HHS Public Access Author manuscript Science. Author manuscript; available in PMC 2016 December 24.
|
|
|
- Scott Robinson
- 5 years ago
- Views:
Transcription
1 Reprogramming of avian neural crest axial identity and cell fate Marcos Simoes-Costa 1 and Marianne E. Bronner 1 1 Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA 91125, USA Abstract HHS Public Access Author manuscript Published in final edited form as: Science June 24; 352(6293): doi: /science.aaf2729. Neural crest populations along the embryonic body axis differ in developmental potential and fate, such that only cranial neural crest can contribute to craniofacial skeleton in vivo. Here, we explore the regulatory program that imbues the cranial crest with its unique features. Using axial-level specific enhancers to isolate and perform genome-wide profiling of cranial versus trunk neural crest in chick embryos, we identify and characterize regulatory relationships between a set of cranial-specific transcription factors. Introducing components of this circuit into neural crest cells of the trunk alters their identity and endows these cells with the ability to give rise to chondroblasts in vivo. Our results demonstrate that gene regulatory circuits that support formation of particular neural crest derivatives may be employed for reprogramming specific neural crest derived cell types. Neural crest cells are characterized by multipotency and migratory ability. During embryonic development, the neural crest differentiates into multiple cell types, including chondrocytes and osteocytes, melanocytes, and neurons and glia of the peripheral nervous system (1,2). Neural crest stem cells are retained postnatally in skin and peripheral nerves, providing a potential target for replacement therapy in regenerative medicine (3 4). However, not all neural crest populations along the body axis are alike. Quail-chick grafting experiments demonstrated that the cranial and trunk neural crest differ in developmental potential: whereas the cranial neural crest forms much of the craniofacial skeleton, the trunk crest fails to contribute to skeletal lineages, even when grafted in vivo to the head (1). Approaches for engineering and replacement of specific cell types depend upon a better understanding of the molecular mechanisms that underlie establishment of specific cell types during embryonic development. Here we take advantage of differences in neural crest subpopulations (1,5 7) to identify the regulatory circuit that controls commitment of the cranial neural crest to a chondrocytic fate. Expression of neural crest specifier genes like FoxD3 and Sox10 is controlled by enhancers specific to particular axial levels, driving onset of their transcription in either the head or trunk neural crest but not both (8,9). The enhancers are activated by different inputs, * Correspondence to: mbronner@caltech.edu. Supplementary Materials: Materials and Methods Figures S1 S7. External Database S1. References (14 26)
2 Simoes-Costa and Bronner Page 2 suggesting that neural crest specification is driven by distinct genetic programs in different subpopulations (2,9). In order to identify the transcriptional program that endows the cranial neural crest with its ability to give rise to ectomesenchyme (cartilage and bone of the face), we utilized the FoxD3 enhancers NC1 and NC2 (9) (Fig. 1A C), active in the cranial and trunk neural crest, respectively, to isolate pure populations of neural crest cells for comparative transcriptional profiling. Embryos were electroporated with expression vectors containing GFP under the control of these enhancers and early-migrating cranial and trunk neural crest cells were obtained by fluorescence activated cell sorting (FACS) (10). RNA-seq analysis comparing these two populations identified 216 genes that were enriched in the cranial neural crest relative to the trunk (Fig. 1D E, Database S1), including 16 transcription factors (listed in Fig. 1F). We confirmed the expression of these regulators in the cranial neural crest by in situ hybridization; while 6 genes were expressed throughout the cranial neural crest (Fig. S1A F), the remainder were detected in specific subsets of cells (Fig. S1G L). Of these, we focused on the first group which were expressed in all cranial neural crest cells, including Brain-Specific Homeobox Protein 3C (Brn3c), LIM Homeobox Protein 5 (Lhx5), Diencephalon/Mesencephalon Homeobox 1 (Dmbx1), Transcription Factor AP-2 Beta (Tfap2b), SRY Box 8 (Sox8) and the V-Ets Avian Erythroblastosis Virus E26 Oncogene Homolog 1 (Ets1). Analysis of the spatiotemporal expression of these cranial-specific regulators demonstrated that Brn3c, Lhx5 and Dmbx1 were first detected in the anterior regions of gastrula-stage embryos at Hamburger Hamilton (HH) stage 4, and persisted through stages of neural crest specification (Fig. 2A, E). These early cranial-specific genes were down-regulated after the neural crest delaminated from the neural tube. Onset of Tfap2b, Sox8 and Ets1 expression was observed later at HH7 and HH8, in neural crest progenitors residing within the cranial neural folds (Fig. 2B, E). These genes were maintained in the migratory neural crest cells during later stages of development (HH10-14; Fig. 2B, E). Co-localization of neural plate border markers Msh Homeobox 1 (Msx1) or Paired Box 7 (Pax7) with Brn3c, Lhx5 and Dmbx1 showed that the latter are expressed by an anterior subset of the neural crest progenitors (Fig. 2C). To verify that the cranial regulators mark the territory that contains cranial neural crest precursors, we analyzed the fate map of the neural plate border at the 3- somite stage using focal injections of a vital lipophilic dye (CM-DiI) to label cells along the anterior-posterior axis. The injected embryos were cultured until the 12-somite stage (HH11) when the labeled cellular progeny were scored with respect to their fate as cranial or vagal/ trunk neural crest cells. The results show that the domain of expression of the early regulators (Brn3c, Lhx5 and Dmbx1) demarcates the territory that contains the progenitors of the cranial neural crest in the early neurula (HH8-) (Fig. 2D). We asked whether cranial-specific regulators are part of a transcriptional circuit that underlies cranial identity by knocking down each regulator individually and assaying for changes in expression levels of the other five genes. This was done via bilateral electroporations (11), with control transfections on the left side and function-blocking morpholinos or dominant negative constructs on the right side of the same embryo (Fig. 3A D). Transfected embryos were analyzed by in situ hybridization (Fig. 3A H) and qpcr (Fig. 3I) for effects on putative targets; loss of the target gene in the experimental side of the embryo indicated the existence of a regulatory link between the two transcription factors.
