FIG S1: Rarefaction analysis of observed richness within Drosophila. All calculations were
|
|
|
- Gabriel Oliver
- 6 years ago
- Views:
Transcription
1 Page 1 of 14 FIG S1: Rarefaction analysis of observed richness within Drosophila. All calculations were performed using mothur (2). OTUs were defined at the 3% divergence threshold using the average neighbor clustering algorithm. Library identifiers are given in Table 1.
2 Page 2 of 14 FIG S2: Number of OTUs as a function of genetic distance. Number of OTUs was calculated at all genetic distances from 0 (unique sequences) to the largest distance between any two sequences (0.217 for yeast and for bacteria). Clustering was performed using the average neighbor algorithm in mothur (2). Only the twelve populations for which both yeast and bacterial data (1) are available are included in this analysis.
3 Page 3 of 14 FIG S3: Phylogenetic tree of Ascomycete yeasts (Uploaded as a separate file due to its size) Representative sequences of each OTU are highlighted in blue. Each representative sequence has a unique identifier corresponding to a sequence within the FASTA files available on BioTorrents Additionally, each representative sequence has been given a taxonomy assignment (Dataset S7). Unhighlighted taxa are from the SILVA parc database and each is followed by its NCBI accession number. The NEWICK tree file is available on BioTorrents
4 Page 4 of 14 FIG S4: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Unweighted UniFrac, P=0.426) bacterial communities. This plot represents unweighted UniFrac data (P-value=0.426). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
5 Page 5 of 14 FIG S5: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Weighted, non-normalized UniFrac, P=0.086) bacterial communities. This plot represents weighted, non-normalized UniFrac data (Pvalue=0.086). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
6 Page 6 of 14 FIG S6: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Jaccard, P=0.672) bacterial communities. This plot represents Jaccard data (P-value=0.672). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
7 Page 7 of 14 FIG S7: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Binary Jaccard, P=0.478) bacterial communities. This plot represents binary Jaccard data (P-value=0.478). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
8 Page 8 of 14 FIG S8: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Bray-Curtis, P=0.282) bacterial communities. This plot represents Bray-Curtis data (P-value=0.282). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
9 Page 9 of 14 FIG S9: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Sorensen-Dice, P=0.428) bacterial communities. This plot represents Sorensen-Dice data (P-value=0.428). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
10 Page 10 of 14 FIG S10: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Euclidean, P=0.270) bacterial communities. This plot represents Euclidean data (P-value=0.270). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
11 Page 11 of 14 FIG S11: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Binary Euclidean, P=0.117) bacterial communities. This plot represents binary Euclidean data (P-value=0.117). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
12 Page 12 of 14 FIG S12: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Gower, P=0.139) bacterial communities. This plot represents Gower data (P-value=0.139). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
13 Page 13 of 14 FIG S13: Plot showing procrustes analyses of transformed PCA of yeast and bacterial communities (Manhattan, P=0.088) bacterial communities. This plot represents Manhattan data (P-value=0.088). The KiNG data file for this and the other comparisons (Table 3) is available through BioTorrents
14 Page 14 of Chandler, J. A., J. M. Lang, S. Bhatnagar, J. A. Eisen, and A. Kopp Bacterial Communities of Diverse Drosophila Species: Ecological Context of a Host-Microbe Model System. Plos Genetics Schloss, P. D., S. L. Westcott, T. Ryabin, J. R. Hall, M. Hartmann, E. B. Hollister, R. A. Lesniewski, B. B. Oakley, D. H. Parks, C. J. Robinson, J. W. Sahl, B. Stres, G. G. Thallinger, D. J. Van Horn, and C. F. Weber Introducing mothur: Open-Source, Platform-Independent, Community-Supported Software for Describing and Comparing Microbial Communities. Applied and Environmental Microbiology 75:
Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
Other resources. Greengenes (bacterial) Silva (bacteria, archaeal and eukarya)
 General QIIME resources http://qiime.org/ Blog (news, updates): http://qiime.wordpress.com/ Support/forum: https://groups.google.com/forum/#!forum/qiimeforum Citing QIIME: Caporaso, J.G. et al., QIIME
General QIIME resources http://qiime.org/ Blog (news, updates): http://qiime.wordpress.com/ Support/forum: https://groups.google.com/forum/#!forum/qiimeforum Citing QIIME: Caporaso, J.G. et al., QIIME
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Detailed overview of the primer-free full-length SSU rrna library preparation.
