Roles of the Mammalian Mitochondrial Fission and Fusion Mediators Fis1, Drp1, and Opa1 in Apoptosis
|
|
|
- Iris Dennis
- 5 years ago
- Views:
Transcription
1 Molecular Biology of the Cell Vol. 15, , November 2004 Roles of the Mammalian Mitochondrial Fission and Fusion Mediators Fis1, Drp1, and Opa1 in Apoptosis Yang-ja Lee,* Seon-Yong Jeong,* Mariusz Karbowski,* Carolyn L. Smith, and Richard J. Youle* *Biochemistry Section, Surgical Neurology Branch, and Light Imaging Facility, National Institute of Neurological Disorders and Stroke, National Institutes of Health, Bethesda, MD Submitted April 8, 2004; Revised August 5, 2004; Accepted August 30, 2004 Monitoring Editor: Keith Yamamoto During apoptosis, the mitochondrial network fragments. Using short hairpin RNAs for RNA interference, we manipulated the expression levels of the proteins hfis1, Drp1, and Opa1 that are involved in mitochondrial fission and fusion in mammalian cells, and we characterized their functions in mitochondrial morphology and apoptosis. Down-regulation of hfis1 powerfully inhibits cell death to an extent significantly greater than down-regulation of Drp1 and at a stage of apoptosis distinct from that induced by Drp1 inhibition. Cells depleted of Opa1 are extremely sensitive to exogenous apoptosis induction, and some die spontaneously by a process that requires hfis1 expression. Wild-type Opa1 may function normally as an antiapoptotic protein, keeping spontaneous apoptosis in check. However, if hfis1 is downregulated, cells do not require Opa1 to prevent apoptosis, suggesting that Opa1 may be normally counteracting the proapoptotic action of hfis1. We also demonstrate in this study that mitochondrial fragmentation per se does not result in apoptosis. However, we provide further evidence that multiple components of the mitochondrial morphogenesis machinery can positively and negatively regulate apoptosis. INTRODUCTION Mitochondria are morphologically dynamic organelles that continuously divide and fuse to form small individual units or interconnected networks within the cell (Bereiter-Hahn, 1990). They reach an equilibrium between these two states in healthy cells by regulating the relative rates of organelle fusion and fission (Nunnari et al., 1997). Proteins controlling mitochondrial fission include Drp1 (DNM1 in yeast) (Otsuga et al., 1998; Bleazard et al., 1999), Fis1 (Mozdy et al., 2000; Jakobs et al., 2003), and up to six other gene products in yeast (Dimmer et al., 2002). Drp1 is a large GTPase similar to dynamin, a protein that spirals around the necks of endocytic vesicles and facilitates their scission from the plasma membrane (Hinshaw, 2000). Inhibition of Drp1 GTPase activity with a dominant negative protein defective in GTP binding (Drp1 K38A ) results in elongated mitochondria, confirming that in mammalian cells (Smirnova et al., 1998, 2001; Frank et al., 2001) Drp1 functions like its yeast ortholog DNM1 in mitochondrial fission (Otsuga et al., 1998; Bleazard et al., 1999). Drp1 exists primarily in the cytoplasm but partially associates into foci on the outer surface of mitochondria that coalesce at sites of organelle fission (Smirnova et al., 2001). Human Fis1 (hfis1) contains a tetratricopeptide Article published online ahead of print. Mol. Biol. Cell / mbc.e Article and publication date are available at These authors contributed equally to this work. Corresponding author. address: youler@ninds.nih.gov. Abbreviations used: Act D, actinomycin D; PARP, poly(adp-ribose) polymerase; RNAi, RNA interference; shrna, short hairpin RNA; STS, staurosporine; zvad-fmk, z-val-ala-asp(ome)-fluoromethyl ketone. repeat (TPR) motif domain (Suzuki et al., 2003; Dohm et al., 2004) facing the cytoplasm and is anchored in the outer mitochondrial membrane via a C-terminal hydrophobic tail (Yoon et al., 2003). hfis1 circumscribes the outer surface of mitochondria and is not localized specifically to mitochondrial scission sites (Suzuki et al., 2003; Yoon et al., 2003). Down-regulating mammalian Fis1 expression by RNA interference (RNAi) induces mitochondrial elongation (Stojanovski et al., 2004), confirming results previously reported in yeast that FIS1 is involved in mitochondrial fission (Mozdy et al., 2000; Jakobs et al., 2003). Mitochondrial fusion in mammalian cells is controlled by the large GTPases Mfn1 (Santel and Fuller, 2001; Ishihara et al., 2003), Mfn2 (Santel and Fuller, 2001; Ishihara et al., 2003), and Opa1 (MGM1 in yeast) (Olichon et al., 2002; Satoh et al., 2003). Elimination of any of these proteins induces mitochondrial fragmentation (Chen et al., 2003; Olichon et al., 2003; Griparic et al., 2004). Mutations in Opa1 were found to cause dominant optic atrophy (Alexander et al., 2000; Delettre et al., 2000), potentially connecting this process (inhibition of mitochondrial fusion) to human disease. During apoptosis, the mitochondrial network fragments, resulting in smaller and more numerous mitochondria (Mancini et al., 1997; De Vos et al., 1998; Zhuang et al., 1998; Desagher et al., 1999; Frank et al., 2001; Karbowski et al., 2002). It has been reported that increased fission, decreased fusion, or both cause this mitochondrial phenotype at a stage of apoptosis upstream of caspase activation and close to that of Bax translocation to mitochondria and cytochrome c release (Karbowski et al., 2004), suggesting a mechanistic link between mitochondrial morphology and apoptosis. Drp1 binding to mitochondria is increased during apoptosis (Frank et al., 2001; Breckenridge et al., 2003), and the Drp1 foci at mitochondrial scission sites colocalize with foci of Bax that occur on mitochondria during apoptosis (Karbowski et 2004 by The American Society for Cell Biology 5001
2 Y. Lee et al. al., 2002). Inhibition of Drp1 GTPase activity with a dominant negative protein (Drp1 K38A ) prevents the mitochondrial fragmentation seen during apoptosis and delays the process of cell death (Frank et al., 2001; Karbowski et al., 2002), suggesting that mitochondrial fission is a required step in apoptosis. Overexpression of hfis1 has been reported to induce apoptosis (James et al., 2003), also suggesting the involvement of mitochondrial fission in apoptosis. Down-regulation of Opa1 expression in cells by RNAi results in spontaneous cell apoptosis (Olichon et al., 2003), suggesting that the promotion of mitochondrial fusion by wild-type levels of Opa1 normally protects cells from apoptosis. To further investigate the link between mitochondrial fission and apoptosis, we have explored the role of hfis1 in these processes. Using short hairpin RNAs (shrnas) for RNAi (Paddison et al., 2002), we have down-regulated three proteins involved in mitochondrial morphology: Drp1, hfis1, and Opa1. Using this technique, we are able to manipulate the expression levels of the proteins and to characterize their functions in mitochondrial morphology and apoptosis. Down-regulation of hfis1 powerfully inhibits cell death by multiple pathways to an extent significantly greater than down-regulation of Drp1 and at a stage of apoptosis distinct from that induced by Drp1 inhibition. Cells depleted of Opa1 are extremely sensitive to exogenous apoptosis induction and spontaneously die by a process that requires hfis1 expression. Thus, multiple proteins that comprise the mitochondrial morphogenesis machinery can positively and negatively regulate apoptosis. MATERIALS AND METHODS Cell Culture, Transfection, DNA Constructs, and RNAi HeLa cells (American Type Culture Collection, Manassas, VA) were cultured in complete DMEM supplemented with 10% heat-inactivated fetal calf serum, 100 U/ml penicillin, and 100 g/ml streptomycin in 5% CO 2 at 37 C. Transfections of HeLa cells were performed using FuGENE 6 (Roche Diagnostics, Indianapolis, IN) according to the manufacturer s instructions. The cdna for hfis1 (GenBank accession no. AF151893) was cloned by reverse transcription-polymerase chain reaction with total human RNA as template. cdna for wild-type hfis1 (hfis1wt) and hfis1 lacking the C-terminal 30 residues (hfis1 C) was cloned into mammalian expression vectors pcdna3.1 and prep4. RNAi was performed using the sh-activated gene silencing system (Paddison et al., 2002). The plasmids expressing shrnas were constructed by synthesizing two cdna oligonucleotides bearing the target sequence, HindIII linker, and U6 terminator, annealing them and ligating them into BseRI and BamHI sites of pshag-1 (Cold Spring Harbor Laboratory, Cold Spring, NY). For long-term (stable) suppression of gene expression by shrna, the region encoding the U6 promoter and shrna in pshag-1 was subcloned into the NotI and BamHI sites of prep4 (Invitrogen, Carlsbad, CA). The target sequences for the hfis1, Opa1, and Drp1 were as follows: 5 -GAGACGCGGGAGCCCACGGAGAACGCTCC-3 for hfis1, 5 -GATGAAGTTATCAGTCTGAGCCAGGTTAC-3 for Opa1, and 5 - TTCAATCCGTGATGAGTATGCTTTTCTTC-3 for Drp1. The sequence for the control shrna was 5 -TCGTACTCATAATCAGCTCTGCATACATC-3, which targets Opa1 3 end noncoding region and has no effect on silencing of Opa1 gene expression. One day after the transfection with these prep4 constructs, HeLa cells were grown in DMEM containing 300 g/ml hygromycin B for 2 d followed by 3 4 d in DMEM containing 50 g/ml hygromycin B for the selection of transfectants. Immunocytochemistry and Confocal Microscopy Cells grown in two-well chamber slides were treated as indicated, fixed with 4% paraformaldehyde for 10 min, permeabilized with 0.2% Triton X-100 for 15 min, and then blocked with 2% bovine serum albumin for 1hatroom temperature (RT). Cells were probed with rabbit polyclonal anti-bax (1:800; Upstate Biotechnology, Lake Placid, NY), mouse monoclonal anti-cytochrome c (1:800; BD Biosciences PharMingen), or mouse monoclonal anti-dlp1/drp1 (1:100; BD Transduction Laboratories, Lexington, KY) overnight at 4 C followed by staining with goat anti-rabbit Alexa Fluor 594 (1:600; Molecular Probes, Eugene, OR) or goat anti-mouse Alexa Fluor 488 antibodies (1:600; Molecular Probes) for 2hatRT. After washing, cells were mounted with SlowFade light antifade reagent (Molecular Probes) and analyzed by confocal microscopy. To visualize the mitochondria in living cells, 50 nm Mitotracker CMXRos (Molecular Probes) was added and incubated for 30 min before confocal microscopy. To analyze the mitochondrial membrane potential, cells were incubated for 20 min with 5 g/ml JC1 (Molecular Probes) in culture medium and observed by confocal microscopy. Images were captured with an LSM 510 Zeiss confocal microscope. Matrix-targeted photoactivable green fluorescent protein (mito-pagfp) based mitochondrial fusion assay was performed as described previously (Karbowski et al., 2004) Subcellular Fractionation and Immunoblotting Cells were permeabilized with digitonin (300 g/ml) in cytosolic extraction buffer (250 mm sucrose, 70 mm KCl, 137 mm NaCl, 4.3 mm Na 2 HPO 4, 1.4 mm KH 2 PO 4, ph 7.2, 100 M phenylmethylsulfonyl fluoride [PMSF], 10 g/ml leupeptin, 2 g/ml aprotinin) for 5 min on ice. Plasma membrane permeabilization of cells was confirmed by staining cells with trypan blue. Permeabilized cells were centrifuged at 1000 g for 5 min at 4 C. The supernatants (cytosolic fractions; S) were saved, and the pellets were solubilized in the same volume of mitochondrial lysis buffer (50 mm Tris-Cl, ph 7.4, 150 mm NaCl, 2 mm EDTA, 2 mm EGTA, 0.2% Triton X-100, 0.3% NP-40, 100 M PMSF, 10 g/ml leupeptin, 2 g/ml aprotinin), followed by centrifugation at 10,000 g for 10 min at 4 