Alternative tools for phylogeny. Identification of unique core sequences
|
|
|
- Ezra McKenzie
- 5 years ago
- Views:
Transcription
1 Alternative tools for phylogeny Identification of unique core sequences Workshop on Whole Genome Sequencing and Analysis, Mar. 2018
2 Learning objective: After this lecture you should be able to account for non-snp based methods for deducing phylogeny account for a method for identification of unique core sequences in a set of strains with a particular phenotype
3 Alternative tools for phylogeny
4 NCBI s Pathogen Detection A centralised system integrating data from bacterial pathogens (food, environment, humans) Public health agencies in US and internationally submit sequence data to NCBI NCBI compare the sequences to those already in database and generates phylogenetic tree based on kmers ( Details of the analysis system will be published at a future date ) The aim is to aid traceback investigations and outbreak response Submitted data are released to the public immediately
5 Database Content (13th of Feb. 2018)......
6
7 Morganella tree
8 Ribosomal MLST (rmlst) Jolly et al Microbiology. 158(pt 4):
9 (16s rrna)
10 core genome MLST (cgmlst) and whole genome MLST (wgmlst) Examining thousands of genes => higher discriminatory power Prefixed set of conserved genomewide genes for each species Potential for standardisation and a name Nisseria meningitides MLST loci versus cgmlst loci. Enterobase (Warwick Medical School): cgmlst and wgmlst for Salmonella, Escherichia/Shigella, and Yersinia CLCBio, BioNumerics, Ridom SeqSphere+ (commercial): offer cgmlst/ wgmlst for a number of species CGE: in the process of implementing cgmlst for Campylobacter, E. coli, Listeria, Salmonella, and Yersinia
11 Campylobacter scheme is from pubmlst Listeria scheme is from Pasteur Institute Salmonella, E. coli, and Yersinia are schemes from Enterobase
12 cgmlstfinder is currently in betamode Only accepts raw reads (fastq files and NOT zipped) For some schemes (e.g., Listeria) it is currently only possible to have alleles identified, no final translation to Sequence Type
13 SNP/ND based methods versus cgmlst/wgmlst Pros Cons SNP/ND based phylogeny Available for all species Free web-services are available (e.g., CGE, NCBI s pipeline) Computationally heavy (slow), if many isolates are included No standardisation, e.g., result may differ if a different reference strain is used No nomenclature cgmlst/ wgmlst Offers standardisation and a name Curated databases for each species must first be made and agreed upon Currently only free solutions for a few species
14 Increasing discriminatory power Analysis Number of genes sampled Example of nomenclature Discriminatory power SNP/ND everything, also - intergenic regions wgmlst >3000 wst543 cgmlst cst832 rmlst 53 rst512 MLST 5-10 ST698 16S rrna 1 E. coli
15 Distinguishing E. coli ST-131 outbreak strains from non-outbreak strains 16 isolates from a nosocomial outbreak of ESBL E. coli ST-131 in Sweden was subjected to WGS (Illumina) after PFGE and LMqPCR HRMA were unable to judge if two isolates (no. 8 and 13) were part of outbreak 3 E. coli control strain were also sequenced (no. 17, 18, and 21) The CGE tools MLST, ResFinder, and PlasmidFinder were used to identify the multilocus sequence type and look for antibiotic resistance genes, and plasmid replicons ST aac(60)ib-cr blaoxa blactx-m blatem-1b IncF II Inc FIA Inc FIB Woksepp et al APMIS 125:
16 cgmlst (SeqSphere) was performed on all strains + reference strain JJ1886, which is also ST-131, but not part of the outbreak. cgmlst could confirm that strains 8 and 13 are not part of outbreak
17 And SNP-based phylogeny (CSIPhylogeny) agree
18 Author discussion cgmlst and SNP-based phylogeny were both able to distinguish outbreak from non-outbreak strains cgmlst has the advantage of making communicating typing results between laboratories possible There is still a need to establish cgmlst as a tool in clinical practice and standardization of cgmlst is urgently needed Although resulting in the same clustering in this case, reference-based SNP analysis of highly diverse organism like E. coli might result in false-negative clustering, but may be favourable for, e.g., M. tuberculosis with more clonal evolution In any case SNP-based phylogeny may not be comparable between laboratories or over time (depend heavily on methodology, e.g., reference)
19 Unique core sequences in genomes with specific phenotype Phenotype+ Phenotype+ Phenotype+ Phenotype- Phenotype- Phenotype-
20 RUCS: Rapid Identification of PCR Primers Pairs for Unique Core Sequences Martin Christen Frølund Thomsen, Henrik Hasman, Henrik Westh, Hülya Kaya, Ole Lund; RUCS: rapid identification of PCR primers for unique core sequences, Bioinformatics, 2017, 33(24):
21
22 Prepare the data for RUCS: Create a directory/folder called positives and add all positive draft genomes in fasta format (not zipped) Create a directory/folder called negatives" and add all negative draft genomes in fasta format (not zipped) Select both folders and create a zip file (on Mac you right click and select Compress 2 items ) The zipped file is what is uploaded as Isolate File Output example: Unique core sequences (sequences found in all the positive genomes and never in the negative genomes)
23 If you run RUCS in full mode, it will even design PCR primers for the unique sequences
24 Summary For some species cgmlst and wgmlst are alternatives to SNP/ND based methods and offer nomenclature The RUCS method identifies unique core sequences in strains with common phenotype.