 Supplementary Figure 1 Detailed overview of the primer-free full-length SSU rrna library preparation. Detailed overview of the primer-free full-length SSU rrna library preparation. Supplementary Figure
Supplementary Figure 1 Detailed overview of the primer-free full-length SSU rrna library preparation. Detailed overview of the primer-free full-length SSU rrna library preparation. Supplementary Figure
Taxonomy and Clustering of SSU rrna Tags. Susan Huse Josephine Bay Paul Center August 5, 2013
 Taxonomy and Clustering of SSU rrna Tags Susan Huse Josephine Bay Paul Center August 5, 2013 Primary Methods of Taxonomic Assignment Bayesian Kmer Matching RDP http://rdp.cme.msu.edu Wang, et al (2007)
Taxonomy and Clustering of SSU rrna Tags Susan Huse Josephine Bay Paul Center August 5, 2013 Primary Methods of Taxonomic Assignment Bayesian Kmer Matching RDP http://rdp.cme.msu.edu Wang, et al (2007)
Microbiome: 16S rrna Sequencing 3/30/2018
 Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Example study for granular bioreactor stratification: three. dimensional evaluation of a sulfate reducing granular bioreactor
 Supplementary Information for Example study for granular bioreactor stratification: three dimensional evaluation of a sulfate reducing granular bioreactor Tian-wei Hao 1, Jing-hai Luo 1, Kui-zu Su 2, Li
Supplementary Information for Example study for granular bioreactor stratification: three dimensional evaluation of a sulfate reducing granular bioreactor Tian-wei Hao 1, Jing-hai Luo 1, Kui-zu Su 2, Li
Amplicon Sequencing. Dr. Orla O Sullivan SIRG Research Fellow Teagasc
 Amplicon Sequencing Dr. Orla O Sullivan SIRG Research Fellow Teagasc What is Amplicon Sequencing? Sequencing of target genes (are regions of ) obtained by PCR using gene specific primers. Why do we do
Amplicon Sequencing Dr. Orla O Sullivan SIRG Research Fellow Teagasc What is Amplicon Sequencing? Sequencing of target genes (are regions of ) obtained by PCR using gene specific primers. Why do we do
Using Topological Data Analysis to find discrimination between microbial states in human microbiome data
 Using Topological Data Analysis to find discrimination between microbial states in human microbiome data Mehrdad Yazdani 1,2, Larry Smarr 1,3 and Rob Knight 4 1 California Institute for Telecommunications
Using Topological Data Analysis to find discrimination between microbial states in human microbiome data Mehrdad Yazdani 1,2, Larry Smarr 1,3 and Rob Knight 4 1 California Institute for Telecommunications
Lecture 2: Diversity, Distances, adonis. Lecture 2: Diversity, Distances, adonis. Alpha- Diversity. Alpha diversity definition(s)
 Lecture 2: Diversity, Distances, adonis Lecture 2: Diversity, Distances, adonis Diversity - alpha, beta (, gamma) Beta- Diversity in practice: Ecological Distances Unsupervised Learning: Clustering, etc
Lecture 2: Diversity, Distances, adonis Lecture 2: Diversity, Distances, adonis Diversity - alpha, beta (, gamma) Beta- Diversity in practice: Ecological Distances Unsupervised Learning: Clustering, etc
Outline Classes of diversity measures. Species Divergence and the Measurement of Microbial Diversity. How do we describe and compare diversity?
 Species Divergence and the Measurement of Microbial Diversity Cathy Lozupone University of Colorado, Boulder. Washington University, St Louis. Outline Classes of diversity measures α vs β diversity Quantitative
Species Divergence and the Measurement of Microbial Diversity Cathy Lozupone University of Colorado, Boulder. Washington University, St Louis. Outline Classes of diversity measures α vs β diversity Quantitative
Chad Burrus April 6, 2010
 Chad Burrus April 6, 2010 1 Background What is UniFrac? Materials and Methods Results Discussion Questions 2 The vast majority of microbes cannot be cultured with current methods Only half (26) out of
Chad Burrus April 6, 2010 1 Background What is UniFrac? Materials and Methods Results Discussion Questions 2 The vast majority of microbes cannot be cultured with current methods Only half (26) out of
Microbial analysis with STAMP
 Microbial analysis with STAMP Conor Meehan cmeehan@itg.be A quick aside on who I am Tangents already! Who I am A postdoc at the Institute of Tropical Medicine in Antwerp, Belgium Mycobacteria evolution
Microbial analysis with STAMP Conor Meehan cmeehan@itg.be A quick aside on who I am Tangents already! Who I am A postdoc at the Institute of Tropical Medicine in Antwerp, Belgium Mycobacteria evolution
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/1/e1500997/dc1 Supplementary Materials for Social behavior shapes the chimpanzee pan-microbiome Andrew H. Moeller, Steffen Foerster, Michael L. Wilson, Anne E.
advances.sciencemag.org/cgi/content/full/2/1/e1500997/dc1 Supplementary Materials for Social behavior shapes the chimpanzee pan-microbiome Andrew H. Moeller, Steffen Foerster, Michael L. Wilson, Anne E.
