Supplementary Figures
|
|
|
- Chester McDaniel
- 6 years ago
- Views:
Transcription
1 Supplementary Figures Supplementary Figure 1 Principal components analysis (PCA) of all samples analyzed in the discovery phase. Colors represent the phenotype of study populations. a) The first sample (GWAS1) was the previously published GWAS dataset of leprosy, consisting of 706 leprosy cases, 1,223 healthy controls, all of northern Chinese Han decent; b-d) The second one (GWAS2) was a newly published dataset of 840 leprosy cases and 924 controls from northern (Chinese Han, b) and southern China (Chinese Han c and ethnic minorities, d); e-f) The third sample was a new GWAS dataset (GWAS3) of 1197 leprosy cases and 1426 controls from northern (Chinese Han, e) and southern China (Chinese Han, f)
2 Supplementary Figure 2 Quantile-quantile plot of the associations Left panel, before removal of SNPs located within known leprosy loci; Right panel, after removal of SNPs located within the known leprosy loci. Dotted vertical line in right panel shows the point where the statistics lift-off from the expected null distribution (between log10(p) of 2 to 3).
3 TSS Repressed DGF SuperEnhancer H3K27ac Promoter 3-PrimeUTR H3K4me3 5-PrimeUTR Enhancer H3K9ac Coding Conserved DHS TFBS PromoterFlanking Transcribed Intron FetalDHS CTCF H3K4me1 WeakEnhancer Immune GI Connective_Bone Adrenal_Pancreas Kidney Cardiovascular SkeletalMuscle CNS Liver Supplementary Figure 3 Overall genetic architecture of leprosy across functional categories and tissues. Enrichment estimates for the main annotations and tissues of LDSC. Error bars represent 95% confidence intervals around the estimate. Categories are sorted by P value, with boxes indicating annotations or tissues that pass the multiple testing significance threshold. CNS, central nervous system;; DHS, DNase hypersenstivity; GI, gastrointestinal; TFBS, transcription factor binding site; Tss, transcription start site; UTR, untranslated region. 24" 22" 20" 18" 16" 14" 12" 10" 8" 6" 4" 2" 0"
4 Supplementary Tables Supplementary Table 1 Baseline characteristics of cases and controls Male/ Mean Mean age N Female age at onset CASES CONTROLS Ethnicity North Sichuan Yunnan Guizhou South N Han Han Han Han Minority Male/ Female Mean North Sichuan age Han Han Ethnicity Yunnan Guizhou Han Han Discovery Study / / Discovery Study / / Discovery Study / / Replication Phase / / Replication Phase / / Total/Mean / / *The location of North Han, Sichuan, Yunan, Guizhou were showed in supplementary figure 3 South Minority
5 Supplementary Table 2 Association results of 127 replicated SNPs in Stage 2 CHR SNP BP A1 A2 F_A F_U OR L95 U95 P 1 rs T C E-01 1 rs A G E-01 1 rs T C E-02 1 rs C T E-01 1 rs A G E-01 1 rs A G E-02 1 rs C T E-01 1 rs A G E-02 1 rs G C E-01 2 rs G T E-01 2 rs G A E-01 2 rs G C E-01 2 rs A G E-01 2 rs G A E-01 2 rs A G E-01 2 rs T C E-01 2 rs A G E-01 2 rs T C E-01 2 rs C T E-01 2 rs G A E-01 2 rs C T E-01 2 rs G C E-01 3 rs G A E-01 3 rs C T E-03 3 rs T C E-06 3 rs A T E-01 3 rs A C E-01 3 rs G A E-01 3 rs C T E-02 3 rs T A inf 9.97E-01 4 rs A G E-04 4 rs A G E-01 4 rs C T E-01 4 rs T C E-02 4 rs T C E-03 4 rs A G E-02 5 rs C A E-01
6 5 rs C G E-03 5 rs G C E-01 5 rs C G E-01 5 rs G A E-01 5 rs T C E-01 5 rs G C inf 9.99E-01 5 rs C T E-01 5 rs A G E-01 5 rs C T E-01 6 rs T C E-01 6 rs G A E-01 6 rs T G E-01 6 rs T C E-03 6 rs C T E-09 6 rs C T E-03 6 rs T C E-01 7 rs T C E-02 7 rs T A E-01 7 rs C T E-01 7 rs T C E-04 7 rs A G E-02 7 rs G A E-02 7 rs C A E-01 7 rs T C E-01 7 rs G A E-01 7 rs T C E-01 7 rs A G E-02 8 rs C A E-01 8 rs A T E-01 8 rs C A E-03 8 rs C T E-01 8 rs G T E-01 8 rs A G E-01 8 rs C T E-02 8 rs A G E-01 8 rs A G E-02 8 rs G C E-01 8 rs T C E-02 9 rs T C E-03 9 rs C T E-02
7 9 rs T C E-02 9 rs G A E-01 9 rs G A E-02 9 rs A C E-02 9 rs G A E-01 9 rs G T E rs T C E rs T C E rs G T E rs T G E rs T C E rs T C E rs T C E rs A G E rs G C E rs C A E rs G A E rs C T E rs A G E rs G A E rs A C E rs G A E rs C T E rs T C E rs G A E rs A G E rs C T E rs C T E rs C T E rs G A E rs G A E rs T C E rs A G E rs G T E rs G A E rs T C E rs T G E rs T C E rs G A E rs T C E-01
8 18 rs A G E rs A G E rs A C E rs A G E rs T C E rs T C E rs C T E rs T C E rs T C E rs A G E-01 A1 is the minor allele, while F_A represents allele frequency in cases and F_U represents allele frequency in controls.
9 Supplementary Table 3 Association results of 21 replicated SNPs in Stage 3 SNP info meta of stage3 meta all CHR BP SNP A1 A2 P OR Q I P OR Q I rs T C 1.56E E rs A G 3.04E E rs A G 8.22E E rs C T 2.84E E rs C T 3.46E E rs T C 6.37E E rs A G 9.08E E rs C G 4.42E E rs T C 6.69E E rs C T 1.09E E rs T C 1.03E E rs C A 2.26E E rs C T 3.09E E rs A G 9.27E E rs T C 2.15E E rs T G 1.96E E rs A G 8.23E E rs G A 8.53E E rs A G 2.12E E rs G A 8.05E E *Rs , rs942793, rs were not reported in the manuscript due to either failed of HWE test or significant Q value.