C, and the supernatants were used as heavy membrane (HM) fractions. Total cell lysates were prepared by solubilizing whole cells in Laemmli sample buffer and boiling them for 10 min. An aliquot of each sample was taken to determine protein concentrations by BCA protein assay kit (Pierce Chemical, Rockford, IL). Equal amounts of samples were run on 4 12% polyacrylamide gradient gels (Invitrogen), transferred to polyvinylidene difluoride membranes (Immobilon-P; Millipore, Billerica, MA), and immunoblotted. The primary antibodies, their dilutions, and their sources are as follows: anti-fis1 (1:500; Axxora, San Diego, CA), anti-opa1 (Zhu et al., 2003), anti-dlp1/drp1 (1:1000; BD Transduction Laboratories), anti-bax (1: 1000; Santa Cruz Biotechnology), anti-cytochrome c (1:1000; BD Biosciences PharMingen), anti-parp (1:1000; BIOMOL Research Laboratories, Plymouth Meeting, PA), and anti-actin (1:1000; Sigma-Aldrich, St. Louis, MO). Horseradish peroxidase-conjugated secondary antibodies (1:10,000) were used. Blots were detected by ECL Plus (Amersham Biosciences, Piscataway, NJ). Assessment of Apoptotic Cell Death By Nuclear Morphology. Cells grown in chamber slides were treated as mentioned in the text or figure legends. Nuclei were stained with Hoechst (Molecular Probes) (1 g/ml; 15 min at RT) and visualized under the fluorescent microscope (for UV excitation), and cells were scored as normal or apoptotic nuclei in several fields. At least 200 cells altogether in each treatment were counted and are shown as a percentage of cells with apoptotic nuclei among total cells counted. By DNA Ladder Formation. Cells grown in a 10-cm culture dish were treated as mentioned above, harvested, and washed once with phosphate-buffered saline, and total DNA was isolated as described previously (Lee and Shacter, 1997). After digestion with proteinase K and RNase A, the DNA was separated in 2% agarose gels and visualized with ethidium bromide under UV light. By Poly(ADP-Ribose) Polymerase (PARP) Cleavage/Caspase-3 Activation. PARP is a well known caspase-3 substrate. During apoptosis, caspase-3 is activated, so PARP cleavage can be used as a marker of apoptosis. Cells grown in a 10-cm culture dish were treated as described above and harvested, and total cell extracts were prepared. Equal amounts of proteins were separated in 4 12% acrylamide gels and immunoblotted with anti-parp antibodies. RESULTS Knockdown of hfis1 or Drp1 Expression Induces Mitochondrial Fusion Mitochondrial division in yeast requires at least seven to eight proteins (Dimmer et al., 2002), including Dnm1p (Otsuga et al., 1998; Bleazard et al., 1999) and Fis1p (Mozdy et al., 2000; Jakobs et al., 2003). The deletion or mutation of either of these genes disrupts proper mitochondrial division and causes mitochondrial membranes to form net-like structures (Dimmer et al., 2002). To explore the roles of the human orthologues of Dnm1p and Fis1p, Drp1 and hfis1, respectively, in mitochondrial fission and apoptosis, we knocked down their expression by using the short hairpin-activated gene silencing system (i.e., SHAGging; Cold Spring Harbor Laboratory) (Paddison et al., 2002). To generate cell lines that stably suppress the endogenous gene, we transferred the U6-shRNA inserts to prep4, an episomally maintained mammalian expression vector with hygromycin as a selective marker. Among 12 different oligonucleotides designed for hfis1 shrna, only one, which corresponds to a sequence situated just outside of the 3 end of the coding sequence, signifi Molecular Biology of the Cell
3 Roles of Fis1 and Opa1 in Apoptosis cells showed a more normal shape (short tubular) (Figure 1, B and C). Normal mitochondria in hfis1 RNAi cells might have been overscored, considering that hygromycin, which was used for the selection, tends to induce slightly shorter mitochondria (our unpublished data). Most of the cells that were transfected with a plasmid carrying control shrna have normal relatively short tubular mitochondria and some fragmented ones, but never elongated mitochondria such as seen in hfis1 shrna transfectants (Figure 1, B and C). Unlike hfis1, four of six sequences targeted to Drp1 could silence Drp1 gene expression, including one reported previously to be effective in mammalian systems (Koch et al., 2003). On Drp1 depletion (Figure 1A) almost 80% of cells had elongated (fused) mitochondria (Figure 1, B and C). Even though mitochondria in hfis1 RNAi cells and Drp1 RNAi cells were both elongated extensively, the shapes of mitochondria were not the same. Many balloon-like or bulbed structures were seen at the base of tubules in the mitochondria of Drp1 RNAi cells but not hfis1 RNAi cells (Figure 1B, arrow). Interestingly, no differences in mitochondrial structure between dnm1 and fis1 cells were observed in yeast; both exhibited net-like structures (Mozdy et al., 2000). However, the larger size of mammalian mitochondria permits better visualization of morphology. Mammalian mitochondria have been reported to display expansions at the ends of mitochondria in cells overexpressing the dominant negative inhibitor Drp1 K38A (Smirnova et al., 2001). Figure 1. Knockdown of hfis1 or Drp1 expression induces mitochondrial fusion. HeLa cells were transfected with prep4 constructs containing shrna of the target sequence of hfis1, Drp1, or control, and the transfectants were selected by growth in media containing hygromycin B. (A) Total cell lysates from the hfis1 RNAi cells, Drp1 RNAi cells, and control RNAi cells were prepared, and the expression levels of hfis1 and Drp1 were analyzed by Western blotting. Actin level also was analyzed for a loading control. (B) Mitochondria of control RNAi cells, hfis1 RNAi cells, or Drp1 RNAi cells were visualized with Mitotracker Red CMXRos and analyzed by confocal microscopy. Enlargements are shown for detailed structure of mitochondria. An arrow shows a typical balloon-like structure in Drp1 RNAi cells. (C) Percentage of cell population with fragmented (dotted), normal (solid), or elongated (striped) mitochondria in control RNAi, hfis1 RNAi, or Drp1 RNAi culture. At least 200 cells in several fields were counted in each experiment. Data represent the mean SD of at least three independent experiments. cantly inhibited hfis1 expression in HeLa cells. Selection of transfected HeLa cells with nearly complete depletion of hfis1 required 1 wk of growth in the presence of hygromycin (Figure 1A). More than 70% of these cells had elongated mitochondria, whereas mitochondria in the remaining Knockdown of hfis1 Did Not Affect the Distribution of Drp1 to Mitochondria Drp1 is known not only to distribute predominantly to the cytosol but also accumulates at foci in the mitochondrial outer membrane that represent future fission sites (Labrousse et al., 1999; Smirnova et al., 2001). Because Drp1 lacks a mitochondrial targeting sequence, some protein(s) may be needed to recruit Drp1 to mitochondria. Human Fis1 is a good candidate for this recruitment because in yeast Fis1p is required for the proper assembly and distribution of Dnm1p-containing complexes on mitochondria (Mozdy et al., 2000; Tieu et al., 2002), and the structure of hfis1 resembles that of certain proteins involved in mitochondrial import (Suzuki et al., 2003). If this were the case, the mitochondrial distribution of Drp1 should be affected by hfis1 depletion. However, there was little or no difference in the Drp1 distribution between control RNAi cells and hfis1 RNAi cells assessed by immunocytochemistry (Figure 2A) or by Western blot analysis of the subcellular-fractionated S and HM (mostly mitochondrial fraction) samples (Figure 2B). Drp1 is distributed to the mitochondria in hfis1 RNAi cells to a similar extent as in control RNAi cells. Fis1 Depletion Produces Greater Resistance to Apoptosis Than Depletion of Drp1 Overexpression of the dominant negative mutant Drp1 K38A induced mitochondrial fusion, and cells became resistant to apoptosis (Frank et al., 2001; Karbowski et al., 2002; Breckenridge et al., 2003), suggesting a close relationship between mitochondrial morphology and sensitivity to cell death. We examined how the knockdown of hfis1 expression affects the sensitivity to cell death by various apoptosis inducers and compared hfis1 RNAi cells to Drp1 RNAi cells. We found that 20% of hfis1 RNAi cells were apoptotic under conditions that produced 40 65% apoptosis of control RNAi cells (assessed by apoptotic nuclei) (Figure 3A). The caspase-3 activity of hfis1 RNAi cells from staurosporine (STS)-, actinomycin D (Act D)-, or anti-fas induced apoptosis was much lower than for control RNAi cells assessed by Vol. 15, November
4 Y. Lee et al. negative and cytochrome c release negative), and upon apoptosis the Bax translocates to mitochondria and cytochrome c is released to the cytosol (Bax translocation positive and cytochrome c release positive). However, we observed an unusual population of Bax translocation positive- and cytochrome c release-negative cells during apoptosis of Drp1-depleted cells (Figure 4C), suggesting that Drp1 acts after Bax translocation to inhibit cytochrome c release. The results of immunocytochemistry were confirmed by Western blot analysis of the subcellular-fractionated S and HM samples (Figure 4D). In control RNAi cells, the majority of cytosolic Bax disappeared (translocated to the mitochondria), and more than one-half of the cytochrome c in mitochondria was released to the cytosol by actinomycin D treatment. In contrast, there was almost no difference in Bax or cytochrome c distribution in hfis1 RNAi cells with or without apoptosis induction. Thus, the absence of hfis1 blocked apoptosis upstream of Bax translocation. In Drp1 RNAi cells, however, cytosolic Bax disappeared almost completely, but cytochrome c release from the mitochondria was significantly lower than in control cells, indicating that the protection from apoptosis by Drp1 RNAi is downstream of Bax translocation and upstream of cytochrome c release. These results confirm the immunocytochemistry results and indicate that hfis1 and Drp1 work at different steps to promote apoptosis. Figure 2. Knockdown of hfis1 did not affect the distribution of Drp1 to mitochondria. (A) Control or hfis1-depleted HeLa cells were incubated with Mitotracker CMXRos (red) for 20 min before fixation and stained with mouse monoclonal anti-dlp1/drp1 followed by antimouse Alexa Fluor 488 antibodies (green). Mitochondria and distribution of Drp1 were analyzed by confocal microscopy. Red, mitochondria; green, Drp1. (B) HeLa cells depleted of hfis1 along with control RNAi were fractionated into S and HM (mainly mitochondria) and analyzed for the distribution of Drp1 by Western blotting. Fractionation quality was verified by the distribution of specific subcellular markers: CoxIV for mitochondria and actin for cytosol. PARP cleavage (Figure 3B). Less inhibition of apoptosis by Drp1 silencing was observed (Figure 3, A and B). To explore the mechanism of the cell death resistance of hfis1 RNAi cells, we examined two early events in apoptosis: the translocation of proapoptotic protein Bax from the cytosol to the mitochondria and the release of cytochrome c from the mitochondria to the cytosol (Kluck et al., 1997; Wolter et al., 1997). Control RNAi, hfis1 RNAi, and Drp1 RNAi cells were treated with actinomycin D in the presence of zvad-fmk for 8 h, before Bax and cytochrome c staining by immunocytochemistry. As shown in Figure 4, A and B, most of the Bax in hfis1 RNAi