BIOINFORMATICS Datamining #1
|
|
|
- Barnaby Simpson
- 6 years ago
- Views:
Transcription
1 BIOINFORMATICS Datamining #1 Mark Gerstein, Yale University gersteinlab.org/courses/452 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
2 Large-scale Datamining Gene Expression Representing Data in a Grid Description of function prediction in abstract context Unsupervised Learning clustering & k-means Local clustering Supervised Learning Discriminants & Decision Tree Bayesian Nets Function Prediction EX Simple Bayesian Approach for Localization Prediction 2 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
3 The recent advent and subsequent onslaught of microarray data 1st generation, Expression Arrays (Brown) 2nd gen., Proteome Chips (Snyder) 3 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
4 4 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu Gene Expression Information and Protein Features
5 Functional Classification COGs (cross-org., just conserved, NCBI Koonin/Lipman) Fly (fly, Ashburner) now extended to GO (cross-org.) GenProtEC (E. coli, Riley) MIPS/PEDANT (yeast, Mewes) ENZYME (SwissProt Bairoch/ Apweiler, just enzymes, cross-org.) Also: Other SwissProt Annotation WIT, KEGG (just pathways) TIGR EGAD (human ESTs) SGD (yeast) 5 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
6 Cor e Prediction of Function on a Genomic Scale from Array Data & Sequence Features Different Aspects of function: molecular action, cellular role, phenotypic manifestation Also: localization, interactions, complexes 6 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
7 Arrange data in a tabulated form, each row representing an example and each column representing a feature, including the dependent experimental quantity to be predicted. predictor1 Predictor2 predictor3 predictor4 response G1 A(1,1) A(1,2) A(1,3) A(1,4) Class A G2 A(2,1) A(2,2) A(2,3) A(2,4) Class A G3 A(3,1) A(3,2) A(3,3) A(3,4) Class B (adapted from Y Kluger) 7 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
8 8 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu Typical Predictors and Response for Yeast
9 Represent predictors in abstract high dimensional space Cor e 9 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
10 Tag Certain Points Cor e 10 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
11 Abstract high-dimensional space representation 11 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
12 Large-scale Datamining Gene Expression Representing Data in a Grid Description of function prediction in abstract context Unsupervised Learning clustering & k-means Local clustering Supervised Learning Discriminants & Decision Tree Bayesian Nets Function Prediction EX Simple Bayesian Approach for Localization Prediction 12 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
13 cluster predictors Cor e 13 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
14 Use clusters to predict Response Cor e 14 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
15 K-means Cor e 15 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
16 K-means Top-down vs. Bottom up Top-down when you know how many subdivisions k-means as an example of top-down 1) Pick ten (i.e. k?) random points as putative cluster centers. 2) Group the points to be clustered by the center to which they are closest. 3) Then take the mean of each group and repeat, with the means now at the cluster center. 4) I suppose you stop when the centers stop moving. 16 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
17 Bottom up clustering Cor e 17 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
18 Large-scale Datamining Gene Expression Representing Data in a Grid Description of function prediction in abstract context Unsupervised Learning clustering & k-means Local clustering Supervised Learning Discriminants & Decision Tree Bayesian Nets Function Prediction EX Simple Bayesian Approach for Localization Prediction 18 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
19 Tag Certain Points Cor e 19 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
20 Find a Division to Separate Tagged Points Cor e 20 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
21 Extrapolate to Untagged Points Cor e 21 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
22 Discriminant to Position Plane Cor e 22 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
23 Fisher discriminant analysis Use the training set to reveal the structure of class distribution by seeking a linear combination y = w 1 x 1 + w 2 x w n x n which maximizes the ratio of the separation of the class means to the sum of each class variance (within class variance). This linear combination is called the first linear discriminant or first canonical variate. Classification of a future case is then determined by choosing the nearest class in the space of the first linear discriminant and significant subsequent discriminants, which maximally separate the class means and are constrained to be uncorrelated with previous ones. 23 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
24 Fischer s Discriminant (Adapted from???) 24 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
25 Fisher cont. Solution of 1 st variate 25 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
26 Find a Division to Separate Tagged Points Cor e 26 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
27 Retrospective Decision Trees Cor e Analysis of the Suitability of 500 M. thermo. proteins to find optimal sequences purification 27 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
28 Retrospective Decision Trees Nomenclature Not Expressible 356 total Cor e Has a hydrophobic stretch? (Y/N) Express -ible 28 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
29 Retrospective Decision Trees Analysis of the Suitability of 500 M. thermo. proteins to find optimal sequences purification Not Express Express 29 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
30 Overfitting, Cross Validation, and Pruning 30 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
31 Decision Trees can handle data that is not linearly separable. A decision tree is an upside down tree in which each branch node represents a choice between a number of alternatives, and each leaf node represents a classification or decision. One classifies instances by sorting them down the tree from the root to some leaf nodes. To classify an instance the tree calls first for a test at the root node, testing the feature indicated on this node and choosing the next node connected to the root branch where the outcome agrees with the value of the feature of that instance. Thereafter a second test on another feature is made on the next node. This process is then repeated until a leaf of the tree is reached. Growing the tree, based on a training set, requires strategies for (a) splitting the nodes and (b) pruning the tree. Maximizing the decrease in average impurity is a common criterion for splitting. In a problem with noisy data (where distribution of observations from the classes overlap) growing the tree will usually over-fit the training set. The strategy in most of the cost-complexity pruning algorithms is to choose the smallest tree whose error rate performance is close to the minimal error rate of the over-fit larger tree. More specifically, growing the trees is based on splitting the node that maximizes the reduction in deviance (or any other impurity-measure of the distribution at a node) over all allowed binary splits of all terminal nodes. Splits are not chosen based on misclassification rate.a binary split for a continuous feature variable v is of the form v<threshold versus v>threshold and for a descriptive factor it divides the factor s levels into two classes. Decision tree-models have been successfully applied in a broad range of domains. Their popularity arises from the following: Decision trees are easy to interpret and use when the predictors are a mix of numeric and nonnumeric (factor) variables. They are invariant to scaling or re-expression of numeric variables. Compared with linear and additive models they are effective in treating missing values and capturing non-additive behavior. They can also be used to predict nonnumeric dependent variables with more than two levels. In addition, decisiontree models are useful to devise prediction rules, screen the variables and summarize the multivariate data set in a comprehensive fashion. We also note that ANN and decision tree learning often have comparable prediction accuracy [Mitchell p. 85] and SVM algorithms are slower compared with decision tree. These facts suggest that the decision tree method should be one of our top candidates to data-mine proteomics datasets. C4.5 and CART are among the most popular decision tree algorithms. Optional: not needed for Quiz (adapted from Y Kluger) 31 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
