Sparse Factorizations of Gene Expression Guided by Binding Data. L. Badea and D. Tilivea. Pacific Symposium on Biocomputing 10: (2005)
|
|
|
- Preston Silas Gardner
- 5 years ago
- Views:
Transcription
1 Sparse actorizations of Gene Expression Guided by Binding Data L. Badea and D. ilivea Pacific Symposium on Biocomputing 10: (005)
2 SPARSE ACORIZAIONS O GENE EXPRESSION DAA GUIDED BY BINDING DAA LIVIU BADEA and DOINA ILIVEA AI Lab, National Institute for Research and Development in Informatics 8-10 Averescu Blvd., Bucharest, Romania, badea@ici.ro Existing clustering methods do not deal well with overlapping clusters, are unstable and do not take into account the robustness of biological systems, or more complex background knowledge such as regulator binding data. Here we describe a nonnegative sparse factorization algorithm dealing with the above problems: cluster overlaps are allowed by design, the nonnegativity constraints implicitly approximate the robustness of biological systems and regulator binding data is used to guide the factorization. Preliminary results show the feasibility of our approach. 1 Introduction and motivation he advent of microarray technology has allowed a revolutionary transition from the exploration of the expression of a handful of genes to that of entire genomes. However, despite its enormous potential, microarray data has proved difficult to analyze, partly due to the significant amount of noise, but also due to the large number of factors that influence gene expression (many of which are not at the mrna/transcriptone level) and the complexity of their interactions. One of the most successful microarray data analysis methods has proved to be clustering (of genes and/or samples), and a large variety of such methods have been proposed and applied to real-life biological data. his large body of work, impossible to extensively review here, has emphasized important limitations of existing clustering algorithms: (1) Most clustering methods produce non-overlapping clusters. However, since genes are typically involved in several biological processes, non-overlapping clustering methods, such as hierarchical clustering (HC) [], self-organizing maps (SOM) [1], k-means clustering, etc., tend to be unstable, producing different gene clusters for only slightly different input samples (e.g. in the case of HC), or depending on the choice of initial conditions (as in the case of SOM [5], or k- means). Algorithms allowing for overlapping clusters, such as fuzzy k-means [4] achieved significant improvements w.r.t. non-overlapping clustering, but they still have the problems discussed below. () Most algorithms perform clustering along a single dimension comparing e.g. genes w.r.t. all the available samples, whereas in reality genes have coordinated expression levels only for certain subsets of conditions. Algorithms dealing with this problem, such as biclustering [13], coupled-two way clustering (CWC) [3], ISA (iterative signature) [1] have other problems mostly related to the control of overlap between biclusters.
3 (3) Although genes are subject to both positive and negative influences from other genes, the robustness of biological systems requires that an observed change in the expression level of a given gene is the result of either a positive or a negative influence rather than a complex combination of positive and negative influences that partly cancel out each other (as in the case of Principal Component Analysis). Nonnegative Matrix actorization (NM) [9] deals with this problem by searching for nonnegative decompositions of (nonnegative) data. he observed localized nature of the decompositions seems to be a biproduct of the nonnegativity constraints [9]. Recently, Brunet at al [5] applied NM for clustering samples in a nonoverlapping mode for three cancer datasets. While oligonucleotide arrays used in that work produce positive data, which lend themselves naturally to nonnegative decompositions, clustering genes in an analogous manner would ignore potential negative influences (i.e. genes downregulating other genes). On the other hand, Kim and idor [6] used NM to cluster genes in the context of a large dataset of yeast perturbation experiments (spotted arrays) [7]. Although NM has the tendency of producing sparse representations, the factorizations obtained were subjected to thresholding and subsequent reoptimization to obtain sufficiently sparse clusters. Unfortunately however, microarray data is noisy and it might be useful to be able to take into account any background knowledge that may be available. or example, Lee et al [8] have published binding (location analysis) data for a large number (106) of transcription factors in the yeast S. Cerevisiae. Elsewhere [in preparation] we have observed that transcription factor () expression levels are not always good predictors for the expression levels of their targets. (he presence of the transcription factor is of course required for the target to be expressed, but very frequently the is activated by a different signaling molecule, e.g. a kinase.) herefore using the binding data directly as background knowledge may not be very helpful in practice. In fact, it seems that although s are not well correlated with their targets, the targets themselves seem to be much better correlated among each other. In the following, we show how binding data can be used as background knowledge in a novel nonnegative factorization algorithm NNSC B (Nonnegative Sparse Coding with Background Knowledge) designed to address the abovementioned problems of existing clustering algorithms. Our algorithm is an improvement of the nonnegative sparse coding (NNSC) algorithm of Hoyer [11]. It produces overlapping clusters that are much more stable than those generated with other algorithms, while also being able to take background knowledge into account.
4 he data sources Since the most extensive background knowledge is available for the yeast S. Cerevisiae, in this paper we use the Rosetta Compendium [7], the largest publicly available gene expression dataset for yeast perturbations, as well as the binding (location analysis) data of Lee et al. [8]. he Rosetta Compendium contains expression profiles over virtually all yeast genes (6315 ORs) corresponding to 300 diverse mutations and chemical treatments (76 deletion mutants, 11 tetracycline-regulatable alleles of essential genes and treatments with 13 well-characterized compounds) of S. Cerevisiae grown under a single (normal) condition. he data contains log-expression ratios g( tf ) log10 r = log, where g(tf ) and g(w) are the mrna concentrations of gene 10 g( W) g in the tf mutant and the wild type respectively. he location analysis data of Lee et al. contains information about binding of 106 transcriptional regulators to upstream regions of target genes. Since log-ratios can be negative, we cannot directly apply a nonnegative factorization algorithm to the log-ratio dataset. On the other hand, although the ratios r (or maybe r-1) are nonnegative, applying nonnegative factorization on them would only uncover the positive influences, while in practice the low level of certain genes is due to them being downregulated by other genes. o address problem (3) mentioned in the Introduction, we separate, as in [6], the up-regulated from the down-regulated part of each gene, i.e. obtain two entries g + and g for each gene g from the original gene expression matrix: + r ( g ) 1 r ( g ) 1 1 r ( g ) = 1 r ( g ) < 1 r ( g ) = 0 otherwise r ( g ) 0 otherwise Note that in this representation, a significantly downregulated gene will require a non-negligible contribution in the factorization. (We use the ratios rather than logratios as in [6], since linear combinations of log-ratios amount to products of powers of ratios rather than additive contributions.) 3 Nonnegative Sparse Coding Hoyer s NNSC algorithm [11] factorizes a n s n g matrix X A S as a product of an n s n c matrix A and a n c n g matrix S by optimizing (minimizing) the following objective function: 1 1 o achieve information compression, the number of internal dimensions n c must verify the constraint n c (n s +n g ) < n s n g.
