonic-enhanced single-molecule fluorescence detection at micromolar concentrations Deep Punj, Raju Regmi, Alexis Devilez, Robin
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Measurements of interaction forces in (biological) model systems
 Measurements of interaction forces in (biological) model systems Marina Ruths Department of Chemistry, UMass Lowell What can force measurements tell us about a system? Depending on the technique, we might
Measurements of interaction forces in (biological) model systems Marina Ruths Department of Chemistry, UMass Lowell What can force measurements tell us about a system? Depending on the technique, we might
Super Resolution Microscopy Structured Illumination
 Super Resolution Microscopy Structured Illumination Bo Huang Department of Pharmaceutical Chemistry, UCSF CSHL Quantitative Microscopy, 10/31/2011 50 years to extend the resolution Confocal microscopy
Super Resolution Microscopy Structured Illumination Bo Huang Department of Pharmaceutical Chemistry, UCSF CSHL Quantitative Microscopy, 10/31/2011 50 years to extend the resolution Confocal microscopy
AP3162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions
 AP162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions Hyun Youk Delft University of Technology (Dated: March 15, 2018) In this lecture, we will derive the Berg-Purcell
AP162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions Hyun Youk Delft University of Technology (Dated: March 15, 2018) In this lecture, we will derive the Berg-Purcell
Gene regulation III Biochemistry 302. Bob Kelm March 2, 2005
 Gene regulation III Biochemistry 302 Bob Kelm March 2, 2005 oncept of transcription ground state Prokaryotes: permissive Eukaryotes: restricted DNA structure: chromatin silencing Requirement for sitespecific
Gene regulation III Biochemistry 302 Bob Kelm March 2, 2005 oncept of transcription ground state Prokaryotes: permissive Eukaryotes: restricted DNA structure: chromatin silencing Requirement for sitespecific
Nuclear import of DNA: genetic modification of plants
 Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
Nature Protocols: doi: /nprot Supplementary Figure 1
 Supplementary Figure 1 Photographs of the 3D-MTC device and the confocal fluorescence microscopy. I: The system consists of a Leica SP8-Confocal microscope (with an option of STED), a confocal PC, a 3D-MTC
Supplementary Figure 1 Photographs of the 3D-MTC device and the confocal fluorescence microscopy. I: The system consists of a Leica SP8-Confocal microscope (with an option of STED), a confocal PC, a 3D-MTC
protein biology cell imaging Automated imaging and high-content analysis
 protein biology cell imaging Automated imaging and high-content analysis Automated imaging and high-content analysis Thermo Fisher Scientific offers a complete portfolio of imaging platforms, reagents
protein biology cell imaging Automated imaging and high-content analysis Automated imaging and high-content analysis Thermo Fisher Scientific offers a complete portfolio of imaging platforms, reagents
Concept review: Binding equilibria
 Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
A STOCHASTIC SIMULATION ALGORITHM FOR ANALYSIS OF FLUORESCENCE RECOVERY AFTER PHOTOBLEACHING KINETICS
 49 A STOCHASTIC SIMULATION ALGORITHM FOR ANALYSIS OF FLUORESCENCE RECOVERY AFTER PHOTOBLEACHING KINETICS Glotsos Dimitris, Ph.D. 1, Spiros Kostopoulos, MS.c. 1, and Cavouras Dionisis, Ph.D. 1 1 Department
49 A STOCHASTIC SIMULATION ALGORITHM FOR ANALYSIS OF FLUORESCENCE RECOVERY AFTER PHOTOBLEACHING KINETICS Glotsos Dimitris, Ph.D. 1, Spiros Kostopoulos, MS.c. 1, and Cavouras Dionisis, Ph.D. 1 1 Department
FOGA II. WHAT DOES A GENOME HAVE TO DO? - GENOME FUNCTION AND ORGANIZATION
 FOGA II. WHAT DOES A GENOME HAVE TO DO? - GENOME FUNCTION AND ORGANIZATION Cognitive, computational context Sophistication of cellular information processing and control regimes Functional requirements
FOGA II. WHAT DOES A GENOME HAVE TO DO? - GENOME FUNCTION AND ORGANIZATION Cognitive, computational context Sophistication of cellular information processing and control regimes Functional requirements
Petra Schwille and Elke Haustein Fluorescence Correlation Spectroscopy 1 TABLE OF CONTENTS 1 1. INTRODUCTION 2 2. EXPERIMENTAL REALIZATION 4
 Petra Schwille and Elke Haustein Fluorescence Correlation Spectroscopy Table of Contents TABLE OF CONTENTS. INTRODUCTION. EXPERIMENTAL REALIZATION 4.. ONE-PHOTON EXCITATION 4.. TWO-PHOTON EXCITATION 6.3.