3 Simoes-Costa and Bronner Page 3 The diagram on Figure 3J contains interactions confirmed by both qpcr and in situ hybridization. Morpholino knockdown efficiency was validated by in vivo translationblocking assays (Fig. S2). By testing 25 putative regulatory links, we found that the early and late cranial-specific genes constitute different hierarchical levels of a gene regulatory network (Fig. 3J). Brn3c, which is placed at the top of the circuit, is necessary for the activation of Dmbx1 in the anterior neural plate border (Fig. 3I; Fig. S3). Subsequently, Lhx5 and Dmbx1 drive expression of Tfap2b and Sox8 in the dorsal neural folds (Fig. 3A C, E G, I). Finally, Tfap2b activates expression of Ets1 as the neural crest becomes specified (Fig 3D, HI). Chromatin immunoprecipitation (ChIP) experiments performed in microdissected neural crest cells showed association of the cranial-specific transcription factors with promoters of predicted downstream target genes, suggesting that these regulatory links are direct (Fig. 3K; Fig. S4). Sox9, Tfap2b and Ets1 are all retained in the migrating cranial crest cells as they move ventrally to give rise to the facial mesenchyme during stages HH To investigate if these genes play a role in the differentiation of neural crest cells into chondroblasts, we assayed the effects of disrupting the terminal module of the circuit on the expression of markers of chondrocytic differentiation. We found that Ets1 was required for the expression of ALX Homeobox 1 (Alx1, also known as Cartilage Paired-Class Homeoprotein 1) in the facial mesenchyme (Fig. S5), indicating a link between cranial identity and chondrocytic differentiation (Fig. S5). Thus, cranial-specific regulators are part of a transcriptional circuit that conveys regulatory information from the anterior neural plate border to the late migratory neural crest. To test if this cranial-specific regulatory circuit could be used to manipulate neural crest identity, we utilized neural crest axial-specific enhancers as reporters of axial level identity. We electroporated expression constructs in the trunk neural tube of stage HH10 embryos, and found that transfection of the late cranial-specific factors (Sox8, Tfap2b and Ets1) robustly activated the cranial enhancer Sox10E2 (9, 11) in the trunk neural crest (n=15/15) (Fig. 4A E), consistent with a shift from trunk to cranial identity. Other axial-specific enhancers were similarly affected; trunk specific enhancers NC2 and Sox10E1 were repressed after electroporation of the late factors (Fig. S6). Early cranial-specific factors (Brn3c, Lhx5 and Dmbx1), or individual late factors, were unable to activate the cranial enhancer in the trunk. To identify changes in the regulatory state of the reprogrammed trunk neural crest, we isolated transfected cells by FACS, and analyzed their expression profile by qpcr, focusing on transcription factors involved in craniofacial differentiation. The results revealed elevated expression of chondrocytic genes Runt-Related Transcription Factor 2 (Runx2) and Alx1, in the trunk Sox10E2+ cells compared with native trunk crest (Fig. 4F). The reprogrammed trunk Sox10E2+ cells also displayed loss of genes enriched in the trunk neural crest like Developing Brain Homeobox 1 (Dbx2) and Hairy And Enhancer Of Split 6 (Hes6) (Fig. 4G). This confirmed that the reprogrammed cells adopt a cranial-like expression profile, and raised the possibility that these cells might display augmented chondrocytic potential.
4 Simoes-Costa and Bronner Page 4 Finally, we tested whether this cranial neural crest circuit could reprogram not only enhancer activity and expression of axial-specific neural crest genes, but also cell fate such that reprogrammed trunk neural crest cells could differentiate into craniofacial cartilage. We cotransfected three constructs encoding late transcription factors Sox8, Tfap2b and Ets1 into the posterior epiblast of HH5 chicken embryos transgenic for GFP by electroporating DNA in the region posterior to the Hensen s node. The GFP+ trunk neural folds then were microdissected at HH11 and immediately transplanted to the cranial regions of HH9 wildtype chick embryos (Fig. S7). The grafted embryos were incubated until host embryonic day (E)7, by which time endogenous cartilage cells have differentiated. The fate of donor tissue was assayed using markers for neuronal, melanocytic and chondrocytic differentiation. By E7, wild type (host) and reprogrammed (donor) trunk neural crest migrated to the proximal part of the jaw. As observed with chick-quail chimeras (5 7), we found that wild type or mock-transfected trunk neural crest cells grafted into the cephalic region gave rise to neurons and melanocytes, but were unable to differentiate into chondroblasts (n=0/5) (Fig. 4H K). The same was observed with trunk neural crest transfected with the early cranialspecific factors (n=0/6). Reprogrammed trunk neural crest, however, acquired chondrogenic potential and formed ectopic cartilage nodules (n=4/7) (Fig. 4L O) in the proximal jaw. Thus, introducing components of the cranial-specific transcriptional circuit is sufficient to reprogram trunk neural crest and to drive them to adopt an additional cartilaginous fate. These results definitively show that the cranial-specific regulatory circuit (Fig. 3J) we have defined confers chondrocytic potential to the trunk neural crest. The development and differentiation of neural crest cells is controlled by a complex gene regulatory network, composed of transcription factors, signaling molecules and epigenetic modifiers (12 13). Here, we have expanded the cranial neural crest gene regulatory network by identifying transcriptional interactions specific to the cranial crest and absent from other subpopulations. By linking anterior identity in the gastrula to the expression of drivers of chondrocytic differentiation, we have identified a cranial-specific circuit (Fig. 3J) that endows the neural crest with its unique potential to differentiate into the craniofacial skeleton of vertebrates. Our results highlight how transcriptional circuits can be rewired to alter progenitor cell identity and fate during embryonic development. Supplementary Material Acknowledgments Refer to Web version on PubMed Central for supplementary material. We thank Joanna Tan-Cabugao, Michael Stone, Brian Jun and Daniel S. E. Koo for technical assistance. The Caltech Millard and Muriel Jacobs Genetics and Genomics Laboratory provided sequencing and bioinformatics support. We are indebted to Diana Perez, Keith Beadle, and Rochelle Diamond for cell-sorting assistance at the Caltech Flow Cytometry Cell Sorting Facility. We also thank Meyer Barembaum for the Sox8 and Tfap2b expression constructs. This work was supported by NIH grants DE and HD to MEB. MS-C was funded by a fellowship from the Pew Fellows Program in Biomedical Sciences and by NIH grant K99DE Supplement contains additional data. References and Notes 1. Le Douarin, N. The neural crest. Cambridge University Press; 1982.