cells was present in the cytosol and the cytochrome c localized to mitochondria under these conditions, in contrast to cells transfected with control RNAi. Interestingly, under the same conditions, in 80% of Drp1 RNAi cells Bax had translocated to mitochondria but in one-half of these cells, cytochrome c was still retained in the mitochondria (Figure 4, A and B). Typically, in healthy cells, Bax is localized in the cytosol and cytochrome c is in the mitochondria (Bax translocation Overexpression of hfis1 Wild-Type (hfis1 wt) but Not C Terminus-truncated Mutant (hfis1 C) Reverts the Apoptosis-resistant hfis1 RNAi Cells To confirm that the apoptosis resistance in hfis1 RNAi cells is the result of hfis1 depletion, we reconstituted hfis1 expression in the hfis1 RNAi cells. The targeted sequence we used for hfis1 RNAi is situated just outside of the 3 end of the coding sequence, so hfis1 cdna constructs can be used to express hfis1 in the hfis1 RNAi cells. We cotransfected HeLa cells with cdna constructs encoding either hfis1 wt or an hfis1 C mutant lacking the membrane anchor along with the hfis1 shrnai construct, selected transfectants with hygromycin, and examined protein expression levels and sensitivity to apoptosis. As shown in Figure 5A, hfis1 (both wt and C) was well expressed in the hfis1 RNAi cells (lanes 5 8). The hfis1 RNAi cells overexpressing hfis1 showed more sensitivity to apoptosis than hfis1 RNAi cells assessed by PARP cleavage (Figure 5A), nuclei fragmentation (Figure 5B), and cytochrome c release (Figure 5, C and D). However, the overexpression of hfis1 C mutant in hfis1 RNAi cells did not affect the sensitivity to apoptosis. The results confirm that hfis1 is required for cell sensitivity to apoptosis and show that mitochondrial targeting of hfis1 is necessary for apoptosis promotion. Knockdown of Opa1 Induces Mitochondrial Fragmentation and Sensitizes Cells to Apoptosis The down-regulation of Opa1 in HeLa cells by RNAi was reported to lead to fragmentation of mitochondria, to the dissipation of the mitochondrial membrane potential, and to a disorganization of the cristae (Olichon et al., 2003). These events were followed by cytochrome c release and caspasedependent apoptotic nuclear events. To study Opa1 function further, especially in conjunction with the fission proteins hfis1 and Drp1, we designed several shrna constructs specific to the Opa1 sequence (coding and noncoding) for gene silencing by the same method we used for hfis1 and Drp1 RNAi. The sequence that was reported to down-regulate Opa1 (Olichon et al., 2003) also worked in the SHAGging system. A sequence that did not produce gene silencing was used as a control RNAi. It took 5 7 d after transfection for effective selection of transfectants, and Opa1 was almost completely depleted in this population (Figure 6A). The 5004 Molecular Biology of the Cell
5 Roles of Fis1 and Opa1 in Apoptosis Figure 3. hfis1 depletion produces greater resistance to apoptosis than depletion of Drp1. (A) HeLa cells depleted of hfis1 or Drp1 by RNAi along with control RNAi cells were treated with STS (1 M; 6 h), Act D (10 M; 8 h), etoposide (100 M; 30 h), or anti-fas antibody (500 ng/ml, 15 h), and the nuclei were stained with Hoechst (1 g/ml; 15 min at RT). Normal or apoptotic nuclei of these cells in several fields were counted under the fluorescent microscope (for UV excitation). At least 200 cells altogether in each treatment were counted and shown as a percentage of cells with apoptotic nuclei among total cells counted. The data are plotted as the mean SD of at least three independent experiments. (B) PARP cleavage was analyzed in the total extracts from HeLa cells depleted of hfis1, Drp1, and control, which were treated with STS (1 M; 0, 3, 6 h), Act D (10 M; 0, 4, 8 h), or anti-fas (500 ng/ml; 0, 6, 15 h). The figure is a representative of at least three independent experiments. lower band, which did not disappear, may be a nonspecific band rather than one of the eight alternative splicing variants (Delettre et al., 2000) because the targeting sequence we used for RNAi exists in all the splicing variants. Under these conditions, almost all the cells displayed extensively fragmented mitochondria, clearly different from the control RNAi cells (Figure 6, B and C). In contrast to hfis1 and Drp1 RNAi cells, Opa1 RNAi cells are more sensitive to apoptosis induced by various stimuli compared with the control RNAi cells, and consistent with a previous report (Olichon et al., 2003), a portion of the cells ( 25 35%) die spontaneously (Figure 6D). Opa1 down-regulated cells die with the typical hallmarks of apoptosis. Nuclear fragmentation (Figure 6D), DNA ladder formation (Figure 6E), Bax translocation, cytochrome c release (Figure 6, F and G), and caspase activation/parp cleavage (Figure 6H) were all observed. The cell death caused by Opa1 depletion was inhibited by the general caspase inhibitor zvad-fmk and by Bcl-2 overexpression (our unpublished data). Cells Depleted of Both hfis1 and Opa1 Show Apoptosis Resistance in Spite of Extensive Mitochondrial Fragmentation The down-regulation of hfis1 caused extensive mitochondrial fusion, and the cells became apoptosis resistant (Figures 1 and 3). In contrast, Opa1 down-regulation caused extensive mitochondrial fragmentation, and cells became more sensitive to death (Figure 6). In yeast, the elimination of fission mediators can be offset by eliminating fusion mediators (Sesaki and Jensen, 1999; Mozdy et al., 2000; Wong et al., 2000; Sesaki et al., 2003). Thus, we down-regulated both hfis1 and Opa1 and examined the effect on mitochondrial morphology and cell sensitivity to apoptosis. More than 70% of the cells with both hfis1 and Opa1 almost completely depleted (hfis1/opa1 RNAi) (Figure 7A) displayed fragmented mitochondria (Figure 7B), similar to results when cells were treated with Opa1 RNAi alone (Figure 6, B and C). This may reflect the rate of disappearance of Opa1 and hfis1 over the 5-d selection period. Therefore, Fis1 was depleted for 5 d and then Opa1 RNAi was transfected into the Fis1-depleted cells. Examining mitochondrial morphology 2 d after the Opa1 depletion showed fragmentation of the mitochondria in contrast to cells that expressed control RNAi in the second transfection (Figure 7C). Thus, even after depletion of hfis1 with RNAi, a finite rate of mitochondrial fission occurs that becomes predominant in the absence of Opa1. Slower but finite mitochondrial fission also occurs after genetic deletion of Fis1 in yeast (Jakobs et al., 2003). Residual fission of mitochondria in hfis/opa1 RNAi was further confirmed by the results obtained using a mitochondrial fusion assay based on the measurement of dilution rates of mito-pagfp, as described previously (Karbowski et al., 2004). We found complete inhibition of mitochondria fusion in hfis1/opa1 double RNAi cells, indistinguishable from that detected in single Opa1 RNAi cells (Figure 7, D and E). hfis1/opa1 RNAi cells were very resistant to apoptosis, almost to the same extent as hfis1 RNAi cells and opposite from Opa1 RNAi cells (Figure 8A). However, the degree of the resistance of these cells varied substantially from experiment to experiment (shown by the larger SE bars in Figure 8A). When the hfis1/opa1 RNAi cells and hfis1 RNAi cells (along with control RNAi cells and Opa1 RNAi cells) were treated longer with anti-fas antibody, the difference in sensitivity to apoptosis between them became larger (Figure 8B). The hfis1/opa1 RNAi cells are more sensitive than the hfis1 RNAi cells, but never as sensitive as the control RNAi cells or the Opa1 RNAi cells. The resistance of hfis1/opa1 RNAi cells to apoptosis also was confirmed by inhibition of Act D-induced cytochrome c release and Bax translocation (Figure 8C). Opa1 is known to be associated with the inner mitochondrial membrane, and it has been reported that the depletion of Opa1 caused a loss in mitochondrial membrane potential (Olichon et al., 2003). Indeed, mitochondrial mem- Vol. 15, November
6 Y. Lee et al. Figure 4. Both Bax translocation and cytochrome c release induced by actinomycin D are inhibited in hfis1- depleted HeLa cells, whereas cytochrome c release, but not Bax translocation, is inhibited in Drp1-depleted cells. (A) HeLa cells depleted of hfis1 or Drp1 by RNAi along with control RNAi cells were treated with Act D (10 M; 8 h) in the presence of zvadfmk (50 M), fixed, and double stained with anti-bax (rabbit polyclonal, red) and anti-cytochrome c (mouse monoclonal, green) antibodies. (B) The number of cells displaying Bax translocation or cytochrome c release was counted and plotted as a percentage of total cells counted in each RNAi cell population that had been treated with Act D and stained with anti-bax and anti-cytochrome c (the same samples as shown in A). At least 200 cells were counted altogether in several fields. Data are plotted as the mean SD of at least three independent experiments. (C) A unique group of cells (Bax had translocated but cytochrome c was not released) was seen in Drp1 RNAi cells. Four groups of cells were counted in each RNAi cell population that had been treated with Act D (the same samples as in A): i) Bax did not translocate and cytochrome c was not released; ii) Bax translocated, but cytochrome c is not released; iii) Bax not translocated, but cytochrome c is released; and iv) Bax translocated and cytochrome c is released were plotted as a percentage of total cells counted in each RNAi cell population. There were no cells categorized as iii, so only i, ii, and iv were plotted. Bars are labeled solid or striped, as in B. Data were plotted as the mean SD of at least three independent experiments. (D) HeLa cells depleted of Drp1 or hfis1 along with control RNAi were incubated with or without Act D (10 M) for 8 h, fractionated into S and HM (mainly mitochondria), and analyzed for the distribution of Bax and cytochrome c by Western blotting. Fractionation quality was verified by the distribution of specific subcellular markers: CoxIV for mitochondria and actin for cytosol. To confirm the depletion of proteins (hfis1 and Drp1) and their localizations, hfis1 and Drp1 also were examined. The figure shown is representative of three independent experiments. brane potential was lost in most of the Opa1-depleted cells, but the loss was prevented by hfis1 codepletion (Figure 8D). Thus, hfis1 is required for both the spontaneous and induced apoptosis caused by loss of Opa1 and functions epistatically to Opa1 in the apoptotic pathway. DISCUSSION Mitochondria fragment into smaller and more numerous units when mammalian cells are stimulated to undergo apoptosis (Mancini et al., 1997; De Vos et al., 1998; Zhuang et al., 1998; Desagher et al., 1999; Frank et al., 2001; Karbowski et al., 2002). When mitochondrial fission is blocked by overexpression of a dominant negative mutant of Drp1 (Drp1 K38A ), the downstream events of mitochondrial-induced cell death, such as cytochrome c release, also are blocked, suggesting that mitochondrial fission might be a required step in apoptosis (Frank et al., 2001; Breckenridge et al., 2003). Bax, a proapoptotic Bcl-2 family member, colocalizes with Drp1 in mitochondrial foci (Karbowski et al., 2002). On apoptosis initiation, Bax translocates from the cytosol to focal structures on the outer membrane of mitochondria (Nechushtan et al., 2001; Valentijn et al., 2003) that become sites of organelle scission (Karbowski et al., 2002), suggesting that Bax may activate the mitochondrial fission machinery. Screening yeast for mutants that suppress the phenotype of cells lacking the mitochondrial fusion gene FZO1 led to the initial identification of the importance of the outer mitochondrial membrane protein Fis1 in mitochondrial fission (Mozdy et al., 2000; Tieu and Nunnari, 2000). To further explore the connection between mitochondrial morphology and apoptosis, we tested the role in apoptosis of another protein involved in mitochondrial fission, hfis1. Human Fis1 has been reported to induce mitochondrial fragmentation upon overexpression (James et al., 2003; Yoon et al., 2003), and loss of hfis1 induces elongation of mammalian mitochondria (Stojanovski et al., 2004), suggesting a conserved role of FIS1 in eukaryotes. In yeast, FIS1 has been shown genetically to interact with DNM1 and seems to be essential for the recruitment of Dnm1p (Drp1) into focal structures on mitochondria (Mozdy et al., 2000; Tieu et al., 2002). Although human Fis1 can be cross-linked to Drp Molecular Biology of the Cell