Other tools for typing and phylogeny
 Other tools for typing and phylogeny Workshop on Whole Genome Sequencing and Analysis, 27-29 Mar. 2017 Learning objective: After this lecture you should be able to account for tools for typing Salmonella
Other tools for typing and phylogeny Workshop on Whole Genome Sequencing and Analysis, 27-29 Mar. 2017 Learning objective: After this lecture you should be able to account for tools for typing Salmonella
Whole genome sequencing (WGS) as a tool for monitoring purposes. Henrik Hasman DTU - Food
 Whole genome sequencing (WGS) as a tool for monitoring purposes Henrik Hasman DTU - Food The Challenge Is to: Continue to increase the power of surveillance and diagnostic using molecular tools Develop
Whole genome sequencing (WGS) as a tool for monitoring purposes Henrik Hasman DTU - Food The Challenge Is to: Continue to increase the power of surveillance and diagnostic using molecular tools Develop
Whole genome sequencing (WGS) - there s a new tool in town. Henrik Hasman DTU - Food
 Whole genome sequencing (WGS) - there s a new tool in town Henrik Hasman DTU - Food Welcome to the NGS world TODAY Welcome Introduction to Next Generation Sequencing DNA purification (Hands-on) Lunch (Sandwishes
Whole genome sequencing (WGS) - there s a new tool in town Henrik Hasman DTU - Food Welcome to the NGS world TODAY Welcome Introduction to Next Generation Sequencing DNA purification (Hands-on) Lunch (Sandwishes
GenomeTrakr: Data Submission and Analysis
 GenomeTrakr: Data Submission and Analysis Ruth Timme and Hugh Rand Center for Food Safety and Applied Nutrition U.S. Food Drug Administration IFSH Whole Genome Sequencing for Food Safety Symposium SEPTEMBER
GenomeTrakr: Data Submission and Analysis Ruth Timme and Hugh Rand Center for Food Safety and Applied Nutrition U.S. Food Drug Administration IFSH Whole Genome Sequencing for Food Safety Symposium SEPTEMBER
Introduction to the SNP/ND concept - Phylogeny on WGS data
 Introduction to the SNP/ND concept - Phylogeny on WGS data Johanne Ahrenfeldt PhD student Overview What is Phylogeny and what can it be used for Single Nucleotide Polymorphism (SNP) methods CSI Phylogeny
Introduction to the SNP/ND concept - Phylogeny on WGS data Johanne Ahrenfeldt PhD student Overview What is Phylogeny and what can it be used for Single Nucleotide Polymorphism (SNP) methods CSI Phylogeny
NRL-Salmonella, Hungary. National Food Chain Safety Office Food and Feed Safety Directorate Erzsébet Adrián 29 May 2018
 NRL-Salmonella, Hungary National Food Chain Safety Office Food and Feed Safety Directorate Erzsébet Adrián 29 May 2018 Structure National Food Chain Safety Office Food and Feed Safety Directorate Official
NRL-Salmonella, Hungary National Food Chain Safety Office Food and Feed Safety Directorate Erzsébet Adrián 29 May 2018 Structure National Food Chain Safety Office Food and Feed Safety Directorate Official
Whole Genome based Phylogeny
 Whole Genome based Phylogeny Johanne Ahrenfeldt PhD student DTU Bioinformatics Short about me Johanne Ahrenfeldt johah@dtu.dk PhD student at DTU Bioinformatics Whole Genome based Phylogeny Graduate Engineer
Whole Genome based Phylogeny Johanne Ahrenfeldt PhD student DTU Bioinformatics Short about me Johanne Ahrenfeldt johah@dtu.dk PhD student at DTU Bioinformatics Whole Genome based Phylogeny Graduate Engineer
By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food Supervised by Henrik Hasman, PhD
 By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food elibi@food.dtu.dk Supervised by Henrik Hasman, PhD 1. Introduction to plasmid biology 2. Plasmid encoded resistance to β- lactams (basic theories)
By Eliza Bielak Bacterial Genomics and Epidemiology, DTU-Food elibi@food.dtu.dk Supervised by Henrik Hasman, PhD 1. Introduction to plasmid biology 2. Plasmid encoded resistance to β- lactams (basic theories)
CRISPR-SeroSeq: A Developing Technique for Salmonella Subtyping
 Department of Biological Sciences Seminar Blog Seminar Date: 3/23/18 Speaker: Dr. Nikki Shariat, Gettysburg College Title: Probing Salmonella population diversity using CRISPRs CRISPR-SeroSeq: A Developing
Department of Biological Sciences Seminar Blog Seminar Date: 3/23/18 Speaker: Dr. Nikki Shariat, Gettysburg College Title: Probing Salmonella population diversity using CRISPRs CRISPR-SeroSeq: A Developing
Comparative Genomics Background & Strategy. Faction 2
 Comparative Genomics Background & Strategy Faction 2 Overview Introduction to comparative genomics Salmonella enterica subsp. enterica serovar Heidelberg Comparative Genomics Faction 2 Objectives Genomic
Comparative Genomics Background & Strategy Faction 2 Overview Introduction to comparative genomics Salmonella enterica subsp. enterica serovar Heidelberg Comparative Genomics Faction 2 Objectives Genomic
Stepping stones towards a new electronic prokaryotic taxonomy. The ultimate goal in taxonomy. Pragmatic towards diagnostics
 Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
Stepping stones towards a new electronic prokaryotic taxonomy - MLSA - Dirk Gevers Different needs for taxonomy Describe bio-diversity Understand evolution of life Epidemiology Diagnostics Biosafety...