5 Simoes-Costa and Bronner Page 5 2. Simoes-Costa M, Bronner ME. Insights into neural crest development and evolution from genomic analysis. Genome research. 2013; 23: [PubMed: ] 3. Uesaka T, Nagashimada M, Enomoto H. Neuronal Differentiation in Schwann Cell Lineage Underlies Postnatal Neurogenesis in the Enteric Nervous System. J Neurosci. 2015; 35: [PubMed: ] 4. Dyachuk V, et al. Parasympathetic neurons originate from nerve associated peripheral glial progenitors. Science. 2014; 345: [PubMed: ] 5. Le Lievre CS, Le Douarin NM. Mesenchymal derivatives of the neural crest: analysis of chimaeric quail and chick embryos. Journal of embryology and experimental morphology. 1975; 34: [PubMed: ] 6. Le Lievre CS, Schweizer GG, Ziller CM, Le Douarin NM. Restrictions of developmental capabilities in neural crest cell derivatives as tested by in vivo transplantation experiments. Developmental biology. 1980; 77: [PubMed: ] 7. Lwigale PY, Conrad GW, Bronner-Fraser M. Graded potential of neural crest to form cornea, sensory neurons and cartilage along the rostrocaudal axis. Development. 2004; 131: [PubMed: ] 8. Betancur P, Bronner-Fraser M, Sauka-Spengler T. Genomic code for Sox10 activation reveals a key regulatory enhancer for cranial neural crest. Proceedings of the National Academy of Sciences of the United States of America. 2010; 107: [PubMed: ] 9. Simoes-Costa MS, McKeown SJ, Tan-Cabugao J, Sauka-Spengler T, Bronner ME. Dynamic and differential regulation of stem cell factor FoxD3 in the neural crest is Encrypted in the genome. PLoS genetics. 2012; 8:e [PubMed: ] 10. Simoes-Costa M, Tan-Cabugao J, Antoshechkin I, Sauka-Spengler T, Bronner ME. Transcriptome analysis reveals novel players in the cranial neural crest gene regulatory network. Genome research. 2014; 24: [PubMed: ] 11. Simoes-Costa M, Stone M, Bronner ME. Axud1 Integrates Wnt Signaling and Transcriptional Inputs to Drive Neural Crest Formation. Developmental cell. 2015; 35(5): Simoes-Costa M, Bronner ME. Establishing neural crest identity: a gene regulatory recipe. Development. 2015; 142: [PubMed: ] 13. Green SA, Simoes-Costa M, Bronner ME. Evolution of vertebrates as viewed from the crest. Nature. 2015; 520: [PubMed: ] 14. Sauka-Spengler T, Barembaum M. Gain- and loss-of-function approaches in the chick embryo. Methods in cell biology. 2008; 87: [PubMed: ] 15. Chapman SC, Collignon J, Schoenwolf GC, Lumsden A. Improved method for chick wholeembryo culture using a filter paper carrier. Developmental dynamics. 2001; 220: [PubMed: ] 16. Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009; 25: [PubMed: ] 17. Trapnell C, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nature biotechnology. 2010; 28:511 U Theveneau E, Duband JL, Altabef M. Ets-1 confers cranial features on neural crest delamination. PloS one. 2007; 2:e1142. [PubMed: ] 19. Wilkinson, DG. In situ hybridization : a practical approach. IRL Press at Oxford University Press; Denkers N, Garcia-Villalba P, Rodesch CK, Nielson KR, Mauch TJ. FISHing for chick genes: Triple-label whole-mount fluorescence in situ hybridization detects simultaneous and overlapping gene expression in avian embryos. Developmental dynamics. 2004; 229: [PubMed: ] 21. Ezin AM, Fraser SE, Bronner-Fraser M. Fate map and morphogenesis of presumptive neural crest and dorsal neural tube. Developmental biology. 2009; 330: [PubMed: ] 22. Kim DS, Matsuda T, Cepko CL. A core paired-type and POU homeodomain containing transcription factor program drives retinal bipolar cell gene expression. Journal of Neuroscience. 2008; 28: [PubMed: ]
6 Simoes-Costa and Bronner Page Kim J, Cantor AB, Orkin SH, Wang J. Use of in vivo biotinylation to study protein protein and protein DNA interactions in mouse embryonic stem cells. Nature Protocols. 2009; 4: [PubMed: ] 24. Kolodziej KE, Pourfarzad F, de Boer E, Krpic S, Grosveld F, Strouboulis J. Optimal use of tandem biotin and V5 tags in ChIP assay. BMC Molecular Biology. 2009; 10:6. [PubMed: ] 25. Nakamura H, Funahashi J. Introduction of DNA into chick embryos by in ovo electroporation. Methods. 2001; 24: [PubMed: ] 26. Barembaum M, Bronner ME. Identification and dissection of a key enhancer mediating cranial neural crest specific expression of transcription factor, Ets-1. Developmental Biology. 2013; 382: [PubMed: ]
7 Simoes-Costa and Bronner Page 7 Figure 1. Identification of cranial-specific regulators by comparative transcriptomics (A) Diagram depicting dorsal view of chick embryo. (B, C) Embryos electroporated with FoxD3 axial-specific enhancers NC1 and NC2, active in cranial and trunk neural crest (NC), respectively. (D) Comparative transcriptome analysis of FACS sorted cranial and trunk neural crest populations identified 216 cranially-enriched genes. (E) Summary of gene ontology analysis for the cranial-enriched genes. (F) Enrichment levels of transcription factors expressed in the cranial neural crest. HH: Hamburger and Hamilton stages.