10 Supplementary Table 4 eqtl analysis of four novel associations Lead SNP rs rs rs rs rs rs rs rs rs Study ID Westra 2013 Westra 2013 Westra 2013 Westra 2013 GTEx20 15_v6 Westra 2013 GTEx20 15_v6 GTEx20 15_v6 Westra 2013 Paper_title Systematic identification of trans eqtls as putative drivers of known disease associations Systematic identification of trans eqtls as putative drivers of known disease associations Systematic identification of trans eqtls as putative drivers of known disease associations Systematic identification of trans eqtls as putative drivers of known disease associations The Genotype-Tissue Expression (GTEx) pilot analysis: Multitissue gene regulation in humans Systematic identification of trans eqtls as putative drivers of known disease associations The Genotype-Tissue Expression (GTEx) pilot analysis: Multitissue gene regulation in humans The Genotype-Tissue Expression (GTEx) pilot analysis: Multitissue gene regulation in humans Systematic identification of trans eqtls as putative drivers of known disease associations Tissue Correlated -gene p-value eqtl SNP r2 with lead SNP D' with lead SNP Whole_Blood SYN2 1.74E-04 rs Whole_Blood SYN2 7.69E-05 rs Whole_Blood SYN2 1.80E-04 rs Whole_Blood SYN2 1.83E-04 rs Thyroid BBS9 2.81E-07 rs Whole_Blood BBS9 3.18E-04 rs Cells_Transfor med_fibroblas ts CTSB 7.48E-18 rs Whole_Blood CTSB 1.35E-09 rs Whole_Blood MED E-05 rs *Lead SNP represents the current reported associations within the genomic region. eqtl SNP represents those SNPs reported in the publications with listed paper title, which were in high LD (r2>0.9 &D >0.9) with lead SNPs
11 Supplementary Table 5 Association results of HLA imputation Classical HLA allele P OR Q P_con OR_con HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E
12 HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_A_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E
13 HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E
14 HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E HLA_B_ E E
15 HLA_B_ E E HLA_B_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E
16 HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_C_ E E HLA_DPA1_ E E HLA_DPA1_ E E HLA_DPA1_ E E HLA_DPA1_ E E HLA_DPA1_ E E HLA_DPA1_ E E HLA_DPA1_ E E HLA_DPA1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E
17 HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E #N/A #N/A HLA_DPB1_ E #N/A #N/A HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DPB1_ E E HLA_DQA1_ E E HLA_DQA1_ E E HLA_DQA1_ E E HLA_DQA1_ E E HLA_DQA1_ E E HLA_DQA1_ E E HLA_DQA1_ E E HLA_DQA1_ E E HLA_DQA1_ E E HLA_DQA1_ E E
18 HLA_DQA1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DQB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E
19 HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E
20 HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E HLA_DRB1_ E E P_con represents the P value after conditioning on HLA-DRB1*15:01 OR_con represents the odds ratio after conditioning on HLA-DRB1*15:01 Supplementary Table 6 Heritability estimates for genome-wide SNPs in leprosy on assumed disease risk. Category Heritability All SNPs SE known region (LD) explained ratio liability scale h2 (prevalence = ) %
Nature Genetics: doi: /ng Supplementary Figure 1. Number of cases and proxy cases required to detect association at designs.
 Supplementary Figure 1 Number of cases and proxy cases required to detect association at designs. = 5 10 8 for case control and proxy case control The ratio of controls to cases (or proxy cases) is 1.
Supplementary Figure 1 Number of cases and proxy cases required to detect association at designs. = 5 10 8 for case control and proxy case control The ratio of controls to cases (or proxy cases) is 1.
Proportional Variance Explained by QLT and Statistical Power. Proportional Variance Explained by QTL and Statistical Power
 Proportional Variance Explained by QTL and Statistical Power Partitioning the Genetic Variance We previously focused on obtaining variance components of a quantitative trait to determine the proportion
Proportional Variance Explained by QTL and Statistical Power Partitioning the Genetic Variance We previously focused on obtaining variance components of a quantitative trait to determine the proportion
Department of Forensic Psychiatry, School of Medicine & Forensics, Xi'an Jiaotong University, Xi'an, China;
 Title: Evaluation of genetic susceptibility of common variants in CACNA1D with schizophrenia in Han Chinese Author names and affiliations: Fanglin Guan a,e, Lu Li b, Chuchu Qiao b, Gang Chen b, Tinglin
Title: Evaluation of genetic susceptibility of common variants in CACNA1D with schizophrenia in Han Chinese Author names and affiliations: Fanglin Guan a,e, Lu Li b, Chuchu Qiao b, Gang Chen b, Tinglin
1 Springer. Nan M. Laird Christoph Lange. The Fundamentals of Modern Statistical Genetics
 1 Springer Nan M. Laird Christoph Lange The Fundamentals of Modern Statistical Genetics 1 Introduction to Statistical Genetics and Background in Molecular Genetics 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
1 Springer Nan M. Laird Christoph Lange The Fundamentals of Modern Statistical Genetics 1 Introduction to Statistical Genetics and Background in Molecular Genetics 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
Lecture 2: Genetic Association Testing with Quantitative Traits. Summer Institute in Statistical Genetics 2017
 Lecture 2: Genetic Association Testing with Quantitative Traits Instructors: Timothy Thornton and Michael Wu Summer Institute in Statistical Genetics 2017 1 / 29 Introduction to Quantitative Trait Mapping
Lecture 2: Genetic Association Testing with Quantitative Traits Instructors: Timothy Thornton and Michael Wu Summer Institute in Statistical Genetics 2017 1 / 29 Introduction to Quantitative Trait Mapping
Association Testing with Quantitative Traits: Common and Rare Variants. Summer Institute in Statistical Genetics 2014 Module 10 Lecture 5