The Effect of Primer Choice and Short Read Sequences on the Outcome of 16S rrna Gene Based Diversity Studies
 The Effect of Primer Choice and Short Read Sequences on the Outcome of 16S rrna Gene Based Diversity Studies Jonas Ghyselinck 1 *., Stefan Pfeiffer 2 *., Kim Heylen 1, Angela Sessitsch 2, Paul De Vos 1
The Effect of Primer Choice and Short Read Sequences on the Outcome of 16S rrna Gene Based Diversity Studies Jonas Ghyselinck 1 *., Stefan Pfeiffer 2 *., Kim Heylen 1, Angela Sessitsch 2, Paul De Vos 1
Microbiota: Its Evolution and Essence. Hsin-Jung Joyce Wu "Microbiota and man: the story about us
 Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Lecture: Mixture Models for Microbiome data
 Lecture: Mixture Models for Microbiome data Lecture 3: Mixture Models for Microbiome data Outline: - - Sequencing thought experiment Mixture Models (tangent) - (esp. Negative Binomial) - Differential abundance
Lecture: Mixture Models for Microbiome data Lecture 3: Mixture Models for Microbiome data Outline: - - Sequencing thought experiment Mixture Models (tangent) - (esp. Negative Binomial) - Differential abundance
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
An Adaptive Association Test for Microbiome Data
 An Adaptive Association Test for Microbiome Data Chong Wu 1, Jun Chen 2, Junghi 1 Kim and Wei Pan 1 1 Division of Biostatistics, School of Public Health, University of Minnesota; 2 Division of Biomedical
An Adaptive Association Test for Microbiome Data Chong Wu 1, Jun Chen 2, Junghi 1 Kim and Wei Pan 1 1 Division of Biostatistics, School of Public Health, University of Minnesota; 2 Division of Biomedical
Lecture 3: Mixture Models for Microbiome data. Lecture 3: Mixture Models for Microbiome data
 Lecture 3: Mixture Models for Microbiome data 1 Lecture 3: Mixture Models for Microbiome data Outline: - Mixture Models (Negative Binomial) - DESeq2 / Don t Rarefy. Ever. 2 Hypothesis Tests - reminder
Lecture 3: Mixture Models for Microbiome data 1 Lecture 3: Mixture Models for Microbiome data Outline: - Mixture Models (Negative Binomial) - DESeq2 / Don t Rarefy. Ever. 2 Hypothesis Tests - reminder
Package LDM. R topics documented: March 19, Type Package
 Type Package Package LDM March 19, 2018 Title Testing Hypotheses about the Microbiome using an Ordination-based Linear Decomposition Model Version 1.0 Date 2018-3-19 Depends GUniFrac, vegan Suggests R.rsp
Type Package Package LDM March 19, 2018 Title Testing Hypotheses about the Microbiome using an Ordination-based Linear Decomposition Model Version 1.0 Date 2018-3-19 Depends GUniFrac, vegan Suggests R.rsp
Phylogenetic trees 07/10/13
 Phylogenetic trees 07/10/13 A tree is the only figure to occur in On the Origin of Species by Charles Darwin. It is a graphical representation of the evolutionary relationships among entities that share
Phylogenetic trees 07/10/13 A tree is the only figure to occur in On the Origin of Species by Charles Darwin. It is a graphical representation of the evolutionary relationships among entities that share
Supplementary Material
 Supplementary Material The impact of logging and forest conversion to oil palm on soil bacterial communities in Borneo Larisa Lee-Cruz 1, David P. Edwards 2,3, Binu Tripathi 1, Jonathan M. Adams 1* 1 Department
Supplementary Material The impact of logging and forest conversion to oil palm on soil bacterial communities in Borneo Larisa Lee-Cruz 1, David P. Edwards 2,3, Binu Tripathi 1, Jonathan M. Adams 1* 1 Department
diversity(datamatrix, index= shannon, base=exp(1))
 Tutorial 11: Diversity, Indicator Species Analysis, Cluster Analysis Calculating Diversity Indices The vegan package contains the command diversity() for calculating Shannon and Simpson diversity indices.
Tutorial 11: Diversity, Indicator Species Analysis, Cluster Analysis Calculating Diversity Indices The vegan package contains the command diversity() for calculating Shannon and Simpson diversity indices.
Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies
 The following supplement accompanies the article Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies Richard L. Hahnke 1, Christina Probian 1, Bernhard M.
The following supplement accompanies the article Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies Richard L. Hahnke 1, Christina Probian 1, Bernhard M.
Probing diversity in a hidden world: applications of NGS in microbial ecology
 Probing diversity in a hidden world: applications of NGS in microbial ecology Guus Roeselers TNO, Microbiology & Systems Biology Group Symposium on Next Generation Sequencing October 21, 2013 Royal Museum
Probing diversity in a hidden world: applications of NGS in microbial ecology Guus Roeselers TNO, Microbiology & Systems Biology Group Symposium on Next Generation Sequencing October 21, 2013 Royal Museum
Assessing and Improving Methods Used in Operational Taxonomic Unit-Based Approaches for 16S rrna Gene Sequence Analysis
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, May 2011, p. 3219 3226 Vol. 77, No. 10 0099-2240/11/$12.00 doi:10.1128/aem.02810-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Assessing
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, May 2011, p. 3219 3226 Vol. 77, No. 10 0099-2240/11/$12.00 doi:10.1128/aem.02810-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Assessing
Carlo Vittorio Cannistraci. Minimum Curvilinear Embedding unveils nonlinear patterns in 16S metagenomic data
 Carlo Vittorio Cannistraci Minimum Curvilinear Embedding unveils nonlinear patterns in 16S metagenomic data Biomedical Cybernetics Group Biotechnology Center (BIOTEC) Technische Universität Dresden (TUD)
Carlo Vittorio Cannistraci Minimum Curvilinear Embedding unveils nonlinear patterns in 16S metagenomic data Biomedical Cybernetics Group Biotechnology Center (BIOTEC) Technische Universität Dresden (TUD)
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Exploring Microbes in the Sea. Alma Parada Postdoctoral Scholar Stanford University
 Exploring Microbes in the Sea Alma Parada Postdoctoral Scholar Stanford University Cruising the ocean to get us some microbes It s all about the Microbe! Microbes = microorganisms an organism that requires
Exploring Microbes in the Sea Alma Parada Postdoctoral Scholar Stanford University Cruising the ocean to get us some microbes It s all about the Microbe! Microbes = microorganisms an organism that requires
An Automated Phylogenetic Tree-Based Small Subunit rrna Taxonomy and Alignment Pipeline (STAP)
 An Automated Phylogenetic Tree-Based Small Subunit rrna Taxonomy and Alignment Pipeline (STAP) Dongying Wu 1 *, Amber Hartman 1,6, Naomi Ward 4,5, Jonathan A. Eisen 1,2,3 1 UC Davis Genome Center, University
An Automated Phylogenetic Tree-Based Small Subunit rrna Taxonomy and Alignment Pipeline (STAP) Dongying Wu 1 *, Amber Hartman 1,6, Naomi Ward 4,5, Jonathan A. Eisen 1,2,3 1 UC Davis Genome Center, University
Bacterial growth efficiency: do consumer and resource diversity influence the fate of carbon in aquatic ecosystems?