5 1 C ( A, S ) = X AS + λ S (1) c, g with respect to the nonnegativity constraints A sc 0, S 0 () he objective function combines a fitness term involving the robenius norm of the error and a size term penalizing the non-zero entries of S. (he robenius norm is given by E = E = r ( E E ). ) s, g sg he Nonnegative Matrix actorization (NM) of Lee and Seung [10] is recovered by setting the size parameter λ to zero, while a non-zero λ would lead to sparser factorizations. he objective function (1) above has an important problem, due to the invariance of the fitness term under diagonal scalings. More precisely we have the following result. Proposition. he fitness term 1 X AS (i.e. the NM objective function) is invariant under the following transformations: A A D -1, S D S, (3) where D = diag(d 1,,d nc ) is a positive diagonal matrix (d c > 0). Note that such positive diagonal matrices are the most general positive matrices whose inverses are also positive (thereby preserving the nonnegativity of A and S under the above transformation). he scaling invariance of the fitness term in (1) makes the size term ineffective, since the latter can be forced as small as needed simply by using a diagonal scaling D with small enough entries. Additional constraints are therefore needed to render the size term operational. Since a diagonal matrix D operates on the rows of S and on the columns of A, we could impose unit norms either for the rows of S, or for the columns of A. Unfortunately, the objective function (1) used in [11] has an important flaw: it produces decompositions that depend on the scale of the original matrix X (i.e. the decompositions of X and αx are essentially different), regardless of the normalization scheme employed. or example, if we constrain the rows of S to unit norm, then we cannot have decompositions of the form X A S and αx αa S, since at least one of these is in general non-optimal due to the dimensional inhomogeneity of the objective function w.r.t. A and X 1 C α X ( α A, S ) = α X AS + λ S. On the other hand, if we constrain the columns of A to unit norm, the decompositions X A S and αx A αs cannot be both optimal, again due to the dimensional inhomogeneity of C, now w.r.t. S and X: c, g
6 1 C αx ( A, αs ) = α X AS + αλ S c, g herefore, as long as the size term depends only on S, we are forced to constrain the columns of A to unit norm, while employing an objective function that is dimensionally homogeneous in S and X. One such dimensionally homogeneous objective function is: 1 C ( A, S ) = X AS + λ S (1 ) which will be minimized w.r.t. the nonnegativity constraints () and the constraints on the norm of the columns of A: A c = 1 (i.e. s A sc = 1 ) (4) It can be easily verified that this produces scale independent decompositions, i.e. if X A S is an optimal decomposition of X, then αx A αs is an optimal decomposition of αx. he constrained optimization problem could be solved with a gradient-based method. However, in the case of NM, faster so-called multiplicative update rules exist [10,11], which we have generalized to the NNSC problem as follows. (hese methods only produce local minima, but the solutions tend to be quite stable see also Section 5 below.) Modified NNSC algorithm Start with random initial matrices A and S loop ( A X ) S S ( A A S + λs ) A A + µ ( X A S ) S normalize the columns of A to unit norm: until convergence. 1 A A D, D = diag( A ) s sc In the following, we assume that the gene expression data is given in an n s n g matrix X, where n s and n g are the numbers of samples and genes respectively, so that X sg represents the expression level of gene g in sample s. A sparse factorization X A S will be interpreted as a generalization of clustering the genes (i.e. the columns X g of X) into overlapping clusters c corresponding to the rows S of S. More precisely, a non-zero value of S (or at least an S larger than a given threshold) will be interpreted as the gene g belonging to cluster c. Note that clusters can be overlapping, since the columns of S may have several significant entries. Although overlaps are allowed, NNSC will not produce highly overlapping clusters, due to the sparseness constraints. his is unlike many other clustering
7 algorithms that allow clusters to overlap, which have to resort to several parameters to keep excessive cluster overlap under control.) Also note that we factorize X rather than X since the sparsity constraint should affect the clusters of genes (i.e. S) rather than the clusters of samples A. (his is unlike NM, for which the factorizations of X and of X are completely symmetrical.) 4 Nonnegative Sparse Coding using Background Knowledge ranscription factor binding data can be represented by a n f n g Boolean matrix B, such that B fg =1 iff the transcription factor f binds to the upstream region of gene g. As already mentioned in the Introduction, although transcription factor expression levels are not always good predictors of the expression levels of their targets, the targets are frequently much better correlated among themselves. his suggests using the co-occurrence matrix K=B B rather than B as background knowledge. K is an n g n g square matrix, in which K g g 0 iff genes g and g are both targets of some common transcription factor f. Our idea of exploiting background knowledge during clustering is quite simple. Normally, non-zero entries in the gene cluster matrix S are penalized for size. However, if certain entries conform to the background knowledge, they will be exempted from size penalization. We thus need to modify the size term in C(A,S) to take into account B. Implementing this simple idea involves however certain subtleties. Assume that some gene cluster c (i.e. set of genes g for which S 0, or at least S > for some given threshold ) contains many genes that are targets of several s (e.g. igure 1b below). Although this cluster is preferable from the point of view of the background knowledge to the one from igure 1a, it is worse than the one from igure 1c, in which all the genes are the targets of a single. tf 1 tf tf 3 tf 4 tf 1 tf tf g 1 g g 3 g 4 g 1 g g 3 g 4 g 1 g g 3 g 4 (a) (b) (c) igure 1. Conformance of clusters to the binding data: (c) is preferable to (b), which is preferable to (a). hus, the size term cannot be simply a sum of overlaps of the clusters (i.e. rows of S) with groups of -targets (rows of B), since S = ( )( ) c f g B fg S g c B f fg does not depend on the way the genes are distributed in groups of -targets for different s.
