Over the past three decades, studies at the
|
|
|
- Lambert Watson
- 5 years ago
- Views:
Transcription
1 Reaction-Diffusion Model as a Framework for Understanding iological Pattern Formation Shigeru Kondo 1 * and Takashi Miura 2 The Turing, or reaction-diffusion (RD), model is one of the best-known theoretical models used to explain self-regulated pattern formation in the developing animal embryo. lthough its real-world relevance was long debated, a number of compelling examples have gradually alleviated much of the skepticism surrounding the model. The RD model can generate a wide variety of spatial patterns, and mathematical studies have revealed the kinds of interactions required for each, giving this model the potential for application as an experimental working hypothesis in awidevarietyofmorphologicalphenomena.inthisreview,wedescribetheessenceofthistheory for experimental biologists unfamiliar with the model, using examples from experimental studies in which the RD model is effectively incorporated. Over the past three decades, studies at the molecular level have revealed that a wide range of physiological phenomena are regulated by complex networks of cellular or molecular interactions (1). The complexity of such networks gives rise to new problems, however, as the behavior of such systems often defies immediate or intuitive understanding. Mathematical approaches can help facilitate the understanding of complex systems, and to date, these approaches have taken two primary forms. The first involves analyzing every element of a network quantitatively and simulating all interactions by computation (1). This strategy is effective in relatively simple systems, such as the metabolic pathway in a single cell, and is extensively explored in the field of systems biology. However, for more complex systems in which spatiotemporal parameters take on importance, it becomes almost impossible to make a meaningful prediction. second strategy, one that includes simple mathematical modeling in which the details of the system are omitted, can be more effective in extracting the nature of the complex system (2). The reactiondiffusion (RD) model (3) proposed by lan Turing is a masterpiece of this sort of mathematical modeling, one that can explain how spatial patterns develop autonomously. In the RD model, Turing used a simple system of two diffusible substances interacting with each other to represent patterning mechanisms in the embryo and found that such systems can generate spatial patterns autonomously. The most revolutionary feature of the RD model is its introduction of a reaction that produces the ligands (morphogens). If diffusion alone is at 1 Graduate School of Frontier iosciences, Osaka University, Suita, Osaka, , Japan. 2 Department of natomy and Developmental iology, Kyoto University Graduate School of Medicine, Kyoto , Japan. *To whom correspondence should be addressed. skondo@fbs.osaka-u.ac.jp One morphogen Two morphogens work, local sources of morphogens are needed to form the gradient. In such cases, the positional information made by the system is dependent on the prepattern (Fig. 1, and ). y introducing the reaction, the system gains the ability to generate various patterns independent of the prepattern (Fig. 1). Unfortunately, Turing died soon after publishing this seminal paper, but simulation studies of the model have shown that this system can replicate most biological spatial patterns (4 6). Later, a number of mathematical models (4) were proposed,but most followed Turing s basic idea that the mutual interaction of elements results in spontaneous pattern formation. The RD model is now recognized as a standard among mathematical theories that deal with biological pattern formation. However, this model has yet to gain wide acceptance among experimental biologists. One reason is the gap between the mathematical simplicity of the model and the complexity of the real world. The hypothetical molecules in the original RD model have been so idealized for the purposes of mathematical analysis that it seems nearly impossible to adapt the model directly to the complexity of real biological systems. However, this is a misunderstanding to which experimental researchers tend to succumb. The logic of pattern formation can be understood with simple models, and by adapting this logic to complex biological phenomena, it becomes easier to extract the essence of the underlying mechanisms. Genomic data and new analytic technologies have shifted the target of developmental research from the identification of molecules to understanding the behavior of complex networks, making the RD model even more important as a tool for theoretical analysis. Gradient 1D horizontal 1D vertical Gradients 2D pattern More complicated Interactions "Wave" Spots and stripes Labyrinth Fig. 1. Schematic drawing showing the difference between the morphogen gradient model and Turing model. ()morphogenmoleculeproducedatoneendofanembryoformsagradientbydiffusion. ells know their position from the concentration of the molecule. The gradient is totally dependent on the prepattern of the morphogen source (boundary condition). () ddingasecondmorphogenproducesa relatively complex pattern; but with no interactions between the morphogens, the system is not selfregulating. () Withadditionoftheinteractionsbetweenthemorphogens, thesystembecomesselfregulating and can form a variety of patterns independent of the prepattern. [rt work by S. Miyazawa] 1616
2 In this review, we describe the RD model and its experimental applications, addressing some of the points biologists tend to question. We hope many biologists gain an appreciation for this beautiful theory and take advantage of it in their experimental work. The Original Turing Model t the beginning of his original paper (3), Turing stated that his aim was to show that the combination of known physical elements is sufficient to explain biological pattern formation. The elements selected by Turing were a theoretical pair of interacting molecules diffused in a continuous field. In his mathematical analysis, Turing revealed that such a system yields six potential steady states, depending on the dynamics of reaction term and wavelength of the pattern [Fig. 2 and Supporting Online Material (SOM) Text 1-1]. In case I, the system converges to a stable and uniform state. In case II, a uniform-phase oscillation of the morphogen concentration occurs. Such phase unification is seen in such systems as circadian rhythms (7)and the contraction of heart muscle cells (8). In case III, the system forms salt-and-pepper patterns, such as are made when differentiated cells inhibit the differentiation of neighboring cells. This is seen, for example, with differentiated neuroprogenitor cells in the epithelium of Drosophila embryos (9). ase IV represents an unusual state in which a pattern of the sort made in case III oscillates. No examples of this have been identified in a developmental system. In case V, a traveling wave is generated. iological traveling waves caused by this mechanism include the spiral patterns formed by the social amoeba Dictyostelium discoideum on aggregation (10), and the wave of calcium ions that traverses the egg of the frog Xenopus laevis on sperm entry (11). In case VI, stationary patterns are made. The finding of this type of wave is the major achievement of Turing s analyses,and these are usually referred to as Turing patterns. Turing pattern is akindofnonlinearwavethatismaintainedbythe dynamic equilibrium of the system. Its wavelength is determined by interactions between molecules and their rates of diffusion. Such patterns arise autonomously, independent of any preexisting positional information. [The supporting materials contain an expanded explanation of the RD model for biologists who are unfamiliar with the mathematical description (SOM Text S1-1), as well as a mathematical explanation of how the periodic pattern arises in the system (SOM Text S1-2).] This property makes it possible to explain how patterns arise precisely in, for example, fertilized oocytes, which present little in the way of positional information. The ability of Turing patterns to regenerate autonomously, even after experimentally induced disturbances, is also important and of great utility in explaining the autonomy shown by pattern-forming developmental processes (4, 6). In addition, through the tuning of parameters and boundary conditions, the system