7 Roles of Fis1 and Opa1 in Apoptosis Figure 5. Overexpression of hfis1 wild-type (hfis1 wt) but not C terminus-truncated mutant (hfis1 C) reverts apoptosis-resistant hfis1 RNAi cells. HeLa cells were transfected with control shrna/prep4 (a), hfis1 shrna/ prep4 (b), hfis1 shrna/prep4 plus hfis1wt cdna/pcdna3.1 (c), or hfis1 shrna/prep4 plus hfis1 C cdna/pcdna3.1 (d), and transfectants were selected as described in Materials and Methods. (A) Transfectants of each group (a d) were treated with or without STS (1 M) for 6 h, lysed, and analyzed for hfis1 expression levels and PARP cleavage by Western blotting. Actin also was analyzed as a loading control. (B) Transfectants of each group (a d) were treated with STS (1 M; 6 h), and the nuclei were stained with Hoechst (1 g/ml; 15 min at RT). Normal or apoptotic nuclei of these cells in several fields were counted under the fluorescent microscope (for UV excitation). At least 200 cells altogether in each treatment were counted and shown as a percentage of cells with apoptotic nuclei among the total cells counted. The data are plotted as the mean SD of at least three independent experiments. (C) Transfectants of each group (a d) were treated with Act D (10 M; 8 h) in the presence of zvad-fmk (50 M), fixed, and stained with anti-cytochrome c (mouse monoclonal, green) antibodies, and images were captured by confocal microscopy. (D) The number of cells whose cytochrome c was released was counted and plotted as a percentage of the total cells counted in each group (a d) of transfectants that had been treated with Act D and stained with anti-cytochrome c (the same samples as shown in C). At least 200 cells were counted altogether in several fields. Data are plotted as the mean SD of at least three independent experiments. (Yoon et al., 2003), the two do not seem to stably associate (James et al., 2003) nor does overexpression of hfis1 increase Drp1 binding to mitochondria (Suzuki et al., 2003). Here, we further show that, in contrast to yeast, loss of hfis1 does not decrease Drp1 binding to mitochondria (Figure 2, A and B). However, it is intriguing that hfis1 is required for Bax recruitment into the same Drp1-containing foci. We have found that loss of hfis1 inhibits induction of apoptosis. Previously, it was reported that overexpression of hfis1 induced cell death that was inhibited by Bcl-xL overexpression (James et al., 2003). Interestingly, the apoptosis induced by hfis1 overexpression was not inhibited by the dominant negative inhibitor Drp1 K38A, whereas cytochrome c release in those cells was blocked. This correlates with our results that hfis1 and Drp1 work at different steps in the apoptosis pathway. Loss of hfis1 inhibits Bax translocation, whereas loss of Drp1 does not. Loss of Drp1, however, prevents cytochrome c release even in cells where Bax has translocated to mitochondria (Figure 4). This represents a novel step in apoptosis inhibition. Typically, Bax translocates to mitochondria very close in time to cytochrome c release so that, within a population, cells exist primarily as Bax translocation negative and cytochrome c release negative or Bax translocation positive and cytochrome c release positive. On loss of Drp1, a large proportion of the cells become Bax translocation positive and cytochrome c release negative, showing a new stage of apoptosis inhibition. In accordance with James et al. (2003), we find that loss of Drp1 produces a more significant inhibition of cytochrome c release than inhibition of apoptosis, hinting that Bax translocation may induce cell death independent of cytochrome c release. Other inhibitors of apoptosis such as Bcl-2 and the loss of hfis1 block both steps, whereas the general caspase inhibitor z-val-ala-asp(ome)-fluoromethyl ketone (zvad-fmk) blocks neither, only inhibiting events downstream of outer mitochondrial membrane permeabilization. Thus, Drp1 seems to function subsequent to Bax translocation in the cytochrome c release process that occurs during apoptosis. How does hfis1 act as a proapoptotic protein? The hfis1 structure (Suzuki et al., 2003) shows that the protein contains four tandem structures resembling the TPR. The TPR motifs are known to facilitate specific protein protein interactions (Blatch and Lassle, 1999), suggesting that hfis1 may bind to other proteins and function as a molecular adaptor (receptor) on the mitochondrial outer membrane. TPR proteins in the outer mitochondrial membrane in addition to hfis1 include Tom20 (Iwahashi et al., 1997) and Tom70 (Young et al., 2003), both of which are involved in recruitment of other proteins to mitochondria. Although hfis1 does not seem to recruit Drp1 to mitochondria, a number of other interesting candidates will be worth examining. The notable differences between yeast and human mitochondrial fission and fusion machinery may offer insights into apoptosis as well as the mechanism of mitochondrial biogenesis. Loss of a protein involved in mitochondrial fusion had the opposite effect on apoptosis from the loss of two proteins involved in mitochondrial fission. When Opa1 was depleted by RNAi, the mitochondria were extensively fragmented, and cells became Vol. 15, November
8 Y. Lee et al. Figure 6. Depletion of Opa1 induces mitochondrial fragmentation, and cells become very sensitive to apoptosis. HeLa cells were transfected with prep4 constructs containing shrna of the target sequence of Opa1 or control, and the transfectants were selected by growing in media containing hygromycin B. (A) total cell lysates from the Opa1 RNAi cells along with control RNAi cells were prepared and the expression level of Opa1 was analyzed by Western blotting. Actin level also was analyzed as a loading control. (B) Mitochondria of control RNAi cells and Opa1 RNAi cells were visualized with Mitotracker Red CMXRos and analyzed by confocal microscopy. Enlargements are shown for detailed structure of mitochondria. (C) Percentage of cell population with fragmented (dotted), normal (solid), or elongated (striped) mitochondria in control RNAi or Opa1 RNAi culture. At least 200 cells in several fields were counted in each experiment. Data represent the mean SD of at least three independent experiments. (D) Opa1-depleted cells along with control RNAi cells were treated with STS (1 M; 6 h), Act D (10 M; 8 h), etoposide (100 M; 30 h), or anti-fas antibody (500 ng/ml; 15 h), and apoptotic nuclei (stained with Hoechst) were scored and plotted as a percentage of total cells (at least 200 cells) counted. Data are shown as the mean SD of three independent experiments. (E) Total DNA was isolated from the cells treated as described above and analyzed by agarose gel electrophoresis and visualized with ethidium bromide. The first lane is a DNA ladder control. The figure shown is a representative of three independent experiments. (F) HeLa cells depleted of Opa1 along with control RNAi cells were incubated with or without Act D (10 M)for8hinthepresence of zvad-fmk, fixed, and stained with anti-bax (red; our unpublished data) and anti-cytochrome c antibodies (green). (G) The number of cells in F displaying Bax translocation or cytochrome c release was counted and plotted as a percentage of total cells counted in each RNAi cell population that had been treated with or without Act D and stained with anti-bax and anti-cytochrome c antibodies (the same samples as shown in F). At least 200 cells were counted altogether in several fields. Data are plotted as the mean SD of at least three independent experiments. (H) PARP cleavage was analyzed in the total extracts from HeLa cells depleted of Opa1 and control that were treated with STS (1 M; 0, 3, 6 h), Act D (10 M; 0, 4, 8 h), or anti-fas (500 ng/ml; 0, 6, 15 h). The figure is a representative of at least three independent experiments Molecular Biology of the Cell
9 Roles of Fis1 and Opa1 in Apoptosis Figure 7. Mitochondria are extensively fragmented in the cells depleted of both hfis1 and Opa1. HeLa cells were transfected with prep4 constructs containing shrna of the target sequence of hfis1, Opa1, or both of them together along with control, and the transfectants were selected by growing in media containing hygromycin B. (A) Total cell lysates from the hfis1 RNAi cells, Opa1 RNAi cells, or both hfis1 and Opa1 RNAi cells along with control RNAi cells were prepared, and the expression levels of hfis1 and Opa1 were analyzed by Western blotting. Actin also was analyzed as a loading control. (B) Left, mitochondria of hfis1/opa1 RNAi cells were visualized with Mitotracker Red CMXRos and analyzed by confocal microscopy. An enlargement is shown for detailed structure of mitochondria. Right, percentage of cell population with fragmented (dotted), normal (solid), or elongated (striped) mitochondria in hfis1/opa1 RNAi cell culture. At least 200 cells in several fields were counted in each experiment. Data represent the mean SD of at least three independent experiments. (C) The elongated mitochondrial morphology in hfis1-depleted cells was reversed by subsequent depletion of Opa1. Hela cells depleted of hfis1 by RNAi were selected for 5 d and transfected again with Opa1 shrna- or a control shrna-construct and examined 2 d later. To identify transfectants, peyfp vector was cotransfected. Mitochondria were visualized with Mitotracker Red CMXRos and analyzed by confocal microscopy. (D and E) Control RNAi, Opa1 RNAi, and hfis1/opa1 RNAi double RNAi cells were analyzed for mitochondrial fusion by using mito-pagf based fusion assay. Cells were transfected with mito-pagfp, followed by the photoactivation of the small regions within the cells (white circles in prepanels) and confocal acquisition of z-series covering the entire thickness of the cell. (E) Changes in the fluorescence intensity within activated regions were measured over time. Vol. 15, November