 Association Testing with Quantitative Traits: Common and Rare Variants Timothy Thornton and Katie Kerr Summer Institute in Statistical Genetics 2014 Module 10 Lecture 5 1 / 41 Introduction to Quantitative
Association Testing with Quantitative Traits: Common and Rare Variants Timothy Thornton and Katie Kerr Summer Institute in Statistical Genetics 2014 Module 10 Lecture 5 1 / 41 Introduction to Quantitative
Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control
 Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control Xiaoquan Wen Department of Biostatistics, University of Michigan A Model
Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control Xiaoquan Wen Department of Biostatistics, University of Michigan A Model
BTRY 7210: Topics in Quantitative Genomics and Genetics
 BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu February 12, 2015 Lecture 3:
BTRY 7210: Topics in Quantitative Genomics and Genetics Jason Mezey Biological Statistics and Computational Biology (BSCB) Department of Genetic Medicine jgm45@cornell.edu February 12, 2015 Lecture 3:
Linear Regression (1/1/17)
 STA613/CBB540: Statistical methods in computational biology Linear Regression (1/1/17) Lecturer: Barbara Engelhardt Scribe: Ethan Hada 1. Linear regression 1.1. Linear regression basics. Linear regression
STA613/CBB540: Statistical methods in computational biology Linear Regression (1/1/17) Lecturer: Barbara Engelhardt Scribe: Ethan Hada 1. Linear regression 1.1. Linear regression basics. Linear regression
Quantitative Genomics and Genetics BTRY 4830/6830; PBSB
 Quantitative Genomics and Genetics BTRY 4830/6830; PBSB.5201.01 Lecture16: Population structure and logistic regression I Jason Mezey jgm45@cornell.edu April 11, 2017 (T) 8:40-9:55 Announcements I April
Quantitative Genomics and Genetics BTRY 4830/6830; PBSB.5201.01 Lecture16: Population structure and logistic regression I Jason Mezey jgm45@cornell.edu April 11, 2017 (T) 8:40-9:55 Announcements I April
Learning Your Identity and Disease from Research Papers: Information Leaks in Genome-Wide Association Study
 Learning Your Identity and Disease from Research Papers: Information Leaks in Genome-Wide Association Study Rui Wang, Yong Li, XiaoFeng Wang, Haixu Tang and Xiaoyong Zhou Indiana University at Bloomington
Learning Your Identity and Disease from Research Papers: Information Leaks in Genome-Wide Association Study Rui Wang, Yong Li, XiaoFeng Wang, Haixu Tang and Xiaoyong Zhou Indiana University at Bloomington
Control of Gene Expression
 Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Quantitative Genomics and Genetics BTRY 4830/6830; PBSB
 Quantitative Genomics and Genetics BTRY 4830/6830; PBSB.5201.01 Lecture 18: Introduction to covariates, the QQ plot, and population structure II + minimal GWAS steps Jason Mezey jgm45@cornell.edu April
Quantitative Genomics and Genetics BTRY 4830/6830; PBSB.5201.01 Lecture 18: Introduction to covariates, the QQ plot, and population structure II + minimal GWAS steps Jason Mezey jgm45@cornell.edu April
Common Variants near MBNL1 and NKX2-5 are Associated with Infantile Hypertrophic Pyloric Stenosis
 Supplementary Information: Common Variants near MBNL1 and NKX2-5 are Associated with Infantile Hypertrophic Pyloric Stenosis Bjarke Feenstra 1*, Frank Geller 1*, Camilla Krogh 1, Mads V. Hollegaard 2,
Supplementary Information: Common Variants near MBNL1 and NKX2-5 are Associated with Infantile Hypertrophic Pyloric Stenosis Bjarke Feenstra 1*, Frank Geller 1*, Camilla Krogh 1, Mads V. Hollegaard 2,
(Genome-wide) association analysis
 (Genome-wide) association analysis 1 Key concepts Mapping QTL by association relies on linkage disequilibrium in the population; LD can be caused by close linkage between a QTL and marker (= good) or by
(Genome-wide) association analysis 1 Key concepts Mapping QTL by association relies on linkage disequilibrium in the population; LD can be caused by close linkage between a QTL and marker (= good) or by
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Expression QTLs and Mapping of Complex Trait Loci. Paul Schliekelman Statistics Department University of Georgia
 Expression QTLs and Mapping of Complex Trait Loci Paul Schliekelman Statistics Department University of Georgia Definitions: Genes, Loci and Alleles A gene codes for a protein. Proteins due everything.
Expression QTLs and Mapping of Complex Trait Loci Paul Schliekelman Statistics Department University of Georgia Definitions: Genes, Loci and Alleles A gene codes for a protein. Proteins due everything.
ChIP seq peak calling. Statistical integration between ChIP seq and RNA seq
 Institute for Computational Biomedicine ChIP seq peak calling Statistical integration between ChIP seq and RNA seq Olivier Elemento, PhD ChIP-seq to map where transcription factors bind DNA Transcription
Institute for Computational Biomedicine ChIP seq peak calling Statistical integration between ChIP seq and RNA seq Olivier Elemento, PhD ChIP-seq to map where transcription factors bind DNA Transcription
Selection-adjusted estimation of effect sizes
 Selection-adjusted estimation of effect sizes with an application in eqtl studies Snigdha Panigrahi 19 October, 2017 Stanford University Selective inference - introduction Selective inference Statistical
Selection-adjusted estimation of effect sizes with an application in eqtl studies Snigdha Panigrahi 19 October, 2017 Stanford University Selective inference - introduction Selective inference Statistical
Lecture 1: Case-Control Association Testing. Summer Institute in Statistical Genetics 2015
 Timothy Thornton and Michael Wu Summer Institute in Statistical Genetics 2015 1 / 1 Introduction Association mapping is now routinely being used to identify loci that are involved with complex traits.
Timothy Thornton and Michael Wu Summer Institute in Statistical Genetics 2015 1 / 1 Introduction Association mapping is now routinely being used to identify loci that are involved with complex traits.