8 Simoes-Costa and Bronner Page 8 Figure 2. Spatial and temporal expression of cranial neural crest transcription factors (TFs) (A, B) Dorsal views of embryos after in situ hybridization for cranial neural crest-specific TFs reveals expression at early (A) or later (B) stages. (C) Double in situ hybridization reveals that cranial regulator Dmbx1 is expressed in the anterior neural plate border, whereas Msx1 is expressed along the entire neural axis. (D) Fate map of neural crest progenitors (red and green dots) at stage HH8- confirms cranial specific expression of Dmbx1 (purple). (E) Diagram summarizing timing of expression of early and late cranial regulators.
9 Simoes-Costa and Bronner Page 9 Figure 3. Cranial-specific transcriptional circuit underlying neural crest axial identity (A D) Whole mount dorsal views of embryos after morpholino (Mo) targeted to indicated transcription factor (TF) was transfected to the right side (green) and control morpholino (CoMo) to the left side (blue) of each embryo. (E H) Same embryos as above after in situ hybridization for indicated downstream TF; blue arrows indicate transcript downregulation. (I) Comparing control to loss of function (LOF) neural folds by qpcr reveals differential regulation of downstream targets with significant changes indicated by * (Student s t-test, P<0.05). (J) Diagram summarizing cranial specific gene regulatory circuit delineated by functional assays. (K) Chromatin immunoprecipitation (ChIP) demonstrates direct association of cranial specific TFs with promoters of their downstream targets. No enrichment was observed for intergenic negative control regions (NCR), or when the procedure was performed with mock-transfected embryos (see Materials and Methods for more information). Error bars on (I) and (K) represent standard deviation.
10 Simoes-Costa and Bronner Page 10 Figure 4. Reprogramming of neural crest axial identity and fate (A) Diagram of chick embryo electroporated with the Sox10E2 enhancer, expressed only in migrating cranial (Cr) neural crest (NC). (B) Transfection of trunk neural crest cells with control RFP expression vector shows no Sox10E2 expression. (C) Reprogramming of trunk neural crest cells with cranial specific regulators Sox8, Tfap2b and Ets1 results in robust expression of the cranial Sox10E2 enhancer in the trunk region (n=15/15). (D, E) Flow cytometry analysis of dissociated embryonic trunks shows a large number of Sox10E2+ trunk neural crest cells after reprogramming. (F, G) Reprogrammed (Rep) trunk neural crest display increased expression of the chondrocytic genes Runx2 and Alx1, while trunk genes Dbx2 and Hes6 are strongly downregulated. Error bars represent standard deviation. (H, I) Histological sections of E7 embryonic heads show that wild type (WT) trunk neural crest cells (GFP+; green) fail to form cartilage (Col9a+; red) after transplantation to the cranial region (n=0/5). (J, K) Inset of (H I), showing absence of GFP+ chondrocytes. (L, M) Reprogrammed trunk neural crest cells (expressing Sox8, Tfap2b and Ets1) from GFP donor embryos transplanted to the head form ectopic cartilage nodules. (N, O) Inset of (L M), showing chondrocytes derived from trunk neural crest cells (GFP+ and Col9a+).
Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome
 Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome Marcos S. Simões-Costa 1., Sonja J. McKeown 1., Joanne Tan-Cabugao 1, Tatjana Sauka-Spengler
Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome Marcos S. Simões-Costa 1., Sonja J. McKeown 1., Joanne Tan-Cabugao 1, Tatjana Sauka-Spengler
NIH Public Access Author Manuscript Histochem Cell Biol. Author manuscript; available in PMC 2013 August 01.
 NIH Public Access Author Manuscript Published in final edited form as: Histochem Cell Biol. 2012 August ; 138(2): 179 186. doi:10.1007/s00418-012-0999-z. Formation and migration of neural crest cells in
NIH Public Access Author Manuscript Published in final edited form as: Histochem Cell Biol. 2012 August ; 138(2): 179 186. doi:10.1007/s00418-012-0999-z. Formation and migration of neural crest cells in
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
NIH Public Access Author Manuscript Dev Biol. Author manuscript; available in PMC 2014 October 15.
 NIH Public Access Author Manuscript Published in final edited form as: Dev Biol. 2013 October 15; 382(2):. doi:10.1016/j.ydbio.2013.08.009. Identification and dissection of a key enhancer mediating cranial
NIH Public Access Author Manuscript Published in final edited form as: Dev Biol. 2013 October 15; 382(2):. doi:10.1016/j.ydbio.2013.08.009. Identification and dissection of a key enhancer mediating cranial
Bio Section III Organogenesis. The Neural Crest and Axonal Specification. Student Learning Objectives. Student Learning Objectives
 Bio 127 - Section III Organogenesis The Neural Crest and Axonal Specification Gilbert 9e Chapter 10 Student Learning Objectives 1. You should understand that the neural crest is an evolutionary advancement
Bio 127 - Section III Organogenesis The Neural Crest and Axonal Specification Gilbert 9e Chapter 10 Student Learning Objectives 1. You should understand that the neural crest is an evolutionary advancement
The role of FGF2 in craniofacial skeletogenesis
 The role of FGF2 in craniofacial skeletogenesis P. Ferretti, S. Sarkar, R. Moore, A. Petiot, C. J. Chan and A. Copp Summary E vidence that the major craniosynostosis syndromes are caused by mutations in
The role of FGF2 in craniofacial skeletogenesis P. Ferretti, S. Sarkar, R. Moore, A. Petiot, C. J. Chan and A. Copp Summary E vidence that the major craniosynostosis syndromes are caused by mutations in
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
HHS Public Access Author manuscript Curr Top Dev Biol. Author manuscript; available in PMC 2016 November 08.