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Microbial Bioinformatics
 1 Microbial Bioinformatics Introduction These exercises are for you to learn how to use bioinformatics tools to explore bacterial genomes, and complements the Microbiology of Safe Food 2 nd Edition (Wiley-
1 Microbial Bioinformatics Introduction These exercises are for you to learn how to use bioinformatics tools to explore bacterial genomes, and complements the Microbiology of Safe Food 2 nd Edition (Wiley-
The emergence of a new phage type of Salmonella Typhimurium in humans and animals in New Zealand
 Introduction The emergence of a new phage type of Salmonella Typhimurium in humans and animals in New Zealand M Dufour AIMS NZIMLS South Pacific Congress Gold Coast, August 2011 New Zealand is a geographically
Introduction The emergence of a new phage type of Salmonella Typhimurium in humans and animals in New Zealand M Dufour AIMS NZIMLS South Pacific Congress Gold Coast, August 2011 New Zealand is a geographically
Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy. Slowly evolving molecules (e.g., rrna) used for large-scale structure; "fast- clock" molecules for fine-structure.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Outbreak of a new serotype Salmonella enterica subsp. enterica, with antigenic formula 11:z 41 : e,n,z 15 in Greece :
 Outbreak of a new serotype Salmonella enterica subsp. enterica, with antigenic formula 11:z 41 : e,n,z 15 in Greece : 2016-2017 An investigation of the Hellenic Centre of Disease Control and Prevention
Outbreak of a new serotype Salmonella enterica subsp. enterica, with antigenic formula 11:z 41 : e,n,z 15 in Greece : 2016-2017 An investigation of the Hellenic Centre of Disease Control and Prevention
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Introduction to Bioinformatics. Shifra Ben-Dor Irit Orr
 Introduction to Bioinformatics Shifra Ben-Dor Irit Orr Lecture Outline: Technical Course Items Introduction to Bioinformatics Introduction to Databases This week and next week What is bioinformatics? A
Introduction to Bioinformatics Shifra Ben-Dor Irit Orr Lecture Outline: Technical Course Items Introduction to Bioinformatics Introduction to Databases This week and next week What is bioinformatics? A
MICROBIAL BIOCHEMISTRY BIOT 309. Dr. Leslye Johnson Sept. 30, 2012
 MICROBIAL BIOCHEMISTRY BIOT 309 Dr. Leslye Johnson Sept. 30, 2012 Phylogeny study of evoluhonary relatedness among groups of organisms (e.g. species, populahons), which is discovered through molecular
MICROBIAL BIOCHEMISTRY BIOT 309 Dr. Leslye Johnson Sept. 30, 2012 Phylogeny study of evoluhonary relatedness among groups of organisms (e.g. species, populahons), which is discovered through molecular
Microbiome: 16S rrna Sequencing 3/30/2018
 Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
CREATING PHYLOGENETIC TREES FROM DNA SEQUENCES
 INTRODUCTION CREATING PHYLOGENETIC TREES FROM DNA SEQUENCES This worksheet complements the Click and Learn developed in conjunction with the 2011 Holiday Lectures on Science, Bones, Stones, and Genes:
INTRODUCTION CREATING PHYLOGENETIC TREES FROM DNA SEQUENCES This worksheet complements the Click and Learn developed in conjunction with the 2011 Holiday Lectures on Science, Bones, Stones, and Genes:
Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
Received 11 February 2010/Accepted 6 July 2010
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Sept. 2010, p. 5947 5959 Vol. 76, No. 17 0099-2240/10/$12.00 doi:10.1128/aem.00377-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. The
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Sept. 2010, p. 5947 5959 Vol. 76, No. 17 0099-2240/10/$12.00 doi:10.1128/aem.00377-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. The
Bioinformatics Exercises
 Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Fitness constraints on horizontal gene transfer
 Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
Fitness constraints on horizontal gene transfer Dan I Andersson University of Uppsala, Department of Medical Biochemistry and Microbiology, Uppsala, Sweden GMM 3, 30 Aug--2 Sep, Oslo, Norway Acknowledgements:
The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome. The minimal prokaryotic genome
 Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Dr. Dirk Gevers 1,2 1 Laboratorium voor Microbiologie 2 Bioinformatics & Evolutionary Genomics The bacterial species in the genomic era CTACCATGAAAGACTTGTGAATCCAGGAAGAGAGACTGACTGGGCAACATGTTATTCAG GTACAAAAAGATTTGGACTGTAACTTAAAAATGATCAAATTATGTTTCCCATGCATCAGG
Chapter 19. Microbial Taxonomy
 Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Microbial Diversity. Yuzhen Ye I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University
 Microbial Diversity Yuzhen Ye (yye@indiana.edu) I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University Contents Microbial diversity Morphological, structural,
Microbial Diversity Yuzhen Ye (yye@indiana.edu) I609 Bioinformatics Seminar I (Spring 2010) School of Informatics and Computing Indiana University Contents Microbial diversity Morphological, structural,
Bio 119 Bacterial Genomics 6/26/10
 BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
Bioinformatics. Dept. of Computational Biology & Bioinformatics
 Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Detailed overview of the primer-free full-length SSU rrna library preparation.