underlying Turing pattern formation can generate a nearly limitless variety of spatial patterns. (Figure 2 shows representative two-dimensional patterns made by simulations with the RD model.) The intricate involutions of seashells (5), the exquisite patterning of feathers (12), and the breathtakingly diverse variety of vertebrate skin patterns (13)haveallbeenmodeledwithintheframework of the Turing model (Fig. 2). The remarkable similarity between prediction and reality in these simulations points strongly to Turing mechanisms as the underlying principle in these and other modes of biological patterning. (User-friendly simulation software is available as supporting online material; a manual of the software is in SOM Text 1-3.) ompatibility of RD and Gradient Models Experimental biologists may recall the gradient model, which has been quite effective in explaining pattern-formation events (14). In the classic gradient model, the fixed source of the morphogen at a specific position provides positional information (14)(Fig.1).lthough the assumption of a morphogen source appears to be different from the assumptions of the RD model, it can be introduced into the RD model quite naturally as a boundary condition. In other words, the classic morphogen gradient model can be thought of as the specific case of the RD model in which the reaction term is removed. In many simulation studies, such boundary conditions are used to make the pattern more realistic (6). Recent experimental studies of morphogen gradients have shown that the precision and the robustness of the gradient are secured by the interactions of molecular elements (15). To model such situations, the authors used a mathematical framework that is essentially identical to that of the RD model. oth reaction and boundary conditions are essential to understanding complex real systems, and the RD model is useful for modeling such cases. pplying the Simple RD Model to a omplex Reality The hypothetical molecules in the original RD model (3)are idealized for the purposes of mathematical analysis. It is assumed that they control their own synthesis and that of their counterparts, and diffuse quickly across spaces that would be divided by cellular membranes. Obviously, it is quite difficult to apply such a model directly to complex living systems. oncerted efforts to align theoretical models to real-world systems, however, have begun to bear fruit, pointing to a much broader range of situations in which the general principles underlying the Turing model might apply. Gierer and Meinhardt (16, 17) showed that a system needs only to include a network that combines ashortrange positive feedback with a long-range negative feedback to generate a Turing pattern. This is now accepted as the basic requirement for Turing pattern formation (14, 16). This refinement leaves the types and numbers of reacting factors unspecified, making it much simpler to envision systems that might fit the requirements. The interacting elements need not be limited to molecules, or even to discrete entities; a circuit of cellular signals will do just as well (18). There is also no need for the stimulus to be provided via diffusion; other modes of transmission can achieve the same end result. Theoretical modeling has shown that a relayed series of direct cell-to-cell signals can form a wave having properties similar to one formed by diffusible factors (19). Other forms of signaling, including chemotactic cell migration (20), mechanochemical activity (21), and neuronal interactions [as in the Swindale model (22) ofoculardominancestripeformation],are also capable of forming Turing-like patterns. For all of these systems, a similar periodic pattern is formed when the condition of short-range positive feedback with long-range negative feedback is satisfied. Why systems represented by apparently different equations behave similarly, and how much the capacity for pattern forming differs among them, are the important subjects from the mathematics perspective. ut if the dynamics of the systems are nearly the same, experimental researchers can select any of the models as their working hypothesis. In the case of fish skin patterning, although experiments apparently suggest the involvement of a nondiffusing signal transduction mechanism, the simplest RD model can predict the movement of the pattern during fish growth (23) and the unusual patterns seen in hybrid fish (24). Finding Turing Patterns in Real Systems During embryogenesis, a great variety of periodic structures develop from various nonperiodic cell or tissue sources, suggesting that waves of the sort generated by Turing or related mechanisms may underlie a wide range of developmental processes. Using modern genetic and molecular techniques, it is possible to identify putative elements of interactive networks that fulfill the criteria of short-range positive feedback and long-range negative feedback, but finding the network alone is not enough. Skeptics rightly point out that just because there is water, it doesn t mean there are waves. No matter how vividly or faithfully a mathematical simulation might replicate an actual biological pattern, this alone does not constitute proof that the simulated state reflects the reality. This has been another major hurdle in identifying compelling examples of Turing patterns in living systems. The solution, however, is not so complicated; to show that a wave exists, we need to identify the dynamic properties of the pattern that is predicted by the computer simulation. Experimental demonstrations have focused on pattern formation in the skin, because the specific characteristics of Turing patterns are more evident in two dimensions than in one. Turing Patterns in Vertebrate Skin Observation of the dynamic properties of Turing patterns in nature was made by Kondo and sai SIENE VOL SEPTEMER
3 Initial condition Six stable states I II III Uniform, stationary Uniform, oscillating Stationary waves with extremely short wavelength oth morphogens diffuse and react with each other IV V VI ase V Oscillatory cases with extremely short wavelength Oscillatory cases with finite wavelength ase VI (Turing pattern) Stationary waves with finite wavelength (Turing pattern) Fig. 2. Schematic drawing showing the mathematical analysis of the RD system and the patterns generated by the simulation. () Six stable states toward which the two-factor RD system can converge. () Two-dimensional patterns generated by the Turing model. These patterns were made by an identical equation with slightly different parameter values. These simulations were calculated by the software provided as supporting online material. () Reproduction of biological patterns created by modified RD mechanisms. With modification, the RD mechanism can generate more complex patterns such as those seen in the real organism. Simulation images are courtesy of H. Meinhardt [sea shell pattern (5)] and. R. Sandersen [fish pattern (13)]. Photos of actual seashells are from ishougai-hp ( Images of popper fish are courtesy of Massimo oyer ( 1618
4 in a study of horizontal stripes in the tropical fish, Pomacanthus imperator (25). They have recently shown that this dynamic nature is shared by many fish species, including the well-established model organism, zebrafish. lthough zebrafish stripes may appear to be stationary, experimental perturbation of the pattern triggers slow changes (23). Following laser ablation of pigment cells in apairofblackhorizontalstripes,the lower line shifts upward before stabilizing in a ell-like curve (Fig. 3) (23). s a result, the spatial interval between the lines is maintained, even when their direction changes. This striking behavior is predicted by simulation (Fig. 3). Fortunately, the zebrafish is amenable to a variety of experimental approaches that may lead to the identification of the circuit of interactions that generates these patterns (26). Work to date has shown that the skin patterns of this fish are set up and maintained by interactions between pigmented cells (26). Nakamasu worked out the interaction network among the pigment cells. lthough the shape of the network is different from that of the original Turing model, it fits the short-range positive, long-range negative feedback description (18). (The mutual inhibition between black and yellow cells behaves as a positive feedback loop, as the expansion of black cells weakens their counterpart.) Identification of the genetic