Petra Schwille and Elke Haustein Fluorescence Correlation Spectroscopy Table of Contents TABLE OF CONTENTS. INTRODUCTION. EXPERIMENTAL REALIZATION 4.. ONE-PHOTON EXCITATION 4.. TWO-PHOTON EXCITATION 6.3.
Protein-Ligand Interactions Are Responsible for Signal Transduction
 Proteinigand Interactions Are Responsible for Signal ransduction ypes of Interactions: 1. ProteinProtein 2. ProteinDNA (RNA) 3. Proteinsmall molecule Dynamic Proteinigand Interaction Dynamic interaction
Proteinigand Interactions Are Responsible for Signal ransduction ypes of Interactions: 1. ProteinProtein 2. ProteinDNA (RNA) 3. Proteinsmall molecule Dynamic Proteinigand Interaction Dynamic interaction
LIST of SUPPLEMENTARY MATERIALS
 LIST of SUPPLEMENTARY MATERIALS Mir et al., Dense Bicoid Hubs Accentuate Binding along the Morphogen Gradient Supplemental Movie S1 (Related to Figure 1). Movies corresponding to the still frames shown
LIST of SUPPLEMENTARY MATERIALS Mir et al., Dense Bicoid Hubs Accentuate Binding along the Morphogen Gradient Supplemental Movie S1 (Related to Figure 1). Movies corresponding to the still frames shown
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
MITOHEALTH Nordic Centre of Excellence Workshop in: Mitochondrial function and metabolic diseases
 MITOHEALTH Nordic Centre of Excellence Workshop in: Mitochondrial function and metabolic diseases Quantitative multi-parameter microscopy of mitochondrial function in living cells Werner J.H. Koopman,
MITOHEALTH Nordic Centre of Excellence Workshop in: Mitochondrial function and metabolic diseases Quantitative multi-parameter microscopy of mitochondrial function in living cells Werner J.H. Koopman,
Laboratory 3: Confocal Microscopy Imaging of Single Emitter Fluorescence and Hanbury Brown, and Twiss Setup for Photon Antibunching
 Laboratory 3: Confocal Microscopy Imaging of Single Emitter Fluorescence and Hanbury Brown, and Twiss Setup for Photon Antibunching Jonathan Papa 1, * 1 Institute of Optics University of Rochester, Rochester,
Laboratory 3: Confocal Microscopy Imaging of Single Emitter Fluorescence and Hanbury Brown, and Twiss Setup for Photon Antibunching Jonathan Papa 1, * 1 Institute of Optics University of Rochester, Rochester,
Combining High Resolution Optical and Scanning Probe Microscopy
 Combining High Resolution Optical and Scanning Probe Microscopy Fernando Vargas WITec, Ulm, Germany www.witec.de Company Background Foundation 1997 by O. Hollricher, J. Koenen, K. Weishaupt WITec = Wissenschaftliche
Combining High Resolution Optical and Scanning Probe Microscopy Fernando Vargas WITec, Ulm, Germany www.witec.de Company Background Foundation 1997 by O. Hollricher, J. Koenen, K. Weishaupt WITec = Wissenschaftliche
Are DNA Transcription Factor Proteins Maxwellian Demons?