10 Y. Lee et al. Figure 8. Cells depleted of both hfis1 and Opa1 show apoptosis resistance like hfis1-depleted cells. (A) HeLa cells depleted of hfis1, Opa1, or both of them by RNAi along with control RNAi cells were treated with STS (1 M; 6 h), Act D (10 M; 8 h), etoposide (100 M; 30 h), or anti-fas antibody (500 ng/ml; 15 h), and the nuclei were stained with Hoechst (1 g/ml; 15 min at RT). Normal or apoptotic nuclei of these cells in several fields were counted under the fluorescent microscope (for UV excitation). At least 200 cells altogether in each treatment were counted and plotted as a percentage of cells with apoptotic nuclei among the total cells counted. The data are shown as the mean SD of at least three independent experiments. (B) HeLa cells depleted of Opa1, hfis1, or both of them along with control RNAi cells were incubated with anti-fas antibody (500 ng/ml) for the periods as indicated, and the nuclei were stained and counted as normal or apoptotic nuclei. At least 200 cells altogether were counted in each sample at each time point and plotted as a percentage of cells with apoptotic nuclei among the total cells counted. The data were plotted as the mean SD of at least three independent experiments. (C) Bax translocation and cytochrome c release induced by Act D treatment are inhibited in hfis/opa1 RNAi cells. Left, HeLa cells depleted of both hfis1 and Opa1 by RNAi along with control RNAi cells were treated with Act D (10 M;8h)in the presence of zvad-fmk (50 M), fixed, and double stained with anti-bax (rabbit polyclonal, red) and anti-cytochrome c (mouse monoclonal, green) antibodies. Right, the number of cells displaying Bax translocation or cytochrome c release was counted and plotted as a percentage of the total cells counted in each RNAi cell population that had been treated with Act D and stained with anti-bax and anti-cytochrome c antibodies (the same samples as shown on the left). At least 200 cells were counted altogether in several fields. Data are plotted as the mean SD of at least three independent experiments. (D) hfis1 depletion prevents mitochondrial membrane potential reduction induced by Opa1 depletion. Left, HeLa cells depleted of hfis1 (b), Opa1 (c), or both of them (d) along with control RNAi cells (a) were incubated with 5 g/ml JC-1 for 20 min and observed by confocal microscopy. JC-1 is a cationic dye that indicates mitochondrial polarization by shifting its fluorescence emission from green to red. Regions of high mitochondrial membrane potential are indicated by red fluorescence, and regions of low mitochondrial membrane potential are indicated by green fluorescence. Right, the number of cells with mitochondria whose membrane potential was lost was counted and plotted as a percentage of the total cells counted in each RNAi cell population. very sensitive to apoptosis, consistent with the results reported by others (Olichon et al., 2003; Griparic et al., 2004), and further correlating mitochondrial fragmentation with sensitivity to apoptosis. Our results show that wild-type Opa1 may function normally as an antiapoptotic protein, keeping spontaneous apoptosis in check. However, if hfis1 is down-regulated, cells do not require Opa1 to prevent apoptosis, suggesting that Opa1 may be normally counteracting the proapoptotic action of hfis1. Thus, although mitochondrial fragmentation per se does not necessarily result in apoptosis, the components of the mitochondrial fissionfusion machinery can positively and negatively regulate apoptosis, and the rate of mitochondrial fission and fusion may be directly connected to the process of programmed cell death. ACKNOWLEDGMENTS We thank Dr. Craig Blackstone (National Institute of Neurological Disorders and Stroke/National Institutes of Health) for providing antibodies against Opa1 and for critical reading of the manuscript and many useful suggestions. We also thank Drs. Sang-Joon Park and Hyeog Kang (National Heart, Lung, and Blood Institute/National Institutes of Health) for much advice on the RNAi experiments. This work was supported by the intramural program of the National Institute of Neurological Disorders and Stroke/National Institutes of Health. REFERENCES Alexander, C., et al. (2000). OPA1, encoding a dynamin-related GTPase, is mutated in autosomal dominant optic atrophy linked to chromosome 3q28. Nat. Genet. 26, Bereiter-Hahn, J. (1990). Behavior of mitochondria in the living cell. Int. Rev. Cytol. 122, Blatch, G.L., and Lassle, M. (1999). The tetratricopeptide repeat: a structural motif mediating protein-protein interactions. Bioessays 21, Bleazard, W., McCaffery, J.M., King, E.J., Bale, S., Mozdy, A., Tieu, Q., Nunnari, J., and Shaw, J.M. (1999). The dynamin-related GTPase Dnm1 regulates mitochondrial fission in yeast. Nat. Cell Biol. 1, Breckenridge, D.G., Stojanovic, M., Marcellus, R.C., and Shore, G.C. (2003). Caspase cleavage product of BAP31 induces mitochondrial fission through 5010 Molecular Biology of the Cell
Roles of the mammalian mitochondrial fission and fusion mediators. Fis1, Drp1 and Opa1 in apoptosis
 Roles of the mammalian mitochondrial fission and fusion mediators Fis1, Drp1 and Opa1 in apoptosis Yang-ja Lee 1#, Seon-Yong Jeong 1#, Mariusz Karbowski 1, Carolyn L Smith 2, Richard J Youle 1* 1 Biochemistry
Roles of the mammalian mitochondrial fission and fusion mediators Fis1, Drp1 and Opa1 in apoptosis Yang-ja Lee 1#, Seon-Yong Jeong 1#, Mariusz Karbowski 1, Carolyn L Smith 2, Richard J Youle 1* 1 Biochemistry
Regulation of mitochondrial fission and apoptosis by the mitochondrial outer membrane protein hfis1
 Research Article 4141 Regulation of mitochondrial fission and apoptosis by the mitochondrial outer membrane protein hfis1 Tianzheng Yu 1, Randall J. Fox 1, Lindsay S. Burwell 2 and Yisang Yoon 1, * 1 Department
Research Article 4141 Regulation of mitochondrial fission and apoptosis by the mitochondrial outer membrane protein hfis1 Tianzheng Yu 1, Randall J. Fox 1, Lindsay S. Burwell 2 and Yisang Yoon 1, * 1 Department
4) Please cite Dagda et al J Biol Chem 284: , for any publications or presentations resulting from use or modification of the macro.
 Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplement Figure S1. Algorithmic quantification of mitochondrial morphology in SH- SY5Y cells treated with known fission/fusion mediators. Parental SH-SY5Y cells were transiently transfected with an empty
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Supporting Information
 Supporting Information Wang et al. 10.1073/pnas.0804871105 SI Materials and Methods Cell Culture and Transfection. Human neuroblastoma M17 cells were grown as described before (1). Transfection was performed
Supporting Information Wang et al. 10.1073/pnas.0804871105 SI Materials and Methods Cell Culture and Transfection. Human neuroblastoma M17 cells were grown as described before (1). Transfection was performed
Role of Mitochondrial Remodeling in Programmed Cell Death in
 Developmental Cell, Vol. 12 Supplementary Data Role of Mitochondrial Remodeling in Programmed Cell Death in Drosophila melanogaster Gaurav Goyal, Brennan Fell, Apurva Sarin, Richard J. Youle, V. Sriram.
Developmental Cell, Vol. 12 Supplementary Data Role of Mitochondrial Remodeling in Programmed Cell Death in Drosophila melanogaster Gaurav Goyal, Brennan Fell, Apurva Sarin, Richard J. Youle, V. Sriram.
7.06 Problem Set
 7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
7.06 Problem Set 5 -- 2006 1. In the first half of the course, we encountered many examples of proteins that entered the nucleus in response to the activation of a cell-signaling pathway. One example of
Levels of human Fis1 at the mitochondrial outer membrane regulate mitochondrial morphology
 Research Article 1201 Levels of human Fis1 at the mitochondrial outer membrane regulate mitochondrial morphology Diana Stojanovski 1, Olga S. Koutsopoulos 1, Koji Okamoto 2 and Michael T. Ryan 1, * 1 Department
Research Article 1201 Levels of human Fis1 at the mitochondrial outer membrane regulate mitochondrial morphology Diana Stojanovski 1, Olga S. Koutsopoulos 1, Koji Okamoto 2 and Michael T. Ryan 1, * 1 Department
TNFα 18hr. Control. CHX 18hr. TNFα+ CHX 18hr. TNFα: 18 18hr (KDa) PARP. Cleaved. Cleaved. Cleaved. Caspase3. Pellino3 shrna. Control shrna.
 Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Supplementary Information
 Supplementary Information MAP2/Hoechst Hyp.-AP ph 6.5 Hyp.-SD ph 7.2 Norm.-SD ph 7.2 Supplementary Figure 1. Mitochondrial elongation in cortical neurons by acidosis. Representative images of neuronal
Supplementary Information MAP2/Hoechst Hyp.-AP ph 6.5 Hyp.-SD ph 7.2 Norm.-SD ph 7.2 Supplementary Figure 1. Mitochondrial elongation in cortical neurons by acidosis. Representative images of neuronal
The Novel Tail-anchored Membrane Protein Mff Controls Mitochondrial and Peroxisomal Fission in Mammalian Cells
 Molecular Biology of the Cell Vol. 19, 2402 2412, June 2008 The Novel Tail-anchored Membrane Protein Mff Controls Mitochondrial and Peroxisomal Fission in Mammalian Cells Shilpa Gandre-Babbe and Alexander
Molecular Biology of the Cell Vol. 19, 2402 2412, June 2008 The Novel Tail-anchored Membrane Protein Mff Controls Mitochondrial and Peroxisomal Fission in Mammalian Cells Shilpa Gandre-Babbe and Alexander
7.06 Cell Biology EXAM #3 April 21, 2005
 7.06 Cell Biology EXAM #3 April 21, 2005 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #3 April 21, 2005 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
Supplemental Figures S1 S5
 Beyond reduction of atherosclerosis: PON2 provides apoptosis resistance and stabilizes tumor cells Ines Witte (1), Sebastian Altenhöfer (1), Petra Wilgenbus (1), Julianna Amort (1), Albrecht M. Clement
Beyond reduction of atherosclerosis: PON2 provides apoptosis resistance and stabilizes tumor cells Ines Witte (1), Sebastian Altenhöfer (1), Petra Wilgenbus (1), Julianna Amort (1), Albrecht M. Clement
Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc
 OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
Mitofusin 1 and 2 play distinct roles in mitochondrial fusion reactions via GTPase activity
 Research Article 6535 Mitofusin 1 and 2 play distinct roles in mitochondrial fusion reactions via GTPase activity Naotada Ishihara, Yuka Eura and Katsuyoshi Mihara* Department of Molecular Biology, Graduate
Research Article 6535 Mitofusin 1 and 2 play distinct roles in mitochondrial fusion reactions via GTPase activity Naotada Ishihara, Yuka Eura and Katsuyoshi Mihara* Department of Molecular Biology, Graduate
Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant
 9 10 11 Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant Makoto Kazama 1 *, Sanae Ogiwara, Takashi Makiuchi 1, Kazuhiro Yoshida 1, Kumiko Nakada-Tsukui, Tomoyoshi
9 10 11 Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant Makoto Kazama 1 *, Sanae Ogiwara, Takashi Makiuchi 1, Kazuhiro Yoshida 1, Kumiko Nakada-Tsukui, Tomoyoshi
NucView TM 488 Caspase-3 Assay Kit for Live Cells
 NucView TM 488 Caspase-3 Assay Kit for Live Cells Catalog Number: 30029 (100-500 assays) Contact Information Address: Biotium, Inc. 3423 Investment Blvd. Suite 8 Hayward, CA 94545 USA Telephone: (510)
NucView TM 488 Caspase-3 Assay Kit for Live Cells Catalog Number: 30029 (100-500 assays) Contact Information Address: Biotium, Inc. 3423 Investment Blvd. Suite 8 Hayward, CA 94545 USA Telephone: (510)
Dynamin-related Protein Drp1 Is Required for Mitochondrial Division in Mammalian Cells
 Vol. 12, 2245 2256, August 2001 Dynamin-related Protein Drp1 Is Required for Mitochondrial Division in Mammalian Cells Elena Smirnova, Lorena Griparic, Dixie-Lee Shurland, and Alexander M. van der Bliek*
Vol. 12, 2245 2256, August 2001 Dynamin-related Protein Drp1 Is Required for Mitochondrial Division in Mammalian Cells Elena Smirnova, Lorena Griparic, Dixie-Lee Shurland, and Alexander M. van der Bliek*
5 Mitochondrial Dynamics in Mammals
 5 Mitochondrial Dynamics in Mammals Hsiuchen Chen and David C. Chan Division of Biology California Institute of Technology Pasadena, California 91125 I. Introduction II. The Central Molecular Players A.