factors involved in the zebrafish pigmentation is under way (26), and it is hoped that this will clarify the details of the signaling network beneath the waves in the skin of fish, and potentially all vertebrates. Many similar surface patterns are seen in invertebrates and plants. We suppose that in these cases an essentially similar mechanism (RD mechanism) is involved, although their molecular basis may be different. Other well-studied examples include the regular disposition of feather buds in chick (27), and of hair follicles in mice (28). Jung et al.showed that the spatially periodic pattern of feather buds regenerates even when the skin is recombined from dissociated cells (27). In the case of mouse hair follicles, alteration of the amount of putative key factors changes the pattern in a manner predicted by computer simulation. Sick et al.used overexpression and inhibition of Wnt and Dkk in fetal mouse to study how such perturbations might affect the patterning of follicle formation Day 13 Day 16 Day 20 Day 23 (28). y simulation, they first predicted that a ringed pattern of Wnt gene expression that does not occur in nature would result if ectopic production of a Wnt protein were to be controlled appropriately, and then confirmed this experimentally. They suggested that Wnt serves as a short-range activator, and Dkk as a long-range inhibitor, in this system (28). This pair of factors Long-range negative feedback Short-range positive feedback Fig. 3. Movement of zebrafish stripes and the interaction network among the pigment cells. The pigment pattern of zebrafish is composed of black pigment cells (melanophores) and yellow pigment cells (xanthophores). The pattern is made by the mutual interaction between these cells. ()Melanophores in the two black stripeswere ablated by laser, and the process of recovery was recorded. () Results of simulation by the Turing model. ()Interactionnetworkbetweenthepigmentcellsdeducedby experiments. The red arrow represents a long-range positive (enhancing) effect, whereas the blue line with the end bar represents a short-range negative (inhibitory) effect. circuit of two negative interactions functions as a positive feedback. functions in various patterning processes as well, making this a critically important result for the intimations it provides of a wider role for Turing patterns in development (Fig. 4). Interestingly, the growth of new hair relying on interactions between neighboring follicles proceeds even in adult mice, and in one case a mutant was identified in which a traveling wave of hair formation gradually moved across the skin over the life of animals carrying this mutation (29). Plikus et al.(29) have shown how the factors FGF (fibroblast growth factor) and MP (bone morphogenetic protein) function to generate the traveling wave (case V in the Turing model). Other Potential Turing-Driven Developmental Phenomena Establishment of right-left asymmetry in vertebrates is triggered by the unidirectional rotation of cilia at the node, followed by the interaction of Nodal and Lefty that amplify and stabilize faint differences in gene expression (30, 31). Nodal enhances both its own expression and that of Lefty, and Lefty inhibits the activity of Nodal. The fact that Lefty spreads further than Nodal suggests that the inhibitory interaction propagates more quickly than does its activating counterpart, which as we have seen, indicates that this system fulfills the fundamental requirements for Turing pattern formation (30) (Fig.4). In vertebrate limb development, precartilage condensation, which is later replaced by skeletal bone, occurs periodically along the anteriorposterior axis of the distal tip region. ecause this patterning occurs without any periodic prepattern, the Turing model has long been suggested to describe the underlying mechanism (32). In this system, transforming growth factor b (TGF-b) has been invoked as a candidate for the activator molecule (33). TGF-b can stimulate its own production and trigger precartilage condensation, and the sites of incipient condensation exert a laterally acting inhibitory effect on chondrogenesis. lthough no candidate inhibitor has been identified, an interaction network comprising TGF-b function and precartilage condensation may satisfy the short-range activation and long-range inhibition criteria (33). Miura and Shiota have shown that nearly periodic spatial patterns of chondrogenesis occur in the culture of dissociated limb cells in vitro, SIENE VOL SEPTEMER
5 Short-range positive feedback Wnt Nodal TGF- or FGF ondensation of cells Long-range negative feedback Dkk Lefty? MP and that the addition of TGF-b changes the pattern in a manner consistent with the predictions of Turing s model(34). Wnt and Dkk are also essential for lung branching in vertebrates (35)andplayakeyrole in head regeneration in Hydra (36), in which involvement of the Turing mechanism has long been suggested by theoretical and experimental studies. Involvement of the same molecules does not directly prove that the same dynamic mechanism is at work. However, because they constitute the self-regulated system that is robust to artificial perturbations, it is reasonable to expect that the Turing mechanism may underlie these phenomena as it does those in the skin. Watanabe et al. have shown that signal transduction via the gap junction (connexin41.8) plays a key role in pigment pattern formation in zebrafish (37). Loss of function of the connexin gene (leopard) reduces the spatial periodicity and changes the pattern from stripes to spots (37). mutation in another gap junction gene, connexin43, shortens each fin ray, resulting in shortened fins (38). lthough the detailed molecular mechanism underlying this phenomenon has yet to be elucidated, it is possible that the same mechanism functions in other processes of animal development. The above examples are by no means an exhaustive list of the candidates currently being examined as potential biological Turingpatterns. More detailed discussions are provided in (4)and(6). Identification of the specific dynamics of the RD system is critical to showing the applicability of the Turing mechanism to the formation of a given pattern. In the case of pigment pattern formation, it was possible to disturb the pattern and observe the process of regeneration. In most other systems, such observation is complicated because experimental perturbations may be lethal. This is one reason why it is difficult to demonstrate RD activity in some biological systems. (In SOM Text 1-4, we have summarized the important points for researchers to keep in mind when they use the RD model as the working hypothesis.) However, recent technological advances in imaging technologies are assisting such studies. Moreover, the artificial generation of Turing patterns in cell culture should be possible in the near future as the result of synthetic biology (39). Turing was born in 1912 and published his RD model in lthough he was unable to witness the impact of his hypothesis on the work of contemporary biologists, we are hopeful that with an increased acceptance among experimental biologists of the principles he first elucidated, we will see Turing s mechanism take its place as a model for the understanding of spatial pattern formation in living systems. Hair follicle spacing Hydra regeneration Lung branching Left-right asymmetry Feather bud spacing in chick Tooth pattern Lung branching Skeletal pattern in limb Fig. 4. Possible networks of protein ligands may give rise to Turing patterns in the embryo. Shown above are candidates for the RD mechanism proposed by molecular experiments. For a detailed explanation of each network, refer to the text and the articles listed in the references. In all these cases, the network is identical to that of the activator-inhibitor model proposed by Gierer and Meinhardt (17). ondensation of cells by migration into a local region causes sparse distribution of cells in a neighboring region. This can also function as long-range inhibition.(note that the involvement of the RD mechanism in some of the phenomena above has not been fully accepted by experimental researchers.) References and Notes 1. U. lon, n Introduction to Systems iology (hapman & Hall/R, oca Raton, FL, 2006) 2. R. M. May, Nature 261, 459(1976). 3.. M. Turing, ull. Math. iol. 52, 153,discussion119 (1990). 4. J. Murray, Mathematical iology (Springer-Verlag, erlin, ed. 4, 2003). 