A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses
|
|
|
- Gregory Berry
- 6 years ago
- Views:
Transcription
1 (2009) 3, & 2009 International Society for Microbial Ecology All rights reserved /09 $ ORIGINAL ARTICLE A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses Ryan T Jones 1, Michael S Robeson 1, Christian L Lauber 2, Micah Hamady 3, Rob Knight 4 and Noah Fierer 1,2 1 Department of Ecology and Evolutionary Biology, University of Colorado, Boulder, CO, USA; 2 Cooperative Institute for Research in Environmental Sciences, University of Colorado, Boulder, CO, USA; 3 Department of Computer Science, University of Colorado, Boulder, CO, USA and 4 Department of Chemistry and Biochemistry, University of Colorado, Boulder, CO, USA Acidobacteria are ubiquitous and abundant members of soil bacterial communities. However, an ecological understanding of this important phylum has remained elusive because its members have been difficult to culture and few molecular investigations have focused exclusively on this group. We generated an unprecedented number of acidobacterial DNA sequence data using pyrosequencing and clone libraries ( and 1787 sequences, respectively) to characterize the relative abundance, diversity and composition of acidobacterial communities across a range of soil types. To gain insight into the ecological characteristics of acidobacterial taxa, we investigated the largescale biogeographic patterns exhibited by acidobacterial communities, and related soil and site characteristics to acidobacterial community assemblage patterns. The 87 soils analyzed by pyrosequencing contained more than 8600 unique acidobacterial phylotypes (at the 97% sequence similarity level). One phylotype belonging to Acidobacteria subgroup 1, but not closely related to any cultured representatives, was particularly abundant, accounting for 7.4% of bacterial sequences and 17.6% of acidobacterial sequences, on average, across the soils. The abundance of Acidobacteria relative to other bacterial taxa was highly variable across the soils examined, but correlated strongly with soil ph (R ¼ 0.80, Po1). Soil ph was also the best predictor of acidobacterial community composition, regardless of how the communities were characterized, and the relative abundances of the dominant Acidobacteria subgroups were readily predictable. Acidobacterial communities were more phylogenetically clustered as soil ph departed from neutrality, suggesting that ph is an effective habitat filter, restricting community membership to progressively more narrowly defined lineages as ph deviates from neutrality. (2009) 3, ; doi:138/ismej ; published online 8 January 2009 Subject Category: microbial population and community ecology Keywords: community phylogenetics; microbial community; community assembly Introduction The founding member of the Acidobacteria, Acidobacterium capsulatum, was isolated from an acidic mineral environment in 1991 (Kishimoto et al., 1991), and was shown to represent a unique bacterial phylum in 1995 (Hiraishi et al., 1995). Molecular surveys of soil bacteria confirmed the existence of the Acidobacteria in soil and gave the first glimpse of the incredible phylogenetic diversity contained within this phylum (Kuske et al., 1997). Correspondence: RT Jones, Department of Ecology and Evolutionary Biology, University of Colorado-Boulder, CB334, Boulder, CO 80309, USA. Ryan.Jones@colorado.edu Received 18 September 2008; revised 13 November 2008; accepted 18 November 2008; published online 8 January 2009 Since then, further molecular surveys have revealed that Acidobacteria are ubiquitous and among the most abundant bacterial phyla in soil (Janssen, 2006). However, we know surprisingly little about the ecological characteristics of this dominant group of soil bacteria. Cultured isolates can give some indication of the ecological characteristics of a lineage, but Acidobacteria have thus far been difficult to culture. Of high-quality 16S rrna gene sequences available from bacterial isolates (Ribosomal Database Project (RDP) II v10.4; index.jsp), only 74 are classified as Acidobacteria. Culturing techniques that have contributed to the successful isolation of Acidobacteria include: media with lowered ph (Sait et al., 2006); increased headspace CO 2 concentrations (Stevenson et al.,
2 2004); substrate amendments (Pankratov et al., 2008); random selection from a wide variety of growth substrates (Joseph et al., 2003); use of a diffusion chamber (Bollmann et al., 2007) and extension of the incubation time (Davis et al., 2005). Despite the successes of these culturing methods, the isolates poorly represent the known diversity of Acidobacteria; of the 74 acidobacterial isolates, nearly 75% (55/74) are in subgroup 1 (RDP v10.4). Nevertheless, the cultured isolates suggest that some Acidobacteria have slow growth rates and either require or tolerate specific environmental conditions. The paucity of cultured isolates poses a significant limitation in understanding soil acidobacterial ecology. As an alternative approach, molecular techniques can be used to describe the biogeographical patterns exhibited by soil acidobacterial communities and such patterns can provide important insight into the ecology, physiology and life history characteristics of the group. Acidobacterialspecific 16S rrna primers developed by Barns et al. (1999) have been used to quantify acidobacterial abundances in specific samples (Felske et al., 2000; Fierer et al., 2005; Fierer et al., 2007a), to target acidobacterial genomic DNA (Quaiser et al., 2003), to develop group-specific denaturing gradient gel electorphoresis (Boon et al., 2002) and to generate clone libraries that specifically target acidobacterial diversity (Barns et al., 2007; Bryant et al., 2008). A recent comprehensive study used quantitative PCR to determine the relative abundances of six bacterial groups in 71 soil samples, and found that the relative abundance of Acidobacteria was negatively correlated with soil carbon availability (Fierer et al., 2007a). Also, two recent studies of microbial succession in soil found the abundance of Acidobacteria to be low in young soils and high in older soils (Nemergut et al., 2007; Tarlera et al., 2008). Additional molecular work has found ph to influence acidobacterial abundances, with the highest abundances found in environments with the lowest ph (Fierer et al., 2007a; Mannisto et al., 2007; Lauber et al., 2008). Taken together, these findings support the idea that Acidobacteria are often slowgrowing oligotrophs and that their abundance within a community may be regulated by ph. We examined acidobacterial communities in 87 diverse soils to gain some insight into the ecological characteristics of the Acidobacteria. More specifically, we wanted to determine which abiotic factors influence the abundance, diversity and composition of soil acidobacterial communities across a range of ecosystem types. We used two complementary methods, pyrosequencing and clone libraries, to generate sequence data and survey acidobacterial communities at an unprecedented breadth (surveying a large number of soils) and depth (detailed surveys of the acidobacterial communities in each of the individual soils). Methods Soil collection, characterization and DNA extraction A total of 87 unique soils representing a diverse array of soil and site characteristics were collected from throughout North and South America. Each soil was collected near the peak of the plant-growing season at each location. We collected the upper 5 cm of mineral soil from each site, collecting soil from 5 to 10 locations within an area of B100 m 2 with the soils from each site combined into a single bulk sample. All samples were sieved to 4 mm and thoroughly homogenized before being stored at 80 1C. Additional details on the soil collection procedures and the measurements of the soil characteristics are provided elsewhere (Fierer and Jackson, 2006; Fierer et al., 2007a). Supplementary Table 1 contains information on the sampling location and soil characteristics for each of the 87 soils included in this study. DNA was isolated from each of the collected soil samples using the MoBio Power Soil DNA extraction kit (MoBio Laboratories, Carlsbad, CA, USA) with the following modifications. Soil samples of approximately 5 g were ground with a mortar and pestle in liquid nitrogen and three 0.5-g subsamples of this ground soil were placed into bead tubes for extraction. The bead tubes were heated to 65 1C for 10 min, then shaken horizontally for 2 min at maximum speed with the MoBio vortex adapter. The remaining steps were performed as directed by the manufacturer. DNA from the three replicate subsamples per site was combined, and samples were stored at 20 1C. Pyrosequencing of bacterial 16S rrna genes Pyrosequencing of the DNA isolated from all 87 samples was conducted using a previously described method (Hamady et al., 2008). Briefly, we amplified the 16S rrna genes from each sample using a primer set identical to that described in Hamady et al., but the reverse primer contained a slightly longer 12-bp error-correcting Golay barcode rather than the originally described 8-bp Hamming barcode. This primer set is universal for bacterial 16S rrna and has been found to be well suited for the phylogenetic analysis of pyrosequencing reads (Liu et al., 2007, 2008). PCR reactions consisted of 0.25 ml (30 mm) of each forward and reverse primers, 3 ml template DNA and 22.5 ml Platinum PCR Super- Mix (Invitrogen, Carlsbad, CA, USA). Samples were initially denatured at 94 1C for 3 min, then amplified using 35 cycles of 94 1C for 45 s, 50 1C for 30 s and 72 1C for 90 s. A final extension of 10 min at 72 1C was added at the end of the program to ensure complete amplification of the target region. All samples were amplified in triplicate and no template controls were included in all steps of the process. 443
3 444 A composite sample for pyrosequencing was prepared by pooling approximately equal amounts of PCR amplicons from each sample. The replicate PCR reactions for each sample were combined and cleaned with the Mobio UltraClean-htp PCR Cleanup kit (Mobio Laboratories) as directed by the manufacturer. Each sample was then quantified using PicoGreen dsdna reagent (Invitrogen) and the appropriate volume of the cleaned PCR amplicons was combined in a sterile, 50 ml polypropylene tube and precipitated on ice with sterile, 5 M NaCl (0.2 M final concentration) and two volumes of ice cold, 100% ethanol for 45 min. The precipitated DNA was centrifuged at 7800 g for 40 min at 4 1C and the resulting pellet was washed with an equal volume of 70% ethanol and centrifuged again at 7800 g for 20 min at 4 1C. The supernatant was removed and the pellet was air dried for 7 min at room temperature, then resuspended in 100 ml DNA nuclease-free water. The sample was sent to the Environmental Genomics Core Facility at the University of South Carolina, Columbia, SC, USA, for pyrosequencing on a 454 Life Sciences Genome Sequencer FLX (Roche Diagnostics, Indianapolis, IN, USA) machine. The average sequence length was 232 bp, and sequences obtained through pyrosequencing have been deposited in the GenBank short read archive. Sequences were processed and analyzed following previously described procedures (Hamady et al., 2008). Phylotypes were defined at the 97% sequence identity level, and a set of representative sequences selected from these phylotypes were aligned using NAST (DeSantis et al., 2006) with a PH lanemask to screen out hypervariable regions of the sequence. Sequence taxonomy assignments were made using RDP 2 classifier (release 10.4), with a minimum support threshold of 60%. Of classified sequences, were classified as Acidobacteria. Sequences were further divided into 26 subgroups: subgroups 1 8 (Hugenholtz et al., 1998); subgroups 9 11 (Zimmermann et al., 2005) and subgroups (Barns et al., 2007). Cloning and sequencing of acidobacterial 16S rrna genes We constructed clone libraries to survey acidobacterial communities in 22 of the 87 soils to provide more detailed phylogenetic information (that is, longer sequence reads) than that offered by pyrosequencing. The clone libraries were constructed by amplifying the DNA samples with the acidobacterial-specific forward primer 31f (5 0 -GATCCTGGCT CAGAATC -3 0 ) developed by Barns et al. (1999) and the universal reverse primer 1492r (5 0 -GGTTAC CTTGTTACGACTT-3 0 ). Each 50-ml PCR reaction contained: 1 PCR