Singular value decomposition for genome-wide expression data processing and modeling. Presented by Jing Qiu
|
|
|
- Jody Kelly
- 6 years ago
- Views:
Transcription
1 Singular value decomposition for genome-wide expression data processing and modeling Presented by Jing Qiu April 23, 2002
2 Outline Biological Background Mathematical Framework:Singular Value Decomposition SVD calculation Pattern Inference Data normalization Data Sorting Biological Data Analysis I: Elutriation-synchronized cell cycle. 1
3 Biological Background Cell cycle regulation Cell cycle: the program for cell growth and division Four broad phases: (and ), S,,and M Cell cycle diagram (cell growth and protein for DNA synthesis) (DNA replication, two daughter cell) (cell growth and protein synthesis) (split apart) Application: cancer(uncontrolled cell growth and proliferation) 2
4 Biological Background Microarray Experiment Diagram of Central Dogma of Molecule Biology 100,000 genes in mammalian genome each cell express 15,000 of these genes each gene is expressed at a different level cell cycle regulated genes have different expression levels across the cycle period Microarrays:Massive parallel analysis of gene expression The process diagram 3
5 Mathematical Framework:Notation denotes a matrix; denotes a column vector denotes a row vector,, all denote inner products denotes outer product. the th column of the th row of the matrix. 4
6 Mathematical Framework:Data of interest Data of interest: (dim=n M)(See Fig 13a) N rows N genes of a model organism the relative expression of the th gene across different arrays. M columns M different samples(arrays) the genome-wide relative expression measured by the th array. 5
7 Mathematical Framework:SVD of the data where,...!, #" $" %&%&%'" )( ". +* the eigenexpression of the, th eigengene in the, th eigenarray. the N-genes L-eigenarray basis sets.- *,the genome-wide expression in the, th eigenarray. 6
8 M-eigengenes M-arrays basis sets *,the, th eigengene across the different arrays The fraction of eigenexpression : * * Shannon entropy,, measure the complexity of the dataset captured by a single eigengene(eigenarray) all eigengenes are equally expressed Figure 13 and Fig14
9 % Mathematical Framework: SVD calculation diagonalizing (dim= ; ) to get and 7
10 Mathematical Framewor:Pattern Inference An eigengene * represents a regulatory process when this pattern is bilogically interpretable..- * the relative amplitude of the, th eigengene pattern in relative to all other genes. The corresponding eigenarray.- * represents the cellular state which corresponds to this process. 8
11 ! Mathematical Framework:Data normalization Normalizing the data by filtering out the eigengenes * which are inferred to represent noise * +*.- * *... * * where +*... 9
12 Mathematical Framework:Data sorting Sort data by similarity in the expression of any chosen subsets of the eigengenes Correlation plot: Scatter plot of *, where : Amplitude of expression of the th gene in the subspace spanned by and * relative to its overall expression: : The phase of the th gene in the transition from the expression pattern * to and back to *. sort the genes according to. 10
13 Biological Data Analysis:Data Elutriation-synchronized cell cycle data of budding yeast genes, of which were classified as cell cycle regulated in Spellman et al.,1998. arrays in interval of 30 minutes. 11
14 Biological Data Analysis:Pattern inference describes the time-invariant relative expression during the cell cycle.(fig 14a, constantly red) Others show oscillation during the cell cycle. Low entropy: % weak perturbation of a steady state of expression(figure 14b) of the overall rela- captures more than tive expression:,, capture % of the overall relative expression, respectively. The time variation of fits a normalized sine function of period T;(fig 14c) 12
15 The time variations of function of period,(fig 14c) and fit a cosine show decreasing expression on transition from to min show increasing expression on transition from to min Inference: represents experimental additive constants superimposed on a steady gene expression state. represents expression oscillation during a cell cycle and represent initial transient increase and decrease in expression in response to the elutriation, respectively.
16 Biological Data Anlalysis: Data Normalization Filter out the first eigengene to remove the steady state of expression: (,.- where and is the variance in the measured expression of the th gene in the th array. Figure 15a displays the eigengenes for ( ( captures more than information in this dataset. of the overall 13
17 It describes a weak initial transient increase superimposed on a time-invariant scale of expression variance (maybe a response to the elutriation). The time-invariant scale of expression variance suggests a steady scale of the data. It also suggests the time-invariant multiplicative constant noise may be superimposed on the data. Filter out (, removing the steady scale of expression variance, ( ( ( (.- (.( ( ( tabulates expression pattern that are approximately centered at the steady state with variance
18 which are approximately normalized by the steady scale of the expression variance.. and. (of similar significance), capture together more than of the overall normalized expression, %.(Figure 1) fit normalized sine and cosine functions of period T and initial phase. The time variations of. and.. Inference: and. represent cell cycle expression oscillations and assume.- and.- represent the corresponding cell cycle cellular states.