 Bacterial growth efficiency: do consumer and resource diversity influence the fate of carbon in aquatic ecosystems? Muscarella M.E. & Lennon J.T. Department of Biology, Indiana University, USA Join the
Bacterial growth efficiency: do consumer and resource diversity influence the fate of carbon in aquatic ecosystems? Muscarella M.E. & Lennon J.T. Department of Biology, Indiana University, USA Join the
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time
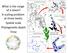 What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Microbial Taxonomy. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Diversity and Assessment (II) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
SUPPLEMENTARY INFORMATION
 City of origin as a confounding variable. The original study was designed such that the city where sampling was performed was perfectly confounded with where the DNA extractions and sequencing was performed.
City of origin as a confounding variable. The original study was designed such that the city where sampling was performed was perfectly confounded with where the DNA extractions and sequencing was performed.
Biological Networks: Comparison, Conservation, and Evolution via Relative Description Length By: Tamir Tuller & Benny Chor
 Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Supplemental Online Results:
 Supplemental Online Results: Functional, phylogenetic, and computational determinants of prediction accuracy using reference genomes A series of tests determined the relationship between PICRUSt s prediction
Supplemental Online Results: Functional, phylogenetic, and computational determinants of prediction accuracy using reference genomes A series of tests determined the relationship between PICRUSt s prediction
Compositional similarity and β (beta) diversity
 CHAPTER 6 Compositional similarity and β (beta) diversity Lou Jost, Anne Chao, and Robin L. Chazdon 6.1 Introduction Spatial variation in species composition is one of the most fundamental and conspicuous
CHAPTER 6 Compositional similarity and β (beta) diversity Lou Jost, Anne Chao, and Robin L. Chazdon 6.1 Introduction Spatial variation in species composition is one of the most fundamental and conspicuous
STATISTICAL LEARNING OF INTEGRATIVE ANALYSIS. Meilei Jiang
 STATISTICAL LEARNING OF INTEGRATIVE ANALYSIS Meilei Jiang A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree
STATISTICAL LEARNING OF INTEGRATIVE ANALYSIS Meilei Jiang A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree
Supplementary Information
 Supplementary Information Table S1. Per-sample sequences, observed OTUs, richness estimates, diversity indices and coverage. Samples codes as follows: YED (Young leaves Endophytes), MED (Mature leaves
Supplementary Information Table S1. Per-sample sequences, observed OTUs, richness estimates, diversity indices and coverage. Samples codes as follows: YED (Young leaves Endophytes), MED (Mature leaves
Multivariate Statistics 101. Ordination (PCA, NMDS, CA) Cluster Analysis (UPGMA, Ward s) Canonical Correspondence Analysis
 Multivariate Statistics 101 Ordination (PCA, NMDS, CA) Cluster Analysis (UPGMA, Ward s) Canonical Correspondence Analysis Multivariate Statistics 101 Copy of slides and exercises PAST software download
Multivariate Statistics 101 Ordination (PCA, NMDS, CA) Cluster Analysis (UPGMA, Ward s) Canonical Correspondence Analysis Multivariate Statistics 101 Copy of slides and exercises PAST software download
Supplementary File 4: Methods Summary and Supplementary Methods
 Supplementary File 4: Methods Summary and Supplementary Methods Methods Summary Constructing and implementing an SDM model requires local measurements of community composition and rasters of environmental
Supplementary File 4: Methods Summary and Supplementary Methods Methods Summary Constructing and implementing an SDM model requires local measurements of community composition and rasters of environmental
Microbial Taxonomy and the Evolution of Diversity
 19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
Lecture 5: Ecological distance metrics; Principal Coordinates Analysis. Univariate testing vs. community analysis
 Lecture 5: Ecological distance metrics; Principal Coordinates Analysis Univariate testing vs. community analysis Univariate testing deals with hypotheses concerning individual taxa Is this taxon differentially
Lecture 5: Ecological distance metrics; Principal Coordinates Analysis Univariate testing vs. community analysis Univariate testing deals with hypotheses concerning individual taxa Is this taxon differentially
Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy. Slowly evolving molecules (e.g., rrna) used for large-scale structure; "fast- clock" molecules for fine-structure.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Lecture 5: Ecological distance metrics; Principal Coordinates Analysis. Univariate testing vs. community analysis