Régression en grande dimension et épistasie par blocs pour les études d association
 Régression en grande dimension et épistasie par blocs pour les études d association V. Stanislas, C. Dalmasso, C. Ambroise Laboratoire de Mathématiques et Modélisation d Évry "Statistique et Génome" 1
Régression en grande dimension et épistasie par blocs pour les études d association V. Stanislas, C. Dalmasso, C. Ambroise Laboratoire de Mathématiques et Modélisation d Évry "Statistique et Génome" 1
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Schematic comparison of linking attacks and detection of a genome in a mixture of attacks. (a) Each box in the figure represents a dataset in the form of a matrix. Multiple boxes
Supplementary Figure 1 Schematic comparison of linking attacks and detection of a genome in a mixture of attacks. (a) Each box in the figure represents a dataset in the form of a matrix. Multiple boxes
Introduction to PLINK H3ABionet Course Covenant University, Nigeria
 UNIVERSITY OF THE WITWATERSRAND, JOHANNESBURG Introduction to PLINK H3ABionet Course Covenant University, Nigeria Scott Hazelhurst H3ABioNet funded by NHGRI grant number U41HG006941 Wits Bioinformatics
UNIVERSITY OF THE WITWATERSRAND, JOHANNESBURG Introduction to PLINK H3ABionet Course Covenant University, Nigeria Scott Hazelhurst H3ABioNet funded by NHGRI grant number U41HG006941 Wits Bioinformatics
Friday Harbor From Genetics to GWAS (Genome-wide Association Study) Sept David Fardo
 Friday Harbor 2017 From Genetics to GWAS (Genome-wide Association Study) Sept 7 2017 David Fardo Purpose: prepare for tomorrow s tutorial Genetic Variants Quality Control Imputation Association Visualization
Friday Harbor 2017 From Genetics to GWAS (Genome-wide Association Study) Sept 7 2017 David Fardo Purpose: prepare for tomorrow s tutorial Genetic Variants Quality Control Imputation Association Visualization
Designer Genes C Test
 Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Latent Variable models for GWAs
 Latent Variable models for GWAs Oliver Stegle Machine Learning and Computational Biology Research Group Max-Planck-Institutes Tübingen, Germany September 2011 O. Stegle Latent variable models for GWAs
Latent Variable models for GWAs Oliver Stegle Machine Learning and Computational Biology Research Group Max-Planck-Institutes Tübingen, Germany September 2011 O. Stegle Latent variable models for GWAs
1. Understand the methods for analyzing population structure in genomes
 MSCBIO 2070/02-710: Computational Genomics, Spring 2016 HW3: Population Genetics Due: 24:00 EST, April 4, 2016 by autolab Your goals in this assignment are to 1. Understand the methods for analyzing population
MSCBIO 2070/02-710: Computational Genomics, Spring 2016 HW3: Population Genetics Due: 24:00 EST, April 4, 2016 by autolab Your goals in this assignment are to 1. Understand the methods for analyzing population
The supplementary document of LLR: A latent low-rank approach to colocalizing genetic risk variants in multiple GWAS
 The supplementary document of LLR: A latent low-rank approach to colocalizing genetic risk variants in multiple GWAS Jin Liu 1, Xiang Wan 2, Chaolong Wang 3, Chao Yang 4, Xiaowei Zhou 5, and Can Yang 6
The supplementary document of LLR: A latent low-rank approach to colocalizing genetic risk variants in multiple GWAS Jin Liu 1, Xiang Wan 2, Chaolong Wang 3, Chao Yang 4, Xiaowei Zhou 5, and Can Yang 6
Case-Control Association Testing. Case-Control Association Testing
 Introduction Association mapping is now routinely being used to identify loci that are involved with complex traits. Technological advances have made it feasible to perform case-control association studies
Introduction Association mapping is now routinely being used to identify loci that are involved with complex traits. Technological advances have made it feasible to perform case-control association studies
A novel fuzzy set based multifactor dimensionality reduction method for detecting gene-gene interaction
 A novel fuzzy set based multifactor dimensionality reduction method for detecting gene-gene interaction Sangseob Leem, Hye-Young Jung, Sungyoung Lee and Taesung Park Bioinformatics and Biostatistics lab
A novel fuzzy set based multifactor dimensionality reduction method for detecting gene-gene interaction Sangseob Leem, Hye-Young Jung, Sungyoung Lee and Taesung Park Bioinformatics and Biostatistics lab
CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS
 CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS * Some contents are adapted from Dr. Hung Huang and Dr. Chengkai Li at UT Arlington Mingon Kang, Ph.D. Computer Science, Kennesaw State University Problems
CS 4491/CS 7990 SPECIAL TOPICS IN BIOINFORMATICS * Some contents are adapted from Dr. Hung Huang and Dr. Chengkai Li at UT Arlington Mingon Kang, Ph.D. Computer Science, Kennesaw State University Problems
p(d g A,g B )p(g B ), g B
 Supplementary Note Marginal effects for two-locus models Here we derive the marginal effect size of the three models given in Figure 1 of the main text. For each model we assume the two loci (A and B)
Supplementary Note Marginal effects for two-locus models Here we derive the marginal effect size of the three models given in Figure 1 of the main text. For each model we assume the two loci (A and B)
Bayesian Partition Models for Identifying Expression Quantitative Trait Loci
 Journal of the American Statistical Association ISSN: 0162-1459 (Print) 1537-274X (Online) Journal homepage: http://www.tandfonline.com/loi/uasa20 Bayesian Partition Models for Identifying Expression Quantitative
Journal of the American Statistical Association ISSN: 0162-1459 (Print) 1537-274X (Online) Journal homepage: http://www.tandfonline.com/loi/uasa20 Bayesian Partition Models for Identifying Expression Quantitative
Figure E1 Manhattan and QQ plots for FEV 1 meta-analyses across all Hispanic ancestry groups
 Figure E1 Manhattan and QQ plots for FEV 1 meta-analyses across all Hispanic ancestry groups Figure E1a: All Participants λgc = 1. Figure E1b: Ever Smokers λgc = 1. Figure E1c: Never Smokers λgc = 1.1
Figure E1 Manhattan and QQ plots for FEV 1 meta-analyses across all Hispanic ancestry groups Figure E1a: All Participants λgc = 1. Figure E1b: Ever Smokers λgc = 1. Figure E1c: Never Smokers λgc = 1.1
Proper Use of Allele-Specific Expression Improves Statistical Power for cis-eqtl Mapping with RNA-Seq Data
 Proper Use of Allele-Specific Expression Improves Statistical Power for cis-eqtl Mapping with RNA-Seq Data Yi-Juan HU, Wei SUN, Jung-Ying TZENG, and Charles M. PEROU Studies of expression quantitative
Proper Use of Allele-Specific Expression Improves Statistical Power for cis-eqtl Mapping with RNA-Seq Data Yi-Juan HU, Wei SUN, Jung-Ying TZENG, and Charles M. PEROU Studies of expression quantitative
Solutions to Even-Numbered Exercises to accompany An Introduction to Population Genetics: Theory and Applications Rasmus Nielsen Montgomery Slatkin
 Solutions to Even-Numbered Exercises to accompany An Introduction to Population Genetics: Theory and Applications Rasmus Nielsen Montgomery Slatkin CHAPTER 1 1.2 The expected homozygosity, given allele
Solutions to Even-Numbered Exercises to accompany An Introduction to Population Genetics: Theory and Applications Rasmus Nielsen Montgomery Slatkin CHAPTER 1 1.2 The expected homozygosity, given allele
Exam 1 PBG430/
 1 Exam 1 PBG430/530 2014 1. You read that the genome size of maize is 2,300 Mb and that in this species 2n = 20. This means that there are 2,300 Mb of DNA in a cell that is a. n (e.g. gamete) b. 2n (e.g.
1 Exam 1 PBG430/530 2014 1. You read that the genome size of maize is 2,300 Mb and that in this species 2n = 20. This means that there are 2,300 Mb of DNA in a cell that is a. n (e.g. gamete) b. 2n (e.g.