 The neural crest migrating into the 21 st century Marianne E. Bronner and Marcos Simões-Costa Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA 91125, USA
The neural crest migrating into the 21 st century Marianne E. Bronner and Marcos Simões-Costa Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA 91125, USA
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
HHS Public Access Author manuscript Differentiation. Author manuscript; available in PMC 2016 July 22.
 Stage-dependent plasticity of the anterior neural folds to form olfactory placode versus neural crest Maxellende Ezin, Meyer Barembaum, and Marianne E. Bronner Division of Biology and Biological Engineering,
Stage-dependent plasticity of the anterior neural folds to form olfactory placode versus neural crest Maxellende Ezin, Meyer Barembaum, and Marianne E. Bronner Division of Biology and Biological Engineering,
Role of Organizer Chages in Late Frog Embryos
 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Neural crest development is regulated by the transcription factor Sox9
 Research article 5681 Neural crest development is regulated by the transcription factor Sox9 Martin Cheung and James Briscoe* Developmental Neurobiology, National Institute for Medical Research, Mill Hill,
Research article 5681 Neural crest development is regulated by the transcription factor Sox9 Martin Cheung and James Briscoe* Developmental Neurobiology, National Institute for Medical Research, Mill Hill,
Biology 218, practise Exam 2, 2011
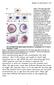 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Name. Biology Developmental Biology Winter Quarter 2013 KEY. Midterm 3
 Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Sonic hedgehog (Shh) signalling in the rabbit embryo
 Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
10/2/2015. Chapter 4. Determination and Differentiation. Neuroanatomical Diversity
 Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
In ovo time-lapse analysis after dorsal neural tube ablation shows rerouting of chick hindbrain neural crest
 In ovo time-lapse analysis after dorsal neural tube ablation shows rerouting of chick hindbrain neural crest Paul Kulesa, Marianne Bronner-Fraser and Scott Fraser (2000) Presented by Diandra Lucia Background
In ovo time-lapse analysis after dorsal neural tube ablation shows rerouting of chick hindbrain neural crest Paul Kulesa, Marianne Bronner-Fraser and Scott Fraser (2000) Presented by Diandra Lucia Background
Geert Geeven. April 14, 2010
 iction of Gene Regulatory Interactions NDNS+ Workshop April 14, 2010 Today s talk - Outline Outline Biological Background Construction of Predictors The main aim of my project is to better understand the
iction of Gene Regulatory Interactions NDNS+ Workshop April 14, 2010 Today s talk - Outline Outline Biological Background Construction of Predictors The main aim of my project is to better understand the
Elucidation of the Evolutionary Origin of the Neural Crest. William K. Nesbitt
 Elucidation of the Evolutionary Origin of the Neural Crest William K. Nesbitt Under the supervision of Dr. David McClay, Department of Biology, Duke University April 22, 2016 Research Supervisor Faculty
Elucidation of the Evolutionary Origin of the Neural Crest William K. Nesbitt Under the supervision of Dr. David McClay, Department of Biology, Duke University April 22, 2016 Research Supervisor Faculty
9/4/2015 INDUCTION CHAPTER 1. Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology. Fig 1.
 INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Origin of mesenchymal stem cell
 Origin of mesenchymal stem cell Takumi Era Department of Cell Modulation, Institute of Molecular embryology and Genetics (IMEG) Kumamoto University Differentiation pathways of in vitro ES cell culture
Origin of mesenchymal stem cell Takumi Era Department of Cell Modulation, Institute of Molecular embryology and Genetics (IMEG) Kumamoto University Differentiation pathways of in vitro ES cell culture
Unit 4 Evaluation Question 1:
 Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
10/15/09. Tetrapod Limb Development & Pattern Formation. Developing limb region is an example of a morphogenetic field
 Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Questions in developmental biology. Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration
 Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
3/8/ Complex adaptations. 2. often a novel trait
 Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Formation of the Cortex
 Formation of the Cortex Neuronal Birthdating with 3 H-thymidine 3H-thymidine is incorporated into the DNA during the S-phase (replication of DNA). It marks all mitotic cells Quantitative technique. (you
Formation of the Cortex Neuronal Birthdating with 3 H-thymidine 3H-thymidine is incorporated into the DNA during the S-phase (replication of DNA). It marks all mitotic cells Quantitative technique. (you
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
Name KEY. Biology Developmental Biology Winter Quarter Midterm 3 KEY
 Name KEY 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2009 Midterm 3 KEY All of the 25 multi-choice questions are single-answer. Choose the best answer. (4 pts each) Place
Name KEY 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2009 Midterm 3 KEY All of the 25 multi-choice questions are single-answer. Choose the best answer. (4 pts each) Place
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Drosophila melanogaster- Morphogen Gradient
 NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NIH Public Access Author Manuscript Dev Biol. Author manuscript; available in PMC 2014 February 15.