buffer; 2.5 mm MgCl 2 ; 0.2 mm of each dntp; 0.2 mm of each primer; 0.5 U of Taq DNA polymerase (Invitrogen) and 2 ml of template DNA. Each DNA sample was amplified with three replicate PCR amplifications (30 cycles each, annealing temperature of 52 1C) with the amplicons from each sample pooled prior to cloning. Amplicons were cleaned using the QIAquick PCR purification kit (Qiagen, Valencia, CA, USA) and cloned using the TOPO TA for Sequencing kit (Invitrogen) following the manufacturer s instructions. Plates were grown overnight at 37 1C before plasmid amplification and sequencing. For each sample, a total of 96 clones were sequenced with single-pass reads at Agencourt Bioscience (Beverly, MA, USA). DNA sequences from the clone libraries were aligned using the NAST aligner (DeSantis et al., 2006). The aligned sequences were imported into ARB v (Ludwig et al., 2004) and edited by hand. Sequences were exported using ARB s Acidobacteria Lane mask to exclude ambiguous base pair positions. A 16S rrna gene sequence of Escherichia coli (accession no. AE016755) was included as an outgroup. After filtering out short sequences (those less than 590 bp), a total of 1787 Acidobacteria sequences remained for analysis (GenBank accession nos: FJ FJ167342). Both ends of each DNA sequence were trimmed to minimize the number of ambiguous characters, and an additional 17 bp that could not be accurately aligned were removed. The alignment file is available upon request. Clone library DNA sequences were classified into subgroups using RDP 2 classifier (release 10.4). Acidobacteria have been divided into 26 subgroups: subgroups 1 8 (Hugenholtz et al., 1998); subgroups 9 11 (Zimmermann et al., 2005) and subgroups (Barns et al., 2007). We compared the relative abundances of Acidobacteria subgroups to environmental characteristics for all individual soils. Sequences were also used in community analyses (detailed below). Community analyses pyrosequencing data We used the pyrosequencing data to estimate the relative abundance of Acidobacteria in each bacterial community by comparing the number of sequences classified as belonging to the Acidobacteria versus the number of classified bacterial sequences per sample. We used Spearman s rank correlation to compare this estimate of acidobacterial relative abundances to the following soil and site characteristics: ph, percent soil organic carbon, mean annual precipitation at location, mean annual temperature at location, soil C/N ratio, soil carbon mineralization rate, and percent silt and clay. We also used Spearman s rank correlations to compare the relative abundances of Acidobacteria subgroups across all of the individual soils, relating the relative abundance estimates for the individual subgroups to the soil and site characteristics. Spearman s rank correlations were performed using an online statistics calculator (Wessa, 2008). To estimate the pairwise similarity in acidobacterial communities, we generated Bray Curtis dissimilarity matrices, as implemented in PRIMER-E v6 (PRIMER-E Ltd, Lutton,
4 UK), using the relative abundance of acidobacterial phylotypes (identified as described above) as an input. Three other Bray Curtis dissimilarity matrices were constructed that excluded singletons (those phylotypes only observed once), singletons and doubletons, and phylotypes not observed at least 10 times. We used the Mantel test in PRIMER-E to compare dissimilarity matrices to pairwise distances in environmental characteristics as estimated using normalized Euclidean distances in the measured soil and site parameters, an approach described by Fierer and Jackson (2006). Community analyses clone library data We compared levels of diversity in individual soils by determining the number of unique phylotypes at four different levels of sequence similarity: 90%, 93%, 95% and 97% using FastGroup II (Yu et al., 2006) to identify unique phylotypes and EstimateS to calculate rarefaction curves (Colwell, 2005). To estimate the pairwise similarity in acidobacterial communities, we generated Bray Curtis dissimilarity matrices, as implemented in PRIMER-E, using the relative abundance of acidobacterial phylotypes as an input. In total, 100 bootstrapped maximum likelihood trees were constructed using RAxML v7.0.4 with the GTRGAMMA model without branch length constraints using the clone library sequence data (Stamatakis, 2006). Duplicate sequences were omitted, and a total of 1652 Acidobacteria 16S rrna gene sequences were analyzed. We used the resultant trees in phylogenetic techniques that, unlike taxonomy-based or phylotype-based methods, incorporate evolutionary relatedness of community members into diversity descriptions and comparisons. Including evolutionary history into community comparisons is valuable because more closely related individuals are more likely to be ecologically similar to one another than distantly related individuals, and thus cannot be considered statistically independent (Felsenstein, 1985; Harvey and Pagel, 1991). We analyzed each bootstrapped tree (n ¼ 100) independently in all phylogenetic analyses, because the true phylogeny is unknown and unknowable and it is important to incorporate phylogenetic uncertainty in phylogenetic analyses (Huelsenbeck et al., 2000). Furthermore, when phylogenetic uncertainty is incorporated into microbial community comparisons, inferences are more conservative and robust (Jones and Martin, 2006). We used three phylogenetic metrics to further explore Acidobacteria diversity and community assembly patterns. The phylogenetic metric, phylodiversity, was used to compare diversity across the acidobacterial communities. Phylodiversity was originally developed to prioritize conservation areas (Faith, 1992), but we use it here to describe the percentage of acidobacterial phylogenetic diversity contained in one soil sample relative to all acidobacterial diversity (that is, the sum of branch lengths in one community relative to the total branch lengths of all communities). Because sampling intensity is not equal among all soil samples and phylodiversity estimates will increase with comparatively higher sampling, we used a bootstrap approach to randomly assemble 100 communities with 65 taxa from each soil sample; each community bootstrap was evaluated with respect to each bootstrapped tree (that is, 100 communities 100 phylogenetic trees ¼ data points). Phylodiversity values were determined using Phylocom v4.0.1 (Webb et al., 2008). We used the net relatedness index (NRI) to describe the phylogenetic evenness of Acidobacteria communities within individual soils, and compared phylogenetic evenness to environmental characteristics. The NRI (Webb, 2000, 2002) compares the pairwise distances (as determined by the total length of branches between two taxa) between all members of a community to pairwise distances of randomly selected taxa from the entire tree, and indicates whether a community s diversity is phylogenetically clumped (members of a community are more related to each other than by chance) or phylogenetically even (members of a community are less related to each other than by chance). Positive NRI values indicate phylogenetically clustered communities and negative values indicate phylogenetically even communities. Because NRI values are affected by sample size, we use the same community bootstrap approach detailed for phylodiversity measurements. NRI values were obtained using the comstruct command in Phylocom v4.0.1 with 1000 randomizations and null model 1 (Webb et al., 2008). We used UniFrac to cluster soils based on the phylogenetic differentiation they contained (Lozupone and Knight, 2005; Lozupone et al., 2006). UniFrac clusters microbial communities based on the fraction of unique branch lengths represented by a community in pairwise comparisons to all other communities. Communities separated by a small fraction of unique evolutionary diversity cluster close together and communities separated by large fractions of evolutionary diversity cluster farther apart. The underlying notion behind this approach is that longer unique branch lengths represent more evolutionary adaptation, which is necessary for survival in one community relative to the other (Lozupone and Knight, 2008). We used Mantel tests to test the significance of correlations between the pairwise UniFrac distances between soil acidobacterial communities and the normalized Euclidean distances in soil and site characteristics. Results We characterized the relative abundance, diversity and composition of soil acidobacterial communities 445
5 446 across a wide range of soil types using two sequencing approaches. Using a pyrosequencing approach with universal bacterial 16S rrna gene primers, we generated acidobacterial sequences from 87 soils. The clone library approach with Acidobacteria-specific 16S rrna gene primers was used to generate 1787 sequences from 22 soils. The pyrosequencing yielded a more extensive survey of acidobacterial communities (more soils, more sequences per soil), but the clone library data allowed us to survey the communities at a higher level of phylogenetic resolution (owing to the longer lengths of the sequence reads). We used the two types of data to characterize Acidobacteria communities in three ways: (1) taxonomy-based, (2) phylotype-based and (3) phylogenetic. We compared community characterizations to soil environmental data (Supplementary Table 1) to gain a better understanding of the factors shaping Acidobacteria communities. Overall abundance of Acidobacteria We used the pyrosequencing data to estimate the phylum-level relative abundance of Acidobacteria in the 87 soils examined. Across all soils, Acidobacteria represented 30.9% of all classified bacterial sequences detected using pyrosequencing ( sequences out of a total of sequences). However, the relative abundance of acidobacterial sequences within an individual soil bacterial community varied from 2.4% to 78.5% (Figure 1). The relative abundance of soil Acidobacteria was strongly correlated with soil ph (R ¼ 0.80, Po1), with Acidobacteria representing a larger portion of the bacterial community in low ph soils (Figure 1). Other measured soil and site characteristics that were also significantly correlated included: mean annual precipitation (R ¼ 0.66, Po1), % soil organic carbon (R ¼ 0.48, Po1) and soil C/N ratio (R ¼ 0.45, Po1). Relative abundance of Acidobacteria in soil R = ph Figure 1 Relationship between ph and the relative abundance of Acidobacteria in 87 soil bacterial communities using pyrosequencing data. Abundance of Acidobacteria is strongly and negatively correlated with ph (Spearman s rank correlation: R ¼ 0.80, Po1). However, it is important to note that these other environmental characteristics were more weakly correlated with acidobacterial abundances than soil ph. Also, these other environmental characteristics are, to varying degrees, also correlated with soil ph (Fierer and Jackson, 2006). Relative abundance of Acidobacteria subgroups We further classified Acidobacteria DNA sequences (from clone libraries and pyrosequencing) into acidobacterial subgroups using the RDP v10.4 classifier. We present the average relative abundances of Acidobacteria subgroups 1 26 for both the pyrosequencing and clone library data in Table 1; we also present the relative abundances of Acidobacteria subgroups 1 26 in individual soils for pyrosequencing data (Supplementary Table 2) and for clone library data (Supplementary Table 3). On the basis of the pyrosequencing results, we find that acidobacterial subgroups 1, 2, 3, 4 and 6 are the most abundant in soil: across all 87 soils, these subgroups accounted for 22.4%, 12.7%, 15.0%, 29.0% and 10.7% of the acidobacterial sequences, respectively. Subgroups 1, 3, 4 and 6 were also found to be Table 1 Abundance of Acidobacteria subgroups detected using pyrosequencing and clone libraries Subgroup Pyrosequencing Clone libraries Average (%) Range (%) Average (%) Range (%) ND a ND ND 9 ND ND ND o ND ND ND ND o ND 20 o ND 21 ND ND ND 23 o ND 24 ND ND ND 26 ND ND Unclassified a ND indicates that the subgroup was not detected. DNA sequences were classified into 26 acidobacterial subgroups using the Ribosomal Database Project v10.4 classifier. The 26 subgroups are classified according to the following designations: subgroups 1 8 according to Hugenholtz et al. (1998); subgroups 9 11 according to Zimmermann et al. (2005) and subgroups according to Barns et al. (2007).