Gene Regula*on, ChIP- X and DNA Mo*fs. Statistics in Genomics Hongkai Ji
 Gene Regula*on, ChIP- X and DNA Mo*fs Statistics in Genomics Hongkai Ji (hji@jhsph.edu) Genetic information is stored in DNA TCAGTTGGAGCTGCTCCCCCACGGCCTCTCCTCACATTCCACGTCCTGTAGCTCTATGACCTCCACCTTTGAGTCCCTCCTC
Gene Regula*on, ChIP- X and DNA Mo*fs Statistics in Genomics Hongkai Ji (hji@jhsph.edu) Genetic information is stored in DNA TCAGTTGGAGCTGCTCCCCCACGGCCTCTCCTCACATTCCACGTCCTGTAGCTCTATGACCTCCACCTTTGAGTCCCTCCTC
Quantile based Permutation Thresholds for QTL Hotspots. Brian S Yandell and Elias Chaibub Neto 17 March 2012
 Quantile based Permutation Thresholds for QTL Hotspots Brian S Yandell and Elias Chaibub Neto 17 March 2012 2012 Yandell 1 Fisher on inference We may at once admit that any inference from the particular
Quantile based Permutation Thresholds for QTL Hotspots Brian S Yandell and Elias Chaibub Neto 17 March 2012 2012 Yandell 1 Fisher on inference We may at once admit that any inference from the particular
Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites
 Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites Paper by: James P. Balhoff and Gregory A. Wray Presentation by: Stephanie Lucas Reviewed
Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites Paper by: James P. Balhoff and Gregory A. Wray Presentation by: Stephanie Lucas Reviewed
Analyzing metabolomics data for association with genotypes using two-component Gaussian mixture distributions
 Analyzing metabolomics data for association with genotypes using two-component Gaussian mixture distributions Jason Westra Department of Statistics, Iowa State University Ames, IA 50011, United States
Analyzing metabolomics data for association with genotypes using two-component Gaussian mixture distributions Jason Westra Department of Statistics, Iowa State University Ames, IA 50011, United States
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
EFFICIENT COMPUTATION WITH A LINEAR MIXED MODEL ON LARGE-SCALE DATA SETS WITH APPLICATIONS TO GENETIC STUDIES
 Submitted to the Annals of Applied Statistics EFFICIENT COMPUTATION WITH A LINEAR MIXED MODEL ON LARGE-SCALE DATA SETS WITH APPLICATIONS TO GENETIC STUDIES By Matti Pirinen, Peter Donnelly and Chris C.A.
Submitted to the Annals of Applied Statistics EFFICIENT COMPUTATION WITH A LINEAR MIXED MODEL ON LARGE-SCALE DATA SETS WITH APPLICATIONS TO GENETIC STUDIES By Matti Pirinen, Peter Donnelly and Chris C.A.
Asymptotic distribution of the largest eigenvalue with application to genetic data
 Asymptotic distribution of the largest eigenvalue with application to genetic data Chong Wu University of Minnesota September 30, 2016 T32 Journal Club Chong Wu 1 / 25 Table of Contents 1 Background Gene-gene
Asymptotic distribution of the largest eigenvalue with application to genetic data Chong Wu University of Minnesota September 30, 2016 T32 Journal Club Chong Wu 1 / 25 Table of Contents 1 Background Gene-gene
Sex-limitation Models
 Sex-limitation Models Meike Bartels (Brad, Sarah, Hermine, Ben, Elizabeth, and most of the rest of the faculty that has contributed bits and pieces to various versions of this talk) COPY FILES FROM: Faculty/meike/2016/heterogeneity
Sex-limitation Models Meike Bartels (Brad, Sarah, Hermine, Ben, Elizabeth, and most of the rest of the faculty that has contributed bits and pieces to various versions of this talk) COPY FILES FROM: Faculty/meike/2016/heterogeneity
Supplementary Figure 1. Nature Genetics: doi: /ng.3848
 Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
TRANSCRIPTOMICS. (or the analysis of the transcriptome) Mario Cáceres. Main objectives of genomics. Determine the entire DNA sequence of an organism
 TRANSCRIPTOMICS (or the analysis of the transcriptome) Mario Cáceres Main objectives of genomics Determine the entire DNA sequence of an organism Identify and annotate the complete set of genes encoded
TRANSCRIPTOMICS (or the analysis of the transcriptome) Mario Cáceres Main objectives of genomics Determine the entire DNA sequence of an organism Identify and annotate the complete set of genes encoded
Coding sequence array Office hours Wednesday 3-4pm 304A Stanley Hall
 Coding sequence array Office hours Wednesday 3-4pm 304A Stanley Hall Review session 5pm Thursday, Dec. 11 GPB100 Fig. 1.13 RNA-seq Expression effects of cancer AAAAA AAAAA AAAAA Solexa sequencing counts
Coding sequence array Office hours Wednesday 3-4pm 304A Stanley Hall Review session 5pm Thursday, Dec. 11 GPB100 Fig. 1.13 RNA-seq Expression effects of cancer AAAAA AAAAA AAAAA Solexa sequencing counts
Statistical Methods in Mapping Complex Diseases
 University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations Summer 8-12-2011 Statistical Methods in Mapping Complex Diseases Jing He University of Pennsylvania, jinghe@mail.med.upenn.edu
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations Summer 8-12-2011 Statistical Methods in Mapping Complex Diseases Jing He University of Pennsylvania, jinghe@mail.med.upenn.edu
Theoretical and computational aspects of association tests: application in case-control genome-wide association studies.
 Theoretical and computational aspects of association tests: application in case-control genome-wide association studies Mathieu Emily November 18, 2014 Caen mathieu.emily@agrocampus-ouest.fr - Agrocampus
Theoretical and computational aspects of association tests: application in case-control genome-wide association studies Mathieu Emily November 18, 2014 Caen mathieu.emily@agrocampus-ouest.fr - Agrocampus
Lesson 4: Understanding Genetics
 Lesson 4: Understanding Genetics 1 Terms Alleles Chromosome Co dominance Crossover Deoxyribonucleic acid DNA Dominant Genetic code Genome Genotype Heredity Heritability Heritability estimate Heterozygous
Lesson 4: Understanding Genetics 1 Terms Alleles Chromosome Co dominance Crossover Deoxyribonucleic acid DNA Dominant Genetic code Genome Genotype Heredity Heritability Heritability estimate Heterozygous
Genotype Imputation. Biostatistics 666
 Genotype Imputation Biostatistics 666 Previously Hidden Markov Models for Relative Pairs Linkage analysis using affected sibling pairs Estimation of pairwise relationships Identity-by-Descent Relatives
Genotype Imputation Biostatistics 666 Previously Hidden Markov Models for Relative Pairs Linkage analysis using affected sibling pairs Estimation of pairwise relationships Identity-by-Descent Relatives
Parts 2. Modeling chromosome segregation
 Genome 371, Autumn 2017 Quiz Section 2 Meiosis Goals: To increase your familiarity with the molecular control of meiosis, outcomes of meiosis, and the important role of crossing over in generating genetic
Genome 371, Autumn 2017 Quiz Section 2 Meiosis Goals: To increase your familiarity with the molecular control of meiosis, outcomes of meiosis, and the important role of crossing over in generating genetic
Chapter 6 Linkage Disequilibrium & Gene Mapping (Recombination)
 12/5/14 Chapter 6 Linkage Disequilibrium & Gene Mapping (Recombination) Linkage Disequilibrium Genealogical Interpretation of LD Association Mapping 1 Linkage and Recombination v linkage equilibrium ²
12/5/14 Chapter 6 Linkage Disequilibrium & Gene Mapping (Recombination) Linkage Disequilibrium Genealogical Interpretation of LD Association Mapping 1 Linkage and Recombination v linkage equilibrium ²
Genome 541! Unit 4, lecture 3! Genomics assays
 Genome 541! Unit 4, lecture 3! Genomics assays Much easier to follow with slides. Good pace.! Having the slides was really helpful clearer to read and easier to follow the trajectory of the lecture.!!