 Supplementary Figure 1 Detailed overview of the primer-free full-length SSU rrna library preparation. Detailed overview of the primer-free full-length SSU rrna library preparation. Supplementary Figure
Supplementary Figure 1 Detailed overview of the primer-free full-length SSU rrna library preparation. Detailed overview of the primer-free full-length SSU rrna library preparation. Supplementary Figure
Science Unit Learning Summary
 Learning Summary Inheritance, variation and evolution Content Sexual and asexual reproduction. Meiosis leads to non-identical cells being formed while mitosis leads to identical cells being formed. In
Learning Summary Inheritance, variation and evolution Content Sexual and asexual reproduction. Meiosis leads to non-identical cells being formed while mitosis leads to identical cells being formed. In
Essentiality in B. subtilis
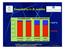 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
11/5/2018. Update on Modern Bacterial Taxonomy for Bench Microbiologists. Why is Taxonomy Important? Bacterial Taxonomy for Clinical Microbiologists
 Update on Modern Bacterial Taxonomy for Bench Microbiologists J. Michael Janda Kern County Public Health Laboratory Bakersfield CA The Name Game Which Ones Different? Why is Taxonomy Important? Bacterial
Update on Modern Bacterial Taxonomy for Bench Microbiologists J. Michael Janda Kern County Public Health Laboratory Bakersfield CA The Name Game Which Ones Different? Why is Taxonomy Important? Bacterial
Microbial Diversity and Assessment (II) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Evaluation of a Stenotrophomonas maltophilia bacteremia cluster in hematopoietic stem cell transplantation recipients using whole genome sequencing
 Kampmeier et al. Antimicrobial Resistance and Infection Control (2017) 6:115 DOI 10.1186/s13756-017-0276-y RESEARCH Evaluation of a Stenotrophomonas maltophilia bacteremia cluster in hematopoietic stem
Kampmeier et al. Antimicrobial Resistance and Infection Control (2017) 6:115 DOI 10.1186/s13756-017-0276-y RESEARCH Evaluation of a Stenotrophomonas maltophilia bacteremia cluster in hematopoietic stem
Introduction to polyphasic taxonomy
 Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Introduction to polyphasic taxonomy Peter Vandamme EUROBILOFILMS - Third European Congress on Microbial Biofilms Ghent, Belgium, 9-12 September 2013 http://www.lm.ugent.be/ Content The observation of diversity:
Evidence That Mutation Is Universally Biased towards AT in Bacteria
 Evidence That Mutation Is Universally Biased towards AT in Bacteria Ruth Hershberg*, Dmitri A. Petrov Department of Biology, Stanford University, Stanford, California, United States of America Abstract
Evidence That Mutation Is Universally Biased towards AT in Bacteria Ruth Hershberg*, Dmitri A. Petrov Department of Biology, Stanford University, Stanford, California, United States of America Abstract
MLVA Update. Eija Trees, Ph.D., D.V.M. PulseNet Methods Development and Reference Unit Enteric Diseases Laboratory Branch CDC, Atlanta, GA
 MLVA Update Eija Trees, Ph.D., D.V.M. PulseNet Methods Development and Reference Unit Enteric Diseases Laboratory Branch CDC, Atlanta, GA Presentation outline Status of the S. Enteritidis MLVA protocol
MLVA Update Eija Trees, Ph.D., D.V.M. PulseNet Methods Development and Reference Unit Enteric Diseases Laboratory Branch CDC, Atlanta, GA Presentation outline Status of the S. Enteritidis MLVA protocol
Genomic Comparison of Bacterial Species Based on Metabolic Characteristics
 Genomic Comparison of Bacterial Species Based on Metabolic Characteristics Gaurav Jain 1, Haozhu Wang 1, Li Liao 1*, E. Fidelma Boyd 2* 1 Department of Computer and Information Sciences University of Delaware
Genomic Comparison of Bacterial Species Based on Metabolic Characteristics Gaurav Jain 1, Haozhu Wang 1, Li Liao 1*, E. Fidelma Boyd 2* 1 Department of Computer and Information Sciences University of Delaware
Genomics links sporadic Salmonella Typhimurium Phage type 9 infections to a recurrent source of outbreak salmonellosis
 Genomics links sporadic Salmonella Typhimurium Phage type 9 infections to a recurrent source of outbreak salmonellosis Zoe Cutcher 1,2, Dieter Bulach 3, Glen Carter 3, Timothy P Stinear 3, Karolina Mercoulia
Genomics links sporadic Salmonella Typhimurium Phage type 9 infections to a recurrent source of outbreak salmonellosis Zoe Cutcher 1,2, Dieter Bulach 3, Glen Carter 3, Timothy P Stinear 3, Karolina Mercoulia
belonging to the Genus Pantoea
 Emerging diseases of maize and onion caused by bacteria belonging to the Genus Pantoea by Teresa Goszczynska Submitted in partial fulfilment of the requirements for the degree Philosophiae Doctoriae in
Emerging diseases of maize and onion caused by bacteria belonging to the Genus Pantoea by Teresa Goszczynska Submitted in partial fulfilment of the requirements for the degree Philosophiae Doctoriae in
Algorithms in Computational Biology (236522) spring 2008 Lecture #1
 Algorithms in Computational Biology (236522) spring 2008 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: 15:30-16:30/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office hours:??