8 Clusters like the one in igure 1c can be highly evaluated if cross-terms between genes controlled by the same are added. We thus encourage genes controlled by the same in the binding data, while penalizing the size of S using an objective function of the form: 1 λ C ( A, S ) = X AS + ( S r ( SKS )) (5) with K = γ ( σ ( B B) diag ( σ ( B ) ), where γ is a factor such that f fg 1 K ' '' = 1 and σ( ) is the Heaviside step function (applied element-wise). g g n g g ', g '' Of course, optimization of (5) is attempted in the context of the constraints () and (4). he algorithm below solves the above optimization problem using combined multiplicative and additive update rules. (he final normalization of the rows of S renders the resulting clusters comparable.) NNSC B algorithm start with random initial matrices A and S loop ( A X + λs K ) S S ( A A S + λs ) A A + µ ( X A S ) S normalize the columns of A to unit norm: until convergence S D 1 S and A A D 1 A A D, D = diag( A ) s sc, where D = diag( S ) o test our approach, we have applied the NNSC B algorithm on a synthetic dataset with several highly overlapping clusters. NNSC B has been able to consistently recover the clusters, or close approximations thereof even in the presence of noise. (See for more details.) g 5 Clustering the Rosetta dataset w.r.t. the binding data of Lee et al. Although the main goal of this paper is the presentation of a new clustering algorithm able to deal with background knowledge rather than obtaining new biological insights, we also briefly discuss our initial attempts at applying our algorithm to yeast microarray data. he binding data of Lee et al. contains the targets of 106 transcription factors, roughly about half the total number of yeast transcription factors. In order not to introduce a bias towards the targets of these s due to the background knowledge,
9 we have selected from the Rosetta dataset only these targets. (We also eliminated the genes that had unreliable measurements in the Rosetta dataset dealing with missing values in our context is a matter of future work.) his left us with a set of 99 s and 099 genes. he matrix X to be factorized was constructed by duplicating genes as described in Section (X has therefore 4198 columns). Duplicating genes g into their positive (g + ) and negative parts (g ) may raise potential problems with possible conflicts between nonzero S + and S entries, as a gene cannot be both up- and down regulated in a given cluster. he fact that our decompositions never have both S + and S nonzero (significant) shows that the approach is biologically sensible. An important parameter of the NNSC B factorization is its internal dimensionality (the number of clusters n c ). A useful estimate of the internal dimensionality of a dataset can be obtained from its singular value decomposition (SVD). A more refined analysis [6] determines the number of dimensions around which the root mean square error (RMSE) change of the real data and that of a randomized dataset become equal. Kim and idor s analysis estimated the internal dimensionality of the Rosetta dataset to be around 50. We performed this analysis for our restricted dataset and obtained a similar dimensionality around 50 (see igure ) relative error randomized data original data internal dimensionality (number of clusters) igure. Determining the internal dimensionality of the Rosetta dataset using the method of Kim and idor We also considered another approach: for a given set of clusters we determined the fraction of intra-cluster correlated pairs of genes (i.e. the number of correlated pairs of genes divided by the total number of correlated pairs of genes). his fraction was estimated for various correlation thresholds r 0. (More precisely, a gene pair (g 1,g ) is called correlated w.r.t. threshold r 0 iff r(g 1,g ) r 0.) igure 3 depicts
10 the r 0 -dependence of the fraction of intra-cluster correlated pairs of genes for various dimensionalities. Notice that n c =50 is a reasonable choice as the above mentioned fraction approaches 90% for a large range of r 0. (his means in other words that most of the correlated gene pairs are within clusters.) igure 3. he correlation-threshold dependence of the fraction of intra-cluster correlated gene pairs. Next we studied the stability of the algorithm w.r.t. the initial starting point. (he algorithm was run disregarding the background knowledge, i.e. taking λ=0, in order to avoid any possible influences of the background knowledge on the stability of some solution fragment.) he able below lists the relative errors ( X-AS / X ) obtained in 7 runs for 7 different initializations. run no relative error o asses the stability of the decomposition in the case of k runs, we determined the best matches among the k sets of clusters. he following able shows the numbers of matching clusters for a progressively larger number of runs. Note the (relative) stabilization of the number of matching clusters w.r.t. the numbers of runs: Numbers of runs Matching clusters (avg.) Next, we studied the influence of the background knowledge on the solution. In order to separate the variability of the results due to the initialization from that due
11 to the background knowledge, we performed all the following tests using the same initialization. We ran NNSC B with several values of the parameter λ, ranging from 0 (background knowledge is not taken into account) to 0.75 (background knowledge has comparable weight to the data fit term) and tested the overlap of the resulting clusters with the background knowledge. Briefly, we observed a clear increase of this overlap with larger λ. More precisely (more details can be found in the able below): - the average fraction of cluster genes controlled by s increased from about 47% for λ=0 to 66% for λ=0.75. (Only the s controlling at least two genes from the cluster were counted.) - the average number of s per cluster controlling at least two genes increased from around 8 for λ=0 to 14 for λ=0.75. λ avg. number of genes in a cluster avg. fraction of cluster genes controlled by s avg. number of s per cluster avg. cluster overlap avg. cluster overlap (when only overlapping pairs are averaged) Relative error Of course, conformance to the data and/or to the background knowledge does not imply the biological relevance of the results. o estimate the latter, we searched for significant Gene Ontology (GO) annotations [14] of the genes in the clusters. More precisely, we employed the hypergeometric distribution to compute p-values representing likelihoods that specific GO annotations and a given cluster share a given number of genes by chance and retained only the annotations with a p-value less than (his p-value threshold was chosen so that not more than 1 or annotations are false positives, given that the genes in our dataset share 4 GO annotations some of which may of course be related by sub- or super-class relationships. We did not use a lower threshold in order to avoid many missing annotations.) o demonstrate the biological relevance of the factorizations using background knowledge, we performed alternative factorizations for randomized background knowledge (more precisely, by randomly permuting the lines of B independently of each other). his lead to a drop in the average number of significant GO annotations per cluster from 8.16 to 4.88 (for λ=0.75). We also looked at a few clusters in more detail. or example, cluster 47 had 18 genes, involved in the SE1 control of pheromone response, among which, for instance, AGA1, IG1, US1 are involved in cell fusion, while GPA1, US3 and PRR are involved in the pheromone signal transduction pathway, KAR4 is a regulatory protein required for pheromone induction of karyogamy genes and SS is involved in desensitization to alpha-factor pheromone. he mating a-factor genes MA1 and MA are also in the cluster. he entire cluster is presented in the
12 Annex, together with the associated significant GO annotations and the cluster coefficients from the S matrix. (he threshold used for extracting clusters from the S matrix was 1/ n g = ) 5 Conclusions Despite their wide-spread use in microarray data analysis, existing clustering algorithms have serious problems, the most important one being related to the fact that biological processes are overlapping rather than isolated. he impact of microarray technology is also limited by the noisy nature of measurements, which can only be compensated by additional background knowledge. Here we have shown how these important problems faced by microarray data analysis can be dealt with in the context of a sparse factorization algorithm capable of dealing with regulator binding data as background knowledge. A key ingredient of this algorithm is the nonnegativity constraint. Such an approach, for example using NM, has been mostly advocated in connection with oligonucleotide array data, which are (at least theoretically) nonnegative. However, this viewpoint is only partially correct, since downregulated genes would not be explained in such a framework. Actually, we argue that nonnegative factorizations are appropriate due to the robustness of biological systems, in which an observed change in a gene s expression level is the result of either a positive or a negative influence rather than a complex combination of the two. Although our preliminary results are encouraging, a more detailed biological analysis should be the focus of subsequent work. (he clusters obtained by our algorithm for various parameter settings can be found online at References 1. Bergmann S., J. Ihmels, N. Barkai. Iterative signature algorithm for the analysis of largescale gene expression data. Physical Review E 67, (003).. Eisen M.B., P.. Spellman, P.O. Brown, D. Botstein. Cluster analysis and display of genome-wide expression patterns, PNAS Vol.95, , Dec Getz G., Levine E., Domany E. Coupled two-way clustering analysis of gene microarray data, PNAS Vol. 97, , Gasch A.P, Eisen M.B. Exploring the conditional coregulation of yeast gene expression through fuzzy k-means clustering. Genome Biol. 00, Oct 10;3(11). 5. Brunet J.P., amayo P., Golub.R., Mesirov J.P. Metagenes and molecular pattern discovery using matrix factorization. PNAS 101(1):4164-9, 004, Mar Kim P.M., idor B. Subsystem identification through dimensionality reduction of largescale gene expression data. Genome Res. 003 Jul;13(7): Hughes.R. et al. unctional discovery via a compendium of expression profiles. Cell. 000 Jul 7;10(1):109-6.