 NIH Public Access Author Manuscript Published in final edited form as: Dev Biol. 2013 February 15; 374(2): 255 263. doi:10.1016/j.ydbio.2012.12.009. Elk3 is essential for the progression from progenitor
NIH Public Access Author Manuscript Published in final edited form as: Dev Biol. 2013 February 15; 374(2): 255 263. doi:10.1016/j.ydbio.2012.12.009. Elk3 is essential for the progression from progenitor
Mesoderm Development
 Quiz rules: Spread out across available tables No phones, text books, or (lecture) notes on your desks No consultation with your colleagues No websites open other than the Quiz page No screen snap shots
Quiz rules: Spread out across available tables No phones, text books, or (lecture) notes on your desks No consultation with your colleagues No websites open other than the Quiz page No screen snap shots
Developmental Biology Biology Ectodermal Organs. November 22, 2005
 Developmental Biology Biology 4361 Ectodermal Organs November 22, 2005 Germinal neuroepithelium external limiting membrane neural tube neuroepithelium (stem cells) Figure 13.3 Figure 13.4 Neuroepithelial
Developmental Biology Biology 4361 Ectodermal Organs November 22, 2005 Germinal neuroepithelium external limiting membrane neural tube neuroepithelium (stem cells) Figure 13.3 Figure 13.4 Neuroepithelial
Bi 117 Final (60 pts) DUE by 11:00 am on March 15, 2012 Box by Beckman Institute B9 or to a TA
 Bi 117 Final (60 pts) DUE by 11:00 am on March 15, 2012 Box by Beckman Institute B9 or to a TA Instructor: Marianne Bronner Exam Length: 6 hours plus one 30-minute break at your discretion. It should take
Bi 117 Final (60 pts) DUE by 11:00 am on March 15, 2012 Box by Beckman Institute B9 or to a TA Instructor: Marianne Bronner Exam Length: 6 hours plus one 30-minute break at your discretion. It should take
Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio
 Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Graded potential of neural crest to form cornea, sensory neurons and cartilage along the rostrocaudal axis
 Research article 1979 Graded potential of neural crest to form cornea, sensory neurons and cartilage along the rostrocaudal axis Peter Y. Lwigale 1, Gary W. Conrad 2 and Marianne Bronner-Fraser 1, * 1
Research article 1979 Graded potential of neural crest to form cornea, sensory neurons and cartilage along the rostrocaudal axis Peter Y. Lwigale 1, Gary W. Conrad 2 and Marianne Bronner-Fraser 1, * 1
presumptiv e germ layers during Gastrulatio n and neurulation Somites
 Vertebrate embryos are similar at the phylotypic stage Patterning the Vertebrate Body Plan II: Mesoderm & Early Nervous System Wolpert L, Beddington R, Jessell T, Lawrence P, Meyerowitz E, Smith J. (2001)
Vertebrate embryos are similar at the phylotypic stage Patterning the Vertebrate Body Plan II: Mesoderm & Early Nervous System Wolpert L, Beddington R, Jessell T, Lawrence P, Meyerowitz E, Smith J. (2001)
Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural crest to definitive roof plate
 Washington University School of Medicine Digital Commons@Becker Open Access Publications 2016 Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural crest
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2016 Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural crest
Principles of Experimental Embryology
 Biology 4361 Developmental Biology Principles of Experimental Embryology June 16, 2008 Overview What forces affect embryonic development? The embryonic environment: external and internal How do forces
Biology 4361 Developmental Biology Principles of Experimental Embryology June 16, 2008 Overview What forces affect embryonic development? The embryonic environment: external and internal How do forces
Developmental Zoology. Ectodermal derivatives (ZOO ) Developmental Stages. Developmental Stages
 Developmental Zoology (ZOO 228.1.0) Ectodermal derivatives 1 Developmental Stages Ø Early Development Fertilization Cleavage Gastrulation Neurulation Ø Later Development Organogenesis Larval molts Metamorphosis
Developmental Zoology (ZOO 228.1.0) Ectodermal derivatives 1 Developmental Stages Ø Early Development Fertilization Cleavage Gastrulation Neurulation Ø Later Development Organogenesis Larval molts Metamorphosis
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
NIH Public Access Author Manuscript Dev Biol. Author manuscript; available in PMC 2012 January 15.
 NIH Public Access Author Manuscript Published in final edited form as: Dev Biol. 2011 January 15; 349(2): 238 249. doi:10.1016/j.ydbio.2010.10.032. Early regulative ability of the neuroepithelium to form
NIH Public Access Author Manuscript Published in final edited form as: Dev Biol. 2011 January 15; 349(2): 238 249. doi:10.1016/j.ydbio.2010.10.032. Early regulative ability of the neuroepithelium to form
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Generation of paraxial mesoderm from the H7 hesc line.
 Supplementary Figure 1 Generation of paraxial mesoderm from the H7 hesc line. H7 hescs were differentiated as shown in Figure 1a. (a) Flow cytometric analyses of the proportion of CD56+, PDGFRα+, and KDR+
Supplementary Figure 1 Generation of paraxial mesoderm from the H7 hesc line. H7 hescs were differentiated as shown in Figure 1a. (a) Flow cytometric analyses of the proportion of CD56+, PDGFRα+, and KDR+
Relationship between spatially restricted Krox-20 gene expression in branchial neural crest and
 The EMBO Journal vol.14 no.8 pp.1697-1710, 1995 Relationship between spatially restricted Krox-20 gene expression in branchial neural crest and segmentation in the chick embryo hindbrain M.Angela Nieto1
The EMBO Journal vol.14 no.8 pp.1697-1710, 1995 Relationship between spatially restricted Krox-20 gene expression in branchial neural crest and segmentation in the chick embryo hindbrain M.Angela Nieto1
A New Mechanistic Scenario for the Origin and Evolution of Vertebrate Cartilage
 A New Mechanistic Scenario for the Origin and Evolution of Vertebrate Cartilage Maria Cattell 1, Su Lai 1, Robert Cerny 2, Daniel Meulemans Medeiros 1 * 1 Department of Ecology and Evolutionary Biology,
A New Mechanistic Scenario for the Origin and Evolution of Vertebrate Cartilage Maria Cattell 1, Su Lai 1, Robert Cerny 2, Daniel Meulemans Medeiros 1 * 1 Department of Ecology and Evolutionary Biology,
Analysis of cranial neural crest migratory pathways in axolotl using cell markers and transplantation
 Development 127, 2751-2761 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV4304 2751 Analysis of cranial neural crest migratory pathways in axolotl using cell markers and transplantation
Development 127, 2751-2761 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV4304 2751 Analysis of cranial neural crest migratory pathways in axolotl using cell markers and transplantation