19 Biological Data Analysis:Data sorting Assume.- and.- approximately represent all cell cycle cellular states All arrays(except ) have at least normalized expression in this subspace. of their Suggest:.- and.- are sufficient to approximate the elutriation array expression The array order according to is similar to the cell cycle time points. (an order of the cell cycle progress) Inferences:.- is associated with the cell cycle cellular state of transition from to. 14
20 -.- is associated with the transition from. to.- is associated with the cell cycle cellular state of transition from to. -.- is associated with the transition from to. The phase of, corresponds to the 30-min delay between the start of the experiment and that of the cell cycle stage. Figure 2b. Recall and. are inferred to approximately represent all cell cycle expression oscillation
21 Expect cell cycle regulated not regulated by the cell cycle at all. 641(classified as cell cycle regulated) have more than of their normalized expression in this subspace. This sortinggives a different classification from that by Spellman et al(the poor quality of the elutriation expression data). With all genes sorted, the gene variations of.- and.- fit normalized sine and cosine functions(figure 3). The sorted and normalized elutriation expression fit approximately a traveling wave of expression varying sinusoidally across both genes and arrays.
22 Some points SVD calculation: don t necessarily exist. Data Sorting What if more then 2 significant eigengenes? or? What smoothing method to fit? The disagreement due to poor quality of the data(too simple)? Are the first two eigengenes enough to explain all?(capture only around and Figure 2b not good projection) 15
Analyzing Microarray Time course Genome wide Data
 OR 779 Functional Data Analysis Course Project Analyzing Microarray Time course Genome wide Data Presented by Xin Zhao April 29, 2002 Cornell University Overview 1. Introduction Biological Background Biological
OR 779 Functional Data Analysis Course Project Analyzing Microarray Time course Genome wide Data Presented by Xin Zhao April 29, 2002 Cornell University Overview 1. Introduction Biological Background Biological
Singular Value Decomposition and Principal Component Analysis (PCA) I
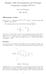 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays
 Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays Prof. Tesler Math 283 Fall 2015 Prof. Tesler Principal Components Analysis Math 283 / Fall 2015
Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays Prof. Tesler Math 283 Fall 2015 Prof. Tesler Principal Components Analysis Math 283 / Fall 2015
Uses of the Singular Value Decompositions in Biology
 Uses of the Singular Value Decompositions in Biology The Singular Value Decomposition is a very useful tool deeply rooted in linear algebra. Despite this, SVDs have found their way into many different
Uses of the Singular Value Decompositions in Biology The Singular Value Decomposition is a very useful tool deeply rooted in linear algebra. Despite this, SVDs have found their way into many different
Basic modeling approaches for biological systems. Mahesh Bule
 Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
BIOINFORMATICS. Geometry of gene expression dynamics. S. A. Rifkin 1 and J. Kim INTRODUCTION SYSTEM
 BIOINFORMATICS Vol. 18 no. 9 2002 Pages 1176 1183 Geometry of gene expression dynamics S. A. Rifkin 1 and J. Kim 1, 2, 3, 1 Department of Ecology and Evolutionary Biology, PO Box 208106, 2 Department of
BIOINFORMATICS Vol. 18 no. 9 2002 Pages 1176 1183 Geometry of gene expression dynamics S. A. Rifkin 1 and J. Kim 1, 2, 3, 1 Department of Ecology and Evolutionary Biology, PO Box 208106, 2 Department of
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
Lecture Notes 1: Vector spaces
 Optimization-based data analysis Fall 2017 Lecture Notes 1: Vector spaces In this chapter we review certain basic concepts of linear algebra, highlighting their application to signal processing. 1 Vector
Optimization-based data analysis Fall 2017 Lecture Notes 1: Vector spaces In this chapter we review certain basic concepts of linear algebra, highlighting their application to signal processing. 1 Vector
Grouping of correlated feature vectors using treelets
 Grouping of correlated feature vectors using treelets Jing Xiang Department of Machine Learning Carnegie Mellon University Pittsburgh, PA 15213 jingx@cs.cmu.edu Abstract In many applications, features
Grouping of correlated feature vectors using treelets Jing Xiang Department of Machine Learning Carnegie Mellon University Pittsburgh, PA 15213 jingx@cs.cmu.edu Abstract In many applications, features
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
December 20, MAA704, Multivariate analysis. Christopher Engström. Multivariate. analysis. Principal component analysis
 .. December 20, 2013 Todays lecture. (PCA) (PLS-R) (LDA) . (PCA) is a method often used to reduce the dimension of a large dataset to one of a more manageble size. The new dataset can then be used to make
.. December 20, 2013 Todays lecture. (PCA) (PLS-R) (LDA) . (PCA) is a method often used to reduce the dimension of a large dataset to one of a more manageble size. The new dataset can then be used to make
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Learning in Bayesian Networks
 Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Cell Division and Reproduction
 Cell Division and Reproduction What do you think this picture shows? If you guessed that it s a picture of two cells, you are right. In fact, the picture shows human cancer cells, and they are nearing
Cell Division and Reproduction What do you think this picture shows? If you guessed that it s a picture of two cells, you are right. In fact, the picture shows human cancer cells, and they are nearing
5.1 Cell Division and the Cell Cycle
 5.1 Cell Division and the Cell Cycle Lesson Objectives Contrast cell division in prokaryotes and eukaryotes. Identify the phases of the eukaryotic cell cycle. Explain how the cell cycle is controlled.