32 Effect of Scaling (adapted from ref?) 32 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
33 Large-scale Datamining Gene Expression Representing Data in a Grid Description of function prediction in abstract context Unsupervised Learning clustering & k-means Local clustering Supervised Learning Discriminants & Decision Tree Bayesian Nets Function Prediction EX Simple Bayesian Approach for Localization Prediction 33 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
34 Represent predictors in abstract high dimensional space 34 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
35 Tagged Data 35 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
36 Probabilistic Predictions of Class 36 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
37 37 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu Yeast Tables for Localization Prediction
38 Large-scale Datamining Gene Expression Representing Data in a Grid Description of function prediction in abstract context Unsupervised Learning clustering & k-means Local clustering Supervised Learning Discriminants & Decision Tree Bayesian Nets Function Prediction EX Simple Bayesian Approach for Localization Prediction 38 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
39 Spectral Methods Outline & Papers Simple background on PCA (emphasizing lingo) More abstract run through on SVD Application to O Alter et al. (2000). "Singular value decomposition for genome-wide expression data processing and modeling." PNAS vol. 97: Y Kluger et al. (2003). "Spectral biclustering of microarray data: coclustering genes and conditions." Genome Res 13: (c) M Gerstein '06, gerstein.info/talks 39
40 PCA (c) M Gerstein '06, gerstein.info/talks 40
41 PCA section will be a "mash up" up a number of PPTs on the web pca-1 - black ---> by Professor Gillian R. Knapp gk@astro.princeton.edu pca-2 - yellow ---> myweb.dal.ca/~hwhitehe/biol4062/pca.ppt by Hal Whitehead. This is the class main url handout4062.htm pca.ppt - what is cov. matrix ----> hebb.mit.edu/courses/9.641/lectures/pca.ppt by Sebastian Seung. Here is the main page of the course from BIIS_05lecture7.ppt ----> by R.S.Gaborski Professor (c) M Gerstein '06, gerstein.info/talks 41
42 abstract Principal component analysis (PCA) is a technique that is useful for the compression and classification of data. The purpose is to reduce the dimensionality of a data set (sample) by finding a new set of variables, smaller than the original set of variables, that nonetheless retains most of the sample's information. By information we mean the variation present in the sample, given by the correlations between the original variables. The new variables, called principal components (PCs), are uncorrelated, and are ordered by the fraction of the total information each retains. Adapted from
43 Geometric picture of principal components (PCs) A sample of n observations in the 2-D space Goal: to account for the variation in a sample in as few variables as possible, to some accuracy Adapted from 43
44 Geometric picture of principal components (PCs) the 1 st PC is a minimum distance fit to a line in space the 2 nd PC is a minimum distance fit to a line in the plane perpendicular to the 1 st PC PCs are a series of linear least squares fits to a sample, each orthogonal to all the previous. 44 Adapted from
45 PCA: General methodology From k original variables: x 1,x 2,...,x k : Produce k new variables: y 1,y 2,...,y k : y 1 = a 11 x 1 + a 12 x a 1k x k y 2 = a 21 x 1 + a 22 x a 2k x k... y k = a k1 x 1 + a k2 x a kk x k such that: y k 's are uncorrelated (orthogonal) y 1 explains as much as possible of original variance in data set y 2 explains as much as possible of remaining variance etc. Adapted from 45
46 PCA: General methodology From k original variables: x 1,x 2,...,x k : Produce k new variables: y 1,y 2,...,y k : y 1 = a 11 x 1 + a 12 x a 1k x k y 2 = a 21 x 1 + a 22 x a 2k x k... y k = a k1 x 1 + a k2 x a kk x k such that: y k 's are uncorrelated (orthogonal) y 1 explains as much as possible of original variance in data set y 2 explains as much as possible of remaining variance etc. Adapted from y k 's are Principal Components 46
47 Principal Components Analysis 2nd Principal Component, y 2 1st Principal Component, y 1 Adapted from 47
48 Principal Components Analysis Rotates multivariate dataset into a new configuration which is easier to interpret Purposes simplify data look at relationships between variables look at patterns of units Adapted from 48
49 Principal Components Analysis Uses: Correlation matrix, or Covariance matrix when variables in same units (morphometrics, etc.) Adapted from 49
50 Adapted from Principal Components Analysis {a 11,a 12,...,a 1k } is 1st Eigenvector of correlation/covariance matrix, and coefficients of first principal component {a 21,a 22,...,a 2k } is 2nd Eigenvector of correlation/covariance matrix, and coefficients of 2nd principal component {a k1,a k2,...,a kk } is kth Eigenvector of correlation/ covariance matrix, and coefficients of kth principal component 50
51 Digression #1: Where do you get covar matrix? {a 11,a 12,...,a 1k } is 1st Eigenvector of correlation/covariance matrix, and coefficients of first principal component {a 21,a 22,...,a 2k } is 2nd Eigenvector of correlation/covariance matrix, and coefficients of 2nd principal component {a k1,a k2,...,a kk } is kth Eigenvector of correlation/ covariance matrix, and coefficients of kth principal component 51
52 Variance A random variable fluctuating about its mean value. Average of the square of the fluctuations. Adapted from hebb.mit.edu/courses/9.641/lectures/pca.ppt 52
53 Covariance Pair of random variables, each fluctuating about its mean value. Average of product of fluctuations. Adapted from hebb.mit.edu/courses/9.641/lectures/pca.ppt 53
54 Covariance examples Adapted from hebb.mit.edu/courses/9.641/lectures/pca.ppt 54
55 Covariance matrix N random variables NxN symmetric matrix Diagonal elements are variances Adapted from hebb.mit.edu/courses/9.641/lectures/pca.ppt 55
56 Adapted from Principal Components Analysis {a 11,a 12,...,a 1k } is 1st Eigenvector of correlation/covariance matrix, and coefficients of first principal component {a 21,a 22,...,a 2k } is 2nd Eigenvector of correlation/covariance matrix, and coefficients of 2nd principal component {a k1,a k2,...,a kk } is kth Eigenvector of correlation/ covariance matrix, and coefficients of kth principal component 56
57 Digression #2: Brief Review of Eigenvectors {a 11,a 12,...,a 1k } is 1st Eigenvector of correlation/covariance matrix, and coefficients of first principal component {a 21,a 22,...,a 2k } is 2nd Eigenvector of correlation/covariance matrix, and coefficients of 2nd principal component {a k1,a k2,...,a kk } is kth Eigenvector of correlation/ covariance matrix, and coefficients of kth principal component 57
58 eigenvalue problem The eigenvalue problem is any problem having the following form: A. v = λ. v A: n x n matrix v: n x 1 non-zero vector λ: scalar Any value of λ for which this equation has a solution is called the eigenvalue of A and vector v which corresponds to this value is called the eigenvector of A. [from BIIS_05lecture7.ppt] Adapted from 58
59 eigenvalue problem x = = 4 x A. v = λ. v Therefore, (3,2) is an eigenvector of the square matrix A and 4 is an eigenvalue of A Given matrix A, how can we calculate the eigenvector and eigenvalues for A? [from BIIS_05lecture7.ppt] Adapted from 59
60 Principal Components Analysis So, principal components are given by: y 1 = a 11 x 1 + a 12 x a 1k x k y 2 = a 21 x 1 + a 22 x a 2k x k... y k = a k1 x 1 + a k2 x a kk x k x j s are standardized if correlation matrix is used (mean 0.0, SD 1.0) Adapted from 60
61 Principal Components Analysis Score of ith unit on jth principal component y i,j = a j1 x i1 + a j2 x i a jk x ik Adapted from 61
62 Adapted from PCA Scores x i2 y i,1 y i,2 x i1 62
63 Principal Components Analysis Amount of variance accounted for by: 1st principal component, λ 1, 1st eigenvalue 2nd principal component, λ 2, 2nd eigenvalue... λ 1 > λ 2 > λ 3 > λ 4 >... Average λ j = 1 (correlation matrix) Adapted from 63
64 Principal Components Analysis: Eigenvalues λ 1 λ 2 Adapted from 64
65 PCA: Terminology jth principal component is jth eigenvector of correlation/covariance matrix coefficients, a jk, are elements of eigenvectors and relate original variables (standardized if using correlation matrix) to components scores are values of units on components (produced using coefficients) amount of variance accounted for by component is given by eigenvalue, λ j proportion of variance accounted for by Adapted from component is given by λ j / Σ λ j 65