6 abundant in the clone libraries, but the Acidobacteria-specific primers used to generate the clone libraries did not amplify sequences from a number of Acidobacteria subgroups (Table 2). Although some of these subgroups appear to be minor and ephemeral constituents of soil Acidobacteria communities, subgroups 2, 7 and 16 were often abundant and represented up to 59.9%, 10.9% and 15.9%, respectively, of Acidobacteria in individual soils as quantified by pyrosequencing (Figure 2). The relative abundance of most Acidobacteria subgroups was strongly correlated with the ph of the individual soil; groups 1, 2, 3, 12, 13 and 15 had negative correlations and groups 4, 6, 7, 10, 11, 16, 17, 18, 22 and 25 had positive correlations with ph (Figure 2; Supplementary Tables 4 and 5). Spearman s correlations between subgroup abundance and all environmental characteristics are also presented (Supplementary Tables 4 and 5). Diversity of soil Acidobacteria The Acidobacteria subgroups represent broad taxonomic groups and each group contains a tremendous amount of diversity. To examine the levels of diversity at finer levels of taxonomic resolution, we binned DNA sequences into phylotypes based on sequence similarity (90%, 93%, 95% and 97% for clone library data; 97% for pyrosequencing data). We constructed rarefaction curves for the clone library data (Supplementary Figure 1) and the absence of clear asymptotes suggests that we did not completely sample Acidobacteria diversity at the 93%, 95% or 97% phylotype designations. Of 1787 DNA sequences generated by the clone libraries, we found the following number of phylotypes across the 22 soils examined: 38 (90%), 102 (93%), 230 (95%) and 317 (97%). Of the acidobacterial DNA sequences generated by pyrosequencing from the 87 soils, we identified 8643 unique phylotypes (97%). It is worth noting that the 97% phylotype designation is not necessarily equivalent between the Sanger sequences and pyrosequences because the region of 16S targeted by pyrosequencing is more variable than the region targeted by Sanger sequencing. To explore the effects of the environment on acidobacterial diversity, we related phylodiversity to soil characteristics. Median phylodiversity values for individual soil communities ranged from 10.4% to 16.8%. However, phylodiversity did not correlate with any of the measured environmental characteristics (P40.1 in all cases), including ph (Supplementary Figure 2). This suggests that, across the wide range of soil types examined, the characteristics of these soils do not restrict or amplify the total amount of diversity contained within a given acidobacterial community. We further explored environmental effects on phylogenetic diversity by comparing environmental characteristics to phylogenetic evenness. Soil ph strongly affected phylogenetic evenness; communities were the most even in soils with close to neutral ph and communities became phylogenetically clustered as ph departed from neutrality (Figure 3). In other words, the more the soil ph departed from neutrality, the more the communities were composed of closely related members. This result may seem at odds with the lack of correlation between phylodiversity and ph (Supplementary Figure 2). However, phylodiversity is a snapshot of diversity and is highly sensitive to the presence or absence of major lineages, whereas the NRI is based on pairwise comparisons between all taxa in a community and is more indicative of broad assemblage patterns. Overall differentiation in Acidobacteria community composition We used the relative abundances of each phylotype in individual samples to characterize differences in 447 Table 2 Mantel correlations between community structure and environmental variables Phylotype definitions No. of phylotypes Pyrosequencing Clone library 97% 97% 95% 93% 90% UniFrac ph (log) (o1) (o1) (o1) (o1) (o1) (o1) MAP a (mm) (o1) (0.336) (0.195) (0.261) (0.274) (0.012) %C b (sqrt) (o1) 3 (0.139) (0.239) (0.149) (0.137) (5) C/N c (log) (o1) (0.021) (7) (8) (0.011) (0.045) C min d (log) (o1) (0.417) (0.263) (0.186) (0.354) (0.203) MAT e (1C) (o1) (0.675) 7 (0.419) (0.483) (0.745) (0.254) % silt and clay (sqrt) (4) (0.066) (0.027) (0.026) (0.027) (0.138) Bold values indicate a significance of o0.05. Significance values in parentheses. a Mean annual precipitation. b % organic carbon. c Carbon to nitrogen ratio. d Carbon mineralization rate. e Mean annual temperature.
7 Subgroup 1 Pyro: R = -0.89** Clone: R = -0.76** Subgroup 5 Pyro: NS Clone: ND Subgroup 2 Pyro: R = -0.89** Clone: ND Subgroup 6 Pyro: R = 0.76** Clone: R = 0.67* Subgroup 3 Pyro: R = -0.40** Clone: R = -0.63* 0.12 Subgroup 7 Pyro: R = 0.52** Clone: ND Subgroup 4 Pyro: R = 0.91** Clone: R = 0.91** Subgroup 16 Pyro: R = 0.74** Clone: R = ND ph Figure 2 Effect of ph on abundance of Acidobacteria subgroups (1 7 and 16) relative to all Acidobacteria using pyrosequencing and clone library data. Triangles (black) represent pyrosequencing data; circles (grey) represent clone library data. Spearman s rank correlations between subgroup abundance and ph are depicted. NS, not significant; ND, not detected. *Po0.05; **Po1. ph acidobacterial community composition as determined by pyrosequencing. Mantel tests between Bray Curtis-transformed community assemblage matrices and pairwise environmental matrices revealed significant correlations between community assembly patterns and ph and C/N ratios at all
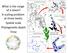
8 Net Relatedness Index Departure from neutral ph Acidic Basic Figure 3 The net relatedness index (NRI) measures phylogenetic evenness. Positive NRI values indicate that communities are phylogenetically clustered and negative values indicate phylogenetic evenness. Here, phylogenetic clustering is positively correlated with departure from neutral ph (R 2 ¼ 0.229, Po0.05); the more the soil ph departs from neutrality, the more the communities are phylogenetically clustered. Median NRI values are plotted. Stress: 0.13 ph < Figure 4 Non-metric multi-dimensional scaling plot of soil community assembly patterns (using Bray Curtis-transformed pyrosequencing data) as related to soil ph. The first principal axis is dominated by soil ph effects, showing the key effect of ph on acidobacterial community assembly. phylotype designations for both pyrosequencing and clone library data (Table 2). The correlation with ph was particularly strong and is clearly depicted in a non-metric multi-dimensional scaling plot of the pyrosequencing phylotype data (Figure 4). We see a similar pattern if we use phylogenetic approaches to compare Acidobacteria communities. We clustered Acidobacteria communities based on their unique phylogenetic diversity using the Uni- Frac algorithm with the clone library data. Mantel tests comparing UniFrac distances and environmental distances showed significant correlations between UniFrac distances and the following environmental parameters: ph, soil moisture, mean annual precipitation, % organic C and C/N ratio (Table 2). UniFrac distances showed more significant associations than phylotype-based approaches, > 8 CF2 HF1 LQ3 LQ1 HI4 BZ1 HJ2 DF1 HJ1 IT2 SK1 CL1 AR1 SK3 CO3 KP1 GB3 MD2 SR2 SA2 <10% # % #34 * 3.5% #33 * 2.0% #55 * 1.9% 10-20% underscoring the importance of using a phylogenetic approach. To further explore these trends, we quantified the abundances of the 20 most common phylotypes identified using the clone library approach in the 22 soils. These 20 phylotypes represented approximately 75% of all acidobacterial DNA sequences in the clone libraries. Again, we found a clear trend of phylotype presence correlated with ph (Figure 5). Importantly, only 4 of the top 20 most common phylotypes shared greater than 97% sequence identity with a cloned isolate (Figure 5). Phylotype no. 32 was particularly abundant: it accounts for 7.4% of all bacterial sequences in soil (17.6% of all Acidobacteria) and represents as much as 23.5% of bacterial sequences (41.3% of acidobacterial sequences) in individual soils. These estimated values were corrected for the clone libraries not detecting some subgroups (for example (abundance of phylotype no. 32 in clone libraries) (proportion of subgroups 1, 3, 4, 5, 6, 11, 13, 15, 17 and 18 relative to all detected Acidobacteria using pyrosequencing data)). Discussion #39 0.3% #1 * 5.4% #9 0.9% #44 1.7% #19 0.3% #2 6.4% 20-30% 30-40% 40-50% Our results show that soil ph strongly regulates the abundance of Acidobacteria (relative to all bacteria) in individual communities (Figure 1). Negative correlations between Acidobacteria phylum-level abundance and ph have also been seen in fine #15 0.7% #16 2.3% #6 2.6% #84 1.1% #4 0.7% #20 3.8% # % #96 2.7% # % # % ph Figure 5 Relative abundances of the 20 most abundant phylotypes (97% OTU designation) in individual soil samples using clone library data. Subgroups are indicated over phylotypes. Soils are arranged from lowest ph (CF2) to highest ph (SA2). Phylotypes with asterisks share 497.0% sequence similarity with a cultured isolate. Percentages at bottom indicate the abundance of each phylotype relative to all detected acidobacterial sequences (values corrected for acidobacterial-specific primers not detecting certain subgroups; see text).