5.1 Cell Division and the Cell Cycle Lesson Objectives Contrast cell division in prokaryotes and eukaryotes. Identify the phases of the eukaryotic cell cycle. Explain how the cell cycle is controlled.
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
Modeling Classes of Shapes Suppose you have a class of shapes with a range of variations: System 2 Overview
 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 Modeling Classes of Shapes Suppose you have a class of shapes with a range of variations: System processes System Overview Previous Systems:
4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 4 4 4 6 Modeling Classes of Shapes Suppose you have a class of shapes with a range of variations: System processes System Overview Previous Systems:
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
Discovering Correlation in Data. Vinh Nguyen Research Fellow in Data Science Computing and Information Systems DMD 7.
 Discovering Correlation in Data Vinh Nguyen (vinh.nguyen@unimelb.edu.au) Research Fellow in Data Science Computing and Information Systems DMD 7.14 Discovering Correlation Why is correlation important?
Discovering Correlation in Data Vinh Nguyen (vinh.nguyen@unimelb.edu.au) Research Fellow in Data Science Computing and Information Systems DMD 7.14 Discovering Correlation Why is correlation important?
Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells
 Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells Understand how a simple biochemical oscillator can drive the
Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells Understand how a simple biochemical oscillator can drive the
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Modeling Gene Expression from Microarray Expression Data with State-Space Equations. F.X. Wu, W.J. Zhang, and A.J. Kusalik
 Modeling Gene Expression from Microarray Expression Data with State-Space Equations FX Wu, WJ Zhang, and AJ Kusalik Pacific Symposium on Biocomputing 9:581-592(2004) MODELING GENE EXPRESSION FROM MICROARRAY
Modeling Gene Expression from Microarray Expression Data with State-Space Equations FX Wu, WJ Zhang, and AJ Kusalik Pacific Symposium on Biocomputing 9:581-592(2004) MODELING GENE EXPRESSION FROM MICROARRAY
Gene expression microarray technology measures the expression levels of thousands of genes. Research Article
 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 7, Number 2, 2 # Mary Ann Liebert, Inc. Pp. 8 DOI:.89/cmb.29.52 Research Article Reducing the Computational Complexity of Information Theoretic Approaches for Reconstructing
JOURNAL OF COMPUTATIONAL BIOLOGY Volume 7, Number 2, 2 # Mary Ann Liebert, Inc. Pp. 8 DOI:.89/cmb.29.52 Research Article Reducing the Computational Complexity of Information Theoretic Approaches for Reconstructing
Linear Algebra (Review) Volker Tresp 2017
 Linear Algebra (Review) Volker Tresp 2017 1 Vectors k is a scalar (a number) c is a column vector. Thus in two dimensions, c = ( c1 c 2 ) (Advanced: More precisely, a vector is defined in a vector space.
Linear Algebra (Review) Volker Tresp 2017 1 Vectors k is a scalar (a number) c is a column vector. Thus in two dimensions, c = ( c1 c 2 ) (Advanced: More precisely, a vector is defined in a vector space.