13 8. Lee.I. et al. ranscriptional regulatory networks in Saccharomyces cerevisiae. Science. 00 Oct 5; 98(5594): Lee D.D., H.S. Seung. Learning the parts of objects by non-negative matrix factorization. Nature, vol. 401, no. 6755, pp , Lee D.D., H.S. Seung. Algorithms for non-negative matrix factorization. In Advances in Neural Information Processing 13 (Proc. NIPS*000), MI Press, P. O. Hoyer. Non-negative sparse coding. Neural Networks for Signal Processing XII, Martigny, Switzerland, amayo P. et al. Interpreting patterns of gene expression with self-organizing maps: methods and application to hematopoietic differentiation, PNAS 96(6):907-1 (1999). 13. Cheng Y, Church GM. Biclustering of expression data. Proc. ISMB 000; 8: Gene Ontology Consortium. Gene Ontology: tool for the unification of biology. Nature Genet. 5:5-9, 000. Annex. he 18 genes of cluster 47 and the s controlling them Significant GO annotations (with p-values): conjugation(0),conjugation with cellular fusion(0),sexual reproduction(0),reproduction(.30e-11),response to pheromone(4.96e-11),response to chemical substance(.4e-9),response to abiotic stimulus(.89e-9),development(1.07e-8),response to pheromone during conjugation with cellular fusion 1.13e-8),response to external stimulus(1.34e-8),cell communication(.51e-8),signal transduction during conjugation with cellular fusion(1.57e-6),response to stimulus(.11e-6),signal transduction(3.85e-6),g-protein coupled receptor protein signaling pathway(4.7e-6),cell surface receptor linked signal transduction(9.4e-6),shmoo tip(9.4e-6),signal transducer activity(4.4e-5),pheromone activity(1.97e-4),receptor binding(1.97e-4),cellular process(3.05e-4),receptor signaling protein serine/threonine kinase activity(3.9e-4),site of polarized growth(9.3e-4),site of polarized growth (sensu ungi)(9.3e-4),site of polarized growth (sensu Saccharomyces)(9.3e-4),receptor signaling protein activity(9.70e-4) SE1 targets: SE1, SS, EC1, US1, KAR4, GPA1, MA MCM1 targets: MA1, SE6, MA, AGA1 PHD1 targets: PRR, GPA1 +: : PRR strong similarity to putative protein kinase NPR1 +: : YDL133W +: : US1 cell fusion protein +: : PRY3 +: : IG1 required for efficient mating +: : AGA1 a-agglutinin anchor subunit +: : HSP1 heat shock protein +: : MA mating pheromone a-factor +: : GPA1 GP-binding protein alpha subunit of the pheromone pathway +: : questionable OR +: : SE6 AP-binding cassette transporter protein +: : SE1 transcriptional activator +: : US3 mitogen-activated protein kinase (MAP kinase) +: : MA1 mating pheromone a-factor 1 +: : SS involved in desensitization to alpha-factor pheromone +: : YLR343W +: : EC1 y transcription activator +: : KAR4 regulatory protein required for pheromone induction of karyogamy genes Parameters: λ=0.1, µ=5 10-6, S-threshold=0.0154, 300 iterations.
Inferring Transcriptional Regulatory Networks from Gene Expression Data II
 Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Modeling Gene Expression from Microarray Expression Data with State-Space Equations. F.X. Wu, W.J. Zhang, and A.J. Kusalik
 Modeling Gene Expression from Microarray Expression Data with State-Space Equations FX Wu, WJ Zhang, and AJ Kusalik Pacific Symposium on Biocomputing 9:581-592(2004) MODELING GENE EXPRESSION FROM MICROARRAY
Modeling Gene Expression from Microarray Expression Data with State-Space Equations FX Wu, WJ Zhang, and AJ Kusalik Pacific Symposium on Biocomputing 9:581-592(2004) MODELING GENE EXPRESSION FROM MICROARRAY
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Cluster Analysis of Gene Expression Microarray Data. BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002
 Cluster Analysis of Gene Expression Microarray Data BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002 1 Data representations Data are relative measurements log 2 ( red
Cluster Analysis of Gene Expression Microarray Data BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002 1 Data representations Data are relative measurements log 2 ( red
Non-Negative Factorization for Clustering of Microarray Data
 INT J COMPUT COMMUN, ISSN 1841-9836 9(1):16-23, February, 2014. Non-Negative Factorization for Clustering of Microarray Data L. Morgos Lucian Morgos Dept. of Electronics and Telecommunications Faculty
INT J COMPUT COMMUN, ISSN 1841-9836 9(1):16-23, February, 2014. Non-Negative Factorization for Clustering of Microarray Data L. Morgos Lucian Morgos Dept. of Electronics and Telecommunications Faculty
NON-NEGATIVE SPARSE CODING
 NON-NEGATIVE SPARSE CODING Patrik O. Hoyer Neural Networks Research Centre Helsinki University of Technology P.O. Box 9800, FIN-02015 HUT, Finland patrik.hoyer@hut.fi To appear in: Neural Networks for
NON-NEGATIVE SPARSE CODING Patrik O. Hoyer Neural Networks Research Centre Helsinki University of Technology P.O. Box 9800, FIN-02015 HUT, Finland patrik.hoyer@hut.fi To appear in: Neural Networks for
Topographic Independent Component Analysis of Gene Expression Time Series Data
 Topographic Independent Component Analysis of Gene Expression Time Series Data Sookjeong Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong,
Topographic Independent Component Analysis of Gene Expression Time Series Data Sookjeong Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong,
Fuzzy Clustering of Gene Expression Data
 Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
AN ENHANCED INITIALIZATION METHOD FOR NON-NEGATIVE MATRIX FACTORIZATION. Liyun Gong 1, Asoke K. Nandi 2,3 L69 3BX, UK; 3PH, UK;
 213 IEEE INERNAIONAL WORKSHOP ON MACHINE LEARNING FOR SIGNAL PROCESSING, SEP. 22 25, 213, SOUHAMPON, UK AN ENHANCED INIIALIZAION MEHOD FOR NON-NEGAIVE MARIX FACORIZAION Liyun Gong 1, Asoke K. Nandi 2,3
213 IEEE INERNAIONAL WORKSHOP ON MACHINE LEARNING FOR SIGNAL PROCESSING, SEP. 22 25, 213, SOUHAMPON, UK AN ENHANCED INIIALIZAION MEHOD FOR NON-NEGAIVE MARIX FACORIZAION Liyun Gong 1, Asoke K. Nandi 2,3
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Keywords: systems biology, microarrays, gene expression, clustering
 Jan H. Ihmels received his PhD in computational biology from the Weizmann Institute of Science, Israel. He is currently a postdoctoral fellow at the Department of Molecular Genetics of the Weizmann Institute.
Jan H. Ihmels received his PhD in computational biology from the Weizmann Institute of Science, Israel. He is currently a postdoctoral fellow at the Department of Molecular Genetics of the Weizmann Institute.