5 Mitochondrial Dynamics in Mammals Hsiuchen Chen and David C. Chan Division of Biology California Institute of Technology Pasadena, California 91125 I. Introduction II. The Central Molecular Players A.
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
DOI: 10.1038/ncb2362 Figure S1 CYLD and CASPASE 8 genes are co-regulated. Analysis of gene expression across 79 tissues was carried out as described previously [Ref: PMID: 18636086]. Briefly, microarray
hfis1, a novel component of the mammalian mitochondrial fission machinery.
 JBC Papers in Press. Published on June 3, 2003 as Manuscript M303758200 hfis1, a novel component of the mammalian mitochondrial fission machinery. Dominic I James, Philippe A Parone, Yves Mattenberger
JBC Papers in Press. Published on June 3, 2003 as Manuscript M303758200 hfis1, a novel component of the mammalian mitochondrial fission machinery. Dominic I James, Philippe A Parone, Yves Mattenberger
Direct Interaction between Survivin and Smac/DIABLO Is Essential for the Anti-apoptotic Activity of Survivin during Taxol-induced Apoptosis*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 25, Issue of June 20, pp. 23130 23140, 2003 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Direct Interaction
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 25, Issue of June 20, pp. 23130 23140, 2003 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Direct Interaction
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL Figure S1. Mitochondrial morphology in Fis1-null, Mff-null and Fis1/Mff-null MEF cells. (A) Western blotting of lysates from Fis1-null, Mff-null and Fis1/Mff-null cells. Lysates were
SUPPLEMENTAL MATERIAL Figure S1. Mitochondrial morphology in Fis1-null, Mff-null and Fis1/Mff-null MEF cells. (A) Western blotting of lysates from Fis1-null, Mff-null and Fis1/Mff-null cells. Lysates were
Supplementary Figure 1. AnnexinV FITC and Sytox orange staining in wild type, Nlrp3 /, ASC / and casp1/11 / TEC treated with TNF /CHX.
 Supplementary Figure 1. AnnexinV FITC and Sytox orange staining in wild type, Nlrp3 /, ASC / and casp1/11 / TEC treated with TNF /CHX. Phase contrast and widefield fluorescence microscopy (20x magnification).
Supplementary Figure 1. AnnexinV FITC and Sytox orange staining in wild type, Nlrp3 /, ASC / and casp1/11 / TEC treated with TNF /CHX. Phase contrast and widefield fluorescence microscopy (20x magnification).
Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with
 Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with DsRed-mito. Right panels are time-course enlarged images
Supplementary Figure 1. CoMIC in 293T, HeLa, and HepG2 cells. (a) Mitochondrial morphology in 293T, HeLa and HepG2 cells. Cells were transfected with DsRed-mito. Right panels are time-course enlarged images
Plant mitochondrial dynamics
 Plant mitochondrial dynamics Peroxisome Chloroplast Nucleus Mitochondria from Alberts et al. 1994 Living Arabidopsis leaf 10 µm Logan & Leaver (2000) Journal of Experimental Botany, 51: 865-871 5 µm S.
Plant mitochondrial dynamics Peroxisome Chloroplast Nucleus Mitochondria from Alberts et al. 1994 Living Arabidopsis leaf 10 µm Logan & Leaver (2000) Journal of Experimental Botany, 51: 865-871 5 µm S.
Co-localization Analysis of Bak-Drp1 on Mitochondria Using Super-resolution Microscopy
 Co-localization Analysis of Bak-Drp1 on Mitochondria Using Super-resolution Microscopy Danzi Song University of Tokyo Research Internship Program Ozawa Laboratory Abstract The mechanistic connection between
Co-localization Analysis of Bak-Drp1 on Mitochondria Using Super-resolution Microscopy Danzi Song University of Tokyo Research Internship Program Ozawa Laboratory Abstract The mechanistic connection between
Technical Bulletin. Caspase 9 Fluorescein (FLICA) Assay. Catalog Number CS0300 Storage Temperature 2-8 C
 1 Caspase 9 Fluorescein (FLICA) Assay Catalog Number CS0300 Storage Temperature 2-8 C Technical Bulletin Product Description Caspases are detected by immunoprecipitation, immunoblotting with caspase specific
1 Caspase 9 Fluorescein (FLICA) Assay Catalog Number CS0300 Storage Temperature 2-8 C Technical Bulletin Product Description Caspases are detected by immunoprecipitation, immunoblotting with caspase specific
Technical Bulletin. Caspase 13 Fluorescein (FLICA) Assay. Catalog Number CS0320 Storage Temperature 2-8 C
 Caspase 13 Fluorescein (FLICA) Assay Catalog Number CS0320 Storage Temperature 2-8 C Technical Bulletin Product Description Caspases are detected by immunoprecipitation, immunoblotting with caspase specific
Caspase 13 Fluorescein (FLICA) Assay Catalog Number CS0320 Storage Temperature 2-8 C Technical Bulletin Product Description Caspases are detected by immunoprecipitation, immunoblotting with caspase specific
Utah School of Medicine, Salt Lake City, UT Tel.: ; Fax: ;
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 281, NO. 25, pp. 17312 17320, June 23, 2006 2006 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Dimeric Dnm1-G385D Interacts
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 281, NO. 25, pp. 17312 17320, June 23, 2006 2006 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Dimeric Dnm1-G385D Interacts
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins
 13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Reconstructing Mitochondrial Evolution?? Morphological Diversity. Mitochondrial Diversity??? What is your definition of a mitochondrion??
 Reconstructing Mitochondrial Evolution?? What is your definition of a mitochondrion?? Morphological Diversity Mitochondria as we all know them: Suprarenal gland Liver cell Plasma cell Adrenal cortex Mitochondrial
Reconstructing Mitochondrial Evolution?? What is your definition of a mitochondrion?? Morphological Diversity Mitochondria as we all know them: Suprarenal gland Liver cell Plasma cell Adrenal cortex Mitochondrial
The novel conserved mitochondrial inner-membrane protein MTGM regulates mitochondrial morphology and cell proliferation
 2252 Research Article The novel conserved mitochondrial inner-membrane protein MTGM regulates mitochondrial morphology and cell proliferation Jian Zhao 1, *, Tong Liu 1, Shao-Bo Jin 2, Nikolay Tomilin
2252 Research Article The novel conserved mitochondrial inner-membrane protein MTGM regulates mitochondrial morphology and cell proliferation Jian Zhao 1, *, Tong Liu 1, Shao-Bo Jin 2, Nikolay Tomilin
The mitochondrial protein MTP18 contributes to mitochondrial fission in mammalian cells
 Research Article 3049 The mitochondrial protein MTP18 contributes to mitochondrial fission in mammalian cells Daniel Tondera*, Frank Czauderna, Katharina Paulick, Rolf Schwarzer, Jörg Kaufmann and Ansgar
Research Article 3049 The mitochondrial protein MTP18 contributes to mitochondrial fission in mammalian cells Daniel Tondera*, Frank Czauderna, Katharina Paulick, Rolf Schwarzer, Jörg Kaufmann and Ansgar
Stress-Induced Nuclear-to-Cytoplasmic Translocation of Cyclin C Promotes Mitochondrial Fission in Yeast
 Article Stress-Induced Nuclear-to-Cytoplasmic Translocation of Cyclin C Promotes Mitochondrial Fission in Yeast Katrina F. Cooper, 1 Svetlana Khakhina, 1 Stephen K. Kim, 1 and Randy Strich 1, * 1 Department
Article Stress-Induced Nuclear-to-Cytoplasmic Translocation of Cyclin C Promotes Mitochondrial Fission in Yeast Katrina F. Cooper, 1 Svetlana Khakhina, 1 Stephen K. Kim, 1 and Randy Strich 1, * 1 Department
Biochimica et Biophysica Acta
 Biochimica et Biophysica Acta 1832 (2013) 453 474 Contents lists available at SciVerse ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbadis PSAP induces a unique
Biochimica et Biophysica Acta 1832 (2013) 453 474 Contents lists available at SciVerse ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbadis PSAP induces a unique
Vibrio cholerae porin OmpU induces caspase-independent programmed cell death upon translocation to the host cell mitochondria
 JBC Papers in Press. Published on November 11, 2015 as Manuscript M115.670182 The latest version is at http://www.jbc.org/cgi/doi/10.1074/jbc.m115.670182 OmpU induces caspase-independent programmed cell
JBC Papers in Press. Published on November 11, 2015 as Manuscript M115.670182 The latest version is at http://www.jbc.org/cgi/doi/10.1074/jbc.m115.670182 OmpU induces caspase-independent programmed cell
Characterization of the mitochondrial protein LETM1, which maintains the mitochondrial tubular shapes and interacts with the AAA-ATPase BCS1L
 2588 Research Article Characterization of the mitochondrial protein LETM1, which maintains the mitochondrial tubular shapes and interacts with the AAA-ATPase BCS1L Shoko Tamai 1, Hiroshi Iida 2, Sadaki
2588 Research Article Characterization of the mitochondrial protein LETM1, which maintains the mitochondrial tubular shapes and interacts with the AAA-ATPase BCS1L Shoko Tamai 1, Hiroshi Iida 2, Sadaki
Fusion and Fission: Interlinked Processes Critical for Mitochondrial Health
 I R E V I E W S E Review in Advance first posted online on August 29, 2012. (Changes may still occur before final publication online and in print.) N C A D V A N Fusion and Fission: Interlinked Processes
I R E V I E W S E Review in Advance first posted online on August 29, 2012. (Changes may still occur before final publication online and in print.) N C A D V A N Fusion and Fission: Interlinked Processes
Protease Inhibitor Cocktail A (1 tablet / 7 10 ml, Roche Cat# ) Protease inhibitor Cocktail B (0.5ml per 250ml, Calbiochem Cat# )
 Protocol for Western Blotting Tissue/Cell Sample Preparation Lysis Buffer 1 (ph8.0) o 50mM Tris-Cl o 150mM NaCl o 1% v/v NP40 o protease inhibitor cocktail A/B Lysis Buffer 2 (RIPA) (ph 8.0) o 50mM Tris-Cl
Protocol for Western Blotting Tissue/Cell Sample Preparation Lysis Buffer 1 (ph8.0) o 50mM Tris-Cl o 150mM NaCl o 1% v/v NP40 o protease inhibitor cocktail A/B Lysis Buffer 2 (RIPA) (ph 8.0) o 50mM Tris-Cl
Carboxyl-terminal modulator protein induces apoptosis by regulating mitochondrial function in lung cancer cells
 INTERNATIONAL JOURNAL OF ONCOLOGY 40: 1515-1524, 2012 Carboxyl-terminal modulator protein induces apoptosis by regulating mitochondrial function in lung cancer cells SOON-KYUNG HWANG 1,2*, ARASH MINAI-TEHRANI
INTERNATIONAL JOURNAL OF ONCOLOGY 40: 1515-1524, 2012 Carboxyl-terminal modulator protein induces apoptosis by regulating mitochondrial function in lung cancer cells SOON-KYUNG HWANG 1,2*, ARASH MINAI-TEHRANI
Myxoma Virus M11L Blocks Apoptosis through Inhibition of Conformational Activation of Bax at the Mitochondria
 JOURNAL OF VIROLOGY, Feb. 2006, p. 1140 1151 Vol. 80, No. 3 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.3.1140 1151.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Myxoma Virus
JOURNAL OF VIROLOGY, Feb. 2006, p. 1140 1151 Vol. 80, No. 3 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.3.1140 1151.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Myxoma Virus
Membrane depolarization activates the mitochondrial protease OMA1 by stimulating self-cleavage