Algorithms in Computational Biology (236522) spring 2008 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: 15:30-16:30/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office hours:??
Satheesh Nair Salmonella Reference Service, GBRU,PHE Colindale. April 2017
 Whole genome sequencing for routine identification, drug resistance detection and epidemiology of Salmonella: A revolution in public health microbiology April 2017 Satheesh Nair Salmonella Reference Service,
Whole genome sequencing for routine identification, drug resistance detection and epidemiology of Salmonella: A revolution in public health microbiology April 2017 Satheesh Nair Salmonella Reference Service,
Introduction to microbiota data analysis
 Introduction to microbiota data analysis Natalie Knox, PhD Head Bacterial Genomics, Bioinformatics Core National Microbiology Laboratory, Public Health Agency of Canada 2 National Microbiology Laboratory
Introduction to microbiota data analysis Natalie Knox, PhD Head Bacterial Genomics, Bioinformatics Core National Microbiology Laboratory, Public Health Agency of Canada 2 National Microbiology Laboratory
Exploring Microbes in the Sea. Alma Parada Postdoctoral Scholar Stanford University
 Exploring Microbes in the Sea Alma Parada Postdoctoral Scholar Stanford University Cruising the ocean to get us some microbes It s all about the Microbe! Microbes = microorganisms an organism that requires
Exploring Microbes in the Sea Alma Parada Postdoctoral Scholar Stanford University Cruising the ocean to get us some microbes It s all about the Microbe! Microbes = microorganisms an organism that requires
RGP finder: prediction of Genomic Islands
 Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
Multivariate analysis of genetic data: an introduction
 Multivariate analysis of genetic data: an introduction Thibaut Jombart MRC Centre for Outbreak Analysis and Modelling Imperial College London XXIV Simposio Internacional De Estadística Bogotá, 25th July
Multivariate analysis of genetic data: an introduction Thibaut Jombart MRC Centre for Outbreak Analysis and Modelling Imperial College London XXIV Simposio Internacional De Estadística Bogotá, 25th July
Microbial Taxonomy. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Handbook of New Bacterial Systematics
 Handbook of New Bacterial Systematics Edited by M. GOODFELLOW Department of Microbiology, The Medical School, Framlington Place, Newcastle upon Tyne, UK and A. G. O'DONNELL Department df Agricultural and
Handbook of New Bacterial Systematics Edited by M. GOODFELLOW Department of Microbiology, The Medical School, Framlington Place, Newcastle upon Tyne, UK and A. G. O'DONNELL Department df Agricultural and
Session 5: Phylogenomics
 Session 5: Phylogenomics B.- Phylogeny based orthology assignment REMINDER: Gene tree reconstruction is divided in three steps: homology search, multiple sequence alignment and model selection plus tree
Session 5: Phylogenomics B.- Phylogeny based orthology assignment REMINDER: Gene tree reconstruction is divided in three steps: homology search, multiple sequence alignment and model selection plus tree
Expansion of Salmonella Typhimurium ST34 clone carrying multiple. resistance determinants in China
 AAC Accepts, published online ahead of print on 24 June 2013 Antimicrob. Agents Chemother. doi:10.1128/aac.01174-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 2 Expansion
AAC Accepts, published online ahead of print on 24 June 2013 Antimicrob. Agents Chemother. doi:10.1128/aac.01174-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 2 Expansion
Comparison of 61 E. coli genomes
 Comparison of 61 E. coli genomes Center for Biological Sequence Analysis Department of Systems Biology Dave Ussery! DTU course 27105 - Comparative Genomics Oksana s 61 E. coli genomes paper! Monday, 23
Comparison of 61 E. coli genomes Center for Biological Sequence Analysis Department of Systems Biology Dave Ussery! DTU course 27105 - Comparative Genomics Oksana s 61 E. coli genomes paper! Monday, 23
Bergey s Manual Classification Scheme. Vertical inheritance and evolutionary mechanisms
 Bergey s Manual Classification Scheme Gram + Gram - No wall Funny wall Vertical inheritance and evolutionary mechanisms a b c d e * * a b c d e * a b c d e a b c d e * a b c d e Accumulation of neutral
Bergey s Manual Classification Scheme Gram + Gram - No wall Funny wall Vertical inheritance and evolutionary mechanisms a b c d e * * a b c d e * a b c d e a b c d e * a b c d e Accumulation of neutral
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
FIG S1: Rarefaction analysis of observed richness within Drosophila. All calculations were
 Page 1 of 14 FIG S1: Rarefaction analysis of observed richness within Drosophila. All calculations were performed using mothur (2). OTUs were defined at the 3% divergence threshold using the average neighbor
Page 1 of 14 FIG S1: Rarefaction analysis of observed richness within Drosophila. All calculations were performed using mothur (2). OTUs were defined at the 3% divergence threshold using the average neighbor
Curriculum Vitae. Farzaneh Firoozeh Assistant Professor of Microbiology
 Curriculum Vitae Farzaneh Firoozeh Assistant Professor of Microbiology PERSONAL First name: Farzaneh Family name: Firoozeh Nationality: Iranian Marital status: Married OFFICE ADDRESS Department of Microbiology