5. H. Meinhardt, The lgorithmic eauty of Seashells (Springer-Verlag, erlin, 1995). 6. H. Meinhardt, Models of iological Pattern Formation (cademic Press, London, 1982). 7. H. R. Ueda, old Spring Harb. Symp. Quant. iol. 72, 365 (2007). 8. L. S. Song et al., nn. N. Y. cad. Sci. 1047, 99 (2005). 9.. V. abrera, Development 110, 733(1990). 10. D. Dormann,. J. Weijer, Development 128, 4535 (2001) Sardet, F. Roegiers, R. Dumollard,. Rouviere,. McDougall, iophys. hem. 72, 131(1998). 12. M. P. Harris, S. Williamson, J. F. Fallon, H. Meinhardt, R. O. Prum, Proc. Natl. cad. Sci. U.S.. 102, (2005) R. Sanderson, R. M. Kirby,. R. Johnson, L. Yang, J. Graphics GPU Game Tools 11, 47 (2006). 14. L. Wolpert, Principles of Development (Oxford Univ. Press, New York, 2006). 15. T. Gregor, D. W. Tank, E. F. Wieschaus, W. ialek, ell 130, 153(2007). 16. H. Meinhardt,. Gierer, ioessays 22, 753(2000). 17. H. Meinhardt,. Gierer, J. ell Sci. 15, 321 (1974) Nakamasu, G. Takahashi,. Kanbe, S. Kondo, Proc. Natl. cad. Sci. U.S.. 106, 8429(2009). 19. E. M. Rauch, M. M. Millonas, J. Theor. iol. 226, 401 (2004). 20. P. K. Maini, M. R. Myerscough, K. H. Winters, J. D. Murray, ull. Math. iol. 53, 701(1991). 21. J. D. Murray, G. F. Oster,. K. Harris, J. Math. iol. 17, 125 (1983). 22. N. V. Swindale, Proc. R. Soc. Lond. iol. Sci. 208, 243 (1980). 23. M. Yamaguchi, E. Yoshimoto, S. Kondo, Proc. Natl. cad. Sci. U.S.. 104, 4790(2007). 24. R. sai, E. Taguchi, Y. Kume, M. Saito, S. Kondo, Mech. Dev. 89, 87 (1999). 25. S. Kondo, R. sai, Nature 376, 765(1995). 26. D. M. Parichy, Semin. ell Dev. iol. 20, 63 (2009). 27. H. S. Jung et al., Dev. iol. 196, 11(1998). 28. S. Sick, S. Reinker, J. Timmer, T. Schlake, Science 314, 1447 (2006). 29. M. V. Plikus et al., Nature 451, 340(2008). 30. T. Nakamura et al., Dev. ell 11, 495(2006) Thisse,. V. Wright,. Thisse, Nature 403, 425 (2000). 32. S.. Newman, H. L. Frisch, Science 205, 662 (1979). 33. S.. Newman, Int. J. Dev. iol. 53, 663(2009). 34. T. Miura, K. Shiota, Dev. Dyn. 217, 241(2000). 35. S. P. De Langhe et al., Dev. iol. 277, 316(2005). 36. R. ugustin et al., Dev. iol. 296, 62 (2006). 37. M. Watanabe et al., EMO Rep. 7, 893(2006). 38. M. K. Iovine, E. P. Higgins,. Hindes,. oblitz, S. L. Johnson, Dev. iol. 278, 208(2005). 39. D. Sprinzak, M.. Elowitz, Nature 438, 443 (2005). 40. We thank D. Sipp for suggestions and help in preparation of the manuscript. We also thank H. Meinhardt,. R. Sandersen, and P. Przemyslaw for figures reproduced from their original works. Supporting Online Material SOM Text Figs. S1 to S4 References omputer Simulation /science
Interactions between zebrafish pigment cells responsible for the generation of Turing patterns
 Interactions between zebrafish pigment cells responsible for the generation of Turing patterns Akiko Nakamasu a, Go Takahashi a, Akio Kanbe a, and Shigeru Kondo a,b,1 a Department of Biological Science,
Interactions between zebrafish pigment cells responsible for the generation of Turing patterns Akiko Nakamasu a, Go Takahashi a, Akio Kanbe a, and Shigeru Kondo a,b,1 a Department of Biological Science,
Synchronous oscillations in biological systems are inevitable yet enigmatic
 Coupling hair follicle cycles to produce oscillating patterns Megan Jeans, Rice University Mentor: G. Bard Ermentrout, University of Pittsburgh June 6 th, 2006 Introduction Synchronous oscillations in
Coupling hair follicle cycles to produce oscillating patterns Megan Jeans, Rice University Mentor: G. Bard Ermentrout, University of Pittsburgh June 6 th, 2006 Introduction Synchronous oscillations in
Drosophila melanogaster- Morphogen Gradient
 NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
DIFFERENTIATION MORPHOGENESIS GROWTH HOW CAN AN IDENTICAL SET OF GENETIC INSTRUCTIONS PRODUCE DIFFERENT TYPES OF CELLS?
 DIFFERENTIATION HOW CAN AN IDENTICAL SET OF GENETIC INSTRUCTIONS PRODUCE DIFFERENT TYPES OF CELLS? MORPHOGENESIS HOW CAN CELLS FORM ORDERED STRUCTURES? GROWTH HOW DO OUR CELLS KNOW WHEN TO STOP DIVIDING
DIFFERENTIATION HOW CAN AN IDENTICAL SET OF GENETIC INSTRUCTIONS PRODUCE DIFFERENT TYPES OF CELLS? MORPHOGENESIS HOW CAN CELLS FORM ORDERED STRUCTURES? GROWTH HOW DO OUR CELLS KNOW WHEN TO STOP DIVIDING
The Chemical Basis of Morphogenesis
 The Chemical Basis of Morphogenesis A. M. Turing Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, Vol. 237, No. 641. (Aug. 14, 1952), pp. 37-72. Early Life: The
The Chemical Basis of Morphogenesis A. M. Turing Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, Vol. 237, No. 641. (Aug. 14, 1952), pp. 37-72. Early Life: The
Diffusion, Reaction, and Biological pattern formation
 Diffusion, Reaction, and Biological pattern formation Morphogenesis and positional information How do cells know what to do? Fundamental questions How do proteins in a cell segregate to front or back?
Diffusion, Reaction, and Biological pattern formation Morphogenesis and positional information How do cells know what to do? Fundamental questions How do proteins in a cell segregate to front or back?
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Morphogens in biological development: Drosophila example
 LSM5194 Morphogens in biological development: Drosophila example Lecture 29 The concept of morphogen gradients The concept of morphogens was proposed by L. Wolpert as a part of the positional information
LSM5194 Morphogens in biological development: Drosophila example Lecture 29 The concept of morphogen gradients The concept of morphogens was proposed by L. Wolpert as a part of the positional information
Simulation of cell-like self-replication phenomenon in a two-dimensional hybrid cellular automata model
 Simulation of cell-like self-replication phenomenon in a two-dimensional hybrid cellular automata model Takeshi Ishida Nippon Institute of Technology ishida06@ecoinfo.jp Abstract An understanding of the
Simulation of cell-like self-replication phenomenon in a two-dimensional hybrid cellular automata model Takeshi Ishida Nippon Institute of Technology ishida06@ecoinfo.jp Abstract An understanding of the
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Exam: Multiscale Mathematical Biology
 Exam: Multiscale Mathematical Biology Roeland Merks 15 januari 2016 Note: Questions are phrased in English. Answers in Dutch or in English are both acceptable. Citations to the literature are given for
Exam: Multiscale Mathematical Biology Roeland Merks 15 januari 2016 Note: Questions are phrased in English. Answers in Dutch or in English are both acceptable. Citations to the literature are given for
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
6.3.4 Action potential
 I ion C m C m dφ dt Figure 6.8: Electrical circuit model of the cell membrane. Normally, cells are net negative inside the cell which results in a non-zero resting membrane potential. The membrane potential
I ion C m C m dφ dt Figure 6.8: Electrical circuit model of the cell membrane. Normally, cells are net negative inside the cell which results in a non-zero resting membrane potential. The membrane potential
Welcome to AP BIOLOGY!!!! NOTES: Chapter 1 Exploring Life
 Welcome to AP BIOLOGY!!!! NOTES: Chapter 1 Exploring Life The phenomenon we call life defies a simple, onesentence definition Exploring LIFE: We recognize life by what living things DO Some Properties
Welcome to AP BIOLOGY!!!! NOTES: Chapter 1 Exploring Life The phenomenon we call life defies a simple, onesentence definition Exploring LIFE: We recognize life by what living things DO Some Properties
Structural Properties of Generative Form by Hormonal Proliferation Algorithm
 Original Paper Forma, 15, 103 107, 2000 Structural Properties of Generative Form by Hormonal Proliferation Algorithm Yasuo YONEZAWA 1 and Keisuke OHTOMO 2 Division of System Engineering, Graduate School
Original Paper Forma, 15, 103 107, 2000 Structural Properties of Generative Form by Hormonal Proliferation Algorithm Yasuo YONEZAWA 1 and Keisuke OHTOMO 2 Division of System Engineering, Graduate School
Fundamentals of Bio-architecture SUMMARY
 Fundamentals of Bio-architecture SUMMARY Melik Demirel, PhD *Pictures and tables in this lecture notes are copied from Internet sources for educational use only. Can order increase without breaking 2 nd
Fundamentals of Bio-architecture SUMMARY Melik Demirel, PhD *Pictures and tables in this lecture notes are copied from Internet sources for educational use only. Can order increase without breaking 2 nd
Big Idea 1: The process of evolution drives the diversity and unity of life.
 Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
AP Curriculum Framework with Learning Objectives
 Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Collective Effects. Equilibrium and Nonequilibrium Physics
 1 Collective Effects in Equilibrium and Nonequilibrium Physics: June 19, 2006 1 Collective Effects in Equilibrium and Nonequilibrium Physics Website: http://cncs.bnu.edu.cn/mccross/course/ Caltech Mirror:
1 Collective Effects in Equilibrium and Nonequilibrium Physics: June 19, 2006 1 Collective Effects in Equilibrium and Nonequilibrium Physics Website: http://cncs.bnu.edu.cn/mccross/course/ Caltech Mirror:
Adventures in Multicellularity
 Adventures in Multicellularity The social amoeba (a.k.a. slime molds) Dictyostelium discoideum Dictyostelium discoideum the most studied of the social amoebae / cellular slime molds predatory soil amoeba
Adventures in Multicellularity The social amoeba (a.k.a. slime molds) Dictyostelium discoideum Dictyostelium discoideum the most studied of the social amoebae / cellular slime molds predatory soil amoeba
Enduring understanding 1.A: Change in the genetic makeup of a population over time is evolution.
 The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Systems Biology Across Scales: A Personal View XXIII. Spatial Patterns in Biology: Turing mechanism. Sitabhra Sinha IMSc Chennai
 Systems Biology Across Scales: A Personal View XXIII. Spatial Patterns in Biology: Turing mechanism Sitabhra Sinha IMSc Chennai The magnificent patterns of Dr Turing Question: How to explain the development
Systems Biology Across Scales: A Personal View XXIII. Spatial Patterns in Biology: Turing mechanism Sitabhra Sinha IMSc Chennai The magnificent patterns of Dr Turing Question: How to explain the development
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
Map of AP-Aligned Bio-Rad Kits with Learning Objectives
 Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Reception The target cell s detection of a signal coming from outside the cell May Occur by: Direct connect Through signal molecules
 Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
PDEs for self-organization in Cellular and Developmental Biology
 REVIEW ARTICLE PDEs for self-organization in Cellular and Developmental Biology Baker RE 1, Gaffney EA 1 and Maini PK 1,2 1 Centre for Mathematical Biology, Mathematical Institute, University of Oxford,
REVIEW ARTICLE PDEs for self-organization in Cellular and Developmental Biology Baker RE 1, Gaffney EA 1 and Maini PK 1,2 1 Centre for Mathematical Biology, Mathematical Institute, University of Oxford,
Plant and animal cells (eukaryotic cells) have a cell membrane, cytoplasm and genetic material enclosed in a nucleus.
 4.1 Cell biology Cells are the basic unit of all forms of life. In this section we explore how structural differences between types of cells enables them to perform specific functions within the organism.
4.1 Cell biology Cells are the basic unit of all forms of life. In this section we explore how structural differences between types of cells enables them to perform specific functions within the organism.
Where Do Bat Wings Come From?
 Where o at Wings ome From? 1 ats are the only mammals that have evolved the power of flight. They can avoid obstacles and slip through tight spaces. Many species are nocturnal and use echolocation to guide
Where o at Wings ome From? 1 ats are the only mammals that have evolved the power of flight. They can avoid obstacles and slip through tight spaces. Many species are nocturnal and use echolocation to guide
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Principles of Experimental Embryology
 Biology 4361 Developmental Biology Principles of Experimental Embryology June 16, 2008 Overview What forces affect embryonic development? The embryonic environment: external and internal How do forces
Biology 4361 Developmental Biology Principles of Experimental Embryology June 16, 2008 Overview What forces affect embryonic development? The embryonic environment: external and internal How do forces
Lectures on Medical Biophysics Department of Biophysics, Medical Faculty, Masaryk University in Brno. Biocybernetics
 Lectures on Medical Biophysics Department of Biophysics, Medical Faculty, Masaryk University in Brno Norbert Wiener 26.11.1894-18.03.1964 Biocybernetics Lecture outline Cybernetics Cybernetic systems Feedback
Lectures on Medical Biophysics Department of Biophysics, Medical Faculty, Masaryk University in Brno Norbert Wiener 26.11.1894-18.03.1964 Biocybernetics Lecture outline Cybernetics Cybernetic systems Feedback
The student might demonstrate the ability to achieve this standard by: Making a chart comparing the similar functions of plant and animal cells.
 1: Life, Cell Biology: All living organisms are composed of cells, from just one to many trillions, whose details are usually visible only through a microscope. s 1.a) Cells function similarly in all living
1: Life, Cell Biology: All living organisms are composed of cells, from just one to many trillions, whose details are usually visible only through a microscope. s 1.a) Cells function similarly in all living
MBios 401/501: Lecture 14.2 Cell Differentiation I. Slide #1. Cell Differentiation
 MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
Essential knowledge 1.A.2: Natural selection
 Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Systems Biology Across Scales: A Personal View XXVI. Waves in Biology: From cells & tissue to populations. Sitabhra Sinha IMSc Chennai
 Systems Biology Across Scales: A Personal View XXVI. Waves in Biology: From cells & tissue to populations Sitabhra Sinha IMSc Chennai Spiral Waves in Biological Excitable Media Ca ++ waves in cytoplasm
Systems Biology Across Scales: A Personal View XXVI. Waves in Biology: From cells & tissue to populations Sitabhra Sinha IMSc Chennai Spiral Waves in Biological Excitable Media Ca ++ waves in cytoplasm
Valley Central School District 944 State Route 17K Montgomery, NY Telephone Number: (845) ext Fax Number: (845)
 Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Cell Biology. AQA Biology topic 1
 Cell Biology AQA Biology topic 1 1.1 Cell Structure Plant and Animal cells (eukaryotic cells) Eukaryotic cells have these features: 1) Cytoplasm 2) Genetic material within a nucleus 3) Cell Membrane Typical
Cell Biology AQA Biology topic 1 1.1 Cell Structure Plant and Animal cells (eukaryotic cells) Eukaryotic cells have these features: 1) Cytoplasm 2) Genetic material within a nucleus 3) Cell Membrane Typical
StarLogo Simulation of Streaming Aggregation. Demonstration of StarLogo Simulation of Streaming. Differentiation & Pattern Formation.