 Are DNA Transcription Factor Proteins Maxwellian Demons? * physics as a design constraint on molecular evolution. Alexander Grosberg, Longhua Hu, U. Minnesota, NYU U. Minnesota Biophysical Journal 2008
Are DNA Transcription Factor Proteins Maxwellian Demons? * physics as a design constraint on molecular evolution. Alexander Grosberg, Longhua Hu, U. Minnesota, NYU U. Minnesota Biophysical Journal 2008
Nonlinear Optics. Single-Molecule Microscopy Group. Physical Optics Maria Dienerowitz.
 Single-Molecule Microscopy Group Nonlinear Optics Physical Optics 21-06-2017 Maria Dienerowitz maria.dienerowitz@med.uni-jena.de www.single-molecule-microscopy.uniklinikum-jena.de Contents Introduction
Single-Molecule Microscopy Group Nonlinear Optics Physical Optics 21-06-2017 Maria Dienerowitz maria.dienerowitz@med.uni-jena.de www.single-molecule-microscopy.uniklinikum-jena.de Contents Introduction
Supplemental material for Bound electron nonlinearity beyond the ionization threshold
 Supplemental material for Bound electron nonlinearity beyond the ionization threshold 1. Experimental setup The laser used in the experiments is a λ=800 nm Ti:Sapphire amplifier producing 42 fs, 10 mj
Supplemental material for Bound electron nonlinearity beyond the ionization threshold 1. Experimental setup The laser used in the experiments is a λ=800 nm Ti:Sapphire amplifier producing 42 fs, 10 mj
Single Molecule Spectroscopy and Imaging
 Single Molecule Spectroscopy and Imaging Ingo Gregor, Thomas Dertinger, Iris von der Hocht, Jan Sykora, Luru Dai, Jörg Enderlein Institute for Biological Information Processing 1 Forschungszentrum Jülich
Single Molecule Spectroscopy and Imaging Ingo Gregor, Thomas Dertinger, Iris von der Hocht, Jan Sykora, Luru Dai, Jörg Enderlein Institute for Biological Information Processing 1 Forschungszentrum Jülich
Lecture 7 FCS, Autocorrelation, PCH, Cross-correlation
 Lecture 7 FCS, Autocorrelation, PCH, Cross-correlation Joachim Mueller Principles of Fluorescence Techniques Laboratory for Fluorescence Dynamics Figure and slide acknowledgements: Enrico Gratton Historic
Lecture 7 FCS, Autocorrelation, PCH, Cross-correlation Joachim Mueller Principles of Fluorescence Techniques Laboratory for Fluorescence Dynamics Figure and slide acknowledgements: Enrico Gratton Historic
Correlation Spectroscopy in Polymer Physics Methodenseminar im Wahlpflichtfach Basics diffusion and brownian motion correlations functions
 Correlation Spectroscopy in Polymer Physics Methodenseminar im Wahlpflichtfach 3 1. Basics diffusion and brownian motion correlations functions 2. Dynamic light scattering (DLS) DLS on cellulose solutions
Correlation Spectroscopy in Polymer Physics Methodenseminar im Wahlpflichtfach 3 1. Basics diffusion and brownian motion correlations functions 2. Dynamic light scattering (DLS) DLS on cellulose solutions
CS-E5880 Modeling biological networks Gene regulatory networks
 CS-E5880 Modeling biological networks Gene regulatory networks Jukka Intosalmi (based on slides by Harri Lähdesmäki) Department of Computer Science Aalto University January 12, 2018 Outline Modeling gene
CS-E5880 Modeling biological networks Gene regulatory networks Jukka Intosalmi (based on slides by Harri Lähdesmäki) Department of Computer Science Aalto University January 12, 2018 Outline Modeling gene
Binding Theory Equations for Affinity and Kinetics Analysis
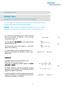 Technology Note #101 Binding Theory Equations for Affinity and Kinetics Analysis This technology note summarizes important equations underlying the theory of binding of solute analytes to surface-tethered
Technology Note #101 Binding Theory Equations for Affinity and Kinetics Analysis This technology note summarizes important equations underlying the theory of binding of solute analytes to surface-tethered
5. 3P PIV Measurements