 Lecture 5: Ecological distance metrics; Principal Coordinates Analysis Univariate testing vs. community analysis Univariate testing deals with hypotheses concerning individual taxa Is this taxon differentially
Lecture 5: Ecological distance metrics; Principal Coordinates Analysis Univariate testing vs. community analysis Univariate testing deals with hypotheses concerning individual taxa Is this taxon differentially
Robert Edgar. Independent scientist
 Robert Edgar Independent scientist robert@drive5.com www.drive5.com "Bacterial taxonomy is a hornets nest that no one, really, wants to get into." Referee #1, UTAX paper Assume prokaryotic species meaningful
Robert Edgar Independent scientist robert@drive5.com www.drive5.com "Bacterial taxonomy is a hornets nest that no one, really, wants to get into." Referee #1, UTAX paper Assume prokaryotic species meaningful
Finding Motifs in Protein Sequences and Marking Their Positions in Protein Structures
 MotifMarker Demo This document contains a demo of MotifMarker used to study the structures of the CDK2,CDC2, CDK4 and CDK6 highlighting the 3 important domains/motifs, namely, the PSTAIRE region, ATP binding
MotifMarker Demo This document contains a demo of MotifMarker used to study the structures of the CDK2,CDC2, CDK4 and CDK6 highlighting the 3 important domains/motifs, namely, the PSTAIRE region, ATP binding
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
08/21/2017 BLAST. Multiple Sequence Alignments: Clustal Omega
 BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
PhyloNet. Yun Yu. Department of Computer Science Bioinformatics Group Rice University
 PhyloNet Yun Yu Department of Computer Science Bioinformatics Group Rice University yy9@rice.edu Symposium And Software School 2016 The University Of Texas At Austin Installation System requirement: Java
PhyloNet Yun Yu Department of Computer Science Bioinformatics Group Rice University yy9@rice.edu Symposium And Software School 2016 The University Of Texas At Austin Installation System requirement: Java
Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics.
 Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics. Tree topologies Two topologies were examined, one favoring
Supplementary material to Whitney, K. D., B. Boussau, E. J. Baack, and T. Garland Jr. in press. Drift and genome complexity revisited. PLoS Genetics. Tree topologies Two topologies were examined, one favoring
LDM Package. 1 Overview. Yi-Juan Hu and Glen A. Satten March 19, 2018
 LDM Package Yi-Juan Hu and Glen A. Satten March 19, 2018 1 Overview The LDM package implements the Linear Decomposition Model (Hu and Satten 2018), which provides a single analysis path that includes distance-based
LDM Package Yi-Juan Hu and Glen A. Satten March 19, 2018 1 Overview The LDM package implements the Linear Decomposition Model (Hu and Satten 2018), which provides a single analysis path that includes distance-based
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature25973 Power Simulations We performed extensive power simulations to demonstrate that the analyses carried out in our study are well powered. Our simulations indicate very high power for
doi:10.1038/nature25973 Power Simulations We performed extensive power simulations to demonstrate that the analyses carried out in our study are well powered. Our simulations indicate very high power for
DEPARTMENT OF MICROBIOLOGY. Microbiology Programme: Bachelor of Science (Honours) Microbiology
 DEPARTMENT OF MICROBIOLOGY Microbiology Programme: Bachelor of Science (Honours) Microbiology Philosophy Microbiology is simply a natural science that deals with the study of microbes: their structure,
DEPARTMENT OF MICROBIOLOGY Microbiology Programme: Bachelor of Science (Honours) Microbiology Philosophy Microbiology is simply a natural science that deals with the study of microbes: their structure,
Introduction to microbiota data analysis
 Introduction to microbiota data analysis Natalie Knox, PhD Head Bacterial Genomics, Bioinformatics Core National Microbiology Laboratory, Public Health Agency of Canada 2 National Microbiology Laboratory
Introduction to microbiota data analysis Natalie Knox, PhD Head Bacterial Genomics, Bioinformatics Core National Microbiology Laboratory, Public Health Agency of Canada 2 National Microbiology Laboratory
Taxonomical Classification using:
 Taxonomical Classification using: Extracting ecological signal from noise: introduction to tools for the analysis of NGS data from microbial communities Bergen, April 19-20 2012 INTRODUCTION Taxonomical
Taxonomical Classification using: Extracting ecological signal from noise: introduction to tools for the analysis of NGS data from microbial communities Bergen, April 19-20 2012 INTRODUCTION Taxonomical