Salt Dome Detection and Tracking Using Texture Analysis and Tensor-based Subspace Learning
 Salt Dome Detection and Tracking Using Texture Analysis and Tensor-based Subspace Learning Zhen Wang*, Dr. Tamir Hegazy*, Dr. Zhiling Long, and Prof. Ghassan AlRegib 02/18/2015 1 /42 Outline Introduction
Salt Dome Detection and Tracking Using Texture Analysis and Tensor-based Subspace Learning Zhen Wang*, Dr. Tamir Hegazy*, Dr. Zhiling Long, and Prof. Ghassan AlRegib 02/18/2015 1 /42 Outline Introduction
Three different fusions led to three basic ideas: 1) If one fuses a cell in mitosis with a cell in any other stage of the cell cycle, the chromosomes
 Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides
 Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data
 GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
Lecture 13. Principal Component Analysis. Brett Bernstein. April 25, CDS at NYU. Brett Bernstein (CDS at NYU) Lecture 13 April 25, / 26
 Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Analysis and Simulation of Biological Systems
 Analysis and Simulation of Biological Systems Dr. Carlo Cosentino School of Computer and Biomedical Engineering Department of Experimental and Clinical Medicine Università degli Studi Magna Graecia Catanzaro,
Analysis and Simulation of Biological Systems Dr. Carlo Cosentino School of Computer and Biomedical Engineering Department of Experimental and Clinical Medicine Università degli Studi Magna Graecia Catanzaro,
Linear Algebra (Review) Volker Tresp 2018
 Linear Algebra (Review) Volker Tresp 2018 1 Vectors k, M, N are scalars A one-dimensional array c is a column vector. Thus in two dimensions, ( ) c1 c = c 2 c i is the i-th component of c c T = (c 1, c
Linear Algebra (Review) Volker Tresp 2018 1 Vectors k, M, N are scalars A one-dimensional array c is a column vector. Thus in two dimensions, ( ) c1 c = c 2 c i is the i-th component of c c T = (c 1, c
Inferring Transcriptional Regulatory Networks from Gene Expression Data II
 Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Network Biology: Understanding the cell s functional organization. Albert-László Barabási Zoltán N. Oltvai
 Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Singular value decomposition (SVD) of large random matrices. India, 2010
 Singular value decomposition (SVD) of large random matrices Marianna Bolla Budapest University of Technology and Economics marib@math.bme.hu India, 2010 Motivation New challenge of multivariate statistics:
Singular value decomposition (SVD) of large random matrices Marianna Bolla Budapest University of Technology and Economics marib@math.bme.hu India, 2010 Motivation New challenge of multivariate statistics:
Lab 2 Worksheet. Problems. Problem 1: Geometry and Linear Equations
 Lab 2 Worksheet Problems Problem : Geometry and Linear Equations Linear algebra is, first and foremost, the study of systems of linear equations. You are going to encounter linear systems frequently in
Lab 2 Worksheet Problems Problem : Geometry and Linear Equations Linear algebra is, first and foremost, the study of systems of linear equations. You are going to encounter linear systems frequently in
COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry.
 North Carolina Draft Standard Course of Study and Grade Level Competencies, Biology BIOLOGY COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry. 1.01
North Carolina Draft Standard Course of Study and Grade Level Competencies, Biology BIOLOGY COMPETENCY GOAL 1: The learner will develop abilities necessary to do and understand scientific inquiry. 1.01
The University of Texas at Austin Department of Electrical and Computer Engineering. EE381V: Large Scale Learning Spring 2013.
 The University of Texas at Austin Department of Electrical and Computer Engineering EE381V: Large Scale Learning Spring 2013 Assignment Two Caramanis/Sanghavi Due: Tuesday, Feb. 19, 2013. Computational
The University of Texas at Austin Department of Electrical and Computer Engineering EE381V: Large Scale Learning Spring 2013 Assignment Two Caramanis/Sanghavi Due: Tuesday, Feb. 19, 2013. Computational
Biological Pathways Representation by Petri Nets and extension
 Biological Pathways Representation by and extensions December 6, 2006 Biological Pathways Representation by and extension 1 The cell Pathways 2 Definitions 3 4 Biological Pathways Representation by and
Biological Pathways Representation by and extensions December 6, 2006 Biological Pathways Representation by and extension 1 The cell Pathways 2 Definitions 3 4 Biological Pathways Representation by and
We use the overhead arrow to denote a column vector, i.e., a number with a direction. For example, in three-space, we write
 1 MATH FACTS 11 Vectors 111 Definition We use the overhead arrow to denote a column vector, ie, a number with a direction For example, in three-space, we write The elements of a vector have a graphical