9 450 organic matter of streams and in Arctic soils (Fierer et al., 2007b; Mannisto et al., 2007). Our results differ, however, from a previous study that found that organic carbon availability was the best predictor of Acidobacteria abundance in soils (Fierer et al., 2007a). This disparity may be related to our finding (evident in Figure 2 and Table 1) that the primer set used by Fierer et al. (2007a), the same primer set used in the clone library analyses described here, excludes some common acidobacterial subgroups. We suggest that our pyrosequencing results (Figure 1) better represent Acidobacteria relative abundances in soil and that the strong correlation between Acidobacteria abundance and ph (R ¼ 0.80) is more robust than the weaker correlation between Acidobacteria abundance and carbon availability (R 2 ¼ 0.26) noted by Fierer et al. (2007a). Our results do not exclude the notion that Acidobacteria are often slow-growing oligotrophs, as suggested by a negative correlation between abundance and carbon availability (Fierer et al., 2007a), culturing studies (Davis et al., 2005) or studies of microbial succession (Nemergut et al., 2007; Tarlera et al., 2008). Nevertheless, based on our comprehensive survey of acidobacterial communities across a wide range of soil types, ph governs phylum-level abundance of Acidobacteria more than any other measured environmental characteristic. This is not unique to Acidobacteria; environmental ph has also been found to significantly correlate with the abundances of Alphaproteobacteria, Bacteroidetes, Flavobacteria and OP10 phyla in lakes (Percent et al., 2008). Within individual Acidobacteria soil communities, composition is also strongly regulated by ph. This finding holds across all characterizations of Acidobacteria communities: taxonomy-based, phylotype-based and phylogenetic. The abundances of most Acidobacteria subgroups significantly correlate with ph: abundances of subgroups 1, 2, 3, 12 and 13 have a negative relationship; abundances of subgroups 4, 6, 7, 10, 11, 16, 17, 18, 22 and 25 have a positive relationship (Figure 2; Supplementary Tables 4 and 5). These results confirm the results of Eichorst et al. (2007) who found that the abundance of subgroup 1 was inversely related to ph in soil and other terrestrial environments and Barns et al. (1999) who found that some subgroups did not occur in low ph soils. Similarly, a higher proportion of Acidobacteria subgroup 1 colonies formed on acidic media versus neutral media (Sait et al., 2006). If we compare community assemblages using a range phylotype definitions (90%, 93%, 95% and 97% sequence similarity for clone library data; 97% sequence similarity for pyrosequencing data), we also find that ph strongly correlates with acidobacterial community composition, regardless of the data set or the phylotype definition (Table 2; Figure 4). As a final means to characterize Acidobacteria communities, we used UniFrac to cluster soils based on the unique fraction of phylodiversity contained in each soil based on pairwise comparisons between all samples (Lozupone et al., 2006; Lozupone and Knight, 2005). Again, we find that soil ph is a strong predictor of acidobacterial community composition (Table 2). These results are similar to other studies that have found ph to regulate fine-scale community composition, such as in the beta-subgroup of proteobacterial ammonia oxidizers (Stephen et al., 1998) and in the aci lineage of freshwater lake Actinobacteria (Newton et al., 2007). Broad-scale surveys comparing entire bacterial communities have also found that bacterial community composition is often well correlated with soil ph (Fierer and Jackson, 2006; Hogberg et al., 2007; Lauber et al., 2008; Singh et al., 2008). As soil ph can be considered a master variable that integrates a number of other soil and site characteristics, we cannot determine whether ph is having a direct influence on acidobacterial communities or whether soil ph is correlated with some other, unmeasured, environmental parameter that, in turn, influences acidobacterial community composition. Regardless, our results confirm results from a range of other studies showing a strong influence of ph on microbial communities and suggest that ph is also a strong predictor of acidobacterial abundance and community composition in soil. Not surprisingly, we found that the Acidobacteria phylum is phylogenetically diverse with even our relatively large libraries failing to capture the full extent of species -level diversity (Supplementary Figure 1). This diversity, however, is not partitioned randomly across individual communities. We used the NRI metric to quantify the phylogenetic evenness of individual Acidobacteria communities and found that communities became more phylogenetically clustered as soil ph departed from neutrality (Figure 3). This suggests that ph acts as an effective habitat filter, restricting community membership to progressively more narrowly defined lineages as ph deviates from neutrality. Similar patterns were noted by Felske et al. (1998), who implicated low ph as a habitat filter for soil bacteria when many duplicate sequences were found in a peaty acidic soil. Intracellular ph of bacteria is close to neutral regardless of environmental ph, and bacterial cells persisting in acidic environments generally cope by having increased buffering capacities or less permeable cell membranes (Booth, 1985). Closely related Acidobacteria likely share similar, phylogenetically constrained, cellular mechanisms to compensate for low ph. For example, closely related Acidobacteria may share similar means of membrane lipid construction and these differences in acidobacterial membrane construction may explain the abundance of a novel group of branched glycerol dialkyl glycerol tetraether membrane lipids found to correlate negatively with soil ph (Weijers et al., 2007). Although Acidobacteria are phylogenetically diverse, the 20 most common phylotypes (as defined
10 by 97% sequence similarity) accounted for 458% of community membership (Figure 5). These abundant Acidobacteria phylotypes should account for a disproportionate amount of biogeochemical cycling compared with the more rare phylotypes. However, the cultured Acidobacteria isolates are not representative of total Acidobacteria diversity or, more importantly, the most common Acidobacteria found in soil (Figure 5). Our results should help identify those specific environmental conditions best suited for targeted cultivation of important and abundant phylotypes of Acidobacteria. Of particular interest is one specific acidobacterial phylotype (phylotype no. 32, as defined by 97% sequence similarity), which was found to be particularly abundant in the clone libraries constructed from many soils (Figure 5). Phylotype no. 32 accounts for 7.4% of all bacterial sequences in soil (17.6% of all Acidobacteria) and represents as much as 23.5% of bacterial sequences (41.3% of acidobacterial sequences) in individual soils. Its abundance suggests that it plays an important role in soil biogeochemistry, and the abundance of phylotype no. 32 in soil may parallel that of the bacterioplankton phylotype SAR11 in oceans. SAR11 was first detected using molecular methods (Giovannoni et al., 1990), and was found to be one of the most abundant bacteria in the oceans (Morris et al., 2002). Over a decade passed after the molecular discovery of SAR11 before a member of the cluster was cultured (Rappe et al., 2002). Culturing, particularly of abundant lineages, is important because it allows genomic surveys that provide more insight into the metabolic potential and biogeochemical function of a lineage (Giovannoni and Stingl, 2007). We suggest that Acidobacteria phylotype no. 32 be targeted for culturing to enable genomic analyses of this numerically dominant soil community member. It is particularly noteworthy that our findings are robust across different data generation methods (clone libraries and pyrosequencing) and across a range of data analyses (taxonomy-based, phylotypebased and phylogenetic-based). There are important differences between results obtained using the two data generation methods. Regrettably, the Acidobacteria-specific primers (Barns et al., 1999) failed to detect a number of Acidobacteria subgroups. Some of these subgroups are rather rare members of soil communities, but subgroup 2, 7 and 16 are often common members of Acidobacteria communities (Figure 2). Nevertheless, analyses of pyrosequencing and clone library data yielded similar results, demonstrating that the composition of soil acidobacterial communities are remarkably predictable across a wide range of soil and ecosystem types if we consider a single soil edaphic characteristic, namely soil ph. This is surprising considering the phylogenetic diversity contained within the soil Acidobacteria and it suggests that there is a lot of value in using these broad-scale biogeographic surveys for revealing important insight into the natural history of bacterial lineages represented by few, if any, cultured isolates. Acknowledgements We thank all the individuals who graciously donated their time and energy to help us collect soil samples from across North and South America. We thank Heather Hamilton for assistance with the molecular analyses. We thank Andrew Martin, Elizabeth Costello and members of the Fierer lab for valuable comments on previous versions of this paper. This study was supported by funding provided to NF from the National Science Foundation (MCB and DBI ) and the Andrew W Mellon Foundation. The NSF Microbial Observatories Program (MCB ) supported MSR. MH was supported by an NSF EAPSI fellowship (OISE ) and a NIH Molecular Biophysics Training Grant (T32GM065103). References Barns SM, Cain EC, Sommerville L, Kuske CR. (2007). Acidobacteria phylum sequences in uranium-contaminated subsurface sediments greatly expand the known diversity