Chapter 2 Embryological Origins and the Identification of Neural Crest Cells
 Chapter 2 Embryological Origins and the Identification of Neural Crest Cells Neural Crest This chapter seeks to answer a twofold question: When and by what mechanisms do the neural crest (NC) and neural
Chapter 2 Embryological Origins and the Identification of Neural Crest Cells Neural Crest This chapter seeks to answer a twofold question: When and by what mechanisms do the neural crest (NC) and neural
Cell Migration I: Neural Crest Cell Migration. Steven McLoon Department of Neuroscience University of Minnesota
 Cell Migration I: Neural Crest Cell Migration Steven McLoon Department of Neuroscience University of Minnesota 1 Types of Cell Movement passive: active: cell sheets flow cilia or flagella ameboid adhesion
Cell Migration I: Neural Crest Cell Migration Steven McLoon Department of Neuroscience University of Minnesota 1 Types of Cell Movement passive: active: cell sheets flow cilia or flagella ameboid adhesion
The Radiata-Bilateria split. Second branching in the evolutionary tree
 The Radiata-Bilateria split Second branching in the evolutionary tree Two very important characteristics are used to distinguish between the second bifurcation of metazoans Body symmetry Germinal layers
The Radiata-Bilateria split Second branching in the evolutionary tree Two very important characteristics are used to distinguish between the second bifurcation of metazoans Body symmetry Germinal layers
Neural crest placode interactions mediate trigeminal ganglion formation and require Slit1 Robo2 signaling
 C h a p t e r 2 Neural crest placode interactions mediate trigeminal ganglion formation and require Slit1 Robo2 signaling 2.1 Abstract 1 The cellular and molecular interactions allowing generation of complex
C h a p t e r 2 Neural crest placode interactions mediate trigeminal ganglion formation and require Slit1 Robo2 signaling 2.1 Abstract 1 The cellular and molecular interactions allowing generation of complex
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis
 18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Building the brain (1): Evolutionary insights
 Building the brain (1): Evolutionary insights Historical considerations! Initial insight into the general role of the brain in human behaviour was already attained in antiquity and formulated by Hippocrates
Building the brain (1): Evolutionary insights Historical considerations! Initial insight into the general role of the brain in human behaviour was already attained in antiquity and formulated by Hippocrates
Measuring TF-DNA interactions
 Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Making Headway: The Roles of Hox Genes and Neural Crest Cells in Craniofacial Development
 Mini-Review TheScientificWorldJOURNAL (2003) 3, 240 264 ISSN 1537-744X; DOI 10.1100/tsw.2003.11 Making Headway: The Roles of Hox Genes and Neural Crest Cells in Craniofacial Development Paul A Trainor
Mini-Review TheScientificWorldJOURNAL (2003) 3, 240 264 ISSN 1537-744X; DOI 10.1100/tsw.2003.11 Making Headway: The Roles of Hox Genes and Neural Crest Cells in Craniofacial Development Paul A Trainor
Segment boundary formation in Drosophila embryos
 Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Embryonic Schwann cell development: the biology of Schwann cell precursors and early Schwann cells
 J. Anat. (1997) 191, pp. 501 505, with 2 figures Printed in the United Kingdom 501 Review Embryonic Schwann cell development: the biology of Schwann cell precursors and early Schwann cells K. R. JESSEN
J. Anat. (1997) 191, pp. 501 505, with 2 figures Printed in the United Kingdom 501 Review Embryonic Schwann cell development: the biology of Schwann cell precursors and early Schwann cells K. R. JESSEN
NEURAL CREST SPECIFICATION: MIGRATING INTO GENOMICS
 NEURAL CREST SPECIFICATION: MIGRATING INTO GENOMICS Laura S. Gammill and Marianne Bronner-Fraser The bones in your face, the pigment in your skin and the neural circuitry that controls your digestive tract
NEURAL CREST SPECIFICATION: MIGRATING INTO GENOMICS Laura S. Gammill and Marianne Bronner-Fraser The bones in your face, the pigment in your skin and the neural circuitry that controls your digestive tract
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
LIST of SUPPLEMENTARY MATERIALS
 LIST of SUPPLEMENTARY MATERIALS Mir et al., Dense Bicoid Hubs Accentuate Binding along the Morphogen Gradient Supplemental Movie S1 (Related to Figure 1). Movies corresponding to the still frames shown
LIST of SUPPLEMENTARY MATERIALS Mir et al., Dense Bicoid Hubs Accentuate Binding along the Morphogen Gradient Supplemental Movie S1 (Related to Figure 1). Movies corresponding to the still frames shown
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
From DNA to Diversity
 From DNA to Diversity Molecular Genetics and the Evolution of Animal Design Sean B. Carroll Jennifer K. Grenier Scott D. Weatherbee Howard Hughes Medical Institute and University of Wisconsin Madison,
From DNA to Diversity Molecular Genetics and the Evolution of Animal Design Sean B. Carroll Jennifer K. Grenier Scott D. Weatherbee Howard Hughes Medical Institute and University of Wisconsin Madison,
Developmental processes Differential gene expression Introduction to determination The model organisms used to study developmental processes
 Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Genes, Development, and Evolution
 14 Genes, Development, and Evolution Chapter 14 Genes, Development, and Evolution Key Concepts 14.1 Development Involves Distinct but Overlapping Processes 14.2 Changes in Gene Expression Underlie Cell
14 Genes, Development, and Evolution Chapter 14 Genes, Development, and Evolution Key Concepts 14.1 Development Involves Distinct but Overlapping Processes 14.2 Changes in Gene Expression Underlie Cell
Zebrafish foxd3 is selectively required for neural crest specification, migration and survival
 Developmental Biology 292 (2006) 174 188 www.elsevier.com/locate/ydbio Zebrafish foxd3 is selectively required for neural crest specification, migration and survival Rodney A. Stewart a, Brigitte L. Arduini
Developmental Biology 292 (2006) 174 188 www.elsevier.com/locate/ydbio Zebrafish foxd3 is selectively required for neural crest specification, migration and survival Rodney A. Stewart a, Brigitte L. Arduini
THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM
 Annu. Rev. Neurosci. 1999. 22:261 94 Copyright c 1999 by Annual Reviews. All rights reserved THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM Kevin J. Lee and Thomas M. Jessell
Annu. Rev. Neurosci. 1999. 22:261 94 Copyright c 1999 by Annual Reviews. All rights reserved THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM Kevin J. Lee and Thomas M. Jessell
Introduction to Embryology. He who sees things grow from the beginning will have the finest view of them.