Changes in bacterial gut community of Reticulitermes flavipes (Kollar) and Reticulitermes tibialis Banks after feeding on termiticidal bait material
 IRG/WP 14-10819 THE INTERNATIONAL RESEARCH GROUP ON WOOD PROTECTION Section 1 Biology Changes in bacterial gut community of Reticulitermes flavipes (Kollar) and Reticulitermes tibialis Banks after feeding
IRG/WP 14-10819 THE INTERNATIONAL RESEARCH GROUP ON WOOD PROTECTION Section 1 Biology Changes in bacterial gut community of Reticulitermes flavipes (Kollar) and Reticulitermes tibialis Banks after feeding
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
BIO 682 Multivariate Statistics Spring 2008
 BIO 682 Multivariate Statistics Spring 2008 Steve Shuster http://www4.nau.edu/shustercourses/bio682/index.htm Lecture 11 Properties of Community Data Gauch 1982, Causton 1988, Jongman 1995 a. Qualitative:
BIO 682 Multivariate Statistics Spring 2008 Steve Shuster http://www4.nau.edu/shustercourses/bio682/index.htm Lecture 11 Properties of Community Data Gauch 1982, Causton 1988, Jongman 1995 a. Qualitative:
The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome
 Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Tree Building Activity
 Tree Building Activity Introduction In this activity, you will construct phylogenetic trees using a phenotypic similarity (cartoon microbe pictures) and genotypic similarity (real microbe sequences). For
Tree Building Activity Introduction In this activity, you will construct phylogenetic trees using a phenotypic similarity (cartoon microbe pictures) and genotypic similarity (real microbe sequences). For
II. Deep insight into plant habitats
 II. Deep insight into plant habitats Sugar Beet The seed microbiome project (ACIB) Christin Zachow, Henry Müller Ralf Tilcher (KWS SAAT AG) Cultivar-specific microbiomes Experimental design Genetic pool
II. Deep insight into plant habitats Sugar Beet The seed microbiome project (ACIB) Christin Zachow, Henry Müller Ralf Tilcher (KWS SAAT AG) Cultivar-specific microbiomes Experimental design Genetic pool
2/19/2018. Dataset: 85,122 islands 19,392 > 1km 2 17,883 with data
 The group numbers are arbitrary. Remember that you can rotate dendrograms around any node and not change the meaning. So, the order of the clusters is not meaningful. Taking a subset of the data changes
The group numbers are arbitrary. Remember that you can rotate dendrograms around any node and not change the meaning. So, the order of the clusters is not meaningful. Taking a subset of the data changes
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST
 Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
Taxonomy and distribution of phenazine-producing. Pseudomonas spp. in the dryland agroecosystem of the. Inland Pacific Northwest (U. S.
 AEM Accepts, published online ahead of print on 12 April 2013 Appl. Environ. Microbiol. doi:10.1128/aem.03945-12 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 Parejko, et al
AEM Accepts, published online ahead of print on 12 April 2013 Appl. Environ. Microbiol. doi:10.1128/aem.03945-12 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 Parejko, et al
Phylogenetic Tree Generation using Different Scoring Methods
 International Journal of Computer Applications (975 8887) Phylogenetic Tree Generation using Different Scoring Methods Rajbir Singh Associate Prof. & Head Department of IT LLRIET, Moga Sinapreet Kaur Student
International Journal of Computer Applications (975 8887) Phylogenetic Tree Generation using Different Scoring Methods Rajbir Singh Associate Prof. & Head Department of IT LLRIET, Moga Sinapreet Kaur Student
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
Structure, function and host control of rhizosphere microbiome
 Structure, function and host control of rhizosphere microbiome Davide Bulgarelli PhD, Microbiome AgBioTech Europe London, September 21st, 2017 Outline The rhizosphere microbiome Barley as a model to study
Structure, function and host control of rhizosphere microbiome Davide Bulgarelli PhD, Microbiome AgBioTech Europe London, September 21st, 2017 Outline The rhizosphere microbiome Barley as a model to study
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Courses offered by the Biology Department- a 5 year plan
 Courses offered by the Biology Department- a 5 year plan Please note that this is a tentative plan. Changes will be made in years to come, but this list can hopefully serve you well when planning your
Courses offered by the Biology Department- a 5 year plan Please note that this is a tentative plan. Changes will be made in years to come, but this list can hopefully serve you well when planning your
Bem Vindo. Amazonian Biodiversity and Systematics in Brazil.