 Scientific Report Membrane depolarization activates the mitochondrial protease OMA1 by stimulating self-cleavage Kuan Zhang, Huihui Li & Zhiyin Song* Abstract Mitochondrial inner membrane fusion depends
Scientific Report Membrane depolarization activates the mitochondrial protease OMA1 by stimulating self-cleavage Kuan Zhang, Huihui Li & Zhiyin Song* Abstract Mitochondrial inner membrane fusion depends
D The WD40 Protein Caf4p is a Component of the Mitochondrial Fission Machinery and Recruits Dnm1p to Mitochondria
 153 D The WD4 Protein Caf4p is a Component of the Mitochondrial Fission Machinery and Recruits Dnm1p to Mitochondria This chapter represents a further example of the use of MudPIT to analyze the polypeptide
153 D The WD4 Protein Caf4p is a Component of the Mitochondrial Fission Machinery and Recruits Dnm1p to Mitochondria This chapter represents a further example of the use of MudPIT to analyze the polypeptide
Mitofusin-1 protein is a generally expressed mediator of mitochondrial fusion in mammalian cells
 Research Article 2763 Mitofusin-1 protein is a generally expressed mediator of mitochondrial fusion in mammalian cells Ansgar Santel 1, *, Stephan Frank 2,4, Brigitte Gaume 2, Michael Herrler 3, Richard
Research Article 2763 Mitofusin-1 protein is a generally expressed mediator of mitochondrial fusion in mammalian cells Ansgar Santel 1, *, Stephan Frank 2,4, Brigitte Gaume 2, Michael Herrler 3, Richard
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Apoptosis Detection Using the BD Accuri C6 Flow Cytometer
 Apoptosis Detection Using the BD Accuri C6 Flow Cytometer Stacey Roys, Marketing Applications Specialist, BD Biosciences Cyndy Lane, Senior Product Manager, BD Biosciences 23-14519-00 Outline What is apoptosis?
Apoptosis Detection Using the BD Accuri C6 Flow Cytometer Stacey Roys, Marketing Applications Specialist, BD Biosciences Cyndy Lane, Senior Product Manager, BD Biosciences 23-14519-00 Outline What is apoptosis?
Apoptosis in Mammalian Cells
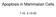 Apoptosis in Mammalian Cells 7.16 2-10-05 Apoptosis is an important factor in many human diseases Cancer malignant cells evade death by suppressing apoptosis (too little apoptosis) Stroke damaged neurons
Apoptosis in Mammalian Cells 7.16 2-10-05 Apoptosis is an important factor in many human diseases Cancer malignant cells evade death by suppressing apoptosis (too little apoptosis) Stroke damaged neurons
Supplemental material
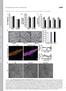 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Measuring Mitochondrial Membrane Potential with JC-1 Using the Cellometer Vision Image Cytometer
 Measuring Mitochondrial Membrane Potential with JC-1 Using the Cellometer Vision Image Cytometer Nexcelom Bioscience LLC. 360 Merrimack Street, Building 9 Lawrence, MA 01843 T: 978.327.5340 F: 978.327.5341
Measuring Mitochondrial Membrane Potential with JC-1 Using the Cellometer Vision Image Cytometer Nexcelom Bioscience LLC. 360 Merrimack Street, Building 9 Lawrence, MA 01843 T: 978.327.5340 F: 978.327.5341
Mitochondria are crucial organelles for life and death of the
 OPA1 requires mitofusin 1 to promote mitochondrial fusion Sara Cipolat, Olga Martins de Brito, Barbara Dal Zilio, and Luca Scorrano* Dulbecco Telethon Institute, Venetian Institute of Molecular Medicine,
OPA1 requires mitofusin 1 to promote mitochondrial fusion Sara Cipolat, Olga Martins de Brito, Barbara Dal Zilio, and Luca Scorrano* Dulbecco Telethon Institute, Venetian Institute of Molecular Medicine,
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
Bypass and interaction suppressors; pathway analysis
 Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Effects of the BH3-only protein human Noxa on mitochondrial dynamics
 FEBS Letters 583 (2009) 2349 2354 journal homepage: www.febsletters.org Effects of the BH3-only protein human Noxa on mitochondrial dynamics Ha-Na Woo a,c,e, Young-Woo Seo b, Ae Ran Moon c, Seon-Yong Jeong
FEBS Letters 583 (2009) 2349 2354 journal homepage: www.febsletters.org Effects of the BH3-only protein human Noxa on mitochondrial dynamics Ha-Na Woo a,c,e, Young-Woo Seo b, Ae Ran Moon c, Seon-Yong Jeong
 http://cwp.embo.org/w09-13/index.html Roads to Ruin is s o t p o ap healthy cell autophagic cell death ne cr os is Thanks to Seamus Martin! Evolution of apoptosis signalling cascades Inhibitor Adopted
http://cwp.embo.org/w09-13/index.html Roads to Ruin is s o t p o ap healthy cell autophagic cell death ne cr os is Thanks to Seamus Martin! Evolution of apoptosis signalling cascades Inhibitor Adopted
HEAVY METAL-INDUCED PROTEINS IN CHLAMYDOMONAS REINHARDTII AND THALASSIOSIRA WEISSFLOGII CELLS
 Proceedings of the 14 th International Conference on Environmental Science and Technology Rhodes, Greece, 3-5 September 2015 HEAVY METAL-INDUCED PROTEINS IN CHLAMYDOMONAS REINHARDTII AND THALASSIOSIRA
Proceedings of the 14 th International Conference on Environmental Science and Technology Rhodes, Greece, 3-5 September 2015 HEAVY METAL-INDUCED PROTEINS IN CHLAMYDOMONAS REINHARDTII AND THALASSIOSIRA
Mitochondrial Fusion and Fission in Mammals
 Annu. Rev. Cell Dev. Biol. 2006. 22:79 99 First published online as a Review in Advance on May 11, 2006 The Annual Review of Cell and Developmental Biology is online at http://cellbio.annualreviews.org
Annu. Rev. Cell Dev. Biol. 2006. 22:79 99 First published online as a Review in Advance on May 11, 2006 The Annual Review of Cell and Developmental Biology is online at http://cellbio.annualreviews.org
Enterovirus 71 2B induces cell apoptosis by directly inducing the conformational activation of the pro-apoptotic protein Bax
 JVI Accepted Manuscript Posted Online 24 August 2016 J. Virol. doi:10.1128/jvi.01499-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
JVI Accepted Manuscript Posted Online 24 August 2016 J. Virol. doi:10.1128/jvi.01499-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Chlamydia trachomatis Infection Inhibits Both Bax and Bak Activation Induced by Staurosporine
 INFECTION AND IMMUNITY, Sept. 2004, p. 5470 5474 Vol. 72, No. 9 0019-9567/04/$08.00 0 DOI: 10.1128/IAI.72.9.5470 5474.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Chlamydia
INFECTION AND IMMUNITY, Sept. 2004, p. 5470 5474 Vol. 72, No. 9 0019-9567/04/$08.00 0 DOI: 10.1128/IAI.72.9.5470 5474.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Chlamydia
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
2. Yeast two-hybrid system
 2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
Cancer Letters 323 (2012) Contents lists available at SciVerse ScienceDirect. Cancer Letters. journal homepage:
 Cancer Letters 323 (212) 62 68 Contents lists available at SciVerse ScienceDirect Cancer Letters journal homepage: www.elsevier.com/locate/canlet Mitofusin 1 inhibits an apoptosis-associated amino-terminal
Cancer Letters 323 (212) 62 68 Contents lists available at SciVerse ScienceDirect Cancer Letters journal homepage: www.elsevier.com/locate/canlet Mitofusin 1 inhibits an apoptosis-associated amino-terminal
M i t o c h o n d r i a
 M i t o c h o n d r i a Dr. Diala Abu-Hassan School of Medicine dr.abuhassand@gmail.com Mitochondria Function: generation of metabolic energy in eukaryotic cells Generation of ATP from the breakdown of
M i t o c h o n d r i a Dr. Diala Abu-Hassan School of Medicine dr.abuhassand@gmail.com Mitochondria Function: generation of metabolic energy in eukaryotic cells Generation of ATP from the breakdown of
According to the manufacture s direction (Pierce), RNA and DNA
 Supplementary method Electrophoretic Mobility-shift assay (EMSA) According to the manufacture s direction (Pierce), RNA and DNA oligonuleotides were firstly labeled by biotin. TAVb (1pM) was incubated
Supplementary method Electrophoretic Mobility-shift assay (EMSA) According to the manufacture s direction (Pierce), RNA and DNA oligonuleotides were firstly labeled by biotin. TAVb (1pM) was incubated
The Caspase System: a potential role in muscle proteolysis and meat quality? Tim Parr
 The Caspase System: a potential role in muscle proteolysis and meat quality? Tim Parr Caroline Kemp, Ron Bardsley,, Peter Buttery Division of Nutritional Sciences, School of Biosciences, University of
The Caspase System: a potential role in muscle proteolysis and meat quality? Tim Parr Caroline Kemp, Ron Bardsley,, Peter Buttery Division of Nutritional Sciences, School of Biosciences, University of
Supporting information for
 Supporting information for Rewiring multi-domain protein switches: transforming a fluorescent Zn 2+ -sensor into a light-responsive Zn 2+ binding protein Stijn J.A. Aper and Maarten Merkx Laboratory of
Supporting information for Rewiring multi-domain protein switches: transforming a fluorescent Zn 2+ -sensor into a light-responsive Zn 2+ binding protein Stijn J.A. Aper and Maarten Merkx Laboratory of
targets. clustering show that different complex pathway
 Supplementary Figure 1. CLICR allows clustering and activation of cytoplasmic protein targets. (a, b) Upon light activation, the Cry2 (red) and LRP6c (green) components co-cluster due to the heterodimeric
Supplementary Figure 1. CLICR allows clustering and activation of cytoplasmic protein targets. (a, b) Upon light activation, the Cry2 (red) and LRP6c (green) components co-cluster due to the heterodimeric
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
mrna Isolation Kit for Blood/Bone Marrow For isolation mrna from blood or bone marrow lysates Cat. No
 For isolation mrna from blood or bone marrow lysates Cat. No. 1 934 333 Principle Starting material Application Time required Results Key advantages The purification of mrna requires two steps: 1. Cells
For isolation mrna from blood or bone marrow lysates Cat. No. 1 934 333 Principle Starting material Application Time required Results Key advantages The purification of mrna requires two steps: 1. Cells
Death-associated Protein 3 Localizes to the Mitochondria and Is Involved in the Process of Mitochondrial Fragmentation during Cell Death*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 279, No. 35, Issue of August 27, pp. 36732 36738, 2004 2004 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Death-associated
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 279, No. 35, Issue of August 27, pp. 36732 36738, 2004 2004 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Death-associated
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
DNA DAMAGE-INDUCED APOPTOSIS: INHIBITION BY CALMODULIN ANTAGONIST, FAS RECEPTOR ANTIBODY AND CASPASE INHIBITORS
 DNA DAMAGE-INDUCED APOPTOSIS: INHIBITION BY CALMODULIN ANTAGONIST, FAS RECEPTOR ANTIBODY AND CASPASE INHIBITORS R. Ray, B. J. Benton, M. E. Burke, T. Rockwood, K. R. Bhat, D. R. Anderson, J. P. Petrali,
DNA DAMAGE-INDUCED APOPTOSIS: INHIBITION BY CALMODULIN ANTAGONIST, FAS RECEPTOR ANTIBODY AND CASPASE INHIBITORS R. Ray, B. J. Benton, M. E. Burke, T. Rockwood, K. R. Bhat, D. R. Anderson, J. P. Petrali,
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
What are mitochondria?