 He who sees things grow from the beginning will have the finest view of them. Aristotle 384 322 B.C. Introduction to Embryology This lecture will introduce you to the science of developmental biology or
He who sees things grow from the beginning will have the finest view of them. Aristotle 384 322 B.C. Introduction to Embryology This lecture will introduce you to the science of developmental biology or
THE PROBLEMS OF DEVELOPMENT. Cell differentiation. Cell determination
 We emphasize these points from Kandel in Bi/CNS 150 Bi/CNS/NB 150: Neuroscience Read Lecture Lecture Friday, October 2, 2015 Development 1: pp 5-10 Introduction Brains evolved All higher animals have brains
We emphasize these points from Kandel in Bi/CNS 150 Bi/CNS/NB 150: Neuroscience Read Lecture Lecture Friday, October 2, 2015 Development 1: pp 5-10 Introduction Brains evolved All higher animals have brains
Neural Crest Development and Craniofacial Morphogenesis Is Coordinated by Nitric Oxide and Histone Acetylation
 Article Neural Crest Development and Craniofacial Morphogenesis Is Coordinated by Nitric Oxide and Histone Acetylation Yawei Kong, 1,2,3 Michael Grimaldi, 1,2,3 Eugene Curtin, 1,2 Max Dougherty, 1,2 Charles
Article Neural Crest Development and Craniofacial Morphogenesis Is Coordinated by Nitric Oxide and Histone Acetylation Yawei Kong, 1,2,3 Michael Grimaldi, 1,2,3 Eugene Curtin, 1,2 Max Dougherty, 1,2 Charles
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm July 28, 2008 Paraxial and Intermediate Mesoderm Overview Development of major mesodermal lineages Somites: formation specification and differentiation Mesodermal
Biology 4361 Paraxial and Intermediate Mesoderm July 28, 2008 Paraxial and Intermediate Mesoderm Overview Development of major mesodermal lineages Somites: formation specification and differentiation Mesodermal
Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural crest to definitive roof plate
 Nitzan et al. BMC Biology (2016) 14:23 DOI 10.1186/s12915-016-0245-6 RESEARCH ARTICLE Open Access Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural
Nitzan et al. BMC Biology (2016) 14:23 DOI 10.1186/s12915-016-0245-6 RESEARCH ARTICLE Open Access Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Role of Cadherin-7 in cranial neural crest development and its potential heterotypic interaction with N-cadherin in trigeminal ganglion formation
 C h a p t e r 4 Role of Cadherin-7 in cranial neural crest development and its potential heterotypic interaction with N-cadherin in trigeminal ganglion formation 4.1 Abstract Cadherins are transmembrane
C h a p t e r 4 Role of Cadherin-7 in cranial neural crest development and its potential heterotypic interaction with N-cadherin in trigeminal ganglion formation 4.1 Abstract Cadherins are transmembrane
Table S1. Sequence of primers used in RT-qPCR
 Table S1. Sequence of primers used in RT-qPCR Primer Name P16Ink4a-F P16Ink4a-R P15Ink4b-F P15Ink4b-R P19Arf-F P19Arf-R P53-F P53-R P21cip1-F P21cip1-R P27kip1-F P27kip1-R P18Ink4c-F P18Ink4c-R P19Ink4d-F
Table S1. Sequence of primers used in RT-qPCR Primer Name P16Ink4a-F P16Ink4a-R P15Ink4b-F P15Ink4b-R P19Arf-F P19Arf-R P53-F P53-R P21cip1-F P21cip1-R P27kip1-F P27kip1-R P18Ink4c-F P18Ink4c-R P19Ink4d-F
The Chick Embryo Model System
 The Chick Embryo Model System Guest Editor Claudio Stern Department of Cell & Developmental Biology, University College London, London, U.K. http://orcid.org/0000-0002-9907-889x Volume 62 Nos. 1/2/3 Special
The Chick Embryo Model System Guest Editor Claudio Stern Department of Cell & Developmental Biology, University College London, London, U.K. http://orcid.org/0000-0002-9907-889x Volume 62 Nos. 1/2/3 Special
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Abstract. Assistant Professor Dr. Lisa Taneyhill, Department of Animal and Avian Sciences
 Abstract Title of Document: THE ROLE OF THE ADHERENS JUNCTION PROTEIN αn-catenin IN NEURAL CREST-DERIVED TRIGEMINAL GANGLIA FORMATION Rachel Hooper, Master of Science 2012 Directed By: Assistant Professor
Abstract Title of Document: THE ROLE OF THE ADHERENS JUNCTION PROTEIN αn-catenin IN NEURAL CREST-DERIVED TRIGEMINAL GANGLIA FORMATION Rachel Hooper, Master of Science 2012 Directed By: Assistant Professor
Supplemental Information
 Molecular Cell, Volume 52 Supplemental Information The Translational Landscape of the Mammalian Cell Cycle Craig R. Stumpf, Melissa V. Moreno, Adam B. Olshen, Barry S. Taylor, and Davide Ruggero Supplemental
Molecular Cell, Volume 52 Supplemental Information The Translational Landscape of the Mammalian Cell Cycle Craig R. Stumpf, Melissa V. Moreno, Adam B. Olshen, Barry S. Taylor, and Davide Ruggero Supplemental
Langman's Medical Embryology
 Langman's Medical Embryology Developmental Biology Differentiation Morphogenesis) Epigenetic landscape (Waddington) ips Langman's Medical Embryology Morphogen gradient FGF8 in mouse limb bud Gilbert "Developmental
Langman's Medical Embryology Developmental Biology Differentiation Morphogenesis) Epigenetic landscape (Waddington) ips Langman's Medical Embryology Morphogen gradient FGF8 in mouse limb bud Gilbert "Developmental
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
UNIVERSITY OF YORK BIOLOGY. Developmental Biology
 Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