 StarLogo Simulation of Streaming Aggregation 1. chemical diffuses 2. if cell is refractory (yellow) 3. then chemical degrades 4. else (it s excitable, colored white) 1. if chemical > movement threshold
StarLogo Simulation of Streaming Aggregation 1. chemical diffuses 2. if cell is refractory (yellow) 3. then chemical degrades 4. else (it s excitable, colored white) 1. if chemical > movement threshold
Plant and animal cells (eukaryotic cells) have a cell membrane, cytoplasm and genetic material enclosed in a nucleus.
 4.1 Cell biology Cells are the basic unit of all forms of life. In this section we explore how structural differences between types of cells enables them to perform specific functions within the organism.
4.1 Cell biology Cells are the basic unit of all forms of life. In this section we explore how structural differences between types of cells enables them to perform specific functions within the organism.
Questions in developmental biology. Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration
 Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Study of life. Premedical course I Biology
 Study of life Premedical course I Biology Biology includes among other two different approaches understanding life via study on the smallest level of life Continuity of life is based on genetic information.
Study of life Premedical course I Biology Biology includes among other two different approaches understanding life via study on the smallest level of life Continuity of life is based on genetic information.
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Revision Based on Chapter 25 Grade 11
 Revision Based on Chapter 25 Grade 11 Biology Multiple Choice Identify the choice that best completes the statement or answers the question. 1. A cell that contains a nucleus and membrane-bound organelles
Revision Based on Chapter 25 Grade 11 Biology Multiple Choice Identify the choice that best completes the statement or answers the question. 1. A cell that contains a nucleus and membrane-bound organelles
TEST SUMMARY AND FRAMEWORK TEST SUMMARY
 Washington Educator Skills Tests Endorsements (WEST E) TEST SUMMARY AND FRAMEWORK TEST SUMMARY BIOLOGY Copyright 2014 by the Washington Professional Educator Standards Board 1 Washington Educator Skills
Washington Educator Skills Tests Endorsements (WEST E) TEST SUMMARY AND FRAMEWORK TEST SUMMARY BIOLOGY Copyright 2014 by the Washington Professional Educator Standards Board 1 Washington Educator Skills
Examples of Excitable Media. Excitable Media. Characteristics of Excitable Media. Behavior of Excitable Media. Part 2: Cellular Automata 9/7/04
 Examples of Excitable Media Excitable Media Slime mold amoebas Cardiac tissue (& other muscle tissue) Cortical tissue Certain chemical systems (e.g., BZ reaction) Hodgepodge machine 9/7/04 1 9/7/04 2 Characteristics
Examples of Excitable Media Excitable Media Slime mold amoebas Cardiac tissue (& other muscle tissue) Cortical tissue Certain chemical systems (e.g., BZ reaction) Hodgepodge machine 9/7/04 1 9/7/04 2 Characteristics
Nervous Systems: Neuron Structure and Function
 Nervous Systems: Neuron Structure and Function Integration An animal needs to function like a coherent organism, not like a loose collection of cells. Integration = refers to processes such as summation
Nervous Systems: Neuron Structure and Function Integration An animal needs to function like a coherent organism, not like a loose collection of cells. Integration = refers to processes such as summation
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Standards Map Basic Comprehensive Program Science Grade Seven Focus on Life Sciences SE/TE: , SE/TE: ,
 Publisher: Pearson Program Title: Biology: Exploring Life 2006 Components: 0-13-250-925-3 (SE), 0-13-250-883-4 (TE) Grade Level(s): 7-12 s Map Basic Comprehensive Program Science Grade Seven Focus on Life
Publisher: Pearson Program Title: Biology: Exploring Life 2006 Components: 0-13-250-925-3 (SE), 0-13-250-883-4 (TE) Grade Level(s): 7-12 s Map Basic Comprehensive Program Science Grade Seven Focus on Life
Plant Stimuli pp Topic 3: Plant Behaviour Ch. 39. Plant Behavioural Responses. Plant Hormones. Plant Hormones pp
 Topic 3: Plant Behaviour Ch. 39 Plants exist in environments that are constantly changing. Like animals, plants must be able to detect and react to stimuli in the environment. Unlike animals, plants can
Topic 3: Plant Behaviour Ch. 39 Plants exist in environments that are constantly changing. Like animals, plants must be able to detect and react to stimuli in the environment. Unlike animals, plants can
1. The basic structural and physiological unit of all living organisms is the A) aggregate. B) organelle. C) organism. D) membrane. E) cell.
 Name: Date: Test File Questions 1. The basic structural and physiological unit of all living organisms is the A) aggregate. B) organelle. C) organism. D) membrane. E) cell. 2. A cell A) can be composed
Name: Date: Test File Questions 1. The basic structural and physiological unit of all living organisms is the A) aggregate. B) organelle. C) organism. D) membrane. E) cell. 2. A cell A) can be composed
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Bio 127 Section I Introduction to Developmental Biology. Cell Cell Communication in Development. Developmental Activities Coordinated in this Way
 Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
growth, the replacement of damaged cells, and development from an embryo into an adult. These cell division occur by means of a process of mitosis
 The Cellular Basis of Reproduction and Inheritance PowerPoint Lectures for Campbell Biology: Concepts & Connections, Seventh Edition Reece, Taylor, Simon, and Dickey Biology 1408 Dr. Chris Doumen Lecture
The Cellular Basis of Reproduction and Inheritance PowerPoint Lectures for Campbell Biology: Concepts & Connections, Seventh Edition Reece, Taylor, Simon, and Dickey Biology 1408 Dr. Chris Doumen Lecture
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
An Introduction to Animal Diversity
 An Introduction to Animal Diversity What defines an animal? Why so many species? The early history of animals life 7 Requirements of Animal Life What is an adaptation? Adapting to different habitats A
An Introduction to Animal Diversity What defines an animal? Why so many species? The early history of animals life 7 Requirements of Animal Life What is an adaptation? Adapting to different habitats A
10/15/09. Tetrapod Limb Development & Pattern Formation. Developing limb region is an example of a morphogenetic field
 Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
Computational Biology Course Descriptions 12-14
 Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
BSC 1010C Biology I. Themes in the Study of Life Chapter 1
 BSC 1010C Biology I Themes in the Study of Life Chapter 1 Objectives Distinguish among the three domains of life. Distinguish between the Levels of Biological Organization. Note the differences in the
BSC 1010C Biology I Themes in the Study of Life Chapter 1 Objectives Distinguish among the three domains of life. Distinguish between the Levels of Biological Organization. Note the differences in the
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
AP Biology Curriculum Framework
 AP Biology Curriculum Framework This chart correlates the College Board s Advanced Placement Biology Curriculum Framework to the corresponding chapters and Key Concept numbers in Campbell BIOLOGY IN FOCUS,
AP Biology Curriculum Framework This chart correlates the College Board s Advanced Placement Biology Curriculum Framework to the corresponding chapters and Key Concept numbers in Campbell BIOLOGY IN FOCUS,
How many lessons is it?