 Bem Vindo Amazonian Biodiversity and Systematics in Brazil. John W. Wenzel Director, Center for Biodiversity and Ecosystems Carnegie Museum of Natural History Pittsburgh, PA. 1800: Alexander von Humbolt
Bem Vindo Amazonian Biodiversity and Systematics in Brazil. John W. Wenzel Director, Center for Biodiversity and Ecosystems Carnegie Museum of Natural History Pittsburgh, PA. 1800: Alexander von Humbolt
G.S. Gilbert, ENVS291 Transition to R vf13 Class 10 Vegan and Picante. Class 10b - Tools for Phylogenetic Ecology
 Class 10b - Tools for Phylogenetic Ecology This is a very brief introduction to some of the R tools available for phylogenetic community ecology and trait analysis. There are lots of levels to this, with
Class 10b - Tools for Phylogenetic Ecology This is a very brief introduction to some of the R tools available for phylogenetic community ecology and trait analysis. There are lots of levels to this, with
2/7/2018. Strata. Strata
 The strata option allows you to control how permutations are done. Specifically, to constrain permutations. Why would you want to do this? In this dataset, there are clear differences in area (A vs. B),
The strata option allows you to control how permutations are done. Specifically, to constrain permutations. Why would you want to do this? In this dataset, there are clear differences in area (A vs. B),
Phylogenetic diversity and conservation
 Phylogenetic diversity and conservation Dan Faith The Australian Museum Applied ecology and human dimensions in biological conservation Biota Program/ FAPESP Nov. 9-10, 2009 BioGENESIS Providing an evolutionary
Phylogenetic diversity and conservation Dan Faith The Australian Museum Applied ecology and human dimensions in biological conservation Biota Program/ FAPESP Nov. 9-10, 2009 BioGENESIS Providing an evolutionary
Generating phylogenetic trees with Phylomatic and dendrograms of functional traits in R
 Generating phylogenetic trees with Phylomatic and dendrograms of functional traits in R Zhang Jinlong IBCAS 2010-8-1 Contents 1. Introduction 2. Phylomatic: a step by step guide 3. Import phylogentic trees
Generating phylogenetic trees with Phylomatic and dendrograms of functional traits in R Zhang Jinlong IBCAS 2010-8-1 Contents 1. Introduction 2. Phylomatic: a step by step guide 3. Import phylogentic trees
Session 5: Phylogenomics
 Session 5: Phylogenomics B.- Phylogeny based orthology assignment REMINDER: Gene tree reconstruction is divided in three steps: homology search, multiple sequence alignment and model selection plus tree
Session 5: Phylogenomics B.- Phylogeny based orthology assignment REMINDER: Gene tree reconstruction is divided in three steps: homology search, multiple sequence alignment and model selection plus tree
Deciphering the Enigma of Undetected Species, Phylogenetic, and Functional Diversity. Based on Good-Turing Theory
 Metadata S1 Deciphering the Enigma of Undetected Species, Phylogenetic, and Functional Diversity Based on Good-Turing Theory Anne Chao, Chun-Huo Chiu, Robert K. Colwell, Luiz Fernando S. Magnago, Robin
Metadata S1 Deciphering the Enigma of Undetected Species, Phylogenetic, and Functional Diversity Based on Good-Turing Theory Anne Chao, Chun-Huo Chiu, Robert K. Colwell, Luiz Fernando S. Magnago, Robin
Characterizing and predicting cyanobacterial blooms in an 8-year
 1 2 3 4 5 Characterizing and predicting cyanobacterial blooms in an 8-year amplicon sequencing time-course Authors Nicolas Tromas 1*, Nathalie Fortin 2, Larbi Bedrani 1, Yves Terrat 1, Pedro Cardoso 4,
1 2 3 4 5 Characterizing and predicting cyanobacterial blooms in an 8-year amplicon sequencing time-course Authors Nicolas Tromas 1*, Nathalie Fortin 2, Larbi Bedrani 1, Yves Terrat 1, Pedro Cardoso 4,
Using Machine Learning to Understand Top-Down Effects in an Ecosystem: Opportunities, Challenges, and Lessons Learned
 Proceedings of the Thirtieth International Florida Artificial Intelligence Research Society Conference Using Machine Learning to Understand Top-Down Effects in an Ecosystem: Opportunities, Challenges,
Proceedings of the Thirtieth International Florida Artificial Intelligence Research Society Conference Using Machine Learning to Understand Top-Down Effects in an Ecosystem: Opportunities, Challenges,
Courses offered by the Biology Department- a 5 year plan
 Courses offered by the Biology Department- a 5 year plan Please note that this is a tentative plan. Changes will be made in years to come, but this list can hopefully serve you well when planning your
Courses offered by the Biology Department- a 5 year plan Please note that this is a tentative plan. Changes will be made in years to come, but this list can hopefully serve you well when planning your
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Microbiology BIOL 202 Lecture Course Outcome Guide (COG) Approved 22 MARCH 2012 Pg.1
 Microbiology BIOL 202 Lecture Course Outcome Guide (COG) Approved 22 MARCH 2012 Pg.1 Course: Credits: 3 Instructor: Course Description: Concepts and Issues 1. Microbial Ecology including mineral cycles.
Microbiology BIOL 202 Lecture Course Outcome Guide (COG) Approved 22 MARCH 2012 Pg.1 Course: Credits: 3 Instructor: Course Description: Concepts and Issues 1. Microbial Ecology including mineral cycles.