within the phylum. Appl Environ Microbiol 73: Barns SM, Takala SL, Kuske CR. (1999). Wide distribution and diversity of members of the bacterial kingdom Acidobacterium in the environment. Appl Environ Microbiol 65: Bollmann A, Lewis K, Epstein SS. (2007). Incubation of environmental samples in a diffusion chamber increases the diversity of recovered isolates. Appl Environ Microbiol 73: Boon N, De Windt W, Verstraete W, Top EM. (2002). Evaluation of nested PCR-DGGE (denaturing gradient gel electrophoresis) with group-specific 16S rrna primers for the analysis of bacterial communities from different wastewater treatment plants. FEMS Microbiol Ecol 39: Booth IR. (1985). Regulation of cytoplasmic ph in bacteria. Microbiol Rev 49: Bryant JA, Lamanna C, Morlon H, Kerkhoff AJ, Enquist BJ, Green JL. (2008). Microbes on mountainsides: contrasting elevational patterns of bacterial and plant diversity. Proc Natl Acad Sci USA 105: Colwell R. (2005). EstimateS: statistical estimation of species richness and shared species from samples, Davis KER, Joseph SJ, Janssen PH. (2005). Effects of growth medium, inoculum size, and incubation time on culturability and isolation of soil bacteria. Appl Environ Microbiol 71: DeSantis TZ, Hugenholtz P, Keller K, Brodie EL, Larsen N, Piceno YM et al. (2006). NAST: a multiple sequence alignment server for comparative analysis of 16S rrna genes. Nucleic Acids Res 34: W394 W399. Eichorst SA, Breznak JA, Schmidt TM. (2007). Isolation and characterization of soil bacteria that define Teniglobus gen.nov., in the phylum Acidobacteria. Appl Environ Microbiol 73: Faith DP. (1992). Conservation evaluation and phylogenetic diversity. Biol Conserv 61:
11 452 Felsenstein J. (1985). Phylogenies and the comparative method. Am Nat 125: Felske A, de Vos WM, Akkermans ADL. (2000). Spatial distribution of 16S rrna levels from uncultured Acidobacteria in soil. Lett Appl Microbiol 31: Felske A, Wolterink A, Van Lis R, Akkermans ADL. (1998). Phylogeny of the main bacterial 16S rrna sequences in Drentse A grassland soils (The Netherlands). Appl Environ Microbiol 64: Fierer N, Bradford M, Jackson R. (2007a). Toward an ecological classification of soil bacteria. Ecology 88: Fierer N, Jackson JA, Vilgalys R, Jackson RB. (2005). Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71: Fierer N, Jackson R. (2006). The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103: Fierer N, Morse JL, Berthrong ST, Bernhardt ES, Jackson RB. (2007b). Environmental controls on the landscapescale biogeography of stream bacterial communities. Ecology 88: Giovannoni S, Stingl U. (2007). The importance of culturing bacterioplankton in the omics age. Nat Rev Microbiol 5: Giovannoni SJ, Britschgi TB, Moyer CL, Field KG. (1990). Genetic diversity in Sargasso sea bacterioplankton. Nature 345: Hamady M, Walker J, Harris J, Gold N, Knight R. (2008). Error-correcting barcoded primers allow hundreds of samples to be pyrosequenced in multiplex. Nat Methods 5: Harvey PH, Pagel MD. (1991). The Comparative Method in Evolutionary Biology. Oxford University Press: Oxford. Hiraishi A, Kishimoto N, Kosako Y, Wakao N, Tano T. (1995). Phylogenetic position of the menaquinonecontaining acidophilic chemo-organotroph Acidobacterium-capsulatum. FEMS Microbiol Lett 132: Hogberg MN, Hogberg P, Myrold DD. (2007). Is microbial community composition in boreal forest soils determined by ph, C-to-N ratio, the trees, or all three? Oecologia 150: Huelsenbeck JP, Rannala B, Masly JP. (2000). Accommodating phylogenetic uncertainty in evolutionary studies. Science 288: Hugenholtz P, Goebel BM, Pace NR. (1998). Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. J Bacteriol 180: Janssen PH. (2006). Identifying the dominant soil bacterial taxa in libraries of 16S rrna and 16S rrna genes. Appl Environ Microbiol 72: Jones RT, Martin AP. (2006). Testing for differentiation of microbial communities using phylogenetic methods: accounting for uncertainty of phylogenetic inference and character state mapping. Microb Ecol 52: Joseph SJ, Hugenholtz P, Sangwan P, Osborne CA, Janssen PH. (2003). Laboratory cultivation of widespread and previously uncultured soil bacteria. Appl Environ Microbiol 69: Kishimoto N, Kosako Y, Tano T. (1991). Acidobacteriumcapsulatum Gen-Nov, Sp-Nov an acidophilic chemoorganotrophic bacterium containing menaquinone from acidic mineral environment. Curr Microbiol 22: 1 7. Kuske CR, Barns SM, Busch JD. (1997). Diverse uncultivated bacterial groups from soils of the arid southwestern United States that are present in many geographic regions. Appl Environ Microbiol 63: Lauber CL, Strickland MS, Bradford MA, Fierer N. (2008). The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40: Liu Z, Lozupone C, Hamady M, Bushman F, Knight R. (2007). Short pyrosequencing reads suffice for accurate microbial community analysis. Nucleic Acids Res 35: e120. Liu ZZ, DeSantis TZ, Andersen GL, Knight R. (2008). Accurate taxonomy assignments from 16S rrna sequences produced by highly parallel pyrosequencers. Nucleic Acids Res 36: e120. Lozupone C, Hamady M, Knight R. (2006). UniFrac an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinformatics 7: 371. Lozupone C, Knight R. (2005). UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71: Lozupone CA, Knight R. (2008). Species divergence and the measurement of microbial diversity. FEMS Microbiol Rev 32: Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar BA et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: Mannisto MK, Tiirola M, Haggblom MM. (2007). Bacterial communities in Arctic fjelds of Finnish Lapland are stable but highly ph-dependent. FEMS Microbiol Ecol 59: Morris RM, Rappe MS, Connon SA, Vergin KL, Siebold WA, Carlson CA et al. (2002). SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420: Nemergut DR, Anderson SP, Cleveland CC, Martin AP, Miller AE, Seimon A et al. (2007). Microbial community succession in an unvegetated, recently deglaciated soil. Microb Ecol 53: Newton RJ, Jones SE, Helmus MR, McMahon KD. (2007). Phylogenetic ecology of the freshwater Actinobacteria aci lineage. Appl Environ Microbiol 73: Pankratov TA, Serkebaeva YM, Kulichevskaya IS, Liesack W, Dedysh SN. (2008). Substrate-induced growth and isolation of Acidobacteria from acidic Sphagnum peat. ISME J 2: Percent SF, Frischer ME, Vescio PA, Duffy EB, Milano V, McLellan M et al. (2008). Bacterial community structure of acid-impacted lakes: what controls diversity? Appl Environ Microbiol 74: Quaiser A, Ochsenreiter T, Lanz C, Schuster SC, Treusch AH, Eck J et al. (2003). Acidobacteria form a coherent but highly diverse group within the bacterial domain: evidence from environmental genomics. Mol Microbiol 50: Rappe MS, Connon SA, Vergin KL, Giovannoni SJ. (2002). Cultivation of the ubiquitous SAR11 marine bacterioplankton clade. Nature 418: Sait M, Davis KER, Janssen PH. (2006). Effect of ph on isolation and distribution of members of subdivision 1 of the phylum Acidobacteria occurring in soil. Appl Environ Microbiol 72:
Outline Classes of diversity measures. Species Divergence and the Measurement of Microbial Diversity. How do we describe and compare diversity?
 Species Divergence and the Measurement of Microbial Diversity Cathy Lozupone University of Colorado, Boulder. Washington University, St Louis. Outline Classes of diversity measures α vs β diversity Quantitative
Species Divergence and the Measurement of Microbial Diversity Cathy Lozupone University of Colorado, Boulder. Washington University, St Louis. Outline Classes of diversity measures α vs β diversity Quantitative
Chad Burrus April 6, 2010
 Chad Burrus April 6, 2010 1 Background What is UniFrac? Materials and Methods Results Discussion Questions 2 The vast majority of microbes cannot be cultured with current methods Only half (26) out of
Chad Burrus April 6, 2010 1 Background What is UniFrac? Materials and Methods Results Discussion Questions 2 The vast majority of microbes cannot be cultured with current methods Only half (26) out of
Phylogenetic diversity and conservation
 Phylogenetic diversity and conservation Dan Faith The Australian Museum Applied ecology and human dimensions in biological conservation Biota Program/ FAPESP Nov. 9-10, 2009 BioGENESIS Providing an evolutionary
Phylogenetic diversity and conservation Dan Faith The Australian Museum Applied ecology and human dimensions in biological conservation Biota Program/ FAPESP Nov. 9-10, 2009 BioGENESIS Providing an evolutionary
Exploring Microbes in the Sea. Alma Parada Postdoctoral Scholar Stanford University
 Exploring Microbes in the Sea Alma Parada Postdoctoral Scholar Stanford University Cruising the ocean to get us some microbes It s all about the Microbe! Microbes = microorganisms an organism that requires
Exploring Microbes in the Sea Alma Parada Postdoctoral Scholar Stanford University Cruising the ocean to get us some microbes It s all about the Microbe! Microbes = microorganisms an organism that requires
Microbiome: 16S rrna Sequencing 3/30/2018
 Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley D.D. Ackerly April 16, 2014. Community Ecology and Phylogenetics Readings: Cavender-Bares,
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley D.D. Ackerly April 16, 2014. Community Ecology and Phylogenetics Readings: Cavender-Bares,
Taxonomy and Clustering of SSU rrna Tags. Susan Huse Josephine Bay Paul Center August 5, 2013
 Taxonomy and Clustering of SSU rrna Tags Susan Huse Josephine Bay Paul Center August 5, 2013 Primary Methods of Taxonomic Assignment Bayesian Kmer Matching RDP http://rdp.cme.msu.edu Wang, et al (2007)
Taxonomy and Clustering of SSU rrna Tags Susan Huse Josephine Bay Paul Center August 5, 2013 Primary Methods of Taxonomic Assignment Bayesian Kmer Matching RDP http://rdp.cme.msu.edu Wang, et al (2007)
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Detailed overview of the primer-free full-length SSU rrna library preparation.