Genome 541! Unit 4, lecture 3! Genomics assays Much easier to follow with slides. Good pace.! Having the slides was really helpful clearer to read and easier to follow the trajectory of the lecture.!!
Inferring Genetic Architecture of Complex Biological Processes
 Inferring Genetic Architecture of Complex Biological Processes BioPharmaceutical Technology Center Institute (BTCI) Brian S. Yandell University of Wisconsin-Madison http://www.stat.wisc.edu/~yandell/statgen
Inferring Genetic Architecture of Complex Biological Processes BioPharmaceutical Technology Center Institute (BTCI) Brian S. Yandell University of Wisconsin-Madison http://www.stat.wisc.edu/~yandell/statgen
Related Courses He who asks is a fool for five minutes, but he who does not ask remains a fool forever.
 CSE 527 Computational Biology http://www.cs.washington.edu/527 Lecture 1: Overview & Bio Review Autumn 2004 Larry Ruzzo Related Courses He who asks is a fool for five minutes, but he who does not ask remains
CSE 527 Computational Biology http://www.cs.washington.edu/527 Lecture 1: Overview & Bio Review Autumn 2004 Larry Ruzzo Related Courses He who asks is a fool for five minutes, but he who does not ask remains
Probability of Detecting Disease-Associated SNPs in Case-Control Genome-Wide Association Studies
 Probability of Detecting Disease-Associated SNPs in Case-Control Genome-Wide Association Studies Ruth Pfeiffer, Ph.D. Mitchell Gail Biostatistics Branch Division of Cancer Epidemiology&Genetics National
Probability of Detecting Disease-Associated SNPs in Case-Control Genome-Wide Association Studies Ruth Pfeiffer, Ph.D. Mitchell Gail Biostatistics Branch Division of Cancer Epidemiology&Genetics National
Binomial Mixture Model-based Association Tests under Genetic Heterogeneity
 Binomial Mixture Model-based Association Tests under Genetic Heterogeneity Hui Zhou, Wei Pan Division of Biostatistics, School of Public Health, University of Minnesota, Minneapolis, MN 55455 April 30,
Binomial Mixture Model-based Association Tests under Genetic Heterogeneity Hui Zhou, Wei Pan Division of Biostatistics, School of Public Health, University of Minnesota, Minneapolis, MN 55455 April 30,
MULTIPLE CHOICE- Select the best answer and write its letter in the space provided.
 Form 1 Key Biol 1400 Quiz 4 (25 pts) RUE-FALSE: If you support the statement circle for true; if you reject the statement circle F for false. F F 1. A bacterial plasmid made of prokaryotic DNA can NO attach
Form 1 Key Biol 1400 Quiz 4 (25 pts) RUE-FALSE: If you support the statement circle for true; if you reject the statement circle F for false. F F 1. A bacterial plasmid made of prokaryotic DNA can NO attach
Proteomics Systems Biology
 Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Network Motifs of Pathogenic Genes in Human Regulatory Network
 Network Motifs of Pathogenic Genes in Human Regulatory Network Michael Colavita Mentor: Soheil Feizi Fourth Annual MIT PRIMES Conference May 18, 2014 Topics Background Genetics Regulatory Networks The
Network Motifs of Pathogenic Genes in Human Regulatory Network Michael Colavita Mentor: Soheil Feizi Fourth Annual MIT PRIMES Conference May 18, 2014 Topics Background Genetics Regulatory Networks The
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00.
 Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
Quantile-based permutation thresholds for QTL hotspot analysis: a tutorial
 Quantile-based permutation thresholds for QTL hotspot analysis: a tutorial Elias Chaibub Neto and Brian S Yandell September 18, 2013 1 Motivation QTL hotspots, groups of traits co-mapping to the same genomic
Quantile-based permutation thresholds for QTL hotspot analysis: a tutorial Elias Chaibub Neto and Brian S Yandell September 18, 2013 1 Motivation QTL hotspots, groups of traits co-mapping to the same genomic
GSBHSRSBRSRRk IZTI/^Q. LlML. I Iv^O IV I I I FROM GENES TO GENOMES ^^^H*" ^^^^J*^ ill! BQPIP. illt. goidbkc. itip31. li4»twlil FIFTH EDITION
 FIFTH EDITION IV I ^HHk ^ttm IZTI/^Q i I II MPHBBMWBBIHB '-llwmpbi^hbwm^^pfc ' GSBHSRSBRSRRk LlML I I \l 1MB ^HP'^^MMMP" jflp^^^^^^^^st I Iv^O FROM GENES TO GENOMES %^MiM^PM^^MWi99Mi$9i0^^ ^^^^^^^^^^^^^V^^^fii^^t^i^^^^^
FIFTH EDITION IV I ^HHk ^ttm IZTI/^Q i I II MPHBBMWBBIHB '-llwmpbi^hbwm^^pfc ' GSBHSRSBRSRRk LlML I I \l 1MB ^HP'^^MMMP" jflp^^^^^^^^st I Iv^O FROM GENES TO GENOMES %^MiM^PM^^MWi99Mi$9i0^^ ^^^^^^^^^^^^^V^^^fii^^t^i^^^^^
G E INTERACTION USING JMP: AN OVERVIEW
 G E INTERACTION USING JMP: AN OVERVIEW Sukanta Dash I.A.S.R.I., Library Avenue, New Delhi-110012 sukanta@iasri.res.in 1. Introduction Genotype Environment interaction (G E) is a common phenomenon in agricultural
G E INTERACTION USING JMP: AN OVERVIEW Sukanta Dash I.A.S.R.I., Library Avenue, New Delhi-110012 sukanta@iasri.res.in 1. Introduction Genotype Environment interaction (G E) is a common phenomenon in agricultural
1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms.