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
Exploratory statistical analysis of multi-species time course gene expression
 Exploratory statistical analysis of multi-species time course gene expression data Eng, Kevin H. University of Wisconsin, Department of Statistics 1300 University Avenue, Madison, WI 53706, USA. E-mail:
Exploratory statistical analysis of multi-species time course gene expression data Eng, Kevin H. University of Wisconsin, Department of Statistics 1300 University Avenue, Madison, WI 53706, USA. E-mail:
COMBINING LOCATION AND EXPRESSION DATA FOR PRINCIPLED DISCOVERY OF GENETIC REGULATORY NETWORK MODELS
 COMBINING LOCATION AND EXPRESSION DATA FOR PRINCIPLED DISCOVERY OF GENETIC REGULATORY NETWORK MODELS ALEXANDER J. HARTEMINK Duke University Department of Computer Science Box 90129, Durham, NC 27708-0129
COMBINING LOCATION AND EXPRESSION DATA FOR PRINCIPLED DISCOVERY OF GENETIC REGULATORY NETWORK MODELS ALEXANDER J. HARTEMINK Duke University Department of Computer Science Box 90129, Durham, NC 27708-0129
Prediction of double gene knockout measurements
 Prediction of double gene knockout measurements Sofia Kyriazopoulou-Panagiotopoulou sofiakp@stanford.edu December 12, 2008 Abstract One way to get an insight into the potential interaction between a pair
Prediction of double gene knockout measurements Sofia Kyriazopoulou-Panagiotopoulou sofiakp@stanford.edu December 12, 2008 Abstract One way to get an insight into the potential interaction between a pair
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data
 GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree
 Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree YOICHI YAMADA, YUKI MIYATA, MASANORI HIGASHIHARA*, KENJI SATOU Graduate School of Natural
Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree YOICHI YAMADA, YUKI MIYATA, MASANORI HIGASHIHARA*, KENJI SATOU Graduate School of Natural
networks in molecular biology Wolfgang Huber
 networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Bioinformatics. Transcriptome
 Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
Inferring Transcriptional Regulatory Networks from High-throughput Data
 Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling
 Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Discovering modules in expression profiles using a network
 Discovering modules in expression profiles using a network Igor Ulitsky 1 2 Protein-protein interactions (PPIs) Low throughput measurements: accurate, scarce High throughput: more abundant, noisy Large,
Discovering modules in expression profiles using a network Igor Ulitsky 1 2 Protein-protein interactions (PPIs) Low throughput measurements: accurate, scarce High throughput: more abundant, noisy Large,
EUSIPCO
 EUSIPCO 2013 1569741067 CLUSERING BY NON-NEGAIVE MARIX FACORIZAION WIH INDEPENDEN PRINCIPAL COMPONEN INIIALIZAION Liyun Gong 1, Asoke K. Nandi 2,3 1 Department of Electrical Engineering and Electronics,
EUSIPCO 2013 1569741067 CLUSERING BY NON-NEGAIVE MARIX FACORIZAION WIH INDEPENDEN PRINCIPAL COMPONEN INIIALIZAION Liyun Gong 1, Asoke K. Nandi 2,3 1 Department of Electrical Engineering and Electronics,
On Spectral Basis Selection for Single Channel Polyphonic Music Separation
 On Spectral Basis Selection for Single Channel Polyphonic Music Separation Minje Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong, Nam-gu
On Spectral Basis Selection for Single Channel Polyphonic Music Separation Minje Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong, Nam-gu
Probabilistic sparse matrix factorization with an application to discovering gene functions in mouse mrna expression data
 Probabilistic sparse matrix factorization with an application to discovering gene functions in mouse mrna expression data Delbert Dueck, Quaid Morris, and Brendan Frey Department of Electrical and Computer
Probabilistic sparse matrix factorization with an application to discovering gene functions in mouse mrna expression data Delbert Dueck, Quaid Morris, and Brendan Frey Department of Electrical and Computer
SPARSE signal representations have gained popularity in recent
 6958 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 57, NO. 10, OCTOBER 2011 Blind Compressed Sensing Sivan Gleichman and Yonina C. Eldar, Senior Member, IEEE Abstract The fundamental principle underlying
6958 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 57, NO. 10, OCTOBER 2011 Blind Compressed Sensing Sivan Gleichman and Yonina C. Eldar, Senior Member, IEEE Abstract The fundamental principle underlying
Grouping of correlated feature vectors using treelets
 Grouping of correlated feature vectors using treelets Jing Xiang Department of Machine Learning Carnegie Mellon University Pittsburgh, PA 15213 jingx@cs.cmu.edu Abstract In many applications, features
Grouping of correlated feature vectors using treelets Jing Xiang Department of Machine Learning Carnegie Mellon University Pittsburgh, PA 15213 jingx@cs.cmu.edu Abstract In many applications, features
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Singular Value Decomposition and Principal Component Analysis (PCA) I
 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
10-810: Advanced Algorithms and Models for Computational Biology. Optimal leaf ordering and classification
 10-810: Advanced Algorithms and Models for Computational Biology Optimal leaf ordering and classification Hierarchical clustering As we mentioned, its one of the most popular methods for clustering gene
10-810: Advanced Algorithms and Models for Computational Biology Optimal leaf ordering and classification Hierarchical clustering As we mentioned, its one of the most popular methods for clustering gene
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Module Based Neural Networks for Modeling Gene Regulatory Networks
 Module Based Neural Networks for Modeling Gene Regulatory Networks Paresh Chandra Barman, Std 1 ID: 20044523 Term Project: BiS732 Bio-Network Department of BioSystems, Korea Advanced Institute of Science
Module Based Neural Networks for Modeling Gene Regulatory Networks Paresh Chandra Barman, Std 1 ID: 20044523 Term Project: BiS732 Bio-Network Department of BioSystems, Korea Advanced Institute of Science
Lecture Notes 10: Matrix Factorization
 Optimization-based data analysis Fall 207 Lecture Notes 0: Matrix Factorization Low-rank models. Rank- model Consider the problem of modeling a quantity y[i, j] that depends on two indices i and j. To
Optimization-based data analysis Fall 207 Lecture Notes 0: Matrix Factorization Low-rank models. Rank- model Consider the problem of modeling a quantity y[i, j] that depends on two indices i and j. To
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Rule learning for gene expression data
 Rule learning for gene expression data Stefan Enroth Original slides by Torgeir R. Hvidsten The Linnaeus Centre for Bioinformatics Predicting biological process from gene expression time profiles Papers:
Rule learning for gene expression data Stefan Enroth Original slides by Torgeir R. Hvidsten The Linnaeus Centre for Bioinformatics Predicting biological process from gene expression time profiles Papers:
Comparative Network Analysis
 Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Physical network models and multi-source data integration
 Physical network models and multi-source data integration Chen-Hsiang Yeang MIT AI Lab Cambridge, MA 02139 chyeang@ai.mit.edu Tommi Jaakkola MIT AI Lab Cambridge, MA 02139 tommi@ai.mit.edu September 30,
Physical network models and multi-source data integration Chen-Hsiang Yeang MIT AI Lab Cambridge, MA 02139 chyeang@ai.mit.edu Tommi Jaakkola MIT AI Lab Cambridge, MA 02139 tommi@ai.mit.edu September 30,
CPSC 340: Machine Learning and Data Mining. Sparse Matrix Factorization Fall 2018
 CPSC 340: Machine Learning and Data Mining Sparse Matrix Factorization Fall 2018 Last Time: PCA with Orthogonal/Sequential Basis When k = 1, PCA has a scaling problem. When k > 1, have scaling, rotation,
CPSC 340: Machine Learning and Data Mining Sparse Matrix Factorization Fall 2018 Last Time: PCA with Orthogonal/Sequential Basis When k = 1, PCA has a scaling problem. When k > 1, have scaling, rotation,
Learning in Bayesian Networks
 Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Clustering & microarray technology
 Clustering & microarray technology A large scale way to measure gene expression levels. Thanks to Kevin Wayne, Matt Hibbs, & SMD for a few of the slides 1 Why is expression important? Proteins Gene Expression
Clustering & microarray technology A large scale way to measure gene expression levels. Thanks to Kevin Wayne, Matt Hibbs, & SMD for a few of the slides 1 Why is expression important? Proteins Gene Expression
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Missing Value Estimation for Time Series Microarray Data Using Linear Dynamical Systems Modeling