 Micro PIV Last Class: 1. Data Validation 2. Vector Field Operator (Differentials & Integrals) 3. Standard Differential Scheme 4. Implementation of Differential & Integral quantities with PIV data 5. 3P
Micro PIV Last Class: 1. Data Validation 2. Vector Field Operator (Differentials & Integrals) 3. Standard Differential Scheme 4. Implementation of Differential & Integral quantities with PIV data 5. 3P
Richik N. Ghosh, Linnette Grove, and Oleg Lapets ASSAY and Drug Development Technologies 2004, 2:
 1 3/1/2005 A Quantitative Cell-Based High-Content Screening Assay for the Epidermal Growth Factor Receptor-Specific Activation of Mitogen-Activated Protein Kinase Richik N. Ghosh, Linnette Grove, and Oleg
1 3/1/2005 A Quantitative Cell-Based High-Content Screening Assay for the Epidermal Growth Factor Receptor-Specific Activation of Mitogen-Activated Protein Kinase Richik N. Ghosh, Linnette Grove, and Oleg
Cellular phenomena are characterized by vast
 Fluorescence correlation spectroscopy and fluorescence cross-correlation spectroscopy Michelle A. Digman 1 and Enrico Gratton 1 This article focuses on methods based on fluctuation correlation spectroscopy
Fluorescence correlation spectroscopy and fluorescence cross-correlation spectroscopy Michelle A. Digman 1 and Enrico Gratton 1 This article focuses on methods based on fluctuation correlation spectroscopy
CHAPTER 3. Cell Structure and Genetic Control. Chapter 3 Outline
 CHAPTER 3 Cell Structure and Genetic Control Chapter 3 Outline Plasma Membrane Cytoplasm and Its Organelles Cell Nucleus and Gene Expression Protein Synthesis and Secretion DNA Synthesis and Cell Division
CHAPTER 3 Cell Structure and Genetic Control Chapter 3 Outline Plasma Membrane Cytoplasm and Its Organelles Cell Nucleus and Gene Expression Protein Synthesis and Secretion DNA Synthesis and Cell Division
Influence of Nuclear Substructure on Gene Regulation and Expression
 Influence of Nuclear Substructure on Gene Regulation and Expression Samuel Isaacson in collaboration with: Carolyn Larabell (UCSF/LBL), David McQueen (NYU), Charles Peskin (NYU) Why is stochasticity in
Influence of Nuclear Substructure on Gene Regulation and Expression Samuel Isaacson in collaboration with: Carolyn Larabell (UCSF/LBL), David McQueen (NYU), Charles Peskin (NYU) Why is stochasticity in
Self-calibrated, line-scan STED-FCS to quantify lipid dynamics in model and cell membranes
 Self-calibrated, line-scan STED-FCS to quantify lipid dynamics in model and cell membranes Aleš Benda, Yuanqing Ma and Katharina Gaus Centre for Vascular Research, Australian Centre for Nanomedicine and
Self-calibrated, line-scan STED-FCS to quantify lipid dynamics in model and cell membranes Aleš Benda, Yuanqing Ma and Katharina Gaus Centre for Vascular Research, Australian Centre for Nanomedicine and
Time-resolved Molecule Counting by Photon Statistics Across the Visible Spectrum
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Time-resolved Molecule Counting by Photon Statistics Across the Visible Spectrum
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Time-resolved Molecule Counting by Photon Statistics Across the Visible Spectrum
arxiv: v1 [q-bio.sc] 13 Sep 2016
![arxiv: v1 [q-bio.sc] 13 Sep 2016 arxiv: v1 [q-bio.sc] 13 Sep 2016](/thumbs/88/117897003.jpg) Sets of FCS experiments to quantify free diffusion coefficients in reaction-diffusion systems. The case of Ca 2+ and its dyes Lorena Sigaut, Cecilia Villarruel, María Laura Ponce and Silvina Ponce Dawson
Sets of FCS experiments to quantify free diffusion coefficients in reaction-diffusion systems. The case of Ca 2+ and its dyes Lorena Sigaut, Cecilia Villarruel, María Laura Ponce and Silvina Ponce Dawson
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Note. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Note File Name: Peer Revie File Description: Supplementary Fig.1. Complete set of ACFs extracted by FLCS analysis.