 Practicing Biology BIG IDEA 3.A 1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms. 2. Using at least 2 well-known experiments, describe which features of DNA and RNA
Practicing Biology BIG IDEA 3.A 1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms. 2. Using at least 2 well-known experiments, describe which features of DNA and RNA
Package KMgene. November 22, 2017
 Type Package Package KMgene November 22, 2017 Title Gene-Based Association Analysis for Complex Traits Version 1.2 Author Qi Yan Maintainer Qi Yan Gene based association test between a
Type Package Package KMgene November 22, 2017 Title Gene-Based Association Analysis for Complex Traits Version 1.2 Author Qi Yan Maintainer Qi Yan Gene based association test between a
Quantitative characters
 Quantitative characters Joe Felsenstein GENOME 453, Autumn 015 Quantitative characters p.1/38 A random mating population with two genes having alleles each, at equal frequencies, symmetrically affecting
Quantitative characters Joe Felsenstein GENOME 453, Autumn 015 Quantitative characters p.1/38 A random mating population with two genes having alleles each, at equal frequencies, symmetrically affecting
The phenotype of this worm is wild type. When both genes are mutant: The phenotype of this worm is double mutant Dpy and Unc phenotype.
 Series 2: Cross Diagrams - Complementation There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Series 2: Cross Diagrams - Complementation There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
The E-M Algorithm in Genetics. Biostatistics 666 Lecture 8
 The E-M Algorithm in Genetics Biostatistics 666 Lecture 8 Maximum Likelihood Estimation of Allele Frequencies Find parameter estimates which make observed data most likely General approach, as long as
The E-M Algorithm in Genetics Biostatistics 666 Lecture 8 Maximum Likelihood Estimation of Allele Frequencies Find parameter estimates which make observed data most likely General approach, as long as
Parts 2. Modeling chromosome segregation
 Genome 371, Autumn 2018 Quiz Section 2 Meiosis Goals: To increase your familiarity with the molecular control of meiosis, outcomes of meiosis, and the important role of crossing over in generating genetic
Genome 371, Autumn 2018 Quiz Section 2 Meiosis Goals: To increase your familiarity with the molecular control of meiosis, outcomes of meiosis, and the important role of crossing over in generating genetic
Quantitative characters
 Quantitative characters Joe Felsenstein GENOME 453, Autumn 013 Quantitative characters p.1/38 A random mating population with two genes having alleles each, at equal frequencies, symmetrically affecting
Quantitative characters Joe Felsenstein GENOME 453, Autumn 013 Quantitative characters p.1/38 A random mating population with two genes having alleles each, at equal frequencies, symmetrically affecting
Test for interactions between a genetic marker set and environment in generalized linear models Supplementary Materials
 Biostatistics (2013), pp. 1 31 doi:10.1093/biostatistics/kxt006 Test for interactions between a genetic marker set and environment in generalized linear models Supplementary Materials XINYI LIN, SEUNGGUEN
Biostatistics (2013), pp. 1 31 doi:10.1093/biostatistics/kxt006 Test for interactions between a genetic marker set and environment in generalized linear models Supplementary Materials XINYI LIN, SEUNGGUEN
Affected Sibling Pairs. Biostatistics 666
 Affected Sibling airs Biostatistics 666 Today Discussion of linkage analysis using affected sibling pairs Our exploration will include several components we have seen before: A simple disease model IBD
Affected Sibling airs Biostatistics 666 Today Discussion of linkage analysis using affected sibling pairs Our exploration will include several components we have seen before: A simple disease model IBD
Androgen-independent prostate cancer
 The following tutorial walks through the identification of biological themes in a microarray dataset examining androgen-independent. Visit the GeneSifter Data Center (www.genesifter.net/web/datacenter.html)
The following tutorial walks through the identification of biological themes in a microarray dataset examining androgen-independent. Visit the GeneSifter Data Center (www.genesifter.net/web/datacenter.html)
Solutions to Problem Set 4
 Question 1 Solutions to 7.014 Problem Set 4 Because you have not read much scientific literature, you decide to study the genetics of garden peas. You have two pure breeding pea strains. One that is tall
Question 1 Solutions to 7.014 Problem Set 4 Because you have not read much scientific literature, you decide to study the genetics of garden peas. You have two pure breeding pea strains. One that is tall
STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK. COURSE OUTLINE BIOL 310 The Human Genome. Prepared By: Ron Tavernier
 STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK COURSE OUTLINE BIOL 310 The Human Genome Prepared By: Ron Tavernier SCHOOL OF SCIENCE, HEALTH & CRIMINAL JUSTICE SCIENCE DEPARTMENT May
STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK COURSE OUTLINE BIOL 310 The Human Genome Prepared By: Ron Tavernier SCHOOL OF SCIENCE, HEALTH & CRIMINAL JUSTICE SCIENCE DEPARTMENT May
Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence
 Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence www.plantcell.org/cgi/doi/10.1105/tpc.110.tt0110 Epigenetics Usually
Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence www.plantcell.org/cgi/doi/10.1105/tpc.110.tt0110 Epigenetics Usually
Sarah Djebali INRA - Toulouse - France Genetics, Physiology and Breeding Systems Laboratory
 Profiling the landscape of transcription, chromatin accessibility and chromosome conformation of cattle, pig, chicken and goat genomes [FAANG pilot project FR-AgENCODE ] Sarah Djebali INRA - Toulouse -
Profiling the landscape of transcription, chromatin accessibility and chromosome conformation of cattle, pig, chicken and goat genomes [FAANG pilot project FR-AgENCODE ] Sarah Djebali INRA - Toulouse -
Package ESPRESSO. August 29, 2013
 Package ESPRESSO August 29, 2013 Type Package Title Power Analysis and Sample Size Calculation Version 1.1 Date 2011-04-01 Author Amadou Gaye, Paul Burton Maintainer Amadou Gaye The package
Package ESPRESSO August 29, 2013 Type Package Title Power Analysis and Sample Size Calculation Version 1.1 Date 2011-04-01 Author Amadou Gaye, Paul Burton Maintainer Amadou Gaye The package
Principles of QTL Mapping. M.Imtiaz
 Principles of QTL Mapping M.Imtiaz Introduction Definitions of terminology Reasons for QTL mapping Principles of QTL mapping Requirements For QTL Mapping Demonstration with experimental data Merit of QTL
Principles of QTL Mapping M.Imtiaz Introduction Definitions of terminology Reasons for QTL mapping Principles of QTL mapping Requirements For QTL Mapping Demonstration with experimental data Merit of QTL
arxiv: v1 [stat.ap] 4 Apr 2017
![arxiv: v1 [stat.ap] 4 Apr 2017 arxiv: v1 [stat.ap] 4 Apr 2017](/thumbs/72/66809022.jpg) Bayesian Model Averaging for the X-Chromosome Inactivation Dilemma in Genetic Association Study arxiv:1704.01207v1 [stat.ap] 4 Apr 2017 Bo Chen, Radu V. Craiu and Lei Sun Department of Statistical Sciences
Bayesian Model Averaging for the X-Chromosome Inactivation Dilemma in Genetic Association Study arxiv:1704.01207v1 [stat.ap] 4 Apr 2017 Bo Chen, Radu V. Craiu and Lei Sun Department of Statistical Sciences
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Single nucleotide variants in transcription factors associate more tightly with phenotype than with gene expression
 Washington University School of Medicine Digital Commons@Becker Open Access Publications 2014 Single nucleotide variants in transcription factors associate more tightly with phenotype than with gene expression
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2014 Single nucleotide variants in transcription factors associate more tightly with phenotype than with gene expression
Genetic polymorphism at the KIR gene locus: determination of gene, genotype, and haplotype frequencies in the Xinjiang Han population
 Genetic polymorphism at the KIR gene locus: determination of gene, genotype, and haplotype frequencies in the Xinjiang Han population G.-Y. Lin 1, B. Yu 2, W.-J. Hu 1, Y.-Z. Zhang 1, X.-J. Zuo 1 and Y.-B.