66 How many components to use? If λ j < 1 then component explains less variance than original variable (correlation matrix) Use 2 components (or 3) for visual ease Scree diagram: Adapted from 66
67 Principal Components Analysis on: Covariance Matrix: Variables must be in same units Emphasizes variables with most variance Mean eigenvalue 1.0 Useful in morphometrics, a few other cases Correlation Matrix: Variables are standardized (mean 0.0, SD 1.0) Variables can be in different units All variables have same impact on analysis Mean eigenvalue = 1.0 Adapted from 67
68 PCA: Potential Problems Lack of Independence NO PROBLEM Lack of Normality Normality desirable but not essential Lack of Precision Precision desirable but not essential Many Zeroes in Data Matrix Problem (use Correspondence Analysis) Adapted from 68
69 PCA applications -Eigenfaces Adapted from the principal eigenface looks like a bland androgynous average human face Image:Eigenfaces.png (c) M Gerstein '06, gerstein.info/talks 69
70 Eigenfaces Face Recognition When properly weighted, eigenfaces can be summed together to create an approximate gray-scale rendering of a human face. Remarkably few eigenvector terms are needed to give a fair likeness of most people's faces Hence eigenfaces provide a means of applying data compression to faces for identification purposes. Adapted from (c) M Gerstein '06, gerstein.info/talks 70
71 SVD Puts together slides prepared by Brandon Xia with images from Alter et al. and Kluger et al. (c) M Gerstein '06, gerstein.info/talks 71
72 SVD A = USV T A (m by n) is any rectangular matrix (m rows and n columns) U (m by n) is an orthogonal matrix S (n by n) is a diagonal matrix V (n by n) is another orthogonal matrix Such decomposition always exists All matrices are real; m n 72
73 SVD for microarray data (Alter et al, PNAS 2000) 73
74 A = USV T A is any rectangular matrix (m n) Row space: vector subspace generated by the row vectors of A Column space: vector subspace generated by the column vectors of A The dimension of the row & column space is the rank of the matrix A: r ( n) A is a linear transformation that maps vector x in row space into vector Ax in column space 74
75 A = USV T U is an orthogonal matrix (m n) Column vectors of U form an orthonormal basis for the column space of A: U T U=I u 1,, u n in U are eigenvectors of AA T AA T =USV T VSU T =US 2 U T Left singular vectors 75
76 A = USV T V is an orthogonal matrix (n by n) Column vectors of V form an orthonormal basis for the row space of A: V T V=VV T =I v 1,, v n in V are eigenvectors of A T A A T A =VSU T USV T =VS 2 V T Right singular vectors 76
77 A = USV T S is a diagonal matrix (n by n) of nonnegative singular values Typically sorted from largest to smallest Singular values are the non-negative square root of corresponding eigenvalues of A T A and AA T 77
78 AV = US Means each Av i = s i u i Remember A is a linear map from row space to column space Here, A maps an orthonormal basis {v i } in row space into an orthonormal basis {u i } in column space Each component of u i is the projection of a row onto the vector v i 78
79 Full SVD We can complete U to a full orthogonal matrix and pad S by zeros accordingly = 79
80 Reduced SVD For rectangular matrices, we have two forms of SVD. The reduced SVD looks like this: The columns of U are orthonormal Cheaper form for computation and storage = 80
81 SVD of A (m by n): recap A = USV T = (big-"orthogonal")(diagonal)(sq-orthogonal) u 1,, u m in U are eigenvectors of AA T v 1,, v n in V are eigenvectors of A T A s 1,, s n in S are nonnegative singular values of A AV = US means each Av i = s i u i Every A is diagonalized by 2 orthogonal matrices 81
82 SVD as sum of rank-1 matrices A = USV T A = s 1 u 1 v 1 T + s 2 u 2 v 2 T + + s n u n v n T s 1 s 2 s n 0 What is the rank-r matrix A that best approximates A? Minimize A = s 1 u 1 v 1 T + s 2 u 2 v 2 T + + s r u r v r T Very useful for matrix approximation 82
83 Examples of (almost) rank-1 matrices Steady states with fluctuations Array artifacts? Signals? 83
84 Geometry of SVD in row space A as a collection of m row vectors (points) in the row space of A s 1 u 1 v 1 T is the best rank-1 matrix approximation for A Geometrically: v 1 is the direction of the best approximating rank-1 subspace that goes through origin s 1 u 1 gives coordinates for row vectors in rank-1 subspace v 1 Gives coordinates for row space basis vectors in rank-1 subspace y 84 v 1 x
85 Geometry of SVD in row space y v 1 A x y y s 1 u 1 v 1 T x x This line segment that goes through origin approximates the original data set The projected data set 85 approximates the original data set
86 Geometry of SVD in row space A as a collection of m row vectors (points) in the row space of A y y x s 1 u 1 v 1 T + s 2 u 2 v 2 T is the best rank-2 matrix approximation for A Geometrically: v 1 and v 2 are the directions of the best approximating rank-2 subspace that goes through origin x s 1 u 1 and s 2 u 2 gives coordinates for row vectors in rank-2 subspace v 1 and v 2 gives coordinates for row space basis vectors in rank-2 subspace 86
87 What about geometry of SVD in A = USV T A T = VSU T column space? The column space of A becomes the row space of A T The same as before, except that U and V are switched 87
88 Geometry of SVD in row and column spaces Row space s i u i gives coordinates for row vectors along unit vector v i v i gives coordinates for row space basis vectors along unit vector v i Column space s i v i gives coordinates for column vectors along unit vector u i u i gives coordinates for column space basis vectors along unit vector u i Along the directions v i and u i, these two spaces look pretty much the same! Up to scale factors s i Switch row/column vectors and row/column space basis vectors Biplot... 88
89 Biplot A biplot is a two-dimensional representation of a data matrix showing a point for each of the n observation vectors (rows of the data matrix) along with a point for each of the p variables (columns of the data matrix). The prefix bi refers to the two kinds of points; not to the dimensionality of the plot. The method presented here could, in fact, be generalized to a threedimensional (or higher-order) biplot. Biplots were introduced by Gabriel (1971) and have been discussed at length by Gower and Hand (1996). We applied the biplot procedure to the following toy data matrix to illustrate how a biplot can be generated and interpreted. See the figure on the next page. Here we have three variables (transcription factors) and ten observations (genomic bins). We can obtain a two-dimensional plot of the observations by plotting the first two principal components of the TF-TF correlation matrix R1. We can then add a representation of the three variables to the plot of principal components to obtain a biplot. This shows each of the genomic bins as points and the axes as linear combination of the factors. The great advantage of a biplot is that its components can be interpreted very easily. First, correlations among the variables are related to the angles between the lines, or more specifically, to the cosines of these angles. An acute angle between two lines (representing two TFs) indicates a positive correlation between the two corresponding variables, while obtuse angles indicate negative correlation. Angle of 0 or 180 degrees indicates perfect positive or negative correlation, respectively. A pair of orthogonal lines represents a correlation of zero. The distances between the points (representing genomic bins) correspond to the similarities between the observation profiles. Two observations that are relatively similar across all the variables will fall relatively close to each other within the two-dimensional space used for the biplot. The value or score for any observation on any variable is related to the perpendicular projection form the point to the line. Refs Gabriel, K. R. (1971), The Biplot Graphical Display of Matrices with Application to Principal Component Analysis, Biometrika, 58, Gower, J. C., and Hand, D. J. (1996), Biplots, London: Chapman & Hall. 89
90 Biplot Ex 90
91 Biplot Ex #2 91
92 Biplot Ex #3 Assuming s=1, Av = u A T u = v 92
93 When is SVD = PCA? Centered data y y x x 93
94 When is SVD different from PCA? PCA y y x x y x SVD Translation is not a linear operation, as it moves the 94 origin!
95 Additional Points Time Complexity Issues with SVD Application of SVD to text mining A 95
96 Conclusion SVD is the absolute high point of linear algebra SVD is difficult to compute; but once we have it, we have many things SVD finds the best approximating subspace, using linear transformation Simple SVD cannot handle translation, nonlinear transformation, separation of labeled data, etc. Good for exploratory analysis; but once we know what we look for, use appropriate tools and model the structure of data explicitly! 96
What is Principal Component Analysis?