1 MATH FACTS 11 Vectors 111 Definition We use the overhead arrow to denote a column vector, ie, a number with a direction For example, in three-space, we write The elements of a vector have a graphical
Principal Components
 Principal Cmpnents Suppse we have N measurements n each f p variables X j, j = 1,..., p. There are several equivalent appraches t principal cmpnents: Given X = (X 1,... X p ), prduce a derived (and small)
Principal Cmpnents Suppse we have N measurements n each f p variables X j, j = 1,..., p. There are several equivalent appraches t principal cmpnents: Given X = (X 1,... X p ), prduce a derived (and small)
Statistics for Differential Expression in Sequencing Studies. Naomi Altman
 Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
Tree-Dependent Components of Gene Expression Data for Clustering
 Tree-Dependent Components of Gene Expression Data for Clustering Jong Kyoung Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong, Nam-gu Pohang
Tree-Dependent Components of Gene Expression Data for Clustering Jong Kyoung Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong, Nam-gu Pohang
AP BIOLOGY SUMMER ASSIGNMENT
 AP BIOLOGY SUMMER ASSIGNMENT Welcome to EDHS Advanced Placement Biology! The attached summer assignment is required for all AP Biology students for the 2011-2012 school year. The assignment consists of
AP BIOLOGY SUMMER ASSIGNMENT Welcome to EDHS Advanced Placement Biology! The attached summer assignment is required for all AP Biology students for the 2011-2012 school year. The assignment consists of
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Lecture 24: Principal Component Analysis. Aykut Erdem May 2016 Hacettepe University
 Lecture 4: Principal Component Analysis Aykut Erdem May 016 Hacettepe University This week Motivation PCA algorithms Applications PCA shortcomings Autoencoders Kernel PCA PCA Applications Data Visualization
Lecture 4: Principal Component Analysis Aykut Erdem May 016 Hacettepe University This week Motivation PCA algorithms Applications PCA shortcomings Autoencoders Kernel PCA PCA Applications Data Visualization
Bioinformatics. Transcriptome
 Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
CS168: The Modern Algorithmic Toolbox Lecture #8: How PCA Works
 CS68: The Modern Algorithmic Toolbox Lecture #8: How PCA Works Tim Roughgarden & Gregory Valiant April 20, 206 Introduction Last lecture introduced the idea of principal components analysis (PCA). The
CS68: The Modern Algorithmic Toolbox Lecture #8: How PCA Works Tim Roughgarden & Gregory Valiant April 20, 206 Introduction Last lecture introduced the idea of principal components analysis (PCA). The
Latent Variable models for GWAs
 Latent Variable models for GWAs Oliver Stegle Machine Learning and Computational Biology Research Group Max-Planck-Institutes Tübingen, Germany September 2011 O. Stegle Latent variable models for GWAs
Latent Variable models for GWAs Oliver Stegle Machine Learning and Computational Biology Research Group Max-Planck-Institutes Tübingen, Germany September 2011 O. Stegle Latent variable models for GWAs
Table of Laplacetransform
 Appendix Table of Laplacetransform pairs 1(t) f(s) oct), unit impulse at t = 0 a, a constant or step of magnitude a at t = 0 a s t, a ramp function e- at, an exponential function s + a sin wt, a sine fun
Appendix Table of Laplacetransform pairs 1(t) f(s) oct), unit impulse at t = 0 a, a constant or step of magnitude a at t = 0 a s t, a ramp function e- at, an exponential function s + a sin wt, a sine fun
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
178 Part 3.2 SUMMARY INTRODUCTION
 178 Part 3.2 Chapter # DYNAMIC FILTRATION OF VARIABILITY WITHIN EXPRESSION PATTERNS OF ZYGOTIC SEGMENTATION GENES IN DROSOPHILA Surkova S.Yu. *, Samsonova M.G. St. Petersburg State Polytechnical University,
178 Part 3.2 Chapter # DYNAMIC FILTRATION OF VARIABILITY WITHIN EXPRESSION PATTERNS OF ZYGOTIC SEGMENTATION GENES IN DROSOPHILA Surkova S.Yu. *, Samsonova M.G. St. Petersburg State Polytechnical University,
Comparative Network Analysis
 Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Differential Modeling for Cancer Microarray Data
 Differential Modeling for Cancer Microarray Data Omar Odibat Department of Computer Science Feb, 01, 2011 1 Outline Introduction Cancer Microarray data Problem Definition Differential analysis Existing
Differential Modeling for Cancer Microarray Data Omar Odibat Department of Computer Science Feb, 01, 2011 1 Outline Introduction Cancer Microarray data Problem Definition Differential analysis Existing
2. Mathematical descriptions. (i) the master equation (ii) Langevin theory. 3. Single cell measurements