 Science Unit Learning Summary Content Eukaryotes and Prokaryotes Cells are the basic unit of all life forms. A eukaryotic cell contains genetic material enclosed within a nucleus. Plant and animal cells
Science Unit Learning Summary Content Eukaryotes and Prokaryotes Cells are the basic unit of all life forms. A eukaryotic cell contains genetic material enclosed within a nucleus. Plant and animal cells
Evolutionary Games and Computer Simulations
 Evolutionary Games and Computer Simulations Bernardo A. Huberman and Natalie S. Glance Dynamics of Computation Group Xerox Palo Alto Research Center Palo Alto, CA 94304 Abstract The prisoner s dilemma
Evolutionary Games and Computer Simulations Bernardo A. Huberman and Natalie S. Glance Dynamics of Computation Group Xerox Palo Alto Research Center Palo Alto, CA 94304 Abstract The prisoner s dilemma
Unit 2: Cellular Chemistry, Structure, and Physiology Module 5: Cellular Reproduction
 Unit 2: Cellular Chemistry, Structure, and Physiology Module 5: Cellular Reproduction NC Essential Standard: 1.2.2 Analyze how cells grow and reproduce in terms of interphase, mitosis, and cytokinesis
Unit 2: Cellular Chemistry, Structure, and Physiology Module 5: Cellular Reproduction NC Essential Standard: 1.2.2 Analyze how cells grow and reproduce in terms of interphase, mitosis, and cytokinesis
5/4/05 Biol 473 lecture
 5/4/05 Biol 473 lecture animals shown: anomalocaris and hallucigenia 1 The Cambrian Explosion - 550 MYA THE BIG BANG OF ANIMAL EVOLUTION Cambrian explosion was characterized by the sudden and roughly simultaneous
5/4/05 Biol 473 lecture animals shown: anomalocaris and hallucigenia 1 The Cambrian Explosion - 550 MYA THE BIG BANG OF ANIMAL EVOLUTION Cambrian explosion was characterized by the sudden and roughly simultaneous
Biology: Life on Earth
 Biology: Life on Earth Eighth Edition Lecture for Chapter 11 The Continuity of Life: Cellular Reproduction Cellular Reproduction Intracellular activity between one cell division to the next is the cell
Biology: Life on Earth Eighth Edition Lecture for Chapter 11 The Continuity of Life: Cellular Reproduction Cellular Reproduction Intracellular activity between one cell division to the next is the cell
Dynamical Modeling in Biology: a semiotic perspective. Junior Barrera BIOINFO-USP
 Dynamical Modeling in Biology: a semiotic perspective Junior Barrera BIOINFO-USP Layout Introduction Dynamical Systems System Families System Identification Genetic networks design Cell Cycle Modeling
Dynamical Modeling in Biology: a semiotic perspective Junior Barrera BIOINFO-USP Layout Introduction Dynamical Systems System Families System Identification Genetic networks design Cell Cycle Modeling
ZEBRAFISH CROSSWORD PUZZLE (LEVEL 1)
 ZEBRAFISH CROSSWORD PUZZLE (LEVEL 1) 1 2 3 4 5 6 7 8 9 1 The defining feature of vertebrates 4 An individual with two identical copies of the 6 A young organism that is physically very 9 How we test our
ZEBRAFISH CROSSWORD PUZZLE (LEVEL 1) 1 2 3 4 5 6 7 8 9 1 The defining feature of vertebrates 4 An individual with two identical copies of the 6 A young organism that is physically very 9 How we test our
Developmental processes Differential gene expression Introduction to determination The model organisms used to study developmental processes
 Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Langman's Medical Embryology
 Langman's Medical Embryology Developmental Biology Differentiation Morphogenesis) Epigenetic landscape (Waddington) ips Langman's Medical Embryology Morphogen gradient FGF8 in mouse limb bud Gilbert "Developmental
Langman's Medical Embryology Developmental Biology Differentiation Morphogenesis) Epigenetic landscape (Waddington) ips Langman's Medical Embryology Morphogen gradient FGF8 in mouse limb bud Gilbert "Developmental
7.32/7.81J/8.591J. Rm Rm (under construction) Alexander van Oudenaarden Jialing Li. Bernardo Pando. Rm.
 Introducing... 7.32/7.81J/8.591J Systems Biology modeling biological networks Lectures: Recitations: ti TR 1:00-2:30 PM W 4:00-5:00 PM Rm. 6-120 Rm. 26-204 (under construction) Alexander van Oudenaarden
Introducing... 7.32/7.81J/8.591J Systems Biology modeling biological networks Lectures: Recitations: ti TR 1:00-2:30 PM W 4:00-5:00 PM Rm. 6-120 Rm. 26-204 (under construction) Alexander van Oudenaarden
Oscillatory Turing Patterns in a Simple Reaction-Diffusion System
 Journal of the Korean Physical Society, Vol. 50, No. 1, January 2007, pp. 234 238 Oscillatory Turing Patterns in a Simple Reaction-Diffusion System Ruey-Tarng Liu and Sy-Sang Liaw Department of Physics,
Journal of the Korean Physical Society, Vol. 50, No. 1, January 2007, pp. 234 238 Oscillatory Turing Patterns in a Simple Reaction-Diffusion System Ruey-Tarng Liu and Sy-Sang Liaw Department of Physics,
Mitosis and Meiosis. 2. The distribution of chromosomes in one type of cell division is shown in the diagram below.
 Name: Date: 1. Jack bought a small turtle. Three months later, the turtle had grown to twice its original size. Which of the following statements best describes why Jack s turtle got bigger? A. Parts of
Name: Date: 1. Jack bought a small turtle. Three months later, the turtle had grown to twice its original size. Which of the following statements best describes why Jack s turtle got bigger? A. Parts of
7.013 Spring 2005 Problem Set 4
 MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel NAME TA 7.013 Spring 2005 Problem Set 4 FRIDAY April 8th,
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel NAME TA 7.013 Spring 2005 Problem Set 4 FRIDAY April 8th,
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Basic modeling approaches for biological systems. Mahesh Bule
 Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
7 th Grade Science Curriculum
 (1 st 9 Weeks- 1 st 4.5 9 Weeks) Date Hobbs Science By being embedded throughout the, these Processing Skills will be addressed throughout the year. NM & 1 Scientific Thinking and Practice Understand the
(1 st 9 Weeks- 1 st 4.5 9 Weeks) Date Hobbs Science By being embedded throughout the, these Processing Skills will be addressed throughout the year. NM & 1 Scientific Thinking and Practice Understand the
Name 8 Cell Cycle and Meiosis Test Date Study Guide You must know: The structure of the replicated chromosome. The stages of mitosis.