 What is Principal Component Analysis? Principal component analysis (PCA) Reduce the dimensionality of a data set by finding a new set of variables, smaller than the original set of variables Retains most
What is Principal Component Analysis? Principal component analysis (PCA) Reduce the dimensionality of a data set by finding a new set of variables, smaller than the original set of variables Retains most
Dimensionality Reduction: PCA. Nicholas Ruozzi University of Texas at Dallas
 Dimensionality Reduction: PCA Nicholas Ruozzi University of Texas at Dallas Eigenvalues λ is an eigenvalue of a matrix A R n n if the linear system Ax = λx has at least one non-zero solution If Ax = λx
Dimensionality Reduction: PCA Nicholas Ruozzi University of Texas at Dallas Eigenvalues λ is an eigenvalue of a matrix A R n n if the linear system Ax = λx has at least one non-zero solution If Ax = λx
PCA & ICA. CE-717: Machine Learning Sharif University of Technology Spring Soleymani
 PCA & ICA CE-717: Machine Learning Sharif University of Technology Spring 2015 Soleymani Dimensionality Reduction: Feature Selection vs. Feature Extraction Feature selection Select a subset of a given
PCA & ICA CE-717: Machine Learning Sharif University of Technology Spring 2015 Soleymani Dimensionality Reduction: Feature Selection vs. Feature Extraction Feature selection Select a subset of a given
Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation)
 Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation) PCA transforms the original input space into a lower dimensional space, by constructing dimensions that are linear combinations
Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation) PCA transforms the original input space into a lower dimensional space, by constructing dimensions that are linear combinations
Principal Component Analysis (PCA)
 Principal Component Analysis (PCA) Additional reading can be found from non-assessed exercises (week 8) in this course unit teaching page. Textbooks: Sect. 6.3 in [1] and Ch. 12 in [2] Outline Introduction
Principal Component Analysis (PCA) Additional reading can be found from non-assessed exercises (week 8) in this course unit teaching page. Textbooks: Sect. 6.3 in [1] and Ch. 12 in [2] Outline Introduction
Robot Image Credit: Viktoriya Sukhanova 123RF.com. Dimensionality Reduction
 Robot Image Credit: Viktoriya Sukhanova 13RF.com Dimensionality Reduction Feature Selection vs. Dimensionality Reduction Feature Selection (last time) Select a subset of features. When classifying novel
Robot Image Credit: Viktoriya Sukhanova 13RF.com Dimensionality Reduction Feature Selection vs. Dimensionality Reduction Feature Selection (last time) Select a subset of features. When classifying novel
Machine Learning. B. Unsupervised Learning B.2 Dimensionality Reduction. Lars Schmidt-Thieme, Nicolas Schilling
 Machine Learning B. Unsupervised Learning B.2 Dimensionality Reduction Lars Schmidt-Thieme, Nicolas Schilling Information Systems and Machine Learning Lab (ISMLL) Institute for Computer Science University
Machine Learning B. Unsupervised Learning B.2 Dimensionality Reduction Lars Schmidt-Thieme, Nicolas Schilling Information Systems and Machine Learning Lab (ISMLL) Institute for Computer Science University
Introduction to Machine Learning. PCA and Spectral Clustering. Introduction to Machine Learning, Slides: Eran Halperin
 1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
Machine Learning. Principal Components Analysis. Le Song. CSE6740/CS7641/ISYE6740, Fall 2012
 Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
CS4495/6495 Introduction to Computer Vision. 8B-L2 Principle Component Analysis (and its use in Computer Vision)
 CS4495/6495 Introduction to Computer Vision 8B-L2 Principle Component Analysis (and its use in Computer Vision) Wavelength 2 Wavelength 2 Principal Components Principal components are all about the directions
CS4495/6495 Introduction to Computer Vision 8B-L2 Principle Component Analysis (and its use in Computer Vision) Wavelength 2 Wavelength 2 Principal Components Principal components are all about the directions
Lecture: Face Recognition and Feature Reduction
 Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab Lecture 11-1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed
Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab Lecture 11-1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed
LEC 2: Principal Component Analysis (PCA) A First Dimensionality Reduction Approach
 LEC 2: Principal Component Analysis (PCA) A First Dimensionality Reduction Approach Dr. Guangliang Chen February 9, 2016 Outline Introduction Review of linear algebra Matrix SVD PCA Motivation The digits
LEC 2: Principal Component Analysis (PCA) A First Dimensionality Reduction Approach Dr. Guangliang Chen February 9, 2016 Outline Introduction Review of linear algebra Matrix SVD PCA Motivation The digits
Introduction to Machine Learning
 10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
Singular Value Decomposition. 1 Singular Value Decomposition and the Four Fundamental Subspaces
 Singular Value Decomposition This handout is a review of some basic concepts in linear algebra For a detailed introduction, consult a linear algebra text Linear lgebra and its pplications by Gilbert Strang
Singular Value Decomposition This handout is a review of some basic concepts in linear algebra For a detailed introduction, consult a linear algebra text Linear lgebra and its pplications by Gilbert Strang
Lecture 13. Principal Component Analysis. Brett Bernstein. April 25, CDS at NYU. Brett Bernstein (CDS at NYU) Lecture 13 April 25, / 26
 Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Unsupervised Learning: K- Means & PCA
 Unsupervised Learning: K- Means & PCA Unsupervised Learning Supervised learning used labeled data pairs (x, y) to learn a func>on f : X Y But, what if we don t have labels? No labels = unsupervised learning
Unsupervised Learning: K- Means & PCA Unsupervised Learning Supervised learning used labeled data pairs (x, y) to learn a func>on f : X Y But, what if we don t have labels? No labels = unsupervised learning
Lecture 24: Principal Component Analysis. Aykut Erdem May 2016 Hacettepe University
 Lecture 4: Principal Component Analysis Aykut Erdem May 016 Hacettepe University This week Motivation PCA algorithms Applications PCA shortcomings Autoencoders Kernel PCA PCA Applications Data Visualization
Lecture 4: Principal Component Analysis Aykut Erdem May 016 Hacettepe University This week Motivation PCA algorithms Applications PCA shortcomings Autoencoders Kernel PCA PCA Applications Data Visualization
CSC 411 Lecture 12: Principal Component Analysis
 CSC 411 Lecture 12: Principal Component Analysis Roger Grosse, Amir-massoud Farahmand, and Juan Carrasquilla University of Toronto UofT CSC 411: 12-PCA 1 / 23 Overview Today we ll cover the first unsupervised
CSC 411 Lecture 12: Principal Component Analysis Roger Grosse, Amir-massoud Farahmand, and Juan Carrasquilla University of Toronto UofT CSC 411: 12-PCA 1 / 23 Overview Today we ll cover the first unsupervised
PCA and admixture models
 PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
Lecture: Face Recognition and Feature Reduction
 Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab 1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed in the
Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab 1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed in the
CS 4495 Computer Vision Principle Component Analysis
 CS 4495 Computer Vision Principle Component Analysis (and it s use in Computer Vision) Aaron Bobick School of Interactive Computing Administrivia PS6 is out. Due *** Sunday, Nov 24th at 11:55pm *** PS7
CS 4495 Computer Vision Principle Component Analysis (and it s use in Computer Vision) Aaron Bobick School of Interactive Computing Administrivia PS6 is out. Due *** Sunday, Nov 24th at 11:55pm *** PS7
Principal Component Analysis
 B: Chapter 1 HTF: Chapter 1.5 Principal Component Analysis Barnabás Póczos University of Alberta Nov, 009 Contents Motivation PCA algorithms Applications Face recognition Facial expression recognition
B: Chapter 1 HTF: Chapter 1.5 Principal Component Analysis Barnabás Póczos University of Alberta Nov, 009 Contents Motivation PCA algorithms Applications Face recognition Facial expression recognition
Statistical Pattern Recognition