 1. Why stochastic?. Mathematical descriptions (i) the master equation (ii) Langevin theory 3. Single cell measurements 4. Consequences Any chemical reaction is stochastic. k P d φ dp dt = k d P deterministic
1. Why stochastic?. Mathematical descriptions (i) the master equation (ii) Langevin theory 3. Single cell measurements 4. Consequences Any chemical reaction is stochastic. k P d φ dp dt = k d P deterministic
NAME MATH 304 Examination 2 Page 1
 NAME MATH 4 Examination 2 Page. [8 points (a) Find the following determinant. However, use only properties of determinants, without calculating directly (that is without expanding along a column or row
NAME MATH 4 Examination 2 Page. [8 points (a) Find the following determinant. However, use only properties of determinants, without calculating directly (that is without expanding along a column or row
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation
 Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation Authors: Fan Zhang, Runsheng Liu and Jie Zheng Presented by: Fan Wu School of Computer Science and
Sig2GRN: A Software Tool Linking Signaling Pathway with Gene Regulatory Network for Dynamic Simulation Authors: Fan Zhang, Runsheng Liu and Jie Zheng Presented by: Fan Wu School of Computer Science and
Androgen-independent prostate cancer
 The following tutorial walks through the identification of biological themes in a microarray dataset examining androgen-independent. Visit the GeneSifter Data Center (www.genesifter.net/web/datacenter.html)
The following tutorial walks through the identification of biological themes in a microarray dataset examining androgen-independent. Visit the GeneSifter Data Center (www.genesifter.net/web/datacenter.html)
CS-E5880 Modeling biological networks Gene regulatory networks
 CS-E5880 Modeling biological networks Gene regulatory networks Jukka Intosalmi (based on slides by Harri Lähdesmäki) Department of Computer Science Aalto University January 12, 2018 Outline Modeling gene
CS-E5880 Modeling biological networks Gene regulatory networks Jukka Intosalmi (based on slides by Harri Lähdesmäki) Department of Computer Science Aalto University January 12, 2018 Outline Modeling gene
Supplementary material (Additional file 1)
 Supplementary material (Additional file 1) Contents I. Bias-Variance Decomposition II. Supplementary Figures: Simulation model 2 and real data sets III. Supplementary Figures: Simulation model 1 IV. Molecule
Supplementary material (Additional file 1) Contents I. Bias-Variance Decomposition II. Supplementary Figures: Simulation model 2 and real data sets III. Supplementary Figures: Simulation model 1 IV. Molecule
Genetics, brain development, and behavior
 Genetics, brain development, and behavior Jan. 13, 2004 Questions: Does it make sense to talk about genes for behavior? How do genes turn into brains? Can environment affect development before birth? What
Genetics, brain development, and behavior Jan. 13, 2004 Questions: Does it make sense to talk about genes for behavior? How do genes turn into brains? Can environment affect development before birth? What
Bioinformatics Chapter 1. Introduction
 Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
2. Review of Linear Algebra
 2. Review of Linear Algebra ECE 83, Spring 217 In this course we will represent signals as vectors and operators (e.g., filters, transforms, etc) as matrices. This lecture reviews basic concepts from linear
2. Review of Linear Algebra ECE 83, Spring 217 In this course we will represent signals as vectors and operators (e.g., filters, transforms, etc) as matrices. This lecture reviews basic concepts from linear
Midterm 1 Solutions Math Section 55 - Spring 2018 Instructor: Daren Cheng
 Midterm 1 Solutions Math 20250 Section 55 - Spring 2018 Instructor: Daren Cheng #1 Do the following problems using row reduction. (a) (6 pts) Let A = 2 1 2 6 1 3 8 17 3 5 4 5 Find bases for N A and R A,
Midterm 1 Solutions Math 20250 Section 55 - Spring 2018 Instructor: Daren Cheng #1 Do the following problems using row reduction. (a) (6 pts) Let A = 2 1 2 6 1 3 8 17 3 5 4 5 Find bases for N A and R A,
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes.
 I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
CELL CYCLE AND DIFFERENTIATION
 CELL CYCLE AND DIFFERENTIATION Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id WHAT IS CELL CYCLE? 09/12/14 d.purnomosari@ugm.ac.id
CELL CYCLE AND DIFFERENTIATION Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id WHAT IS CELL CYCLE? 09/12/14 d.purnomosari@ugm.ac.id
GENOMIC SIGNAL PROCESSING. Lecture 2. Classification of disease subtype based on microarray data
 GENOMIC SIGNAL PROCESSING Lecture 2 Classification of disease subtype based on microarray data 1. Analysis of microarray data (see last 15 slides of Lecture 1) 2. Classification methods for microarray