 Name 8 Cell Cycle and Meiosis Test Date Study Guide You must know: The structure of the replicated chromosome. The stages of mitosis. The role of kinases and cyclin in the regulation of the cell cycle.
Name 8 Cell Cycle and Meiosis Test Date Study Guide You must know: The structure of the replicated chromosome. The stages of mitosis. The role of kinases and cyclin in the regulation of the cell cycle.
Growth & Development. Characteristics of Living Things. What is development? Movement. What is a cell?
 Characteristics of Living Things made of cells growth acquire and use energy reproduction movement adaptation respond to stimuli/homeostasis interdependence organization What is development? What are some
Characteristics of Living Things made of cells growth acquire and use energy reproduction movement adaptation respond to stimuli/homeostasis interdependence organization What is development? What are some
The Environment and Change Over Time
 The Environment and Change Over Time Biological Evidence of Evolution What do you think? Read the two statements below and decide whether you agree or disagree with them. Place an A in the Before column
The Environment and Change Over Time Biological Evidence of Evolution What do you think? Read the two statements below and decide whether you agree or disagree with them. Place an A in the Before column
Chetek-Weyerhaeuser Middle School
 Chetek-Weyerhaeuser Middle School Science 7 Units and s Science 7A Unit 1 Nature of Science Scientific Explanations (12 days) s 1. I can make an informed decision using a scientific decision-making model
Chetek-Weyerhaeuser Middle School Science 7 Units and s Science 7A Unit 1 Nature of Science Scientific Explanations (12 days) s 1. I can make an informed decision using a scientific decision-making model
Biology Unit Overview and Pacing Guide
 This document provides teachers with an overview of each unit in the Biology curriculum. The Curriculum Engine provides additional information including knowledge and performance learning targets, key
This document provides teachers with an overview of each unit in the Biology curriculum. The Curriculum Engine provides additional information including knowledge and performance learning targets, key
Diffusion and cellular-level simulation. CS/CME/BioE/Biophys/BMI 279 Nov. 7 and 9, 2017 Ron Dror
 Diffusion and cellular-level simulation CS/CME/BioE/Biophys/BMI 279 Nov. 7 and 9, 2017 Ron Dror 1 Outline How do molecules move around in a cell? Diffusion as a random walk (particle-based perspective)
Diffusion and cellular-level simulation CS/CME/BioE/Biophys/BMI 279 Nov. 7 and 9, 2017 Ron Dror 1 Outline How do molecules move around in a cell? Diffusion as a random walk (particle-based perspective)
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Cells: 3 Star. Which row in the chart below best explains the movement of some molecules between the model cell and the solution in the beaker?
 ells: 3 Star 1. ase your answer(s) to the following question(s) on the diagram below and on your knowledge of biology. The diagram represents a model cell setup. The locations of three different substances
ells: 3 Star 1. ase your answer(s) to the following question(s) on the diagram below and on your knowledge of biology. The diagram represents a model cell setup. The locations of three different substances
Chapter 37 Active Reading Guide Neurons, Synapses, and Signaling
 Name: AP Biology Mr. Croft Section 1 1. What is a neuron? Chapter 37 Active Reading Guide Neurons, Synapses, and Signaling 2. Neurons can be placed into three groups, based on their location and function.
Name: AP Biology Mr. Croft Section 1 1. What is a neuron? Chapter 37 Active Reading Guide Neurons, Synapses, and Signaling 2. Neurons can be placed into three groups, based on their location and function.
Tigard-Tualatin School District Science Grade Level Priority Standards
 Sixth Grade Science Physical Science 6.1 Structure and Function: Living and non-living systems are organized groups of related parts that function together and have characteristics and properties. 6.1P.1
Sixth Grade Science Physical Science 6.1 Structure and Function: Living and non-living systems are organized groups of related parts that function together and have characteristics and properties. 6.1P.1
!!!!!!!! DB3230 Midterm 2 12/13/2013 Name:
 1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
Variation of Traits. genetic variation: the measure of the differences among individuals within a population
 Genetic variability is the measure of the differences among individuals within a population. Because some traits are more suited to certain environments, creating particular niches and fits, we know that
Genetic variability is the measure of the differences among individuals within a population. Because some traits are more suited to certain environments, creating particular niches and fits, we know that
Diffusion. CS/CME/BioE/Biophys/BMI 279 Nov. 15 and 20, 2016 Ron Dror
 Diffusion CS/CME/BioE/Biophys/BMI 279 Nov. 15 and 20, 2016 Ron Dror 1 Outline How do molecules move around in a cell? Diffusion as a random walk (particle-based perspective) Continuum view of diffusion
Diffusion CS/CME/BioE/Biophys/BMI 279 Nov. 15 and 20, 2016 Ron Dror 1 Outline How do molecules move around in a cell? Diffusion as a random walk (particle-based perspective) Continuum view of diffusion
The Role of Retinoic Acid and Notch in the Symmetry Breaking Instabilities for Axon Formation
 The Role of Retinoic Acid and Notch in the Symmetry Breaking Instabilities for Axon Formation MAJID BANI-YAGHOB DAVID E. AMNDSEN School of Mathematics and Statistics, Carleton niversity 1125 Colonel By
The Role of Retinoic Acid and Notch in the Symmetry Breaking Instabilities for Axon Formation MAJID BANI-YAGHOB DAVID E. AMNDSEN School of Mathematics and Statistics, Carleton niversity 1125 Colonel By
Biology 218, practise Exam 2, 2011
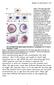 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
FUNDAMENTALS of SYSTEMS BIOLOGY From Synthetic Circuits to Whole-cell Models
 FUNDAMENTALS of SYSTEMS BIOLOGY From Synthetic Circuits to Whole-cell Models Markus W. Covert Stanford University 0 CRC Press Taylor & Francis Group Boca Raton London New York Contents /... Preface, xi
FUNDAMENTALS of SYSTEMS BIOLOGY From Synthetic Circuits to Whole-cell Models Markus W. Covert Stanford University 0 CRC Press Taylor & Francis Group Boca Raton London New York Contents /... Preface, xi
Encyclopedia of Applied and Computational Mathematics
 Björn Engquist Editor Encyclopedia of Applied and Computational Mathematics Volume L Z With 31 Figures and 33 Tables Editor Björn Engquist University of Texas at Austin Austin, TX, USA ISBN 97-3-5-75-
Björn Engquist Editor Encyclopedia of Applied and Computational Mathematics Volume L Z With 31 Figures and 33 Tables Editor Björn Engquist University of Texas at Austin Austin, TX, USA ISBN 97-3-5-75-
EvolutionIntro.notebook. May 13, Do Now LE 1: Copy Now. May 13 12:28 PM. Apr 21 6:33 AM. May 13 7:22 AM. May 13 7:00 AM.
 Different interpretations of cetacean evolutionary history 4/19/10 Aim: What is Evolution by Natural Selection Do Now: How do we know all life on earth is related? Homework Read pp. 375 379 p. 379 # 1,2,3
Different interpretations of cetacean evolutionary history 4/19/10 Aim: What is Evolution by Natural Selection Do Now: How do we know all life on earth is related? Homework Read pp. 375 379 p. 379 # 1,2,3