Measurement and Data
 Measurement and Data Data describes the real world Data maps entities in the domain of interest to symbolic representation by means of a measurement procedure Numerical relationships between variables
Measurement and Data Data describes the real world Data maps entities in the domain of interest to symbolic representation by means of a measurement procedure Numerical relationships between variables
Detecting bacterial associations in the human microbiome
 Karoline Faust Raes Lab (Bioinformatics and (Eco-)Systems Biology) 10 th October 2012 Bertinoro Computational Biology Detecting bacterial associations in the human microbiome 1. Introduction Examples for
Karoline Faust Raes Lab (Bioinformatics and (Eco-)Systems Biology) 10 th October 2012 Bertinoro Computational Biology Detecting bacterial associations in the human microbiome 1. Introduction Examples for
Stepping stones towards a new electronic prokaryotic taxonomy. The ultimate goal in taxonomy. Pragmatic towards diagnostics
 Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
arxiv: v1 [q-bio.pe] 7 Jul 2014
![arxiv: v1 [q-bio.pe] 7 Jul 2014 arxiv: v1 [q-bio.pe] 7 Jul 2014](/thumbs/83/88400281.jpg) PHYLOGENETICS AND THE HUMAN MICROBIOME. FREDERICK A. MATSEN IV arxiv:1407.1794v1 [q-bio.pe] 7 Jul 2014 ABSTRACT. The human microbiome is the ensemble of genes in the microbes that live inside and on the
PHYLOGENETICS AND THE HUMAN MICROBIOME. FREDERICK A. MATSEN IV arxiv:1407.1794v1 [q-bio.pe] 7 Jul 2014 ABSTRACT. The human microbiome is the ensemble of genes in the microbes that live inside and on the
Supplementary Information
 Supplementary Information Altitudinal patterns of diversity and functional traits of metabolically active microorganisms in stream biofilms Linda Wilhelm 1, Katharina Besemer 2, Lena Fragner 3, Hannes
Supplementary Information Altitudinal patterns of diversity and functional traits of metabolically active microorganisms in stream biofilms Linda Wilhelm 1, Katharina Besemer 2, Lena Fragner 3, Hannes
Comparing Prokaryotic and Eukaryotic Cells
 A prokaryotic cell Basic unit of living organisms is the cell; the smallest unit capable of life. Features found in all cells: Ribosomes Cell Membrane Genetic Material Cytoplasm ATP Energy External Stimuli
A prokaryotic cell Basic unit of living organisms is the cell; the smallest unit capable of life. Features found in all cells: Ribosomes Cell Membrane Genetic Material Cytoplasm ATP Energy External Stimuli
Istituto di Microbiologia. Università Cattolica del Sacro Cuore, Roma. Gut Microbiota assessment and the Meta-HIT program.
 Istituto di Microbiologia Università Cattolica del Sacro Cuore, Roma Gut Microbiota assessment and the Meta-HIT program Giovanni Delogu 1 Most of the bacteria species living in the gut cannot be cultivated
Istituto di Microbiologia Università Cattolica del Sacro Cuore, Roma Gut Microbiota assessment and the Meta-HIT program Giovanni Delogu 1 Most of the bacteria species living in the gut cannot be cultivated
Correlation Analysis of Binary Similarity and Distance Measures on Different Binary Database Types
 Correlation Analysis of Binary Similarity and Distance Measures on Different Binary Database Types Seung-Seok Choi, Sung-Hyuk Cha, Charles C. Tappert Department of Computer Science, Pace University, New
Correlation Analysis of Binary Similarity and Distance Measures on Different Binary Database Types Seung-Seok Choi, Sung-Hyuk Cha, Charles C. Tappert Department of Computer Science, Pace University, New
Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction. Lesser Tenrec (Echinops telfairi)
 Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction Lesser Tenrec (Echinops telfairi) Goals: 1. Use phylogenetic experimental design theory to select optimal taxa to
Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction Lesser Tenrec (Echinops telfairi) Goals: 1. Use phylogenetic experimental design theory to select optimal taxa to
Using Bioinformatics to Study Evolutionary Relationships Instructions
 3 Using Bioinformatics to Study Evolutionary Relationships Instructions Student Researcher Background: Making and Using Multiple Sequence Alignments One of the primary tasks of genetic researchers is comparing
3 Using Bioinformatics to Study Evolutionary Relationships Instructions Student Researcher Background: Making and Using Multiple Sequence Alignments One of the primary tasks of genetic researchers is comparing
Assigning Taxonomy to Marker Genes. Susan Huse Brown University August 7, 2014
 Assigning Taxonomy to Marker Genes Susan Huse Brown University August 7, 2014 In a nutshell Taxonomy is assigned by comparing your DNA sequences against a database of DNA sequences from known taxa Marker
Assigning Taxonomy to Marker Genes Susan Huse Brown University August 7, 2014 In a nutshell Taxonomy is assigned by comparing your DNA sequences against a database of DNA sequences from known taxa Marker
University of Florida CISE department Gator Engineering. Clustering Part 1
 Clustering Part 1 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville What is Cluster Analysis? Finding groups of objects such that the objects
Clustering Part 1 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville What is Cluster Analysis? Finding groups of objects such that the objects
Microbial Diversity. Yuzhen Ye I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University
 Microbial Diversity Yuzhen Ye (yye@indiana.edu) I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University Contents Microbial diversity Morphological, structural,
Microbial Diversity Yuzhen Ye (yye@indiana.edu) I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University Contents Microbial diversity Morphological, structural,
Let s get started. So, what is science?
 Let s get started So, what is science? Well Science Science is the observation of phenomena and the theoretical explanation of it. Simply, it is the state of knowing. Biology Biology is the study of life.
Let s get started So, what is science? Well Science Science is the observation of phenomena and the theoretical explanation of it. Simply, it is the state of knowing. Biology Biology is the study of life.