 22nd International Conference on Advanced Information Networking and Applications - Workshops Missing Value Estimation for Time Series Microarray Data Using Linear Dynamical Systems Modeling Connie Phong
22nd International Conference on Advanced Information Networking and Applications - Workshops Missing Value Estimation for Time Series Microarray Data Using Linear Dynamical Systems Modeling Connie Phong
Approximating the Covariance Matrix with Low-rank Perturbations
 Approximating the Covariance Matrix with Low-rank Perturbations Malik Magdon-Ismail and Jonathan T. Purnell Department of Computer Science Rensselaer Polytechnic Institute Troy, NY 12180 {magdon,purnej}@cs.rpi.edu
Approximating the Covariance Matrix with Low-rank Perturbations Malik Magdon-Ismail and Jonathan T. Purnell Department of Computer Science Rensselaer Polytechnic Institute Troy, NY 12180 {magdon,purnej}@cs.rpi.edu
Predicting causal effects in large-scale systems from observational data
 nature methods Predicting causal effects in large-scale systems from observational data Marloes H Maathuis 1, Diego Colombo 1, Markus Kalisch 1 & Peter Bühlmann 1,2 Supplementary figures and text: Supplementary
nature methods Predicting causal effects in large-scale systems from observational data Marloes H Maathuis 1, Diego Colombo 1, Markus Kalisch 1 & Peter Bühlmann 1,2 Supplementary figures and text: Supplementary
Inferring Gene Regulatory Networks from Time-Ordered Gene Expression Data of Bacillus Subtilis Using Differential Equations
 Inferring Gene Regulatory Networks from Time-Ordered Gene Expression Data of Bacillus Subtilis Using Differential Equations M.J.L. de Hoon, S. Imoto, K. Kobayashi, N. Ogasawara, S. Miyano Pacific Symposium
Inferring Gene Regulatory Networks from Time-Ordered Gene Expression Data of Bacillus Subtilis Using Differential Equations M.J.L. de Hoon, S. Imoto, K. Kobayashi, N. Ogasawara, S. Miyano Pacific Symposium
Network Biology-part II
 Network Biology-part II Jun Zhu, Ph. D. Professor of Genomics and Genetic Sciences Icahn Institute of Genomics and Multi-scale Biology The Tisch Cancer Institute Icahn Medical School at Mount Sinai New
Network Biology-part II Jun Zhu, Ph. D. Professor of Genomics and Genetic Sciences Icahn Institute of Genomics and Multi-scale Biology The Tisch Cancer Institute Icahn Medical School at Mount Sinai New
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
c Springer, Reprinted with permission.
 Zhijian Yuan and Erkki Oja. A FastICA Algorithm for Non-negative Independent Component Analysis. In Puntonet, Carlos G.; Prieto, Alberto (Eds.), Proceedings of the Fifth International Symposium on Independent
Zhijian Yuan and Erkki Oja. A FastICA Algorithm for Non-negative Independent Component Analysis. In Puntonet, Carlos G.; Prieto, Alberto (Eds.), Proceedings of the Fifth International Symposium on Independent
Analyzing Microarray Time course Genome wide Data
 OR 779 Functional Data Analysis Course Project Analyzing Microarray Time course Genome wide Data Presented by Xin Zhao April 29, 2002 Cornell University Overview 1. Introduction Biological Background Biological
OR 779 Functional Data Analysis Course Project Analyzing Microarray Time course Genome wide Data Presented by Xin Zhao April 29, 2002 Cornell University Overview 1. Introduction Biological Background Biological
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Breker et al., http://www.jcb.org/cgi/content/full/jcb.201301120/dc1 Figure S1. Single-cell proteomics of stress responses. (a) Using
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Breker et al., http://www.jcb.org/cgi/content/full/jcb.201301120/dc1 Figure S1. Single-cell proteomics of stress responses. (a) Using
Differential Modeling for Cancer Microarray Data
 Differential Modeling for Cancer Microarray Data Omar Odibat Department of Computer Science Feb, 01, 2011 1 Outline Introduction Cancer Microarray data Problem Definition Differential analysis Existing
Differential Modeling for Cancer Microarray Data Omar Odibat Department of Computer Science Feb, 01, 2011 1 Outline Introduction Cancer Microarray data Problem Definition Differential analysis Existing
A New Method to Build Gene Regulation Network Based on Fuzzy Hierarchical Clustering Methods
 International Academic Institute for Science and Technology International Academic Journal of Science and Engineering Vol. 3, No. 6, 2016, pp. 169-176. ISSN 2454-3896 International Academic Journal of
International Academic Institute for Science and Technology International Academic Journal of Science and Engineering Vol. 3, No. 6, 2016, pp. 169-176. ISSN 2454-3896 International Academic Journal of
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Lecture 5: November 19, Minimizing the maximum intracluster distance
 Analysis of DNA Chips and Gene Networks Spring Semester, 2009 Lecture 5: November 19, 2009 Lecturer: Ron Shamir Scribe: Renana Meller 5.1 Minimizing the maximum intracluster distance 5.1.1 Introduction
Analysis of DNA Chips and Gene Networks Spring Semester, 2009 Lecture 5: November 19, 2009 Lecturer: Ron Shamir Scribe: Renana Meller 5.1 Minimizing the maximum intracluster distance 5.1.1 Introduction
Evidence for dynamically organized modularity in the yeast protein-protein interaction network
 Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Automatic Reconstruction of the Building Blocks of Molecular Interaction Networks
 Automatic Reconstruction of the Building Blocks of Molecular Interaction Networks Corban G. Rivera Dissertation submitted to the Faculty of the Virginia Polytechnic Institute and State University in partial
Automatic Reconstruction of the Building Blocks of Molecular Interaction Networks Corban G. Rivera Dissertation submitted to the Faculty of the Virginia Polytechnic Institute and State University in partial
Chapter 16. Clustering Biological Data. Chandan K. Reddy Wayne State University Detroit, MI
 Chapter 16 Clustering Biological Data Chandan K. Reddy Wayne State University Detroit, MI reddy@cs.wayne.edu Mohammad Al Hasan Indiana University - Purdue University Indianapolis, IN alhasan@cs.iupui.edu
Chapter 16 Clustering Biological Data Chandan K. Reddy Wayne State University Detroit, MI reddy@cs.wayne.edu Mohammad Al Hasan Indiana University - Purdue University Indianapolis, IN alhasan@cs.iupui.edu
Ensemble Non-negative Matrix Factorization Methods for Clustering Protein-Protein Interactions
 Belfield Campus Map Ensemble Non-negative Matrix Factorization Methods for Clustering Protein-Protein Interactions
Belfield Campus Map Ensemble Non-negative Matrix Factorization Methods for Clustering Protein-Protein Interactions
Genetic assimilation can occur in the absence of selection for the assimilating phenotype, suggesting a role for the canalization heuristic
 doi: 10.1111/j.1420-9101.2004.00739.x Genetic assimilation can occur in the absence of selection for the assimilating phenotype, suggesting a role for the canalization heuristic J. MASEL Department of
doi: 10.1111/j.1420-9101.2004.00739.x Genetic assimilation can occur in the absence of selection for the assimilating phenotype, suggesting a role for the canalization heuristic J. MASEL Department of
An Introduction to the spls Package, Version 1.0
 An Introduction to the spls Package, Version 1.0 Dongjun Chung 1, Hyonho Chun 1 and Sündüz Keleş 1,2 1 Department of Statistics, University of Wisconsin Madison, WI 53706. 2 Department of Biostatistics
An Introduction to the spls Package, Version 1.0 Dongjun Chung 1, Hyonho Chun 1 and Sündüz Keleş 1,2 1 Department of Statistics, University of Wisconsin Madison, WI 53706. 2 Department of Biostatistics
Singular value decomposition for genome-wide expression data processing and modeling. Presented by Jing Qiu
 Singular value decomposition for genome-wide expression data processing and modeling Presented by Jing Qiu April 23, 2002 Outline Biological Background Mathematical Framework:Singular Value Decomposition
Singular value decomposition for genome-wide expression data processing and modeling Presented by Jing Qiu April 23, 2002 Outline Biological Background Mathematical Framework:Singular Value Decomposition
Data visualization and clustering: an application to gene expression data
 Data visualization and clustering: an application to gene expression data Francesco Napolitano Università degli Studi di Salerno Dipartimento di Matematica e Informatica DAA Erice, April 2007 Thanks to
Data visualization and clustering: an application to gene expression data Francesco Napolitano Università degli Studi di Salerno Dipartimento di Matematica e Informatica DAA Erice, April 2007 Thanks to
V19 Metabolic Networks - Overview
 V19 Metabolic Networks - Overview There exist different levels of computational methods for describing metabolic networks: - stoichiometry/kinetics of classical biochemical pathways (glycolysis, TCA cycle,...