 Statistical Pattern Recognition Feature Extraction Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi, Payam Siyari Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Dimensionality Reduction
Statistical Pattern Recognition Feature Extraction Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi, Payam Siyari Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Dimensionality Reduction
PCA, Kernel PCA, ICA
 PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
Principal Component Analysis. Applied Multivariate Statistics Spring 2012
 Principal Component Analysis Applied Multivariate Statistics Spring 2012 Overview Intuition Four definitions Practical examples Mathematical example Case study 2 PCA: Goals Goal 1: Dimension reduction
Principal Component Analysis Applied Multivariate Statistics Spring 2012 Overview Intuition Four definitions Practical examples Mathematical example Case study 2 PCA: Goals Goal 1: Dimension reduction
Karhunen-Loève Transform KLT. JanKees van der Poel D.Sc. Student, Mechanical Engineering
 Karhunen-Loève Transform KLT JanKees van der Poel D.Sc. Student, Mechanical Engineering Karhunen-Loève Transform Has many names cited in literature: Karhunen-Loève Transform (KLT); Karhunen-Loève Decomposition
Karhunen-Loève Transform KLT JanKees van der Poel D.Sc. Student, Mechanical Engineering Karhunen-Loève Transform Has many names cited in literature: Karhunen-Loève Transform (KLT); Karhunen-Loève Decomposition
Machine Learning 2nd Edition
 INTRODUCTION TO Lecture Slides for Machine Learning 2nd Edition ETHEM ALPAYDIN, modified by Leonardo Bobadilla and some parts from http://www.cs.tau.ac.il/~apartzin/machinelearning/ The MIT Press, 2010
INTRODUCTION TO Lecture Slides for Machine Learning 2nd Edition ETHEM ALPAYDIN, modified by Leonardo Bobadilla and some parts from http://www.cs.tau.ac.il/~apartzin/machinelearning/ The MIT Press, 2010
Principal Component Analysis (PCA)
 Principal Component Analysis (PCA) Salvador Dalí, Galatea of the Spheres CSC411/2515: Machine Learning and Data Mining, Winter 2018 Michael Guerzhoy and Lisa Zhang Some slides from Derek Hoiem and Alysha
Principal Component Analysis (PCA) Salvador Dalí, Galatea of the Spheres CSC411/2515: Machine Learning and Data Mining, Winter 2018 Michael Guerzhoy and Lisa Zhang Some slides from Derek Hoiem and Alysha
Linear Dimensionality Reduction
 Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Principal Component Analysis 3 Factor Analysis
Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Principal Component Analysis 3 Factor Analysis
Vectors To begin, let us describe an element of the state space as a point with numerical coordinates, that is x 1. x 2. x =
 Linear Algebra Review Vectors To begin, let us describe an element of the state space as a point with numerical coordinates, that is x 1 x x = 2. x n Vectors of up to three dimensions are easy to diagram.
Linear Algebra Review Vectors To begin, let us describe an element of the state space as a point with numerical coordinates, that is x 1 x x = 2. x n Vectors of up to three dimensions are easy to diagram.
Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function.
 Bayesian learning: Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function. Let y be the true label and y be the predicted
Bayesian learning: Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function. Let y be the true label and y be the predicted
Deriving Principal Component Analysis (PCA)
 -0 Mathematical Foundations for Machine Learning Machine Learning Department School of Computer Science Carnegie Mellon University Deriving Principal Component Analysis (PCA) Matt Gormley Lecture 11 Oct.
-0 Mathematical Foundations for Machine Learning Machine Learning Department School of Computer Science Carnegie Mellon University Deriving Principal Component Analysis (PCA) Matt Gormley Lecture 11 Oct.
Machine Learning (BSMC-GA 4439) Wenke Liu
 Machine Learning (BSMC-GA 4439) Wenke Liu 02-01-2018 Biomedical data are usually high-dimensional Number of samples (n) is relatively small whereas number of features (p) can be large Sometimes p>>n Problems
Machine Learning (BSMC-GA 4439) Wenke Liu 02-01-2018 Biomedical data are usually high-dimensional Number of samples (n) is relatively small whereas number of features (p) can be large Sometimes p>>n Problems
Advanced Introduction to Machine Learning CMU-10715
 Advanced Introduction to Machine Learning CMU-10715 Principal Component Analysis Barnabás Póczos Contents Motivation PCA algorithms Applications Some of these slides are taken from Karl Booksh Research
Advanced Introduction to Machine Learning CMU-10715 Principal Component Analysis Barnabás Póczos Contents Motivation PCA algorithms Applications Some of these slides are taken from Karl Booksh Research
Principal Component Analysis
 Principal Component Analysis Yingyu Liang yliang@cs.wisc.edu Computer Sciences Department University of Wisconsin, Madison [based on slides from Nina Balcan] slide 1 Goals for the lecture you should understand
Principal Component Analysis Yingyu Liang yliang@cs.wisc.edu Computer Sciences Department University of Wisconsin, Madison [based on slides from Nina Balcan] slide 1 Goals for the lecture you should understand
Singular Value Decomposition and Principal Component Analysis (PCA) I
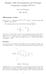 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA
 Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA Principle Components Analysis: Uses one group of variables (we will call this X) In
Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA Principle Components Analysis: Uses one group of variables (we will call this X) In
PRINCIPAL COMPONENTS ANALYSIS
 121 CHAPTER 11 PRINCIPAL COMPONENTS ANALYSIS We now have the tools necessary to discuss one of the most important concepts in mathematical statistics: Principal Components Analysis (PCA). PCA involves
121 CHAPTER 11 PRINCIPAL COMPONENTS ANALYSIS We now have the tools necessary to discuss one of the most important concepts in mathematical statistics: Principal Components Analysis (PCA). PCA involves
Machine Learning. CUNY Graduate Center, Spring Lectures 11-12: Unsupervised Learning 1. Professor Liang Huang.
 Machine Learning CUNY Graduate Center, Spring 2013 Lectures 11-12: Unsupervised Learning 1 (Clustering: k-means, EM, mixture models) Professor Liang Huang huang@cs.qc.cuny.edu http://acl.cs.qc.edu/~lhuang/teaching/machine-learning
Machine Learning CUNY Graduate Center, Spring 2013 Lectures 11-12: Unsupervised Learning 1 (Clustering: k-means, EM, mixture models) Professor Liang Huang huang@cs.qc.cuny.edu http://acl.cs.qc.edu/~lhuang/teaching/machine-learning
Machine Learning - MT & 14. PCA and MDS
 Machine Learning - MT 2016 13 & 14. PCA and MDS Varun Kanade University of Oxford November 21 & 23, 2016 Announcements Sheet 4 due this Friday by noon Practical 3 this week (continue next week if necessary)
Machine Learning - MT 2016 13 & 14. PCA and MDS Varun Kanade University of Oxford November 21 & 23, 2016 Announcements Sheet 4 due this Friday by noon Practical 3 this week (continue next week if necessary)
ECE 521. Lecture 11 (not on midterm material) 13 February K-means clustering, Dimensionality reduction
 ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
Data Preprocessing Tasks
 Data Tasks 1 2 3 Data Reduction 4 We re here. 1 Dimensionality Reduction Dimensionality reduction is a commonly used approach for generating fewer features. Typically used because too many features can
Data Tasks 1 2 3 Data Reduction 4 We re here. 1 Dimensionality Reduction Dimensionality reduction is a commonly used approach for generating fewer features. Typically used because too many features can
Data Mining. Dimensionality reduction. Hamid Beigy. Sharif University of Technology. Fall 1395
 Data Mining Dimensionality reduction Hamid Beigy Sharif University of Technology Fall 1395 Hamid Beigy (Sharif University of Technology) Data Mining Fall 1395 1 / 42 Outline 1 Introduction 2 Feature selection
Data Mining Dimensionality reduction Hamid Beigy Sharif University of Technology Fall 1395 Hamid Beigy (Sharif University of Technology) Data Mining Fall 1395 1 / 42 Outline 1 Introduction 2 Feature selection
CPSC 340: Machine Learning and Data Mining. More PCA Fall 2017
 CPSC 340: Machine Learning and Data Mining More PCA Fall 2017 Admin Assignment 4: Due Friday of next week. No class Monday due to holiday. There will be tutorials next week on MAP/PCA (except Monday).