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Note File Name: Peer Revie File Description: Supplementary Fig.1. Complete set of ACFs extracted by FLCS analysis.
Bi 1x Spring 2014: LacI Titration
 Bi 1x Spring 2014: LacI Titration 1 Overview In this experiment, you will measure the effect of various mutated LacI repressor ribosome binding sites in an E. coli cell by measuring the expression of a
Bi 1x Spring 2014: LacI Titration 1 Overview In this experiment, you will measure the effect of various mutated LacI repressor ribosome binding sites in an E. coli cell by measuring the expression of a
Slide 1 / 7. Free Response
 Slide 1 / 7 Free Response Slide 2 / 7 Slide 3 / 7 1 The above diagrams illustrate the experiments carried out by Griffith and Hershey and Chaserespectively. Describe the hypothesis or conclusion that each
Slide 1 / 7 Free Response Slide 2 / 7 Slide 3 / 7 1 The above diagrams illustrate the experiments carried out by Griffith and Hershey and Chaserespectively. Describe the hypothesis or conclusion that each
High Resolution Laser Microscopy: a fascinating method to explore the molecular world
 High Resolution Laser Microscopy: a fascinating method to explore the molecular world Alfred J. Meixner Physical and Theoretical Chemistry Laboratory University of Siegen Single-molecule spectroscopy and
High Resolution Laser Microscopy: a fascinating method to explore the molecular world Alfred J. Meixner Physical and Theoretical Chemistry Laboratory University of Siegen Single-molecule spectroscopy and
Supplemental Data. Lermontova et al. (2013). Plant Cell /tpc
 Supplemental Figure 1. Multiple alignment of the N-terminal parts of plant KNL2 orthologs. Multiple sequence alignment was performed by the MUSCLE method and visualized by the JalView program. Highly conserved
Supplemental Figure 1. Multiple alignment of the N-terminal parts of plant KNL2 orthologs. Multiple sequence alignment was performed by the MUSCLE method and visualized by the JalView program. Highly conserved
STM: Scanning Tunneling Microscope
 STM: Scanning Tunneling Microscope Basic idea STM working principle Schematic representation of the sample-tip tunnel barrier Assume tip and sample described by two infinite plate electrodes Φ t +Φ s =
STM: Scanning Tunneling Microscope Basic idea STM working principle Schematic representation of the sample-tip tunnel barrier Assume tip and sample described by two infinite plate electrodes Φ t +Φ s =
I. Proteomics by Mass Spectrometry 1. What is an internal standard and what does it accomplish analytically?
 Name I. Proteomics by Mass Spectrometry 1. What is an internal standard and what does it accomplish analytically? Internal standards are standards added intentionally to all samples, standards and blanks.
Name I. Proteomics by Mass Spectrometry 1. What is an internal standard and what does it accomplish analytically? Internal standards are standards added intentionally to all samples, standards and blanks.