Curriculum Vitae Farzaneh Firoozeh Assistant Professor of Microbiology PERSONAL First name: Farzaneh Family name: Firoozeh Nationality: Iranian Marital status: Married OFFICE ADDRESS Department of Microbiology
PlasmidFinder and pmlst: in silico detection and typing of
 AAC Accepts, published online ahead of print on 28 April 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02412-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 PlasmidFinder
AAC Accepts, published online ahead of print on 28 April 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02412-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 PlasmidFinder
NEWSLETTER. European Union Reference Laboratory for Salmonella. Vol. 24 No. 2 June 2018 ISSN
 NEWSLETTER European Union Reference Laboratory for Salmonella Vol. 24 No. 2 June 2018 ISSN 2211-6877 Continuation of Newsletter Community Reference Laboratory for Salmonella ISSN 1572-3836 Page 2 of 63
NEWSLETTER European Union Reference Laboratory for Salmonella Vol. 24 No. 2 June 2018 ISSN 2211-6877 Continuation of Newsletter Community Reference Laboratory for Salmonella ISSN 1572-3836 Page 2 of 63
Horizontal transfer and pathogenicity
 Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
Horizontal transfer and pathogenicity Victoria Moiseeva Genomics, Master on Advanced Genetics UAB, Barcelona, 2014 INDEX Horizontal Transfer Horizontal gene transfer mechanisms Detection methods of HGT
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Microbiota: Its Evolution and Essence. Hsin-Jung Joyce Wu "Microbiota and man: the story about us
 Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
1. HyperLogLog algorithm
 SUPPLEMENTARY INFORMATION FOR KRAKENHLL (BREITWIESER AND SALZBERG, 2018) 1. HyperLogLog algorithm... 1 2. Database building and reanalysis of the patient data (Salzberg, et al., 2016)... 7 3. Enabling
SUPPLEMENTARY INFORMATION FOR KRAKENHLL (BREITWIESER AND SALZBERG, 2018) 1. HyperLogLog algorithm... 1 2. Database building and reanalysis of the patient data (Salzberg, et al., 2016)... 7 3. Enabling
Probing diversity in a hidden world: applications of NGS in microbial ecology
 Probing diversity in a hidden world: applications of NGS in microbial ecology Guus Roeselers TNO, Microbiology & Systems Biology Group Symposium on Next Generation Sequencing October 21, 2013 Royal Museum
Probing diversity in a hidden world: applications of NGS in microbial ecology Guus Roeselers TNO, Microbiology & Systems Biology Group Symposium on Next Generation Sequencing October 21, 2013 Royal Museum
Genomes and Their Evolution
 Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Centrifuge: rapid and sensitive classification of metagenomic sequences
 Centrifuge: rapid and sensitive classification of metagenomic sequences Daehwan Kim, Li Song, Florian P. Breitwieser, and Steven L. Salzberg Supplementary Material Supplementary Table 1 Supplementary Note
Centrifuge: rapid and sensitive classification of metagenomic sequences Daehwan Kim, Li Song, Florian P. Breitwieser, and Steven L. Salzberg Supplementary Material Supplementary Table 1 Supplementary Note
Vital Statistics Derived from Complete Genome Sequencing (for E. coli MG1655)
 We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
We still consider the E. coli genome as a fairly typical bacterial genome, and given the extensive information available about this organism and it's lifestyle, the E. coli genome is a useful point of
Whole Genome Alignment. Adam Phillippy University of Maryland, Fall 2012
 Whole Genome Alignment Adam Phillippy University of Maryland, Fall 2012 Motivation cancergenome.nih.gov Breast cancer karyotypes www.path.cam.ac.uk Goal of whole-genome alignment } For two genomes, A and
Whole Genome Alignment Adam Phillippy University of Maryland, Fall 2012 Motivation cancergenome.nih.gov Breast cancer karyotypes www.path.cam.ac.uk Goal of whole-genome alignment } For two genomes, A and
Pipelining RDP Data to the Taxomatic Background Accomplishments vs objectives
 Pipelining RDP Data to the Taxomatic Timothy G. Lilburn, PI/Co-PI George M. Garrity, PI/Co-PI (Collaborative) James R. Cole, Co-PI (Collaborative) Project ID 0010734 Grant No. DE-FG02-04ER63932 Background
Pipelining RDP Data to the Taxomatic Timothy G. Lilburn, PI/Co-PI George M. Garrity, PI/Co-PI (Collaborative) James R. Cole, Co-PI (Collaborative) Project ID 0010734 Grant No. DE-FG02-04ER63932 Background
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction. Lesser Tenrec (Echinops telfairi)
 Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction Lesser Tenrec (Echinops telfairi) Goals: 1. Use phylogenetic experimental design theory to select optimal taxa to
Phylogenetics - Orthology, phylogenetic experimental design and phylogeny reconstruction Lesser Tenrec (Echinops telfairi) Goals: 1. Use phylogenetic experimental design theory to select optimal taxa to
Taxonomy and Clustering of SSU rrna Tags. Susan Huse Josephine Bay Paul Center August 5, 2013
 Taxonomy and Clustering of SSU rrna Tags Susan Huse Josephine Bay Paul Center August 5, 2013 Primary Methods of Taxonomic Assignment Bayesian Kmer Matching RDP http://rdp.cme.msu.edu Wang, et al (2007)
Taxonomy and Clustering of SSU rrna Tags Susan Huse Josephine Bay Paul Center August 5, 2013 Primary Methods of Taxonomic Assignment Bayesian Kmer Matching RDP http://rdp.cme.msu.edu Wang, et al (2007)
Use of the 3M Molecular Detection System for Salmonella and Listeria spp.