CPSC 340: Machine Learning and Data Mining More PCA Fall 2017 Admin Assignment 4: Due Friday of next week. No class Monday due to holiday. There will be tutorials next week on MAP/PCA (except Monday).
Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations.
 Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
EECS490: Digital Image Processing. Lecture #26
 Lecture #26 Moments; invariant moments Eigenvector, principal component analysis Boundary coding Image primitives Image representation: trees, graphs Object recognition and classes Minimum distance classifiers
Lecture #26 Moments; invariant moments Eigenvector, principal component analysis Boundary coding Image primitives Image representation: trees, graphs Object recognition and classes Minimum distance classifiers
Face Detection and Recognition
 Face Detection and Recognition Face Recognition Problem Reading: Chapter 18.10 and, optionally, Face Recognition using Eigenfaces by M. Turk and A. Pentland Queryimage face query database Face Verification
Face Detection and Recognition Face Recognition Problem Reading: Chapter 18.10 and, optionally, Face Recognition using Eigenfaces by M. Turk and A. Pentland Queryimage face query database Face Verification
Image Analysis. PCA and Eigenfaces
 Image Analysis PCA and Eigenfaces Christophoros Nikou cnikou@cs.uoi.gr Images taken from: D. Forsyth and J. Ponce. Computer Vision: A Modern Approach, Prentice Hall, 2003. Computer Vision course by Svetlana
Image Analysis PCA and Eigenfaces Christophoros Nikou cnikou@cs.uoi.gr Images taken from: D. Forsyth and J. Ponce. Computer Vision: A Modern Approach, Prentice Hall, 2003. Computer Vision course by Svetlana
Structure in Data. A major objective in data analysis is to identify interesting features or structure in the data.
 Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Face Recognition. Face Recognition. Subspace-Based Face Recognition Algorithms. Application of Face Recognition
 ace Recognition Identify person based on the appearance of face CSED441:Introduction to Computer Vision (2017) Lecture10: Subspace Methods and ace Recognition Bohyung Han CSE, POSTECH bhhan@postech.ac.kr
ace Recognition Identify person based on the appearance of face CSED441:Introduction to Computer Vision (2017) Lecture10: Subspace Methods and ace Recognition Bohyung Han CSE, POSTECH bhhan@postech.ac.kr
Multivariate Statistics (I) 2. Principal Component Analysis (PCA)
 Multivariate Statistics (I) 2. Principal Component Analysis (PCA) 2.1 Comprehension of PCA 2.2 Concepts of PCs 2.3 Algebraic derivation of PCs 2.4 Selection and goodness-of-fit of PCs 2.5 Algebraic derivation
Multivariate Statistics (I) 2. Principal Component Analysis (PCA) 2.1 Comprehension of PCA 2.2 Concepts of PCs 2.3 Algebraic derivation of PCs 2.4 Selection and goodness-of-fit of PCs 2.5 Algebraic derivation
7. Variable extraction and dimensionality reduction
 7. Variable extraction and dimensionality reduction The goal of the variable selection in the preceding chapter was to find least useful variables so that it would be possible to reduce the dimensionality
7. Variable extraction and dimensionality reduction The goal of the variable selection in the preceding chapter was to find least useful variables so that it would be possible to reduce the dimensionality
Regularized Discriminant Analysis and Reduced-Rank LDA
 Regularized Discriminant Analysis and Reduced-Rank LDA Department of Statistics The Pennsylvania State University Email: jiali@stat.psu.edu Regularized Discriminant Analysis A compromise between LDA and
Regularized Discriminant Analysis and Reduced-Rank LDA Department of Statistics The Pennsylvania State University Email: jiali@stat.psu.edu Regularized Discriminant Analysis A compromise between LDA and
Example: Face Detection
 Announcements HW1 returned New attendance policy Face Recognition: Dimensionality Reduction On time: 1 point Five minutes or more late: 0.5 points Absent: 0 points Biometrics CSE 190 Lecture 14 CSE190,
Announcements HW1 returned New attendance policy Face Recognition: Dimensionality Reduction On time: 1 point Five minutes or more late: 0.5 points Absent: 0 points Biometrics CSE 190 Lecture 14 CSE190,
PATTERN CLASSIFICATION
 PATTERN CLASSIFICATION Second Edition Richard O. Duda Peter E. Hart David G. Stork A Wiley-lnterscience Publication JOHN WILEY & SONS, INC. New York Chichester Weinheim Brisbane Singapore Toronto CONTENTS
PATTERN CLASSIFICATION Second Edition Richard O. Duda Peter E. Hart David G. Stork A Wiley-lnterscience Publication JOHN WILEY & SONS, INC. New York Chichester Weinheim Brisbane Singapore Toronto CONTENTS
The Singular Value Decomposition (SVD) and Principal Component Analysis (PCA)
 Chapter 5 The Singular Value Decomposition (SVD) and Principal Component Analysis (PCA) 5.1 Basics of SVD 5.1.1 Review of Key Concepts We review some key definitions and results about matrices that will
Chapter 5 The Singular Value Decomposition (SVD) and Principal Component Analysis (PCA) 5.1 Basics of SVD 5.1.1 Review of Key Concepts We review some key definitions and results about matrices that will
14 Singular Value Decomposition
 14 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
14 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
Statistical Machine Learning
 Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides
 Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization
 Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization Haiping Lu 1 K. N. Plataniotis 1 A. N. Venetsanopoulos 1,2 1 Department of Electrical & Computer Engineering,
Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization Haiping Lu 1 K. N. Plataniotis 1 A. N. Venetsanopoulos 1,2 1 Department of Electrical & Computer Engineering,
CS 340 Lec. 6: Linear Dimensionality Reduction
 CS 340 Lec. 6: Linear Dimensionality Reduction AD January 2011 AD () January 2011 1 / 46 Linear Dimensionality Reduction Introduction & Motivation Brief Review of Linear Algebra Principal Component Analysis
CS 340 Lec. 6: Linear Dimensionality Reduction AD January 2011 AD () January 2011 1 / 46 Linear Dimensionality Reduction Introduction & Motivation Brief Review of Linear Algebra Principal Component Analysis
Singular Value Decomposition
 Singular Value Decomposition Motivatation The diagonalization theorem play a part in many interesting applications. Unfortunately not all matrices can be factored as A = PDP However a factorization A =
Singular Value Decomposition Motivatation The diagonalization theorem play a part in many interesting applications. Unfortunately not all matrices can be factored as A = PDP However a factorization A =
CS168: The Modern Algorithmic Toolbox Lecture #8: How PCA Works
 CS68: The Modern Algorithmic Toolbox Lecture #8: How PCA Works Tim Roughgarden & Gregory Valiant April 20, 206 Introduction Last lecture introduced the idea of principal components analysis (PCA). The
CS68: The Modern Algorithmic Toolbox Lecture #8: How PCA Works Tim Roughgarden & Gregory Valiant April 20, 206 Introduction Last lecture introduced the idea of principal components analysis (PCA). The
CS 143 Linear Algebra Review
 CS 143 Linear Algebra Review Stefan Roth September 29, 2003 Introductory Remarks This review does not aim at mathematical rigor very much, but instead at ease of understanding and conciseness. Please see
CS 143 Linear Algebra Review Stefan Roth September 29, 2003 Introductory Remarks This review does not aim at mathematical rigor very much, but instead at ease of understanding and conciseness. Please see
L11: Pattern recognition principles
 L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
December 20, MAA704, Multivariate analysis. Christopher Engström. Multivariate. analysis. Principal component analysis
 .. December 20, 2013 Todays lecture. (PCA) (PLS-R) (LDA) . (PCA) is a method often used to reduce the dimension of a large dataset to one of a more manageble size. The new dataset can then be used to make
.. December 20, 2013 Todays lecture. (PCA) (PLS-R) (LDA) . (PCA) is a method often used to reduce the dimension of a large dataset to one of a more manageble size. The new dataset can then be used to make
CoDa-dendrogram: A new exploratory tool. 2 Dept. Informàtica i Matemàtica Aplicada, Universitat de Girona, Spain;
 CoDa-dendrogram: A new exploratory tool J.J. Egozcue 1, and V. Pawlowsky-Glahn 2 1 Dept. Matemàtica Aplicada III, Universitat Politècnica de Catalunya, Barcelona, Spain; juan.jose.egozcue@upc.edu 2 Dept.