V19 Metabolic Networks - Overview There exist different levels of computational methods for describing metabolic networks: - stoichiometry/kinetics of classical biochemical pathways (glycolysis, TCA cycle,...
Exam: high-dimensional data analysis January 20, 2014
 Exam: high-dimensional data analysis January 20, 204 Instructions: - Write clearly. Scribbles will not be deciphered. - Answer each main question not the subquestions on a separate piece of paper. - Finish
Exam: high-dimensional data analysis January 20, 204 Instructions: - Write clearly. Scribbles will not be deciphered. - Answer each main question not the subquestions on a separate piece of paper. - Finish
V14 extreme pathways
 V14 extreme pathways A torch is directed at an open door and shines into a dark room... What area is lighted? Instead of marking all lighted points individually, it would be sufficient to characterize
V14 extreme pathways A torch is directed at an open door and shines into a dark room... What area is lighted? Instead of marking all lighted points individually, it would be sufficient to characterize
Using Knowledge Driven Matrix Factorization to Reconstruct Modular Gene Regulatory Network
 Using Knowledge Driven Matrix Factorization to Reconstruct Modular Gene Regulatory Network Yang Zhou 1 and Zheng Li and Xuerui Yang and Linxia Zhang and Shireesh Srivastava and Rong Jin 1 and Christina
Using Knowledge Driven Matrix Factorization to Reconstruct Modular Gene Regulatory Network Yang Zhou 1 and Zheng Li and Xuerui Yang and Linxia Zhang and Shireesh Srivastava and Rong Jin 1 and Christina
Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation
 Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation Authors: Fan Zhang, Runsheng Liu and Jie Zheng Presented by: Fan Wu School of Computer Science and
Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation Authors: Fan Zhang, Runsheng Liu and Jie Zheng Presented by: Fan Wu School of Computer Science and
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
Tree-Dependent Components of Gene Expression Data for Clustering
 Tree-Dependent Components of Gene Expression Data for Clustering Jong Kyoung Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong, Nam-gu Pohang
Tree-Dependent Components of Gene Expression Data for Clustering Jong Kyoung Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong, Nam-gu Pohang
Network Biology: Understanding the cell s functional organization. Albert-László Barabási Zoltán N. Oltvai
 Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Hub Gene Selection Methods for the Reconstruction of Transcription Networks
 for the Reconstruction of Transcription Networks José Miguel Hernández-Lobato (1) and Tjeerd. M. H. Dijkstra (2) (1) Computer Science Department, Universidad Autónoma de Madrid, Spain (2) Institute for
for the Reconstruction of Transcription Networks José Miguel Hernández-Lobato (1) and Tjeerd. M. H. Dijkstra (2) (1) Computer Science Department, Universidad Autónoma de Madrid, Spain (2) Institute for
Automatic Rank Determination in Projective Nonnegative Matrix Factorization
 Automatic Rank Determination in Projective Nonnegative Matrix Factorization Zhirong Yang, Zhanxing Zhu, and Erkki Oja Department of Information and Computer Science Aalto University School of Science and
Automatic Rank Determination in Projective Nonnegative Matrix Factorization Zhirong Yang, Zhanxing Zhu, and Erkki Oja Department of Information and Computer Science Aalto University School of Science and
Contingency Table Analysis via Matrix Factorization
 Contingency Table Analysis via Matrix Factorization Kumer Pial Das 1, Jay Powell 2, Myron Katzoff 3, S. Stanley Young 4 1 Department of Mathematics,Lamar University, TX 2 Better Schooling Systems, Pittsburgh,
Contingency Table Analysis via Matrix Factorization Kumer Pial Das 1, Jay Powell 2, Myron Katzoff 3, S. Stanley Young 4 1 Department of Mathematics,Lamar University, TX 2 Better Schooling Systems, Pittsburgh,
Variable Selection in Structured High-dimensional Covariate Spaces
 Variable Selection in Structured High-dimensional Covariate Spaces Fan Li 1 Nancy Zhang 2 1 Department of Health Care Policy Harvard University 2 Department of Statistics Stanford University May 14 2007
Variable Selection in Structured High-dimensional Covariate Spaces Fan Li 1 Nancy Zhang 2 1 Department of Health Care Policy Harvard University 2 Department of Statistics Stanford University May 14 2007
Understanding Science Through the Lens of Computation. Richard M. Karp Nov. 3, 2007
 Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
a Short Introduction
 Collaborative Filtering in Recommender Systems: a Short Introduction Norm Matloff Dept. of Computer Science University of California, Davis matloff@cs.ucdavis.edu December 3, 2016 Abstract There is a strong
Collaborative Filtering in Recommender Systems: a Short Introduction Norm Matloff Dept. of Computer Science University of California, Davis matloff@cs.ucdavis.edu December 3, 2016 Abstract There is a strong
ESL Chap3. Some extensions of lasso
 ESL Chap3 Some extensions of lasso 1 Outline Consistency of lasso for model selection Adaptive lasso Elastic net Group lasso 2 Consistency of lasso for model selection A number of authors have studied
ESL Chap3 Some extensions of lasso 1 Outline Consistency of lasso for model selection Adaptive lasso Elastic net Group lasso 2 Consistency of lasso for model selection A number of authors have studied
Case story: Analysis of the Cell Cycle
 DNA microarray analysis, January 2 nd 2006 Case story: Analysis of the Cell Cycle Center for Biological Sequence Analysis Outline Introduction Cell division and cell cycle regulation Experimental studies
DNA microarray analysis, January 2 nd 2006 Case story: Analysis of the Cell Cycle Center for Biological Sequence Analysis Outline Introduction Cell division and cell cycle regulation Experimental studies
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón
 GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
Optimality, Robustness, and Noise in Metabolic Network Control
 Optimality, Robustness, and Noise in Metabolic Network Control Muxing Chen Gal Chechik Daphne Koller Department of Computer Science Stanford University May 18, 2007 Abstract The existence of noise, or
Optimality, Robustness, and Noise in Metabolic Network Control Muxing Chen Gal Chechik Daphne Koller Department of Computer Science Stanford University May 18, 2007 Abstract The existence of noise, or
Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling
 Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
V15 Flux Balance Analysis Extreme Pathways
 V15 Flux Balance Analysis Extreme Pathways Stoichiometric matrix S: m n matrix with stochiometries of the n reactions as columns and participations of m metabolites as rows. The stochiometric matrix is
V15 Flux Balance Analysis Extreme Pathways Stoichiometric matrix S: m n matrix with stochiometries of the n reactions as columns and participations of m metabolites as rows. The stochiometric matrix is
IEEE TRANSACTIONS ON SIGNAL PROCESSING, VOL. 54, NO. 2, FEBRUARY
 IEEE TRANSACTIONS ON SIGNAL PROCESSING, VOL 54, NO 2, FEBRUARY 2006 423 Underdetermined Blind Source Separation Based on Sparse Representation Yuanqing Li, Shun-Ichi Amari, Fellow, IEEE, Andrzej Cichocki,
IEEE TRANSACTIONS ON SIGNAL PROCESSING, VOL 54, NO 2, FEBRUARY 2006 423 Underdetermined Blind Source Separation Based on Sparse Representation Yuanqing Li, Shun-Ichi Amari, Fellow, IEEE, Andrzej Cichocki,
Biological Networks. Gavin Conant 163B ASRC
 Biological Networks Gavin Conant 163B ASRC conantg@missouri.edu 882-2931 Types of Network Regulatory Protein-interaction Metabolic Signaling Co-expressing General principle Relationship between genes Gene/protein/enzyme
Biological Networks Gavin Conant 163B ASRC conantg@missouri.edu 882-2931 Types of Network Regulatory Protein-interaction Metabolic Signaling Co-expressing General principle Relationship between genes Gene/protein/enzyme
Sparse, stable gene regulatory network recovery via convex optimization
 Sparse, stable gene regulatory network recovery via convex optimization Arwen Meister June, 11 Gene regulatory networks Gene expression regulation allows cells to control protein levels in order to live
Sparse, stable gene regulatory network recovery via convex optimization Arwen Meister June, 11 Gene regulatory networks Gene expression regulation allows cells to control protein levels in order to live
25 : Graphical induced structured input/output models
 10-708: Probabilistic Graphical Models 10-708, Spring 2013 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Meghana Kshirsagar (mkshirsa), Yiwen Chen (yiwenche) 1 Graph
10-708: Probabilistic Graphical Models 10-708, Spring 2013 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Meghana Kshirsagar (mkshirsa), Yiwen Chen (yiwenche) 1 Graph
Generalized Power Method for Sparse Principal Component Analysis
 Generalized Power Method for Sparse Principal Component Analysis Peter Richtárik CORE/INMA Catholic University of Louvain Belgium VOCAL 2008, Veszprém, Hungary CORE Discussion Paper #2008/70 joint work
Generalized Power Method for Sparse Principal Component Analysis Peter Richtárik CORE/INMA Catholic University of Louvain Belgium VOCAL 2008, Veszprém, Hungary CORE Discussion Paper #2008/70 joint work
Single-channel source separation using non-negative matrix factorization
 Single-channel source separation using non-negative matrix factorization Mikkel N. Schmidt Technical University of Denmark mns@imm.dtu.dk www.mikkelschmidt.dk DTU Informatics Department of Informatics
Single-channel source separation using non-negative matrix factorization Mikkel N. Schmidt Technical University of Denmark mns@imm.dtu.dk www.mikkelschmidt.dk DTU Informatics Department of Informatics
Supplemental Information for Pramila et al. Periodic Normal Mixture Model (PNM)
 Supplemental Information for Pramila et al. Periodic Normal Mixture Model (PNM) The data sets alpha30 and alpha38 were analyzed with PNM (Lu et al. 2004). The first two time points were deleted to alleviate
Supplemental Information for Pramila et al. Periodic Normal Mixture Model (PNM) The data sets alpha30 and alpha38 were analyzed with PNM (Lu et al. 2004). The first two time points were deleted to alleviate
Associative Clustering by Maximizing a Bayes Factor
 Associative Clustering by Maximizing a Bayes Factor Janne Sinkkonen, Janne Nikkilä, Leo Lahti, and Samuel Kaski Neural Networks Research Centre Helsinki University of Technology P.O. Box 9800, FIN-02015
Associative Clustering by Maximizing a Bayes Factor Janne Sinkkonen, Janne Nikkilä, Leo Lahti, and Samuel Kaski Neural Networks Research Centre Helsinki University of Technology P.O. Box 9800, FIN-02015
Basic modeling approaches for biological systems. Mahesh Bule
 Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Tiffany Samaroo MB&B 452a December 8, Take Home Final. Topic 1
 Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Lecture 3: A basic statistical concept
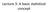 Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Recovery of Low-Rank Plus Compressed Sparse Matrices with Application to Unveiling Traffic Anomalies
 July 12, 212 Recovery of Low-Rank Plus Compressed Sparse Matrices with Application to Unveiling Traffic Anomalies Morteza Mardani Dept. of ECE, University of Minnesota, Minneapolis, MN 55455 Acknowledgments:
July 12, 212 Recovery of Low-Rank Plus Compressed Sparse Matrices with Application to Unveiling Traffic Anomalies Morteza Mardani Dept. of ECE, University of Minnesota, Minneapolis, MN 55455 Acknowledgments:
Robustness of Principal Components
 PCA for Clustering An objective of principal components analysis is to identify linear combinations of the original variables that are useful in accounting for the variation in those original variables.
PCA for Clustering An objective of principal components analysis is to identify linear combinations of the original variables that are useful in accounting for the variation in those original variables.
27: Case study with popular GM III. 1 Introduction: Gene association mapping for complex diseases 1
 10-708: Probabilistic Graphical Models, Spring 2015 27: Case study with popular GM III Lecturer: Eric P. Xing Scribes: Hyun Ah Song & Elizabeth Silver 1 Introduction: Gene association mapping for complex
10-708: Probabilistic Graphical Models, Spring 2015 27: Case study with popular GM III Lecturer: Eric P. Xing Scribes: Hyun Ah Song & Elizabeth Silver 1 Introduction: Gene association mapping for complex
A Multiobjective GO based Approach to Protein Complex Detection
 Available online at www.sciencedirect.com Procedia Technology 4 (2012 ) 555 560 C3IT-2012 A Multiobjective GO based Approach to Protein Complex Detection Sumanta Ray a, Moumita De b, Anirban Mukhopadhyay
Available online at www.sciencedirect.com Procedia Technology 4 (2012 ) 555 560 C3IT-2012 A Multiobjective GO based Approach to Protein Complex Detection Sumanta Ray a, Moumita De b, Anirban Mukhopadhyay