GENOMIC SIGNAL PROCESSING Lecture 2 Classification of disease subtype based on microarray data 1. Analysis of microarray data (see last 15 slides of Lecture 1) 2. Classification methods for microarray
I. Specialization. II. Autonomous signals
 Multicellularity Up to this point in the class we have been discussing individual cells, or, at most, populations of individual cells. But some interesting life forms (for example, humans) consist not
Multicellularity Up to this point in the class we have been discussing individual cells, or, at most, populations of individual cells. But some interesting life forms (for example, humans) consist not
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Cell cycle regulation in the budding yeast
 Cell cycle regulation in the budding yeast Bởi: TS. Nguyen Cuong Introduction The cell cycle is the sequence of events by which a growing cell duplicates all its components and then divides into two daughter
Cell cycle regulation in the budding yeast Bởi: TS. Nguyen Cuong Introduction The cell cycle is the sequence of events by which a growing cell duplicates all its components and then divides into two daughter
Problem # Max points possible Actual score Total 120
 FINAL EXAMINATION - MATH 2121, FALL 2017. Name: ID#: Email: Lecture & Tutorial: Problem # Max points possible Actual score 1 15 2 15 3 10 4 15 5 15 6 15 7 10 8 10 9 15 Total 120 You have 180 minutes to
FINAL EXAMINATION - MATH 2121, FALL 2017. Name: ID#: Email: Lecture & Tutorial: Problem # Max points possible Actual score 1 15 2 15 3 10 4 15 5 15 6 15 7 10 8 10 9 15 Total 120 You have 180 minutes to
BIOLOGY Grades Summer Units: 10 high school credits UC Requirement Category: d. General Description:
 Summer 2015 Units: 10 high school credits UC Requirement Category: d General Description: BIOLOGY Grades 9-12 Summer session biology will be an intense, fast paced course. Students will gain an understanding
Summer 2015 Units: 10 high school credits UC Requirement Category: d General Description: BIOLOGY Grades 9-12 Summer session biology will be an intense, fast paced course. Students will gain an understanding
Cluster Analysis of Gene Expression Microarray Data. BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002
 Cluster Analysis of Gene Expression Microarray Data BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002 1 Data representations Data are relative measurements log 2 ( red
Cluster Analysis of Gene Expression Microarray Data BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002 1 Data representations Data are relative measurements log 2 ( red
Dimension Reduction and Iterative Consensus Clustering
 Dimension Reduction and Iterative Consensus Clustering Southeastern Clustering and Ranking Workshop August 24, 2009 Dimension Reduction and Iterative 1 Document Clustering Geometry of the SVD Centered
Dimension Reduction and Iterative Consensus Clustering Southeastern Clustering and Ranking Workshop August 24, 2009 Dimension Reduction and Iterative 1 Document Clustering Geometry of the SVD Centered
Unravelling the biochemical reaction kinetics from time-series data
 Unravelling the biochemical reaction kinetics from time-series data Santiago Schnell Indiana University School of Informatics and Biocomplexity Institute Email: schnell@indiana.edu WWW: http://www.informatics.indiana.edu/schnell
Unravelling the biochemical reaction kinetics from time-series data Santiago Schnell Indiana University School of Informatics and Biocomplexity Institute Email: schnell@indiana.edu WWW: http://www.informatics.indiana.edu/schnell
Biol478/ August
 Biol478/595 29 August # Day Inst. Topic Hwk Reading August 1 M 25 MG Introduction 2 W 27 MG Sequences and Evolution Handouts 3 F 29 MG Sequences and Evolution September M 1 Labor Day 4 W 3 MG Database
Biol478/595 29 August # Day Inst. Topic Hwk Reading August 1 M 25 MG Introduction 2 W 27 MG Sequences and Evolution Handouts 3 F 29 MG Sequences and Evolution September M 1 Labor Day 4 W 3 MG Database
CHAPTER 12 - THE CELL CYCLE (pgs )
 CHAPTER 12 - THE CELL CYCLE (pgs. 228-245) CHAPTER SEVEN TARGETS I. Describe the importance of mitosis in single-celled and multi-cellular organisms. II. Explain the organization of DNA molecules and their
CHAPTER 12 - THE CELL CYCLE (pgs. 228-245) CHAPTER SEVEN TARGETS I. Describe the importance of mitosis in single-celled and multi-cellular organisms. II. Explain the organization of DNA molecules and their
DS-GA 1002 Lecture notes 10 November 23, Linear models
 DS-GA 2 Lecture notes November 23, 2 Linear functions Linear models A linear model encodes the assumption that two quantities are linearly related. Mathematically, this is characterized using linear functions.
DS-GA 2 Lecture notes November 23, 2 Linear functions Linear models A linear model encodes the assumption that two quantities are linearly related. Mathematically, this is characterized using linear functions.