CoDa-dendrogram: A new exploratory tool J.J. Egozcue 1, and V. Pawlowsky-Glahn 2 1 Dept. Matemàtica Aplicada III, Universitat Politècnica de Catalunya, Barcelona, Spain; juan.jose.egozcue@upc.edu 2 Dept.
Linear Algebra Methods for Data Mining
 Linear Algebra Methods for Data Mining Saara Hyvönen, Saara.Hyvonen@cs.helsinki.fi Spring 2007 Linear Discriminant Analysis Linear Algebra Methods for Data Mining, Spring 2007, University of Helsinki Principal
Linear Algebra Methods for Data Mining Saara Hyvönen, Saara.Hyvonen@cs.helsinki.fi Spring 2007 Linear Discriminant Analysis Linear Algebra Methods for Data Mining, Spring 2007, University of Helsinki Principal
20 Unsupervised Learning and Principal Components Analysis (PCA)
 116 Jonathan Richard Shewchuk 20 Unsupervised Learning and Principal Components Analysis (PCA) UNSUPERVISED LEARNING We have sample points, but no labels! No classes, no y-values, nothing to predict. Goal:
116 Jonathan Richard Shewchuk 20 Unsupervised Learning and Principal Components Analysis (PCA) UNSUPERVISED LEARNING We have sample points, but no labels! No classes, no y-values, nothing to predict. Goal:
Robustness of Principal Components
 PCA for Clustering An objective of principal components analysis is to identify linear combinations of the original variables that are useful in accounting for the variation in those original variables.
PCA for Clustering An objective of principal components analysis is to identify linear combinations of the original variables that are useful in accounting for the variation in those original variables.
Principal Component Analysis and Linear Discriminant Analysis
 Principal Component Analysis and Linear Discriminant Analysis Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/29
Principal Component Analysis and Linear Discriminant Analysis Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/29
Dimensionality Reduction Using PCA/LDA. Hongyu Li School of Software Engineering TongJi University Fall, 2014
 Dimensionality Reduction Using PCA/LDA Hongyu Li School of Software Engineering TongJi University Fall, 2014 Dimensionality Reduction One approach to deal with high dimensional data is by reducing their
Dimensionality Reduction Using PCA/LDA Hongyu Li School of Software Engineering TongJi University Fall, 2014 Dimensionality Reduction One approach to deal with high dimensional data is by reducing their
Least Squares Optimization
 Least Squares Optimization The following is a brief review of least squares optimization and constrained optimization techniques. Broadly, these techniques can be used in data analysis and visualization
Least Squares Optimization The following is a brief review of least squares optimization and constrained optimization techniques. Broadly, these techniques can be used in data analysis and visualization
Discriminant analysis and supervised classification
 Discriminant analysis and supervised classification Angela Montanari 1 Linear discriminant analysis Linear discriminant analysis (LDA) also known as Fisher s linear discriminant analysis or as Canonical
Discriminant analysis and supervised classification Angela Montanari 1 Linear discriminant analysis Linear discriminant analysis (LDA) also known as Fisher s linear discriminant analysis or as Canonical
7. Symmetric Matrices and Quadratic Forms
 Linear Algebra 7. Symmetric Matrices and Quadratic Forms CSIE NCU 1 7. Symmetric Matrices and Quadratic Forms 7.1 Diagonalization of symmetric matrices 2 7.2 Quadratic forms.. 9 7.4 The singular value
Linear Algebra 7. Symmetric Matrices and Quadratic Forms CSIE NCU 1 7. Symmetric Matrices and Quadratic Forms 7.1 Diagonalization of symmetric matrices 2 7.2 Quadratic forms.. 9 7.4 The singular value
Machine Learning (Spring 2012) Principal Component Analysis
 1-71 Machine Learning (Spring 1) Principal Component Analysis Yang Xu This note is partly based on Chapter 1.1 in Chris Bishop s book on PRML and the lecture slides on PCA written by Carlos Guestrin in
1-71 Machine Learning (Spring 1) Principal Component Analysis Yang Xu This note is partly based on Chapter 1.1 in Chris Bishop s book on PRML and the lecture slides on PCA written by Carlos Guestrin in
Dimensionality reduction
 Dimensionality Reduction PCA continued Machine Learning CSE446 Carlos Guestrin University of Washington May 22, 2013 Carlos Guestrin 2005-2013 1 Dimensionality reduction n Input data may have thousands
Dimensionality Reduction PCA continued Machine Learning CSE446 Carlos Guestrin University of Washington May 22, 2013 Carlos Guestrin 2005-2013 1 Dimensionality reduction n Input data may have thousands
CSE 554 Lecture 7: Alignment
 CSE 554 Lecture 7: Alignment Fall 2012 CSE554 Alignment Slide 1 Review Fairing (smoothing) Relocating vertices to achieve a smoother appearance Method: centroid averaging Simplification Reducing vertex
CSE 554 Lecture 7: Alignment Fall 2012 CSE554 Alignment Slide 1 Review Fairing (smoothing) Relocating vertices to achieve a smoother appearance Method: centroid averaging Simplification Reducing vertex
Dimensionality Reduction
 Lecture 5 1 Outline 1. Overview a) What is? b) Why? 2. Principal Component Analysis (PCA) a) Objectives b) Explaining variability c) SVD 3. Related approaches a) ICA b) Autoencoders 2 Example 1: Sportsball
Lecture 5 1 Outline 1. Overview a) What is? b) Why? 2. Principal Component Analysis (PCA) a) Objectives b) Explaining variability c) SVD 3. Related approaches a) ICA b) Autoencoders 2 Example 1: Sportsball
I L L I N O I S UNIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN
 Introduction Edps/Psych/Stat/ 584 Applied Multivariate Statistics Carolyn J Anderson Department of Educational Psychology I L L I N O I S UNIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN c Board of Trustees,
Introduction Edps/Psych/Stat/ 584 Applied Multivariate Statistics Carolyn J Anderson Department of Educational Psychology I L L I N O I S UNIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN c Board of Trustees,
Vector Space Models. wine_spectral.r
 Vector Space Models 137 wine_spectral.r Latent Semantic Analysis Problem with words Even a small vocabulary as in wine example is challenging LSA Reduce number of columns of DTM by principal components
Vector Space Models 137 wine_spectral.r Latent Semantic Analysis Problem with words Even a small vocabulary as in wine example is challenging LSA Reduce number of columns of DTM by principal components
Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes. October 3, Statistics 202: Data Mining
 Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Combinations of features Given a data matrix X n p with p fairly large, it can
Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Combinations of features Given a data matrix X n p with p fairly large, it can
Eigenfaces. Face Recognition Using Principal Components Analysis
 Eigenfaces Face Recognition Using Principal Components Analysis M. Turk, A. Pentland, "Eigenfaces for Recognition", Journal of Cognitive Neuroscience, 3(1), pp. 71-86, 1991. Slides : George Bebis, UNR
Eigenfaces Face Recognition Using Principal Components Analysis M. Turk, A. Pentland, "Eigenfaces for Recognition", Journal of Cognitive Neuroscience, 3(1), pp. 71-86, 1991. Slides : George Bebis, UNR
ECE 661: Homework 10 Fall 2014
 ECE 661: Homework 10 Fall 2014 This homework consists of the following two parts: (1) Face recognition with PCA and LDA for dimensionality reduction and the nearest-neighborhood rule for classification;
ECE 661: Homework 10 Fall 2014 This homework consists of the following two parts: (1) Face recognition with PCA and LDA for dimensionality reduction and the nearest-neighborhood rule for classification;
Linear & Non-Linear Discriminant Analysis! Hugh R. Wilson
 Linear & Non-Linear Discriminant Analysis! Hugh R. Wilson PCA Review! Supervised learning! Fisher linear discriminant analysis! Nonlinear discriminant analysis! Research example! Multiple Classes! Unsupervised
Linear & Non-Linear Discriminant Analysis! Hugh R. Wilson PCA Review! Supervised learning! Fisher linear discriminant analysis! Nonlinear discriminant analysis! Research example! Multiple Classes! Unsupervised
Classification. The goal: map from input X to a label Y. Y has a discrete set of possible values. We focused on binary Y (values 0 or 1).