Transient kinetic methods. Biophysics seminars Kinga Futó
 Transient kinetic methods Biophysics seminars Kinga Futó Fast kinetics Reaction kinetics: temporal description of reactions reactionmechanisms reaction dynamics (processes on molecular level) rate of reaction
Transient kinetic methods Biophysics seminars Kinga Futó Fast kinetics Reaction kinetics: temporal description of reactions reactionmechanisms reaction dynamics (processes on molecular level) rate of reaction
Supplementary table I. Table of contact angles of the different solutions on the surfaces used here. Supplementary Notes
 1 Supplementary Figure 1. Sketch of the experimental setup (not to scale) : it consists of a thin mylar sheet (0, 02 4 3cm 3 ) held fixed vertically. The spacing y 0 between the glass plate and the upper
1 Supplementary Figure 1. Sketch of the experimental setup (not to scale) : it consists of a thin mylar sheet (0, 02 4 3cm 3 ) held fixed vertically. The spacing y 0 between the glass plate and the upper
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Epigenetics in Yeast. Dom Helmlinger CRBM, Montpellier
 Epigenetics in Yeast Dom Helmlinger CRBM, Montpellier Outline Genetic and epigenetic regulation of gene expression. Mating-type switching in budding yeast. Positive and negative regulation of mating-type
Epigenetics in Yeast Dom Helmlinger CRBM, Montpellier Outline Genetic and epigenetic regulation of gene expression. Mating-type switching in budding yeast. Positive and negative regulation of mating-type
Challenges of Single Ion-Channel Biosensors
 Vanderbilt Institute for Integrative Biosystems Research and Education Instrumenting and Control the Single Cell Challenges of Single Ion-Channel Biosensors Fredrick Sachs (SUNY Buffalo) John P. Wikswo
Vanderbilt Institute for Integrative Biosystems Research and Education Instrumenting and Control the Single Cell Challenges of Single Ion-Channel Biosensors Fredrick Sachs (SUNY Buffalo) John P. Wikswo
Introduction to FCS. Enrico Gratton. Laboratory for Fluorescence Dynamics Department of Biomedical Engineering University of California, Irvine
 Introduction to FCS Enrico Gratton Laboratory for Fluorescence Dynamics Department of Biomedical Engineering University of California, Irvine Outline What is diffusion? Diffusion of molecules Diffusion
Introduction to FCS Enrico Gratton Laboratory for Fluorescence Dynamics Department of Biomedical Engineering University of California, Irvine Outline What is diffusion? Diffusion of molecules Diffusion
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray
 CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
BME Engineering Molecular Cell Biology. Review: Basics of the Diffusion Theory. The Cytoskeleton (I)
 BME 42-620 Engineering Molecular Cell Biology Lecture 08: Review: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 08, September 22, 2011 1 Outline Background: FRAP & SPT Review: microscopic
BME 42-620 Engineering Molecular Cell Biology Lecture 08: Review: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 08, September 22, 2011 1 Outline Background: FRAP & SPT Review: microscopic
Measuring TF-DNA interactions
 Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
ECE280: Nano-Plasmonics and Its Applications. Week8
 ECE280: Nano-Plasmonics and Its Applications Week8 Surface Enhanced Raman Scattering (SERS) and Surface Plasmon Amplification by Stimulated Emission of Radiation (SPASER) Raman Scattering Chandrasekhara
ECE280: Nano-Plasmonics and Its Applications Week8 Surface Enhanced Raman Scattering (SERS) and Surface Plasmon Amplification by Stimulated Emission of Radiation (SPASER) Raman Scattering Chandrasekhara
Manipulating and Probing Enzymatic Conformational Fluctuations and Enzyme-Substrate Interactions by Single-Molecule FRET- Magnetic Tweezers Microscopy
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2014 Supporting Information (SI) Manipulating and Probing Enzymatic Conformational Fluctuations
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2014 Supporting Information (SI) Manipulating and Probing Enzymatic Conformational Fluctuations
Diffusion in biological systems and Life at low Reynolds number
 Diffusion in biological systems and Life at low Reynolds number Aleksandra Radenovic EPFL Ecole Polytechnique Federale de Lausanne Bioengineering Institute Laboratory of Nanoscale Biology Lausanne September
Diffusion in biological systems and Life at low Reynolds number Aleksandra Radenovic EPFL Ecole Polytechnique Federale de Lausanne Bioengineering Institute Laboratory of Nanoscale Biology Lausanne September