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Properties of Matrices and Operations on Matrices
 Properties of Matrices and Operations on Matrices A common data structure for statistical analysis is a rectangular array or matris. Rows represent individual observational units, or just observations,
Properties of Matrices and Operations on Matrices A common data structure for statistical analysis is a rectangular array or matris. Rows represent individual observational units, or just observations,
PCA, Kernel PCA, ICA
 PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
Curriculum Links. AQA GCE Biology. AS level
 Curriculum Links AQA GCE Biology Unit 2 BIOL2 The variety of living organisms 3.2.1 Living organisms vary and this variation is influenced by genetic and environmental factors Causes of variation 3.2.2
Curriculum Links AQA GCE Biology Unit 2 BIOL2 The variety of living organisms 3.2.1 Living organisms vary and this variation is influenced by genetic and environmental factors Causes of variation 3.2.2
Total
 Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
Advanced Introduction to Machine Learning CMU-10715
 Advanced Introduction to Machine Learning CMU-10715 Principal Component Analysis Barnabás Póczos Contents Motivation PCA algorithms Applications Some of these slides are taken from Karl Booksh Research
Advanced Introduction to Machine Learning CMU-10715 Principal Component Analysis Barnabás Póczos Contents Motivation PCA algorithms Applications Some of these slides are taken from Karl Booksh Research
Networks in systems biology
 Networks in systems biology Matthew Macauley Department of Mathematical Sciences Clemson University http://www.math.clemson.edu/~macaule/ Math 4500, Spring 2017 M. Macauley (Clemson) Networks in systems
Networks in systems biology Matthew Macauley Department of Mathematical Sciences Clemson University http://www.math.clemson.edu/~macaule/ Math 4500, Spring 2017 M. Macauley (Clemson) Networks in systems
Correlations to Next Generation Science Standards. Life Sciences Disciplinary Core Ideas. LS-1 From Molecules to Organisms: Structures and Processes
 Correlations to Next Generation Science Standards Life Sciences Disciplinary Core Ideas LS-1 From Molecules to Organisms: Structures and Processes LS1.A Structure and Function Systems of specialized cells
Correlations to Next Generation Science Standards Life Sciences Disciplinary Core Ideas LS-1 From Molecules to Organisms: Structures and Processes LS1.A Structure and Function Systems of specialized cells
BIOINFORMATICS Datamining #1
 BIOINFORMATICS Datamining #1 Mark Gerstein, Yale University gersteinlab.org/courses/452 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu Large-scale Datamining Gene Expression Representing Data in
BIOINFORMATICS Datamining #1 Mark Gerstein, Yale University gersteinlab.org/courses/452 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu Large-scale Datamining Gene Expression Representing Data in
Keywords: systems biology, microarrays, gene expression, clustering
 Jan H. Ihmels received his PhD in computational biology from the Weizmann Institute of Science, Israel. He is currently a postdoctoral fellow at the Department of Molecular Genetics of the Weizmann Institute.
Jan H. Ihmels received his PhD in computational biology from the Weizmann Institute of Science, Israel. He is currently a postdoctoral fellow at the Department of Molecular Genetics of the Weizmann Institute.
Chapter: Life's Structure and Classification
 Table of Contents Chapter: Life's Structure and Classification Section 1: Living Things 1- What is an organism? Any living thing is called an organism. Organisms vary in size: 1)one-celled or unicellular
Table of Contents Chapter: Life's Structure and Classification Section 1: Living Things 1- What is an organism? Any living thing is called an organism. Organisms vary in size: 1)one-celled or unicellular
International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February ISSN
 International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February-2015 273 Analogizing And Investigating Some Applications of Metabolic Pathway Analysis Methods Gourav Mukherjee 1,
International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February-2015 273 Analogizing And Investigating Some Applications of Metabolic Pathway Analysis Methods Gourav Mukherjee 1,
Subspace sampling and relative-error matrix approximation
 Subspace sampling and relative-error matrix approximation Petros Drineas Rensselaer Polytechnic Institute Computer Science Department (joint work with M. W. Mahoney) For papers, etc. drineas The CUR decomposition
Subspace sampling and relative-error matrix approximation Petros Drineas Rensselaer Polytechnic Institute Computer Science Department (joint work with M. W. Mahoney) For papers, etc. drineas The CUR decomposition
Non-Negative Factorization for Clustering of Microarray Data
 INT J COMPUT COMMUN, ISSN 1841-9836 9(1):16-23, February, 2014. Non-Negative Factorization for Clustering of Microarray Data L. Morgos Lucian Morgos Dept. of Electronics and Telecommunications Faculty
INT J COMPUT COMMUN, ISSN 1841-9836 9(1):16-23, February, 2014. Non-Negative Factorization for Clustering of Microarray Data L. Morgos Lucian Morgos Dept. of Electronics and Telecommunications Faculty
L26: Advanced dimensionality reduction
 L26: Advanced dimensionality reduction The snapshot CA approach Oriented rincipal Components Analysis Non-linear dimensionality reduction (manifold learning) ISOMA Locally Linear Embedding CSCE 666 attern
L26: Advanced dimensionality reduction The snapshot CA approach Oriented rincipal Components Analysis Non-linear dimensionality reduction (manifold learning) ISOMA Locally Linear Embedding CSCE 666 attern
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
6.435, System Identification
 SET 6 System Identification 6.435 Parametrized model structures One-step predictor Identifiability Munther A. Dahleh 1 Models of LTI Systems A complete model u = input y = output e = noise (with PDF).
SET 6 System Identification 6.435 Parametrized model structures One-step predictor Identifiability Munther A. Dahleh 1 Models of LTI Systems A complete model u = input y = output e = noise (with PDF).