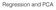 Regression and PCA Classification The goal: map from input X to a label Y. Y has a discrete set of possible values We focused on binary Y (values 0 or 1). But we also discussed larger number of classes
Regression and PCA Classification The goal: map from input X to a label Y. Y has a discrete set of possible values We focused on binary Y (values 0 or 1). But we also discussed larger number of classes
System 1 (last lecture) : limited to rigidly structured shapes. System 2 : recognition of a class of varying shapes. Need to:
 System 2 : Modelling & Recognising Modelling and Recognising Classes of Classes of Shapes Shape : PDM & PCA All the same shape? System 1 (last lecture) : limited to rigidly structured shapes System 2 :
System 2 : Modelling & Recognising Modelling and Recognising Classes of Classes of Shapes Shape : PDM & PCA All the same shape? System 1 (last lecture) : limited to rigidly structured shapes System 2 :
Principal Component Analysis, A Powerful Scoring Technique
 Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
Eigenvalues, Eigenvectors, and an Intro to PCA
 Eigenvalues, Eigenvectors, and an Intro to PCA Eigenvalues, Eigenvectors, and an Intro to PCA Changing Basis We ve talked so far about re-writing our data using a new set of variables, or a new basis.
Eigenvalues, Eigenvectors, and an Intro to PCA Eigenvalues, Eigenvectors, and an Intro to PCA Changing Basis We ve talked so far about re-writing our data using a new set of variables, or a new basis.
STATISTICAL LEARNING SYSTEMS
 STATISTICAL LEARNING SYSTEMS LECTURE 8: UNSUPERVISED LEARNING: FINDING STRUCTURE IN DATA Institute of Computer Science, Polish Academy of Sciences Ph. D. Program 2013/2014 Principal Component Analysis
STATISTICAL LEARNING SYSTEMS LECTURE 8: UNSUPERVISED LEARNING: FINDING STRUCTURE IN DATA Institute of Computer Science, Polish Academy of Sciences Ph. D. Program 2013/2014 Principal Component Analysis
Eigenvalues, Eigenvectors, and an Intro to PCA
 Eigenvalues, Eigenvectors, and an Intro to PCA Eigenvalues, Eigenvectors, and an Intro to PCA Changing Basis We ve talked so far about re-writing our data using a new set of variables, or a new basis.
Eigenvalues, Eigenvectors, and an Intro to PCA Eigenvalues, Eigenvectors, and an Intro to PCA Changing Basis We ve talked so far about re-writing our data using a new set of variables, or a new basis.
15 Singular Value Decomposition
 15 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
15 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
The Singular Value Decomposition
 The Singular Value Decomposition Philippe B. Laval KSU Fall 2015 Philippe B. Laval (KSU) SVD Fall 2015 1 / 13 Review of Key Concepts We review some key definitions and results about matrices that will
The Singular Value Decomposition Philippe B. Laval KSU Fall 2015 Philippe B. Laval (KSU) SVD Fall 2015 1 / 13 Review of Key Concepts We review some key definitions and results about matrices that will
Classification 2: Linear discriminant analysis (continued); logistic regression
 Classification 2: Linear discriminant analysis (continued); logistic regression Ryan Tibshirani Data Mining: 36-462/36-662 April 4 2013 Optional reading: ISL 4.4, ESL 4.3; ISL 4.3, ESL 4.4 1 Reminder:
Classification 2: Linear discriminant analysis (continued); logistic regression Ryan Tibshirani Data Mining: 36-462/36-662 April 4 2013 Optional reading: ISL 4.4, ESL 4.3; ISL 4.3, ESL 4.4 1 Reminder:
Principal Component Analysis
 Machine Learning Michaelmas 2017 James Worrell Principal Component Analysis 1 Introduction 1.1 Goals of PCA Principal components analysis (PCA) is a dimensionality reduction technique that can be used
Machine Learning Michaelmas 2017 James Worrell Principal Component Analysis 1 Introduction 1.1 Goals of PCA Principal components analysis (PCA) is a dimensionality reduction technique that can be used
Data Mining Techniques
 Data Mining Techniques CS 6220 - Section 3 - Fall 2016 Lecture 12 Jan-Willem van de Meent (credit: Yijun Zhao, Percy Liang) DIMENSIONALITY REDUCTION Borrowing from: Percy Liang (Stanford) Linear Dimensionality
Data Mining Techniques CS 6220 - Section 3 - Fall 2016 Lecture 12 Jan-Willem van de Meent (credit: Yijun Zhao, Percy Liang) DIMENSIONALITY REDUCTION Borrowing from: Percy Liang (Stanford) Linear Dimensionality
Probabilistic Latent Semantic Analysis
 Probabilistic Latent Semantic Analysis Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr
Probabilistic Latent Semantic Analysis Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr
Dimensionality Reduction and Principal Components
 Dimensionality Reduction and Principal Components Nuno Vasconcelos (Ken Kreutz-Delgado) UCSD Motivation Recall, in Bayesian decision theory we have: World: States Y in {1,..., M} and observations of X
Dimensionality Reduction and Principal Components Nuno Vasconcelos (Ken Kreutz-Delgado) UCSD Motivation Recall, in Bayesian decision theory we have: World: States Y in {1,..., M} and observations of X
Introduction to SVD and Applications
 Introduction to SVD and Applications Eric Kostelich and Dave Kuhl MSRI Climate Change Summer School July 18, 2008 Introduction The goal of this exercise is to familiarize you with the basics of the singular
Introduction to SVD and Applications Eric Kostelich and Dave Kuhl MSRI Climate Change Summer School July 18, 2008 Introduction The goal of this exercise is to familiarize you with the basics of the singular
Table of Contents. Multivariate methods. Introduction II. Introduction I
 Table of Contents Introduction Antti Penttilä Department of Physics University of Helsinki Exactum summer school, 04 Construction of multinormal distribution Test of multinormality with 3 Interpretation
Table of Contents Introduction Antti Penttilä Department of Physics University of Helsinki Exactum summer school, 04 Construction of multinormal distribution Test of multinormality with 3 Interpretation