 Use of the 3M Molecular Detection System for Salmonella and Listeria spp. March 11, 213 Prof Steve Forsythe Pathogen Research Centre, School of Science and Technology Nottingham Trent University Clifton
Use of the 3M Molecular Detection System for Salmonella and Listeria spp. March 11, 213 Prof Steve Forsythe Pathogen Research Centre, School of Science and Technology Nottingham Trent University Clifton
Benchmarking of Methods for Genomic Taxonomy.
 JCM Accepts, published online ahead of print on 26 February 2014 J. Clin. Microbiol. doi:10.1128/jcm.02981-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 Benchmarking of Methods
JCM Accepts, published online ahead of print on 26 February 2014 J. Clin. Microbiol. doi:10.1128/jcm.02981-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 Benchmarking of Methods
DATA ACQUISITION FROM BIO-DATABASES AND BLAST. Natapol Pornputtapong 18 January 2018
 DATA ACQUISITION FROM BIO-DATABASES AND BLAST Natapol Pornputtapong 18 January 2018 DATABASE Collections of data To share multi-user interface To prevent data loss To make sure to get the right things
DATA ACQUISITION FROM BIO-DATABASES AND BLAST Natapol Pornputtapong 18 January 2018 DATABASE Collections of data To share multi-user interface To prevent data loss To make sure to get the right things
Microbiology Helmut Pospiech
 Microbiology http://researchmagazine.uga.edu/summer2002/bacteria.htm 05.04.2018 Helmut Pospiech The Species Concept in Microbiology No universally accepted concept of species for prokaryotes Current definition
Microbiology http://researchmagazine.uga.edu/summer2002/bacteria.htm 05.04.2018 Helmut Pospiech The Species Concept in Microbiology No universally accepted concept of species for prokaryotes Current definition
CLASSIFYING LABORATORY TEST RESULTS USING MACHINE LEARNING
 CLASSIFYING LABORATORY TEST RESULTS USING MACHINE LEARNING Joy (Sizhe) Chen, Kenny Chiu, William Lu, Nilgoon Zarei AUGUST 31, 2018 TEAM Joy (Sizhe) Chen Kenny Chiu William Lu Nelly (Nilgoon) Zarei 2 AGENDA
CLASSIFYING LABORATORY TEST RESULTS USING MACHINE LEARNING Joy (Sizhe) Chen, Kenny Chiu, William Lu, Nilgoon Zarei AUGUST 31, 2018 TEAM Joy (Sizhe) Chen Kenny Chiu William Lu Nelly (Nilgoon) Zarei 2 AGENDA
Comparative Genomics II
 Comparative Genomics II Advances in Bioinformatics and Genomics GEN 240B Jason Stajich May 19 Comparative Genomics II Slide 1/31 Outline Introduction Gene Families Pairwise Methods Phylogenetic Methods
Comparative Genomics II Advances in Bioinformatics and Genomics GEN 240B Jason Stajich May 19 Comparative Genomics II Slide 1/31 Outline Introduction Gene Families Pairwise Methods Phylogenetic Methods
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Review of Bacterial Source Tracking in Texas. Kevin Wagner, George DiGiovanni, Terry Gentry, Elizabeth Casarez, Emily Martin
 Review of Bacterial Source Tracking in Texas Kevin Wagner, George DiGiovanni, Terry Gentry, Elizabeth Casarez, Emily Martin Bacteria The #1 Cause of Water Quality Impairment in Texas Where did the Bacteria
Review of Bacterial Source Tracking in Texas Kevin Wagner, George DiGiovanni, Terry Gentry, Elizabeth Casarez, Emily Martin Bacteria The #1 Cause of Water Quality Impairment in Texas Where did the Bacteria
Outline. I. Methods. II. Preliminary Results. A. Phylogeny Methods B. Whole Genome Methods C. Horizontal Gene Transfer
 Comparative Genomics Preliminary Results April 4, 2016 Juan Castro, Aroon Chande, Cheng Chen, Evan Clayton, Hector Espitia, Alli Gombolay, Walker Gussler, Ken Lee, Tyrone Lee, Hari Prasanna, Carlos Ruiz,
Comparative Genomics Preliminary Results April 4, 2016 Juan Castro, Aroon Chande, Cheng Chen, Evan Clayton, Hector Espitia, Alli Gombolay, Walker Gussler, Ken Lee, Tyrone Lee, Hari Prasanna, Carlos Ruiz,
MOLECULAR PHYLOGENY AND GENETIC DIVERSITY ANALYSIS. Masatoshi Nei"
 MOLECULAR PHYLOGENY AND GENETIC DIVERSITY ANALYSIS Masatoshi Nei" Abstract: Phylogenetic trees: Recent advances in statistical methods for phylogenetic reconstruction and genetic diversity analysis were
MOLECULAR PHYLOGENY AND GENETIC DIVERSITY ANALYSIS Masatoshi Nei" Abstract: Phylogenetic trees: Recent advances in statistical methods for phylogenetic reconstruction and genetic diversity analysis were
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Ribosomal multilocus sequence typing: universal characterization of bacteria from domain to strain
 Microbiology (2012), 158, 1005 1015 DOI 10.1099/mic.0.055459-0 Ribosomal multilocus sequence typing: universal characterization of bacteria from domain to strain Keith A. Jolley, Carly M. Bliss, Julia
Microbiology (2012), 158, 1005 1015 DOI 10.1099/mic.0.055459-0 Ribosomal multilocus sequence typing: universal characterization of bacteria from domain to strain Keith A. Jolley, Carly M. Bliss, Julia
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett. Introduction
 The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
Introduction to Microbiology. CLS 212: Medical Microbiology Miss Zeina Alkudmani
 Introduction to Microbiology CLS 212: Medical Microbiology Miss Zeina Alkudmani Microbiology Micro- means very small (that needs a microscope to see). Microbiology is the study of very small living organisms.