 Supplementary Figure 1 Detailed overview of the primer-free full-length SSU rrna library preparation. Detailed overview of the primer-free full-length SSU rrna library preparation. Supplementary Figure
Supplementary Figure 1 Detailed overview of the primer-free full-length SSU rrna library preparation. Detailed overview of the primer-free full-length SSU rrna library preparation. Supplementary Figure
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy. Slowly evolving molecules (e.g., rrna) used for large-scale structure; "fast- clock" molecules for fine-structure.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder
 Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder nrpace@colorado.edu Outline What is the Tree of Life? -- Historical Conceptually
Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder nrpace@colorado.edu Outline What is the Tree of Life? -- Historical Conceptually
Bacterial Community Variation in Human Body Habitats Across Space and Time
 Bacterial Community Variation in Human Body Habitats Across Space and Time Elizabeth K. Costello, 1 Christian L. Lauber, 2 Micah Hamady, 3 Noah Fierer, 2,4 Jeffrey I. Gordon, 5 Rob Knight 1* 1 Department
Bacterial Community Variation in Human Body Habitats Across Space and Time Elizabeth K. Costello, 1 Christian L. Lauber, 2 Micah Hamady, 3 Noah Fierer, 2,4 Jeffrey I. Gordon, 5 Rob Knight 1* 1 Department
8/23/2014. Phylogeny and the Tree of Life
 Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
PHYLOGENY AND SYSTEMATICS
 AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
Comparison of Three Fugal ITS Reference Sets. Qiong Wang and Jim R. Cole
 RDP TECHNICAL REPORT Created 04/12/2014, Updated 08/08/2014 Summary Comparison of Three Fugal ITS Reference Sets Qiong Wang and Jim R. Cole wangqion@msu.edu, colej@msu.edu In this report, we evaluate the
RDP TECHNICAL REPORT Created 04/12/2014, Updated 08/08/2014 Summary Comparison of Three Fugal ITS Reference Sets Qiong Wang and Jim R. Cole wangqion@msu.edu, colej@msu.edu In this report, we evaluate the
Bacterial Communities in Women with Bacterial Vaginosis: High Resolution Phylogenetic Analyses Reveal Relationships of Microbiota to Clinical Criteria
 Bacterial Communities in Women with Bacterial Vaginosis: High Resolution Phylogenetic Analyses Reveal Relationships of Microbiota to Clinical Criteria Seminar presentation Pierre Barbera Supervised by:
Bacterial Communities in Women with Bacterial Vaginosis: High Resolution Phylogenetic Analyses Reveal Relationships of Microbiota to Clinical Criteria Seminar presentation Pierre Barbera Supervised by:
Bacillus anthracis. Last Lecture: 1. Introduction 2. History 3. Koch s Postulates. 1. Prokaryote vs. Eukaryote 2. Classifying prokaryotes
 Last Lecture: Bacillus anthracis 1. Introduction 2. History 3. Koch s Postulates Today s Lecture: 1. Prokaryote vs. Eukaryote 2. Classifying prokaryotes 3. Phylogenetics I. Basic Cell structure: (Fig.
Last Lecture: Bacillus anthracis 1. Introduction 2. History 3. Koch s Postulates Today s Lecture: 1. Prokaryote vs. Eukaryote 2. Classifying prokaryotes 3. Phylogenetics I. Basic Cell structure: (Fig.
Rank-abundance. Geometric series: found in very communities such as the
 Rank-abundance Geometric series: found in very communities such as the Log series: group of species that occur _ time are the most frequent. Useful for calculating a diversity metric (Fisher s alpha) Most
Rank-abundance Geometric series: found in very communities such as the Log series: group of species that occur _ time are the most frequent. Useful for calculating a diversity metric (Fisher s alpha) Most
Supplementary Information
 Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Howard Hughes Medical Institute, Chevy Chase, Maryland USA
 Reports Ecology, 92(4), 2011, pp. 797 804 Ó 2011 by the Ecological Society of America Microbes do not follow the elevational diversity patterns of plants and animals NOAH FIERER, 1,2,8 CHRISTY M. MCCAIN,
Reports Ecology, 92(4), 2011, pp. 797 804 Ó 2011 by the Ecological Society of America Microbes do not follow the elevational diversity patterns of plants and animals NOAH FIERER, 1,2,8 CHRISTY M. MCCAIN,
Microbiota: Its Evolution and Essence. Hsin-Jung Joyce Wu "Microbiota and man: the story about us
 Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics.
 Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics. Tree topologies Two topologies were examined, one favoring
Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics. Tree topologies Two topologies were examined, one favoring
UoN, CAS, DBSC BIOL102 lecture notes by: Dr. Mustafa A. Mansi. The Phylogenetic Systematics (Phylogeny and Systematics)
 - Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
- Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
Impact of training sets on classification of high-throughput bacterial 16s rrna gene surveys
 (2012) 6, 94 103 & 2012 International Society for Microbial Ecology All rights reserved 1751-7362/12 www.nature.com/ismej ORIGINAL ARTICLE Impact of training sets on classification of high-throughput bacterial
(2012) 6, 94 103 & 2012 International Society for Microbial Ecology All rights reserved 1751-7362/12 www.nature.com/ismej ORIGINAL ARTICLE Impact of training sets on classification of high-throughput bacterial
Microbial Taxonomy and the Evolution of Diversity
 19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
Supplementary Material
 Supplementary Material The impact of logging and forest conversion to oil palm on soil bacterial communities in Borneo Larisa Lee-Cruz 1, David P. Edwards 2,3, Binu Tripathi 1, Jonathan M. Adams 1* 1 Department
Supplementary Material The impact of logging and forest conversion to oil palm on soil bacterial communities in Borneo Larisa Lee-Cruz 1, David P. Edwards 2,3, Binu Tripathi 1, Jonathan M. Adams 1* 1 Department
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Microbial Diversity and Assessment (II) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Taxonomical Classification using:
 Taxonomical Classification using: Extracting ecological signal from noise: introduction to tools for the analysis of NGS data from microbial communities Bergen, April 19-20 2012 INTRODUCTION Taxonomical
Taxonomical Classification using: Extracting ecological signal from noise: introduction to tools for the analysis of NGS data from microbial communities Bergen, April 19-20 2012 INTRODUCTION Taxonomical
Supporting online material
 Supporting online material Materials and Methods Target proteins All predicted ORFs in the E. coli genome (1) were downloaded from the Colibri data base (2) (http://genolist.pasteur.fr/colibri/). 737 proteins
Supporting online material Materials and Methods Target proteins All predicted ORFs in the E. coli genome (1) were downloaded from the Colibri data base (2) (http://genolist.pasteur.fr/colibri/). 737 proteins
Microbial Taxonomy. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Final Report- Anna-Louise Reysenbach, Portland State University. Project ID Number: Project Title: Lead Principal Investigator:
 Final Report- Anna-Louise Reysenbach, Portland State University. This material is based upon work supported by the U.S. Department of Energy under award #70206. Any opinions, findings and conclusions or
Final Report- Anna-Louise Reysenbach, Portland State University. This material is based upon work supported by the U.S. Department of Energy under award #70206. Any opinions, findings and conclusions or
Chapter 19. Microbial Taxonomy
 Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
The Role of the Horizontal Gene Pool and Lateral Gene Transfer in Enhancing Microbial Activities in Marine Sediments
 The Role of the Horizontal Gene Pool and Lateral Gene Transfer in Enhancing Microbial Activities in Marine Sediments Patricia A. Sobecky School of Biology Georgia Institute of Technology 310 Ferst Drive
The Role of the Horizontal Gene Pool and Lateral Gene Transfer in Enhancing Microbial Activities in Marine Sediments Patricia A. Sobecky School of Biology Georgia Institute of Technology 310 Ferst Drive
RNA Transport. R preps R preps
 RNA Transport R0527-00 5 preps R0527-01 50 preps July 2014 RNA Transport Table of Contents Introduction...2 Kit Contents/Storage and Stability...3 Protocol...4 Storage Procedure...4 Recovery Procedure...5
RNA Transport R0527-00 5 preps R0527-01 50 preps July 2014 RNA Transport Table of Contents Introduction...2 Kit Contents/Storage and Stability...3 Protocol...4 Storage Procedure...4 Recovery Procedure...5
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time
 What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
Bergey s Manual Classification Scheme. Vertical inheritance and evolutionary mechanisms
 Bergey s Manual Classification Scheme Gram + Gram - No wall Funny wall Vertical inheritance and evolutionary mechanisms a b c d e * * a b c d e * a b c d e a b c d e * a b c d e Accumulation of neutral
Bergey s Manual Classification Scheme Gram + Gram - No wall Funny wall Vertical inheritance and evolutionary mechanisms a b c d e * * a b c d e * a b c d e a b c d e * a b c d e Accumulation of neutral
NOVABEADS FOOD 1 DNA KIT
 NOVABEADS FOOD 1 DNA KIT NOVABEADS FOOD DNA KIT is the new generation tool in molecular biology techniques and allows DNA isolations from highly processed food products. The method is based on the use
NOVABEADS FOOD 1 DNA KIT NOVABEADS FOOD DNA KIT is the new generation tool in molecular biology techniques and allows DNA isolations from highly processed food products. The method is based on the use
Soil microbial responses to climate perturbations Sarah E. Evans Kellogg Biological Station Michigan State University
 Soil microbial responses to climate perturbations Sarah E. Evans Kellogg Biological Station Michigan State University The soil environment Harsh environment, many abiotic stressors Highly diverse, many
Soil microbial responses to climate perturbations Sarah E. Evans Kellogg Biological Station Michigan State University The soil environment Harsh environment, many abiotic stressors Highly diverse, many
A biogeographic distribution of magnetotactic bacteria influenced by salinity
 (2012) 6, 475 479 & 2012 International Society for Microbial Ecology All rights reserved 1751-7362/12 www.nature.com/ismej SHORT COMMUNICATION A biogeographic distribution of magnetotactic bacteria influenced
(2012) 6, 475 479 & 2012 International Society for Microbial Ecology All rights reserved 1751-7362/12 www.nature.com/ismej SHORT COMMUNICATION A biogeographic distribution of magnetotactic bacteria influenced
Origins of Life. Fundamental Properties of Life. Conditions on Early Earth. Evolution of Cells. The Tree of Life
 The Tree of Life Chapter 26 Origins of Life The Earth formed as a hot mass of molten rock about 4.5 billion years ago (BYA) -As it cooled, chemically-rich oceans were formed from water condensation Life
The Tree of Life Chapter 26 Origins of Life The Earth formed as a hot mass of molten rock about 4.5 billion years ago (BYA) -As it cooled, chemically-rich oceans were formed from water condensation Life
Mag-Bind Soil DNA Kit. M preps M preps M preps
 Mag-Bind Soil DNA Kit M5635-00 5 preps M5635-01 50 preps M5635-02 200 preps January 2013 Mag-Bind Soil DNA Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Preparing
Mag-Bind Soil DNA Kit M5635-00 5 preps M5635-01 50 preps M5635-02 200 preps January 2013 Mag-Bind Soil DNA Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Preparing
Chapter 26: Phylogeny and the Tree of Life Phylogenies Show Evolutionary Relationships
 Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 19: Taxonomy, Systematics, and Phylogeny
 Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
SPECIATION. REPRODUCTIVE BARRIERS PREZYGOTIC: Barriers that prevent fertilization. Habitat isolation Populations can t get together
 SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
This is a repository copy of Microbiology: Mind the gaps in cellular evolution.