Genetic polymorphism at the KIR gene locus: determination of gene, genotype, and haplotype frequencies in the Xinjiang Han population G.-Y. Lin 1, B. Yu 2, W.-J. Hu 1, Y.-Z. Zhang 1, X.-J. Zuo 1 and Y.-B.
Computational Approaches to Statistical Genetics
 Computational Approaches to Statistical Genetics GWAS I: Concepts and Probability Theory Christoph Lippert Dr. Oliver Stegle Prof. Dr. Karsten Borgwardt Max-Planck-Institutes Tübingen, Germany Tübingen
Computational Approaches to Statistical Genetics GWAS I: Concepts and Probability Theory Christoph Lippert Dr. Oliver Stegle Prof. Dr. Karsten Borgwardt Max-Planck-Institutes Tübingen, Germany Tübingen
VCE BIOLOGY Relationship between the key knowledge and key skills of the Study Design and the Study Design
 VCE BIOLOGY 2006 2014 Relationship between the key knowledge and key skills of the 2000 2005 Study Design and the 2006 2014 Study Design The following table provides a comparison of the key knowledge (and
VCE BIOLOGY 2006 2014 Relationship between the key knowledge and key skills of the 2000 2005 Study Design and the 2006 2014 Study Design The following table provides a comparison of the key knowledge (and
Multi-trait analysis of genome-wide association summary statistics using MTAG
 SUPPLEMENTARY INFORMATION Articles https://doi.org/0.038/s4588-07-0009-4 In the format provided by the authors and unedited. Multi-trait analysis of genome-wide association summary statistics using MTAG
SUPPLEMENTARY INFORMATION Articles https://doi.org/0.038/s4588-07-0009-4 In the format provided by the authors and unedited. Multi-trait analysis of genome-wide association summary statistics using MTAG
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Statistical Genetics I: STAT/BIOST 550 Spring Quarter, 2014
 Overview - 1 Statistical Genetics I: STAT/BIOST 550 Spring Quarter, 2014 Elizabeth Thompson University of Washington Seattle, WA, USA MWF 8:30-9:20; THO 211 Web page: www.stat.washington.edu/ thompson/stat550/
Overview - 1 Statistical Genetics I: STAT/BIOST 550 Spring Quarter, 2014 Elizabeth Thompson University of Washington Seattle, WA, USA MWF 8:30-9:20; THO 211 Web page: www.stat.washington.edu/ thompson/stat550/
Incorporating Functional Annotations for Fine-Mapping Causal Variants in a. Bayesian Framework using Summary Statistics
 Genetics: Early Online, published on September 21, 2016 as 10.1534/genetics.116.188953 1 Incorporating Functional Annotations for Fine-Mapping Causal Variants in a Bayesian Framework using Summary Statistics
Genetics: Early Online, published on September 21, 2016 as 10.1534/genetics.116.188953 1 Incorporating Functional Annotations for Fine-Mapping Causal Variants in a Bayesian Framework using Summary Statistics
Science Department-High School
 Science Department-High School Course Description SUBJECT: Biology Course Title: HEREDITY Grade Level: 12 Course Number: Bio II NUMBER OF CREDITS: 1 Reference Book and online resources: Holt McDougal MICHIGAN
Science Department-High School Course Description SUBJECT: Biology Course Title: HEREDITY Grade Level: 12 Course Number: Bio II NUMBER OF CREDITS: 1 Reference Book and online resources: Holt McDougal MICHIGAN
Supplemental material
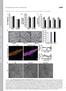 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Variance Components: Phenotypic, Environmental and Genetic
 Variance Components: Phenotypic, Environmental and Genetic You should keep in mind that the Simplified Model for Polygenic Traits presented above is very simplified. In many cases, polygenic or quantitative
Variance Components: Phenotypic, Environmental and Genetic You should keep in mind that the Simplified Model for Polygenic Traits presented above is very simplified. In many cases, polygenic or quantitative
Association studies and regression
 Association studies and regression CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar Association studies and regression 1 / 104 Administration
Association studies and regression CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar Association studies and regression 1 / 104 Administration
Intro Gene regulation Synteny The End. Today. Gene regulation Synteny Good bye!
 Today Gene regulation Synteny Good bye! Gene regulation What governs gene transcription? Genes active under different circumstances. Gene regulation What governs gene transcription? Genes active under
Today Gene regulation Synteny Good bye! Gene regulation What governs gene transcription? Genes active under different circumstances. Gene regulation What governs gene transcription? Genes active under
. Also, in this case, p i = N1 ) T, (2) where. I γ C N(N 2 2 F + N1 2 Q)
 Supplementary information S7 Testing for association at imputed SPs puted SPs Score tests A Score Test needs calculations of the observed data score and information matrix only under the null hypothesis,
Supplementary information S7 Testing for association at imputed SPs puted SPs Score tests A Score Test needs calculations of the observed data score and information matrix only under the null hypothesis,
Figure S2. The distribution of the sizes (in bp) of syntenic regions of humans and chimpanzees on human chromosome 21.
 Frequency 0 1000 2000 3000 4000 5000 0 2 4 6 8 10 Distance Figure S1. The distribution of human-chimpanzee sequence divergence for syntenic regions of humans and chimpanzees on human chromosome 21. Distance
Frequency 0 1000 2000 3000 4000 5000 0 2 4 6 8 10 Distance Figure S1. The distribution of human-chimpanzee sequence divergence for syntenic regions of humans and chimpanzees on human chromosome 21. Distance