Introduction to Microbiology CLS 212: Medical Microbiology Miss Zeina Alkudmani Microbiology Micro- means very small (that needs a microscope to see). Microbiology is the study of very small living organisms.
Supplemental Materials
 JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
West Windsor-Plainsboro Regional School District AP Biology Grades 11-12
 West Windsor-Plainsboro Regional School District AP Biology Grades 11-12 Unit 1: Chemistry of Life Content Area: Science Course & Grade Level: AP Biology, 11 12 Summary and Rationale The structural levels
West Windsor-Plainsboro Regional School District AP Biology Grades 11-12 Unit 1: Chemistry of Life Content Area: Science Course & Grade Level: AP Biology, 11 12 Summary and Rationale The structural levels
Whole-Genome Sequencing of Drug-Resistant Salmonella enterica Isolated from Dairy
 AEM Accepted Manuscript Posted Online 7 April 2017 Appl. Environ. Microbiol. doi:10.1128/aem.00140-17 Copyright 2017 American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11 12 13
AEM Accepted Manuscript Posted Online 7 April 2017 Appl. Environ. Microbiol. doi:10.1128/aem.00140-17 Copyright 2017 American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11 12 13
The Pennsylvania State University. The Graduate School. Department of Food Science VIRULENCE GENE AND CRISPR MULTILOCUS SEQUENCE TYPING
 The Pennsylvania State University The Graduate School Department of Food Science VIRULENCE GENE AND CRISPR MULTILOCUS SEQUENCE TYPING SCHEME FOR SUBTYPING THE MAJOR SEROVARS OF SALMONELLA ENTERICA SUBSPECIES
The Pennsylvania State University The Graduate School Department of Food Science VIRULENCE GENE AND CRISPR MULTILOCUS SEQUENCE TYPING SCHEME FOR SUBTYPING THE MAJOR SEROVARS OF SALMONELLA ENTERICA SUBSPECIES
Lecture WS Evolutionary Genetics Part I 1
 Quantitative genetics Quantitative genetics is the study of the inheritance of quantitative/continuous phenotypic traits, like human height and body size, grain colour in winter wheat or beak depth in
Quantitative genetics Quantitative genetics is the study of the inheritance of quantitative/continuous phenotypic traits, like human height and body size, grain colour in winter wheat or beak depth in
Genetic Basis of Variation in Bacteria
 Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
Mechanisms of Infectious Disease Fall 2009 Genetics I Jonathan Dworkin, PhD Department of Microbiology jonathan.dworkin@columbia.edu Genetic Basis of Variation in Bacteria I. Organization of genetic material
The Genetic Epidemiology of Antibiotic Resistance
 The Genetic Epidemiology of Antibiotic Resistance Professor Neil Woodford Antimicrobial Resistance & Healthcare Associated Infections (AMRHAI) Reference Unit Crown copyright The forensics of AMR Resistance
The Genetic Epidemiology of Antibiotic Resistance Professor Neil Woodford Antimicrobial Resistance & Healthcare Associated Infections (AMRHAI) Reference Unit Crown copyright The forensics of AMR Resistance
IR Biotyper. Innovation with Integrity. Microbial typing for real-time epidemiology FT-IR
 IR Biotyper Microbial typing for real-time epidemiology Innovation with Integrity FT-IR IR Biotyper - Proactive hospital hygiene and infection control Fast, easy-to-apply and economical typing methods
IR Biotyper Microbial typing for real-time epidemiology Innovation with Integrity FT-IR IR Biotyper - Proactive hospital hygiene and infection control Fast, easy-to-apply and economical typing methods
Evolutionary Genetics: Part 0.2 Introduction to Population genetics
 Evolutionary Genetics: Part 0.2 Introduction to Population genetics S. chilense S. peruvianum Winter Semester 2012-2013 Prof Aurélien Tellier FG Populationsgenetik Population genetics Evolution = changes
Evolutionary Genetics: Part 0.2 Introduction to Population genetics S. chilense S. peruvianum Winter Semester 2012-2013 Prof Aurélien Tellier FG Populationsgenetik Population genetics Evolution = changes