 This is a repository copy of Microbiology: Mind the gaps in cellular evolution. White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/114978/ Version: Accepted Version Article:
This is a repository copy of Microbiology: Mind the gaps in cellular evolution. White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/114978/ Version: Accepted Version Article:
Microbes and mountains: metagenetics on Mount Fuji, Japan. Jonathan Adams, Biology Department, SNU, Korea
 Microbes and mountains: metagenetics on Mount Fuji, Japan Jonathan Adams, Biology Department, SNU, Korea jonadams@snu.ac.kr Until about a decade ago, culturing could only yield 8,000 described species
Microbes and mountains: metagenetics on Mount Fuji, Japan Jonathan Adams, Biology Department, SNU, Korea jonadams@snu.ac.kr Until about a decade ago, culturing could only yield 8,000 described species
Chapter 8. Biogeographic Processes. Upon completion of this chapter the student will be able to:
 Chapter 8 Biogeographic Processes Chapter Objectives Upon completion of this chapter the student will be able to: 1. Define the terms ecosystem, habitat, ecological niche, and community. 2. Outline how
Chapter 8 Biogeographic Processes Chapter Objectives Upon completion of this chapter the student will be able to: 1. Define the terms ecosystem, habitat, ecological niche, and community. 2. Outline how
Amplicon Sequencing. Dr. Orla O Sullivan SIRG Research Fellow Teagasc
 Amplicon Sequencing Dr. Orla O Sullivan SIRG Research Fellow Teagasc What is Amplicon Sequencing? Sequencing of target genes (are regions of ) obtained by PCR using gene specific primers. Why do we do
Amplicon Sequencing Dr. Orla O Sullivan SIRG Research Fellow Teagasc What is Amplicon Sequencing? Sequencing of target genes (are regions of ) obtained by PCR using gene specific primers. Why do we do
Marine Resources Development Foundation/MarineLab Grades: 9, 10, 11, 12 States: AP Biology Course Description Subjects: Science
 Marine Resources Development Foundation/MarineLab Grades: 9, 10, 11, 12 States: AP Biology Course Description Subjects: Science Highlighted components are included in Tallahassee Museum s 2016 program
Marine Resources Development Foundation/MarineLab Grades: 9, 10, 11, 12 States: AP Biology Course Description Subjects: Science Highlighted components are included in Tallahassee Museum s 2016 program
Lowndes County Biology II Pacing Guide Approximate
 Lowndes County Biology II Pacing Guide 2009-2010 MS Frameworks Pacing Guide Worksheet Grade Level: Biology II Grading Period: 1 st 9 weeks Chapter/Unit Lesson Topic Objective Number 1 The Process of 1.
Lowndes County Biology II Pacing Guide 2009-2010 MS Frameworks Pacing Guide Worksheet Grade Level: Biology II Grading Period: 1 st 9 weeks Chapter/Unit Lesson Topic Objective Number 1 The Process of 1.
Text of objective. Investigate and describe the structure and functions of cells including: Cell organelles
 This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
Global patterns in the biogeography of bacterial taxaemi_2315
 1..10 Environmental Microbiology (2010) doi:10.1111/j.1462-2920.2010.02315.x Global patterns in the biogeography of bacterial taxaemi_2315 Diana R. Nemergut, 1,2 * Elizabeth K. Costello, 3 Micah Hamady,
1..10 Environmental Microbiology (2010) doi:10.1111/j.1462-2920.2010.02315.x Global patterns in the biogeography of bacterial taxaemi_2315 Diana R. Nemergut, 1,2 * Elizabeth K. Costello, 3 Micah Hamady,
The Effect of Primer Choice and Short Read Sequences on the Outcome of 16S rrna Gene Based Diversity Studies
 The Effect of Primer Choice and Short Read Sequences on the Outcome of 16S rrna Gene Based Diversity Studies Jonas Ghyselinck 1 *., Stefan Pfeiffer 2 *., Kim Heylen 1, Angela Sessitsch 2, Paul De Vos 1
The Effect of Primer Choice and Short Read Sequences on the Outcome of 16S rrna Gene Based Diversity Studies Jonas Ghyselinck 1 *., Stefan Pfeiffer 2 *., Kim Heylen 1, Angela Sessitsch 2, Paul De Vos 1
MICROBIOLOGY ECOLOGY. Chronic N-amended soils exhibit an altered bacterial community structure in Harvard Forest, MA, USA.
 RESEARCH ARTICLE Chronic N-amended soils exhibit an altered bacterial community structure in Harvard Forest, MA, USA Swathi A. Turlapati 1, Rakesh Minocha 2, Premsai S. Bhiravarasa 1, Louis S. Tisa 3,
RESEARCH ARTICLE Chronic N-amended soils exhibit an altered bacterial community structure in Harvard Forest, MA, USA Swathi A. Turlapati 1, Rakesh Minocha 2, Premsai S. Bhiravarasa 1, Louis S. Tisa 3,
Map of AP-Aligned Bio-Rad Kits with Learning Objectives
 Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Culturing captures members of the soil rare biosphereemi_
 bs_bs_banner Environmental Microbiology (2012) doi:10.1111/j.1462-2920.2012.02817.x Correspondence Culturing captures members of the soil rare biosphereemi_2817 1..6 Ashley Shade, 1 Clifford S. Hogan,
bs_bs_banner Environmental Microbiology (2012) doi:10.1111/j.1462-2920.2012.02817.x Correspondence Culturing captures members of the soil rare biosphereemi_2817 1..6 Ashley Shade, 1 Clifford S. Hogan,
Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae
 Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae Frederik Schulz, Ilias Lagkouvardos, Florian Wascher, Karin Aistleitner, Rok Kostanjšek, Matthias Horn Supplementary
Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae Frederik Schulz, Ilias Lagkouvardos, Florian Wascher, Karin Aistleitner, Rok Kostanjšek, Matthias Horn Supplementary
Supplementary Information
 Supplementary Information For the article"comparable system-level organization of Archaea and ukaryotes" by J. Podani, Z. N. Oltvai, H. Jeong, B. Tombor, A.-L. Barabási, and. Szathmáry (reference numbers
Supplementary Information For the article"comparable system-level organization of Archaea and ukaryotes" by J. Podani, Z. N. Oltvai, H. Jeong, B. Tombor, A.-L. Barabási, and. Szathmáry (reference numbers
Valley Central School District 944 State Route 17K Montgomery, NY Telephone Number: (845) ext Fax Number: (845)
 Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
ADVANCED PLACEMENT BIOLOGY
 ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
(This is a sample cover image for this issue. The actual cover is not yet available at this time.)
 (This is a sample cover image for this issue. The actual cover is not yet available at this time.) This article appeared in a journal published by Elsevier. The attached copy is furnished to the author
(This is a sample cover image for this issue. The actual cover is not yet available at this time.) This article appeared in a journal published by Elsevier. The attached copy is furnished to the author
DETECTING BIOLOGICAL AND ENVIRONMENTAL CHANGES: DESIGN AND ANALYSIS OF MONITORING AND EXPERIMENTS (University of Bologna, 3-14 March 2008)
 Dipartimento di Biologia Evoluzionistica Sperimentale Centro Interdipartimentale di Ricerca per le Scienze Ambientali in Ravenna INTERNATIONAL WINTER SCHOOL UNIVERSITY OF BOLOGNA DETECTING BIOLOGICAL AND
Dipartimento di Biologia Evoluzionistica Sperimentale Centro Interdipartimentale di Ricerca per le Scienze Ambientali in Ravenna INTERNATIONAL WINTER SCHOOL UNIVERSITY OF BOLOGNA DETECTING BIOLOGICAL AND
A. Incorrect! In the binomial naming convention the Kingdom is not part of the name.
 Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
Post-doc fellowships to non-eu researchers FINAL REPORT. Home Institute: Centro de Investigaciones Marinas, Universidad de La Habana, CUBA
 Recipient: Maickel Armenteros Almanza. Post-doc fellowships to non-eu researchers FINAL REPORT Home Institute: Centro de Investigaciones Marinas, Universidad de La Habana, CUBA Promoter: Prof. Dr. Wilfrida
Recipient: Maickel Armenteros Almanza. Post-doc fellowships to non-eu researchers FINAL REPORT Home Institute: Centro de Investigaciones Marinas, Universidad de La Habana, CUBA Promoter: Prof. Dr. Wilfrida
Links between Plant and Fungal Diversity in Habitat Fragments of Coastal Sage Scrub
 26-211 Mission Kearney Foundation of Soil Science: Understanding and Managing Soil-Ecosystem Functions Across Spatial and Temporal Scales Final Report: 271, 1/1/29-12/31/29 Links between Plant and Fungal
26-211 Mission Kearney Foundation of Soil Science: Understanding and Managing Soil-Ecosystem Functions Across Spatial and Temporal Scales Final Report: 271, 1/1/29-12/31/29 Links between Plant and Fungal
Supplementary Figure S1
 Supplementary Figure S1 KRO STO R-p R-p3 RUD R-nt L-n L-h L-l ph 7 WEON [mg/kg dw soil] 3 WEOC [mg/kg dw soil] 3 Soil type sandy sandy loam silty clay GSF B77V B77T E16 Figure S1:
Supplementary Figure S1 KRO STO R-p R-p3 RUD R-nt L-n L-h L-l ph 7 WEON [mg/kg dw soil] 3 WEOC [mg/kg dw soil] 3 Soil type sandy sandy loam silty clay GSF B77V B77T E16 Figure S1:
An introduction to the picante package
 An introduction to the picante package Steven Kembel (steve.kembel@gmail.com) April 2010 Contents 1 Installing picante 1 2 Data formats in picante 2 2.1 Phylogenies................................ 2 2.2
An introduction to the picante package Steven Kembel (steve.kembel@gmail.com) April 2010 Contents 1 Installing picante 1 2 Data formats in picante 2 2.1 Phylogenies................................ 2 2.2
Biology 211 (2) Week 1 KEY!
 Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Stepping stones towards a new electronic prokaryotic taxonomy. The ultimate goal in taxonomy. Pragmatic towards diagnostics
 Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
Introductory Microbiology Dr. Hala Al Daghistani
 Introductory Microbiology Dr. Hala Al Daghistani Why Study Microbes? Microbiology is the branch of biological sciences concerned with the study of the microbes. 1. Microbes and Man in Sickness and Health
Introductory Microbiology Dr. Hala Al Daghistani Why Study Microbes? Microbiology is the branch of biological sciences concerned with the study of the microbes. 1. Microbes and Man in Sickness and Health
Using Topological Data Analysis to find discrimination between microbial states in human microbiome data
 Using Topological Data Analysis to find discrimination between microbial states in human microbiome data Mehrdad Yazdani 1,2, Larry Smarr 1,3 and Rob Knight 4 1 California Institute for Telecommunications
Using Topological Data Analysis to find discrimination between microbial states in human microbiome data Mehrdad Yazdani 1,2, Larry Smarr 1,3 and Rob Knight 4 1 California Institute for Telecommunications
Other resources. Greengenes (bacterial) Silva (bacteria, archaeal and eukarya)
 General QIIME resources http://qiime.org/ Blog (news, updates): http://qiime.wordpress.com/ Support/forum: https://groups.google.com/forum/#!forum/qiimeforum Citing QIIME: Caporaso, J.G. et al., QIIME
General QIIME resources http://qiime.org/ Blog (news, updates): http://qiime.wordpress.com/ Support/forum: https://groups.google.com/forum/#!forum/qiimeforum Citing QIIME: Caporaso, J.G. et al., QIIME
An Automated Phylogenetic Tree-Based Small Subunit rrna Taxonomy and Alignment Pipeline (STAP)
 An Automated Phylogenetic Tree-Based Small Subunit rrna Taxonomy and Alignment Pipeline (STAP) Dongying Wu 1 *, Amber Hartman 1,6, Naomi Ward 4,5, Jonathan A. Eisen 1,2,3 1 UC Davis Genome Center, University
An Automated Phylogenetic Tree-Based Small Subunit rrna Taxonomy and Alignment Pipeline (STAP) Dongying Wu 1 *, Amber Hartman 1,6, Naomi Ward 4,5, Jonathan A. Eisen 1,2,3 1 UC Davis Genome Center, University
Supplementary Information
 Supplementary Information Table S1. Per-sample sequences, observed OTUs, richness estimates, diversity indices and coverage. Samples codes as follows: YED (Young leaves Endophytes), MED (Mature leaves
Supplementary Information Table S1. Per-sample sequences, observed OTUs, richness estimates, diversity indices and coverage. Samples codes as follows: YED (Young leaves Endophytes), MED (Mature leaves
ADDITIONAL STATISTICAL ANALYSES. The data were not normally distributed (Kolmogorov-Smirnov test; Legendre &
 DDITIONL STTISTICL NLYSES The data were not normally distributed (Kolmogorov-Smirnov test; Legendre & Legendre, 1998) and violate the assumption of equal variances (Levene test; Rohlf & Sokal, 1994) for
DDITIONL STTISTICL NLYSES The data were not normally distributed (Kolmogorov-Smirnov test; Legendre & Legendre, 1998) and violate the assumption of equal variances (Levene test; Rohlf & Sokal, 1994) for
II. Deep insight into plant habitats
 II. Deep insight into plant habitats Sugar Beet The seed microbiome project (ACIB) Christin Zachow, Henry Müller Ralf Tilcher (KWS SAAT AG) Cultivar-specific microbiomes Experimental design Genetic pool
II. Deep insight into plant habitats Sugar Beet The seed microbiome project (ACIB) Christin Zachow, Henry Müller Ralf Tilcher (KWS SAAT AG) Cultivar-specific microbiomes Experimental design Genetic pool
FIG S1: Rarefaction analysis of observed richness within Drosophila. All calculations were
 Page 1 of 14 FIG S1: Rarefaction analysis of observed richness within Drosophila. All calculations were performed using mothur (2). OTUs were defined at the 3% divergence threshold using the average neighbor
Page 1 of 14 FIG S1: Rarefaction analysis of observed richness within Drosophila. All calculations were performed using mothur (2). OTUs were defined at the 3% divergence threshold using the average neighbor
Comparisons of Microbial Communities in a Sequencing Batch Reactor (Cromaglass Corporation) at Two Time Increments
 Comparisons of Microbial Communities in a Sequencing Batch Reactor (Cromaglass Corporation) at Two Time Increments By: Brittane Miller and Jeff Newman Lycoming College Williamsport, Pennsylvania What is
Comparisons of Microbial Communities in a Sequencing Batch Reactor (Cromaglass Corporation) at Two Time Increments By: Brittane Miller and Jeff Newman Lycoming College Williamsport, Pennsylvania What is
Studying the effect of species dominance on diversity patterns using Hill numbers-based indices
 Studying the effect of species dominance on diversity patterns using Hill numbers-based indices Loïc Chalmandrier Loïc Chalmandrier Diversity pattern analysis November 8th 2017 1 / 14 Introduction Diversity
Studying the effect of species dominance on diversity patterns using Hill numbers-based indices Loïc Chalmandrier Loïc Chalmandrier Diversity pattern analysis November 8th 2017 1 / 14 Introduction Diversity
Name: Class: Date: ID: A
 Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
Globin-Zero Gold Kit
 Cat. No. GZG1206 (Contains 1 box of Cat. No. GZRR1306 and 1 box of Cat. No. MRZ116C) Cat. No. GZG1224 (Contains 1 box of Cat. No. GZRR1324 and 1 box of Cat. No. MRZ11124C) Connect with Epicentre on our
Cat. No. GZG1206 (Contains 1 box of Cat. No. GZRR1306 and 1 box of Cat. No. MRZ116C) Cat. No. GZG1224 (Contains 1 box of Cat. No. GZRR1324 and 1 box of Cat. No. MRZ11124C) Connect with Epicentre on our
Lecture 3: Mixture Models for Microbiome data. Lecture 3: Mixture Models for Microbiome data
 Lecture 3: Mixture Models for Microbiome data 1 Lecture 3: Mixture Models for Microbiome data Outline: - Mixture Models (Negative Binomial) - DESeq2 / Don t Rarefy. Ever. 2 Hypothesis Tests - reminder
Lecture 3: Mixture Models for Microbiome data 1 Lecture 3: Mixture Models for Microbiome data Outline: - Mixture Models (Negative Binomial) - DESeq2 / Don t Rarefy. Ever. 2 Hypothesis Tests - reminder
Supplementary Information
 Supplementary Information Altitudinal patterns of diversity and functional traits of metabolically active microorganisms in stream biofilms Linda Wilhelm 1, Katharina Besemer 2, Lena Fragner 3, Hannes
Supplementary Information Altitudinal patterns of diversity and functional traits of metabolically active microorganisms in stream biofilms Linda Wilhelm 1, Katharina Besemer 2, Lena Fragner 3, Hannes
Bacterial Communities Associated with the Lichen Symbiosis
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Feb. 2011, p. 1309 1314 Vol. 77, No. 4 0099-2240/11/$12.00 doi:10.1128/aem.02257-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Bacterial
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Feb. 2011, p. 1309 1314 Vol. 77, No. 4 0099-2240/11/$12.00 doi:10.1128/aem.02257-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Bacterial
A Study of the Moss Parasite Eocronartium muscicola By: Alicia Knudson Advisor: Dr. Elizabeth Frieders
 A Study of the Moss Parasite Eocronartium muscicola By: Alicia Knudson Advisor: Dr. Elizabeth Frieders Abstract The genus Eocronartium contains a single described species of parasitic fungus on moss plants
A Study of the Moss Parasite Eocronartium muscicola By: Alicia Knudson Advisor: Dr. Elizabeth Frieders Abstract The genus Eocronartium contains a single described species of parasitic fungus on moss plants
Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles
 Electronic Supplementary Material Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles Ignacio Morales-Castilla 1,2 *, Jonathan T. Davies 3 and
Electronic Supplementary Material Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles Ignacio Morales-Castilla 1,2 *, Jonathan T. Davies 3 and
Cladistics and Bioinformatics Questions 2013
 AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Big Idea 1: The process of evolution drives the diversity and unity of life.
 Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies
 The following supplement accompanies the article Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies Richard L. Hahnke 1, Christina Probian 1, Bernhard M.
The following supplement accompanies the article Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies Richard L. Hahnke 1, Christina Probian 1, Bernhard M.
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
Phylogenetic organization of bacterial activity
 (2016) 10, 2336 2340 2016 International Society for Microbial Ecology All rights reserved 1751-7362/16 www.nature.com/ismej SHORT COMMUNICATION of bacterial activity OPEN Ember M Morrissey 1,6, Rebecca
(2016) 10, 2336 2340 2016 International Society for Microbial Ecology All rights reserved 1751-7362/16 www.nature.com/ismej SHORT COMMUNICATION of bacterial activity OPEN Ember M Morrissey 1,6, Rebecca
INSTITUT FÜR ANGEWANDTE LABORANALYSEN GMBH. First-Beer Magnetic DNA Kit. Extraktion von Hefe- und Bakterien-DNA aus Bier und anderen Getränken
 First-Beer INSTITUT FÜR ANGEWANDTE LABORANALYSEN GMBH First-Beer Magnetic DNA Kit Extraktion von Hefe- und Bakterien-DNA aus Bier und anderen Getränken Extraction of bacteria and yeast DNA from beer and
First-Beer INSTITUT FÜR ANGEWANDTE LABORANALYSEN GMBH First-Beer Magnetic DNA Kit Extraktion von Hefe- und Bakterien-DNA aus Bier und anderen Getränken Extraction of bacteria and yeast DNA from beer and