 What are mitochondria? What are mitochondria? An intracellular organelle. There are 100 to 1000s of mitochondria/cell. Most mitochondria come from the mother. Mitochondria have their own DNA Mitochondria
What are mitochondria? What are mitochondria? An intracellular organelle. There are 100 to 1000s of mitochondria/cell. Most mitochondria come from the mother. Mitochondria have their own DNA Mitochondria
Apoptosis: Death Comes for the Cell
 Apoptosis: Death Comes for the Cell Joe W. Ramos joeramos@hawaii.edu From Ingmar Bergman s The Seventh Seal 1 2 Mutations in proteins that regulate cell proliferation, survival and death can contribute
Apoptosis: Death Comes for the Cell Joe W. Ramos joeramos@hawaii.edu From Ingmar Bergman s The Seventh Seal 1 2 Mutations in proteins that regulate cell proliferation, survival and death can contribute
Bax/Bak-Dependent Release of DDP/TIMM8a Promotes Drp1-Mediated Mitochondrial Fission and Mitoptosis during Programmed Cell Death
 Current Biology, Vol. 15, 2112 2118, December 6, 2005, ª2005 Elsevier Ltd All rights reserved. DOI 10.1016/j.cub.2005.10.041 Bax/Bak-Dependent Release of DDP/TIMM8a Promotes Drp1-Mediated Mitochondrial
Current Biology, Vol. 15, 2112 2118, December 6, 2005, ª2005 Elsevier Ltd All rights reserved. DOI 10.1016/j.cub.2005.10.041 Bax/Bak-Dependent Release of DDP/TIMM8a Promotes Drp1-Mediated Mitochondrial
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Scale in the biological world
 Scale in the biological world 2 A cell seen by TEM 3 4 From living cells to atoms 5 Compartmentalisation in the cell: internal membranes and the cytosol 6 The Origin of mitochondria: The endosymbion hypothesis
Scale in the biological world 2 A cell seen by TEM 3 4 From living cells to atoms 5 Compartmentalisation in the cell: internal membranes and the cytosol 6 The Origin of mitochondria: The endosymbion hypothesis
Mitochondrial dynamics in ischemia and reperfusion
 Mitochondrial dynamics in ischemia and reperfusion Derek J Hausenloy Reader in Cardiovascular Medicine, British Heart Foundation Senior Clinical Research Fellow, The Hatter Cardiovascular Institute, University
Mitochondrial dynamics in ischemia and reperfusion Derek J Hausenloy Reader in Cardiovascular Medicine, British Heart Foundation Senior Clinical Research Fellow, The Hatter Cardiovascular Institute, University
SUPPLEMENTARY INFORMATION
 GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
GP2 Type I-piliated bacteria FAE M cell M cell pocket idc T cell mdc Generation of antigenspecific T cells Induction of antigen-specific mucosal immune response Supplementary Figure 1 Schematic diagram
Quantification of Protein Half-Lives in the Budding Yeast Proteome
 Supporting Methods Quantification of Protein Half-Lives in the Budding Yeast Proteome 1 Cell Growth and Cycloheximide Treatment Three parallel cultures (17 ml) of each TAP-tagged strain were grown in separate
Supporting Methods Quantification of Protein Half-Lives in the Budding Yeast Proteome 1 Cell Growth and Cycloheximide Treatment Three parallel cultures (17 ml) of each TAP-tagged strain were grown in separate
Copyright 2013 The Authors. Deposited on: 16 January 2015
 nn Lefebvre, J. et al. (2013) Caspase-generated fragment of the Met receptor favors apoptosis via the intrinsic pathway independently of its tyrosine kinase activity., 4 (10). e871. ISSN 2041-4889 Copyright
nn Lefebvre, J. et al. (2013) Caspase-generated fragment of the Met receptor favors apoptosis via the intrinsic pathway independently of its tyrosine kinase activity., 4 (10). e871. ISSN 2041-4889 Copyright
SEPTEMBER 10, 2010 VOLUME 285 NUMBER 37 JOURNAL OF BIOLOGICAL CHEMISTRY 28749
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 37, pp. 28749 28763, September 10, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Bcl-2 and Bax Interact
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 37, pp. 28749 28763, September 10, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Bcl-2 and Bax Interact
Downloaded from at University of Washington Health Sciences Libraries, on November 12, 2010
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 41, pp. 31590 31602, October 8, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Rab32 Modulates Apoptosis
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 285, NO. 41, pp. 31590 31602, October 8, 2010 2010 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Rab32 Modulates Apoptosis
Supplemental Data. Chen and Thelen (2010). Plant Cell /tpc
 Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
2 The abbreviations used are: FADD, Fas-associated death domain; tbid, truncated
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 287, NO. 34, pp. 29125 29133, August 17, 2012 2012 by The American Society for Biochemistry and Molecular Biology, Inc. Published in the U.S.A. Death Receptor 6
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 287, NO. 34, pp. 29125 29133, August 17, 2012 2012 by The American Society for Biochemistry and Molecular Biology, Inc. Published in the U.S.A. Death Receptor 6
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1
 Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
horseradish peroxidase-labeled anti-mouse secondary antibody were procured from
 SUPPORTING INFORMATION Characterization of anti-platelet properties of silver nanoparticles Siddhartha Shrivastava, Tanmay Bera, Sunil K. Singh, Gajendra Singh, P. Ramachandrarao and Debabrata Dash 1.
SUPPORTING INFORMATION Characterization of anti-platelet properties of silver nanoparticles Siddhartha Shrivastava, Tanmay Bera, Sunil K. Singh, Gajendra Singh, P. Ramachandrarao and Debabrata Dash 1.
A novel mitochondrial ubiquitin ligase plays a critical role in mitochondrial dynamics
 The EMBO Journal (2006) 25, 3618 3626 & 2006 European Molecular Biology Organization All Rights Reserved 0261-4189/06 www.embojournal.org A novel mitochondrial ubiquitin ligase plays a critical role in
The EMBO Journal (2006) 25, 3618 3626 & 2006 European Molecular Biology Organization All Rights Reserved 0261-4189/06 www.embojournal.org A novel mitochondrial ubiquitin ligase plays a critical role in
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1121356/dc1 Supporting Online Material for Polar PIN Localization Directs Auxin Flow in Plants Justyna Wiśniewska, Jian Xu, Daniela Seifertová, Philip B. Brewer, Kamil
www.sciencemag.org/cgi/content/full/1121356/dc1 Supporting Online Material for Polar PIN Localization Directs Auxin Flow in Plants Justyna Wiśniewska, Jian Xu, Daniela Seifertová, Philip B. Brewer, Kamil
Molecular Cell Biology 5068 In Class Exam 1 September 30, Please print your name:
 Molecular Cell Biology 5068 In Class Exam 1 September 30, 2014 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your
Molecular Cell Biology 5068 In Class Exam 1 September 30, 2014 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11419 Supplementary Figure 1 Schematic representation of innate immune signaling pathways induced by intracellular Salmonella in cultured macrophages. a, During the infection Salmonella
doi:10.1038/nature11419 Supplementary Figure 1 Schematic representation of innate immune signaling pathways induced by intracellular Salmonella in cultured macrophages. a, During the infection Salmonella
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
Mitochondria associate with the cytoskeleton. Figure 14-5 Molecular Biology of the Cell ( Garland Science 2008)
 Mitochondria associate with the cytoskeleton Figure 14-5 Molecular Biology of the Cell ( Garland Science 2008) Special arrangements of mitochondria Figure 14-6 Molecular Biology of the Cell ( Garland Science
Mitochondria associate with the cytoskeleton Figure 14-5 Molecular Biology of the Cell ( Garland Science 2008) Special arrangements of mitochondria Figure 14-6 Molecular Biology of the Cell ( Garland Science
cgmp ELISA Kit (Direct Competitive) Based on Monoclonal Anti-cGMP Antibody
 (FOR RESEARCH USE ONLY. DO NOT USE IT IN CLINICAL DIAGNOSIS!) cgmp ELISA Kit (Direct Competitive) Based on Monoclonal Anti-cGMP Antibody Catalog No: E-EL-DS02 96T This manual must be read attentively and
(FOR RESEARCH USE ONLY. DO NOT USE IT IN CLINICAL DIAGNOSIS!) cgmp ELISA Kit (Direct Competitive) Based on Monoclonal Anti-cGMP Antibody Catalog No: E-EL-DS02 96T This manual must be read attentively and
